Revised: February 14, 2013
Accepted: April 9, 2013
Published online: May 12, 2013
Processing time: 155 Days and 3.4 Hours
The paramyxoviruses are a family of > 30 viruses that variously infect humans, other mammals and fish to cause diverse outcomes, ranging from asymptomatic to lethal disease, with the zoonotic paramyxoviruses Nipah and Hendra showing up to 70% case-fatality rate in humans. The capacity to evade host immunity is central to viral infection, and paramyxoviruses have evolved multiple strategies to overcome the host interferon (IFN)-mediated innate immune response through the activity of their IFN-antagonist proteins. Although paramyxovirus IFN antagonists generally target common factors of the IFN system, including melanoma differentiation associated factor 5, retinoic acid-inducible gene-I, signal transducers and activators of transcription (STAT)1 and STAT2, and IFN regulatory factor 3, the mechanisms of antagonism show remarkable diversity between different genera and even individual members of the same genus; the reasons for this diversity, however, are not currently understood. Here, we review the IFN antagonism strategies of paramyxoviruses, highlighting mechanistic differences observed between individual species and genera. We also discuss potential sources of this diversity, including biological differences in the host and/or tissue specificity of different paramyxoviruses, and potential effects of experimental approaches that have largely relied on in vitro systems. Importantly, recent studies using recombinant virus systems and animal infection models are beginning to clarify the importance of certain mechanisms of IFN antagonism to in vivo infections, providing important indications not only of their critical importance to virulence, but also of their potential targeting for new therapeutic/vaccine approaches.
Core tip: The paramyxoviruses are a family of > 30 viruses that variously infect humans, other mammals and fish to cause diverse outcomes, ranging from asymptomatic to lethal disease, with the zoonotic paramyxoviruses Nipah and Hendra showing up to 70% case-fatality rate in humans. Here, we review the interferon antagonism strategies of paramyxoviruses, highlighting mechanistic differences observed between individual species and genera. We also discuss potential sources of this diversity, including biological differences in the host and/or tissue specificity of different paramyxoviruses, and potential effects of experimental approaches that have largely relied on in vitro systems.
- Citation: Audsley MD, Moseley GW. Paramyxovirus evasion of innate immunity: Diverse strategies for common targets. World J Virol 2013; 2(2): 57-70
- URL: https://www.wjgnet.com/2220-3249/full/v2/i2/57.htm
- DOI: https://dx.doi.org/10.5501/wjv.v2.i2.57
Since the discovery more than 50 years ago of type I interferons (IFNs) as the principal mediators of mammalian innate antiviral responses, it has become increasingly evident that infection by viruses depends on the capacity to counteract host cell IFN responses. Viruses have evolved diverse strategies to antagonise IFN responses, often by hijacking and modifying cellular regulatory pathways through the activity of specific viral IFN-antagonist proteins. Among the best-studied viruses in this respect are the paramyxoviruses, which include established human pathogens such as measles virus (MeV) and mumps virus (MuV), and emerging zoonotic viruses such as the henipaviruses Nipah virus (NiV) and Hendra virus (HeV). Although effective vaccines are available for MeV, it remains a leading cause of fatalities in children, with almost 140000 human deaths globally in 2010[1], while the henipaviruses show remarkable pathogenicity, with case-fatality rates between 40%-70% in humans[2-5].
The paramyxoviruses are a subfamily of the Paramyxoviridae family [order Mononegavirales (MNV)] of enveloped, non-segmented negative-strand RNA viruses (NNSV), which also includes the Pneumovirus subfamily[6,7]. Based largely on antigenic cross-reactivity and neuramidase activity paramyxoviruses are currently classified into seven genera[6,7]: Rubulavirus, Avulavirus, Henipavirus, Morbillivirus, Respirovirus, Ferlavirus and Aquaparamyxovirus (Table 1). Members of the paramyxovirus family show diverse tissue tropism and infect a variety of species in a fashion that does not appear to be specific to genus classification (Table 1). Because their relatively small genomes lack dedicated IFN-antagonist genes, paramyxoviruses generally encode IFN-antagonists as accessory protein isoforms encoded within their conserved P genes, another factor in genus classification[6]. These IFN antagonists broadly target several members of a select group of signalling molecules of the IFN system, including melanoma differentiation associated factor 5 (MDA5), retinoic acid-inducible gene-I (RIG)-I, IFN regulatory factor (IRF)-3, and signal transducers and activators of transcription (STAT)1 and STAT2, but use diverse mechanisms including proteosomal degradation, inhibition of phosphorylation, and subcellular mis-localisation. Intriguingly, the mechanisms can vary significantly between different genera and, in some cases, different species of the same genera.
| Genus | Virus | Major host |
| Morbillivirus | Measles virus1 | Human |
| Canine distemper virus | Canine | |
| Rinderpest virus | Bovine | |
| Peste-des-petits-ruminants virus | Caprine | |
| Phocine distemper virus | Phocine | |
| Cetacean morbillivirus | Cetacean | |
| Rubulavirus | Mumps virus1 | Human |
| Parainfluenza virus 5 (previously, Simian virus 5) | Human | |
| Human parainfluenza virus 2, 4a and 4b | Human | |
| Mapuera virus | Chiropteran2 | |
| Porcine rubulavirus | Porcine | |
| Respirovirus | Sendai virus1 | Murine |
| Human parainfluenza virus 1, 3 | Human | |
| Bovine parainfluenza virus 3 | Bovine | |
| Avulavirus | Newcastle disease virus1 | Avian |
| Avian paramyxoviruses 2-9 | Avian | |
| Henipavirus | Hendra virus1 | Chiropteran/equine /human3 |
| Nipah virus | Chiropteran/porcine/human3 | |
| Cedar virus | Chiropteran2 | |
| Aquaparamyxovirus | Atlantic salmon paramyxovirus1 | Piscine |
| Ferlavirus | Fer-de-Lance virus1 | Serpentine |
| Unassigned | J-virus | Murine |
| Beilong virus | Murine | |
| Tailam virus | Murine | |
| Menangle virus | Porcine | |
| Tioman virus | Chiropteran2 | |
| Tupaia virus | Chiropteran2 | |
| Salem virus | Chiropteran2 | |
| Mossman virus | Chiropteran2 | |
| Nariva virus | Chiropteran2 | |
| Pigeon paramyxovirus 1 | Avian |
Here we review the mechanistic data relating to paramyxovirus IFN antagonism with a focus on common and distinct features within the family, before discussing possible origins of the diversity within the family. Although much of the available research on paramyxovirus IFN antagonism has been restricted to in vitro studies, recent findings using in vivo infection and recombinant virus systems point to a pivotal role in pathogenicity that may provide potent targets for the development of new vaccines/antiviral therapeutics.
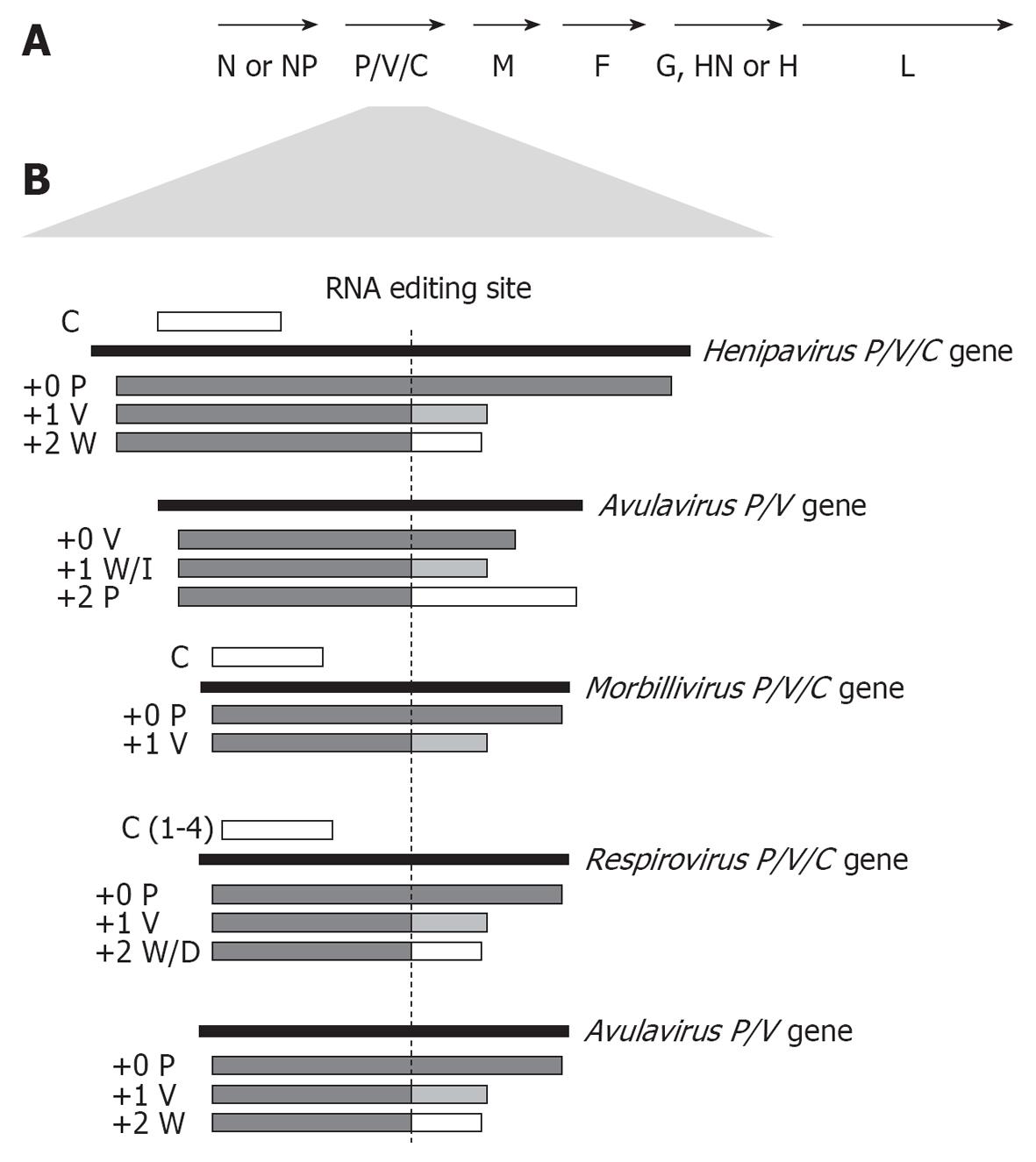
While viruses with large genomes can encode dedicated IFN-antagonist proteins, the high error rates of the RNA-dependent RNA polymerase means that RNA viruses generally have restricted genome sizes, with the paramyxovirus genome containing only six principal genes to express essential structural/replication factors, specifically M (matrix), G/HN/H (attachment), F (fusion), L (polymerase), N/NP (nucleocapsid) and P (phosphoprotein) (Figure 1A). Thus the IFN-antagonists of RNA viruses are often encoded as “accessory” protein isoforms within one or more of the conserved genes[8,9]; in paramyxoviruses up to 9 proteins are encoded in the P gene, including V, C, and P proteins and a protein variously named W, D or I, which have established IFN antagonist functions.
Isoform expression from the paramyxovirus P gene is variously achieved by a conserved RNA-editing mechanism, and through the use of internal start codons and alternate open reading frames (ORFs). RNA editing is mediated by the viral RNA-dependent RNA polymerase through the insertion of additional non-coded guanosine (G) nucleotides into P gene mRNA transcripts at a predetermined purine rich editing site. This causes a +1 or +2 frameshift in the downstream ORF[10-12] which results in the generation of two or three distinct proteins (P, V and W/D/I), which have common N-terminal sequences but unique C-termini (Figure 1B). A comparable editing process is used by Ebolavirus of the Filovirus family to produce isoforms from its G gene[13]. This mechanism is conserved among all paramyxoviruses examined except human parainfluenza virus (hPIV) 1 and the recently discovered cedar virus[14,15].
P protein, the polymerase cofactor essential to genome transcription/replication processes, is usually generated from the unedited ORF as the principal P gene product, with the production of edited RNA varying in a broadly genus-specific fashion (Figure 1B), although the +1 frameshift commonly encodes V protein and the +2 frameshift W/D/I[16,17]. Members of the Rubulavirus genus uniquely encode V protein in the unedited transcript, with P protein expression requiring editing (Figure 1B)[9], with c. 63% of the P gene mRNA transcribed unedited by the rubulavirus MuV, indicative of a particular requirement for high levels of V protein by these viruses[9]. The henipaviruses have the highest editing frequency of the paramyxoviruses: 66% to 94% of transcripts are edited, compared with c. 42% for MeV (Morbillivirus genus), and c. 31% for Sendai virus (SeV) (Respirovirus genus)[18-21]. Henipaviruses insert up to 11 additional G nucleotides[18], and in NiV-infected cells P transcripts are detected at the highest levels (c. 60%-100%) early in infection, with V and W transcripts peaking between 9.5-24 h post-infection (up to 59% and 37% respectively). This suggests that editing is regulated to enable particularly important roles for V and W late in infection[18], although other factors such as mRNA/protein stability are likely to affect the final levels of protein.
Henipaviruses, morbilliviruses, and respiroviruses use a start codon within the P gene in an alternate ORF to produce a C protein (Figure 1B), while the SeV P gene encodes up to five proteins other than P, V and W: four C proteins encoded by overlapping sequences in the +1 reading frame, and X protein, a truncated version of P protein translated from an internal start site[11,22,23]. HeV, but not NiV, encodes a putative SB (short basic) protein, homologous to SB of several viruses of other MNV families[21]. These differences in P gene coding capacity indicate different requirements of specific viruses for accessory proteins, possibly due to host/tissue specific aspects of IFN signalling (see below).
The V proteins are generally considered the principal IFN-antagonists of paramyxoviruses, and are the best studied of the P gene accessory proteins. However, there is increasing evidence that P, W, or C proteins of paramyxoviruses including NiV, MeV, and SeV play important roles in IFN antagonism by distinct mechanisms. Thus, it seems that most if not all P gene accessory proteins have evolved for roles in immune evasion as important pathogenicity factors[24-28]. Consistent with important roles in infection, V proteins show high conservation in the unique C-terminal region (Figure 2)[29-31], including absolute conservation of seven conserved cysteine residues and a histidine, which form a zinc-finger domain (highlighted in Figure 2). In the parainfluenza virus 5 (PIV5) V protein (Rubulavirus genus), two zinc atoms are coordinated by two loops, incorporating V residues H171, C190, C215, C218, and C194, C206, C208, C211 respectively[31,32], and mutations of these residues disrupt certain IFN inhibitory functions (see below), although the role of zinc-binding is not known. The C-terminal region is also important to the formation of oligomeric structures of V proteins and certain innate immune factors important to IFN antagonism[33].
Mammalian cell responses to infection depend on the detection of pathogen-associated molecular patterns (PAMPs) produced during microbial infection and replication, such as single-stranded RNA (ssRNA), double-stranded RNA (dsRNA) and RNA with exposed/uncapped 5’ triphosphates that are generated by RNA viruses[34,35]. Detection of virus components is principally mediated by three types of PAMP-recognition-receptors (PRRs): Toll-like receptors (TLRs) and RIG-I-like receptors (RLRs), thought to be the main receptors responsible for type I IFN (IFNα/β) induction, and nucleotide-oligomerisation domain-like receptors[36].
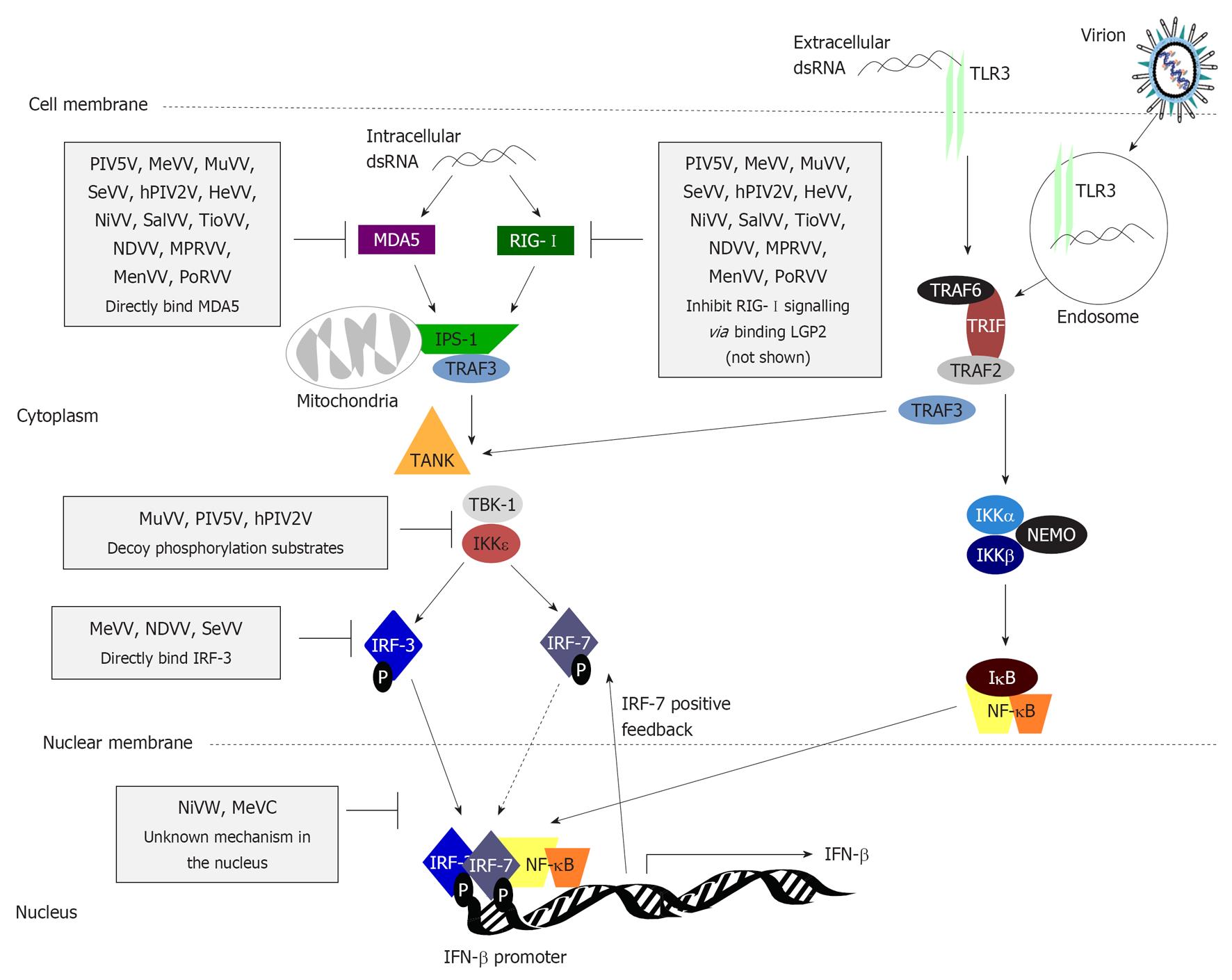
TLRs are trans-membrane proteins expressed at the plasma membrane or on intracellular structures such as endosomes and the endoplasmic reticulum[37,38] to detect extracellular viral nucleic acids such as dsRNA (TLR3)[37-40] and G/U-rich ssRNAs (TLR7)[38]. By contrast, the almost ubiquitously expressed RLR helicases RIG-I and MDA5 detect viral dsRNA in the cytoplasm of infected cells[36,41-47]; RIG-I also recognises cytoplasmic 5’ tri-phosphorylated and uncapped viral ssRNA[48-50]. RNA-activated MDA5 and RIG-I interact with the mitochondrial membrane-associated adaptor protein IFNβ promoter stimulator 1 (IPS-1, also known as MAVS, VISA, or CARDIF) via their caspase activation and recruitment domains (CARDs) to trigger downstream signalling (Figure 3). TLRs activate distinct pathways (Figure 3), but RLR and TLR signalling converges with the phosphorylation of the constitutively expressed cytoplasmic transcription factors IRF-3, as well as nuclear factor κB (NF-κB), causing their translocation into the nucleus to activate the transcription of early type I IFNs (IFNβ and IFNα4)[36,51-56]. Most human cell types can produce type I IFNs in response to infection, with “professional” IFN-producing immune cells including plasmacytoid DCs and macrophages being major producers during infection, due to constitutive expression of IRF-7 (which requires induction in other cell types) and the use of alternative TLR-9 pathways[57]. Importantly, paramyxoviruses can induce type I IFN expression through RIG-I, MDA5 and TLR pathways (Figure 3)[42,58,59].
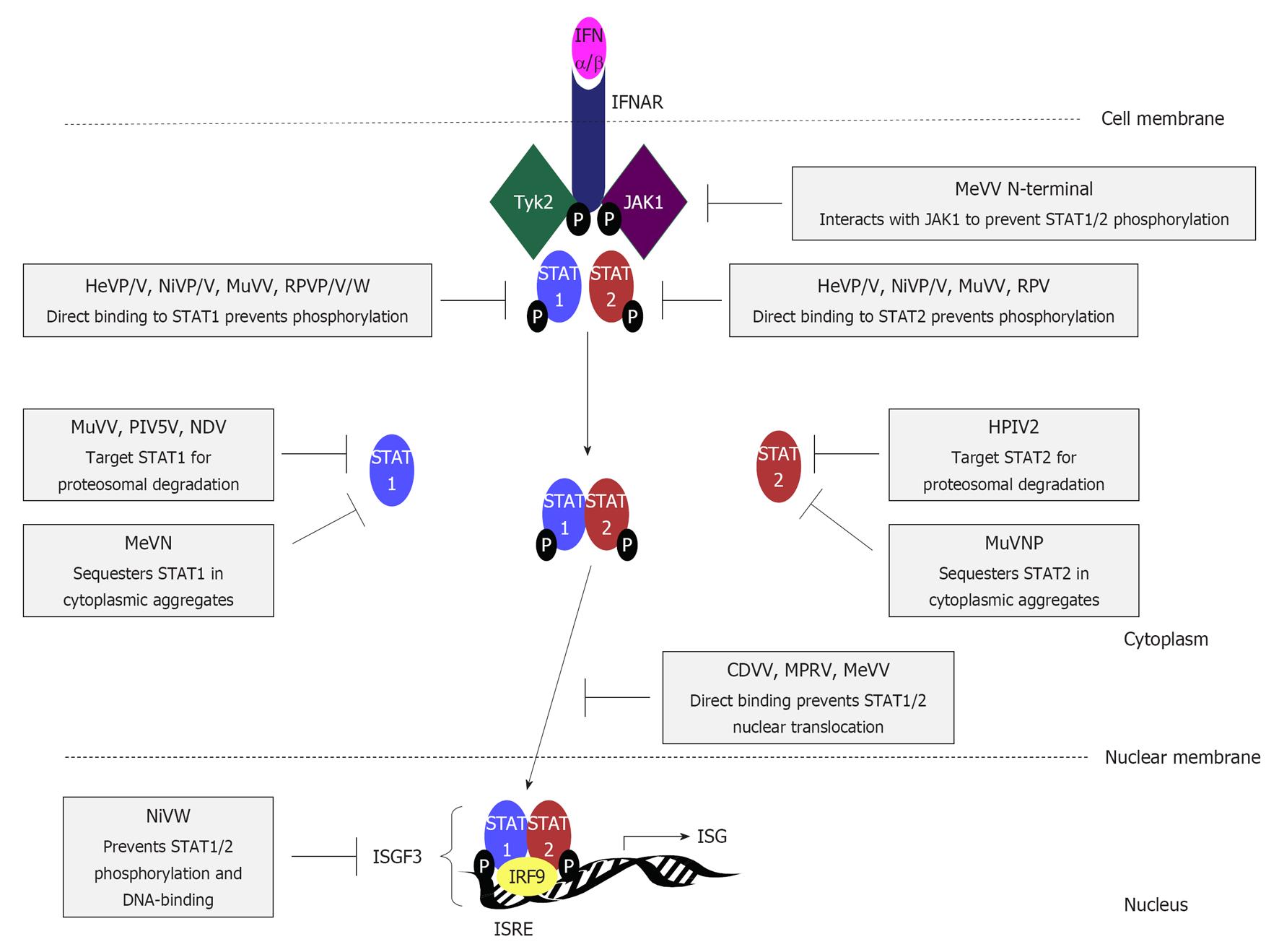
Type I IFNs signal in autocrine and paracrine fashion, binding to the ubiquitously expressed IFNα/β receptor (IFNAR) to activate the Janus kinase (JAK)/STAT signalling pathway (Figure 4), resulting in the phosphorylation and nuclear translocation of STAT1 and STAT2 proteins. In the form of a heterotrimeric complex [IFN-stimulated gene factor 3 (ISGF3)] which incorporates IRF-9, STAT1 and STAT2 trans-activate hundreds of IFN-stimulated genes, many of which encode known antiviral proteins including protein kinase R, which inhibits translation of mRNAs[60]; 2’5’-oligoadenylate synthetase, which activates RNase L to effect degradation of ssRNA[60]; Mx GTPase proteins that interfere with the growth of certain viruses including the paramyxoviruses[52]; and PML, which has antiviral properties but with unresolved mechanisms[52]. IRF-7 is also up regulated to activate a positive feedback loop by forming heterodimers with IRF-3 (Figure 3) inducing “late” IFNα subtypes for prolonged responses to infection[61].
Although signalling through STAT1/2 heterodimers is essential to type I IFN responses, type I IFN activates other complexes including homodimers of STAT1 and STAT3 and STAT1-STAT3 heterodimers, which have different gene specificity or regulatory roles[62], and recent data suggest that STAT2 can also effect STAT1-independent antiviral functions[63]. Thus, type I IFN activation can affect diverse gene expression through distinct pathways. STATs are also critical to signalling by type II and III IFNs and various other cytokines[54,64] such as interleukin (IL)-6[65,66], presenting potential targets for viral inhibition of several immune signalling systems.
A large body of evidence indicates that viruses/IFN-antagonist proteins generally target multiple steps in the IFN system[52,67,68]. The requirement for this broad targeting probably relates to factors such as differences in the kinetics of viral IFN-antagonist expression compared with the mounting of IFN responses, the contribution of infected cells and non-infected professional IFN producing cells, and the overall antiviral potency of the IFN system. Most paramyxoviruses can inhibit both IFN induction and signalling by targeting several cellular proteins. Intriguingly, although paramyxoviruses generally target common factors including MDA5, IRF-3 and STATs, the mechanisms of inhibition show significant divergence between different viruses.
The V proteins of at least 13 paramyxoviruses tested bind to MDA5 to inhibit IFN induction[32,69-71]. Rinderpest virus (RPV) may differ, as it appears to use the C protein rather than V to inhibit MDA5 signalling, although the binding of RPV V to MDA5 has not been examined[72]. The V proteins of PIV5, hPIV2, MuV, MeV, NiV, HeV, SeV, Mapuera virus (MPRV), Menangle virus (MenV) and Salem virus (SalV) were shown to bind a specific region within/proximal to residues 701-816 of the MDA5 helicase domain, independently of the MDA5 ligand dsRNA[70,71,73], thereby blocking dsRNA-MDA5 interaction[70]. Although a recent study of PIV5 V identified a change in the dsRNA-binding properties of MDA5 when V was co-expressed, rather than a complete inhibition, suggesting that V may still allow non-cooperative dsRNA binding[74]. In addition, the V proteins of PIV5, MenV, and SalV might have further specialist antagonistic functions, as yeast two-hybrid assays indicated that they interacted with multiple distinct regions of MDA5[70]. A crystal structure of PIV5 V in complex with MDA5 has demonstrated that V unfolds the ATPase domain of MDA5, which allows it to bind a region normally hidden beneath the helicase fold[74]. This unfolding disturbs the ATPase hydrolysis site, and it was shown using MeV V that increasing concentrations of V correlate with decreasing ATPase activity[74].
The MDA5 binding site has been mapped to the C-terminal region of the V proteins of PIV5, MeV, MuV, Newcastle disease virus (NDV), NiV, HeV and SeV[32,69-71,75,76], with conserved residues of the zinc-finger critical to the interaction. However, the precise molecular details differ between specific paramyxoviruses, with conserved cysteine residues in the large zinc finger loop, but not the smaller loop, of PIV5 V and NiV V dispensable for antagonism of IFN induction[32], whereas MuV V and MeV V required all conserved cysteine residues[32]. A crystal structure of MDA5:PIV5 has shown PIV5 V to have six residues (174, 175, 177, 179, 184 and 197) involved in forming the interface with MDA5, only some of which are conserved with other paramyxovirus V proteins[74].
In contrast to MDA5, V proteins do not bind directly to RIG-I, nor inhibit RIG-I oligomersation or dsRNA-binding[70], which has been assumed to indicate that they have no direct role in inhibiting RIG-I activation, but rather target downstream signalling components such as IRF-3 (see below). However, recent data has indicated that V proteins can inhibit RIG-I by interaction with another cellular helicase, the laboratory of genetics and physiology 2 (LGP2)[73], via a region of LGP2 homologous to the V protein binding region in MDA5[71,73]. The interaction appears to be dependent on the unique C-terminal domain of V protein, as PIV5 P protein did not bind to LGP2, but the C-terminal domains of MeV and MuV V proteins were necessary and sufficient for the interaction[71,73]. Importantly, V proteins were able to inhibit RIG-I signalling only in cells where LGP2 was coexpressed[73], and RIG-I-LGP2 interaction was detected only in cells expressing V protein, suggesting that V facilitates or mediates this interaction to shutdown RIG-I activation[73]. Because LGP2 is homologous to RIG-I and MDA5, but lacks the CARD domain to activate downstream signalling, it is thought to be a negative regulator of IFN induction, consistent with the inhibitory effects of V protein expression. However, there is evidence that LGP2 can positively regulate IFN induction under some conditions[77-80], so the precise mechanisms of V protein/LGP2 antagonism of RIG-I remain to be determined.
In addition to inhibition of PRRs, paramyxoviruses target downstream signalling components to prevent activation of IRF-3, potentially as a mechanism to inhibit signalling by both RLRs and TLRs (Figure 3). Rubulaviruses including MuV, hPIV2, and PIV5 use V protein as a decoy substrate for the IRF-3 kinases TANK-binding kinase 1 (TBK-1) and inhibitor of NF-κB kinase (IKK)ε (Figure 3), both inhibiting phosphorylation of IRF-3 and facilitating IKKε/TBK-1 polyubiquitination and degradation to prevent further signalling[81].
Henipavirus V proteins do not cause IKKε/TBK-1 degradation[81] or block TLR-3/IRF-3 dependent signalling[76,81]. For henipaviruses, this appears to be a function of the W protein, as NiV W, although having no effect on MDA5 signalling, inhibited TLR-3-dependent phosphorylation of IRF-3[82]. It is possible this is due to binding and sequestration of inactive IRF-3 in the nucleus where NiV W localises, to prevent interaction with cytoplasmic IKKε/TBK-1[82]. This model is consistent with the reported importance of NiV W protein nuclear localisation to its inhibition of TLR3-dependent IFN induction[82]. MeV C protein also inhibits IFN induction, correlating with its nuclear localisation[83], although MeV C does not affect IRF-3 directly, and appears to have an undetermined nuclear target[83]. By contrast, cytoplasmic NDV and SeV V protein bind directly to IRF-3, thereby preventing its nuclear translocation[76]. Thus, paramyxovirus targeting of IRF-3-mediated signalling involves mechanisms that appear to differ significantly between species.
Almost all rubulavirus V proteins target STAT1 or STAT2 for degradation by the host-cell proteosomal pathways[84-87] through assembly of a V-degradation complex (VDC) containing V protein, STAT1, STAT2, and components of an E3 ubiquitin ligase complex, specifically the UV damage-specific DNA binding protein 1 (DDB1), and Cul4A[88-92], which likely mediate the STAT1/2 polyubiquitination[93]. In vitro studies/crystallographic analysis of the PIV5 V-DBB1 complex have indicated that both the N-terminal and unique C-terminal regions of PIV5 V are required for VDC assembly and STAT1 degradation[33,88,93,94]. Intriguingly, although some rubulaviruses target only STAT1 or STAT2 for degradation (see Figure 4 for details)[95,96], both STATs are required, with the non-degraded STAT acting as a “co-factor”[97,98].
The MuV V protein VDC polyubiquitinates and degrades not only STAT1, but also STAT3[84,99], such that MuV V protein can inhibit STAT3-dependent transcriptional activation by IL-6 and v-Src[99]. MuV targeting of STAT3 is independent of STAT1 targeting, as a point mutation abrogating targeting of STAT3 did not affect STAT1[100], and STAT3 degradation does not require the STAT2 “cofactor”[99]. STAT3 targeting by the V protein of MuV is also highly specific to this species, as the V proteins of the rubulaviruses MPRV, hPIV2 and hPIV4 do not reduce cellular levels of STAT3[87,101,102].
Intriguingly, the V proteins of hPIV4a and hPIV4b do not degrade STATs or measurably affect their localisation or phosphorylation, but still bind to STAT1, STAT2 and other VDC components[101]. While these viruses appear to lack the ability to antagonise STAT signalling, the specific binding capacity of the proteins is suggestive of a previous role in STAT antagonism, which may have been lost due to changes in selective pressures[101].
MPRV V protein, by contrast with those of other rubulaviruses, binds to STAT1 and STAT2 to prevent their nuclear translocation without inducing degradation[102]. This is similar to reports for the V proteins of the henipaviruses and morbilliviruses (see below), except in that MPRV V does not inhibit STAT1 phosphorylation and can bind to STAT1 and STAT2 independently[102]. A similar mechanism may be used by the MuV NP protein, which co-localises with STAT2 in punctate aggregates in the cytoplasm of infected cells[99], indicating that NP protein, like P protein, can mediate both replication and IFN antagonist functions.
In common with rubulaviruses such as PIV5, the avulavirus NDV targets STAT1, but not STAT2, for degradation. Deletion of the C-terminal region of V protein, or deletion of both V and W C-termini by disruption of the RNA editing site, prevented STAT1 degradation by recombinant NDV[95]. As little difference was observed between virus deleted for both V and W, and virus deleted for the V protein C-terminal domain alone, V protein appears to be the major player, and consistent with this, NDV V but not NDV W degraded STAT1 in transfected cells[95].
MeV V binds STAT1 and STAT2 through distinct sites in its N-terminal and C-terminal regions[103], respectively, indicating that targeting of STAT2 independently of STAT1 is important to this virus. MeV V protein does not degrade STATs[104], but has been reported by different laboratories to use several distinct mechanisms, including inhibition of STAT nuclear translocation without affecting STAT phosphorylation[103-105], and inhibition of STAT1 and STAT2 phosphorylation due to interaction of its N-terminal domain with JAK1[106,107]. Canine distemper virus (CDV) and RPV V proteins also inhibit IFN-activated STAT1/STAT2 nuclear import[108,109], with RPV V protein, but not that of CDV, inhibiting STAT1/2 phosphorylation.
MeV V also interacts with IRF-9, which is likely to affect ISGF3 formation (Figure 4)[104], and with STAT3[104], a property thus far restricted in the paramyxovirus family to MeV and MuV V proteins[99,100,104,110]. HeV V and PIV5 V have been shown to lack STAT3 binding function, and while SeV infection can inhibit IFNα-dependent STAT3 phosphorylation, this appears to relate to upstream effects on Tyk2 rather than STAT3 directly[111]. STAT3 binding by other paramyxovirus V proteins, however, has not been investigated.
MeV N protein also inhibits STAT1/2 signalling[112], indicating a particular importance of STAT inhibition to MeV, and co-localises with STAT1 in cytoplasmic aggregates in infected cells, analogously to MuV NP[99,104]. STAT2 also co-localised with MeV N in aggregates, but with reduced frequency compared with STAT1[104].
STAT targeting by respiroviruses differs significantly from other paramyxoviruses, due to the expression of additional proteins from the P gene (Figure 2), including four C proteins by hPIV1[14,113], which does not express V or W. The C’ protein of hPIV1 binds and sequesters STAT1 in perinuclear aggregates, suggesting that the C proteins may be sufficient for IFN antagonism by this virus[114]. SeV C proteins (C’, C, Y1 and Y2), also bind to STAT1 and prevent signalling and, importantly, the functions of the individual C proteins appear non-redundant, as knockout of all four proteins is required to completely prevent IFN antagonism in infected cells[115,116]. Data regarding the mechanisms of SeV C proteins activity are conflicting[116-121], with some reports suggesting that C and C’, but not Y1 or Y2, cause STAT1 mono-ubiquitination/degradation[116,117] dependent on the C protein N-termini[118,119], while others reported no reduction in STAT1 expression but indicated inhibition of STAT1 and STAT2 phosphorylation by the C proteins, independently of their N-termini[120,121].
The henipavirus P, V and W proteins can bind to STAT1 and STAT2 through the shared N-terminal region[122,123] to prevent STAT1/2 phosphorylation and activation by holding them in high molecular weight complexes[110,123-125]. Transfection studies indicate that P, V and W have differing capacities to inhibit STAT signalling, with P protein the least effective[125]. This is consistent with the hypothesis that the V and W accessory proteins have evolved to enable specific, distinct roles as IFN-antagonists, sequestering STATs in the cytoplasm and the nucleus, respectively[82,122,125], whereas P protein functions principally as the polymerase cofactor, but can arrest STATs in the cytoplasm. Mutation of the shared G121 residue was found to specifically ablate STAT1 binding by V, W and P, without affecting P protein polymerase co-factor function, enabling the production of recombinant NiV impaired for STAT antagonistic functions to confirm that inhibition of STAT1 phosphorylation in NiV infected cells is due to P/V/W binding[122,126]. In wild-type NiV-infected cells, but not those infected with the mutant NiV, unphosphorylated STAT1 localised exclusively to the nucleus, similar to cells expressing W protein alone, suggesting NiV W has the predominant role in blocking STAT signalling in infected cells[122].
Although there is abundant evidence that paramyxovirus P gene-encoded proteins can antagonise IFN responses by diverse species-specific/genera-specific mechanisms, the source of this diversity is currently unclear. A major caveat of the available data is its heavy reliance on in vitro studies, particularly transfection studies of single IFN-antagonist proteins. Although these approaches enable highly specific analyses of the properties of particular IFN-antagonists, including mapping/mutagenesis studies, the potential to generate artefactual data due to the absence of other viral factors and/or non-physiological expression levels is a significant concern. Indeed, several transfection studies in different laboratories have generated conflicting mechanistic data for the same viral protein, including SeV C and MeV V protein[103-107,116,117,120,121], suggesting that some reported differences between IFN antagonists of different paramyxovirus species/genera might arise from experimental rather than biological differences. Importantly, however, recent studies comparing in parallel the functions of V proteins from panels of paramyxoviruses have confirmed clear divergence in specific mechanisms/interactions[70,71,76], indicating genuine divergence at the molecular level.
Recent studies have also directly compared IFN-antagonist protein expression/functions in transfected and infected cells, identifying clear differences. Notably, one study reported that while henipavirus V and W proteins profoundly inhibit IFN/STAT signalling in transfected cells, no inhibition was apparent in infected cells, which appeared to relate to the higher expression of V/W proteins in transfected cells[127]. This suggested that STAT inhibition by V and W does not have significant roles in infected cells, but it seems unlikely that viruses would evolve proteins that can specifically target factors of the IFN response and impede their function by sophisticated mechanisms were this not important at some stage of infection. While in vitro infection approaches are clearly closer to natural infection than transfection, they also use controlled in vitro conditions including the inoculation of cultured monolayers of specific cell types with precise multiplicities of infection, and treatments with specific concentrations of IFNs. By contrast, in natural infection the kinetics of viral protein expression and induction of the IFN system is highly dynamic, involving both infected cells and professional IFN-producing cells, and factors such as the infectious dose, route of infection, host species, and infectious spread to specific tissues can vary greatly, significantly affecting requirements for IFN antagonism and the disease outcome[128]. Thus, the diverse mechanisms of IFN antagonism identified in transfection studies may have vital roles in infection in vivo.
Importantly, IFN antagonism has been implicated as a key factor in host and tissue specificity, with PIV5 showing limited host range dependent on the capacity of the V protein to bind to STAT2 from different species[129-132], whereas NiV V blocks IFN signalling in cells of many species, consistent with its broad infectious range[82,110,124,126]. Tissue-specific antagonism of IFN has also been reported for NiV, which induces an IFN response in endothelial but not neuronal cells, correlating with differential subcellular localisation of NiV W[133].
A genuine appreciation of the importance of specific IFN-antagonistic mechanisms to pathogenicity, however, requires the use of recombinant virus systems and in vivo pathogenicity models. Recent advances in this area include reports that recombinant hPIV2 impaired for V protein antagonism of MDA5 is attenuated in rhesus monkeys[134-136], and that the severity of clinical signs in MeV-infected monkeys was reduced by mutation of the P/V proteins to prevent inhibition of STAT1[137]. In addition, the deletion of V or C proteins from MeV caused attenuation in mice, but V deletion alone resulted in restricted spread in the brain[138], supporting the hypothesis that specific mechanisms of IFN-antagonism are important to infection of certain tissues. Deletion of the V C-terminal domain in recombinant NiV also reduced pathogenicity in a hamster model[123-125,139], possibly due to IFN-antagonist functions of the V C-terminal domain, such as the targeting of MDA5.
Of paramount importance to delineating the roles of specific mechanisms of IFN antagonism in pathogenicity will be the extension of in vivo studies to include genetically modified animals deficient in specific IFN signalling processes. For example, recent research indicated that SeV pathogenicity is increased in MDA5 knockout mice[140], suggesting that this might provide a useful model to investigate the importance of MDA5 antagonism in in vivo infection.
A substantial body of data from the past c. 15 years has provided key insights into the immune evasion strategies of paramyxovirus IFN-antagonists, indicating that they employ a remarkable array of mechanisms to target essential factors of the IFN response, with the limited in vivo infection data indicating that these functions are essential to pathogenicity. However, as much of the current mechanistic data comes from in vitro transfection approaches, their importance to natural infection remains largely unresolved. Future studies employing in vivo infection models, recombinant virus systems and genetically modified animals should begin to unravel in detail the interactions of paramyxoviruses with the IFN system in vivo. This is likely to result in the identification of new potential targets for the development of vaccines and antivirals required for the treatment of established prolific human pathogens such as MeV, as well as emerging zoonotic threats including NiV and HeV.
P- Reviewer Guo HT S- Editor Song XX L- Editor A E- Editor Zheng XM
| 1. | World Health Organization. Measles: Fact sheet N¡ã286. 2012; Available from: http://www.who.int/mediacentre/factsheets/fs286/en/index.html. |
| 2. | Hossain MJ, Gurley ES, Montgomery JM, Bell M, Carroll DS, Hsu VP, Formenty P, Croisier A, Bertherat E, Faiz MA. Clinical presentation of nipah virus infection in Bangladesh. Clin Infect Dis. 2008;46:977-984. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 165] [Cited by in RCA: 179] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 3. | Ali R, Mounts AW, Parashar UD, Sahani M, Lye MS, Isa MM, Balathevan K, Arif MT, Ksiazek TG. Nipah virus among military personnel involved in pig culling during an outbreak of encephalitis in Malaysia, 1998-1999. Emerg Infect Dis. 2001;7:759-761. [PubMed] |
| 4. | Marsh GA, Todd S, Foord A, Hansson E, Davies K, Wright L, Morrissy C, Halpin K, Middleton D, Field HE. Genome sequence conservation of Hendra virus isolates during spillover to horses, Australia. Emerg Infect Dis. 2010;16:1767-1769. [PubMed] |
| 5. | Chua KB, Bellini WJ, Rota PA, Harcourt BH, Tamin A, Lam SK, Ksiazek TG, Rollin PE, Zaki SR, Shieh W. Nipah virus: a recently emergent deadly paramyxovirus. Science. 2000;288:1432-1435. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 878] [Cited by in RCA: 889] [Article Influence: 35.6] [Reference Citation Analysis (0)] |
| 6. | Lamb RA, Parks GD. Paramyxoviridae: the viruses and their replication. Fields virology. 5th ed. Philadelphia: Lippincott Williams and Wilkins 2006; 1449-1496. |
| 7. | King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ. Virus taxonomy: classification and nomenclature of viruses: Ninth Report of the International Committee on Taxonomy of Viruses. San Diego: Elsevier Academic Press 2011; . |
| 8. | Chenik M, Chebli K, Blondel D. Translation initiation at alternate in-frame AUG codons in the rabies virus phosphoprotein mRNA is mediated by a ribosomal leaky scanning mechanism. J Virol. 1995;69:707-712. [PubMed] |
| 9. | Paterson RG, Lamb RA. RNA editing by G-nucleotide insertion in mumps virus P-gene mRNA transcripts. J Virol. 1990;64:4137-4145. [PubMed] |
| 10. | Jacques JP, Hausmann S, Kolakofsky D. Paramyxovirus mRNA editing leads to G deletions as well as insertions. EMBO J. 1994;13:5496-5503. [PubMed] |
| 11. | Vidal S, Curran J, Kolakofsky D. Editing of the Sendai virus P/C mRNA by G insertion occurs during mRNA synthesis via a virus-encoded activity. J Virol. 1990;64:239-246. [PubMed] |
| 12. | Vidal S, Curran J, Kolakofsky D. A stuttering model for paramyxovirus P mRNA editing. EMBO J. 1990;9:2017-2022. [PubMed] |
| 13. | Sanchez A, Trappier SG, Mahy BW, Peters CJ, Nichol ST. The virion glycoproteins of Ebola viruses are encoded in two reading frames and are expressed through transcriptional editing. Proc Natl Acad Sci USA. 1996;93:3602-3607. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 422] [Cited by in RCA: 421] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 14. | Matsuoka Y, Curran J, Pelet T, Kolakofsky D, Ray R, Compans RW. The P gene of human parainfluenza virus type 1 encodes P and C proteins but not a cysteine-rich V protein. J Virol. 1991;65:3406-3410. [PubMed] |
| 15. | Marsh GA, de Jong C, Barr JA, Tachedjian M, Smith C, Middleton D, Yu M, Todd S, Foord AJ, Haring V. Cedar virus: a novel Henipavirus isolated from Australian bats. PLoS Pathog. 2012;8:e1002836. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 197] [Cited by in RCA: 240] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 16. | Fontana JM, Bankamp B, Rota PA. Inhibition of interferon induction and signaling by paramyxoviruses. Immunol Rev. 2008;225:46-67. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 83] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 17. | Hausmann S, Jacques JP, Kolakofsky D. Paramyxovirus RNA editing and the requirement for hexamer genome length. RNA. 1996;2:1033-1045. [PubMed] |
| 18. | Kulkarni S, Volchkova V, Basler CF, Palese P, Volchkov VE, Shaw ML. Nipah virus edits its P gene at high frequency to express the V and W proteins. J Virol. 2009;83:3982-3987. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 73] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 19. | Lo MK, Harcourt BH, Mungall BA, Tamin A, Peeples ME, Bellini WJ, Rota PA. Determination of the henipavirus phosphoprotein gene mRNA editing frequencies and detection of the C, V and W proteins of Nipah virus in virus-infected cells. J Gen Virol. 2009;90:398-404. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 61] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 20. | Yu X, Tsibane T, McGraw PA, House FS, Keefer CJ, Hicar MD, Tumpey TM, Pappas C, Perrone LA, Martinez O. Neutralizing antibodies derived from the B cells of 1918 influenza pandemic survivors. Nature. 2008;455:532-536. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 355] [Cited by in RCA: 328] [Article Influence: 19.3] [Reference Citation Analysis (0)] |
| 21. | Wang LF, Michalski WP, Yu M, Pritchard LI, Crameri G, Shiell B, Eaton BT. A novel P/V/C gene in a new member of the Paramyxoviridae family, which causes lethal infection in humans, horses, and other animals. J Virol. 1998;72:1482-1490. [PubMed] |
| 22. | Curran J, Kolakofsky D. Scanning independent ribosomal initiation of the Sendai virus X protein. EMBO J. 1988;7:2869-2874. [PubMed] |
| 23. | Curran J, Boeck R, Kolakofsky D. The Sendai virus P gene expresses both an essential protein and an inhibitor of RNA synthesis by shuffling modules via mRNA editing. EMBO J. 1991;10:3079-3085. [PubMed] |
| 24. | Gotoh B, Komatsu T, Takeuchi K, Yokoo J. Paramyxovirus accessory proteins as interferon antagonists. Microbiol Immunol. 2001;45:787-800. [PubMed] |
| 25. | Gotoh B, Komatsu T, Takeuchi K, Yokoo J. Paramyxovirus strategies for evading the interferon response. Rev Med Virol. 2002;12:337-357. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 103] [Cited by in RCA: 106] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 26. | Goodbourn S, Randall RE. The regulation of type I interferon production by paramyxoviruses. J Interferon Cytokine Res. 2009;29:539-547. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 72] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 27. | Horvath CM. Weapons of STAT destruction. Interferon evasion by paramyxovirus V protein. Eur J Biochem. 2004;271:4621-4628. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 133] [Cited by in RCA: 132] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 28. | Horvath CM. Silencing STATs: lessons from paramyxovirus interferon evasion. Cytokine Growth Factor Rev. 2004;15:117-127. [PubMed] |
| 29. | Huang C, Kiyotani K, Fujii Y, Fukuhara N, Kato A, Nagai Y, Yoshida T, Sakaguchi T. Involvement of the zinc-binding capacity of Sendai virus V protein in viral pathogenesis. J Virol. 2000;74:7834-7841. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 71] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 30. | Liston P, Briedis DJ. Measles virus V protein binds zinc. Virology. 1994;198:399-404. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 79] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 31. | Paterson RG, Leser GP, Shaughnessy MA, Lamb RA. The paramyxovirus SV5 V protein binds two atoms of zinc and is a structural component of virions. Virology. 1995;208:121-131. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 121] [Cited by in RCA: 120] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 32. | Ramachandran A, Horvath CM. Dissociation of paramyxovirus interferon evasion activities: universal and virus-specific requirements for conserved V protein amino acids in MDA5 interference. J Virol. 2010;84:11152-11163. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 53] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 33. | Ulane CM, Kentsis A, Cruz CD, Parisien JP, Schneider KL, Horvath CM. Composition and assembly of STAT-targeting ubiquitin ligase complexes: paramyxovirus V protein carboxyl terminus is an oligomerization domain. J Virol. 2005;79:10180-10189. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 93] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 34. | Akira S, Uematsu S, Takeuchi O. Pathogen recognition and innate immunity. Cell. 2006;124:783-801. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8061] [Cited by in RCA: 8811] [Article Influence: 463.7] [Reference Citation Analysis (0)] |
| 35. | Medzhitov R. Recognition of microorganisms and activation of the immune response. Nature. 2007;449:819-826. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1864] [Cited by in RCA: 1940] [Article Influence: 107.8] [Reference Citation Analysis (0)] |
| 36. | Takeuchi O, Akira S. Innate immunity to virus infection. Immunol Rev. 2009;227:75-86. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 925] [Cited by in RCA: 953] [Article Influence: 59.6] [Reference Citation Analysis (0)] |
| 37. | Matsumoto M, Oshiumi H, Seya T. Antiviral responses induced by the TLR3 pathway. Rev Med Virol. 2011;Epub ahead of print. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 134] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 38. | Carty M, Bowie AG. Recent insights into the role of Toll-like receptors in viral infection. Clin Exp Immunol. 2010;161:397-406. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 82] [Cited by in RCA: 107] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 39. | Alexopoulou L, Holt AC, Medzhitov R, Flavell RA. Recognition of double-stranded RNA and activation of NF-kappaB by Toll-like receptor 3. Nature. 2001;413:732-738. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4510] [Cited by in RCA: 4642] [Article Influence: 193.4] [Reference Citation Analysis (0)] |
| 40. | Johnsen IB, Nguyen TT, Ringdal M, Tryggestad AM, Bakke O, Lien E, Espevik T, Anthonsen MW. Toll-like receptor 3 associates with c-Src tyrosine kinase on endosomes to initiate antiviral signaling. EMBO J. 2006;25:3335-3346. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 150] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 41. | Takeuchi O, Akira S. MDA5/RIG-I and virus recognition. Curr Opin Immunol. 2008;20:17-22. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 427] [Cited by in RCA: 439] [Article Influence: 25.8] [Reference Citation Analysis (0)] |
| 42. | Yoneyama M, Kikuchi M, Natsukawa T, Shinobu N, Imaizumi T, Miyagishi M, Taira K, Akira S, Fujita T. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat Immunol. 2004;5:730-737. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2934] [Cited by in RCA: 3122] [Article Influence: 148.7] [Reference Citation Analysis (0)] |
| 43. | Kato H, Sato S, Yoneyama M, Yamamoto M, Uematsu S, Matsui K, Tsujimura T, Takeda K, Fujita T, Takeuchi O. Cell type-specific involvement of RIG-I in antiviral response. Immunity. 2005;23:19-28. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1024] [Cited by in RCA: 1059] [Article Influence: 53.0] [Reference Citation Analysis (0)] |
| 44. | Luthra P, Sun D, Silverman RH, He B. Activation of IFN-& amp; #946; expression by a viral mRNA through RNase L and MDA5. Proc Natl Acad Sci USA. 2011;108:2118-2123. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 75] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 45. | Loo YM, Fornek J, Crochet N, Bajwa G, Perwitasari O, Martinez-Sobrido L, Akira S, Gill MA, García-Sastre A, Katze MG. Distinct RIG-I and MDA5 signaling by RNA viruses in innate immunity. J Virol. 2008;82:335-345. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 786] [Cited by in RCA: 830] [Article Influence: 46.1] [Reference Citation Analysis (0)] |
| 46. | Baum A, Sachidanandam R, García-Sastre A. Preference of RIG-I for short viral RNA molecules in infected cells revealed by next-generation sequencing. Proc Natl Acad Sci USA. 2010;107:16303-16308. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 332] [Cited by in RCA: 348] [Article Influence: 23.2] [Reference Citation Analysis (0)] |
| 47. | Kato H, Takeuchi O, Mikamo-Satoh E, Hirai R, Kawai T, Matsushita K, Hiiragi A, Dermody TS, Fujita T, Akira S. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J Exp Med. 2008;205:1601-1610. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1112] [Cited by in RCA: 1245] [Article Influence: 73.2] [Reference Citation Analysis (0)] |
| 48. | Pichlmair A, Schulz O, Tan CP, Näslund TI, Liljeström P, Weber F, Reis e Sousa C. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5’-phosphates. Science. 2006;314:997-1001. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1735] [Cited by in RCA: 1736] [Article Influence: 91.4] [Reference Citation Analysis (0)] |
| 49. | Habjan M, Andersson I, Klingström J, Schümann M, Martin A, Zimmermann P, Wagner V, Pichlmair A, Schneider U, Mühlberger E. Processing of genome 5’ termini as a strategy of negative-strand RNA viruses to avoid RIG-I-dependent interferon induction. PLoS One. 2008;3:e2032. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 244] [Cited by in RCA: 249] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 50. | Nallagatla SR, Toroney R, Bevilacqua PC. A brilliant disguise for self RNA: 5’-end and internal modifications of primary transcripts suppress elements of innate immunity. RNA Biol. 2008;5:140-144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 61] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 51. | Kawai T, Akira S. Antiviral signaling through pattern recognition receptors. J Biochem. 2007;141:137-145. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 320] [Cited by in RCA: 343] [Article Influence: 18.1] [Reference Citation Analysis (0)] |
| 52. | Randall RE, Goodbourn S. Interferons and viruses: an interplay between induction, signalling, antiviral responses and virus countermeasures. J Gen Virol. 2008;89:1-47. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1165] [Cited by in RCA: 1235] [Article Influence: 72.6] [Reference Citation Analysis (0)] |
| 53. | Takeuchi O, Akira S. Recognition of viruses by innate immunity. Immunol Rev. 2007;220:214-224. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 256] [Cited by in RCA: 262] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 54. | Baum A, García-Sastre A. Induction of type I interferon by RNA viruses: cellular receptors and their substrates. Amino Acids. 2010;38:1283-1299. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 101] [Cited by in RCA: 107] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 55. | Pestka S, Krause CD, Walter MR. Interferons, interferon-like cytokines, and their receptors. Immunol Rev. 2004;202:8-32. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1150] [Cited by in RCA: 1235] [Article Influence: 61.8] [Reference Citation Analysis (0)] |
| 56. | Díaz MO, Testa D. Type I interferon genes and proteins. Biotherapy. 1996;8:157-162. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 3] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 57. | Tailor P, Tamura T, Ozato K. IRF family proteins and type I interferon induction in dendritic cells. Cell Res. 2006;16:134-140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 124] [Cited by in RCA: 142] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 58. | Kato H, Takeuchi O, Sato S, Yoneyama M, Yamamoto M, Matsui K, Uematsu S, Jung A, Kawai T, Ishii KJ. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature. 2006;441:101-105. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2766] [Cited by in RCA: 2971] [Article Influence: 156.4] [Reference Citation Analysis (0)] |
| 59. | Melchjorsen J, Jensen SB, Malmgaard L, Rasmussen SB, Weber F, Bowie AG, Matikainen S, Paludan SR. Activation of innate defense against a paramyxovirus is mediated by RIG-I and TLR7 and TLR8 in a cell-type-specific manner. J Virol. 2005;79:12944-12951. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 147] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 60. | Hovanessian AG. On the discovery of interferon-inducible, double-stranded RNA activated enzymes: the 2’-5’oligoadenylate synthetases and the protein kinase PKR. Cytokine Growth Factor Rev. 2007;18:351-361. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 103] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 61. | Marié I, Durbin JE, Levy DE. Differential viral induction of distinct interferon-alpha genes by positive feedback through interferon regulatory factor-7. EMBO J. 1998;17:6660-6669. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 833] [Cited by in RCA: 840] [Article Influence: 31.1] [Reference Citation Analysis (0)] |
| 62. | Ho HH, Ivashkiv LB. Role of STAT3 in type I interferon responses. Negative regulation of STAT1-dependent inflammatory gene activation. J Biol Chem. 2006;281:14111-14118. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 221] [Cited by in RCA: 256] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 63. | Perry ST, Buck MD, Lada SM, Schindler C, Shresta S. STAT2 mediates innate immunity to Dengue virus in the absence of STAT1 via the type I interferon receptor. PLoS Pathog. 2011;7:e1001297. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 108] [Cited by in RCA: 119] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 64. | Zhou Z, Hamming OJ, Ank N, Paludan SR, Nielsen AL, Hartmann R. Type III interferon (IFN) induces a type I IFN-like response in a restricted subset of cells through signaling pathways involving both the Jak-STAT pathway and the mitogen-activated protein kinases. J Virol. 2007;81:7749-7758. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 348] [Cited by in RCA: 383] [Article Influence: 21.3] [Reference Citation Analysis (0)] |
| 65. | Yu CL, Meyer DJ, Campbell GS, Larner AC, Carter-Su C, Schwartz J, Jove R. Enhanced DNA-binding activity of a Stat3-related protein in cells transformed by the Src oncoprotein. Science. 1995;269:81-83. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 731] [Cited by in RCA: 755] [Article Influence: 25.2] [Reference Citation Analysis (0)] |
| 66. | Zhong Z, Wen Z, Darnell JE. Stat3: a STAT family member activated by tyrosine phosphorylation in response to epidermal growth factor and interleukin-6. Science. 1994;264:95-98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1520] [Cited by in RCA: 1637] [Article Influence: 52.8] [Reference Citation Analysis (0)] |
| 67. | Oksayan S, Ito N, Moseley G, Blondel D. Subcellular trafficking in rhabdovirus infection and immune evasion: a novel target for therapeutics. Infect Disord Drug Targets. 2012;12:38-58. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 30] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 68. | Basler CF, Amarasinghe GK. Evasion of interferon responses by Ebola and Marburg viruses. J Interferon Cytokine Res. 2009;29:511-520. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 129] [Cited by in RCA: 120] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 69. | Childs K, Stock N, Ross C, Andrejeva J, Hilton L, Skinner M, Randall R, Goodbourn S. mda-5, but not RIG-I, is a common target for paramyxovirus V proteins. Virology. 2007;359:190-200. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 260] [Cited by in RCA: 249] [Article Influence: 13.8] [Reference Citation Analysis (0)] |
| 70. | Childs KS, Andrejeva J, Randall RE, Goodbourn S. Mechanism of mda-5 Inhibition by paramyxovirus V proteins. J Virol. 2009;83:1465-1473. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 112] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 71. | Parisien JP, Bamming D, Komuro A, Ramachandran A, Rodriguez JJ, Barber G, Wojahn RD, Horvath CM. A shared interface mediates paramyxovirus interference with antiviral RNA helicases MDA5 and LGP2. J Virol. 2009;83:7252-7260. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 101] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 72. | Boxer EL, Nanda SK, Baron MD. The rinderpest virus non-structural C protein blocks the induction of type 1 interferon. Virology. 2009;385:134-142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 73. | Childs K, Randall R, Goodbourn S. Paramyxovirus V proteins interact with the RNA Helicase LGP2 to inhibit RIG-I-dependent interferon induction. J Virol. 2012;86:3411-3421. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 108] [Cited by in RCA: 116] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 74. | Motz C, Schuhmann KM, Kirchhofer A, Moldt M, Witte G, Conzelmann KK, Hopfner KP. Paramyxovirus V proteins disrupt the fold of the RNA sensor MDA5 to inhibit antiviral signaling. Science. 2013;339:690-693. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 103] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 75. | Andrejeva J, Childs KS, Young DF, Carlos TS, Stock N, Goodbourn S, Randall RE. The V proteins of paramyxoviruses bind the IFN-inducible RNA helicase, mda-5, and inhibit its activation of the IFN-beta promoter. Proc Natl Acad Sci USA. 2004;101:17264-17269. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 788] [Cited by in RCA: 807] [Article Influence: 38.4] [Reference Citation Analysis (0)] |
| 76. | Irie T, Kiyotani K, Igarashi T, Yoshida A, Sakaguchi T. Inhibition of interferon regulatory factor 3 activation by paramyxovirus V protein. J Virol. 2012;86:7136-7145. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 47] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 77. | Satoh T, Kato H, Kumagai Y, Yoneyama M, Sato S, Matsushita K, Tsujimura T, Fujita T, Akira S, Takeuchi O. LGP2 is a positive regulator of RIG-I- and MDA5-mediated antiviral responses. Proc Natl Acad Sci USA. 2010;107:1512-1517. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 456] [Cited by in RCA: 504] [Article Influence: 33.6] [Reference Citation Analysis (0)] |
| 78. | Yoneyama M, Kikuchi M, Matsumoto K, Imaizumi T, Miyagishi M, Taira K, Foy E, Loo YM, Gale M, Akira S. Shared and unique functions of the DExD/H-box helicases RIG-I, MDA5, and LGP2 in antiviral innate immunity. J Immunol. 2005;175:2851-2858. [PubMed] |
| 79. | Saito T, Hirai R, Loo YM, Owen D, Johnson CL, Sinha SC, Akira S, Fujita T, Gale M. Regulation of innate antiviral defenses through a shared repressor domain in RIG-I and LGP2. Proc Natl Acad Sci USA. 2007;104:582-587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 565] [Cited by in RCA: 573] [Article Influence: 31.8] [Reference Citation Analysis (0)] |
| 80. | Rothenfusser S, Goutagny N, DiPerna G, Gong M, Monks BG, Schoenemeyer A, Yamamoto M, Akira S, Fitzgerald KA. The RNA helicase Lgp2 inhibits TLR-independent sensing of viral replication by retinoic acid-inducible gene-I. J Immunol. 2005;175:5260-5268. [PubMed] |
| 81. | Lu LL, Puri M, Horvath CM, Sen GC. Select paramyxoviral V proteins inhibit IRF3 activation by acting as alternative substrates for inhibitor of kappaB kinase epsilon (IKKe)/TBK1. J Biol Chem. 2008;283:14269-14276. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 84] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 82. | Shaw ML, Cardenas WB, Zamarin D, Palese P, Basler CF. Nuclear localization of the Nipah virus W protein allows for inhibition of both virus- and toll-like receptor 3-triggered signaling pathways. J Virol. 2005;79:6078-6088. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 157] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 83. | Sparrer KM, Pfaller CK, Conzelmann KK. Measles virus C protein interferes with Beta interferon transcription in the nucleus. J Virol. 2012;86:796-805. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 61] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 84. | Kubota T, Yokosawa N, Yokota S, Fujii N. C terminal CYS-RICH region of mumps virus structural V protein correlates with block of interferon alpha and gamma signal transduction pathway through decrease of STAT 1-alpha. Biochem Biophys Res Commun. 2001;283:255-259. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 114] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 85. | Kubota T, Yokosawa N, Yokota S, Fujii N, Tashiro M, Kato A. Mumps virus V protein antagonizes interferon without the complete degradation of STAT1. J Virol. 2005;79:4451-4459. [PubMed] |
| 86. | Nishio M, Garcin D, Simonet V, Kolakofsky D. The carboxyl segment of the mumps virus V protein associates with Stat proteins in vitro via a tryptophan-rich motif. Virology. 2002;300:92-99. [PubMed] |
| 87. | Yokosawa N, Yokota S, Kubota T, Fujii N. C-terminal region of STAT-1alpha is not necessary for its ubiquitination and degradation caused by mumps virus V protein. J Virol. 2002;76:12683-12690. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 61] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 88. | Andrejeva J, Poole E, Young DF, Goodbourn S, Randall RE. The p127 subunit (DDB1) of the UV-DNA damage repair binding protein is essential for the targeted degradation of STAT1 by the V protein of the paramyxovirus simian virus 5. J Virol. 2002;76:11379-11386. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 91] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 89. | Lin GY, Paterson RG, Richardson CD, Lamb RA. The V protein of the paramyxovirus SV5 interacts with damage-specific DNA binding protein. Virology. 1998;249:189-200. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 107] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 90. | Andrejeva J, Young DF, Goodbourn S, Randall RE. Degradation of STAT1 and STAT2 by the V proteins of simian virus 5 and human parainfluenza virus type 2, respectively: consequences for virus replication in the presence of alpha/beta and gamma interferons. J Virol. 2002;76:2159-2167. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 133] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 91. | Sarikas A, Hartmann T, Pan ZQ. The cullin protein family. Genome Biol. 2011;12:220. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 421] [Cited by in RCA: 409] [Article Influence: 29.2] [Reference Citation Analysis (0)] |
| 92. | Leung-Pineda V, Huh J, Piwnica-Worms H. DDB1 targets Chk1 to the Cul4 E3 ligase complex in normal cycling cells and in cells experiencing replication stress. Cancer Res. 2009;69:2630-2637. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 103] [Cited by in RCA: 107] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 93. | Ulane CM, Horvath CM. Paramyxoviruses SV5 and HPIV2 assemble STAT protein ubiquitin ligase complexes from cellular components. Virology. 2002;304:160-166. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 185] [Cited by in RCA: 187] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 94. | Li T, Chen X, Garbutt KC, Zhou P, Zheng N. Structure of DDB1 in complex with a paramyxovirus V protein: viral hijack of a propeller cluster in ubiquitin ligase. Cell. 2006;124:105-117. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 197] [Cited by in RCA: 203] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 95. | Huang Z, Krishnamurthy S, Panda A, Samal SK. Newcastle disease virus V protein is associated with viral pathogenesis and functions as an alpha interferon antagonist. J Virol. 2003;77:8676-8685. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 169] [Cited by in RCA: 168] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 96. | Precious B, Young DF, Andrejeva L, Goodbourn S, Randall RE. In vitro and in vivo specificity of ubiquitination and degradation of STAT1 and STAT2 by the V proteins of the paramyxoviruses simian virus 5 and human parainfluenza virus type 2. J Gen Virol. 2005;86:151-158. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 42] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 97. | Precious B, Childs K, Fitzpatrick-Swallow V, Goodbourn S, Randall RE. Simian virus 5 V protein acts as an adaptor, linking DDB1 to STAT2, to facilitate the ubiquitination of STAT1. J Virol. 2005;79:13434-13441. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 93] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 98. | Parisien JP, Lau JF, Rodriguez JJ, Ulane CM, Horvath CM. Selective STAT protein degradation induced by paramyxoviruses requires both STAT1 and STAT2 but is independent of alpha/beta interferon signal transduction. J Virol. 2002;76:4190-4198. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 123] [Cited by in RCA: 121] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 99. | Ulane CM, Rodriguez JJ, Parisien JP, Horvath CM. STAT3 ubiquitylation and degradation by mumps virus suppress cytokine and oncogene signaling. J Virol. 2003;77:6385-6393. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 143] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 100. | Puri M, Lemon K, Duprex WP, Rima BK, Horvath CM. A point mutation, E95D, in the mumps virus V protein disengages STAT3 targeting from STAT1 targeting. J Virol. 2009;83:6347-6356. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 20] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 101. | Nishio M, Tsurudome M, Ito M, Ito Y. Human parainfluenza virus type 4 is incapable of evading the interferon-induced antiviral effect. J Virol. 2005;79:14756-14768. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 18] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 102. | Hagmaier K, Stock N, Precious B, Childs K, Wang LF, Goodbourn S, Randall RE. Mapuera virus, a rubulavirus that inhibits interferon signalling in a wide variety of mammalian cells without degrading STATs. J Gen Virol. 2007;88:956-966. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 22] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 103. | Caignard G, Bouraï M, Jacob Y, Tangy F, Vidalain PO. Inhibition of IFN-alpha/beta signaling by two discrete peptides within measles virus V protein that specifically bind STAT1 and STAT2. Virology. 2009;383:112-120. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 61] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 104. | Palosaari H, Parisien JP, Rodriguez JJ, Ulane CM, Horvath CM. STAT protein interference and suppression of cytokine signal transduction by measles virus V protein. J Virol. 2003;77:7635-7644. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 233] [Cited by in RCA: 237] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 105. | Ramachandran A, Parisien JP, Horvath CM. STAT2 is a primary target for measles virus V protein-mediated alpha/beta interferon signaling inhibition. J Virol. 2008;82:8330-8338. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 99] [Cited by in RCA: 95] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 106. | Takeuchi K, Kadota SI, Takeda M, Miyajima N, Nagata K. Measles virus V protein blocks interferon (IFN)-alpha/beta but not IFN-gamma signaling by inhibiting STAT1 and STAT2 phosphorylation. FEBS Lett. 2003;545:177-182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 164] [Cited by in RCA: 163] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 107. | Caignard G, Guerbois M, Labernardière JL, Jacob Y, Jones LM, Wild F, Tangy F, Vidalain PO. Measles virus V protein blocks Jak1-mediated phosphorylation of STAT1 to escape IFN-alpha/beta signaling. Virology. 2007;368:351-362. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 105] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 108. | Röthlisberger A, Wiener D, Schweizer M, Peterhans E, Zurbriggen A, Plattet P. Two domains of the V protein of virulent canine distemper virus selectively inhibit STAT1 and STAT2 nuclear import. J Virol. 2010;84:6328-6343. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 46] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 109. | Nanda SK, Baron MD. Rinderpest virus blocks type I and type II interferon action: role of structural and nonstructural proteins. J Virol. 2006;80:7555-7568. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 51] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 110. | Rodriguez JJ, Wang LF, Horvath CM. Hendra virus V protein inhibits interferon signaling by preventing STAT1 and STAT2 nuclear accumulation. J Virol. 2003;77:11842-11845. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 121] [Cited by in RCA: 120] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 111. | Komatsu T, Takeuchi K, Yokoo J, Tanaka Y, Gotoh B. Sendai virus blocks alpha interferon signaling to signal transducers and activators of transcription. J Virol. 2000;74:2477-2480. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 90] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 112. | Takayama I, Sato H, Watanabe A, Omi-Furutani M, Sugai A, Kanki K, Yoneda M, Kai C. The nucleocapsid protein of measles virus blocks host interferon response. Virology. 2012;424:45-55. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 28] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 113. | Power UF, Ryan KW, Portner A. The P genes of human parainfluenza virus type 1 clinical isolates are polycistronic and microheterogeneous. Virology. 1992;189:340-343. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 26] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 114. | Schomacker H, Hebner RM, Boonyaratanakornkit J, Surman S, Amaro-Carambot E, Collins PL, Schmidt AC. The C proteins of human parainfluenza virus type 1 block IFN signaling by binding and retaining Stat1 in perinuclear aggregates at the late endosome. PLoS One. 2012;7:e28382. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 20] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 115. | Gotoh B, Takeuchi K, Komatsu T, Yokoo J, Kimura Y, Kurotani A, Kato A, Nagai Y. Knockout of the Sendai virus C gene eliminates the viral ability to prevent the interferon-alpha/beta-mediated responses. FEBS Lett. 1999;459:205-210. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 111] [Cited by in RCA: 113] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 116. | Garcin D, Marq JB, Strahle L, le Mercier P, Kolakofsky D. All four Sendai Virus C proteins bind Stat1, but only the larger forms also induce its mono-ubiquitination and degradation. Virology. 2002;295:256-265. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 95] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 117. | Garcin D, Curran J, Kolakofsky D. Sendai virus C proteins must interact directly with cellular components to interfere with interferon action. J Virol. 2000;74:8823-8830. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 72] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 118. | Garcin D, Marq JB, Iseni F, Martin S, Kolakofsky D. A short peptide at the amino terminus of the Sendai virus C protein acts as an independent element that induces STAT1 instability. J Virol. 2004;78:8799-8811. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 27] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 119. | Garcin D, Marq JB, Goodbourn S, Kolakofsky D. The amino-terminal extensions of the longer Sendai virus C proteins modulate pY701-Stat1 and bulk Stat1 levels independently of interferon signaling. J Virol. 2003;77:2321-2329. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 44] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 120. | Kato A, Cortese-Grogan C, Moyer SA, Sugahara F, Sakaguchi T, Kubota T, Otsuki N, Kohase M, Tashiro M, Nagai Y. Characterization of the amino acid residues of sendai virus C protein that are critically involved in its interferon antagonism and RNA synthesis down-regulation. J Virol. 2004;78:7443-7454. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 42] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 121. | Kato A, Ohnishi Y, Hishiyama M, Kohase M, Saito S, Tashiro M, Nagai Y. The amino-terminal half of Sendai virus C protein is not responsible for either counteracting the antiviral action of interferons or down-regulating viral RNA synthesis. J Virol. 2002;76:7114-7124. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 31] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 122. | Ciancanelli MJ, Volchkova VA, Shaw ML, Volchkov VE, Basler CF. Nipah virus sequesters inactive STAT1 in the nucleus via a P gene-encoded mechanism. J Virol. 2009;83:7828-7841. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 101] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 123. | Rodriguez JJ, Cruz CD, Horvath CM. Identification of the nuclear export signal and STAT-binding domains of the Nipah virus V protein reveals mechanisms underlying interferon evasion. J Virol. 2004;78:5358-5367. [PubMed] |
| 124. | Rodriguez JJ, Parisien JP, Horvath CM. Nipah virus V protein evades alpha and gamma interferons by preventing STAT1 and STAT2 activation and nuclear accumulation. J Virol. 2002;76:11476-11483. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 234] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 125. | Shaw ML, García-Sastre A, Palese P, Basler CF. Nipah virus V and W proteins have a common STAT1-binding domain yet inhibit STAT1 activation from the cytoplasmic and nuclear compartments, respectively. J Virol. 2004;78:5633-5641. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 172] [Cited by in RCA: 190] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 126. | Hagmaier K, Stock N, Goodbourn S, Wang LF, Randall R. A single amino acid substitution in the V protein of Nipah virus alters its ability to block interferon signalling in cells from different species. J Gen Virol. 2006;87:3649-3653. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 34] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 127. | Virtue ER, Marsh GA, Wang LF. Interferon signaling remains functional during henipavirus infection of human cell lines. J Virol. 2011;85:4031-4034. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 35] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 128. | Geisbert TW, Feldmann H, Broder CC. Animal challenge models of henipavirus infection and pathogenesis. Curr Top Microbiol Immunol. 2012;359:153-177. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 55] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 129. | Park MS, García-Sastre A, Cros JF, Basler CF, Palese P. Newcastle disease virus V protein is a determinant of host range restriction. J Virol. 2003;77:9522-9532. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 179] [Cited by in RCA: 186] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 130. | Young DF, Chatziandreou N, He B, Goodbourn S, Lamb RA, Randall RE. Single amino acid substitution in the V protein of simian virus 5 differentiates its ability to block interferon signaling in human and murine cells. J Virol. 2001;75:3363-3370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 78] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 131. | Iwasaki M, Yanagi Y. Expression of the Sendai (murine parainfluenza) virus C protein alleviates restriction of measles virus growth in mouse cells. Proc Natl Acad Sci USA. 2011;108:15384-15389. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 17] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 132. | Parisien JP, Lau JF, Horvath CM. STAT2 acts as a host range determinant for species-specific paramyxovirus interferon antagonism and simian virus 5 replication. J Virol. 2002;76:6435-6441. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 89] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 133. | Lo MK, Miller D, Aljofan M, Mungall BA, Rollin PE, Bellini WJ, Rota PA. Characterization of the antiviral and inflammatory responses against Nipah virus in endothelial cells and neurons. Virology. 2010;404:78-88. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 65] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 134. | Schaap-Nutt A, Higgins C, Amaro-Carambot E, Nolan SM, D’Angelo C, Murphy BR, Collins PL, Schmidt AC. Identification of human parainfluenza virus type 2 (HPIV-2) V protein amino acid residues that reduce binding of V to MDA5 and attenuate HPIV-2 replication in nonhuman primates. J Virol. 2011;85:4007-4019. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 13] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 135. | Schaap-Nutt A, D’Angelo C, Amaro-Carambot E, Nolan SM, Davis S, Wise SM, Higgins C, Bradley K, Kim O, Mayor R. Recombinant human parainfluenza virus type 2 with mutations in V that permit cellular interferon signaling are not attenuated in non-human primates. Virology. 2010;406:65-79. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 8] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 136. | Schaap-Nutt A, D’Angelo C, Scull MA, Amaro-Carambot E, Nishio M, Pickles RJ, Collins PL, Murphy BR, Schmidt AC. Human parainfluenza virus type 2 V protein inhibits interferon production and signaling and is required for replication in non-human primates. Virology. 2010;397:285-298. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 15] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 137. | Devaux P, Hudacek AW, Hodge G, Reyes-Del Valle J, McChesney MB, Cattaneo R. A recombinant measles virus unable to antagonize STAT1 function cannot control inflammation and is attenuated in rhesus monkeys. J Virol. 2011;85:348-356. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 48] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 138. | Patterson JB, Thomas D, Lewicki H, Billeter MA, Oldstone MB. V and C proteins of measles virus function as virulence factors in vivo. Virology. 2000;267:80-89. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 126] [Cited by in RCA: 116] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 139. | Yoneda M, Guillaume V, Sato H, Fujita K, Georges-Courbot MC, Ikeda F, Omi M, Muto-Terao Y, Wild TF, Kai C. The nonstructural proteins of Nipah virus play a key role in pathogenicity in experimentally infected animals. PLoS One. 2010;5:e12709. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 78] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 140. | Gitlin L, Benoit L, Song C, Cella M, Gilfillan S, Holtzman MJ, Colonna M. Melanoma differentiation-associated gene 5 (MDA5) is involved in the innate immune response to Paramyxoviridae infection in vivo. PLoS Pathog. 2010;6:e1000734. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 99] [Cited by in RCA: 108] [Article Influence: 7.2] [Reference Citation Analysis (0)] |