Published online Jun 7, 2020. doi: 10.3748/wjg.v26.i21.2702
Peer-review started: January 21, 2020
First decision: March 15, 2020
Revised: March 26, 2020
Accepted: May 13, 2020
Article in press: May 13, 2020
Published online: June 7, 2020
Processing time: 136 Days and 19.5 Hours
Inflammatory bowel disease (IBD) is an inflammatory disorder of the gastrointestinal tract that affects millions of patients worldwide. It has a complex and multifactorial etiology leading to excessive exposure of intestinal epithelium to microbial antigens, inappropriate activation of the immune system and ultimately to the damage of intestinal tissues. Although numerous efforts have been made to improve the disease management, IBD remains persistently recurring and beyond cure. This is due largely to the gaps in our understanding of the pathogenesis of IBD that hamper the development of timely diagnoses and effective treatment. However, some recent discoveries, including the beneficial effects of interleukin-22 (IL-22) on the inflamed intestine, have shed light on a self-protective mechanism in IBD. Regenerating islet-derived (REG/Reg) proteins are small secretory proteins which function as IL-22’s downstream effectors. Mounting studies have demonstrated that IBD patients have significantly increased REG expressions in the injured intestine, but with undefined mechanisms and roles. The reported functions of REG/Reg proteins in intestinal homeostasis, such as those of antibacterial, anti-inflammatory and tissue repair, lead us to discuss their potential mechanisms and clinical relevance in IBD in order to advance IBD research and management.
Core tip: The clinical management of inflammatory bowel disease (IBD) remains a significant challenge due to the knowledge gap in its pathogenesis. In this review paper, we have discussed the literature regarding increased expressions of regenerating islet-derived proteins in IBD and proposed the potential clinical relevance of these proteins based on their known protective activities in the inflamed intestine. We therefore provide insight from a new perspective in order to advance IBD research and clinical management.
- Citation: Edwards JA, Tan N, Toussaint N, Ou P, Mueller C, Stanek A, Zinsou V, Roudnitsky S, Sagal M, Dresner L, Schwartzman A, Huan C. Role of regenerating islet-derived proteins in inflammatory bowel disease. World J Gastroenterol 2020; 26(21): 2702-2714
- URL: https://www.wjgnet.com/1007-9327/full/v26/i21/2702.htm
- DOI: https://dx.doi.org/10.3748/wjg.v26.i21.2702
Regenerating islet-derived (REG/Reg) proteins are small secretory C-type-like lectins. In 1979, De Caro et al[1] found that pancreatic stones in patients with chronic calcifying pancreatitis were made largely of a type of proteins. This pancreatic stone protein was later independently reported as pancreatic thread protein by Gross et al[2] in 1985, and as regenerating islet-derived protein by Terazono et al[3] in 1988. After the recognition of its different isoforms and the corresponding homologs across species in mammals, it was given the current name REG1α in a sequence and structure-based classification of the family proteins[4,5]. The tissue distributions and physiological activities of this protein family are studied primarily in humans and mice. The human REG family consists of REG1α, REG1β, REG3α, REG3γ and REG4, and these family members share between 30 to 80 percent homologous sequences. In mice, the family members of Reg1, Reg2, Reg3α, Reg3β, Reg3γ, Reg3δ and Reg4 share more than 50 percent homologous sequences, and are structurally and functionally conserved with their human counterparts[5]. Studies of REG/Reg proteins have focused on their functions in the pancreas and intestine since REG/Reg proteins are predominantly expressed in these two digestive organs. We and others have shown that in acute pancreatitis, the production of REG/Reg proteins is significantly induced in pancreatic acinar cells to inhibit inflammatory cell infiltration and to promote tissue repair[6-11]. In the inflamed intestine, REG/Reg expression is highly upregulated in different crypt cells including Paneth cells, deep secreting cells and enteroendocrine cells, and extend to other intestinal epithelial cells depending on the isoform[12-16]. Although the precise roles of REG proteins in human inflammatory bowel disease (IBD) have not been defined, mouse studies have confirmed that Reg proteins are required to maintain intestinal homeostasis in both physiological and pathological conditions.
Ose et al[13] reported that Reg1 deficient mice had reduced stem cell proliferation in the crypts and impaired epithelial migration along the villi in the small intestine, revealing the role of Reg1 in the preservation and renewal of intestinal villous structure. In line with these findings, Sun et al[17] showed that Reg1 protein protected against intestinal damage caused by indomethacin, a nonsteroidal anti-inflammatory drug. In the same disease model, Kitayama et al[18] found that Reg1 deficiency severely attenuated the expression of tight junction proteins including claudins 3 and 4. Therefore, in addition to promoting the recovery of villous structure, Reg1 may enhance the integrity of the epithelial barrier during intestinal injury.
Like Reg1 protein, Reg3 proteins also have a protective role in injured intestine. Studies showed that Reg3β and Reg3γ could promote the growth of cultured colonic epithelial cells[19], and their intestinal expression was highly upregulated in dextran sulfate sodium (DSS)- or pathogenic bacteria-induced mouse colitis[12,20,21]. Reg3β deficiency consistently exacerbated symptoms of colitis in DSS-treated mice[22]. Similarly, in a mouse model of graft-vs-host disease, intestinal expressions of Reg3α and Reg3γ were upregulated in response to IL-22 signaling to enhance the survival of intestinal stem cells and Paneth cells[23,24]. Furthermore, compensation of Reg3γ deficiencies in IL-22 deficient mice by intraperitoneal injections of REG3γ or Reg3γ blocked the intestinal epithelial invasion of orally introduced C. rodentium and saved the animals’ lives[25].
Notably, in addition to Reg3 proteins’ trophic effect on the epithelial barrier, the contribution of their direct inhibitory effect on bacterial invasiveness cannot be excluded[26,27]. Cash et al[28] showed that Reg3γ and its human counterpart REG3α are antimicrobial proteins that bind to the peptidoglycan carbohydrate on Gram-positive bacteria. Further studies showed that the interaction was mediated by a Glu-Pro-Asn motif in the long loop region of REG3α[29]. Upon interacting with peptidoglycan, REG3α oligomerizes to form hexameric transmembrane pores that kill bacteria by increasing their membrane permeability[30]. Consistently, Reg3γ deficient mice have more Gram-positive bacteria in the intestinal mucosa[31]. In contrast to REG3α/Reg3γ, Reg3β may attach to lipopolysaccharide to cause osmotic rupture in Gram-negative bacteria[26,27,32-34], in line with the increased proportion of intestinal Gram-negative bacteria in Reg3β deficient mice[35].
Unlike the expressions of Reg1/3 proteins in both the small and large intestines, Reg4 proteins are distributed predominantly in the colon. It has been shown that Reg4 is produced by deep crypt secretory cells and intestinal enteroendocrine cells, and expands to epithelial cells of the upper colonic crypts during inflammation[15,16].
IBD comprises two major disorders: Crohn’s disease (CD) and ulcerative colitis (UC). CD and UC share the features of chronic and destructive mucosal inflammation, but they are pathogenically and clinically distinct. CD patients have abdominal cramps, pain, diarrhea and weight loss, and their intestinal injury is characterized by transmural and segmental lesions that may occur at any level of the gastrointestinal tract “from mouth to anus”. UC patients frequently present with bloody stools and diarrhea, and their intestinal inflammation is localized to colonic mucosa. However, the mechanisms underlying the different pathogenesis in CD and UC remain poorly understood[36,37].
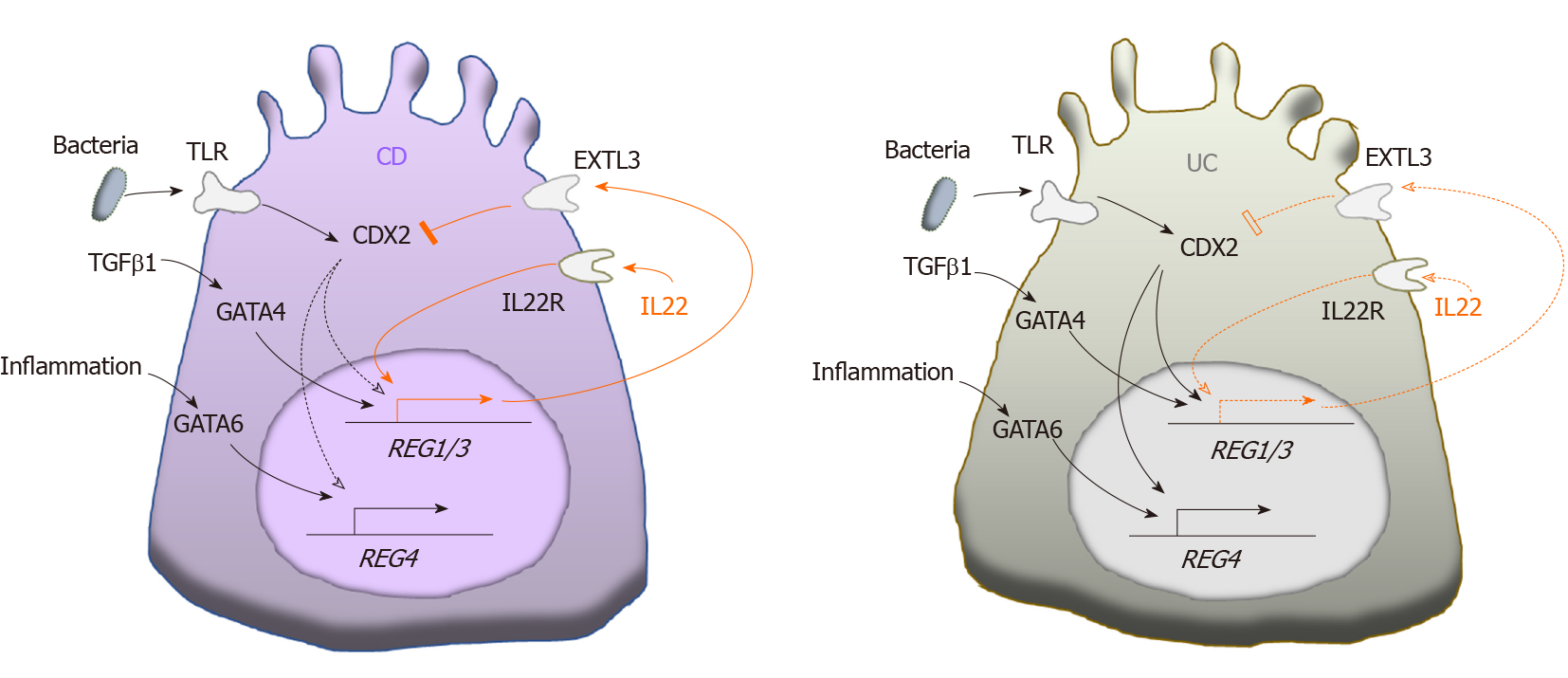
Over the past decade, increased REG expression has been detected by various techniques in both CD and UC patients[12,14,38-52] (Table 1). Interestingly, the increased levels of different REG isoforms vary significantly in CD and UC, and in active and remissive phases of each disease, which could be attributed to the different transcriptional regulations of REG isoforms by inflammatory cytokines such as IL22 and TGFβ illustrated in Figure 1. CD and UC are characterized by different cytokine responses. In CD, the T-helper type 1 (Th1) cytokine interferon-γ and the Th17 cytokines IL-17/IL-22 are the main inflammation regulators. In contrast, UC has a Th2-like cytokine response that activates IL-13/IL-5, producing natural killer T cells[53]. These distinct cytokine patterns are associated with relatively high IL-22 levels in patients with CD compared to those with UC[54-56]. Animal studies have shown comparable results. For example, in the Th17 response-mediated mouse model of CD generated by the transfer of CD45RBhigh T cells, IL-22 levels are high; but in the Th2 response-regulated UC model of T cell receptor α-chain deficient mice, IL-22 levels are low[56]. A recent study by Leung et al[57] provided an explanation. They showed that in active UC, a Th22 subset of helper T cells that produces IL-22, but not IL-17, was depleted by increased TGFβ in patients’ colonic tissues[57].
| Ref. | REGs | Samples (size) | Relevant findings |
| Lawrance et al[38], 2001 | REG1α/1β/3α | CD (6) UC (6) control (6) | Increased intestinal REG1α/1β/3α in IBD detected by microarray |
| Shinozaki et al[39], 2001 | REG1α | CD (9) UC (21) control (5 non-IBD, 6 normal) | Increased intestinal REG1α in IBD detected by RT-PCR and ISH |
| Desjeux et al[40], 2002 | REG3α | CD (124) normal control (54) | Increased serum REG3α in active CD detected by ELISA |
| Dieckgraefe et al[41], 2002 | REG1α/1β/3γ | CD (3) UC (5) control (4) | Increased intestinal REG1α/1β/3γ in IBD detected by microarray and IHC |
| Ogawa et al[12], 2003 | REG3α | CD (20) UC (23) control (18) | Increased intestinal REG3α in IBD detected by ISH and Northern blot |
| Kämäräinen et al[42], 2003 | REG4 | CD (N/A) UC (N/A) | By ISH and IHC, REG4 constitutively expressed in neuroendocrine cells, and upregulated in inflamed epithelial cells |
| Gironella et al[43], 2005 | REG3α | IBD (171) control (14 non-IBD, 29 normal) | Increased serum REG3α correlated with IBD severity detected by ELISA. Higher REG3α in CD than UC. REG3α localized to colonic Paneth cells |
| Wu et al[44], 2007 | REG1β/REG3α | CD (9) UC (5) control (4) | Increased intestinal REG1β in CD and REG3α in CD and UC detected by microarray |
| Nanakin et al[45], 2007 | REG4 | UC (22) normal control (5) | Increased intestinal REG4 in UC detected by RT-PCR, ISH and IHC |
| Sekikawa et al[46], 2010 | REG1α | UC (60) control (10) | Increased intestinal REG1α in UC detected by RT-PCR and IHC |
| Tanaka et al[47], 2011 | REG1α | UC (31) control (5) | Increased intestinal REG1α in UC detected by IHC |
| Granlund et al[14], 2011 | REG1α/1β/3α/4 | CD/control (12/21) UC/control (32/34) | Increased intestinal REG1α/1β/3α/4 in IBD detected by microarray. Different cellular localizations of REG1α and REG4 detected by IHC |
| van Beelen Granlund et al[48], 2013 | REG1α/1β/3α/4 | CD (N/A) UC (N/A) | By ISH, REG1α/1β/3α localized to Paneth cells in the crypt base, REG4 localized to enteroendocrine cells towards the luminal face |
| Planell et al[49], 2013 | REG1α/4 | Microarray: UC (15 active/8 remissive), Non-IBD (13); RT-PCR: UC (8 active/12 remissive), non-IBD (10) | Comparably increased intestinal REG4 in active and remissive UC, and significantly increased REG1α in active UC but not in remissive UC, detected by microarray and RT-PCR |
| Marafini et al[50], 2014 | REG3α | CD (72) UC (22) | Infliximab treatment decreased the high serum REG3α in CD and UC |
| Nunes et al[51], 2014 | REG3α | CD (66) UC (74) | Increased serum REG3α serum in active CD but not UC detected by ELISA |
| Tsuchida et al[52], 2017 | REG1α/1β/3α/4 | CD (49) UC (39) control (44) | Increased intestinal REG1α/1β/4 in CD, and REG4 in UC detected by RT-PCR |
Tsuchida et al[52] showed that REG1α, REG1β, and REG4 were all highly overexpressed in CD, while only REG4 was significantly overexpressed in UC. This could be caused by the limited IL-22 production in the colon because IL-22 signaling is essential for transcriptional activation of REG1/3 genes[25,46,52,58], while REG4 transcription could be driven by the mechanisms other than IL-22 signaling[52]. For example, REG4 transcription is known to be activated by caudal type homeobox 2 (CDX2), a transcription factor that regulates multiple genes for maintaining intestinal homeostasis in response to the Toll-like receptor signaling[59-62]. Interestingly in this regard, CDX2 expression has been found to be inhibited by phosphoinositide 3-kinase[63], which can be activated by exostosin-like glycosyltransferase 3 (EXTL3), a cell surface enzyme expressed in multiple organs including the intestine[64,65]. Since EXTL3 has been identified as a receptor for REG1/Reg1 and REG3/Reg3 proteins[64-67], it is possible that as illustrated in Figure 1, CDX2-activated colonic REG4 transcription is inhibited in CD, but activated in UC, due to higher REG1/3 levels in CD than UC. In addition, GATA binding proteins (GATAs) could possibly regulate REG genes transcription in an IL-22 independent manner. Studies have shown that Reg1/3 transcriptions can be activated by GATA4, which is normally expressed in the proximal small intestine[68,69], but abnormally expressed in the inflammatory lesions of the distal small intestine and colon[70]. On the other hand, REG4 transcription is specifically activated by GATA6[52,71], which is expressed in both the small and large intestines[68]. Deficiency of GATA4 or GATA6 causes abnormal alterations in intestinal cells including Paneth cells, enteroendocrine and goblet cells[72,73]. Both GATA4- and GATA6-regulated REG transcriptions could be mediated by inflammation in IBD. Haveri et al[70] suggested the upregulation of GATA4 in inflamed intestine by activated signaling of TGFβ, whose expression is elevated in active but not remissive CD and UC[74,75]. This explains the significant increase of intestinal REG1α in active phase but not remissive phase of UC, which is in contrast to the disease status-independent increase of REG4 in UC[49]. As discussed before, REG4 transcription could be activated by CDX2 in UC when IL-22 is diminished. In addition, the activation of GATA6 by inflammation also possibly contributes to REG4 expression in UC (Figure 1). In support of this view, Mustfa et al[76] showed that in IBD, inflammation globally decreased SUMOylation, a post-translational modification that inhibits GATA6 transcription, in colonic cells[77].
Of note, other factors, such as exclusive enteral nutrition diet commonly recommended for CD patients, influence the bacteria population, inflammation and mucosal healing in IBD[78]. Therefore, they could also have a direct and/or indirect regulatory effect on intestinal REG expressions, which should be investigated in future studies.
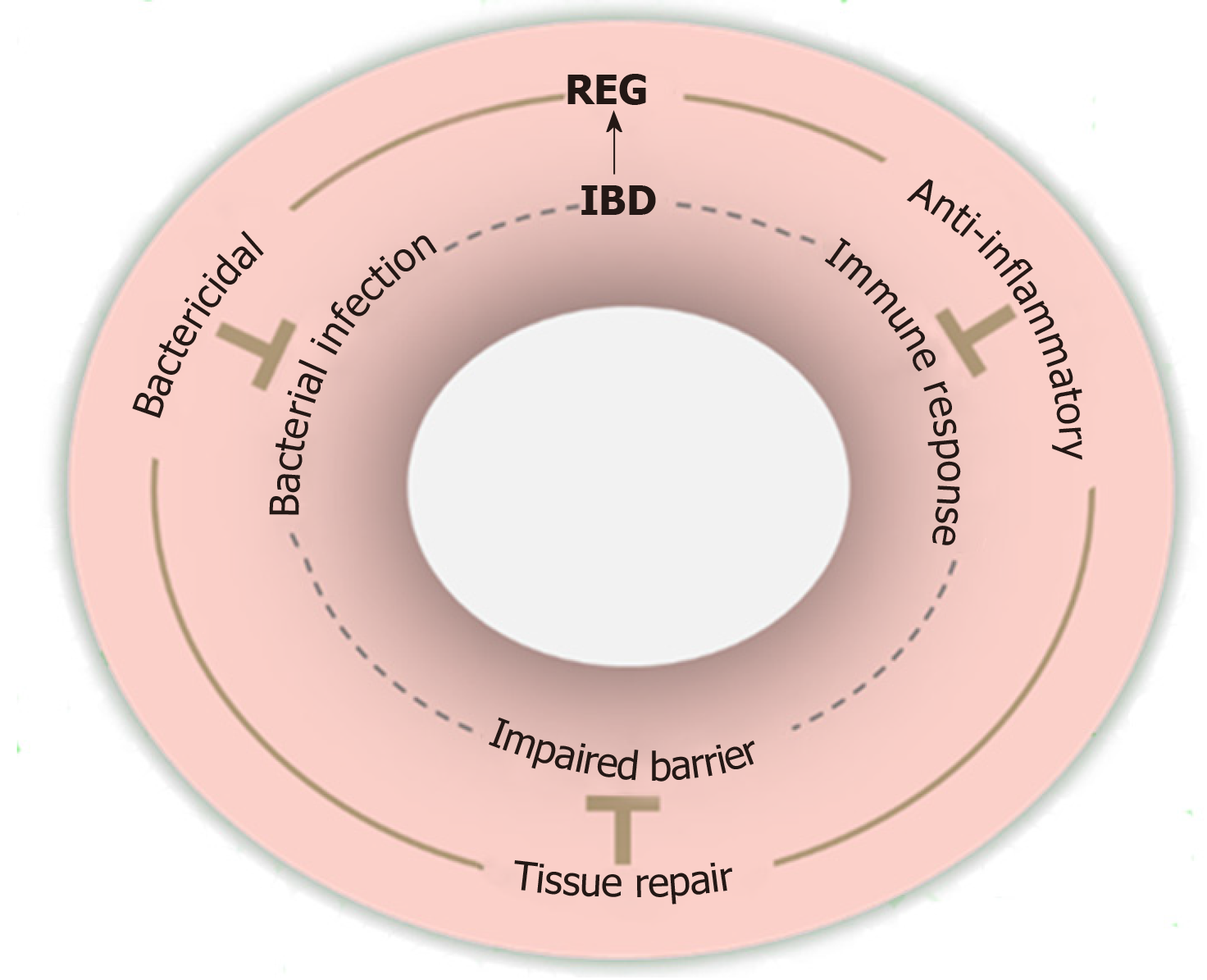
Studies have supported that IBD progression is driven by defective bacterial clearance, aberrant immune responses, and impaired epithelial barrier[37]. Notably, REG proteins appear to have the corresponding protective effects that can counteract these defects in IBD as illustrated in Figure 2.
In line with REG/Reg proteins’ bactericidal activities, mice carrying genetically modified REG/Reg genes have altered compositions of intestinal bacterial microbiota[16,31,35,79,80] (Table 2), indicating the regulatory roles played by REG/Reg proteins in the gut microbiome. Notable among these are that mice with REG3α overexpression and mice with Reg4 deficiency were resistant to DSS-induced colitis, suggesting the importance of REG/Reg-regulated intestinal microbiota in IBD pathogenesis[16,79]. Furthermore, mice with Reg3γ overexpression had an enriched fraction of beneficial Lactobacilli in the gut microbiome, suggesting the positive selection of “good bacteria” by Reg3γ[80]. In support of the importance of an altered intestinal microbiome in IBD pathogenesis, a systematic review and meta-analysis of CD and UC patients showed different intestinal microbiome compositions in patients with active disease as compared to those in remission[81]. Given the fact that patients with active IBD have higher levels of REG proteins, further studies are needed to define the significance between increased REG expression and altered microbiome composition observed in these patients.
| Class | Order | Family | Genera |
| Actinobacteria | Bifidobacteriales | Bifidobacteriaceae | Bifidobacterium ↑2 |
| Coriobacteriales | Coriobacteriaceae ↓1↓3 | ||
| Eggerthellales | Eggerthellaceae | Enterorhabdus ↓3 | |
| Alphaproteobacteria | Caulobacterales | Caulobacteraceae | Brevundimonas ↓3 |
| Rhizobiales | Bartonellaceae | Bartonella ↑3 | |
| Bacilli | Bacillales | Bacillaceae | Oceanobacillus ↑3 |
| Staphylococcaceae | Staphylococcus ↓2 | ||
| Lactococcus ↑3 | |||
| N/A | Gemella ↑3 | ||
| Lactobacillales ↑3 | Lactobacillaceae −1 | Lactobacillus ↓2↑3−4↓5 | |
| Enterococcaceae | Enterococcus ↑2↑3 | ||
| Aerococcaceae | Facklamia ↓3 | ||
| Carnobacteriaceae | Carnobacterium ↑3 | ||
| Bacteroidia | Bacteroidales ↑1↑3↓5 | Prevotellaceae ↓1↑3 | Prevotella ↓1↑2↑3 |
| Rikenellaceae −1 | |||
| Porphyromonadaceae ↓1↓5 | Parabacteroides ↓1↑3↑5 | ||
| Barnesiella ↓1 | |||
| Bacteroidaceae ↓1 | Bacteroides ↓1−2−3−4↑5 | ||
| Betaproteobacteria | Burkholderiales | Sutterellaceae ↓1 | Parasutterella ↑1↓3 |
| Clostridia ↑1 | Clostridiales ↓5 | Lachnospiraceae ↑1↑3−5 | Roseburia ↑3 |
| Ruminococcaceae −2↑3−5 | Faecalibacterium ↑3 | ||
| Clostridiaceae | Clostridium ↑2↑3(XI Cluster) ↓5(XIVa Cluster) ↑5 | ||
| Candidatus Arthromitus ↑3 | |||
| Candidatus Savagella ↑4 | |||
| Eubacteriaceae | Eubacterium (rectale) ↑4 | ||
| Oscillospiraceae | Oscillibacter ↑1 | ||
| Peptococcaceae ↑3 | |||
| Delta Proteobacteria | Desulfovibrionales ↑1 | Desulfovibrionaceae −5 | Lawsonia −5 |
| Desulfovibrio −5 | |||
| Epsilonproteobacteria | Campylobacterales | Helicobacteraceae | Helicobacter ↑2↑3 |
| Erysipelotrichia | Erysipelotrichales | Erysipelotrichaceae ↑3↑5 | Turicibacter ↑3 |
| Allobaculum −3↓5 | |||
| Gammaproteobacteria | Enterobacterales | Enterobacteriaceae ↑3 | Escherichia ↓2 |
| Enterobacter ↓1 | |||
| Morganellaceae | Proteus ↓2 | ||
| Pseudomonadales | Moraxellaceae | Psychrobacter ↑3 | |
| Acinetobacter ↑3 | |||
| Xanthomonadales | Xanthomonadaceae | Stenotrophomonas ↑3 | |
| Negativicutes | Acidaminococcales | Acidaminococcaceae | Phascolarctobacterium ↑3 |
| Veillonellales | Veillonellaceae ↑3 | ||
| Verrucomicrobiae | Verrucomicrobiales | Akkermansiaceae | Akkermansia ↓1↑3↑5 |
| Verrucomicrobiaceae ↓1 |
In addition to their bactericidal activities, REG3/Reg3 proteins are anti-inflammatory since they can inhibit proinflammatory cytokine secretion, inflammatory cell activation and infiltration in inflammatory diseases including IBD[8-10,12,82-84]. REG3α incubation inhibited the proinflammatory cytokine secretion in intestinal mucosa harvested from patients with active CD in a dose dependent manner, and decreased the adhesive molecules, such as E-selectin, ICAM-1 and VCAM-1, which were found to be upregulated on endothelial cells to promote inflammatory infiltration[12]. Additionally, REG3/Reg3 proteins regulate the activities of macrophages, which regulate inflammatory injury in IBD[84-87]. It is thus possible that REG3/Reg3 proteins may also alleviate IBD inflammation via macrophages.
As previously mentioned, REG/Reg proteins have trophic effects on intestinal epithelium in both physiological and pathological conditions. These trophic effects have been attributed to the activation of pro-survival and pro-proliferative signaling pathways, such as MEK1/2, ERK1/2, phosphoinositide 3-kinase-Akt and JAK2-STAT3[5]. Therefore, the direct tissue protection and repair of mucosa in IBD by REG/Reg proteins cannot be excluded. Indeed, transgenic overexpression of REG3α or intrarectal administration of REG3α alleviated the epithelial damage in 2,4,6-trinitrobenzene sulphonic acid-induced mouse colitis[79], while Reg3β deficiency worsen DSS colitis in mice[22]. Similarly, administration of REG3γ /Reg3γ improved epithelial integrity in C. rodentium-induced mouse colitis[25]. Additionally, Reg4+ deep crypt secretory cells promoted the formation of organoids derived from Lgr5+ colonic stem cells, and Reg4 stimulated the growth of colonic organoids isolated from mice with DSS-induced colitis[15,16].
Despite the recent advancements, there are unmet needs in current IBD management, particularly including disease activity detection and treatment[88,89]. REG proteins have been considered as potential diagnostic markers and/or therapeutic targets for immune-mediated diseases[90]. The recognized upregulation of REG proteins in IBD and REG proteins’ beneficial activities provide unique opportunities to address some of these unmet needs for improving IBD management, such as serving as biomarkers for disease activity, shifting the composition of bacterial microbiota, and enhancing the repair of intestinal epithelium.
The clinical diagnosis of IBD remission/relapse depends on the endoscopic biopsy, which has its own limitations including invasiveness, financial burden and inter-user variability. Non-invasive imaging such as ultrasound, and CT and MR enterography are useful modalities but also have drawbacks such as inter-operator variability, radiation exposure, and financial burden[91,92]. Therefore, efforts have been made to study potential IBD biomarkers in serum and feces including C-reactive protein (CRP), anti-Saccharomyces cerevisiae antibody, which is chiefly linked with CD, and perinuclear antineutrophil cytoplasmic antibody, which is linked with UC[93-96]. However, these biomarkers have high specificities but limited sensitivities. Of note, inhibition of tumor necrosis factor alpha by Infliximab reduced serum REG3α levels in CD and UC patients[50], supporting the potential use of REG3α for evaluating the response of treatment. Furthermore, a multicenter prospective study showed increased serum REG3α with 94% specificity and 60% sensitivity for active CD[38]. Similarly, a recent study showed increased serum REG3α in CD patients 3 mo prior to relapse, but with only 73% specificity and 50% sensitivity[51]. The authors explained that the lower specificity and sensitivity in their study compared with those from the previously mentioned report was likely due to the patients’ mild disease activity and low relapse rate. Given the functional redundancy among REG family members and confirmed upregulated REG1α/β in CD patients (Table 1), it is possible that CD patients with lower REG3α levels may have higher serum REG1α/β levels. If so, combining REG3α and REG1α/β measurements in each CD patient could collectively improve the sensitivity for evaluating the relapse. On the other hand, this study showed that serum REG3α levels were not correlative with UC activity[51]. This could be due to the relatively attenuated upregulation of REG3 in UC (Figure 1). Therefore, it would be more informative to assess the use of REG4 as a biomarker for UC, since it is more specific and highly upregulated in UC[45,49,52].
Multiple clinical trials have shown that fecal microbiota transplantation is a promising treatment to induce remission in active UC[97-100]. However, the specific bacteria that protect against UC have not been identified. Studying altered intestinal bacteria populations in DSS colitis resistant REG3α transgenic mice and Reg4 deficient mice therefore could help to identify the protective bacteria and associated REG/Reg regulation in UC [16,79].
Bowel resection is considered for the patients that are refractory to medical therapy or with serious complications of the medications[101-103]. However, in addition to the risk of short bowel syndrome, septic complications are commonly associated with anastomotic leaks[103]. The early diagnosis of anastomotic leaks enables a timelier intervention essential for a better outcome. It has been shown that a low CRP on postoperative day 4 is a reliable biomarker for excluding postoperative infectious complications in abdominal surgery, while high CRP levels should prompt aggressive imaging for possible anastomotic failure[104]. Like CRP, REG/Reg proteins have been considered to be acute phase proteins that could be used as markers of septic complications in patients including those undergoing abdominal surgery[105-110]. Therefore, determining whether REG proteins can serve as a more sensitive and specific sensor of postoperative anastomotic leak could have significant clinical potential.
It is also worthy of note that Reg proteins do not lead to immune suppression or immunogenicity, the side-effects that are associated with risks of infection and immune dysregulation in some agents used in IBD treatment[88,89]. Given that REG proteins have bactericidal, anti-inflammatory and tissue repair functions in the inflamed intestine (Figure 2), the use of REG proteins as an adjunct to reduce the doses of current medications for IBD could potentially minimize complications. Despite these benefits, however, concerns of long-term application of REG proteins remain. For example, REG/Reg proteins may potentially overactivate the oncogenic STAT3 signaling pathway[5,27], even though the gastrointestinal tract administration may decrease the oncogenic risk in other organs. Additionally, CD patients are prone to bowel stricturing/stenosis formation[111]. Based on our finding of Reg1 as an activator of stellate cells, the predominant producer of collagen in pancreatitis[67], the potential of REG1 for bowel stricturing/stenosis should be clarified, which could argue against its use in patients with CD.
IBD remains a significantly challenging disorder due to as yet unresolved issues in its pathogenesis, diagnosis and clinical management. In this review, by discussing the expressions and activities of REG proteins in the inflamed intestine, we have attempted to illuminate potential applications of these REG proteins that may help to improve detection and treatment of the disease, but further comprehensive studies are necessary to clarify and confirm these benefits in IBD.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: United States
Peer-review report’s scientific quality classification
Grade A (Excellent): A
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): D
Grade E (Poor): 0
P-Reviewer: Huang Z, Sitkin S, Skrautvol K S-Editor: Dou Y L-Editor: A E-Editor: Ma YJ
| 1. | De Caro A, Lohse J, Sarles H. Characterization of a protein isolated from pancreatic calculi of men suffering from chronic calcifying pancreatitis. Biochem Biophys Res Commun. 1979;87:1176-1182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 113] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 2. | Gross J, Carlson RI, Brauer AW, Margolies MN, Warshaw AL, Wands JR. Isolation, characterization, and distribution of an unusual pancreatic human secretory protein. J Clin Invest. 1985;76:2115-2126. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 94] [Cited by in RCA: 95] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 3. | Terazono K, Yamamoto H, Takasawa S, Shiga K, Yonemura Y, Tochino Y, Okamoto H. A novel gene activated in regenerating islets. J Biol Chem. 1988;263:2111-2114. [PubMed] |
| 4. | Parikh A, Stephan AF, Tzanakakis ES. Regenerating proteins and their expression, regulation and signaling. Biomol Concepts. 2012;3:57-70. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 81] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 5. | Chen Z, Downing S, Tzanakakis ES. Four Decades After the Discovery of Regenerating Islet-Derived (Reg) Proteins: Current Understanding and Challenges. Front Cell Dev Biol. 2019;7:235. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 63] [Article Influence: 10.5] [Reference Citation Analysis (1)] |
| 6. | Orelle B, Keim V, Masciotra L, Dagorn JC, Iovanna JL. Human pancreatitis-associated protein. Messenger RNA cloning and expression in pancreatic diseases. J Clin Invest. 1992;90:2284-2291. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 94] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 7. | Kemppainen E, Sand J, Puolakkainen P, Laine S, Hedström J, Sainio V, Haapiainen R, Nordback I. Pancreatitis associated protein as an early marker of acute pancreatitis. Gut. 1996;39:675-678. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 24] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 8. | Zhang H, Kandil E, Lin YY, Levi G, Zenilman ME. Targeted inhibition of gene expression of pancreatitis-associated proteins exacerbates the severity of acute pancreatitis in rats. Scand J Gastroenterol. 2004;39:870-881. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 46] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 9. | Gironella M, Folch-Puy E, LeGoffic A, Garcia S, Christa L, Smith A, Tebar L, Hunt SP, Bayne R, Smith AJ, Dagorn JC, Closa D, Iovanna JL. Experimental acute pancreatitis in PAP/HIP knock-out mice. Gut. 2007;56:1091-1097. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 67] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 10. | Lin YY, Viterbo D, Mueller CM, Stanek AE, Smith-Norowitz T, Drew H, Wadgaonkar R, Zenilman ME, Bluth MH. Small-interference RNA gene knockdown of pancreatitis-associated proteins in rat acute pancreatitis. Pancreas. 2008;36:402-410. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 11. | Hassanain E, Huan C, Mueller CM, Stanek A, Quan W, Viterbo D, Bluth MH, Zenilman ME. Pancreatitis-associated proteins' regulation of inflammation is correlated with their ability to aggregate. Pancreas. 2011;40:1151-1153. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 12. | Ogawa H, Fukushima K, Naito H, Funayama Y, Unno M, Takahashi K, Kitayama T, Matsuno S, Ohtani H, Takasawa S, Okamoto H, Sasaki I. Increased expression of HIP/PAP and regenerating gene III in human inflammatory bowel disease and a murine bacterial reconstitution model. Inflamm Bowel Dis. 2003;9:162-170. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 121] [Cited by in RCA: 133] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 13. | Ose T, Kadowaki Y, Fukuhara H, Kazumori H, Ishihara S, Udagawa J, Otani H, Takasawa S, Okamoto H, Kinoshita Y. Reg I-knockout mice reveal its role in regulation of cell growth that is required in generation and maintenance of the villous structure of small intestine. Oncogene. 2007;26:349-359. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 46] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 14. | Granlund Av, Beisvag V, Torp SH, Flatberg A, Kleveland PM, Ostvik AE, Waldum HL, Sandvik AK. Activation of REG family proteins in colitis. Scand J Gastroenterol. 2011;46:1316-1323. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 42] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 15. | Sasaki N, Sachs N, Wiebrands K, Ellenbroek SI, Fumagalli A, Lyubimova A, Begthel H, van den Born M, van Es JH, Karthaus WR, Li VS, López-Iglesias C, Peters PJ, van Rheenen J, van Oudenaarden A, Clevers H. Reg4+ deep crypt secretory cells function as epithelial niche for Lgr5+ stem cells in colon. Proc Natl Acad Sci USA. 2016;113:E5399-E5407. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 221] [Article Influence: 24.6] [Reference Citation Analysis (0)] |
| 16. | Xiao Y, Lu Y, Wang Y, Yan W, Cai W. Deficiency in intestinal epithelial Reg4 ameliorates intestinal inflammation and alters the colonic bacterial composition. Mucosal Immunol. 2019;12:919-929. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 10] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 17. | Sun C, Fukui H, Hara K, Kitayama Y, Eda H, Yang M, Yamagishi H, Tomita T, Oshima T, Watari J, Takasawa S, Chiba T, Miwa H. Expression of Reg family genes in the gastrointestinal tract of mice treated with indomethacin. Am J Physiol Gastrointest Liver Physiol. 2015;308:G736-G744. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 21] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 18. | Kitayama Y, Fukui H, Hara K, Eda H, Kodani M, Yang M, Sun C, Yamagishi H, Tomita T, Oshima T, Watari J, Takasawa S, Miwa H. Role of regenerating gene I in claudin expression and barrier function in the small intestine. Transl Res. 2016;173:92-100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 17] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 19. | Moucadel V, Soubeyran P, Vasseur S, Dusetti NJ, Dagorn JC, Iovanna JL. Cdx1 promotes cellular growth of epithelial intestinal cells through induction of the secretory protein PAP I. Eur J Cell Biol. 2001;80:156-163. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 39] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 20. | Pull SL, Doherty JM, Mills JC, Gordon JI, Stappenbeck TS. Activated macrophages are an adaptive element of the colonic epithelial progenitor niche necessary for regenerative responses to injury. Proc Natl Acad Sci USA. 2005;102:99-104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 466] [Cited by in RCA: 482] [Article Influence: 24.1] [Reference Citation Analysis (0)] |
| 21. | Keilbaugh SA, Shin ME, Banchereau RF, McVay LD, Boyko N, Artis D, Cebra JJ, Wu GD. Activation of RegIIIbeta/gamma and interferon gamma expression in the intestinal tract of SCID mice: an innate response to bacterial colonisation of the gut. Gut. 2005;54:623-629. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 73] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 22. | Shindo R, Katagiri T, Komazawa-Sakon S, Ohmuraya M, Takeda W, Nakagawa Y, Nakagata N, Sakuma T, Yamamoto T, Nishiyama C, Nishina T, Yamazaki S, Kameda H, Nakano H. Regenerating islet-derived protein (Reg)3β plays a crucial role in attenuation of ileitis and colitis in mice. Biochem Biophys Rep. 2020;21:100738. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 12] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 23. | Ferrara JL, Harris AC, Greenson JK, Braun TM, Holler E, Teshima T, Levine JE, Choi SW, Huber E, Landfried K, Akashi K, Vander Lugt M, Reddy P, Chin A, Zhang Q, Hanash S, Paczesny S. Regenerating islet-derived 3-alpha is a biomarker of gastrointestinal graft-versus-host disease. Blood. 2011;118:6702-6708. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 229] [Cited by in RCA: 261] [Article Influence: 18.6] [Reference Citation Analysis (0)] |
| 24. | Zhao D, Kim YH, Jeong S, Greenson JK, Chaudhry MS, Hoepting M, Anderson ER, van den Brink MR, Peled JU, Gomes AL, Slingerland AE, Donovan MJ, Harris AC, Levine JE, Ozbek U, Hooper LV, Stappenbeck TS, Ver Heul A, Liu TC, Reddy P, Ferrara JL. Survival signal REG3α prevents crypt apoptosis to control acute gastrointestinal graft-versus-host disease. J Clin Invest. 2018;128:4970-4979. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 106] [Article Influence: 15.1] [Reference Citation Analysis (0)] |
| 25. | Zheng Y, Valdez PA, Danilenko DM, Hu Y, Sa SM, Gong Q, Abbas AR, Modrusan Z, Ghilardi N, de Sauvage FJ, Ouyang W. Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat Med. 2008;14:282-289. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1508] [Cited by in RCA: 1550] [Article Influence: 91.2] [Reference Citation Analysis (0)] |
| 26. | Mukherjee S, Hooper LV. Antimicrobial defense of the intestine. Immunity. 2015;42:28-39. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 185] [Cited by in RCA: 237] [Article Influence: 23.7] [Reference Citation Analysis (0)] |
| 27. | Shin JH, Seeley RJ. Reg3 Proteins as Gut Hormones? Endocrinology. 2019;160:1506-1514. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 71] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 28. | Cash HL, Whitham CV, Behrendt CL, Hooper LV. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science. 2006;313:1126-1130. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1159] [Cited by in RCA: 1076] [Article Influence: 56.6] [Reference Citation Analysis (0)] |
| 29. | Lehotzky RE, Partch CL, Mukherjee S, Cash HL, Goldman WE, Gardner KH, Hooper LV. Molecular basis for peptidoglycan recognition by a bactericidal lectin. Proc Natl Acad Sci USA. 2010;107:7722-7727. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 122] [Cited by in RCA: 115] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 30. | Mukherjee S, Zheng H, Derebe MG, Callenberg KM, Partch CL, Rollins D, Propheter DC, Rizo J, Grabe M, Jiang QX, Hooper LV. Antibacterial membrane attack by a pore-forming intestinal C-type lectin. Nature. 2014;505:103-107. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 210] [Cited by in RCA: 241] [Article Influence: 20.1] [Reference Citation Analysis (0)] |
| 31. | Vaishnava S, Yamamoto M, Severson KM, Ruhn KA, Yu X, Koren O, Ley R, Wakeland EK, Hooper LV. The antibacterial lectin RegIIIgamma promotes the spatial segregation of microbiota and host in the intestine. Science. 2011;334:255-258. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1124] [Cited by in RCA: 1080] [Article Influence: 77.1] [Reference Citation Analysis (0)] |
| 32. | Miki T, Holst O, Hardt WD. The bactericidal activity of the C-type lectin RegIIIβ against Gram-negative bacteria involves binding to lipid A. J Biol Chem. 2012;287:34844-34855. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 71] [Cited by in RCA: 91] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 33. | Stelter C, Käppeli R, König C, Krah A, Hardt WD, Stecher B, Bumann D. Salmonella-induced mucosal lectin RegIIIβ kills competing gut microbiota. PLoS One. 2011;6:e20749. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 103] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 34. | van Ampting MT, Loonen LM, Schonewille AJ, Konings I, Vink C, Iovanna J, Chamaillard M, Dekker J, van der Meer R, Wells JM, Bovee-Oudenhoven IM. Intestinally secreted C-type lectin Reg3b attenuates salmonellosis but not listeriosis in mice. Infect Immun. 2012;80:1115-1120. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 85] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 35. | Wang L, Fouts DE, Stärkel P, Hartmann P, Chen P, Llorente C, DePew J, Moncera K, Ho SB, Brenner DA, Hooper LV, Schnabl B. Intestinal REG3 Lectins Protect against Alcoholic Steatohepatitis by Reducing Mucosa-Associated Microbiota and Preventing Bacterial Translocation. Cell Host Microbe. 2016;19:227-239. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 209] [Cited by in RCA: 293] [Article Influence: 32.6] [Reference Citation Analysis (0)] |
| 36. | Mahadevan U, Silverberg MS. Inflammatory Bowel Disease-Gastroenterology Diamond Jubilee Review. Gastroenterology. 2018;154:1555-1558. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 15] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 37. | de Souza HS, Fiocchi C. Immunopathogenesis of IBD: current state of the art. Nat Rev Gastroenterol Hepatol. 2016;13:13-27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1185] [Cited by in RCA: 1120] [Article Influence: 124.4] [Reference Citation Analysis (1)] |
| 38. | Lawrance IC, Fiocchi C, Chakravarti S. Ulcerative colitis and Crohn's disease: distinctive gene expression profiles and novel susceptibility candidate genes. Hum Mol Genet. 2001;10:445-456. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 278] [Cited by in RCA: 292] [Article Influence: 12.2] [Reference Citation Analysis (1)] |
| 39. | Shinozaki S, Nakamura T, Iimura M, Kato Y, Iizuka B, Kobayashi M, Hayashi N. Upregulation of Reg 1alpha and GW112 in the epithelium of inflamed colonic mucosa. Gut. 2001;48:623-629. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 78] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 40. | Desjeux A, Barthet M, Barthellemy S, Dagorn JC, Hastier P, Heresbach D, Bernard JP, Grimaud JC. Serum measurements of pancreatitis associated protein in active Crohn's disease with ileal location. Gastroenterol Clin Biol. 2002;26:23-28. [PubMed] |
| 41. | Dieckgraefe BK, Crimmins DL, Landt V, Houchen C, Anant S, Porche-Sorbet R, Ladenson JH. Expression of the regenerating gene family in inflammatory bowel disease mucosa: Reg Ialpha upregulation, processing, and antiapoptotic activity. J Investig Med. 2002;50:421-434. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 57] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 42. | Kämäräinen M, Heiskala K, Knuutila S, Heiskala M, Winqvist O, Andersson LC. RELP, a novel human REG-like protein with up-regulated expression in inflammatory and metaplastic gastrointestinal mucosa. Am J Pathol. 2003;163:11-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 42] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 43. | Gironella M, Iovanna JL, Sans M, Gil F, Peñalva M, Closa D, Miquel R, Piqué JM, Panés J. Anti-inflammatory effects of pancreatitis associated protein in inflammatory bowel disease. Gut. 2005;54:1244-1253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 89] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 44. | Wu F, Dassopoulos T, Cope L, Maitra A, Brant SR, Harris ML, Bayless TM, Parmigiani G, Chakravarti S. Genome-wide gene expression differences in Crohn's disease and ulcerative colitis from endoscopic pinch biopsies: insights into distinctive pathogenesis. Inflamm Bowel Dis. 2007;13:807-821. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 199] [Cited by in RCA: 217] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 45. | Nanakin A, Fukui H, Fujii S, Sekikawa A, Kanda N, Hisatsune H, Seno H, Konda Y, Fujimori T, Chiba T. Expression of the REG IV gene in ulcerative colitis. Lab Invest. 2007;87:304-314. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 59] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 46. | Sekikawa A, Fukui H, Suzuki K, Karibe T, Fujii S, Ichikawa K, Tomita S, Imura J, Shiratori K, Chiba T, Fujimori T. Involvement of the IL-22/REG Ialpha axis in ulcerative colitis. Lab Invest. 2010;90:496-505. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 76] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 47. | Tanaka H, Fukui H, Fujii S, Sekikawa A, Yamagishi H, Ichikawa K, Tomita S, Imura J, Yasuda Y, Chiba T, Fujimori T. Immunohistochemical analysis of REG Iα expression in ulcerative colitis-associated neoplastic lesions. Digestion. 2011;83:204-209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 48. | van Beelen Granlund A, Østvik AE, Brenna Ø, Torp SH, Gustafsson BI, Sandvik AK. REG gene expression in inflamed and healthy colon mucosa explored by in situ hybridisation. Cell Tissue Res. 2013;352:639-646. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 54] [Cited by in RCA: 71] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 49. | Planell N, Lozano JJ, Mora-Buch R, Masamunt MC, Jimeno M, Ordás I, Esteller M, Ricart E, Piqué JM, Panés J, Salas A. Transcriptional analysis of the intestinal mucosa of patients with ulcerative colitis in remission reveals lasting epithelial cell alterations. Gut. 2013;62:967-976. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 157] [Cited by in RCA: 212] [Article Influence: 17.7] [Reference Citation Analysis (0)] |
| 50. | Marafini I, Di Sabatino A, Zorzi F, Monteleone I, Sedda S, Cupi ML, Antenucci C, Biancheri P, Giuffrida P, Di Stefano M, Corazza GR, Pallone F, Monteleone G. Serum regenerating islet-derived 3-alpha is a biomarker of mucosal enteropathies. Aliment Pharmacol Ther. 2014;40:974-981. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 23] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 51. | Nunes T, Etchevers MJ, Sandi MJ, Pinó Donnay S, Grandjean T, Pellisé M, Panés J, Ricart E, Iovanna JL, Dagorn JC, Chamaillard M, Sans M. Pancreatitis-associated protein does not predict disease relapse in inflammatory bowel disease patients. PLoS One. 2014;9:e84957. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 52. | Tsuchida C, Sakuramoto-Tsuchida S, Taked M, Itaya-Hironaka A, Yamauchi A, Misu M, Shobatake R, Uchiyama T, Makino M, Pujol-Autonell I, Vives-Pi M, Ohbayashi C, Takasawa S. Expression of REG family genes in human inflammatory bowel diseases and its regulation. Biochem Biophys Rep. 2017;12:198-205. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 22] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 53. | Strober W, Fuss IJ. Proinflammatory cytokines in the pathogenesis of inflammatory bowel diseases. Gastroenterology. 2011;140:1756-1767. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 887] [Cited by in RCA: 853] [Article Influence: 60.9] [Reference Citation Analysis (0)] |
| 54. | Andoh A, Zhang Z, Inatomi O, Fujino S, Deguchi Y, Araki Y, Tsujikawa T, Kitoh K, Kim-Mitsuyama S, Takayanagi A, Shimizu N, Fujiyama Y. Interleukin-22, a member of the IL-10 subfamily, induces inflammatory responses in colonic subepithelial myofibroblasts. Gastroenterology. 2005;129:969-984. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 379] [Cited by in RCA: 398] [Article Influence: 19.9] [Reference Citation Analysis (0)] |
| 55. | Brand S, Beigel F, Olszak T, Zitzmann K, Eichhorst ST, Otte JM, Diepolder H, Marquardt A, Jagla W, Popp A, Leclair S, Herrmann K, Seiderer J, Ochsenkühn T, Göke B, Auernhammer CJ, Dambacher J. IL-22 is increased in active Crohn's disease and promotes proinflammatory gene expression and intestinal epithelial cell migration. Am J Physiol Gastrointest Liver Physiol. 2006;290:G827-G838. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 417] [Cited by in RCA: 447] [Article Influence: 23.5] [Reference Citation Analysis (0)] |
| 56. | Sugimoto K, Ogawa A, Mizoguchi E, Shimomura Y, Andoh A, Bhan AK, Blumberg RS, Xavier RJ, Mizoguchi A. IL-22 ameliorates intestinal inflammation in a mouse model of ulcerative colitis. J Clin Invest. 2008;118:534-544. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 183] [Cited by in RCA: 566] [Article Influence: 33.3] [Reference Citation Analysis (0)] |
| 57. | Leung JM, Davenport M, Wolff MJ, Wiens KE, Abidi WM, Poles MA, Cho I, Ullman T, Mayer L, Loke P. IL-22-producing CD4+ cells are depleted in actively inflamed colitis tissue. Mucosal Immunol. 2014;7:124-133. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 103] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 58. | Sanos SL, Vonarbourg C, Mortha A, Diefenbach A. Control of epithelial cell function by interleukin-22-producing RORγt+ innate lymphoid cells. Immunology. 2011;132:453-465. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 92] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 59. | Ikeda H, Sasaki M, Ishikawa A, Sato Y, Harada K, Zen Y, Kazumori H, Nakanuma Y. Interaction of Toll-like receptors with bacterial components induces expression of CDX2 and MUC2 in rat biliary epithelium in vivo and in culture. Lab Invest. 2007;87:559-571. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 46] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 60. | Coskun M, Troelsen JT, Nielsen OH. The role of CDX2 in intestinal homeostasis and inflammation. Biochim Biophys Acta. 2011;1812:283-289. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 76] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 61. | Koh I, Nosaka S, Sekine M, Sugimoto J, Hirata E, Kudo Y. Regulation of REG4 Expression and Prediction of 5-Fluorouracil Sensitivity by CDX2 in Ovarian Mucinous Carcinoma. Cancer Genomics Proteomics. 2019;16:481-490. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 8] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 62. | Naito Y, Oue N, Hinoi T, Sakamoto N, Sentani K, Ohdan H, Yanagihara K, Sasaki H, Yasui W. Reg IV is a direct target of intestinal transcriptional factor CDX2 in gastric cancer. PLoS One. 2012;7:e47545. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 29] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 63. | Kim S, Domon-Dell C, Wang Q, Chung DH, Di Cristofano A, Pandolfi PP, Freund JN, Evers BM. PTEN and TNF-alpha regulation of the intestinal-specific Cdx-2 homeobox gene through a PI3K, PKB/Akt, and NF-kappaB-dependent pathway. Gastroenterology. 2002;123:1163-1178. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 99] [Cited by in RCA: 102] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 64. | Kobayashi S, Akiyama T, Nata K, Abe M, Tajima M, Shervani NJ, Unno M, Matsuno S, Sasaki H, Takasawa S, Okamoto H. Identification of a receptor for reg (regenerating gene) protein, a pancreatic beta-cell regeneration factor. J Biol Chem. 2000;275:10723-10726. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 135] [Cited by in RCA: 130] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 65. | Lai Y, Li D, Li C, Muehleisen B, Radek KA, Park HJ, Jiang Z, Li Z, Lei H, Quan Y, Zhang T, Wu Y, Kotol P, Morizane S, Hata TR, Iwatsuki K, Tang C, Gallo RL. The antimicrobial protein REG3A regulates keratinocyte proliferation and differentiation after skin injury. Immunity. 2012;37:74-84. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 170] [Cited by in RCA: 195] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 66. | Acquatella-Tran Van Ba I, Marchal S, François F, Silhol M, Lleres C, Michel B, Benyamin Y, Verdier JM, Trousse F, Marcilhac A. Regenerating islet-derived 1α (Reg-1α) protein is new neuronal secreted factor that stimulates neurite outgrowth via exostosin Tumor-like 3 (EXTL3) receptor. J Biol Chem. 2012;287:4726-4739. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 34] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 67. | LaFonte MW, Stanek A, Mueller C, Zenilman ME, Sugawara A, Alfonso AE, Huan C. Identification of Reg1 as a novel stellate cell activator in regenerating pancreas. J Am Coll Surg. 2013;217:S18. [RCA] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 68. | Lentjes MH, Niessen HE, Akiyama Y, de Bruïne AP, Melotte V, van Engeland M. The emerging role of GATA transcription factors in development and disease. Expert Rev Mol Med. 2016;18:e3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 138] [Cited by in RCA: 172] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 69. | Lepage D, Bruneau J, Brouillard G, Jones C, Lussier CR, Rémillard A, Lemieux É, Asselin C, Boudreau F. Identification of GATA-4 as a novel transcriptional regulatory component of regenerating islet-derived family members. Biochim Biophys Acta. 2015;1849:1411-1422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 70. | Haveri H, Ashorn M, Iltanen S, Wilson DB, Andersson LC, Heikinheimo M. Enhanced expression of transcription factor GATA-4 in inflammatory bowel disease and its possible regulation by TGF-beta1. J Clin Immunol. 2009;29:444-453. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 71. | Kawasaki Y, Matsumura K, Miyamoto M, Tsuji S, Okuno M, Suda S, Hiyoshi M, Kitayama J, Akiyama T. REG4 is a transcriptional target of GATA6 and is essential for colorectal tumorigenesis. Sci Rep. 2015;5:14291. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 36] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 72. | Beuling E, Baffour-Awuah NY, Stapleton KA, Aronson BE, Noah TK, Shroyer NF, Duncan SA, Fleet JC, Krasinski SD. GATA factors regulate proliferation, differentiation, and gene expression in small intestine of mature mice. Gastroenterology. 2011;140:1219-1229.e1-2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 90] [Cited by in RCA: 85] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 73. | Bosse T, Piaseckyj CM, Burghard E, Fialkovich JJ, Rajagopal S, Pu WT, Krasinski SD. Gata4 is essential for the maintenance of jejunal-ileal identities in the adult mouse small intestine. Mol Cell Biol. 2006;26:9060-9070. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 112] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 74. | Babyatsky MW, Rossiter G, Podolsky DK. Expression of transforming growth factors alpha and beta in colonic mucosa in inflammatory bowel disease. Gastroenterology. 1996;110:975-984. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 291] [Cited by in RCA: 285] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 75. | McCabe RP, Secrist H, Botney M, Egan M, Peters MG. Cytokine mRNA expression in intestine from normal and inflammatory bowel disease patients. Clin Immunol Immunopathol. 1993;66:52-58. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 56] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 76. | Mustfa SA, Singh M, Suhail A, Mohapatra G, Verma S, Chakravorty D, Rana S, Rampal R, Dhar A, Saha S, Ahuja V, Srikanth CV. SUMOylation pathway alteration coupled with downregulation of SUMO E2 enzyme at mucosal epithelium modulates inflammation in inflammatory bowel disease. Open Biol. 2017;7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 37] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 77. | Kang X, Qi Y, Zuo Y, Wang Q, Zou Y, Schwartz RJ, Cheng J, Yeh ET. SUMO-specific protease 2 is essential for suppression of polycomb group protein-mediated gene silencing during embryonic development. Mol Cell. 2010;38:191-201. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 184] [Cited by in RCA: 176] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 78. | Damas OM, Garces L, Abreu MT. Diet as Adjunctive Treatment for Inflammatory Bowel Disease: Review and Update of the Latest Literature. Curr Treat Options Gastroenterol. 2019;17:313-325. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 44] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 79. | Darnaud M, Dos Santos A, Gonzalez P, Augui S, Lacoste C, Desterke C, De Hertogh G, Valentino E, Braun E, Zheng J, Boisgard R, Neut C, Dubuquoy L, Chiappini F, Samuel D, Lepage P, Guerrieri F, Doré J, Bréchot C, Moniaux N, Faivre J. Enteric Delivery of Regenerating Family Member 3 alpha Alters the Intestinal Microbiota and Controls Inflammation in Mice With Colitis. Gastroenterology. 2018;154:1009-1023.e14. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 120] [Article Influence: 17.1] [Reference Citation Analysis (0)] |
| 80. | Huang Y, Qi H, Zhang Z, Wang E, Yun H, Yan H, Su X, Liu Y, Tang Z, Gao Y, Shang W, Zhou J, Wang T, Che Y, Zhang Y, Yang R. Gut REG3γ-Associated Lactobacillus Induces Anti-inflammatory Macrophages to Maintain Adipose Tissue Homeostasis. Front Immunol. 2017;8:1063. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 20] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 81. | Prosberg M, Bendtsen F, Vind I, Petersen AM, Gluud LL. The association between the gut microbiota and the inflammatory bowel disease activity: a systematic review and meta-analysis. Scand J Gastroenterol. 2016;51:1407-1415. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 171] [Cited by in RCA: 153] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 82. | Yang X, Jin H, Liu K, Gu Q, Xu X. A novel peptide derived from human pancreatitis-associated protein inhibits inflammation in vivo and in vitro and blocks NF-kappa B signaling pathway. PLoS One. 2011;6:e29155. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 22] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 83. | Folch-Puy E, Granell S, Dagorn JC, Iovanna JL, Closa D. Pancreatitis-associated protein I suppresses NF-kappa B activation through a JAK/STAT-mediated mechanism in epithelial cells. J Immunol. 2006;176:3774-3779. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 55] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 84. | Vasseur S, Folch-Puy E, Hlouschek V, Garcia S, Fiedler F, Lerch MM, Dagorn JC, Closa D, Iovanna JL. p8 improves pancreatic response to acute pancreatitis by enhancing the expression of the anti-inflammatory protein pancreatitis-associated protein I. J Biol Chem. 2004;279:7199-7207. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 93] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 85. | Viterbo D, Bluth MH, Lin YY, Mueller CM, Wadgaonkar R, Zenilman ME. Pancreatitis-associated protein 2 modulates inflammatory responses in macrophages. J Immunol. 2008;181:1948-1958. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 26] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 86. | Viterbo D, Bluth MH, Mueller CM, Zenilman ME. Mutational characterization of pancreatitis-associated protein 2 domains involved in mediating cytokine secretion in macrophages and the NF-kappaB pathway. J Immunol. 2008;181:1959-1968. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 87. | Na YR, Stakenborg M, Seok SH, Matteoli G. Macrophages in intestinal inflammation and resolution: a potential therapeutic target in IBD. Nat Rev Gastroenterol Hepatol. 2019;16:531-543. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 272] [Cited by in RCA: 594] [Article Influence: 99.0] [Reference Citation Analysis (0)] |
| 88. | Click B, Regueiro M. A Practical Guide to the Safety and Monitoring of New IBD Therapies. Inflamm Bowel Dis. 2019;25:831-842. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 43] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 89. | Su HJ, Chiu YT, Chiu CT, Lin YC, Wang CY, Hsieh JY, Wei SC. Inflammatory bowel disease and its treatment in 2018: Global and Taiwanese status updates. J Formos Med Assoc. 2019;118:1083-1092. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 44] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 90. | Takasawa S. Regenerating gene (REG) product and its potential clinical usage. Expert Opin Ther Targets. 2016;20:541-550. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 40] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 91. | Dietrich CF. Significance of abdominal ultrasound in inflammatory bowel disease. Dig Dis. 2009;27:482-493. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 44] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 92. | Deepak P, Fletcher JG, Fidler JL, Bruining DH. Computed Tomography and Magnetic Resonance Enterography in Crohn's Disease: Assessment of Radiologic Criteria and Endpoints for Clinical Practice and Trials. Inflamm Bowel Dis. 2016;22:2280-2288. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 33] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 93. | Norouzinia M, Chaleshi V, Alizadeh AHM, Zali MR. Biomarkers in inflammatory bowel diseases: insight into diagnosis, prognosis and treatment. Gastroenterol Hepatol Bed Bench. 2017;10:155-167. [PubMed] |
| 94. | Iskandar HN, Ciorba MA. Biomarkers in inflammatory bowel disease: current practices and recent advances. Transl Res. 2012;159:313-325. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 128] [Cited by in RCA: 146] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 95. | Rogler G, Biedermann L. Clinical Utility of Biomarkers in IBD. Curr Gastroenterol Rep. 2015;17:26. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 21] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 96. | Bennike T, Birkelund S, Stensballe A, Andersen V. Biomarkers in inflammatory bowel diseases: current status and proteomics identification strategies. World J Gastroenterol. 2014;20:3231-3244. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 78] [Cited by in RCA: 87] [Article Influence: 7.9] [Reference Citation Analysis (1)] |
| 97. | Moayyedi P, Surette MG, Kim PT, Libertucci J, Wolfe M, Onischi C, Armstrong D, Marshall JK, Kassam Z, Reinisch W, Lee CH. Fecal Microbiota Transplantation Induces Remission in Patients With Active Ulcerative Colitis in a Randomized Controlled Trial. Gastroenterology. 2015;149:102-109.e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 930] [Cited by in RCA: 1087] [Article Influence: 108.7] [Reference Citation Analysis (1)] |
| 98. | Paramsothy S, Kamm MA, Kaakoush NO, Walsh AJ, van den Bogaerde J, Samuel D, Leong RWL, Connor S, Ng W, Paramsothy R, Xuan W, Lin E, Mitchell HM, Borody TJ. Multidonor intensive faecal microbiota transplantation for active ulcerative colitis: a randomised placebo-controlled trial. Lancet. 2017;389:1218-1228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 710] [Cited by in RCA: 876] [Article Influence: 109.5] [Reference Citation Analysis (0)] |
| 99. | Costello SP, Hughes PA, Waters O, Bryant RV, Vincent AD, Blatchford P, Katsikeros R, Makanyanga J, Campaniello MA, Mavrangelos C, Rosewarne CP, Bickley C, Peters C, Schoeman MN, Conlon MA, Roberts-Thomson IC, Andrews JM. Effect of Fecal Microbiota Transplantation on 8-Week Remission in Patients With Ulcerative Colitis: A Randomized Clinical Trial. JAMA. 2019;321:156-164. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 414] [Cited by in RCA: 621] [Article Influence: 103.5] [Reference Citation Analysis (0)] |
| 100. | Narula N, Kassam Z, Yuan Y, Colombel JF, Ponsioen C, Reinisch W, Moayyedi P. Systematic Review and Meta-analysis: Fecal Microbiota Transplantation for Treatment of Active Ulcerative Colitis. Inflamm Bowel Dis. 2017;23:1702-1709. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 145] [Article Influence: 18.1] [Reference Citation Analysis (0)] |
| 101. | Feinberg AE, Valente MA. Elective Abdominal Surgery for Inflammatory Bowel Disease. Surg Clin North Am. 2019;99:1123-1140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 102. | Fuglestad MA, Thompson JS. Inflammatory Bowel Disease and Short Bowel Syndrome. Surg Clin North Am. 2019;99:1209-1221. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 15] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 103. | Hwang JM, Varma MG. Surgery for inflammatory bowel disease. World J Gastroenterol. 2008;14:2678-2690. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 63] [Cited by in RCA: 77] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 104. | Adamina M, Steffen T, Tarantino I, Beutner U, Schmied BM, Warschkow R. Meta-analysis of the predictive value of C-reactive protein for infectious complications in abdominal surgery. Br J Surg. 2015;102:590-598. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 98] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 105. | Fisher OM, Oberkofler CE, Raptis DA, Soll C, Béchir M, Schiesser M, Graf R. Pancreatic stone protein (PSP) and pancreatitis-associated protein (PAP): a protocol of a cohort study on the diagnostic efficacy and prognostic value of PSP and PAP as postoperative markers of septic complications in patients undergoing abdominal surgery (PSP study). BMJ Open. 2014;4:e004914. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 106. | Klein HJ, Csordas A, Falk V, Slankamenac K, Rudiger A, Schönrath F, Rodriguez Cetina Biefer H, Starck CT, Graf R. Pancreatic stone protein predicts postoperative infection in cardiac surgery patients irrespective of cardiopulmonary bypass or surgical technique. PLoS One. 2015;10:e0120276. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 23] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 107. | Reding T, Palmiere C, Pazhepurackel C, Schiesser M, Bimmler D, Schlegel A, Süss U, Steiner S, Mancina L, Seleznik G, Graf R. The pancreas responds to remote damage and systemic stress by secretion of the pancreatic secretory proteins PSP/regI and PAP/regIII. Oncotarget. 2017;8:30162-30174. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 22] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 108. | Rass AA, Talat MA, Arafa MA, El-Saadany HF, Amin EK, Abdelsalam MM, Mansour MA, Khalifa NA, Kamel LM. The Role of Pancreatic Stone Protein in Diagnosis of Early Onset Neonatal Sepsis. Biomed Res Int. 2016;2016:1035856. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 7] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 109. | Keel M, Härter L, Reding T, Sun LK, Hersberger M, Seifert B, Bimmler D, Graf R. Pancreatic stone protein is highly increased during posttraumatic sepsis and activates neutrophil granulocytes. Crit Care Med. 2009;37:1642-1648. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 78] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 110. | Que YA, Delodder F, Guessous I, Graf R, Bain M, Calandra T, Liaudet L, Eggimann P. Pancreatic stone protein as an early biomarker predicting mortality in a prospective cohort of patients with sepsis requiring ICU management. Crit Care. 2012;16:R114. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 47] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 111. | Bettenworth D, Nowacki TM, Cordes F, Buerke B, Lenze F. Assessment of stricturing Crohn's disease: Current clinical practice and future avenues. World J Gastroenterol. 2016;22:1008-1016. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 37] [Cited by in RCA: 37] [Article Influence: 4.1] [Reference Citation Analysis (0)] |