Published online Jan 15, 2024. doi: 10.4251/wjgo.v16.i1.30
Peer-review started: July 29, 2023
First decision: September 25, 2023
Revised: October 7, 2023
Accepted: November 2, 2023
Article in press: November 2, 2023
Published online: January 15, 2024
Processing time: 165 Days and 22.5 Hours
Pachymic acid (PA) is derived from Poria cocos. PA has a variety of pharmacological and inhibitory effects on various tumors. However, the mechanism of action of PA in gastric cancer (GC) remains unclear.
To investigate the mechanism of PA in treating GC via the combination of network pharmacology and experimental verification.
The GeneCards and OMIM databases were used to derive the GC targets, while the Pharm Mapper database provided the PA targets. Utilizing the STRING database, a protein-protein interaction network was constructed and core targets were screened. The analyses of Gene Ontology, Kyoto Encyclopedia of Genes and Genomes (KEGG), and gene set enrichment analysis were conducted, and molecular docking and clinical correlation analyses were performed on the core targets. Ultimately, the network pharmacology findings were validated through in vitro cell assays, encompassing assessments of cell viability, apoptosis, cell cycle, cloning, and western blot analysis.
According to network pharmacology analysis, the core targets were screened, and the PI3K/AKT signaling pathway is likely to be the mechanism by which PA effectively treats GC, according to KEGG enrichment analysis. The experimental findings showed that PA could control PI3K/AKT signaling to prevent GC cell proliferation, induce apoptosis, and pause the cell cycle.
Network pharmacology demonstrated that PA could treat GC by controlling a variety of signaling pathways and acting on a variety of targets. This has also been supported by in vitro cell studies, which serve as benchmarks for further research.
Core Tip: Pachymic acid (PA) is an important bioactive component of Poria cocos. Network pharmacology analysis showed that the core targets of PA in treating gastric cancer (GC) were proto-oncogene tyrosine-protein kinase Src, mitogen-activated protein kinase 1, phosphatidylinositol 3-kinase regulatory subunit alpha, heat shock protein 90-alpha, and tyrosine-protein phosphatase non-receptor type 11. Molecular docking results showed that PA could combine well with the core targets. Gene Ontology, Kyoto Encyclopedia of Genes and Genomes, and gene set enrichment analysis suggested that the PI3K/AKT signaling pathway was possible for treating GC with PA. The experiment results demonstrated that PA could decrease the survival rate of HGC-27 cells, halt the progression of the cell cycle, enhance cell apoptosis, control the PI3K/AKT signaling pathway to stimulate apoptosis, and impede cell growth.
- Citation: Du YH, Zhao JJ, Li X, Huang SC, Ning N, Chen GQ, Yang Y, Nan Y, Yuan L. Mechanism of pachymic acid in the treatment of gastric cancer based on network pharmacology and experimental verification. World J Gastrointest Oncol 2024; 16(1): 30-50
- URL: https://www.wjgnet.com/1948-5204/full/v16/i1/30.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v16.i1.30
As a malignant digestive tract tumor that arises from gastric epithelial cells, gastric cancer (GC) is one of the most common causes of cancer-related deaths worldwide. It is the fifth most frequent malignant tumor[1]. It is unusual for patients with early GC to present obvious symptoms. The middle and late stages may present with weight loss, upper abdominal pain, and appetite loss[2].
GC is usually diagnosed at an advanced stage. The current treatment methods include surgery, radiotherapy, chemotherapy and targeted drugs[3,4]. Surgical treatment may not be suitable for patients with advanced disease. Surgery is associated with poor prognosis. Radiotherapy and chemotherapy have certain curative effects but have significant side effects and are easily resistant to drugs[5,6]. Identifying drugs that effectively treat GC is therefore urgently needed. The treatment of tumors with natural pharmaceutical ingredients has made great advances[7]. A rising number of clinical investigations have focused on the anti-tumor properties of traditional Chinese medicine or its constituent parts[8,9]. In traditional Chinese medicine, Poria cocos is regarded as a pharmacologically beneficial food with sedative, diuretic, and tonic effects and can also be used to treat dyspepsia and other gastrointestinal disorders[10-12]. It contains pachymic acid (PA), a triterpenoid found naturally in Poria cocos.
PA exerts various pharmacological effects in recent studies. According to Huang et al[13] PA exerts a hypoglycemic effect by enhancing the expression of glucose transporter type 4 to facilitate glucose uptake. Feng et al[14] revealed that PA reduced the levels of interleukin IL-6, IL-1, and lactate dehydrogenase in the bladder, down-regulated tumor necrosis factor-α, and up-regulated TP53 protein in bladder samples to produce anti-inflammatory effects in an animal model. In mice treated with pentobarbital, PA also inhibited motor activity via chloride channel activation and GABAergic mechanisms, prolonged sleep duration, and enhanced hypnotic effects[15]. Furthermore, PA has anti-inflammatory, hypoglycemic, sedative, and hypnotic effects as well as antitumor properties, and can significantly inhibit lung cancer[16], pancreatic cancer[17], breast cancer[18], ovarian cancer[19], GC[20] and other tumor cells, as well as cell proliferation, cell cycle, apoptosis, and other phenotypes.
Our team previously reported that 18β-glycyrrhetinic acid, a natural drug component, affects the proliferation and metastasis of GC[21]. PA also appears to be useful as a natural product for the treatment of GC. Earlier research has indicated that PA hinders the proliferation and capacity of GC cells to form colonies while also causing a notable halt in the G2/M phase and triggering apoptosis. As a result, it effectively restrains the progression of GC[20]. Previous research indicates that PA has the ability to curb the migration and metastasis ability of GC cells by reducing the matrix metalloproteinase proteins expression and suppressing the epithelial-mesenchymal transition process. SGC-7901 tumor growth in nude mice was significantly suppressed by PA[22]. In addition, PA can increase the sensitivity of GC cells by upregulating BAX protein expression[23].
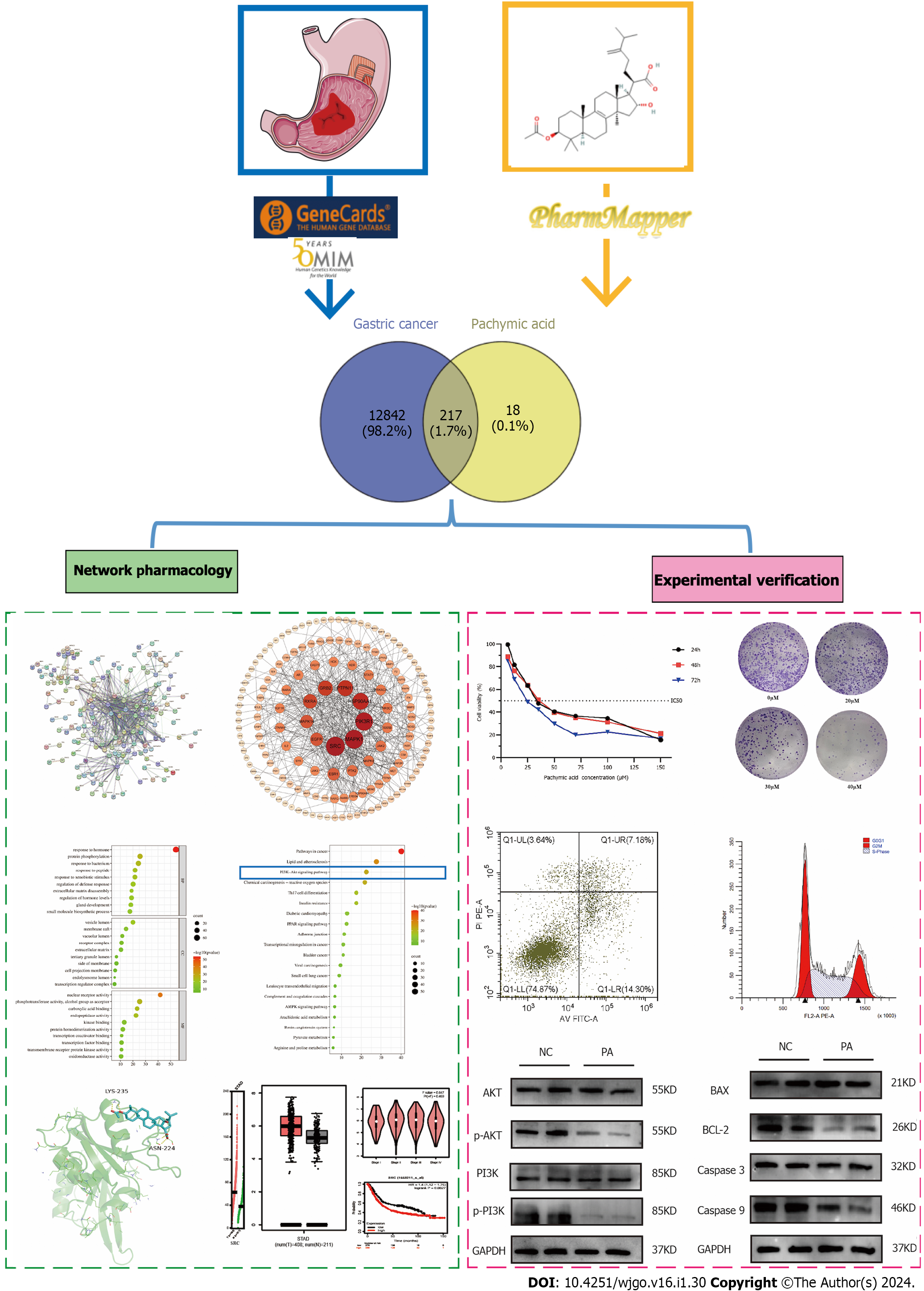
At present, limited information is available regarding the utilization of PA in the treatment of GC, and the underlying mechanism behind this phenomenon is not yet understood. In this work, we examined the healing effect of PA on GC. Further, we studied its underlying mechanisms using network pharmacology, bioinformatic analysis, and experimental validation in the human GC cell line HGC-27 (Figure 1). This study may offer new perspectives on the treatment of PA in GC.
The composition of Poria cocos can be found in the TCMSP database (https://old.tcmsp-e.com/tcmsp.php). By accessing the live 3D structure of PA from PubChem (https://pubchem.ncbi.nlm.nih.gov/), the related targets of PA were obtained by 3D structure on PharmMapper (http://www. lilab-ecust.cn/pharmmapper/). Targets for GC are available from GeneCards (http://www. genecards. org) and the OMIM database (http://omim.org/). Finally, we obtained targets for common targets of GC and PA used Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/index.html).
Set the parameters and import the common target into the STRING database (https://string-db.org) and export TSV format files. Cytoscape3.8.2 was used for visualization and arranged according to degree, with the depth of color and size of the dots representing the magnitude of the degree value. Core targets were chosen based on degree values of > 10.
We conducted analyses for biological process (BP), cellular component (CC), and molecular function as part of the Gene Ontology (GO) analysis. According to the -log10 (P value), an online platform for data analysis and visualization (https://www.bioinformatics.com.cn/) was used for the top 10 for visual analysis. The top 20 Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed for visual analysis. The network diagram of the pathway and the top 12 core targets were performed using cytoscape3.8.2 according to the degree value. GEO in the NCBI database (https://www.ncbi.nlm.nih.gov/geo/) to download GES79973 chip original gene expression data (data from the expression of GC and normal tissue), using the Weishengxin website (https://www.bioinformatics.com.cn), gene set enrichment analysis (GSEA) analysis results are obtained.
Using the PubChem database, the 2D structure of PA was downloaded and imported into the Chem3D software to transform and optimize it into a 3D structure. The RCSB PDB database (http://www.rcsb.org/) was used to download the 3D structure of the core target, which was then imported into PyMOL for processing using the species set for humans. Subsequently, the ligands and receptors were processed using the AutoDockTools program, molecular docking was performed using the AutoDockVina program, and the outcomes were displayed using the PyMOL program.
To acquire the expression levels of the leading ten core genes in GC and normal tissues, as well as to examine the variation in expression levels and copy number at various stages, the Gene Expression Profiling Interactive Analysis (GEPIA) database (http://GEPIA.cancer-pku.cn//) was utilized. We utilized the Kaplan-Meier plotter database (http://kmplot.com/analysis/) to examine the impact of ten genes on the prognosis of GC patients.
MCE provided PA (CAT: HY-N0371, United States). Punosai Co., Ltd. provided HGC-27 (CAT: CL-0107, China) and GES-1 (CAT: CL-0563, China) cells, which were confirmed by STR to be free of cross-contamination. Gemini was used to sell fetal bovine serum (CAT: 900108, China). Gibco provided RPMI-1640 medium (CAT: 1187517, United States). Dojindo Cell Counting Kit-8 (CAT: CK04, Japan). The Annexin-V FITC/PI double-staining apoptosis detection kit (CAT: KGA1023; China) and cell cycle detection kit (CAT: KGA511, China) were provided by Jiangsu KeyGEN Bio TECH Corp., Ltd. Antibodies against PI3K (CAT: ab133595, United Kingdom), p-PI3K (CAT: ab278545, United Kingdom), BAX (CAT: ab32503, United Kingdom), BCL-2 (CAT: ab182858, United Kingdom), Caspase 3 (CAT: ab32351, United Kingdom), Caspase 9 (CAT: ab32539, United Kingdom), and GADPH (CAT: ab8245, United States) were purchased from Abcam. Antibodies against AKT (CAT: AF6261, China) and p-AKT (CAT: AF0016, China) were purchased from Affinity.
HGC-27 cells were cultured in RPMI-1640 medium containing 10% fetal calf serum and 1% mixture of penicillin-streptomycin, whereas GES-1 cells were cultured in DMEM medium containing 10% fetal calf serum and 1% mixture of penicillin-streptomycin. The culture was maintained at 37 °C with 5% CO2.
The HGC-27 and GES-1 cells (6 × 103 cells per well) were seeded in 96-well plates and treated with various doses of PA. (6.25 μM, 12.5 μM, 25 μM, 35 μM, 50 μM, 70 μM, 100 μM, 150 μM) in the incubator at 37 °C for 24 h, 48 h, and 72 h. Each group consisted of three replicates. After incubation, 10 μL of CCK-8 was added to every 100 μL of the medium for 2 h, and the OD was measured at 450 nm.
According to the cell counting results, 800 cells per well were inoculated into 6-well plates, and three duplicate wells were established. The cells were then cultured for 14 d. When each clone contained more than 50 cells, 1 mL 4% paraformaldehyde and 500 μL crystal violet solution were added. The staining process was carried out for 15 min at ambient temperature, the staining solution was washed, dried at room temperature, photographed, and analyzed. Clones had a diameter greater than 1 mm were counted.
The cells were seeded in cell culture dishes, the supernatant was removed, and drug-containing medium was added. Each group was set up in triplicate. After a 24 h drug intervention, the cells were digested and washed in prechilled phosphate buffered saline (PBS). They were then supplemented with 500 μL binding buffer, 5 μL annexin V-FITC and 5 μL propidium iodide. Finally, they were left to respond in the dark for 10 min. The analysis of flow cytometry was finished.
Different concentrations of drug-containing media were introduced after the cells had been cultured under optimal growth conditions for a while, and each group was set up in triplicates. After the 24 h intervention period, the cells were removed, cleaned with PBS, and fixed with 70% ethanol at 4 °C. After washing the fixative with PBS and adding 500ul of cell cycle working solution (RNase A/PI = 1:9) to each well, the reaction was performed in the dark for 30-60 min. The proportion of cells in each cycle was calculated using flow cytometry.
Total protein was extracted from normal HGC-27 cells and PA-treated cells. The BCA assay was performed to assess protein concentration. After the total proteins were separated by electrophoresis, the target proteins were transferred to polyvinylidene fluoride (PVDF) membranes and blocked with skim milk powder for 2 h. After washing with TBST, the culture plates were incubated with primary antibodies overnight at a temperature of 4 °C. After washing with TBST, PVDF membranes were exposed to appropriate secondary antibodies and incubated for 1 h at room temperature. Finally, an ultrasensitive multifunctional imager and a luminous solution were used for detection[24].
All data were statistically analyzed using GraphPad Prism 8. Data were shown as mean ± SD, and the differences between different groups were analyzed using a one-way ANOVA or t-test. The level of significance was set at P < 0.05.
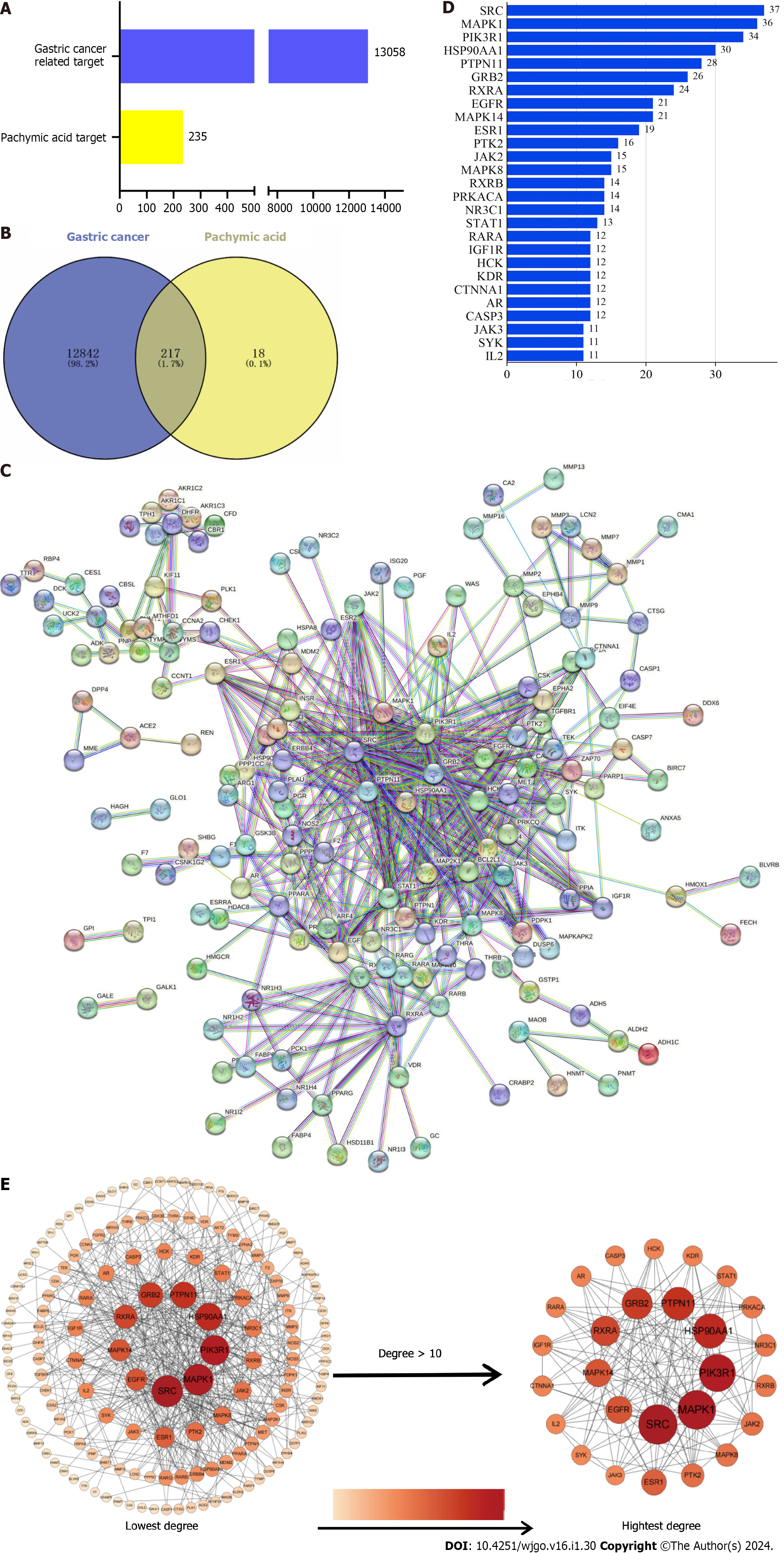
Among the components of Poria cocos found in the TCMSP database, oral bioavailability = 33.63 and drug-likeliss = 0.81 for PA (Table 1 and Supplementary Figure 1). The PA targets were obtained from the PharmMapper database and standardized using the Uniport database to remove duplicates and non-human targets. In total, 235 PA targets were identified. 12706 GC targets were obtained by eliminating duplicates from the Genecards database and 506 GC targets were obtained by deleting duplicates from the OMIM database (Figure 2A). Duplicate GC entries in the two databases were merged and deleted to obtain 13058 GC targets. The 235 targets of PA and 13058 targets of GC were interposed to obtain 217 possible targets of PA for GC treatment (Figure 2B).
| ID | Phytochemical name | OB | DL |
| MOL000273 | 16alpha-Hydroxydehydrotrametenolic acid | 30.93 | 0.81 |
| MOL000274 | 3β-hydroxylanosta-7,9(11),24-trien-21-oic acid | 24.92 | 0.8 |
| MOL000275 | Trametenolic acid | 38.71 | 0.8 |
| MOL000276 | 7,9(11)-dehydropachymic acid | 35.11 | 0.81 |
| MOL000277 | Tumulosic acid | 15.95 | 0.81 |
| MOL000278 | Beta-glucan | 0.73 | 0.7 |
| MOL000279 | Cerevisterol | 37.96 | 0.77 |
| MOL000280 | Dehydrotumulosic acid | 31.07 | 0.82 |
| MOL000281 | Dimethyl L-malate | 8.59 | 0.03 |
| MOL000282 | Stellasterol | 43.51 | 0.72 |
| MOL000283 | Ergosterol peroxide | 40.36 | 0.81 |
| MOL000284 | L-uridine | 23.4 | 0.11 |
| MOL000285 | (2R)-2-[(5R,10S,13R,14R,16R,17R)-16-hydroxy-3-keto-4,4,10,13,14-pentamethyl-1,2,5,6,12,15,16,17-octahydrocyclopenta[a]phenanthren-17-yl]-5-isopropyl-hex-5-enoic acid | 38.26 | 0.82 |
| MOL000286 | β-amyrin acetate | 9.11 | 0.74 |
| MOL000287 | 3-beta-hydroxy-24-methylene-8-lanostene-21-oic acid | 38.7 | 0.81 |
| MOL000288 | Pachyman | 0.45 | 0.68 |
| MOL000289 | Pachymic acid | 33.63 | 0.81 |
| MOL000290 | Poricoic acid A | 30.61 | 0.76 |
| MOL000291 | Poricoic acid B | 30.52 | 0.75 |
| MOL000292 | Poricoic acid C | 38.15 | 0.75 |
| MOL000293 | Poricoic acid D | 22.38 | 0.78 |
| MOL000294 | Poricoic acid DM | 29.32 | 0.78 |
| MOL000295 | Alexandrin | 20.63 | 0.63 |
| MOL000296 | Hederagenin | 36.91 | 0.75 |
| MOL000297 | Tumulosic acid | 29.88 | 0.81 |
| MOL000298 | Ergosterol | 14.29 | 0.72 |
| MOL000299 | Trimethyl citrate | 67.61 | 0.07 |
| MOL000300 | Dehydroeburicoic acid | 44.17 | 0.83 |
| MOL000301 | 2-lauroleic acid | 31.42 | 0.04 |
| MOL000302 | Undekansaeure | 30.14 | 0.03 |
| MOL000303 | Caprylic acid | 16.4 | 0.02 |
| MOL000304 | Ethyl glucoside | 16.4 | 0.02 |
| MOL000305 | Lauric acid | 23.59 | 0.04 |
| MOL000069 | Palmitic acid | 19.3 | 0.1 |
To create the protein-protein interaction network diagram, 217 intersection targets from the STRING database of PA and GC were imported (Figure 2C). The results were imported into Cytospace3.8.2 for analysis and arranged according to the degree value. The larger the degree value, the darker the color, the larger the circle, and the closer it is to the center. The top 27 core targets were selected based on a degree value of > 10 (Figure 2D). The degree values of the 27 core targets were arranged as bars (Figure 2E). The top 10 targets for the degree value were proto-oncogene tyrosine-protein kinase Src (SRC), mitogen-activated protein kinase 1 (MAPK1), phosphatidylinositol 3-kinase regulatory subunit alpha (PIK3R1), heat shock protein 90-alpha (HSP90AA1), tyrosine-protein phosphatase non-receptor type 11 (PTPN11), growth factor receptor-bound protein 2 (GRB2), retinoic acid receptor RXR-alpha (RXRA), mitogen-activated protein kinase 14 (MAPK14), epidermal growth factor receptor (EGFR), and estrogen receptor (ESR1).
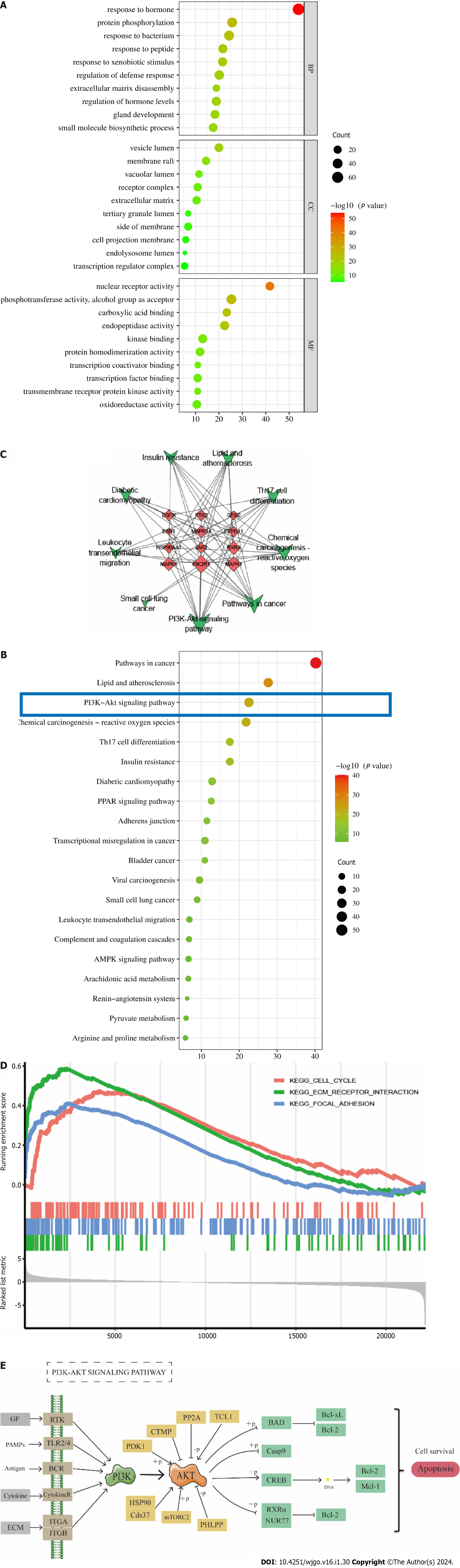
For GO and KEGG analyses, common targets were loaded into Metascape. GO enrichment analysis showed that PA treatment of GC affected response to hormone, protein phosphorylation, response to bacterium, response to peptide, response to xenobiotic stimulus, and other BPs. CCs include vesicle lumen, membrane raft, vacuolar lumen, receptor complex, and extracellular matrix. Molecular features included nuclear receptor activity, phosphotransferase activity, alcohol group as acceptor, carboxylic acid binding, endopeptidase activity, and kinase binding (Figure 3A). The top 20 most important pathways were analyzed using KEGG enrichment. PA exerted anti-GC effects through cancer pathways and the PI3K-Akt signaling pathway (Figure 3B). The top 12 targets were mainly enriched in the PI3K-Akt signaling pathway (Figure 3C). The GSEA results showed that genes involved in the cell cycle, ECM receptor interaction, and focal adhesion pathways were significantly downregulated in GC (Figure 3D). The findings from the analysis of KEGG, GO, and GSEA, along with the discovery of key targets in the PPI network, indicate that the PI3K/AKT signaling pathway could potentially be utilized for treating GC using PA.
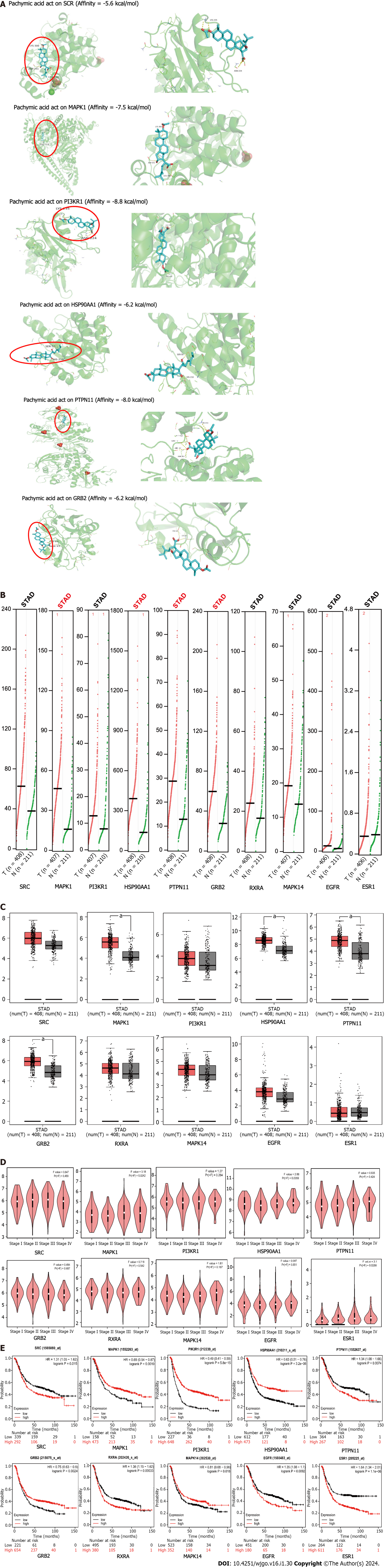
PA was utilized to conduct molecular docking of the top six core targets. When the binding energy is below -4.25 kcal/mol, there exists a certain capacity for the two to form a bond. The two exhibit a stronger affinity when the binding energy is below -5.0 kcal/mol. If the binding energy is lower than -7.0 kcal/mol, it is considered to be extremely high. The binding energy of PA with SRC, MAPK1, PI3KR1, and HSP90AA1 was -5.6 kcal/mol, -7.5 kcal/mol, -8.8 kcal/mol, and -6.2 kcal/mol, respectively. The binding energy was -8.0 kcal/mol for PTPN11 and -6.2 kcal/mol for GRB2 (Figure 4A).
The core genes MAPK14, HSP90AA1, PTPN11, and GRB2 transcription levels were significantly upregulated in the GEPIA database (Figure 4B). Comparison between GC and normal tissues showed that MAPK1, HSP90AA1, PTPN11, and GRB2 were up-regulated considerably (P < 0.05) (Figure 4C). MAPK1 (P = 0.0242), HSP90AA1 (P = 0.0359), and ESR1 (P = 0.0268) were significantly associated with tumor staging in patients (Figure 4D).
The survival of patients with high SRC, PTPN11, RXRA, EGFR, and ESR1 expression was worse than that of patients with low expression, whereas MAPK1, PIK3R1, HSP90AA1, GRB2, and MAPK14 showed the opposite trend. A P value of less than 0.05 was considered statistically significant. A P value of less than 0.001 indicates that the gene can be used as a marker for the prognosis of GC. According to the P value, PIK3R1 (P = 5.8e-15), HSP90AA1 (P = 3.2e-06), RXRA (P = 0.00033), and ESR1 (P = 1.1e-06) could be used as prognostic markers for GC. MAPK1 (P = 0.0019), PTPN11 (P = 0.0074), GRB2 (P = 0.0024), and EGFR (P = 0.0092) expression levels affected the prognosis of GC. SRC (P = 0.015) and MAPK14 (P = 0.018) were statistically significant for the prognosis of GC (Figure 4E).
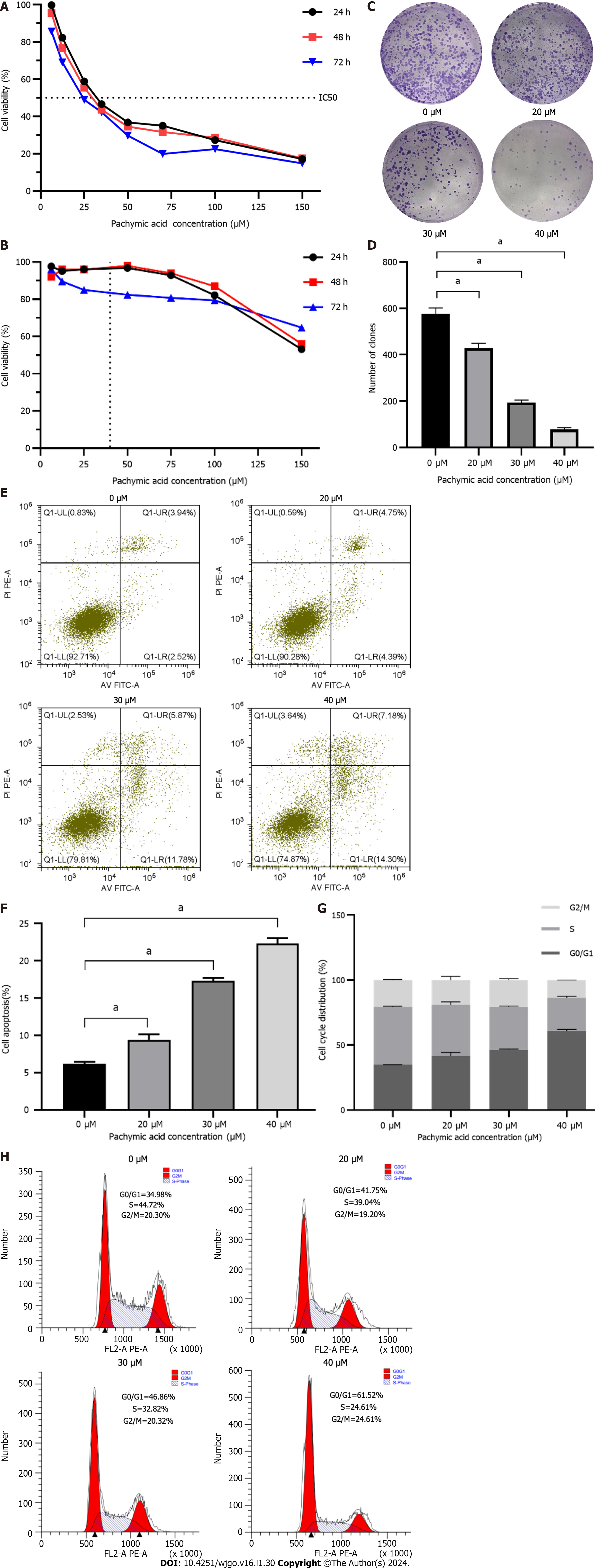
HGC-27 and GES-1 cells were treated with different concentrations of PA (6.25 μM, 12.5 μM, 25 μM, 35 μM, 50 μM, 70 μM, 100 μM, 150 μM) for 24 h, 48 h, and 72 h. These findings demonstrate that PA, in a time- and concentration-dependent manner, may dramatically diminish the viability of HGC-27 cells. The inhibitory effect eventually became stronger as the medication concentration and treatment duration increased. The proliferation of HGC-27 cells was significantly inhibited by drug intervention for 24 h. Therefore, 24 h was selected as the duration of drug intervention. Finally, according to the IC50 of 24 h = 37.32 μM, the high, medium, and low concentrations of the drugs (20 μM, 30 μM and 40 μM) were determined as the concentrations in subsequent experiments (Figure 5A). PA at a concentration of 37.32 μM had little effect on the viability of GES-1 (Figure 5B). When the GC cells were half dead, the damage of PA to normal gastric mucosal cells was very small, indicating that PA has the potential to be used in the treatment of GC. This result is similar to that the study by Wang et al[22] on the inhibition of invasion and metastasis of GC cells by PA.
The results showed that PA (20, 30, and 40 μM) considerably reduced the number of clones produced in HGC-27 cells compared to the control group, and that the inhibitory effect increased as the concentration increased. The difference was statistically significant (P < 0.001) (Figures 5C and D).
The findings from the apoptosis experiment indicated that the rates of cell death in HGC-27 cells exposed to varying concentrations of PA for 24 h were 9.39% ± 0.72%, 17.28% ± 0.42%, and 22.27% ± 0.74% for low, medium, and high concentrations, respectively. These rates were observed to be higher compared to the control group’s rate of 6.15% ± 0.27%. Significant statistical differences were observed in each group compared with the control group (P < 0.001) (Figures 5E and F).
Analysis of the cell cycle showed a significant alteration in the percentage of HGC-27 cells in the G0/G1 phase following 24 h exposure to varying concentrations of PA: Low 41.70% ± 2.65%, medium 46.40% ± 0.43%, and high 60.80% ± 1.29%. The values observed in the experimental group were more significant than those in the control group 34.83% ± 0.13%. Significant statistical differences were observed in each group compared with the control group (P < 0.001) (Figures 5G and H).
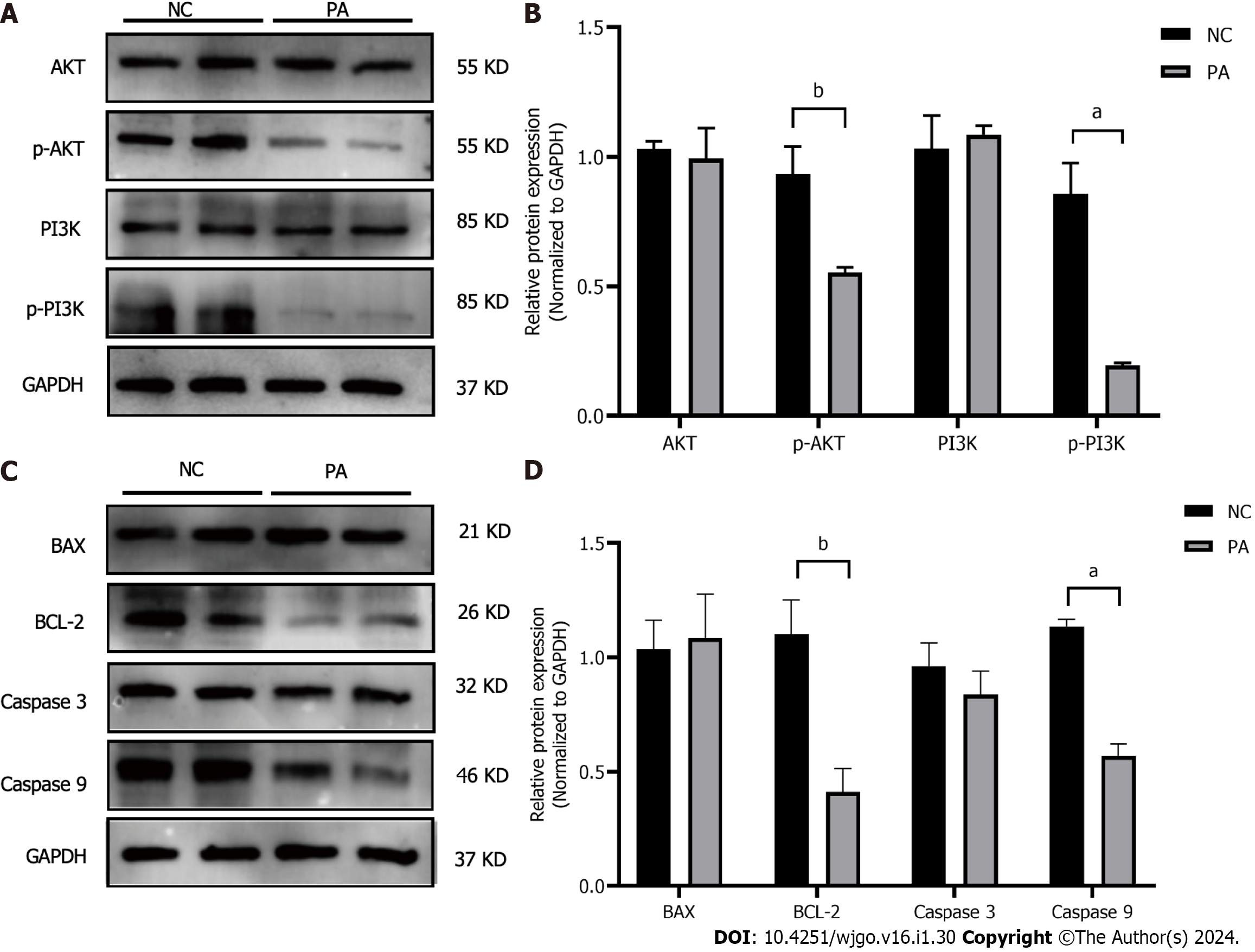
PI3K, AKT, and their phosphorylated forms, along with the apoptosis-related proteins BAX, BCL-2, CASPASE3, and CASPASE9, were detected using western blotting. The findings showed a notable decline in phosphorylated PI3K and AKT levels compared to the control group and it was statistically significant (P < 0.01, P < 0.001). In contrast, the overall protein levels of PI3K and AKT were not impacted (Figures 6A and B). There were no noticeable disparities in the protein manifestation of BAX and Caspase3, while the manifestation of BCL-2 and Caspase9 exhibited a substantial decline (P < 0.01, P < 0.001) (Figures 6C and D).
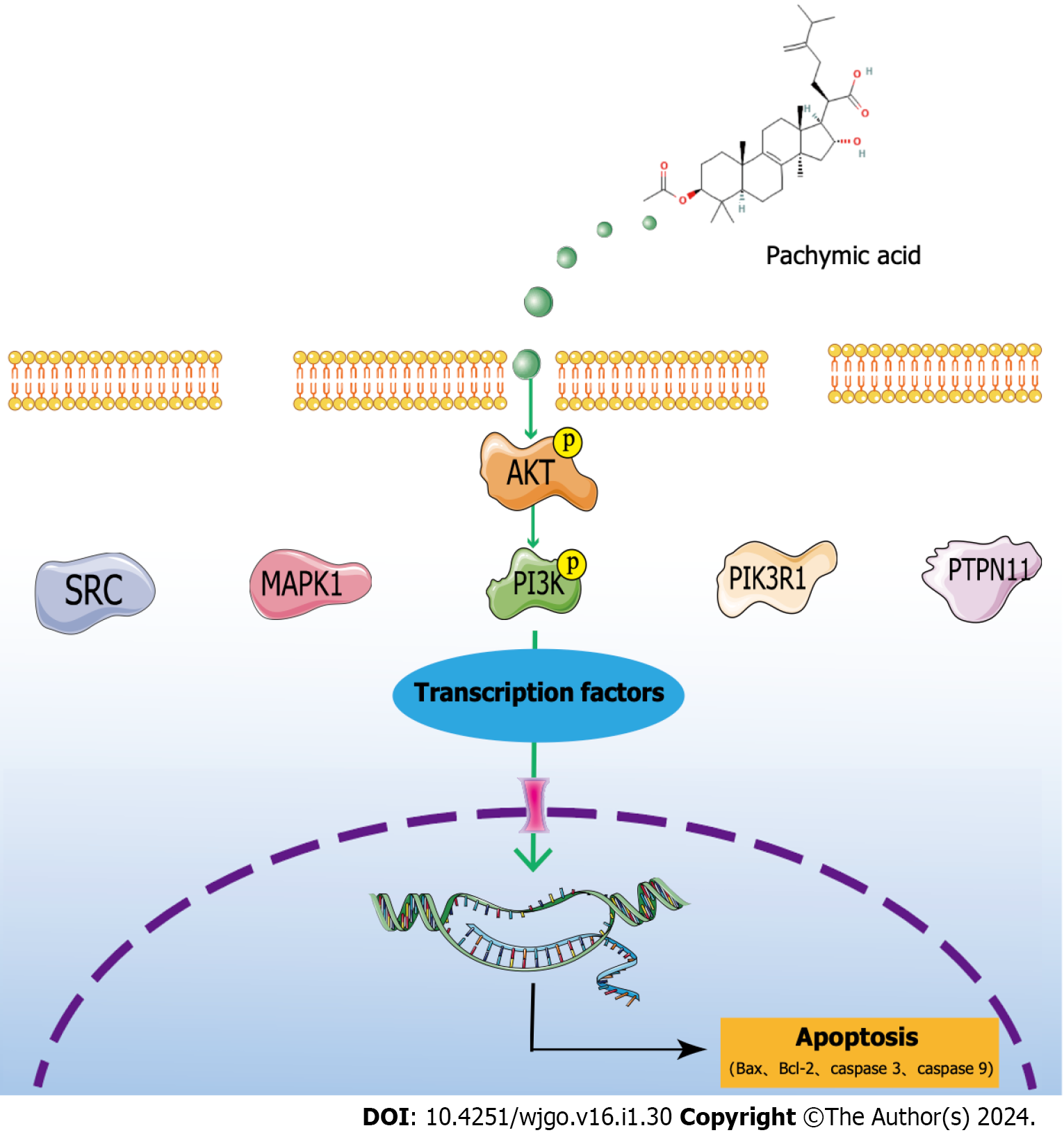
The analysis of network pharmacology revealed that SRC, MAPK1, PIK3R1, HSP90AA1, and PTPN11 were the critical targets of PA in the treatment of GC. The molecular docking results demonstrated PA’s strong binding ability with these key targets. Through a variety of database analysis, these core targets are closely related to the clinical relevance of GC. Through GO, KEGG, and GSEA analyses, the potential therapeutic mechanism for GC with PA was identified as the PI3K/AKT signaling pathway. The experiments showed that PA reduced the viability of HGC-27 cells, halted the cell cycle, enhanced cell apoptosis, regulated the PI3K/AKT signaling pathway to promote apoptosis, and restricted cell proliferation (Figure 7). At present, PA has not been used in clinical practice. Through this study, it can be clearly seen that in cell experiments, PA has a clear effect on GC, with the characteristics of high efficiency and low toxicity. Combined with network pharmacology analysis and clinical correlation analysis, it was confirmed that PA may have good clinical value and can provide a new treatment strategy for patients with GC.
GC cells SGC-7901, MGC-803, and BGC-823 may be contaminated by HELA cells. Ye et al[25] reported that STR typing results of SGC-7901 were consistent with those of HELA cells. Huang et al[26] performed STR typing on SGC-7901 from different laboratories, and the results showed that one strain was HELA cells and the other was non-human. In the same year, Bian et al[27] reported the cross-infection of 482 human tumor cell lines, of which 25 were contaminated with HELA cells, including SGC-7901. Therefore, Sun and Xia[28] reported that PA inhibits the growth of SGC-7901 cells and induces cell cycle arrest and apoptosis may need to be further verified using other GC cell lines. In this study, we used HGC-27 cells to increase the reliability of the inhibitory effect of PA on GC proliferation. In addition, compared with others, we have added the content of network pharmacological prediction and clinical correlation research, which provides the support of bioinformatics analysis for the treatment of GC with PA, and provides a more reliable basis for the application of PA in clinical treatment in the future.
The P13K/AKT signaling pathway is essential in multiple processes associated with tumor cell proliferation, apoptosis, invasion, metastasis, and tumor formation[29,30]. It is considered a key signaling pathway in tumor cells and has significant implications for tumor treatment and prognosis. In their study, Zhu et al[31] demonstrated that salvianol has the potential to inhibit the proliferation, migration, and invasion of hepatocellular carcinoma cells by modulating the PI3K/AKT signaling pathway. Similarly, Rong et al[32] noted that salidroside triggers apoptosis and protective autophagy in AGS cells via the PI3K/AKT/mTOR pathway. According to Zhou et al[33], epithelial-mesenchymal transition is induced by hypoxia in human GC cells via activation of the PI3K/AKT signaling pathway. The inhibition of tumor progression results by regulating the PI3K/AKT signaling pathway, which promotes tumor proliferation and metastasis by providing nutrition for tumor cells, reducing adhesion force between cells, influencing the extracellular matrix, and regulating growth factor receptors[34,35]. GSEA analysis of the GC microarray showed that the cell cycle, ECM-receptor interaction, and focal adhesion pathways were activated in GC, which were associated with the PI3K/AKT signaling pathway to promote GC development. Simultaneously, the PI3K/AKT signaling pathway exhibits a strong association with apoptosis[36]. Anomalous activation of the PI3K/AKT signaling pathway is observed in various tumor tissues, wherein it impedes tumor cell proliferation and fosters apoptosis through negative regulation[37]. By modulating the expression of the PI3K/AKT signaling pathway, PA has the potential to induce apoptosis in GC cells and suppress their proliferation.
In this study, only network pharmacological analysis and cell phenotype experiments were used to predict and verify the targets and possible molecular mechanisms of PA in GC treatment. Next, we conducted tumor formation experiments in nude mice to verify the efficacy of PA in vivo. Through in vivo and in vitro double verification, we provide reliable evidence for the treatment of GC with PA. In addition, high-throughput screening methods can be used for proteomics, genomics, and metabolomics analyses, and gene silencing was performed by lentivirus technology. In order to investigate the interplay between signal proteins during the treatment of GC using immunoprecipitation, and to elucidate the molecular mechanism of PA in GC treatment. The research offers fresh perspectives on the management of GC using PA.
At present, there is a lack of high-efficiency and low-toxicity drugs for the treatment of GC. However, there is still a long way to go for the application of PA in the clinical treatment of GC. This study only predicts the strong clinical relevance of PA in GC at the bioinformatics level, and has not actually been developed into a relevant drug for clinical trials. The current study suggests that PA has the potential for clinical application in GC, although the process is difficult. In the future, we can confirm this through more studies and make the treatment of GC further.
The inhibition of GC cell proliferation by PA and its potential to induce apoptosis in HGC-27 cells were confirmed through network pharmacological analysis and experimental verification. It was found that PA may target SRC, MAPK1, PIK3R1, HSP90AA1, PTPN11 proteins, and PI3K/AKT signaling pathways.
Gastric cancer (GC) is one of the most serious malignant tumors of the digestive tract, with high morbidity and mortality. In recent years, more and more evidence has shown that natural ingredients from traditional Chinese medicine can prevent and inhibit the occurrence and development of GC, showing great therapeutic potential.
Network pharmacology is used to predict the targets and pathways of drugs in the treatment of diseases, and the method of experimental verification is used to verify them, which provides an effective reference for further elucidating the molecular mechanism of natural products in the treatment of GC.
To find the possible targets and pathways of pachymic acid (PA) in the treatment of GC, and to explore the molecular mechanism of PA promoting apoptosis and arresting cell cycle of GC cells by regulating PI3K/AKT signal transduction.
Using the TCMSP database to find the components of Poria cocos, we identified PA in Poria cocos as a potential effective component in the treatment of GC. The PharmMapper database was used to find the related targets of PA, and the GENECARDS and OMIM databases were used to find the related targets of GC, and the common targets were found by intersection. The protein-protein interaction network was constructed, and the top five core targets were screened according to the degree value. The Gene Ontology, Kyoto Encyclopedia of Genes and Genomes (KEGG), and gene set enrichment analysis were used to identify related pathways. Molecular docking and clinical correlation studies were used to explore the effects of core targets on GC. Cell Counting Kit-8, colony formation assay and flow cytometry were used to detect cell function indexes. Western blot was used to determine the effect of PA on PI3K/AKT signaling pathway and apoptosis pathway.
A total of 217 possible targets of PA in the treatment of GC were identified by network pharmacology, and the top five core targets were found: Proto-oncogene tyrosine-protein kinase Src, mitogen-activated protein kinase 1, phosphatidylinositol 3-kinase regulatory subunit alpha, heat shock protein 90-alpha, tyrosine-protein phosphatase non-receptor type 11. KEGG pathway analysis showed that PI3K/AKT signaling pathway was a possible pathway for the treatment of GC. Molecular docking results showed that the binding energy between PA and PI3KR1 was the highest (-8.8 kcal/mol). Experiments on HGC-27 cells confirmed that PA could inhibit the proliferation of GC cells, promote the apoptosis of cells and arrest their G0/G1 phase. Western blot results showed that PA decreased the expression of phosphorylated PI3K and AKT in GC cells, and also had an effect on apoptosis-related proteins.
The inhibitory effect of PA on GC proliferation and its potential to induce apoptosis of GC cells were confirmed by network pharmacological analysis and experimental verification.
PA have shown high efficiency and low toxicity in the treatment of GC. However, PA have not been truly developed into relevant drugs for clinical trials, and there is still a long way to go for the application of PA in the clinical treatment of GC. In the future, we look forward to making the treatment of GC more in-depth through more studies on the effectiveness of PA on GC and providing more treatment options for patients with GC.
The authors would like to acknowledge Li-Qun Wang for statistical analysis assistance. Thanks to Joanna Tibenda for revising the article.
Provenance and peer review: Unsolicited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Oncology
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Bugaj AM, Poland; Gupta L, Indonesia S-Editor: Wang JJ L-Editor: A P-Editor: Zhang XD
| 1. | Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75126] [Cited by in RCA: 64649] [Article Influence: 16162.3] [Reference Citation Analysis (176)] |
| 2. | Smyth EC, Nilsson M, Grabsch HI, van Grieken NC, Lordick F. Gastric cancer. Lancet. 2020;396:635-648. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1150] [Cited by in RCA: 2858] [Article Influence: 571.6] [Reference Citation Analysis (5)] |
| 3. | Xu W, Yang Z, Lu N. Molecular targeted therapy for the treatment of gastric cancer. J Exp Clin Cancer Res. 2016;35:1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 93] [Cited by in RCA: 148] [Article Influence: 16.4] [Reference Citation Analysis (0)] |
| 4. | Rausei S, Lianos GD. Treatment of Gastric Cancer. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 5] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 5. | Moehler M, Baltin CT, Ebert M, Fischbach W, Gockel I, Grenacher L, Hölscher AH, Lordick F, Malfertheiner P, Messmann H, Meyer HJ, Palmqvist A, Röcken C, Schuhmacher C, Stahl M, Stuschke M, Vieth M, Wittekind C, Wagner D, Mönig SP. International comparison of the German evidence-based S3-guidelines on the diagnosis and multimodal treatment of early and locally advanced gastric cancer, including adenocarcinoma of the lower esophagus. Gastric Cancer. 2015;18:550-563. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 57] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 6. | Kim WH, Gomez-Izquierdo L, Vilardell F, Chu KM, Soucy G, Dos Santos LV, Monges G, Viale G, Brito MJ, Osborne S, Noé J, Du X. HER2 Status in Gastric and Gastroesophageal Junction Cancer: Results of the Large, Multinational HER-EAGLE Study. Appl Immunohistochem Mol Morphol. 2018;26:239-245. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 32] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 7. | Zubair H, Khan MA, Anand S, Srivastava SK, Singh S, Singh AP. Modulation of the tumor microenvironment by natural agents: implications for cancer prevention and therapy. Semin Cancer Biol. 2022;80:237-255. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 33] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 8. | Wang J, Qi F, Wang Z, Zhang Z, Pan N, Huai L, Qu S, Zhao L. A review of traditional Chinese medicine for treatment of glioblastoma. Biosci Trends. 2020;13:476-487. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 56] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 9. | Sun Q, He M, Zhang M, Zeng S, Chen L, Zhao H, Yang H, Liu M, Ren S, Xu H. Traditional Chinese Medicine and Colorectal Cancer: Implications for Drug Discovery. Front Pharmacol. 2021;12:685002. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 34] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 10. | Ríos JL. Chemical constituents and pharmacological properties of Poria cocos. Planta Med. 2011;77:681-691. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 250] [Article Influence: 17.9] [Reference Citation Analysis (0)] |
| 11. | Nie A, Chao Y, Zhang X, Jia W, Zhou Z, Zhu C. Phytochemistry and Pharmacological Activities of Wolfiporia cocos (F.A. Wolf) Ryvarden & Gilb. Front Pharmacol. 2020;11:505249. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 66] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 12. | Feng YL, Zhao YY, Ding F, Xi ZH, Tian T, Zhou F, Du X, Chen DQ, Wei F, Cheng XL, Lin RC. [Chemical constituents of surface layer of Poria cocos and their pharmacological properties (I)]. Zhongguo Zhong Yao Za Zhi. 2013;38:1098-1102. [PubMed] |
| 13. | Huang YC, Chang WL, Huang SF, Lin CY, Lin HC, Chang TC. Pachymic acid stimulates glucose uptake through enhanced GLUT4 expression and translocation. Eur J Pharmacol. 2010;648:39-49. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 64] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 14. | Feng Z, Shi H, Liang B, Ge T, Cai M, Liu F, Huang K, Wen J, Chen Q, Ge B. Bioinformatics and experimental findings reveal the therapeutic actions and targets of pachymic acid against cystitis glandularis. Biofactors. 2021;47:665-673. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 6] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 15. | Shah VK, Choi JJ, Han JY, Lee MK, Hong JT, Oh KW. Pachymic Acid Enhances Pentobarbital-Induced Sleeping Behaviors via GABAA-ergic Systems in Mice. Biomol Ther (Seoul). 2014;22:314-320. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 46] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 16. | Ma J, Liu J, Lu C, Cai D. Pachymic acid induces apoptosis via activating ROS-dependent JNK and ER stress pathways in lung cancer cells. Cancer Cell Int. 2015;15:78. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 51] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 17. | Cheng S, Eliaz I, Lin J, Thyagarajan-Sahu A, Sliva D. Triterpenes from Poria cocos suppress growth and invasiveness of pancreatic cancer cells through the downregulation of MMP-7. Int J Oncol. 2013;42:1869-1874. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 48] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 18. | Hong R, Shen MH, Xie XH, Ruan SM. Inhibition of breast cancer metastasis via PITPNM3 by pachymic acid. Asian Pac J Cancer Prev. 2012;13:1877-1880. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 24] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 19. | Gao AH, Zhang L, Chen X, Chen Y, Xu ZZ, Liu YN, Zhang H. Inhibition of ovarian cancer proliferation and invasion by pachymic acid. Int J Clin Exp Pathol. 2015;8:2235-2241. [PubMed] |
| 20. | Lu C, Ma J, Cai D. Pachymic acid inhibits the tumorigenicity of gastric cancer cells by the mitochondrial pathway. Anticancer Drugs. 2017;28:170-179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 17] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 21. | Yuan L, Yang Y, Li X, Zhou X, Du YH, Liu WJ, Zhang L, Yu L, Ma TT, Li JX, Chen Y, Nan Y. 18β-glycyrrhetinic acid regulates mitochondrial ribosomal protein L35-associated apoptosis signaling pathways to inhibit proliferation of gastric carcinoma cells. World J Gastroenterol. 2022;28:2437-2456. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 5] [Cited by in RCA: 11] [Article Influence: 3.7] [Reference Citation Analysis (1)] |
| 22. | Wang H, Luo Y, Chu Z, Ni T, Ou S, Dai X, Zhang X, Liu Y. Poria Acid, Triterpenoids Extracted from Poria cocos, Inhibits the Invasion and Metastasis of Gastric Cancer Cells. Molecules. 2022;27. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 24] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 23. | Lu C, Cai D, Ma J. Pachymic Acid Sensitizes Gastric Cancer Cells to Radiation Therapy by Upregulating Bax through Hypoxia. Am J Chin Med. 2018;46:875-890. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 17] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 24. | Pillai-Kastoori L, Schutz-Geschwender AR, Harford JA. A systematic approach to quantitative Western blot analysis. Anal Biochem. 2020;593:113608. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 236] [Article Influence: 47.2] [Reference Citation Analysis (0)] |
| 25. | Ye F, Chen C, Qin J, Liu J, Zheng C. Genetic profiling reveals an alarming rate of cross-contamination among human cell lines used in China. FASEB J. 2015;29:4268-4272. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 94] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 26. | Huang Y, Liu Y, Zheng C, Shen C. Investigation of Cross-Contamination and Misidentification of 278 Widely Used Tumor Cell Lines. PLoS One. 2017;12:e0170384. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 64] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 27. | Bian X, Yang Z, Feng H, Sun H, Liu Y. A Combination of Species Identification and STR Profiling Identifies Cross-contaminated Cells from 482 Human Tumor Cell Lines. Sci Rep. 2017;7:9774. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 53] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 28. | Sun KX, Xia HW. Pachymic acid inhibits growth and induces cell cycle arrest and apoptosis in gastric cancer SGC-7901 cells. Oncol Lett. 2018;16:2517-2524. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 29. | Fattahi S, Amjadi-Moheb F, Tabaripour R, Ashrafi GH, Akhavan-Niaki H. PI3K/AKT/mTOR signaling in gastric cancer: Epigenetics and beyond. Life Sci. 2020;262:118513. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 269] [Cited by in RCA: 248] [Article Influence: 49.6] [Reference Citation Analysis (0)] |
| 30. | Noorolyai S, Shajari N, Baghbani E, Sadreddini S, Baradaran B. The relation between PI3K/AKT signalling pathway and cancer. Gene. 2019;698:120-128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 143] [Cited by in RCA: 412] [Article Influence: 68.7] [Reference Citation Analysis (0)] |
| 31. | Zhu P, Liu Z, Zhou J, Chen Y. Tanshinol inhibits the growth, migration and invasion of hepatocellular carcinoma cells via regulating the PI3K/AKT signaling pathway. Onco Targets Ther. 2019;12:87-99. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 21] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 32. | Rong L, Li Z, Leng X, Li H, Ma Y, Chen Y, Song F. Salidroside induces apoptosis and protective autophagy in human gastric cancer AGS cells through the PI3K/Akt/mTOR pathway. Biomed Pharmacother. 2020;122:109726. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 151] [Article Influence: 25.2] [Reference Citation Analysis (0)] |
| 33. | Zhou X, Ma W, Li X, Xu J. Glaucocalyxin a prevents hypoxia-induced epithelial-mesenchymal transition in human gastric cancer cells through the PI3K/Akt signaling pathway. J Recept Signal Transduct Res. 2022;42:109-116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 34. | Baghery Saghchy Khorasani A, Pourbagheri-Sigaroodi A, Pirsalehi A, Safaroghli-Azar A, Zali MR, Bashash D. The PI3K/Akt/mTOR signaling pathway in gastric cancer; from oncogenic variations to the possibilities for pharmacologic interventions. Eur J Pharmacol. 2021;898:173983. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 51] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 35. | Hoxhaj G, Manning BD. The PI3K/AKT network at the interface of oncogenic signalling and cancer metabolism. Nat Rev Cancer. 2020;20:74-88. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1028] [Cited by in RCA: 1345] [Article Influence: 269.0] [Reference Citation Analysis (0)] |
| 36. | Fresno Vara JA, Casado E, de Castro J, Cejas P, Belda-Iniesta C, González-Barón M. PI3K/Akt signalling pathway and cancer. Cancer Treat Rev. 2004;30:193-204. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1407] [Cited by in RCA: 1787] [Article Influence: 85.1] [Reference Citation Analysis (0)] |
| 37. | Bishnupuri KS, Alvarado DM, Khouri AN, Shabsovich M, Chen B, Dieckgraefe BK, Ciorba MA. IDO1 and Kynurenine Pathway Metabolites Activate PI3K-Akt Signaling in the Neoplastic Colon Epithelium to Promote Cancer Cell Proliferation and Inhibit Apoptosis. Cancer Res. 2019;79:1138-1150. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 150] [Article Influence: 25.0] [Reference Citation Analysis (0)] |