Published online Jun 26, 2019. doi: 10.12998/wjcc.v7.i12.1522
Peer-review started: January 28, 2019
First decision: March 9, 2019
Revised: March 20, 2019
Accepted: April 18, 2019
Article in press: May 2, 2019
Published online: June 26, 2019
Processing time: 152 Days and 3.2 Hours
To screen for possible pathogenic loci in a patient with Gitelman syndrome by high-throughput exome sequencing and to explore the relationship between genotype and phenotype.
The clinical data of the patient were collected. Peripheral blood samples were obtained to isolate white blood cells and extract genomic DNA. High-throughput whole exome sequencing for candidate pathogenic genes in the proband was completed by the Huada Gene Technology Co. Ltd (Shenzhen, China). Sequencing showed a novel heterozygous missense mutation (a G to A transition at nucleotide 2582) in exon 22 of the SLC12A3 gene, which resulted in a substitution of histidine for arginine at position 816 of the LRP1B protein and caused the occurrence of disease.
This is the first report of a new pathogenic mutation in SLC12A3. Further functional studies are particularly necessary to explore potential molecular mechanisms.
Core tip: To screen for possible pathogenic loci in a patient with Gitelman syndrome by high-throughput exome sequencing and to explore the relationship between the genotype and phenotype. Sequencing showed a novel heterozygous missense mutation (a G to A transition at nucleotide 2582) in exon 22 of SLC12A3 gene, which resulted in a substitution of histidine for arginine at position 816 of the LRP1B protein and caused the occurrence of disease.
- Citation: Wang CL. Novel heterozygous missense mutation of SLC12A3 gene in Gitelman syndrome: A case report. World J Clin Cases 2019; 7(12): 1522-1528
- URL: https://www.wjgnet.com/2307-8960/full/v7/i12/1522.htm
- DOI: https://dx.doi.org/10.12998/wjcc.v7.i12.1522
Gitelman syndrome (GS) is an inherited autosomal recessive renal tubular disorder that was first described by Gitelman in 1966. The main clinical manifestations include hypokalemia, hypomagnesia, hypocalciuria, and hypochloremic metabolic alkalosis[1-3]. GS is often found in infants and young children with growth retardation and convulsions. Patients usually have normal blood pressure. The prevalence of GS ranges from 1/1000 to 9/10000. It is easily neglected due to their mild clinical manifestations and good prognosis. Several studies have shown that GS may be associated with chondrocalcinosis and dysglycemia; in severe cases, the patients may also develop ventricular arrhythmia and progressive renal insufficiency, which can be highly dangerous.
The main pathogenic gene in GS is SLC12A3, which encodes for thiazide-sensitive NaCl cotransporter. The rapid development of gene sequencing technology in recent years has facilitated the gene diagnosis[4-6]. According to expert consensus, the sequencing gene panels for GS should include the SLC12A3, CLCNKB, and HNF1B genes[7]. Whole exome sequencing (WES) can detect exon regions of over 20000 genes at a time. With the decrease in its price, WES has been increasingly used in clinical diagnosis[8]. Therefore, we applied WES for the genetic analysis in a clinically confirmed GS patient. In addition to SLC12A3, the most common gene associated with GS, we also detected CLCNKB and HNF1B[9-11]. We report a patient with clinically confirmed GS and determined the relevant gene mutation loci in an attempt to further improve our understanding of this disease.
A sudden onset of limb weakness without obvious cause, followed by limb numbness/stiffness, which was accompanied by palpitation.
The patient was a 16-year-old male. He was admitted in January 2018 due to limb weakness and stiffness for two years. Two years ago, the patient had a sudden onset of limb weakness without obvious cause, followed by limb numbness/stiffness, which was accompanied by palpitation. Examination in a local hospital revealed hypokalemia, which was improved after potassium supplementation. However, the above symptoms recurred 2 mo ago due to cold, and the patient was admitted to our hospital for further treatment.
He denied any other medical conditions.
There was no history of consanguineous marriage in the pedigree of three generations. The study was approved by the Ethics Committee of Shanxi Provincial People’s Hospital, Taiyuan, China. The proband and his family members signed the informed consent.
The thyroid gland was not large. There was no obvious abnormality in the heart and lungs.
Blood analysis: potassium, 2.64 mmol/L; sodium, 133.10 mmol/L; chlorine, 96.20 mmol/L; magnesium, 0.510 mmol/L; triglycerides, 1.64 mmol/L; blood pH, 7.35; standard bicarbonate, 25.60 mmol/L; and total carbon dioxide, 20.00 mmol/L. Urine analysis showed: calcium, 0.12 mmol/24 h; magnesium 2.200 mmol/24 h, phosphorus, 2.19 mmol/24 h; during the same period the blood potassium was 3.05 mmol/L and magnesium was 0.562 mmol/L. Circadian and pulsatile secretion of adrenocorticotropic hormone and cortisol were normal. Baseline renin-angiotensin-aldosterone system test: Angiotensin I (37 °C), 49.94 μg/L; angiotensin I (4 °C), 6.87 μg/L; aldosterone, 149.05 ng/L; renin activity, 31.87 UG/L per hour, and aldosterone/renin activity 0.47. The average 24-h ambulatory blood pressure was 105/71 mmHg.
No abnormality was seen on X-ray chest film, abdominal ultrasound, thyroid ultrasound, bilateral kidney and renal vascular ultrasound, adrenal ultrasound, and adrenal thin-slice computed tomography. Electrocardiogram showed sinus tachycardia at 105 beats/min.
DNA extraction: Peripheral venous blood (2 mL) was collected with heparin as anticoagulant. Genomic DNA was isolated from peripheral blood lymphocytes using OMEGA SE Blood DNA Kit and then sent to the Shenzhen Huada Gene Technology Co. Ltd for WES.
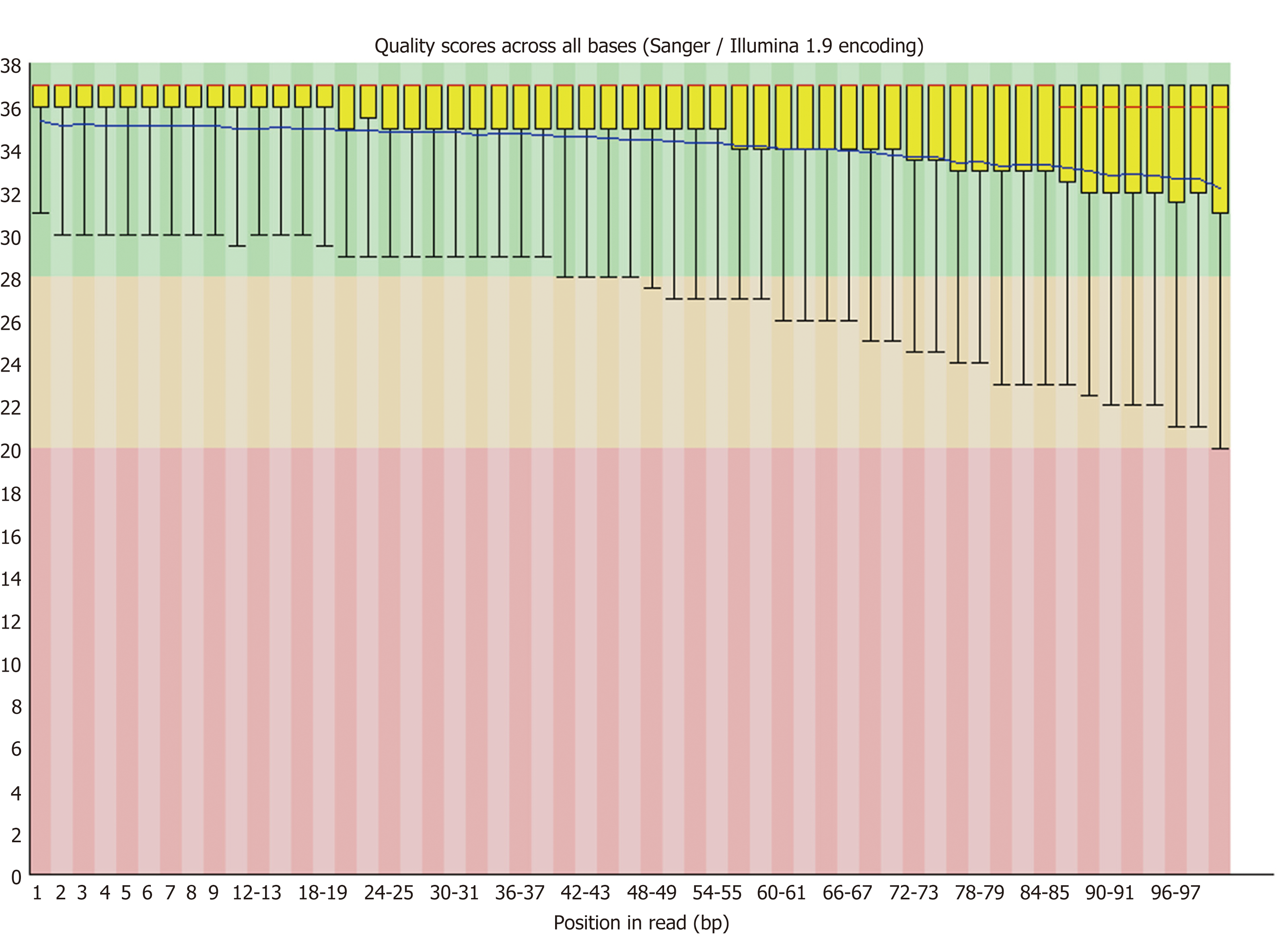
Bioinformatics analysis: Quality control of the raw reads was managed via FastQC[12]. Sequences were aligned to human reference genome hg19 using the Burrows-Wheeler Aligner[13]. The duplicate reads were removed by the Samblaster[14]. The INDEL was re-aligned using GATK realignment and base quality score recalibration was performed. We used five kinds of software to analyze variation, including GATK, Samtools, Freebayes, Platypus, and Varscan2, to ensure the accuracy of identification. Marginal variants were annotated in databases including dbSNP, 1000 Genomes Project, dbNSFP, and ClinVar[15-17]. The possible pathogenic mutations on SLC12A3, CLCNKB, and HNF1B genes were analyzed, and the relevant literature was searched according to these loci.
The quality control results of the raw reads (Fastq) are shown in Figures 1 and 2. The average value of base qualities was larger than 30 (accuracy: 99.9%).
WES identified a total of 214137288 reads, among which 99.83% could be mapped to the human reference genome, and the duplicate reads accounted for 11.81%. The mean depth was 282X, which exceeded the general exome sequencing depth (Table 1).
| Total reads | Mapped reads | Duplicate reads | Mean depth |
| 214137288 | 99.83% | 11.81% | 282X |
A total of 67537 mutations were identified by bioinformatics analysis, including 55184 SNPs and 12353 INDELs (Figure 3). After dbSNP annotation, 94% of the SNPs were annotated in dbSNP, while only 35% of the INDELs could be annotated in dbSNP.
Mutations in SLC12A3, CLCNKB, and HNF1B3 genes were filtered based on the following conditions: (1) The variant is located on an exon; (2) The variation does not belong to synonymous mutation; and (3) Population frequency is greater than 0.001.
After filtering, only one missense heterozygous mutation in the SLC12A3 gene was left. Its population frequency was unknown. Most mutation prediction software such as Polyphen2 HDIV, SIFT, and FATHMM predicted it as a harmful mutation. The mutation information is shown in Table 2, and the mutation of exon 22 reported by another paper is shown in Table 3.
| Type | Information |
| Gene | SLC12A3 |
| RNA | NM_000339 |
| Exon | exon22 |
| DNA mutation | G2582A |
| AA mutation | R861H |
| Mutation frequency | 50% |
| Population frequency | Unknown |
| Polyphen2_HDIV | D, D, D |
| FATHMM | D |
| MutationTaster | D |
| MutationAssessor | L |
| LRT | D |
| SIFT | T |
| Exon | Mutation | Pmid |
| exon22 | Glv876Ser | 17654016 |
| exon22 | Leu849His | 17873326, 20229814 |
| exon22 | Arg852His | 17873326, 20229814 |
| exon22 | Arg861Cys | 27872838 |
| exon22 | Arg871His | 21051746 |
| exon22 | Leu859Pro | 21753071 |
| exon22 | Arg861Cys | 21753071 |
| exon22 | Arg861His | Present study |
According to the typical symptoms, laboratory tests, and gene analysis, the patient was diagnosed with GS.
The patient was given potassium therapy with antisterone.
The patient recovered well and was discharged 7 d later. Regular detection of potassium is necessary.
WES can detect the exon information of all genes at one time. With the decreased cost of high-throughput next-generation sequencing, WES has been increasingly applied in clinical diagnoses. In the present study, we used WES to further clarify the gene mutations in our patient. After bioinformatics analysis and population frequency filtering, we found a non-synonymous mutation in SLC12A3 gene. A G2582A heterozygous mutation has also been reported in this site in the literature[18].
Mutation analysis of the SLC12A3 gene in our patient and his family members revealed a heterozygous missense mutation of G-to-A transition at nucleotide position 2582 within exon 22. An autosomal recessive disease does not present its traits in the heterozygous state. It occurs only when a pair of alleles is homozygous or compound heterozygotes of a recessive pathogenic gene. However, Balavoine et al[19] detected two mutation sites in the SLC12A3 gene in most GS patients and only one mutation site in a small number of GS patients. In addition, patients with two mutation sites have more severe clinical symptoms than those with only one mutation site. GS is an autosomal recessive hereditary disease, and it does not occur in carriers. Current clinical studies have not found a significant correlation between GS genotype and phenotype.
With the decreased cost of sequencing and better understanding of diseases, the concept of precision medicine has been widely recognized over the past two years. Precision medicine represents the future direction of medical development. The core of precision medicine is to precisely identify pathogenic gene sites or pathogenic loci by gene sequencing and carry out targeted therapy according to pathogenic genes or pathogenic sites.
A novel heterozygous missense mutation (a G to A transition at nucleotide 2582) in exon 22 of the SLC12A3 gene is the first report of a new pathogenic mutation in SLC12A3. Further functional studies are particularly necessary to explore potential molecular mechanisms.
Manuscript source: Unsolicited manuscript
Specialty type: Medicine, research and experimental
Country of origin: China
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): 0
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Cheungpasitporn W, Stavroulopoulos A S-Editor: Ji FF L-Editor: Filipodia E-Editor: Liu JH
| 1. | van der Merwe PD, Rensburg MA, Haylett WL, Bardien S, Davids MR. Gitelman syndrome in a South African family presenting with hypokalaemia and unusual food cravings. BMC Nephrol. 2017;18:38. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 2. | Chen Q, Wu Y, Zhao J, Jia Y, Wang W. A case of hypokalemia and proteinuria with a new mutation in the SLC12A3 Gene. BMC Nephrol. 2018;19:275. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 6] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 3. | Gu X, Su Z, Chen M, Xu Y, Wang Y. Acquired Gitelman syndrome in a primary Sjögren syndrome patient with a SLC12A3 heterozygous mutation: A case report and literature review. Nephrology (Carlton). 2017;22:652-655. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 29] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 4. | Chen Y, Zhang Z, Lin X, Pan Q, Zheng F, Li H. A novel compound heterozygous variant of the SLC12A3 gene in Gitelman syndrome pedigree. BMC Med Genet. 2018;19:17. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 6] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 5. | Xia MF, Bian H, Liu H, Wu HJ, Zhang ZG, Lu ZQ, Gao X. Hypokalemia, hypomagnesemia, hypocalciuria, and recurrent tetany: Gitelman syndrome in a Chinese pedigree and literature review. Clin Case Rep. 2017;5:578-586. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 7] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 6. | Takeuchi Y, Mishima E, Shima H, Akiyama Y, Suzuki C, Suzuki T, Kobayashi T, Suzuki Y, Nakayama T, Takeshima Y, Vazquez N, Ito S, Gamba G, Abe T. Exonic mutations in the SLC12A3 gene cause exon skipping and premature termination in Gitelman syndrome. J Am Soc Nephrol. 2015;26:271-279. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 34] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 7. | Zhou H, Liang X, Qing Y, Meng B, Zhou J, Huang S, Lu S, Huang Z, Yang H, Ma Y, Luo Z. Complicated Gitelman syndrome and autoimmune thyroid disease: a case report with a new homozygous mutation in the SLC12A3 gene and literature review. BMC Endocr Disord. 2018;18:82. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 19] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 8. | Lee JW, Lee J, Heo NJ, Cheong HI, Han JS. Mutations in SLC12A3 and CLCNKB and Their Correlation with Clinical Phenotype in Patients with Gitelman and Gitelman-like Syndrome. J Korean Med Sci. 2016;31:47-54. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 34] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 9. | Subasinghe CJ, Sirisena ND, Herath C, Berge KE, Leren TP, Bulugahapitiya U, Dissanayake VHW. Novel mutation in the SLC12A3 gene in a Sri Lankan family with Gitelman syndrome & coexistent diabetes: a case report. BMC Nephrol. 2017;18:140. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 10] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 10. | Peng X, Jiang L, Chen C, Qin Y, Yuan T, Wang O, Xing X, Li X, Nie M, Chen L. Increased urinary prostaglandin E2 metabolite: A potential therapeutic target of Gitelman syndrome. PLoS One. 2017;12:e0180811. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 11. | Kusuda T, Hosoya T, Mori T, Ihara K, Nishida H, Chiga M, Sohara E, Rai T, Koike R, Uchida S, Kohsaka H. Acquired Gitelman Syndrome in an Anti-SSA Antibody-positive Patient with a SLC12A3 Heterozygous Mutation. Intern Med. 2016;55:3201-3204. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 13] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 12. | Peng X, Zhao B, Zhang L, Jiang L, Yuan T, Wang Y, Wang H, Ma J, Li N, Zheng K, Nie M, Li X, Xing X, Chen L. Hydrochlorothiazide Test as a Tool in the Diagnosis of Gitelman Syndrome in Chinese Patients. Front Endocrinol (Lausanne). 2018;9:559. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 9] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 13. | Al-Shibli A, Yusuf M, Abounajab I, Willems PJ. Mixed Bartter-Gitelman syndrome: an inbred family with a heterogeneous phenotype expression of a novel variant in the CLCNKB gene. Springerplus. 2014;3:96. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 14. | Makino S, Tajima T, Shinozuka J, Ikumi A, Awaguni H, Tanaka S, Maruyama R, Imashuku S. Gitelman Syndrome in a School Boy Who Presented with Generalized Convulsion and Had a R642H/R642W Mutation in the SLC12A3 Gene. Case Rep Pediatr. 2014;2014:279389. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 15. | Fujimura J, Nozu K, Yamamura T, Minamikawa S, Nakanishi K, Horinouchi T, Nagano C, Sakakibara N, Nakanishi K, Shima Y, Miyako K, Nozu Y, Morisada N, Nagase H, Ninchoji T, Kaito H, Iijima K. Clinical and Genetic Characteristics in Patients With Gitelman Syndrome. Kidney Int Rep. 2018;4:119-125. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 44] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 16. | Gug C, Mihaescu A, Mozos I. Two mutations in the thiazide-sensitive NaCl co-transporter gene in a Romanian Gitelman syndrome patient: case report. Ther Clin Risk Manag. 2018;14:149-155. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 12] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 17. | Mishima E, Mori T, Sohara E, Uchida S, Abe T, Ito S. Inherited, not acquired, Gitelman syndrome in a patient with Sjögren's syndrome: importance of genetic testing to distinguish the two forms. CEN Case Rep. 2017;6:180-184. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 11] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 18. | Yang W, Zhao S, Xie Y, Mo Z. A novel SLC12A3 homozygous c2039delG mutation in Gitelman syndrome with hypocalcemia. BMC Nephrol. 2018;19:362. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 19. | Balavoine AS, Bataille P, Vanhille P, Azar R, Noël C, Asseman P, Soudan B, Wémeau JL, Vantyghem MC. Phenotype-genotype correlation and follow-up in adult patients with hypokalaemia of renal origin suggesting Gitelman syndrome. Eur J Endocrinol. 2011;165:665-673. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 36] [Article Influence: 2.6] [Reference Citation Analysis (0)] |