Published online Jan 19, 2021. doi: 10.5498/wjp.v11.i1.1
Peer-review started: November 24, 2020
First decision: December 12, 2020
Revised: December 15, 2020
Accepted: December 26, 2020
Article in press: December 26, 2020
Published online: January 19, 2021
Processing time: 48 Days and 2.2 Hours
Psychiatry remains in a permanent state of crisis, which fragmented psychiatry from the field of medicine. The crisis in psychiatry is evidenced by the many different competing approaches to psychiatric illness including psychodynamic, biological, molecular, pan-omics, precision, cognitive and phenomenological psychiatry, folk psychology, mind-brain dualism, descriptive psychopathology, and postpsychiatry. The current “gold standard” Diagnostic and Statistical Manual of Mental Disorders/International Classification of Diseases taxonomies of mood disorders and schizophrenia are unreliable and preclude to employ a deductive reasoning approach. Therefore, it is not surprising that mood disorders and schizophrenia research was unable to revise the conventional classifications and did not provide more adequate therapeutic approaches. The aim of this paper is to explain the new nomothetic network psychiatry (NNP) approach, which uses machine learning methods to build data-driven causal models of mental illness by assembling risk-resilience, adverse outcome pathways (AOP), cognitome, brainome, staging, symptomatome, and phenomenome latent scores in a causal model. The latter may be trained, tested and validated with Partial Least Squares analysis. This approach not only allows to compute pathway-phenotypes or biosignatures, but also to construct reliable and replicable nomothetic networks, which are, therefore, generalizable as disease models. After integrating the validated feature vectors into a well-fitting nomothetic network, clustering analysis may be applied on the latent variable scores of the R/R, AOP, cognitome, brainome, and phenome latent vectors. This pattern recognition method may expose new (transdiagnostic) classes of patients which if cross-validated in independent samples may constitute new (transdiagnostic) nosological categories.
Core Tip: The nomothetic network psychiatry approach is a new method which aims to construct causal models of schizophrenia and mood disorders by integrating all features of those mental illnesses into a data-driven model. These features comprise data on risk-resilience, adverse outcome pathways, the cognitome, brainome, symptomatome, staging, and the phenomenome. Partial Least Squares analysis may be employed to train, test, and validate those models and to build pathway-phenotypes or biosignatures. Clustering analysis performed on all illness features, reduced into latent traits scores, may expose relevant new transdiagnostic classes.
- Citation: Stoyanov D, Maes MH. How to construct neuroscience-informed psychiatric classification? Towards nomothetic networks psychiatry. World J Psychiatr 2021; 11(1): 1-12
- URL: https://www.wjgnet.com/2220-3206/full/v11/i1/1.htm
- DOI: https://dx.doi.org/10.5498/wjp.v11.i1.1
For the past 200 years, psychiatry remained a discipline plagued by conceptual controversies, whose roots go back to its poor ontological and epistemological foundations[1]. Psychiatry remained in a permanent state of crisis due to methodological mistrust in psychiatric case definitions, which fragmented psychiatry from the field of medicine[2]. The crisis is psychiatry is further evidenced by the many different competing approaches and ways to understand mental and psychiatric disorders including the etiological approach of psychodynamic psychiatry, biological, molecular, pan-omics, and precision psychiatry, cognitive psychiatry, folk psychology, the mind-brain dualism, descriptive psychopathology, postpsychiatry, and phenomenological psychiatry. Moreover, the gold standard taxonomies used to diagnose mood disorders and schizophrenia are not reliable[3,4].
Recently, we employed a new approach, namely the nomothetic network psychiatry (NNP) approach, which uses machine learning methods to build new data-driven models of mood disorders and schizophrenia using all features of those disorders including etiological, context centered hermeneutic, biological, molecular, cognitive, descriptive psychopathological, and phenomenological features[3-7]. The aim of this opinion paper is to review how to build nomothetic networks using Partial least Squares (PLS) analysis and how to expose new classifications of these disorders using unsupervised pattern recognition techniques.
Folk or commonsense psychology tries to explain the mental state of individuals including symptoms, cognitions, or behaviors as the outcome of everyday life psychology and daily life experiences such as pleasure, sensations, pain, common beliefs, perceptions, etc.[8]. Folk psychology narratives are embodied in psychiatric inventories, either observational interviews or self-evaluation scales, which are supposed to deliver meaningful information intended to contribute to the diagnostic criteria of the conventional psychiatric classification systems or to rating scales that measure severity of illness. Thus, structural components of current psychiatric inventories are decomposed into items (statements and questions) some of which are formulated in a folk psychology-like language. For example, items such as “I cry easily” or “I feel down and depressed” are borrowed from folk psychology. In a futile effort to translate these symptoms into a more technical and medical jargon, such as depressive mood for instance (anhedonia and dysthymia), those items are then scored on a 4- or 7-degrees Likert scale so that the total score may resemble a statistically digestible entity. In this manner, common sense folk psychology expressions are converted into “diagnostic” statements without any reference to independent validators. The most common “state” dependent clinical measures or inventories in psychiatry remain folk psychology narratives with some window dressing for statistical purposes. What is missing in such perspective is the biological, neuronal, and cognitive basis to better understand the existing phenomena in psychopathology, which is declared on the agenda of post-modern psychiatry.
There are two main intellectual frameworks which outline the rationale behind the scientific enterprise in psychiatry. The first is psycho-physical dualism which is supposed to drive the advances in psychotherapy and psychosocial interventions in mental illness. Mind-body dualism is the theory which proposes that mental phenomena are non-physical or that not all mental processes are physical. As such, mind and body would be, at least in part, separable entities[9,10]. The common psychiatric approach is essentially focused on what might be described as “mind” in terms of the mind-brain debate.
The second is the physicalism stance, which considers that everything is physical[11]. Physicalism and materialism are implicated as a primary assumption in influential advances in psychiatric research including biological, molecular, and pan-omics psychiatry, functional neuroimaging, and cognitive science[12].
Early efforts initiated by the Wilhelm Griesinger and the Wernicke-Kleist-Leonhard schools tried to consolidate psychiatric nosology using organic etiological factors[13]. These early theories, associated with the notions of localizationism, culminated in the works of Karl Wernicke and Karl Kleist and Maynert-Wernicke’s connectionism. Psychodynamic theories, initiated by Sigmund Freud, and later versions of psychoanalysis, applied a conceptual organization of psychiatric syndromes based on a psychodynamic etiologic approach but remained grounded on the tacit assumptions of psychophysical dualism. However, attempts to bring together the disparate etiological explanatory models of psychodynamic paradigms and localizationism and clinical diagnoses proved to be inefficient. Neither of those views offered consolidated and sustainable pictures of psychiatric diagnoses applicable in the medical practice.
Descriptive psychopathology or psychiatry focusses on readily observable behaviors and symptoms, rather than on underlying psychoanalytic or organic etiologies. Phenomenological psychopathology focuses on the patients subjective, own lived experiences of selfhood, space, time, body, and mind[14].
Current gold standard psychiatric classifications are based on descriptive and phenomenological psychopathology, including the Diagnostic and Statistical Manual of Mental Disorders (DSM) and the International Classification of Diseases (ICD) taxonomies. These case definitions are derived from cross-culturally diverse, even sometimes unique criteria[15], established ex convention by professional bodies like the American Psychiatric Association (APA) and the World Health Association (WHO). These ex consensus-based case definitions of nosological psychiatric classes use de-contextualized narratives and descriptive features of the disorder derived from folk psychology-like self-reports by the patient and observer-based interviews[16].
Due to the missing etiological and biomarker foundations, one crucial limitation of the current classifications, such as DSM-5 and ICD-10, is their top-down manner of generation[3-5,16]. The structured interviews, which are used to construct diagnostic categories, pre-define the clinical diagnosis before other tests are performed, including etiologic, state and trait-biomarkers, brain imaging, and cognitive probes. In that regard the diagnosis remains based on controversial and value laden statements, whereas the causal and biological measures are supposed to be concomitant data, either supporting the diagnosis or not. However, no falsification or dispute of the diagnostic assumption is possible based on information from outside the data source of the clinical interview, thus precluding a top-down deductive approach[3-5]. It should be added that those professional bodies are most often under the influence of para-motivation from the pharmaceutical industry or other confounds which leads to deeply controversial and in the end of the day counter-productive debates on the existence of specific case definitions of psychiatric illness.
Moreover, the taxonomies used to make the diagnosis of psychotic disorders show inadequate reliability validity as for example indicated by significant differences in the diagnoses of DSM-III-R, DSM-IV, ICD-8, ICD-9, and ICD-10 classifications[16,17]. There is considerable inter-departmental diagnostic variability in the ICD-8 and ICD-10 diagnosis[18] explaining that schizophrenia may be often overdiagnosed or underdiagnosed[17]. In addition, the DSM suffers from a poor demarcation of the clinical heterogeneity present in schizophrenia[4,19]. For example, using machine learning techniques we discovered that schizophrenia consists of qualitatively distinct categories (including deficit vs non-deficit schizophrenia)[20], indicating that schizophrenia biomarker research which does not take this distinction into account is bound to fail. Also, the DSM case definitions of mood disorders including major depressive disorder (MDD) lack reliability validity[21], with MDD taxonomies showing minimal agreement between psychiatrists[22]. Furthermore, there was limited or no unification and harmonization of the DSM case definitions[18]. All in all, these taxonomies lack reliability and validity and are therefore counterproductive for research purposes[3-5,16,23-25].
A more radical physicalism theory, namely eliminative materialism, has been outlined in the past decades, especially under the influence of Churchland[26]. This theory applied to psychiatry relies on neuroscience and aims to replace the “folk” psychology vocabulary and methods on a systematic level by material concepts, namely aberrations in brain functions and neurocircuitry. Biological psychiatry aims to explain mental illness in terms of biological aberrations in neuronal functions; molecular psychiatry explains mental illness based on molecular pathways including the effects of genes and intracellular networks; and cognitive psychiatry explains mental illness though effects of cognitive impairments and their neuronal substrates. Nevertheless, the biological, molecular and cognitive approaches turned out to be insufficient to delineate biomarker or cognitive tools that externally validate the case definitions.
Biological, molecular and cognitive psychiatry research generally uses the “gold standard” DSM/ICD case definitions of mood and psychotic disorders in top-down research[3,4,16]. These methods commonly enter diagnosis as explanatory variable in GLM analysis or analysis of variance to analyze alterations in causome (e.g., early lifetime trauma), biomarker levels, brainome data, and cognitive probe scores. The latter are entered as the dependent variables even when causal reasoning shows that they should be employed as the explanatory variables in logistic regression or other machine learning techniques, including neural networks. Consequently, most biological and cognitive psychiatry research projects employ unreliable diagnostic classes applied in inadequate model assumptions and tested with inappropriate statistical tests[3-5]. Also, molecular psychiatry uses a similar approach when examining pathways and networks or when conducting studies which associate genetic markers with the DSM/ICD taxonomies. A newer method, namely the Research Diagnostic criteria (RDoC), developed by the NIH, tries to integrate genetic, neurodevelopmental, environmental factors, with social, regulatory, cognitive and social domains, with negative and positive valence[25]. However, also the RDoC is largely a top-down concept driven by ex-consensus commitments by experts.
Another critical point is that the entire hypostasis of eliminative materialism of biological, molecular, and cognitive psychiatry is a fragmented or “patchy” reductionist approach[27] whereby psychiatric diagnoses tend to be reduced to neuronal entities, genetic markers, plasma biomarkers, intracellular signaling molecules, or functional MRI responses to emotional tasks. What is missing is an integrated model with precise mapping of genomics data, specific (causal and protective) and generalized (e.g., context centered and lifestyle) environmental factors, and phenome features. All in all, the current “gold standard” DSM/ICD taxonomies are unreliable constructs and preclude using a deductive reasoning approach and, therefore, it is not surprising that biological, molecular, and cognitive psychiatry research was unable to revise the conventional classifications and did not provide valid predictions from a therapeutic perspective.
Pan-omics psychiatry proposes to use systems biomedicine to decipher the complex non-linear interactions between pathways and intracellular networks that govern those pathways, and the multifactorial factors including genes and environmental factors that may trigger those pathways/networks[28]. Pan-omics psychiatry proposes to use a data-driven bottom-up approach to compute biosignatures consisting of molecular pathways and networks and symptoms as well as environmental features thereby developing pathway-phenotypes[28,29]. A related field is precision psychiatry, which is based on precision medicine defined as “an emerging approach for treatment and prevention that takes into account each person’s variability in genes, environment, and lifestyle”[30]. Precision psychiatry aims to transform the psychiatric landscape through a bottom-up approach applied to pan-omics using system biology and computer science to compute a biosignature, which in turn may be used in a top-down approach to help to understand domains, which differ from components but allow to construct endophenotypes[31]. Both pan-omics and precision psychiatry propose to combine cognitive neuroscience, neural circuits, big data, molecular biosignatures, individual characteristics, physiology, and environment into a biosignature, which is a feature set defining an endophenotype[28,31]. These methods[31], however, do not aim to integrate all features of complex psychiatric disorders into a model characterized by causal paths linking causome (all possible causal factors), protectome (all possible protective factors), adverse output pathways (AOPs, namely biological, molecular, pan-omics, brain imaging features) and phenome (cognitome, symptomatome and phenomenome) feature sets.
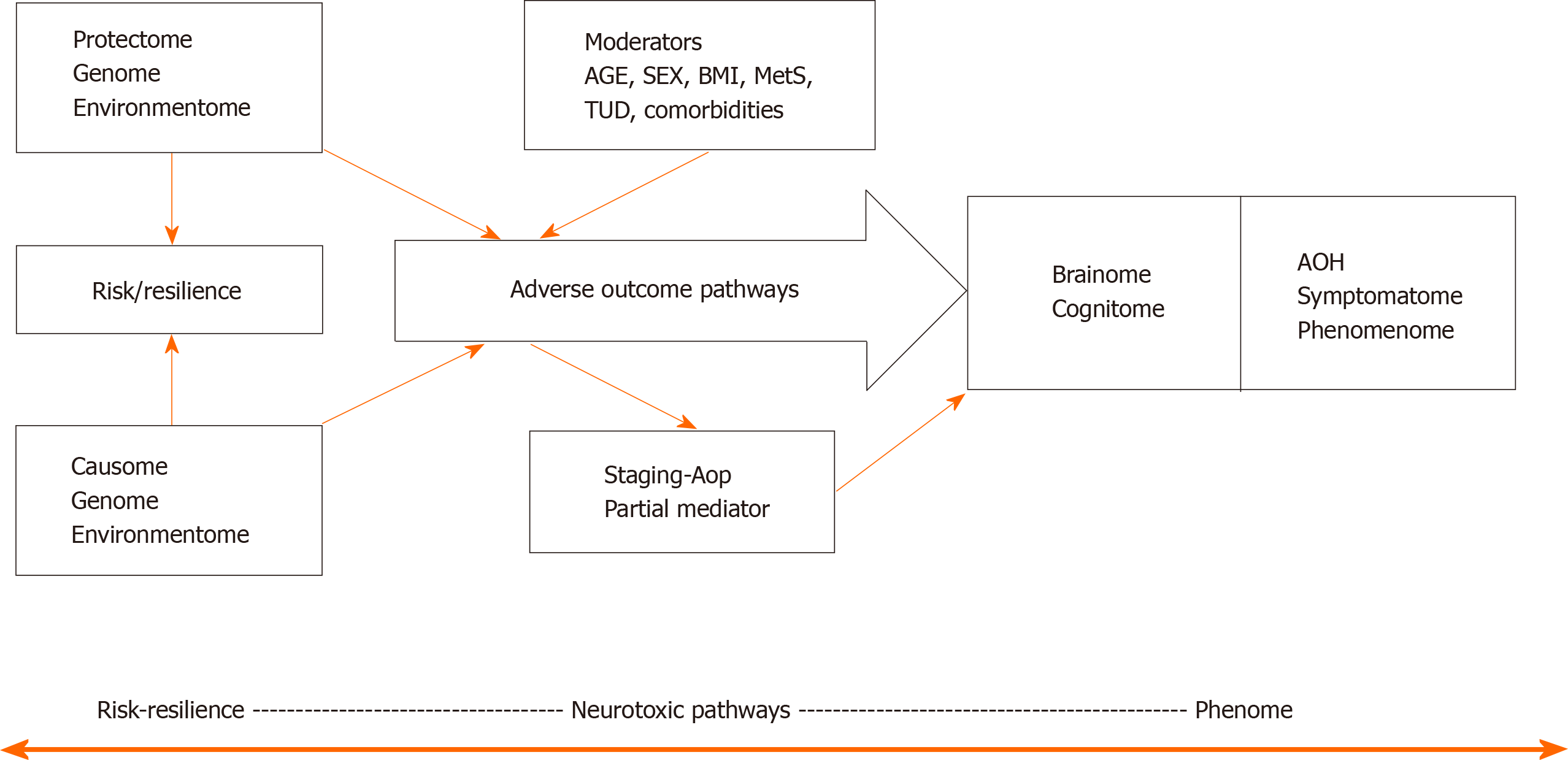
Figure 1 shows a causal theoretical model applicable to mood and psychotic disorders[3-7]. Causal reasoning based on state-of-the-art knowledge of these mental disorders indicates that causome features including genes and its products including enzymatic activity as well as specific environmentome factors (e.g., early lifetime trauma) predict AOPs and the phenome of the illness[32]. Moreover, the generalized environmentome (including lifestyle, nutrition, toxins, context centered social, cultural, and political factors) should be added. Pan-omics may be employed to measure causome (genomics) and AOPs (e.g., immunomics, epigenomics, transcriptomics, metabolomics, and proteomics). In psychiatry, another important AOP component is the brainome, which may be assessed using in vivo histology spectroscopy and magnetic resonance imaging. Also, the phenome of psychiatric disorders is very complex and consists of various feature sets including (1) staging of the disorder, as defined by recurrence of episodes and suicidal attempts, chronicity, etc.[32]; (2) the cognitome, namely the aggregate of cognitive features of the illness including in memory, executive functions, and attention; (3) the symptomatome, namely the aggregate of observed clinical symptoms, illness severity, subtypes, treatment responsivity; and (4) the phenomenome, namely the illness features as experienced by the patient[3,4].
One of the aims of our new nomothetic network psychiatry (NNP) approach is to reunify such data (1, 2, 3 and 4) into an illness model, which integrates different approaches including etiological, biological, molecular, pan-omics, cognitive, descriptive and phenomenological psychiatry, as well as folk psychology and postpsychiatry. In fact, machine learning conducted on these data will extract and select the most important features in a process referred to as feature re-engineering, selection, and learning to make the most accurate models of mood and psychotic disorders.
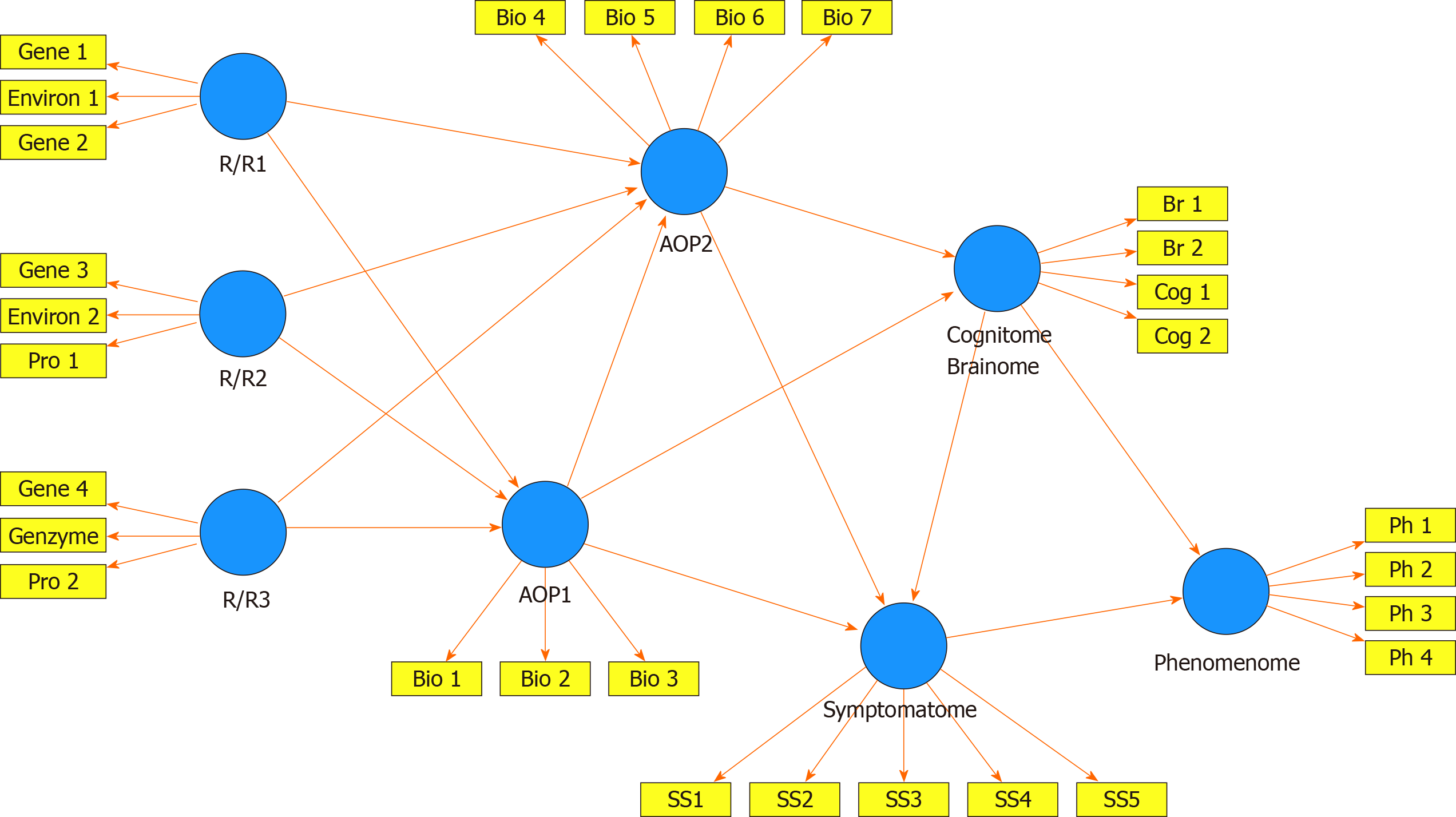
Figure 2 shows a framework or general structure which is based on scientific evidence linking a multitude of causome, protectome, AOPs, cognitome, brainome, staging, symptomatome, and phenomenome data. The selection of variables (indicators) and the concept of the framework are guided by the available theoretical knowledge, expertise, and information accumulated over the past decades and by formal causal reasoning. Nevertheless, the framework shown in Figure 2 is far for complete as for example socio-demographic data and generalized environmentome factors (e.g., those relevant to postpsychiatry) should be added to the model but are deleted from the figure for reasons of clarity. Consequently, data (facts) are accumulated to test this theoretical framework[33].
The multitude of data to be entered should first be reduced (dimensionality reduction) to a smaller number of relevant feature sets or vectors using feature construction processes[3,4]. The first step is to re-engineer the causome and protectome data into one of more new feature sets reflecting risk-resilience (R/R), namely the balance between causal and risk factors[3,4]. These R/R feature sets can consequently be used as input variables (predictors) in logistic models, regression analysis, and neural networks to expose their effects on the downstream features of the framework (AOP and phenome data). For example, in schizophrenia, we established that R/R indices re-engineered from genome data, i.e., paraoxonase 1 (PON1) Q192R genotype combined with PON1 enzymatic activity, zonulin levels (a product of the haptoglobin 2-2 genotype), and lowered natural IgM (a protectome factor) predict AOPs (neuro-immune and neuro-oxidative toxicity pathways), cognitome (episodic and semantic memory and executive functions), symptomatome (psychosis, hostility, excitation, mannerism, negative symptoms) and phenomenome (self-rated quality of life) features[4]. In mood disorders, a new R/R index consisting of the PON1 Q192R genotype combined with PON1 enzymatic activity and early lifetime trauma predicts AOPs (antioxidant defenses and neuro-oxidative stress biomarkers), the symptomatome (depression severity, suicidal ideation, and mood disorders subtypes such as treatment resistance and melancholia) and the phenomenome (self-rated quality of life and disabilities)[3].
In the framework displayed in Figure 2, the newly re-engineered R/R feature sets are entered as input variables and can predict downstream feature sets as delineated through formal causal reasoning. The indicators of all downstream concepts are represented as latent vectors extracted from a set of features in reflective models because the aim is to construct a single underlying trait (e.g., the symptomatome) which explains its manifestations (e.g., all different symptom domains and phenotypes)[3-7]. The phenome feature sets are entered as output variables, whereas AOPs, cognitomone, and brainome feature sets predict the phenome and are predicted by the R/R features. It should be underscored that this method allows to reduce many features into a few relevant single traits. As such, the framework displayed in Figure 2 comprises one dependent variable (namely a latent vector reflecting the phenomenome) which is predicted by seven input variables, namely the RR, AOP, brainome, and symptomatome latent vectors. This parsimonious formal causal framework can then be trained, tested and validated using PLS structural equation modeling[3-7,34].
PLS analysis allows to build pathway-phenotypes or biosignatures and train and evaluate a novel nomothetic model based on a combination of factor and regression analysis. Pathway-phenotypes may be exposed by combining for example AOPs with cognitome features into new reflective indicators of molecular paths and cognitive functions that underpin the illness[35]. Causal-pathway-phenotypes may be exposed by combining causome (e.g., number of transfusions in transfusion-dependent thalassemia or TDT), AOPs (iron overload biomarkers and neuro-immune pathways as a consequence of the transfusions) with the phenome (depressive symptoms) into reflective indicators of a single latent trait, namely “depression due to immune activation as a consequence of transfusions and iron overload in TDT”[7]. Interestingly, in mood disorders, but not schizophrenia, symptomatome and phenomenome features may be combined as reflective manifestations of a single latent trait, namely the clinical – phenomenological phenome[3,4].
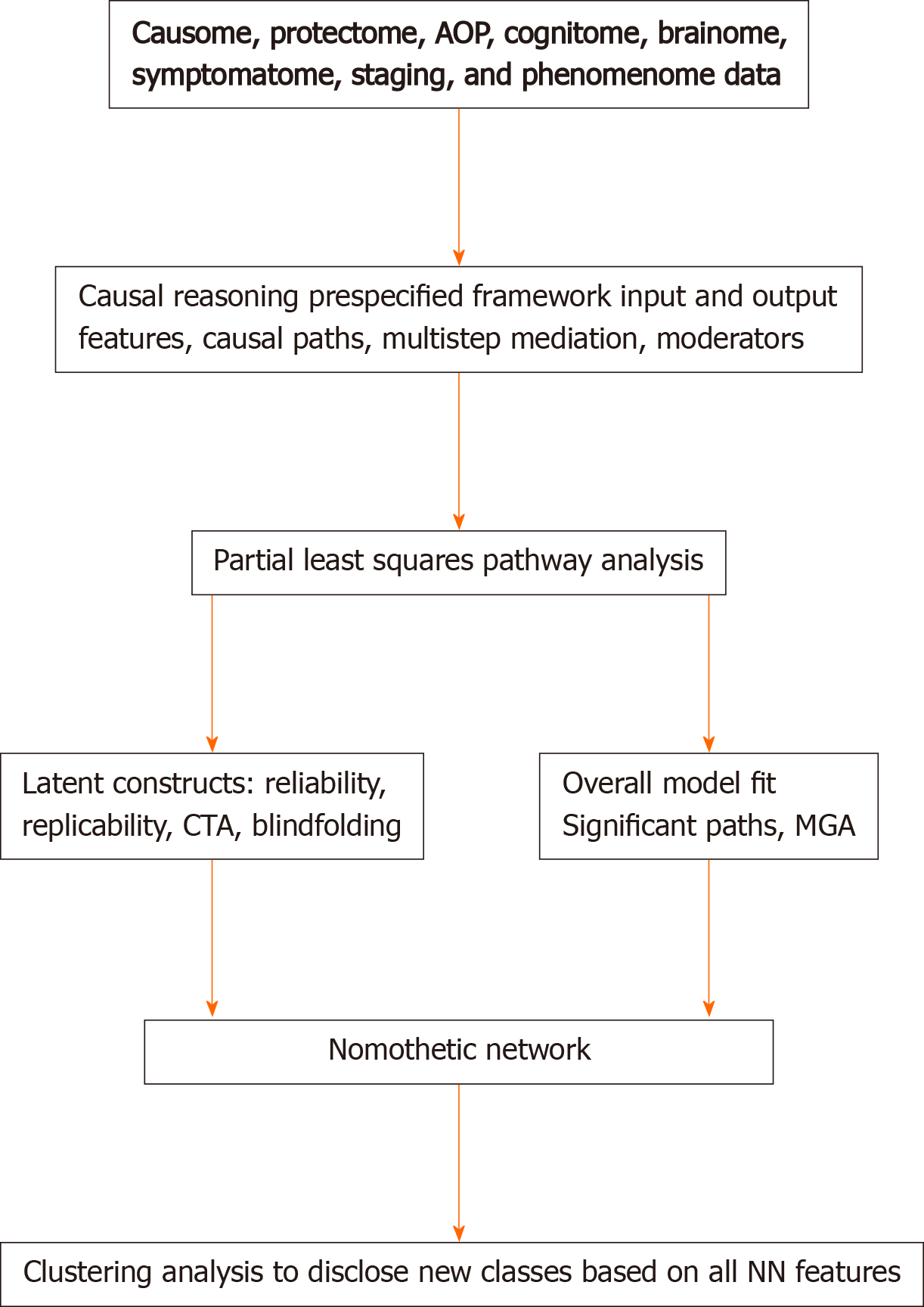
Figure 3 shows that the theoretical framework including new pathway-phenotypes can be trained and tested employing PLS on bootstrapped samples (e.g., 5.000)[36]. Goodness of fit should be assessed using standardized root mean square residuals to avoid model misspecifications. The validity reliability of the latent vectors should be evaluated using psychometric properties such as composite reliability, rho-A, Cronbach’s alpha, and average variance extracted values. All indicators of the LVs should show adequate loadings > 0.5 or by preference > 0.66[36]. Moreover, confirmatory tetrad analysis (CTA) should be used to ascertain whether the LV models are not mis-specified as reflective models, and blindfolding is used to test the construct cross-validated redundancies, which test the predictive relevance of the output LVs in the model[3,4]. Sample size determination and statistical power estimation should be performed based on (1) the psychometric properties of the vectors (factor loadings) and the strength of the intercorrelations among the vectors, (2) the explained variance and the maximum number of arrows pointing to a construct, or (3) power analysis specific to multiple regression analysis[37,38]. These methods show that to achieve a power of 0.8 in the PLS model displayed in Maes et al[3] a relatively small sample size of n = 70-127 is sufficient. Nevertheless, larger sample sizes will yield more stable parameter estimates.
Consequently, PLS is conducted on bootstrapped samples which expose the path coefficients with exact p-values of all significant links (paths), as well as the total direct and indirect and specific indirect effects. Importantly, the indirect effects indicate the mediating effects of upstream on downstream indicators including in multistep mediating models. For example, in Figure 2, the R/R feature sets may have significant indirect effects on the phenomenome, which are mediated by the paths from AOP1 to the symptomatome or by the path from AOP2 to the cognitome to the symptomatome. In addition, also moderator (interaction) effects between 2 or more downstream indicators on upstream indicators may be added to the model which may account for possible moderating effects of age, sex, metabolic syndrome, and comorbidities. Finally, PLS allows to establish possible group differences in the model or paths using Multi-Group-Analysis (PLS-MGA) or permutations, which can be employed to examine differences in the model or paths, for example between different genotypes and between men and women. The latter is important to examine in schizophrenia and mood disorders because sexual dimorphisms were detected in those disorders[39,40]. Using PLS-MGA in schizophrenia we found significant differences between both women and men in the path from AOP to the phenomenome (quality of life) with a significant impact in women, but not in men[4]. On the other hand, no significant sex-related differences in the nomothetic network or in any of the pathways could be detected in mood disorders[3,5].
In summary, a bottom-up, data-driven model of mood disorders and schizophrenia may be constructed using the knowledge-based causal framework shown in Figure 2 and by assembling R/R, AOPs, and phenome feature sets into an explicit data model, namely the PLS nomothetic network. Nomothetic indicates the tendency to generalize and to derive models (“laws”) from independent variables, which explain variations in phenomena[41]. As such, the nomothetic network approach objectivates the symptomatome and phenomenome of mood disorders and schizophrenia[3-5], and, therefore, translates R/R, AOP, brainome, and cognitome feature sets into relevant descriptive narratives. The process which reifies the abstract concepts of descriptive narratives to realize a more concrete and material concept using computer science is named “reification of clinical diagnosis”. It is important to note that in contrast to pan-omics and precision psychiatry, the aim of our nomothetic network approach is not only to compute pathways-phenotypes or biosignatures, but especially to make a nomothetic network, with causal links between the building blocks of the disease.
It should be added that this nomothetic network approach, in contrast to biological psychiatry models[42-44], may pass Karl Popper’s critical rationalism test[33]. Indeed, our nomothetic networks are progressive (the model is based on all available knowledge), parsimonious (through feature reduction), changeable (other researchers can elaborate on the model and add more indicators or delete less robust features), provisional (the latent variable scores of the network will change when more pan-omics and brainome data are added), and falsifiable (the network can be refuted or corroborated). In this respect, our nomothetic networks deserve validation in more heterogeneous study groups consisting of individuals with comorbid psychiatric and medical disorders.
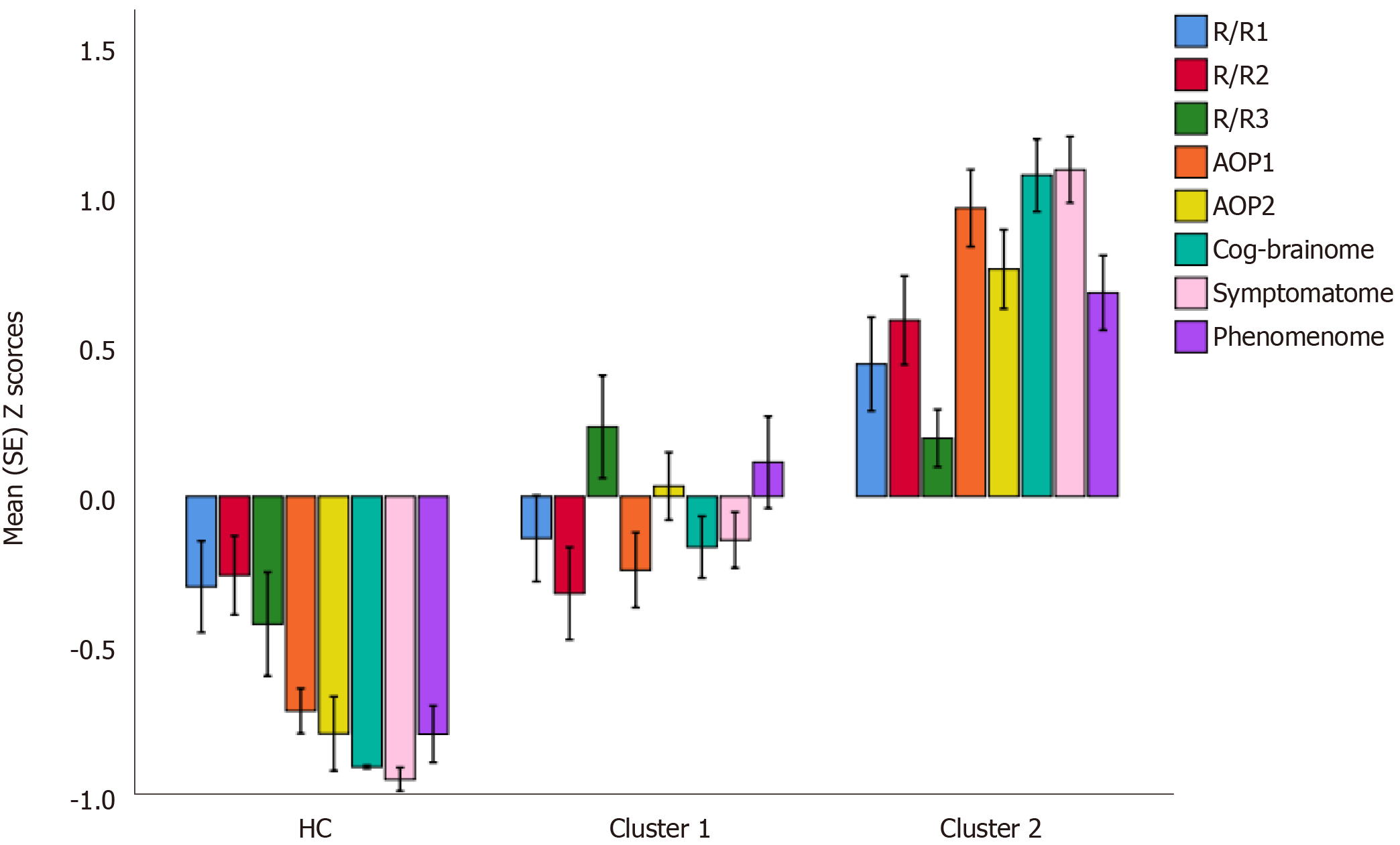
After integrating the validated feature vectors into a well-fitting nomothetic network, latent variable scores may be computed which reflect the severity of the various R/R, AOP, brainome, cognitome, staging, symptomatome, and phenomenome feature sets. The latter may be employed in unsupervised pattern recognition methods, including clustering analysis, to expose new categories (Figure 3). Previously, we employed different clustering techniques on such latent variable scores including K-mean, K-median, and Ward’s and Forgy’s methods[3-5]. Figure 4 shows a hypothetical example of cluster analysis-generated classes, with the latent variable scores (in z transformation) displayed in a clustered bar graph. This figure shows a normal cluster with healthy control subjects and two patients clusters. The second patient cluster may be discriminated from the first cluster (and from controls) by higher R/R, AOP, cognitive and brainome, symptomatome and phenomenome scores. The first patient cluster may be discriminated from controls by increased R/R3, AOP, and phenome scores. Previously, we showed that, in mood disorders, these new bottom-up cluster analysis-derived classes are more influential for classification purposes than the top-down classification into bipolar type 1 and type 2 and major depression. As such, new mechanistic, biosignature-based, and/or transdiagnostic classes may be discovered[3,5].
Nevertheless, these new classes should be cross-validated in independent samples using other machine learning methods including support vector machine with 10-fold cross-validation or soft independent modeling of class analogy (SIMCA)[7]. It is interesting to note that our nomothetic networks computed in mood disorders and schizophrenia contain self-rated phenomenological features (including self-rated quality of life and severity of disabilities), and, therefore, may comprise idiographic features[3,4]. Therefore, the latent variable scores not only delineate an objective nomothetic network and new diagnostic classes, but also shape an idiomatic feature profile, which is unique for every individual. As such, adequate treatments of mood disorders and schizophrenia should target the components of the nomothetic networks (R/R, AOP, brainome) constructed in those disorders. In addition, the individualized feature profile allows a more personalized treatment targeting aberrations in specific R/R, AOP, cognitome, brainome, and staging latent variable scores.
In this paper we explained how to use the new nomothetic network psychiatry (NNP) approach to construct new causal models of mental illness by machine learning techniques, which assemble all features of mental illness, namely risk-resilience, AOPs, cognitome, brainome, symptomatome, staging, and phenomenome scores. PLS analysis may successfully be used to train, test and validate those models, to build pathway-phenotypes or biosignatures, and to construct comprehensive models of mood disorders and schizophrenia which objectivate the clinical phenome of those disorders. Clustering analysis performed on all illness features reduced into latent traits may expose relevant new (transdiagnostic) classes. The reification of the clinical diagnosis of mood disorders and schizophrenia (and by inference other psychiatric disorders) using the nomothetic network psychiatry approach is an awaited achievement which constitutes a major paradigm shift in psychiatry[16].
Manuscript source: Invited manuscript
Specialty type: Psychiatry
Country/Territory of origin: Bulgaria
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B, B, B, B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Cheng J, Tcheremissine OV S-Editor: Gao CC L-Editor: A P-Editor: Li JH
| 1. | Stoyanov D, Di Nicola V. Psychiatry in crisis: Epistemological and ontological concerns. Am J Psychiatry Neurosci. 2017;5:6. |
| 2. | Di Nicola V, Stoyanov D. Psychiatry in Crisis: at the Crossroads of Social Sciences, the Humanities, and Neuroscience, Springer: 2021. |
| 3. | Maes M, Brum J, Bonifacio K, Barbosa D, Vargas H, Michelin AP, Nunes S. Towards A New Model and Classification of Mood Disorders based on Risk Resilience, Neuro-Affective Toxicity, Staging, and Phenome Features Using the Nomothetic Network Psychiatry Approach. Preprints 2020, 2020090610. [DOI] [Full Text] |
| 4. | Maes M, Vojdani A, Galecki P, Kanchanatawan B. How to Construct a Bottom-Up Nomothetic Network Model and Disclose Novel Nosological Classes by Integrating Risk Resilience and Adverse Outcome Pathways with the Phenome of Schizophrenia. Brain Sci. 2020;10. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 5. | Simeonova D, Stoyanov D, Leunis JC, Murdjeva M, Maes M. Construction of a nitro-oxidative stress-driven, mechanistic model of mood disorders: A nomothetic network approach. Nitric Oxide. 2021;106:45-54. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 28] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 6. | Mousa R, Smesam H, Qazmooz H, Al-Hakeim H, Maes M. A Nomothetic Network Model Disclosing the Comorbidity of Depression and Unstable Angina: Effects of Atherogenicity, Insulin Resistance, Immune Activation, Antioxidants, the Endogenous Opioid System, Trace Elements, and Macrominerals. Preprints 2020. [DOI] [Full Text] |
| 7. | Al-Hakeim H, Najm A, Moustafa S, Maes M. Construction of an Exposure-Pathway-Phenotype in Children with Depression due to Transfusion-Dependent Thalassemia: Results of (Un)supervised Machine Learning. Preprints 2020. [DOI] [Full Text] |
| 8. | Ravenscroft I. Folk Psychology as a Theory. In: Stanford Encyclopedia of Philosophy, 2016.. |
| 9. | Available from: https://plato.stanford.edu/entries/folkpsych-theory/. |
| 10. | Hart WD. "Dualism." In: A Companion to the Philosophy of Mind, edited by S. Guttenplan, Oxford, Blackwell, 1996: 265-267. |
| 11. | Crane T, Patterson S. History of the Mind-Body Problem. Taylor and Francis, London Studies in the History of Philosophy. 2002: 1-264. |
| 12. | Stoljar D. "Physicalism". In: Edward N. Zalta. The Stanford Encyclopedia of Philosophy, Fall 2009 Edition. 2009. Retrieved 2014-08-07. |
| 13. | Stoyanov DS. An essay on the mind-brain problem and legal proof. Balkan J Philosophy. 2018;10:27-36. |
| 14. | Stoyanov D, Telles-Correia D, Cuthbert BN. The Research Domain Criteria (RDoC) and the historical roots of psychopathology: A viewpoint. Eur Psychiatry. 2019;57:58-60. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 17] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 15. | Messas G, Tamelini M, Mancini M, Stanghellini G. New Perspectives in Phenomenological Psychopathology: Its Use in Psychiatric Treatment. Front Psychiatry. 2018;9:466. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 22] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 16. | Stoyanov D, Fulford B, Stanghellini G, Van Staden W, Wong M. International Perspectives in Values-Based Mental Health Practice: Case Studies and Commentaries, Springer International, eBook, 2021. |
| 17. | Stoyanov D. The Reification of Diagnosis in Psychiatry. Neurotox Res. 2020;37:772-774. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 18. | Pihlajamaa J, Suvisaari J, Henriksson M, Heilä H, Karjalainen E, Koskela J, Cannon M, Lönnqvist J. The validity of schizophrenia diagnosis in the Finnish Hospital Discharge Register: findings from a 10-year birth cohort sample. Nord J Psychiatry. 2008;62:198-203. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 76] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 19. | Nordgaard J, Jessen K, Sæbye D, Parnas J. Variability in clinical diagnoses during the ICD-8 and ICD-10 era. Soc Psychiatry Psychiatr Epidemiol. 2016;51:1293-1299. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 14] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 20. | Tandon R. Schizophrenia and Other Psychotic Disorders in Diagnostic and Statistical Manual of Mental Disorders (DSM)-5: Clinical Implications of Revisions from DSM-IV. Indian J Psychol Med. 2014;36:223-225. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 20] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 21. | Kanchanatawan B, Sriswasdi S, Thika S, Sirivichayakul S, Carvalho AF, Geffard M, Kubera M, Maes M. Deficit schizophrenia is a discrete diagnostic category defined by neuro-immune and neurocognitive features: results of supervised machine learning. Metab Brain Dis. 2018;33:1053-1067. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 34] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 22. | Lieblich SM, Castle DJ, Pantelis C, Hopwood M, Young AH, Everall IP. High heterogeneity and low reliability in the diagnosis of major depression will impair the development of new drugs. BJPsych Open. 2015;1:e5-e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 30] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 23. | Regier DA, Kuhl EA, Kupfer DJ. The DSM-5: Classification and criteria changes. World Psychiatry. 2013;12:92-98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 362] [Cited by in RCA: 528] [Article Influence: 44.0] [Reference Citation Analysis (0)] |
| 24. | Hyman SE. The diagnosis of mental disorders: the problem of reification. Annu Rev Clin Psychol. 2010;6:155-179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 530] [Cited by in RCA: 495] [Article Influence: 33.0] [Reference Citation Analysis (0)] |
| 25. | Hyman SE. Diagnosing the DSM: Diagnostic Classification Needs Fundamental Reform. Cerebrum. 2011;2011:6. [PubMed] |
| 26. | Insel T, Cuthbert B, Garvey M, Heinssen R, Pine DS, Quinn K, Sanislow C, Wang P. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am J Psychiatry. 2010;167:748-751. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4107] [Cited by in RCA: 4317] [Article Influence: 287.8] [Reference Citation Analysis (0)] |
| 27. | Churchland PS. Neurophilosophy: Toward a unified science of the mind-brain. A Bradford Book (Reprint Edition), 1989: 1-560. |
| 28. | Kendler KS. Toward a philosophical structure for psychiatry. Am J Psychiatry. 2005;162:433-440. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 286] [Cited by in RCA: 231] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 29. | Maes M, Nowak G, Caso JR, Leza JC, Song C, Kubera M, Klein H, Galecki P, Noto C, Glaab E, Balling R, Berk M. Toward Omics-Based, Systems Biomedicine, and Path and Drug Discovery Methodologies for Depression-Inflammation Research. Mol Neurobiol. 2016;53:2927-2935. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 36] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 30. | Maes M, Rief W. Diagnostic classifications in depression and somatization should include biomarkers, such as disorders in the tryptophan catabolite (TRYCAT) pathway. Psychiatry Res. 2012;196:243-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 63] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 31. | National Research Council (US) Committee on A Framework for Developing a New Taxonomy of Disease. 2011. [PubMed] [DOI] [Full Text] |
| 32. | Fernandes BS, Williams LM, Steiner J, Leboyer M, Carvalho AF, Berk M. The new field of 'precision psychiatry'. BMC Med. 2017;15:80. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 310] [Cited by in RCA: 312] [Article Influence: 39.0] [Reference Citation Analysis (0)] |
| 33. | Maes M, Moraes JB, Congio A, Bonifacio KL, Barbosa DS, Vargas HO, Michelin AP, Carvalho AF, Nunes SOV. Development of a Novel Staging Model for Affective Disorders Using Partial Least Squares Bootstrapping: Effects of Lipid-Associated Antioxidant Defenses and Neuro-Oxidative Stress. Mol Neurobiol. 2019;56:6626-6644. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 43] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 34. | Popper K. On the Structure of Scientific Revolution, Chicago University Press, Chicago, IL, 1962. |
| 35. | Kanchanatawan B, Sriswasdi S, Maes M. Supervised machine learning to decipher the complex associations between neuro-immune biomarkers and quality of life in schizophrenia. Metab Brain Dis. 2019;34:267-282. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 23] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 36. | Al-Hakeim HK, Almulla AF, Al-Dujaili AH, Maes M. Construction of a Neuro-Immune-Cognitive Pathway-Phenotype Underpinning the Phenome of Deficit Schizophrenia. Curr Top Med Chem. 2020;20:747-758. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 21] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 37. | Ringle CM, da Silva D, Bido D. Structural equation modeling with the SmartPLS. Brazilian Journal of Marketing - BJM Revista Brasileira de Marketing – ReMark Edição Especial 2014; 13: n2. [DOI] [Full Text] |
| 38. | Marcoulides GA, Saunders C. PLS: A Silver Bullet? MIS Quarterly, 2006; 30: 3-9. |
| 39. | Hair Jr JF, Hult GTM, Ringle C, Sarstedt, M. A primer on partial least squares structural equation modeling (PLS-SEM). Sage Publications, 2013. |
| 40. | Lewine RR, Gulley LR, Risch SC, Jewart R, Houpt JL. Sexual dimorphism, brain morphology, and schizophrenia. Schizophr Bull. 1990;16:195-203. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 39] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 41. | Seney ML, Sibille E. Sex differences in mood disorders: perspectives from humans and rodent models. Biol Sex Differ. 2014;5:17. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 85] [Cited by in RCA: 100] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 42. | Cone JD. Idiographic, nomothetic, and related perspectives in behavioral assessment. In: Nelson RO, Hayes SC. Conceptual Foundations of Behavioral Assessment. New York, Guilford Press, 1986: 111-128. [RCA] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 43. | Ross C, Pam A. Pseudoscience in Biological Psychiatry: Blaming the Body. Wiley, New York, 1995: 1-294. |
| 44. | Modrow J. How to Become a Schizophrenic: The Case Against Biological Psychiatry. Apollyon Press (a revised and expanded 2003 edition published by Writers Club Press is on print).1996: 239. |