Published online Jul 6, 2022. doi: 10.12998/wjcc.v10.i19.6750
Peer-review started: February 28, 2022
First decision: April 8, 2022
Revised: April 19, 2022
Accepted: April 28, 2022
Article in press: April 28, 2022
Published online: July 6, 2022
Processing time: 115 Days and 22.6 Hours
Tumor-to-tumor metastasis (TTM) is an uncommon condition. Only a few cases of renal cell carcinoma (RCC) as donor tumor of TTM have been reported in literature, and none of these studies have described RCC metastasizing to synchronous pheochromocytoma (PCC).
The patient was a 54-year-old woman who presented with recurrent dull abdominal pain for six months, which was further aggravated for one more month. Enhanced computed tomography revealed a tumor mass in the right kidney and another mass in the left retroperitoneum/adrenal gland. Histopa
This is the first case showing metastasis of CCRCC to PCC, thus leading to tumor-to-tumor metastasis.
Core Tip: Tumor-to-tumor metastasis (TTM) is uncommon. Only a limited number of cases in which renal cell carcinoma (RCC) is the donor tumor of TTM have been reported in the literature, and none of these studies have described RCC metastasizing to synchronous pheochromocytoma (PCC). Here, we report a unique case of clear cell renal cell carcinoma (CCRCC) metastasizing to contralateral retroperi
- Citation: Wen HY, Hou J, Zeng H, Zhou Q, Chen N. Tumor-to-tumor metastasis of clear cell renal cell carcinoma to contralateral synchronous pheochromocytoma: A case report. World J Clin Cases 2022; 10(19): 6750-6758
- URL: https://www.wjgnet.com/2307-8960/full/v10/i19/6750.htm
- DOI: https://dx.doi.org/10.12998/wjcc.v10.i19.6750
Renal cell carcinoma (RCC) is one of the most frequently observed malignancies of the urinary system[1]. One-third to one-fourth of RCC patients present with metastasis at the initial diagnosis[1]. RCC often metastasizes to the lungs, liver, lymph nodes, and bones but less frequently to the adrenal glands[1,2].
Tumor-to-tumor metastasis (TTM), in which one primary cancer metastasizes to another primary tumor at a different anatomical site, is uncommon but has been recognized for a long time[3,4]. This phenomenon is different from “collision tumors”, which are defined as two anatomically neighboring neoplasms that directly invade one another[3-5].
Only a few cases in which RCC as a donor tumor of TTM has been reported in literature[6-12] (Table 1), while clear cell renal cell carcinoma (CCRCC) metastasizing to synchronous pheochromocytoma (PCC) has not been described, till date.
| Case number | Year | Donar neoplasm | Recepient neoplasm | Ref. |
| 1 | 2003 | RCC | Oncocytic carcinoma of the thyroid | Ales Ryska et al[6] |
| 2 | 2009 | RCC | Meningioma | Gutiérrez Morales JC et al[7] |
| 3 | 2011 | RCC | pNEN | Michael Cenkowski et al[8] |
| 4 | 2014 | RCC | Breast cancer | Tai-Di Chen et al[9] |
| 5 | 2020 | RCC | Meningioma | Johannes Dietterle et al[10] |
| 6 | 2022 | RCC | pNEN | Shunryo Minezaki et al[11] |
| 7 | 2022 | RCC | Meningioma | Pirlog R et al[12] |
Here, we report a unique case of CCRCC metastasizing to PCC, as shown by molecular characterization using whole genome sequencing (WES), mutational analysis by Sanger sequencing, and fluorescence in situ hybridization (FISH) analysis for chromosome 3p.
A 54-year-old woman presented with recurrent dull abdominal pain for six months, which became aggravated during the last one month.
The patient had recurrent chronic dull pain in her left abdomen for six months without showing any obvious causes. She did not receive any treatment until the pain became unexpectedly aggravated in the last one month, with nausea and difficulty in defecating. Enhanced computed tomography (CT) of the abdomen revealed a tumor in the right kidney and a vast left retroperitoneal mass. The patient was admitted to the West China Hospital, Sichuan University.
The patient did not have a history of previous diseases.
The patient did not have personal or familial risk factors for renal malignancies.
On physical examination, hard, fixed, and tender abdominal masses were palpated in her right and left abdomen. The patient’s blood pressure was normal (116/73 mmHg).
Before surgery, laboratory tests showed that dopamine level was increased to 6.26 nmol/L, which was much higher than the standard value of 0.31 nmol/L. Normetanephrine level was 0.84 nmol/L, much higher than the standard value of 0.71 nmol/L. The 3-methoxytyramine level was increased to 239.82 pg/mL, exceeding the 18.4 pg/mL standard value. The patient’s serum potassium was 3.86 mmol/L, which was within the standard range.
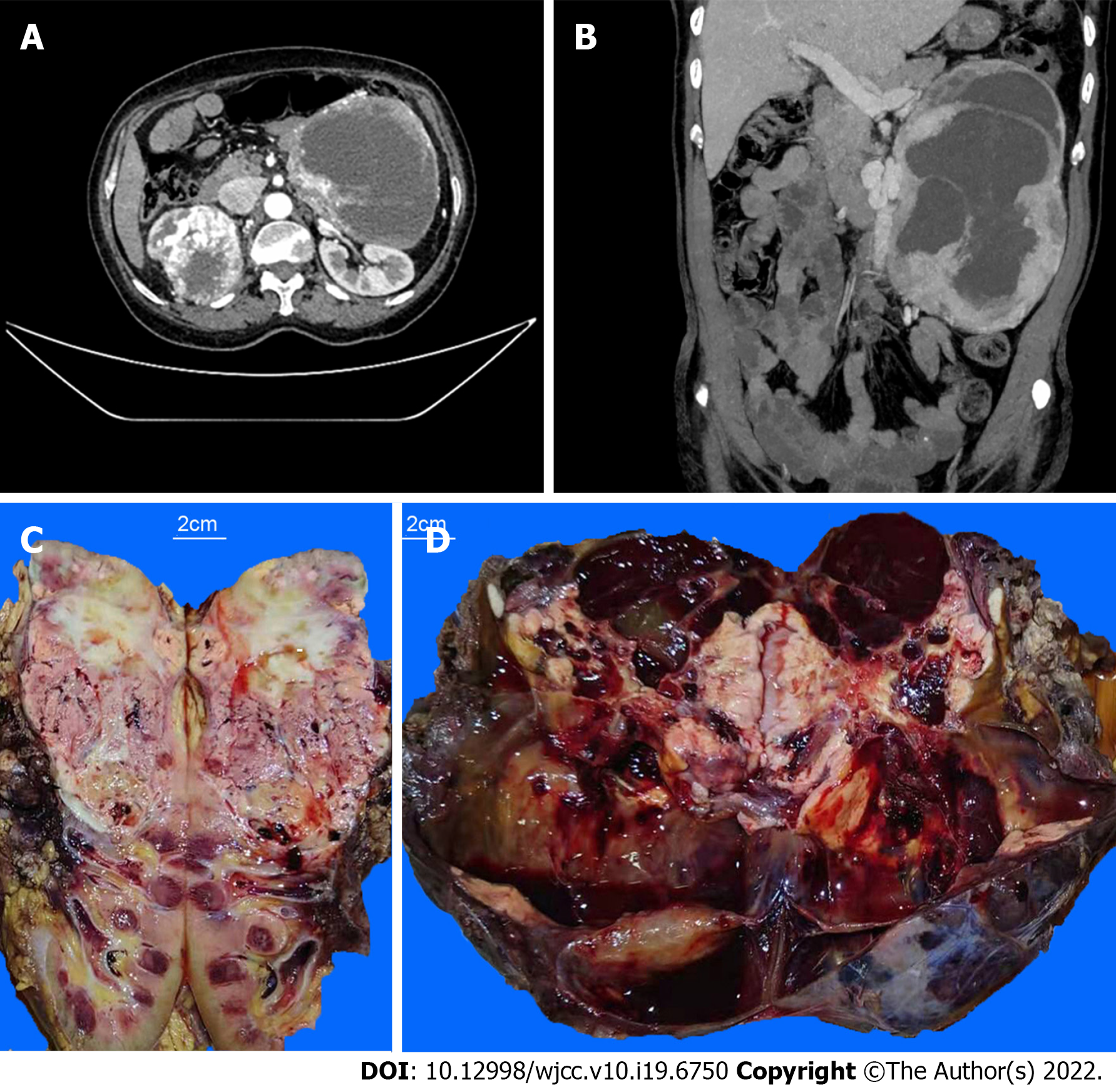
An enhanced computed tomography (CT) scan of the upper and lower abdomen showed a huge solid cyst mass in the patient’s left retroperitoneum. Maximum diameter of the mass was 20.6 cm, which presented apparent intensification with plenty of blood supply. A solid irregular tumor was found in the right kidney with a diameter of 9.2 cm, which showed uneven and apparent high density, with partial necrosis. The vein of the right kidney was strengthened and curved. An irregular, dense nodule was detected in the vein lumen. (Figures 1A and B).
The upper part of the resected right kidney contained a tumor with a maximum diameter of 10.5 cm. The cut surface of the specimen was tan to whitish or yellowish in color, solid, and showed hemorrhage and necrosis. The tumor invaded the capsule of the kidney (Figure 1C). The left retroperitoneal tumor had a maximum diameter of 21.5 cm. The cut section appeared dark red, and cystic changes, hemorrhage, and necrosis were observed. Distinct, confluent whitish to yellowish nodules were observed within the tumor mass (Figure 1D).
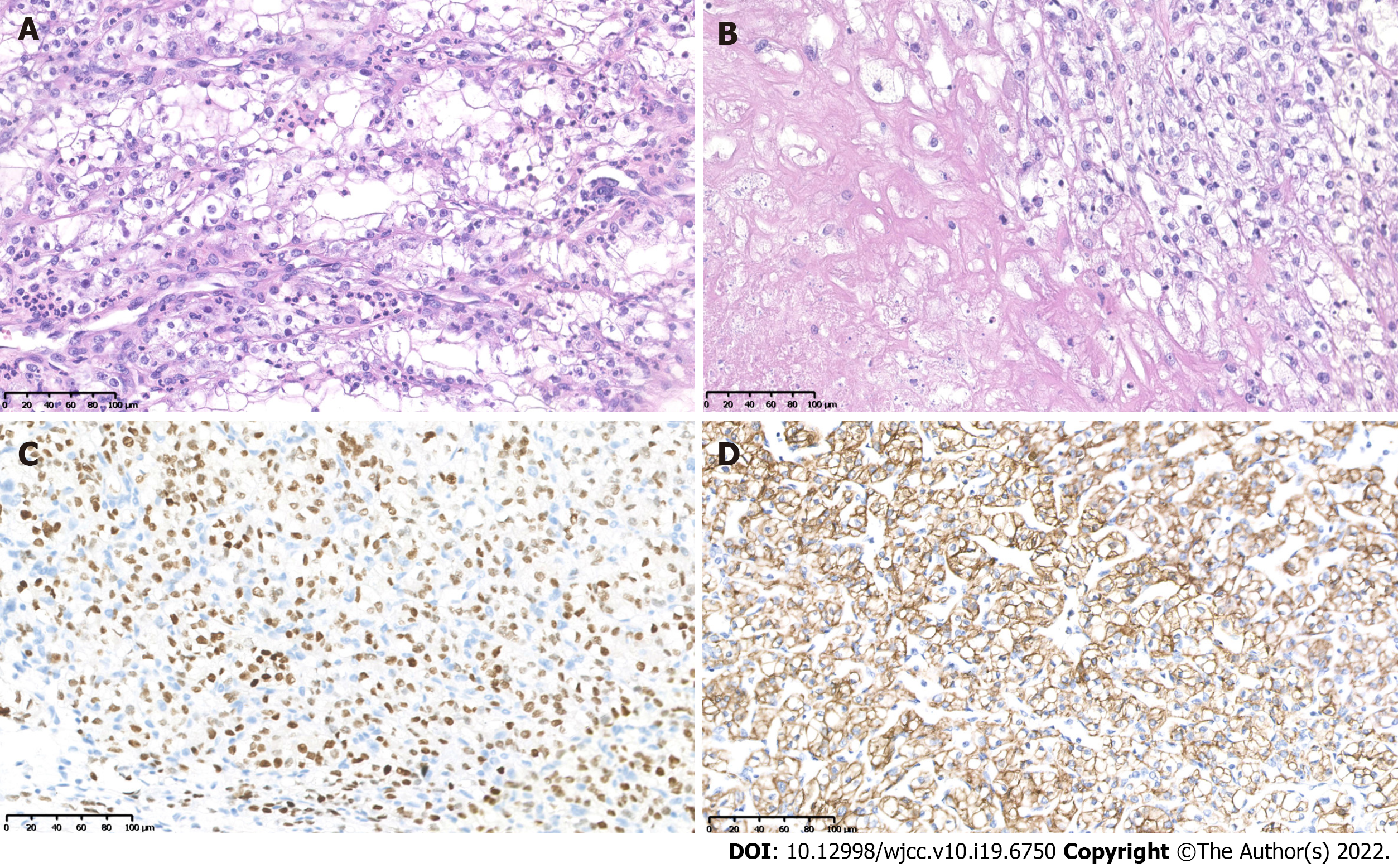
Histopathological examination of the right kidney tumor showed a typical World Health Organization (WHO) grade 3 CCRCC morphology, characterized by clear tumor cells with distinct cell membranes and focal necrosis (Figures 2A and B). Immunohistochemistry (IHC) showed characteristic positivity for PAX8 (Figure 2C), CAIX (Figure 2D), and EMA, as well as intact expression of fumarate hydratase and succinate dehydrogenase (SDHB). The results were negative for CK7 and TFE3 expression. The final diagnosis was CCRCC with WHO/ISUP grade 3.
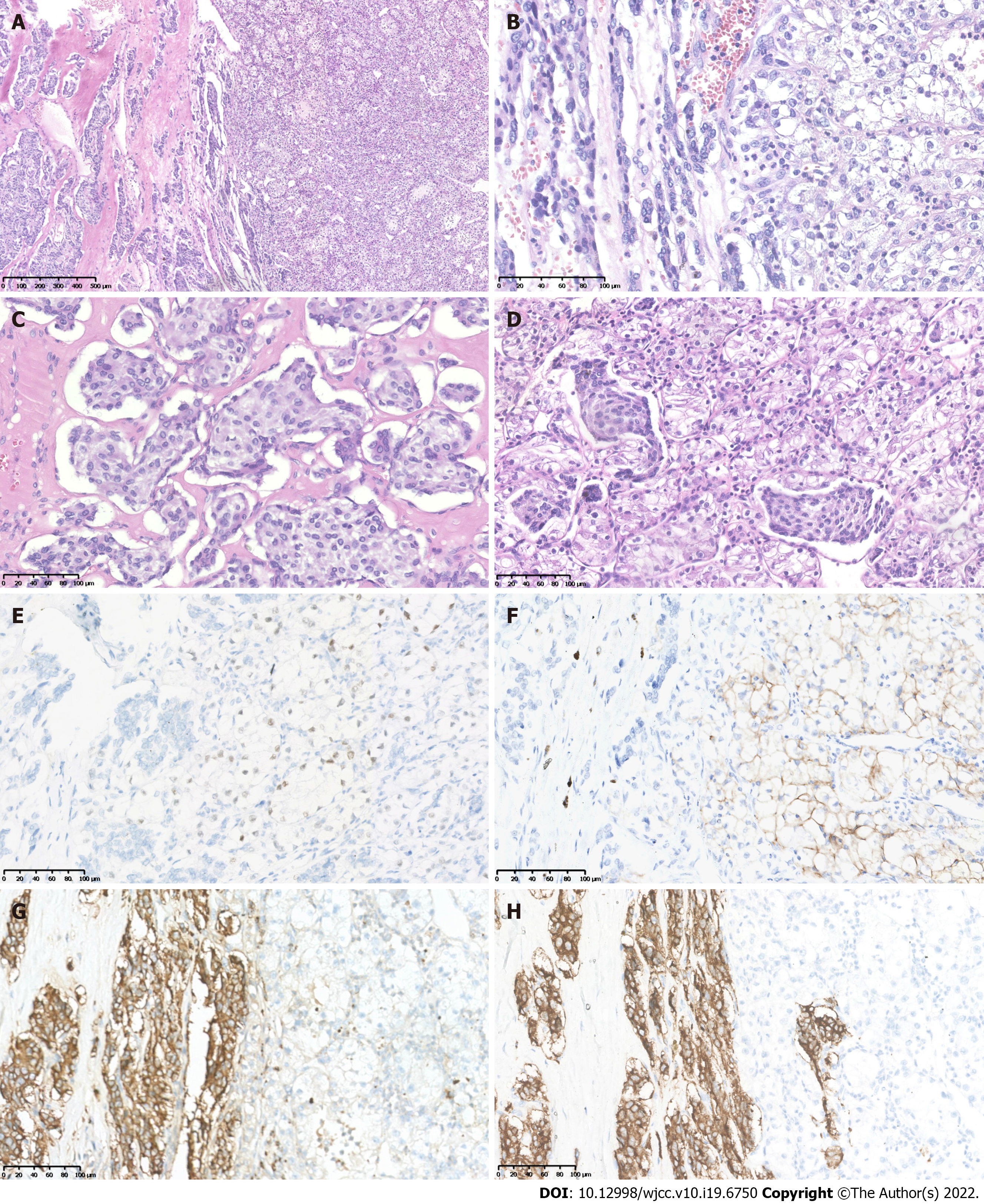
Two components were observed in the left retroperitoneal tumor mass. The main tumor component (grossly dark-red areas) showed typical pheochromocytoma morphology with amphophilic tumor cells arranged in organoid nests, and cords and rich stromal capillary vasculature (Figures 3A and D). IHC showed positive staining for CgA (Figure 3G), Syn (Figure 3H), and intact expression of SDHB, but was negative for PAX8 (Figure 3E), CAIX (Figure 3F), CR, inhibin-α, MART(A103), TFE3, and EMA, with a Ki67 index of approximately 2%. The other tumor component from the whitish to yellowish areas showed CCRCC morphology similar to that of the right kidney tumor (Figure 3A and D), and was immunohistochemically positive for PAX8 (Figure 3E), CAIX (Figure 3F), EMA, and SDHB, but negative for CgA (Figure 3G), Syn (Figure 3H), S-100, CR, inhibin-α, MART-190 (A103), and TFE3, with a Ki67 index of approximately 5%, consistent with the diagnosis of metastatic CCRCC.
WES was first performed on the CCRCC component tissue within the left retroperitoneal PCC, together with normal right kidney tissue, using a HiSeq next-generation sequencer (Illumina, San Diego, CA, United States) at GloriousMed Technology (Beijing, China). DNA extraction and quantification, library preparation, and sequencing were performed in accordance with the manufacturer’s protocol. Next generation sequencing results identified a c.529A>T (p.Arg177Ter) somatic mutation in the VHL gene.
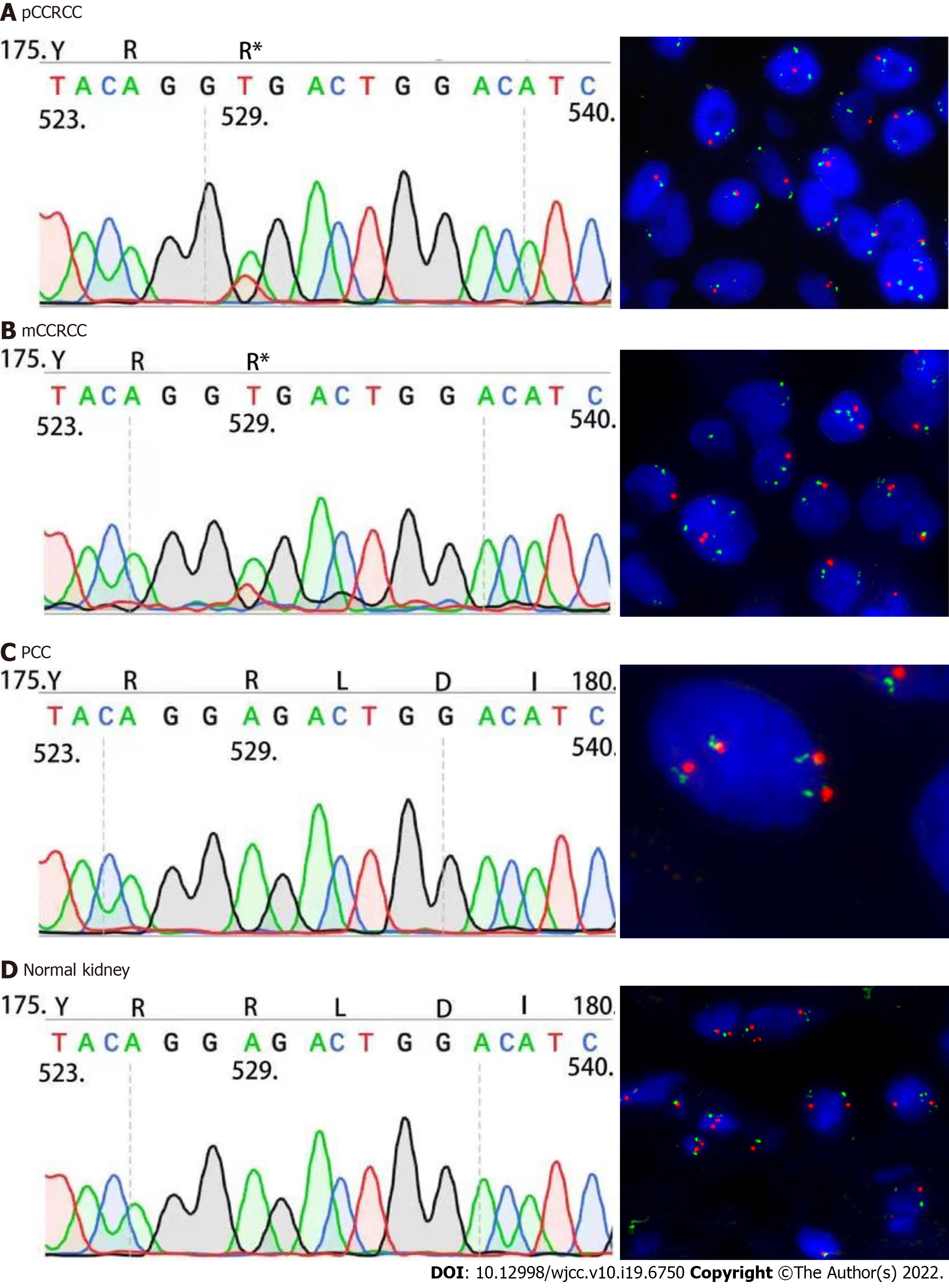
Sanger sequencing was performed using the following primers: 5'- TCT GAA AGA GCG ATG CCT CC-3' (Forward primer) and 5'- TCT CCC ATC CGT TGA TGT GC-3' (Reverse primer). The 169bp PCR products were sequenced in both sense and antisense directions using an automated sequencer (ABI PRISM 3100 Genetic Analyzer; Applied Biosystems, United States). Sanger sequencing was performed on the right kidney CCRCC (Figure 4A), CCRCC tumor component within the left PCC tumor mass (Figure 4B), PCC (Figure 4C), and right normal kidney tissue (Figure 4D). These results confirmed that the right kidney CCRCC (Figure 4A) and left metastatic CCRCC tumor (Figure 4B) components had an identical VHL c.529A>T mutation. This mutation was not detected in the left retroperitoneal PCC (Figure 4C) or in normal tissue samples obtained from the right kidney resection specimen (Figure 4D).
We also analyzed the tissue samples using FISH, which showed a loss of chromosome 3p in the right kidney CCRCC (Figure 4A), as well as in the metastatic CCRCC component within the left PCC (Figure 4B). Loss of 3p was not observed in the left retroperitoneal PCC (Figure 4C) or in normal tissue samples obtained from the right kidney (Figure 4D).
Sequencing and FISH analyses demonstrated that identical molecular features were shared by the primary right kidney CCRCC and the CCRCC component within the left retroperitoneal PCC, further supporting the final diagnosis of the right kidney CCRCC metastasizing to PCC.
Based on the laboratory and imaging examination results, the surgeons initially diagnosed primary synchronous tumors of the right CCRCC and left retroperitoneal PCC. The above histopathological, immunohistochemical, and molecular analyses suggested that the final diagnosis was right kidney CCRCC that metastasized to the left PCC.
In light of the results of laboratory and image analyses, clinicians considered the left retroperitoneal mass was PCC. The patient was administered phenoxybenzamine hydrochloride tablets at a dose of 10 mg twice daily for seven days before surgery. Surgical resection was performed to excise both the tumor masses, and the patient received sunitinib treatment after surgery.
No recurrence or metastasis was observed in the six-month follow-up period up to the time of manuscript preparation.
Tumor-to-tumor metastasis (TTM) is a rare phenomenon that requires at least two separate primary tumors, with the donor tumor metastasizing to the parenchyma of the recipient tumor[4]. Collision tumors, tumor spread to lymph nodes, or tumor thrombosis cannot be considered tumor-to-tumor metastasis[4].
The most frequent TTM donor tumors include lung cancer (40%-50%), breast cancer, prostate cancer, renal cancer, gallbladder carcinoma, melanoma, and major salivary gland cancers[3]. Involvement of solid tumors in hematopoietic neoplasms has also been reported. The most common recipient tumors in TTM include RCCs (40%-70%), sarcomas, meningiomas, thyroid tumors, and pituitary adenomas[3,13]. CCRCC is a more frequent recipient of TTM, presumably due to the rich amount of glycogen and lipids, which creates a nutrient-rich microenvironment for the homing of metastatic tumor cells[14,15]. The rich vasculature of CCRCC may provide a basis for a large number of circulating neoplastic cells shed from the donor cancer to enter the recipient tumor[14,15].
Synchronous or metachronous PCC and RCC may suggest VHL syndrome/VHL disease[16-20]. The present case lacked germline mutations, but carried a somatic c.529A>T mutation. Somatic mutation of the tumor suppressor gene VHL[16-20] has been reported in as many as 80-90% of sporadic CCRCCs[20]. Chromosome 3p loss may also occur in up to 90% of sporadic CCRCCs[21].
The c.529A>T nonsense mutation in the VHL gene leads to a change in the codon for the Arg 177 residue to a stop codon (Arg177Ter), resulting in premature termination of translation within the α domain and a truncated, non-functional VHL protein with a disrupted α domain; functional VHL is required for interaction with elongin that is important for HIF1α polyubiquitination and degradation[18-21]. Inactivation of VHL and loss of chromosome 3p are major oncogenic driver events in CCRCCs[18-21]. c.529A>T mutation and consequent Arg177Ter change do not occur frequently in CCRCC but have been reported in CCRCC associated with VHL disease[22]. A VHL-associated CCRCC case in which a deletion of c.530-536 resulted in an Arg177Thr change has also been reported[23]. In addition, one study reported a rare VHL somatic c.529A>T mutation in a MEN1-associated metastatic pancreatic neuroendocrine tumor[24].
Although ipsilateral and metachronous CCRCC and PCC have been reported in the literature[16], there are only three reported cases of synchronous bilateral CCRCCs and PCCs[16,17], in which VHL gene mutations or related family history have not been found[16,17,25].
Imaging techniques may be beneficial for initial diagnosis of multiple tumors. Patients with multiple primary tumors should undergo 18F-fluorodeoxyglucose (F-FDG) positron emission tomography (PET)/CT to detect a wide range of unexpected malignant tumors[26]. Some special imaging techniques are particularly helpful for diagnosing CCRCC and PCC. A recent study shows that MRI is a feasible tool for the diagnosis of CCRCC and may predict the Fuhrman grade of CCRCC[27]. Radiotracers such as radioiodine labelling metaiodoenzylguanidine or 68Ga labelling somatostatin analogs could help diagnose PCC[28,29]. In our study, the patient showed typical imaging features of CCRCC and PCC on an enhanced CT scan, but regrettably she refused to undergo PET/CT and MIGB examinations, although the surgeon recommended these tests. In the present case, although examination of images was very beneficial for diagnosing this condition before surgery, the rare diagnosis of TTM was still based on pathological investigations.
The present study reports a unique case of a patient with right kidney CCRCC that metastasized to left retroperitoneal PCC. Both the primary and metastatic CCRCC carried a rare c.529A>T mutation and an Arg177Ter change in the VHL protein. These findings expand the scope of tumor-to-tumor metastasis.
Provenance and peer review: Unsolicited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Medicine, research and experimental
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Maurea S, Italy; Zavaleta MJC, Peru S-Editor: Wang LL L-Editor: A P-Editor: Wang LL
| 1. | Bukowski RM. Natural history and therapy of metastatic renal cell carcinoma: the role of interleukin-2. Cancer. 1997;80:1198-1220. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 2. | von Knobloch R, Schrader AJ, Walthers EM, Hofmann R. Simultaneous adrenalectomy during radical nephrectomy for renal cell carcinoma will not cure patients with adrenal metastasis. Urology. 2009;73:333-336. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 10] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 3. | Bartoš V. Tumor-to-Tumor Metastasis - a Unique Case of Clear Cell Renal Cell Carcinoma Harboring Metastasis of Adenocarcinoma of Unknown Origin. Klin Onkol. 31:366-370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 4. | Campbell LV Jr, Gilbert E, Chamberlain CR Jr, Watne AL. Metastases of cancer to cancer. Cancer. 1968;22:635-643. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |
| 5. | Wimmer JL, Coffey DM, Kaplan AL, Ayala AG, Ro JY. Tumor-to-tumor metastasis with endometrial carcinoma metastatic to squamous cell carcinoma of vulva: the first reported case. Arch Pathol Lab Med. 2013;137:1825-1828. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 6. | Ryska A, Cáp J. Tumor-to-tumor metastasis of renal cell carcinoma into oncocytic carcinoma of the thyroid. Report of a case and review of the literature. Pathol Res Pract. 2003;199:101-106. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 14] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 7. | Gutiérrez Morales JC, Gutiérrez Morales SE, Astudillo González A. 72 year-old man with new seizure while dancing. Brain Pathol. 2009;19:347-348. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 8. | Cenkowski M, Gibson IW, Lategan B, Czaykowski PM. Tumor-to-tumor metastasis: report of a case of renal cell carcinoma metastasizing to a pancreatic endocrine neoplasm. J Clin Oncol. 2011;29:e303-e304. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 10] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 9. | Chen TD, Lee LY. A case of renal cell carcinoma metastasizing to invasive ductal breast carcinoma. J Formos Med Assoc. 2014;113:133-136. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 10. | Dietterle J, Frydrychowicz C, Müller W, Hoffmann KT, Jähne K, Meixensberger J. Tumor-to-Tumor Metastasis of Multiple Meningiomas and Clear Cell Renal Cell Carcinoma Metastasis as First Clinical Appearance of Kidney Cancer: A Case Report and Analysis. J Neurol Surg Rep. 2020;81:e10-e14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 11. | Minezaki S, Misawa T, Tsukayama H, Shibuya M, Wada K, Sano K, Mochizuki M, Sasajima Y, Kondo H. Tumor-to-tumor metastasis: an extremely rare combination with renal cell carcinoma as the donor and a pancreatic neuroendocrine tumor as the recipient. Surg Case Rep. 2022;8:8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 12. | Pirlog R, Sirbu OM, Laquerrière A, Billaud-Porte E, Curey S, Lozouet M, Marguet F, Derrey S. Tumor-to-tumor metastases: Latent renal cell carcinoma discovered after elective surgical resection of a convexity meningioma. Neurochirurgie. 2022;68:196-201. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 13. | Sella A, Ro JY. Renal cell cancer: best recipient of tumor-to-tumor metastasis. Urology. 1987;30:35-38. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 95] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 14. | Shin T, Kan T, Sato F, Mimata H. Tumor-to-Tumor Metastasis to Chromophobe Renal Cell Carcinoma: A First Report. Case Rep Urol. 2011;2011:520839. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 8] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 15. | Huo Z, Gao Y, Yu Z, Zuo W, Zhang Y. Metastasis of breast cancer to renal cancer: report of a rare case. Int J Clin Exp Pathol. 2015;8:15417-15421. [PubMed] |
| 16. | Bahrami A, Truong LD, Shen SS, Krishnan B. Synchronous renal and adrenal masses: an analysis of 80 cases. Ann Diagn Pathol. 2009;13:9-15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 15] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 17. | Chen HH, Wu ST, Lin YC, Lin CS. Synchronous renal cell carcinoma and pheochromocytoma presenting as acute decompensated heart failure. J Postgrad Med. 2019;65:44-46. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 18. | Chittiboina P, Lonser RR. Von Hippel-Lindau disease. Handb Clin Neurol. 2015;132:139-156. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 125] [Cited by in RCA: 132] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 19. | Lonser RR, Glenn GM, Walther M, Chew EY, Libutti SK, Linehan WM, Oldfield EH. von Hippel-Lindau disease. Lancet. 2003;361:2059-2067. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1083] [Cited by in RCA: 1012] [Article Influence: 46.0] [Reference Citation Analysis (0)] |
| 20. | Arjumand W, Sultana S. Role of VHL gene mutation in human renal cell carcinoma. Tumour Biol. 2012;33:9-16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 68] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 21. | Hsieh JJ, Le VH, Oyama T, Ricketts CJ, Ho TH, Cheng EH. Chromosome 3p Loss-Orchestrated VHL, HIF, and Epigenetic Deregulation in Clear Cell Renal Cell Carcinoma. J Clin Oncol. 2018;JCO2018792549. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 125] [Article Influence: 17.9] [Reference Citation Analysis (0)] |
| 22. | Maher ER, Webster AR, Richards FM, Green JS, Crossey PA, Payne SJ, Moore AT. Phenotypic expression in von Hippel-Lindau disease: correlations with germline VHL gene mutations. J Med Genet. 1996;33:328-332. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 143] [Cited by in RCA: 145] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 23. | Wang J, Cao W, Wang Z, Zhu H. Novel gene mutation in von Hippel-Lindau disease - a report of two cases. BMC Med Genet. 2019;20:194. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 24. | Shell J, Patel D, Powers A, Quezado M, Killian K, Meltzer P, Zhu J, Gaitanidis A, Karzai F, Neychev V, Green P, Kebebew E. Somatic VHL Mutation in a Patient With MEN1-Associated Metastatic Pancreatic Neuroendocrine Tumor Responding to Sunitinib Treatment: A Case Report. J Endocr Soc. 2017;1:1124-1134. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 25. | Fairchild RS, Kyner JL, Hermreck A, Schimke RN. Neuroblastoma, pheochromocytoma, and renal cell carcinoma. Occurrence in a single patient. JAMA. 1979;242:2210-2211. [PubMed] |
| 26. | Klain M, Maurea S, Gaudieri V, Zampella E, Volpe F, Manganelli M, Piscopo L, De Risi M, Cuocolo A. The diagnostic role of total-body 18F-FDG PET/CT in patients with multiple tumors: a report of the association of thyroid cancer with lung or renal tumors. Quant Imaging Med Surg. 2021;11:4211-4215. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 27. | Stanzione A, Ricciardi C, Cuocolo R, Romeo V, Petrone J, Sarnataro M, Mainenti PP, Improta G, De Rosa F, Insabato L, Brunetti A, Maurea S. MRI Radiomics for the Prediction of Fuhrman Grade in Clear Cell Renal Cell Carcinoma: a Machine Learning Exploratory Study. J Digit Imaging. 2020;33:879-887. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 32] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 28. | Maurea S, Caracò C, Klain M, Mainolfi C, Salvatore M. Imaging characterization of non-hypersecreting adrenal masses. Comparison between MR and radionuclide techniques. Q J Nucl Med Mol Imaging. 2004;48:188-197. [PubMed] |
| 29. | Lastoria S, Maurea S, Vergara E, Acampa W, Varrella P, Klain M, Muto P, Bernardy JD, Salvatore M. Comparison of labeled MIBG and somatostatin analogs in imaging neuroendocrine tumors. Q J Nucl Med. 1995;39:145-149. [PubMed] |