Published online Jan 7, 2015. doi: 10.3748/wjg.v21.i1.311
Peer-review started: April 12, 2014
First decision: May 13, 2014
Revised: June 3, 2014
Accepted: July 11, 2014
Article in press: July 11, 2014
Published online: January 7, 2015
Processing time: 269 Days and 17.6 Hours
AIM: To determine the mutation status of human telomerase reverse transcriptase gene (TERT) promoter region in hepatocellular carcinoma (HCC) from different geographical regions.
METHODS: We analyzed the genomic DNA sequences of 59 HCC samples comprising 15 cell lines and 44 primary tumors, collected from patients living in Asia, Europe and Africa. We amplified a 474 bp DNA fragment of the promoter region of TERT gene including the 1295228 and 1295250 sequence of chromosome 5 by using PCR. Amplicons were then sequenced by Sanger technique and the sequence data were analyzed with by using DNADynamo software in comparison with wild type TERT gene sequence as a reference.
RESULTS: The TERT mutations were found highly frequent in HCC. Eight of the fifteen tested cell lines displayed C228T mutation, and one had C250T mutation with a mutation frequency up to 60%. All of the mutations were heterozygous and mutually exclusive. Ten out of forty-four tumors displayed C228T mutation, and additional five tumors had C250T mutation providing evidence for mutation frequency of 34% in primary tumors. Considering the geographic origins of HCC tumors tested, TERT promoter mutation frequencies were higher in African (53%), when compared to non-African (24%) tumors (P = 0.056). There was also a weak inverse correlation between TERT promoter mutations and murine double minute 2 single nucleotide polymorphism 309 TG polymorphism (P = 0.058). Mutation frequency was nearly two times higher in established HCC cell lines (60%) compared to the primary tumors (34%).
CONCLUSION: TERT promoter is one of most frequent mutational targets in liver cancer, and hepatocellular carcinogenesis is highly associated with the loss of telomere-dependent cellular senescence control.
Core tip: Our study demonstrated that telomerase reverse transcriptase (TERT) promoter mutations are present in hepatocellular carcinomas (HCCs) from different geographical regions, and the highest frequency was observed in tumors from Africa. These mutations occur both primarily as C228T mutation and as C250T mutation. These results also provide evidence for TERT mutations as a common trait of HCC regardless of their geographical location.
- Citation: Cevik D, Yildiz G, Ozturk M. Common telomerase reverse transcriptase promoter mutations in hepatocellular carcinomas from different geographical locations. World J Gastroenterol 2015; 21(1): 311-317
- URL: https://www.wjgnet.com/1007-9327/full/v21/i1/311.htm
- DOI: https://dx.doi.org/10.3748/wjg.v21.i1.311
Hepatocellular carcinoma (HCC) is one of the most common and fatal cancers with a heterogeneous incidence throughout different regions of the world[1]. HCC, whose incidence has been vigorously increasing in western countries, has the highest incidence in China, Middle Africa, and Japan. Epidemiology of HCC differs among different geographical regions. Hepatitis B and C are the main risk factors in Asia and Africa while alcohol intake is the main driving force in Europe and the United States[2]. Overall survival rate of HCC patients is very low due to inefficient treatment options. HCC is resistant to most of the conventional therapies, thus the only plausible treatment is liver transplantation that is restricted to early-diagnosed cases[3]. In order to provide a more effective therapeutic approach to HCC patients, genetic mechanisms underlying liver carcinogenesis have been studied for years; however, most of the mutations identified so far are “loss-of-function” type, thus they are not suitable to be used for targeted therapy[4]. The only molecularly targeted drug for HCC treatment is Sorafenib whose efficacy is not satisfactory[5].
Tumor cells need to overcome the telomere shortening problem, one of the most crucial obstacles during the transformation process. This can be achieved either by up-regulating telomerase activity or with alternative lengthening of telomeres[6]. Integration of hepatitis B viral DNA into the telomerase reverse transcriptase (TERT) gene is observed in HCC patients with hepatitis B viral (HBV) infection and considered found as one of the paths to increase telomere length[7-9]. However, there are many HCC cases without HBV involvement and in which telomere length is still an issue for those. Recently, many groups reported the presence of two frequent mutations in TERT promoter region in different tumors including HCC[10-16]. These promoter mutations are claimed to upregulate the TERT transcription by creating a binding site for ETS (E-twenty six)[10] and ternary complex factor (TCF) transcription factors[11]. Reported HCC tumors with TERT promoter mutations were from United States[12] and France[15] and mutation frequencies were 44% and 59%, respectively. Highly frequent TERT mutations may serve not only as novel diagnostic markers but also as and potential therapeutic targets for HCC. However, it is still unknown whether TERT promoter mutations occur in diverse HCCs worldwide, regardless of their geographical origin. As these tumors occur less frequently in western populations, but quite commonly then in Asian and African, TERT promoter status in Asian and African HCC patients is worth to know. In this study, we analyzed 15 HCC cell lines, as well as 44 HCC tumors from three different continents in search for two hotspot mutations in TERT promoter.
We used archival HCC tumor DNA samples (n = 44) that have been described previously in terms of hepatitis B viral DNA testing, TP53 mutations and murine double minute 2 (MDM2) polymorphism[17,18].
Huh7, HepG2, Hep3B, Hep40, PLC/PRF/5, FOCUS, Mahlavu, FLC4, and SK-HEP-1 cells were cultured in Dulbecco’s modified Eagles medium, whereas SNU182, SNU387, SNU398, SNU423, SNU449, and SNU475 cell lines are grown in RPMI. Both media were supplemented with 10% fetal calf serum, 2 mmol/L L-glutamine, 1× non essential amino acids, and 100 units of penicillin/streptomycin (all from Life Technologies™). Cells were grown up to 70% confluency before genomic DNA extraction.
Genomic DNA samples were isolated by using Purelink Genomic DNA Kit (Life Technologies™) according to manufacturer’s instructions, then DNA concentrations were measured with Nanodrop Spectrometer (Thermo Scientific). 100 ng of genomic DNA was used to amplify a 474 bp region of TERT promoter flanking hotspot mutations that are found at positions 1295228 and 1295250 of chromosome 5 by using AccuPrime GC-rich DNA polymerase kit (Life Technologies™) with forward primer 5’-ACGAACGTGGCCAGCGGCAG-3’ and reverse primer 5’- CTGGCGTCCCTGCACCCTGG-3’[11]. Amplicons were sequenced with Sanger technique, and data were analyzed with DNADynamo software (BlueTractor Software Ltd) by comparing TERT sequence from UCSC Genome Browser as a reference.
Fisher exact test was used to compare statistical differences (P-values; one-tailed) among clinical samples holding and lacking TERT promoter mutation using Wassar Statistics Tool available online (http://vassarstats.net). A P-value of less than 0.05 was considered to be significant.
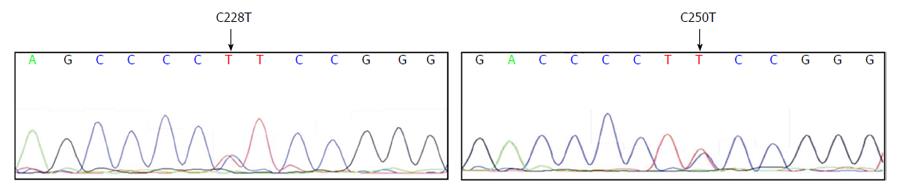
We tested a panel of 15 HCC cell lines composed of six epithelial-like (Huh7, HepG2, Hep3B, Hep40, PLC/PRF/5, and FLC4) and nine mesenchymal-like (FOCUS, Mahlavu, SNU182, SNU387, SNU398, SNU423, SNU449, and SNU475, SKHEP1) cell types[19] for mutations at TERT gene promoter. Nine cell lines carried C228T mutation but only one cell line, Mahlavu, carried C250T mutation; all mutations were heterozygous (Table 1). Two examples of sequence chromatograms representing C228T and C250T mutations are given in Figure 1. In sum, 67% (10 out of 15) of HCC cell lines displayed a TERT promoter mutation. In all HCC cell lines tested, C228T and C250T mutations were found in a mutually exclusive manner. Both epithelial-like and mesenchymal-like cells had these mutations with similar frequencies (4 out of 6, and 6 out of 9 respectively). We concluded that TERT promoter mutations occur frequently in HCC cell lines, regardless of their differentiation status.
| Cell lines | TERT promoter status |
| Epitheliel-like | |
| Huh7 | C228T |
| HepG2 | C228T |
| Hep3B | C228T |
| FLC4 | C228T |
| Hep40 | Wild-type |
| PLC/PRF/5 | Wild-type |
| Mesenchymal-like | |
| FOCUS | C228T |
| SNU387 | C228T |
| SNU398 | C228T |
| SNU423 | C228T |
| SNU475 | C228T |
| Mahlavu | C250T |
| SNU182 | Wild-type |
| SNU449 | Wild-type |
| SKHEP1 | Wild-type |
To determine TERT promoter mutation frequency in HCC tumors, we tested an archival collection of 44 HCC tumor DNAs (Table 2) collected from different countries around the world including Japan (11 patients), China (8), Germany (7), France (2), Israel (1), Mozambique (6), Transkei (4), Lesotho (2), Swaziland (1), and South Africa (2). Based on tumor viral DNA testin[16,17] the etiology for 23 out of 44 (52.3%) of these tumors was hepatitis B virus infection. The etiology of other tumors was unknown. We identified 15 mutations in 44 tumors, 10 of which were C228T and the other 5 were C250T mutations. C228T mutations (23%) were again more frequent than C250T mutations (11%) and they were mutually exclusive, as observed in HCC cell lines (Table 2).
| Country | TERT mutations | p53 mutations | MDM2 | HBV | Stage | ||
| C228T | C250T | Codon | Amino Acid change | SNP 309 | |||
| Japan | C228T | WT | 6 bp del | Del (AGCTAC) | G/G | Minus1 | Unknown |
| Japan | C228T | WT | WT | T/G | Minus1 | Unknown | |
| Japan | C228T | WT | WT | T/G | Minus1 | Unknown | |
| Japan | WT | C250T | WT | G/G | Minus1 | Unknown | |
| Japan | WT | WT | WT | T/G | Minus1 | Unknown | |
| Japan | WT | WT | WT | T/G | Minus1 | Unknown | |
| Japan | WT | WT | WT | T/G | Minus1 | Unknown | |
| Japan | WT | WT | WT | T/G | Minus1 | Unknown | |
| Japan | WT | WT | WT | T/G | plus1 | Unknown | |
| Japan | WT | WT | WT | G/G | Minus1 | Unknown | |
| Japan | WT | WT | WT | T/G | Minus1 | Unknown | |
| China | WT | WT | 281 | C > A Asp > Glu | T/T | plus1 | Unknown |
| China | WT | WT | WT | G/G | Plus1 | Unknown | |
| China | WT | WT | WT | T/G | Plus1 | Unknown | |
| China | WT | WT | WT | T/T | Plus1 | Unknown | |
| China | WT | WT | WT | G/G | Plus1 | Unknown | |
| China | WT | WT | WT | T/T | Plus1 | Unknown | |
| China | WT | WT | WT | G/G | Plus2 | Unknown | |
| China | WT | WT | WT | G/G | plus1 | Unknown | |
| Israel | WT | WT | WT | T/G | Minus1 | Unknown | |
| Mozambique | C228T | WT | 157 | G > T Val > Phe | T/T | Plus1 | Late |
| Mozambique | C228T | WT | WT | T/T | Plus2 | Late | |
| Mozambique | WT | C250T | WT | T/T | Plus2 | Late | |
| Mozambique | WT | C250T | 249 | G > T Arg > Ser | T/T | Minus1 | Early |
| Mozambique | WT | WT | WT | T/T | Plus2 | Late | |
| Mozambique | WT | WT | 249 | G > T Arg > Ser | T/T | Plus1 | Late |
| Transkei | C228T | WT | WT | T/T | NT | Late | |
| Transkei | C228T | WT | WT | T/T | NT | Late | |
| Transkei | WT | WT | WT | T/T | NT | Early | |
| Transkei | WT | WT | WT | T/T | Plus1 | Late | |
| Lesotho | WT | C250T | WT | T/T | Plus2 | Early | |
| Lesotho | WT | WT | WT | T/G | Plus2 | late | |
| Swaziland | WT | WT | WT | T/T | Plus | Early | |
| South Africa | C228T | WT | WT | T/T | Plus2 | Late | |
| South Africa | WT | WT | WT | T/G | Plus2 | Late | |
| Germany | C228T | WT | WT | G/G | Minus1 | Metastasis | |
| Germany | C228T | WT | 273 | C > T Arg > Cys | T/T | Minus1 | HCC |
| Germany | WT | C250T | WT | G/G | Plus2 | HCC | |
| Germany | WT | WT | WT | G/G | Minus1 | Unknown | |
| Germany | WT | WT | WT | G/G | Minus1 | Metastasis | |
| Germany | WT | WT | WT | T/T | Plus2 | Unknown | |
| Germany | WT | WT | WT | T/G | Plus2 | HCC | |
| France | WT | WT | WT | T/G | Minus1 | Unknown | |
| France | WT | WT | WT | T/G | Minus1 | Unknown | |
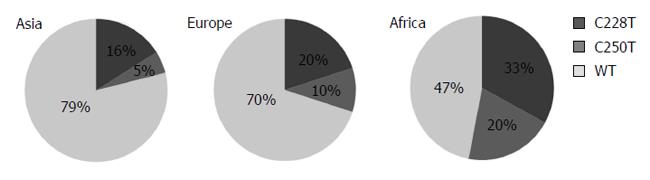
Figure 2 displays the distribution of TERT mutations in different continents. Tumors from Africa with the highest mutation frequency (53%) were followed by tumors from Europe (30%) and Asia (21%), respectively. We were not able to test whether these mutations were germ-line or somatically acquired, however, all reported C228T and C250T mutations in HCC were acquired somatic mutations[12,15]. Thus; we assume that mutations reported here are also somatic.
Table 3 compares patient characteristics such as gender, age, geographical status, tumor HBV DNA and TP53 mutation as well as patient MDM2 single nucleotide polymorphism (SNP) 309 status with the mutational status of TERT promoter. There was no significant difference found in patient gender and age, but a weak association (P = 0.056) was found in geographical origin. Tumors from African patients displayed TERT promoter mutations two-fold more frequently (53%) than non-African patients (24%). Tumors with HBV DNA displayed less frequent TERT promoter mutations (26%) as compared to HBV-negative tumors (39%), but the difference did not reach to a significance (P = 0.295). Similarly, tumors with wild-type TP53 displayed less frequent TERT promoter mutations (29%) as compared to those with a mutation (50%). However, this difference did not reach to a significant level (P = 0.280). In contrast, we found a week association between TERT promoter mutations and MDM2 SNP 309 TG polymorphism (P = 0.058). Patients with SNP309 TT polymorphism displayed 44% TERT promoter mutation, in contrast to those with TG polymorphism which displayed only 13% TERT promoter mutations. Indeed, TERT promoter mutations were over 3-fold more frequent in patients with MDM2 SNP 309 TT status, than those with a TG status.
| Variable | Overall series | TERT promoter mutated | TERT promoter non-mutated | P value |
| (n = 44) | (n = 15) | (n = 29) | ||
| Gender | ||||
| Male | 27 | 10 | 17 | 0.2059 |
| Female | 1 | 1 | 0 | |
| Age | ||||
| ≥ 60 yr | 9 | 3 (33) | 6 (67) | 0.6547 |
| < 60 yr | 19 | 8 (42) | 11 (58) | |
| Geographical origin | ||||
| African | 15 | 8 (53) | 7 (47) | 0.0528 |
| Non-African | 29 | 7 (24) | 22 (76) | |
| HBV DNA | ||||
| Positive | 23 | 6 (26) | 17 (74) | 0.2950 |
| Negative | 18 | 7 (39) | 11 (61) | |
| TP53 | ||||
| Mutated | 6 | 3 (50) | 3 (50) | 0.6315 |
| Wild-type | 38 | 11 (29) | 27 (71) | |
| MDM2 SNP 309 | ||||
| TT | 18 | 8 (44) | 10 (56) | |
| TG | 15 | 2 (13) | 13 (87) | 0.0528 (vs TT) |
| GG | 11 | 4 (36) | 7 (64) |
The TERT gene, encoding the catalytic subunit of telomerase reverse transcriptase enzyme, is a limiting factor for unlimited proliferation of most human somatic cells including hepatocytes. Lack of TERT gene expression in these cells leads to a progressive erosion of telomeres during successive cell divisions culminating with a permanent cell cycle arrest when telomere DNA reaches a critically short stature. Cancer cells such as HCC cells overcome this arrest by reactivating TERT gene expression with ill-known mechanisms. TERT reactivation is so far the most frequently observed (80%-90%) aberration in HCC tumors[20,21]. Several mechanisms have been reported for the activation of TERT expression in cancer cells, including myc and Wnt/β-catenin signaling-mediated activation[22-24], alternative splicing, and epigenetic alterations[25,26]. Whether these mechanisms are involved in hepatocellular carcinogenesis is still unknown.
TERT reactivation is associated with HBV DNA integration near the TERT gene in rare cases of HCC, providing a clue about viral reactivation of TERT expression[7]. In addition, TERT promoter mutations have been reported recently as frequent events in some cancers such as melanoma, sarcomas, urothelial carcinoma, bladder cancer, glioblastoma, thyroid cancer, and HCC[10-16]. Although it is not clear yet whether such mutations are necessary and sufficient for TERT reactivation in cancer cells, it appears that somatic mutations of TERT promoter are among the most frequent aberrations observed in some tumor types. Our studies in HCC cell lines reiterate this striking finding. With 60% frequency, TERT mutation is the most frequent mutational event observed in these cell lines together with TP53 mutations so far[27]. Thus, it is very likely that TERT promoter mutations facilitate the establishment of HCC cell lines by overcoming telomere shortening during in vitro culture. We have found similar mutation frequencies for both epithelial-like and mesenchymal-like cell lines suggesting that mutagenesis of the TERT promoter is independent of the differentiation status of the cell lines. Early HCCs display epithelial like morphology whereas advanced HCCs may display mesenchymal-like morphology associated with epithelial to mesenchymal transition that is often observed during tumor progression[28,29]. Our findings suggest that TERT mutations are early events during hepatocellular carcinogenesis in confirmation with a recent report[15]. The mutations observed in cell lines are the same type of mutations observed in primary tumors. This suggests that cell line mutations did not occur spontaneously during cell culture. Their high frequency may indicate that tumor cells with such mutations are established more easily.
TERT promoter mutations that are observed in 34% of primary HCC tumors are quite high, albeit less frequent than those observed in cell lines. This lower frequency in tumors may be expected because of the potential bias due to a selective advantage during cell culture as stated above. Additionally, heterozygous TERT promoter mutations may be more difficult to detect due to the contamination of tumor DNA with the DNA coming from non-cancer cells into tumor tissues. Despite these limitations, the existence of TERT promoter mutations in at least one-third of primary tumors indicates that this gene is one of the most frequent targets for mutation in liver cancer. Our recently published findings pinpointed TERT as a critical gene involved in HCC cell immortality, which itself is viewed as a central mechanism of hepatocellular carcinogenesis in humans[30,31]. This present study, together with a recent study[15] clearly establishes that TERT promoter mutation is a hallmark of liver cancer. Our findings provide further evidence for a global incidence of TERT promoter mutations in liver cancer regardless of their geographical origin. Moreover, we provide preliminary evidence for a higher frequency of these mutations in patients from Africa. Thus TERT mutations restricted to two hotspots at its promoter, are universal markers for liver cancer and thus they may serve as easy cancer biomarkers in high risk populations such as those chronically infected with hepatitis viruses, as well as cirrhosis. Finally, higher manifestation of TERT promoter mutations in HCC patients with MDM2 SNP309 TG status strongly suggests that there is a cross talk between TP53-MDM2 axis and TERT functions in liver cancer. Further research is needed to confirm these initial observations.
In conclusion, TERT promoter mutations that are widely observed in liver cancers from around the world provide sufficient evidence for the critical role of telomere biology and cellular immortality in these cancers.
Hepatocellular carcinoma (HCC) is one of the most fatal cancers over the world with an increasing incidence in western countries, so it is of great importance to reveal genetic mechanisms that may play an important role in liver tumor formation. Telomeres are repetitive DNA sequences found at both ends of each chromosome. In normal somatic cells, they get shorter after each cell division and cells can no longer divide when telomere length becomes too short. Tumor cells require mechanisms to overcome telomere shortening problem to be able to divide infinitely. One way to solve telomere shortening problem is to reactivate telomerase reverse transcriptase (TERT) to synthesize telomeric DNA and prevent telomeres from shortening. TERT may be activated via promoter mutations. Here we determine mutation status of TERT promoter in established liver cancer cell lines and patient tumor samples.
TERT promoter mutations have first been defined in melanoma and they are claimed to create new binding sites for specific transcription factors and increase TERT expression. This may be used by tumor cells as a mechanism to overcome telomere shortening problem so it is important to show the presence of the same mutations in the promoter region of TERT and determine their frequencies in HCC. Deficiencies of early diagnosis and systemic therapy of liver cancer are the major causes of its high mortality. Screening of TERT promoter status may help early diagnosis of tumor formation in patients with chronic liver disease. In addition, targeting of TERT promoter mutations may open new horizons for specific therapies of liver cancer.
Telomerase reactivation is common to liver cancer samples, and TERT promoter mutations have been reported recently. Tumor samples were collected from hospitals from counties such as France and United States, where liver cancer is not a major disease contrary to some other countries located in Asia (China and Japan) and Africa (southern African countries) with a very high incidence. Thus, it was not clear how common TERT promoter mutations were over the world, especially in Africa and Asia. In this present study, we tried to show that TERT promoter mutations are common in hepatocellular carcinoma (HCC), regardless of geographical location. Moreover, this research showed that HCCs from Africa are more likely to carry TERT promoter mutations, in comparison with Non-African tumors.
The high frequency of TERT promoter mutations resulting from the present study suggests that these mutations are critical or may be necessary for liver tumor formation. Therefore, they can be used for diagnostic or prognostic purposes for patient care. Furthermore, if such mutations are causing tumor-specific reactivation of telomerase activity, they may serve as tumor-selective targets for novel therapies.
Hepatocellular carcinoma is a primary liver cancer. Telomeres are DNA sequences located on the tips of chromosomes. TERT gene encodes for an enzyme responsible for the synthesis of telomeric DNA.
The authors determined mutation status of human TERT promoter region in HCCs from different geographical regions. Although some articles have the same scope but the new item is the effect of different geographical locations, the article is well-organized and is perfectly written.
P- Reviewer: Abdel-Raheem IT, Chen GY, Liu ZH, Santoro N S- Editor: Ding Y L- Editor: A E- Editor: Liu XM
| 1. | Lin S, Hoffmann K, Schemmer P. Treatment of hepatocellular carcinoma: a systematic review. Liver Cancer. 2012;1:144-158. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 216] [Cited by in RCA: 279] [Article Influence: 21.5] [Reference Citation Analysis (0)] |
| 2. | Venook AP, Papandreou C, Furuse J, de Guevara LL. The incidence and epidemiology of hepatocellular carcinoma: a global and regional perspective. Oncologist. 2010;15 Suppl 4:5-13. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 626] [Cited by in RCA: 736] [Article Influence: 49.1] [Reference Citation Analysis (0)] |
| 3. | Villanueva A, Hernandez-Gea V, Llovet JM. Medical therapies for hepatocellular carcinoma: a critical view of the evidence. Nat Rev Gastroenterol Hepatol. 2013;10:34-42. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 234] [Cited by in RCA: 253] [Article Influence: 21.1] [Reference Citation Analysis (0)] |
| 4. | Ozen C, Yildiz G, Dagcan AT, Cevik D, Ors A, Keles U, Topel H, Ozturk M. Genetics and epigenetics of liver cancer. N Biotechnol. 2013;30:381-384. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 71] [Cited by in RCA: 77] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 5. | Siegel AB, Olsen SK, Magun A, Brown RS. Sorafenib: where do we go from here? Hepatology. 2010;52:360-369. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 94] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 6. | Cesare AJ, Reddel RR. Alternative lengthening of telomeres: models, mechanisms and implications. Nat Rev Genet. 2010;11:319-330. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 653] [Cited by in RCA: 738] [Article Influence: 49.2] [Reference Citation Analysis (0)] |
| 7. | Paterlini-Bréchot P, Saigo K, Murakami Y, Chami M, Gozuacik D, Mugnier C, Lagorce D, Bréchot C. Hepatitis B virus-related insertional mutagenesis occurs frequently in human liver cancers and recurrently targets human telomerase gene. Oncogene. 2003;22:3911-3916. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 239] [Cited by in RCA: 247] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 8. | Fujimoto A, Totoki Y, Abe T, Boroevich KA, Hosoda F, Nguyen HH, Aoki M, Hosono N, Kubo M, Miya F. Whole-genome sequencing of liver cancers identifies etiological influences on mutation patterns and recurrent mutations in chromatin regulators. Nat Genet. 2012;44:760-764. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 631] [Cited by in RCA: 713] [Article Influence: 54.8] [Reference Citation Analysis (0)] |
| 9. | Sung WK, Zheng H, Li S, Chen R, Liu X, Li Y, Lee NP, Lee WH, Ariyaratne PN, Tennakoon C. Genome-wide survey of recurrent HBV integration in hepatocellular carcinoma. Nat Genet. 2012;44:765-769. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 619] [Cited by in RCA: 722] [Article Influence: 55.5] [Reference Citation Analysis (0)] |
| 10. | Huang FW, Hodis E, Xu MJ, Kryukov GV, Chin L, Garraway LA. Highly recurrent TERT promoter mutations in human melanoma. Science. 2013;339:957-959. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1525] [Cited by in RCA: 1451] [Article Influence: 120.9] [Reference Citation Analysis (0)] |
| 11. | Horn S, Figl A, Rachakonda PS, Fischer C, Sucker A, Gast A, Kadel S, Moll I, Nagore E, Hemminki K. TERT promoter mutations in familial and sporadic melanoma. Science. 2013;339:959-961. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1305] [Cited by in RCA: 1398] [Article Influence: 116.5] [Reference Citation Analysis (0)] |
| 12. | Killela PJ, Reitman ZJ, Jiao Y, Bettegowda C, Agrawal N, Diaz LA, Friedman AH, Friedman H, Gallia GL, Giovanella BC. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc Natl Acad Sci USA. 2013;110:6021-6026. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 979] [Cited by in RCA: 1119] [Article Influence: 93.3] [Reference Citation Analysis (0)] |
| 13. | Liu X, Wu G, Shan Y, Hartmann C, von Deimling A, Xing M. Highly prevalent TERT promoter mutations in bladder cancer and glioblastoma. Cell Cycle. 2013;12:1637-1638. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 100] [Cited by in RCA: 116] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 14. | Nonoguchi N, Ohta T, Oh JE, Kim YH, Kleihues P, Ohgaki H. TERT promoter mutations in primary and secondary glioblastomas. Acta Neuropathol. 2013;126:931-937. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 171] [Cited by in RCA: 186] [Article Influence: 15.5] [Reference Citation Analysis (0)] |
| 15. | Nault JC, Mallet M, Pilati C, Calderaro J, Bioulac-Sage P, Laurent C, Laurent A, Cherqui D, Balabaud C, Zucman-Rossi J. High frequency of telomerase reverse-transcriptase promoter somatic mutations in hepatocellular carcinoma and preneoplastic lesions. Nat Commun. 2013;4:2218. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 414] [Cited by in RCA: 511] [Article Influence: 46.5] [Reference Citation Analysis (1)] |
| 16. | Landa I, Ganly I, Chan TA, Mitsutake N, Matsuse M, Ibrahimpasic T, Ghossein RA, Fagin JA. Frequent somatic TERT promoter mutations in thyroid cancer: higher prevalence in advanced forms of the disease. J Clin Endocrinol Metab. 2013;98:E1562-E1566. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 306] [Cited by in RCA: 329] [Article Influence: 27.4] [Reference Citation Analysis (0)] |
| 17. | Acun T, Terzioğlu-Kara E, Konu O, Ozturk M, Yakicier MC. Mdm2 Snp309 G allele displays high frequency and inverse correlation with somatic P53 mutations in hepatocellular carcinoma. Mutat Res. 2010;684:106-108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 9] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 18. | Unsal H, Yakicier C, Marçais C, Kew M, Volkmann M, Zentgraf H, Isselbacher KJ, Ozturk M. Genetic heterogeneity of hepatocellular carcinoma. Proc Natl Acad Sci USA. 1994;91:822-826. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 116] [Cited by in RCA: 109] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 19. | Yuzugullu H, Benhaj K, Ozturk N, Senturk S, Celik E, Toylu A, Tasdemir N, Yilmaz M, Erdal E, Akcali KC. Canonical Wnt signaling is antagonized by noncanonical Wnt5a in hepatocellular carcinoma cells. Mol Cancer. 2009;8:90. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 142] [Cited by in RCA: 169] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 20. | Tahara H, Nakanishi T, Kitamoto M, Nakashio R, Shay JW, Tahara E, Kajiyama G, Ide T. Telomerase activity in human liver tissues: comparison between chronic liver disease and hepatocellular carcinomas. Cancer Res. 1995;55:2734-2736. [PubMed] |
| 21. | Ozturk M, Arslan-Ergul A, Bagislar S, Senturk S, Yuzugullu H. Senescence and immortality in hepatocellular carcinoma. Cancer Lett. 2009;286:103-113. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 64] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 22. | Wu KJ, Grandori C, Amacker M, Simon-Vermot N, Polack A, Lingner J, Dalla-Favera R. Direct activation of TERT transcription by c-MYC. Nat Genet. 1999;21:220-224. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 658] [Cited by in RCA: 672] [Article Influence: 25.8] [Reference Citation Analysis (0)] |
| 23. | Hoffmeyer K, Raggioli A, Rudloff S, Anton R, Hierholzer A, Del Valle I, Hein K, Vogt R, Kemler R. Wnt/β-catenin signaling regulates telomerase in stem cells and cancer cells. Science. 2012;336:1549-1554. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 387] [Cited by in RCA: 413] [Article Influence: 31.8] [Reference Citation Analysis (0)] |
| 24. | Greider CW. Molecular biology. Wnt regulates TERT--putting the horse before the cart. Science. 2012;336:1519-1520. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 29] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 25. | Kyo S, Inoue M. Complex regulatory mechanisms of telomerase activity in normal and cancer cells: how can we apply them for cancer therapy? Oncogene. 2002;21:688-697. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 130] [Cited by in RCA: 128] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 26. | Bernardes de Jesus B, Blasco MA. Telomerase at the intersection of cancer and aging. Trends Genet. 2013;29:513-520. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 141] [Cited by in RCA: 162] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 27. | Cagatay T, Ozturk M. P53 mutation as a source of aberrant beta-catenin accumulation in cancer cells. Oncogene. 2002;21:7971-7980. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 87] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 28. | Young AP, Sigman DS. Conformational effects of volatile anesthetics on the membrane-bound acetylcholine receptor protein: facilitation of the agonist-induced affinity conversion. Biochemistry. 1983;22:2155-2162. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 21] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 29. | van Zijl F, Zulehner G, Petz M, Schneller D, Kornauth C, Hau M, Machat G, Grubinger M, Huber H, Mikulits W. Epithelial-mesenchymal transition in hepatocellular carcinoma. Future Oncol. 2009;5:1169-1179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 236] [Cited by in RCA: 270] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 30. | Ozturk N, Erdal E, Mumcuoglu M, Akcali KC, Yalcin O, Senturk S, Arslan-Ergul A, Gur B, Yulug I, Cetin-Atalay R. Reprogramming of replicative senescence in hepatocellular carcinoma-derived cells. Proc Natl Acad Sci USA. 2006;103:2178-2183. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 47] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 31. | Yildiz G, Arslan-Ergul A, Bagislar S, Konu O, Yuzugullu H, Gursoy-Yuzugullu O, Ozturk N, Ozen C, Ozdag H, Erdal E. Genome-wide transcriptional reorganization associated with senescence-to-immortality switch during human hepatocellular carcinogenesis. PLoS One. 2013;8:e64016. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 48] [Article Influence: 4.0] [Reference Citation Analysis (0)] |