Published online Jul 14, 2019. doi: 10.3748/wjg.v25.i26.3392
Peer-review started: March 14, 2019
First decision: April 30, 2019
Revised: May 9, 2019
Accepted: May 31, 2019
Article in press: June 1, 2019
Published online: July 14, 2019
Processing time: 123 Days and 8.4 Hours
DNA methylation, acknowledged as a key modification in the field of epigenetics, regulates gene expression at the transcriptional level. Aberrant methylation in DNA regulatory regions could upregulate oncogenes and downregulate tumor suppressor genes without changing the sequences. However, studies of methylation in the control of gene expression are still inadequate. In the present research, we performed bioinformatics analysis to clarify the function of methylation and supply candidate methylation-related biomarkers and drivers for colon cancer.
To identify and analyze methylation-regulated differentially expressed genes (MeDEGs) in colon cancer by bioinformatics analysis.
We downloaded RNA expression profiles, Illumina Human Methylation 450K BeadChip data, and clinical data of colon cancer from The Cancer Genome Atlas project. MeDEGs were identified by analyzing the gene expression and methylation levels using the edgeR and limma package in R software. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed in the DAVID database and KEGG Orthology-Based Annotation System 3.0, respectively. We then conducted Kaplan–Meier survival analysis to explore the relationship between methylation and expression and prognosis. Gene set enrichment analysis (GSEA) and investigation of protein-protein interactions (PPI) were performed to clarify the function of prognosis-related genes.
A total of 5 up-regulated and 81 down-regulated genes were identified as MeDEGs. GO and KEGG pathway analyses indicated that MeDEGs were enriched in multiple cancer-related terms. Furthermore, Kaplan–Meier survival analysis showed that the prognosis was negatively associated with the methylation status of glial cell-derived neurotrophic factor (GDNF) and reelin (RELN). In PPI networks, GDNF and RELN interact with neural cell adhesion molecule 1. Besides, GDNF can interact with GDNF family receptor alpha (GFRA1), GFRA2, GFRA3, and RET. RELN can interact with RAFAH1B1, disabled homolog 1, very low-density lipoprotein receptor, lipoprotein receptor-related protein 8, and NMDA 2B. Based on GSEA, hypermethylation of GDNF and RELN were both significantly associated with pathways including “RNA degradation,” “ribosome,” “mismatch repair,” “cell cycle” and “base excision repair.”
Aberrant DNA methylation plays an important role in colon cancer progression. MeDEGs that are associated with the overall survival of patients may be potential targets in tumor diagnosis and treatment.
Core tip: We acquired high-throughput data from The Cancer Genome Atlas database and identified 5 up-regulated and 81 down-regulated methylation-regulated differentially expressed gene (MeDEGs). Then, we performed gene ontology and Kyoto Encyclopedia of Genes and Genomes pathway analyses to clarify the function of MeDEGs. Furthermore, methylation status and expression of glial cell-derived neurotrophic factor and reelin were found to be associated with overall survival. Our study revealed that methylation is critical in colon cancer development and the prognosis-related MeDEGs are worth exploring further.
- Citation: Liang Y, Zhang C, Dai DQ. Identification of differentially expressed genes regulated by methylation in colon cancer based on bioinformatics analysis. World J Gastroenterol 2019; 25(26): 3392-3407
- URL: https://www.wjgnet.com/1007-9327/full/v25/i26/3392.htm
- DOI: https://dx.doi.org/10.3748/wjg.v25.i26.3392
Global cancer statistics for 185 countries reported that there were 1091601 new cases and 551269 deaths from colon cancer in 2018. The incidence and mortality were both ranked fourth worldwide[1]. Epidemiological studies have confirmed that the occurrence of colon cancer is related to multiple factors including smoking, alcohol abuse, insufficient activity, and high-fat food[2]. Although surgery-based com-prehensive treatment is considered an effective treatment, a large number of patients still die from postoperative recurrence and metastasis[3]. Hence, research into specific biomarkers and therapeutic pathways is still of great value for improving patient prognosis.
Accumulating studies indicate that epigenetic modifications could promote the initiation and progression of colon cancer via multiple signaling pathways[4]. DNA methylation has been discovered and verified as a critical modification in the field of epigenetics. Numerous oncogenes and cancer-suppressor genes exhibit irregular expression, which is due to aberrant cytosine-phosphate-guanine (CpG)-island methylation in DNA regulatory regions rather than changes in the sequences[5]. For instance, the anti-oncogene ZNF350 undergoes epigenetic silencing due to hypermethylation of three sites in the promoter[6]. However, studies of DNA methylation in the individual genes remain inadequate. Identification of methylation-regulated differentially expressed genes (MeDEGs) based on high-throughput data will be of profound significance for clarifying the role of methylation and identifying candidate directions for future research.
In recent years, with the development of gene-sequencing platforms, accumulating differentially expressed genes (DEGs) and epigenetic alterations have been revealed by bioinformatic analysis. For instance, Naumov et al[7] applied an Illumina Human Methylation 450K BeadChip to detect DNA methylation profiles in colorectal cancer tissues and colon tissues from cancer-free donors. As a result, 10342 hypermethylated and 5325 hypomethylated CpG sites were identified. Dumenil et al[8] further indicated that the Illumina Human Methylation 450K BeadChip could be utilized to detect and obtain high quality data from paraffin-embedded tissues. Multiple studies have shown that the Illumina Methylation 450K BeadChip has important application value in methylation research. Almost 450000 methylation sites and 96% of the CpG islands can be detected using the Illumina Human Methylation 450K BeadChip platform[9]. However, there is still a lack of conjoint analysis of colon cancer methylation and studies on the correlation between methylation and patient prognosis in large cohorts.
In the present research, we downloaded in silico data and clinical data of colon cancer from The Cancer Genome Atlas (TCGA, http://cancergenome.nih.gov) project[10]. MeDEGs were identified and related enrichment analysis was then performed. Moreover, we analyzed the correlation between the methylation status and overall survival of colon cancer patients. Protein-protein interaction (PPI) networks were constructed and gene set enrichment analysis (GSEA) of prognosis-related MeDEGs was further performed.
We downloaded colon cancer RNA expression profiles and clinical data from TCGA using the Genomic Data Commons Data Transfer Tool 1.3.0[10]. Up to December 2018, the public database included the expression profiles of 473 colon cancer tissues and 41 normal tissues (level 3) derived by RNA-seq. A total of 314 tumor tissues and 37 normal tissues had also been analyzed in the Illumina Human Methylation 450K BeadChip platform, and we also downloaded homologous methylation profiles from TCGA database. As 303 tumor tissues and 19 normal tissues were analyzed using both platforms, we merged the methylation and expression data together for correlation analysis. Based on the guidelines released by the National Cancer Institute in December 2015 (https://cancergenome.nih.gov/publications/publication guidelines), our research did not require the approval of an ethics committee.
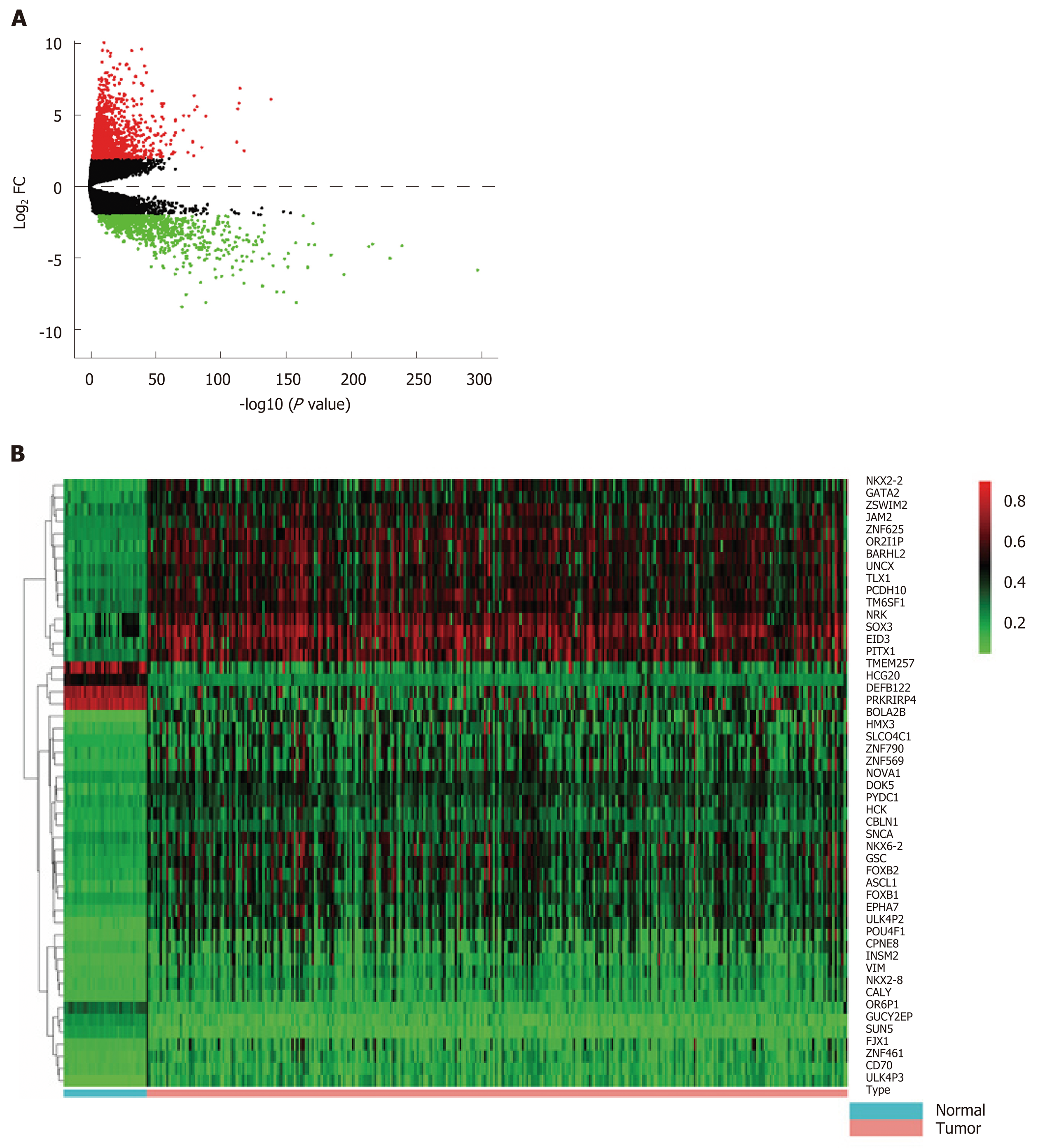
We constructed an RNA matrix and methylation data matrix containing the expression and methylation profiles using PERL software. Then, the coding gene IDs were converted to gene names based on information in the Ensembl database (Homo sapiens) (http://asia.ensembl.org/index.html). DEGs were identified with the edgeR package in R software with a threshold log2 fold change (FC) > 2.0 and P < 0.01. Moreover, differentially methylated genes (DMGs) were identified using the limma package with a threshold log2 FC > 1.0 and P < 0.01. Additionally, we analyzed the correlation between gene expression and methylation data through Spearman’s correlation analysis. The upregulated-hypomethylated and downregulated-hypermethylated genes were identified as MeDEGs when they satisfied the cut-off criteria including correlation coefficient < -0.3 and P < 0.01. A volcano plot and heat map were drawn using R software.
To explore the function of the MeDEGs in colon cancer carcinogenesis and progression, gene ontology (GO)[11] enrichment analysis was performed using the DAVID database (https://david.ncifcrf.gov/). The GO enrichment analysis included three categories: cellular component, molecular function, and biological process. Furthermore, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways[12] were analyzed using KEGG Orthology-Based Annotation System 3.0 (http://koba s.cbi.pku.edu.cn/). Differences with P < 0.05 were regarded as statistically significant.
We divided the 314 colon cancer patients into two groups according to the median methylation value of the MeDEGs. Moreover, the patients were also divided into a hypermethylation and low-expression MeDEG (Hyper-LG) group and a hypomethylation and high-expression MeDEG (Hypo-HG) group according to the median value of methylation and expression of MeDEGs. Comparison of the overall survival between the two groups was then analyzed. Kaplan–Meier’s method and the log-rank test were performed to assess survival rate. Differences with P < 0.05 were regarded as statistically significant.
GSEA of prognosis-related MeDEGs was performed using GSEA 3.0 software with gene set c2 (cp.kegg.v.6.2.symbols.gmt). High throughput RNA expression of 303 colon cancer genes from TCGA were utilized as the dataset. Each sample was defined as either “H” or “L,” depending on whether it was greater than the median methylation value of prognosis-related MeDEGs or not. The number and type of permutations was set at “1000” and “phenotype,” respectively. An enrichment score > 0.4 and P < 0.05 were regarded as statistically significant.
PPI analysis was conducted to reveal the molecular mechanisms of the prognosis-related MeDEGs in colon cancer. We utilized the STRING protein database 11.0 (http://string-db.org/) to construct the PPI networks. An interaction score > 0.4 was regarded as the cut-off criterion.
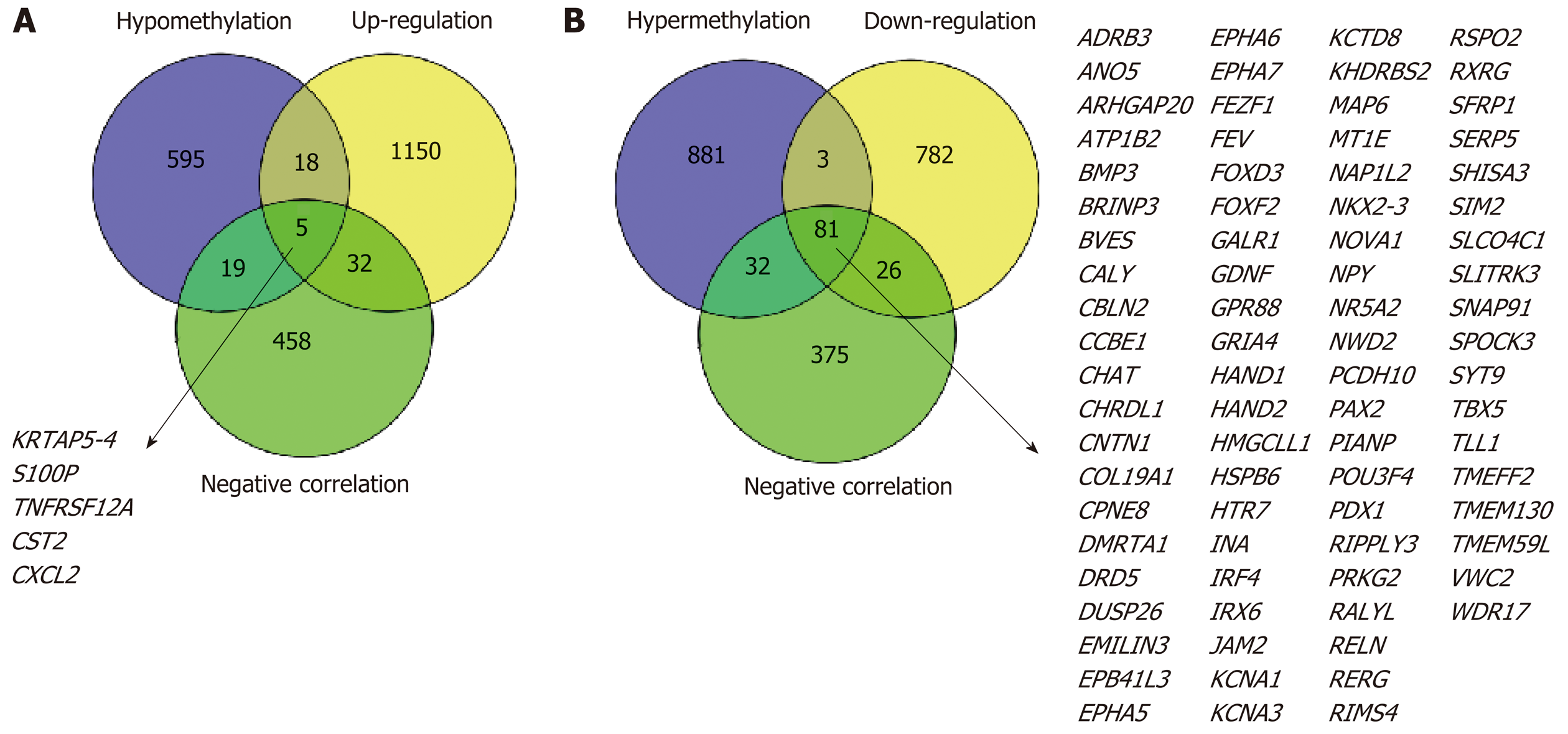
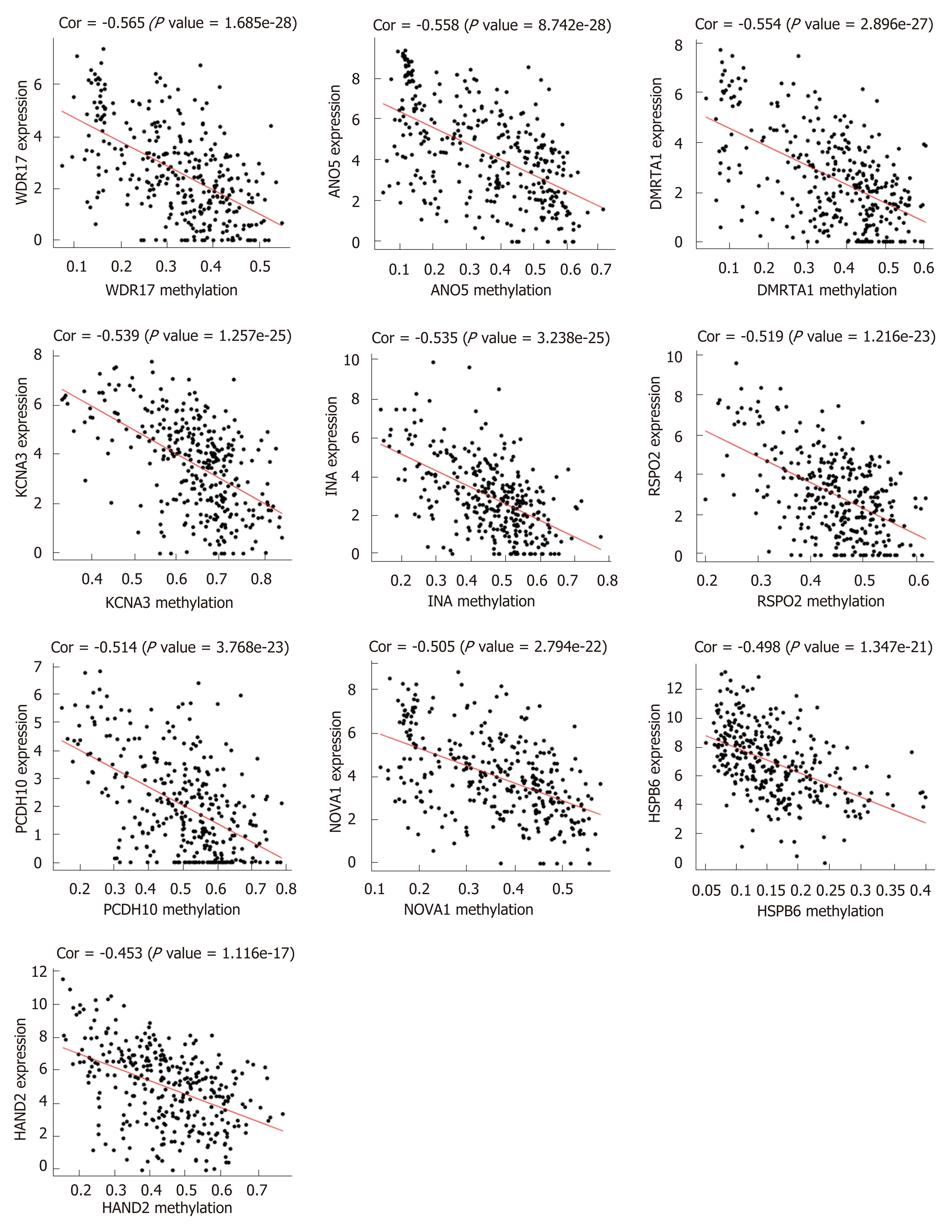
First, we identified 1205 up-regulated and 892 down-regulated DEGs from 473 colon cancer and 41 normal tissues. The DEGs are shown as a volcano plot in Figure 1A. Second, 997 hypermethylated and 637 hypomethylated DMGs were identified from 314 colon cancer and 37 normal tissues. The heat map of the top 50 DMGs is shown in Figure 1B. Third, 514 genes were identified that showed a negative correlation between methylation and expression value. Finally, 5 up-regulated and 81 down-regulated MeDEGs were identified that met the above three conditions. The Venn diagram and MeDEGs list are shown in Figure 2. The top 10 MeDEGs with the highest Spearman’s correlation coefficients are shown in Figure 3.
To further investigate the function of MeDEGs in colon cancer, we subjected the 86 MeDEGs to GO and KEGG pathway analyses using DAVID 6.8. The enrichment analyses of GO are summarized in Table 1. A total of 13 biological processes, 5 cellular components, and 8 molecular functions were enriched among the MeDEGs. As for biological process and molecular function, “positive regulation of neuron differentiation” and “transcription factor binding” were the most enriched terms in the respective categories. KEGG pathway enrichment analysis suggested that MeDEGs predominantly participate in cancer-related pathways including “transcriptional misregulation in cancer”, “cAMP signal pathway”, and “cGMP-PKG signaling pathway” (Table 2).
| Category | Term | Gene name | P-value |
| BP | GO:0045666-positive regulation of neuron differentiation | FEZF1, VWC2, NAP1L2, NKX2-5, BRINP3 | 4.08E-04 |
| BP | GO:0007268-chemical synaptic transmission | DRD5, NOVA1, GRIA4, KCNA1, SNAP91, NPY, HTR7 | 7.08E-04 |
| BP | GO:0006366-transcription from RNA polymerase II promoter | HAND1, FEV, PAX2, POU3F4, NKX2-5, IRF4, HAND2, PDX1, FOXF2 | 2.01E-03 |
| BP | GO:0045944-positive regulation of transcription from RNA polymerase II promoter | HAND1, NR5A2, TBX5, PAX2, GDNF, NKX2-5, FOXD3, GALR1, IRF4, HAND2, PDX1, FOXF2 | 4.04E-03 |
| BP | GO:0030154-cell differentiation | FEV, TLL1, COL19A1, NKX2-5, CHRDL1, SFRP5, SIM2, INA | 4.52E-03 |
| BP | GO:0045893-positive regulation of transcription, DNA-templated | FEZF1, NR5A2, TBX5, PAX2, NKX2-5, IRF4, HAND2, FOXF2 | 8.06E-03 |
| BP | GO:0007189-adenylate cyclase-activating G-protein coupled receptor signaling pathway | ADRB3, DRD5, GALR1 | 2.10E-02 |
| BP | GO:0051891-positive regulation of cardioblast differentiation | TBX5, NKX2-5 | 2.21E-02 |
| BP | GO:0043433-negative regulation of sequence-specific DNA binding transcription factor activity | HAND1, SFRP5, HAND2 | 2.95E-02 |
| BP | GO:0003337-mesenchymal to epithelial transition involved in metanephros morphogenesis | PAX2, GDNF | 3.09E-02 |
| BP | GO:0018108-peptidyl-tyrosine phosphorylation | EPHA7, EPHA6, RELN, EPHA5 | 3.11E-02 |
| BP | GO:0008285-negative regulation of cell proliferation | FEZF1, TBX5, SFRP5, RERG, RIPPLY3, PDX1 | 3.20E-02 |
| BP | GO:0000122-negative regulation of transcription from RNA polymerase II promoter | FEZF1, HAND1, NKX2-5, SIM2, RIPPLY3, DUSP26, FOXD3, PDX1 | 4.15E-02 |
| BP | GO:0030509-BMP signaling pathway | BMP3, NKX2-5, CHRDL1 | 4.54E-02 |
| CC | GO:0030425-dendrite | EPHA7, RELN, GRIA4, KCNA1, BRINP3, EPHA5, HTR7 | 4.25E-03 |
| CC | GO:0090575-RNA polymerase II transcription factor complex | HAND1, NR5A2, NKX2-5 | 9.87E-03 |
| CC | GO:0005578-proteinaceous extracellular matrix | RELN, COL19A1, CCBE1, EMILIN3, SPOCK3 | 3.40E-02 |
| CC | GO:0044224-juxtaparanode region of axon | KCNA1, EPB41L3 | 4.46E-02 |
| CC | GO:0005887-integral component of plasma membrane | JAM2, EPHA7, CALY, EPHA6, ADRB3, DRD5, GPR88, TMEM130, KCNA1, GALR1, EPHA5, HTR7 | 4.77E-02 |
| MF | GO:0008134-transcription factor binding | HAND1, TBX5, PAX2, NKX2-5, IRF4, HAND2, PDX1, FOXF2 | 1.43E-04 |
| MF | GO:0003700-transcription factor activity, sequence-specific DNA binding | DMRTA1, FEV, NR5A2, TBX5, POU3F4, NKX2-5, RXRG, SIM2, FOXD3, IRF4, PDX1, FOXF2 | 1.49E-03 |
| MF | GO:0000977-RNA polymerase II regulatory region sequence-specific DNA binding | TBX5, POU3F4, RXRG, FOXD3, HAND2, FOXF2 | 1.50E-03 |
| MF | GO:0043565-sequence-specific DNA binding | FEV, NR5A2, TBX5, NKX2-5, RXRG, IRX6, IRF4, FOXF2 | 4.74E-03 |
| MF | GO:0044212-transcription regulatory region DNA binding | HAND1, NR5A2, PAX2, NKX2-5, HAND2 | 1.07E-02 |
| MF | GO:0046982-protein heterodimerization activity | JAM2, HAND1, NKX2-5, SIM2, HAND2, PDX1, KHDRBS2 | 1.10E-02 |
| MF | GO:0000978-RNA polymerase II core promoter proximal region sequence-specific DNA binding | FEZF1, HAND1, NR5A2, NKX2-5, IRF4, PDX1 | 1.41E-02 |
| MF | GO:0005004-GPI-linked ephrin receptor activity | EPHA7, EPHA5 | 2.79E-02 |
| Pathway ID | Description | Gene name | P-value |
| hsa04970 | Salivary secretion | CST2, ADRB3, ATP1B2, PRKG2 | 2.80e-04 |
| hsa04923 | Regulation of lipolysis in adipocytes | NPY, ADRB3, PRKG2 | 1.12e-03 |
| hsa04080 | Neuroactive ligand-receptor interaction | GRIA4, ADRB3, DRD5, GALR1, HTR7 | 2.59e-03 |
| hsa04024 | cAMP signaling pathway | NPY, GRIA4, DRD5, ATP1B2 | 4.95e-03 |
| hsa04728 | Dopaminergic synapse | GRIA4, DRD5, CALY | 9.85e-03 |
| hsa04978 | Mineral absorption | MT1E, ATP1B2 | 1.41e-02 |
| hsa05202 | Transcriptional misregulation in cancer | FEZ, RXRG | 2.05e-02 |
| hsa04022 | cGMP-PKG signaling pathway | ADRB3, ATP1B2, PRKG2 | 2.12e-02 |
| hsa04360 | Axon guidance | EPHA7, EPHA5, EPHA6 | 2.28e-02 |
| hsa04020 | Calcium signaling pathway | ADRB3, DRD5, HTR7 | 2.31e-02 |
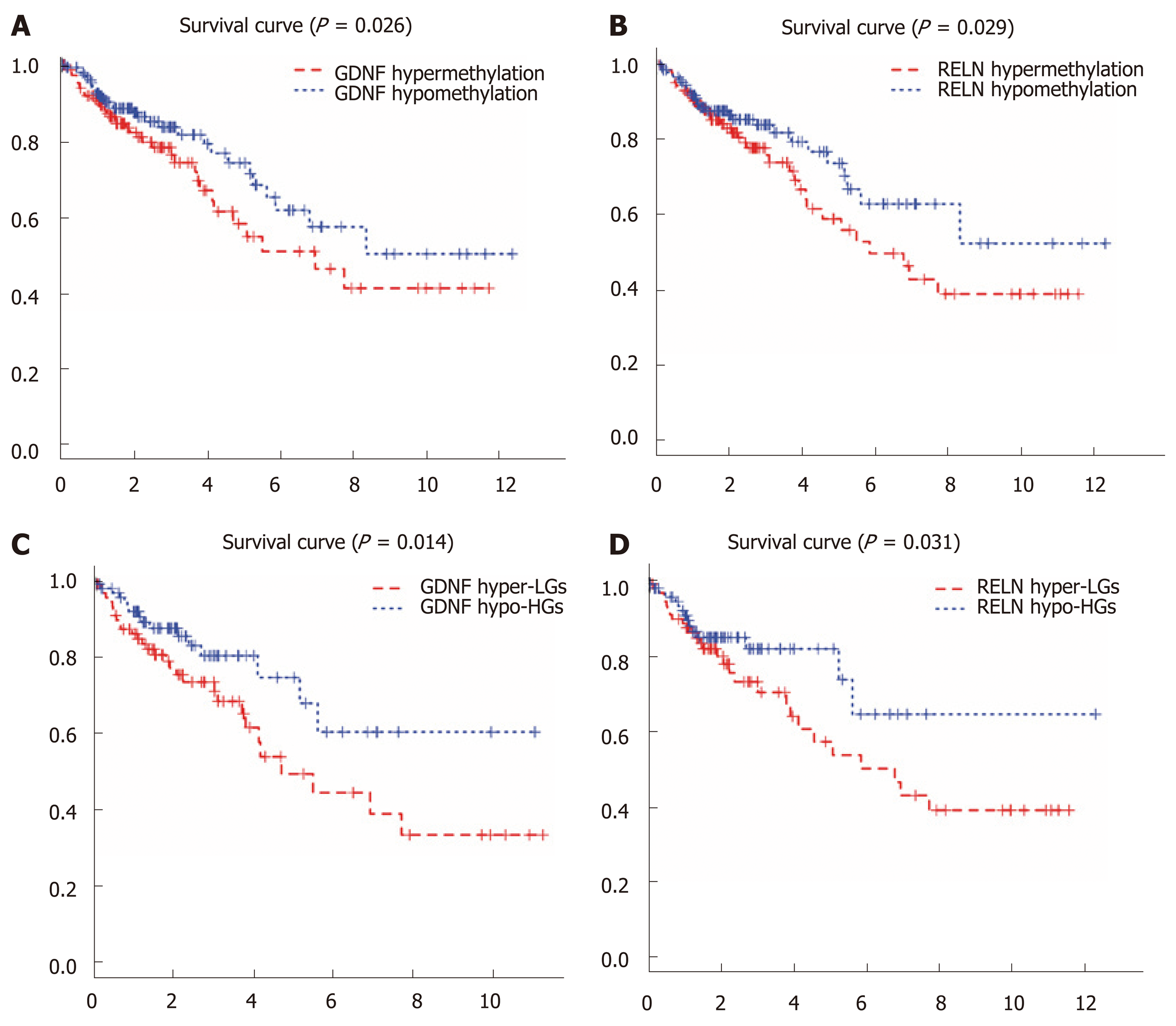
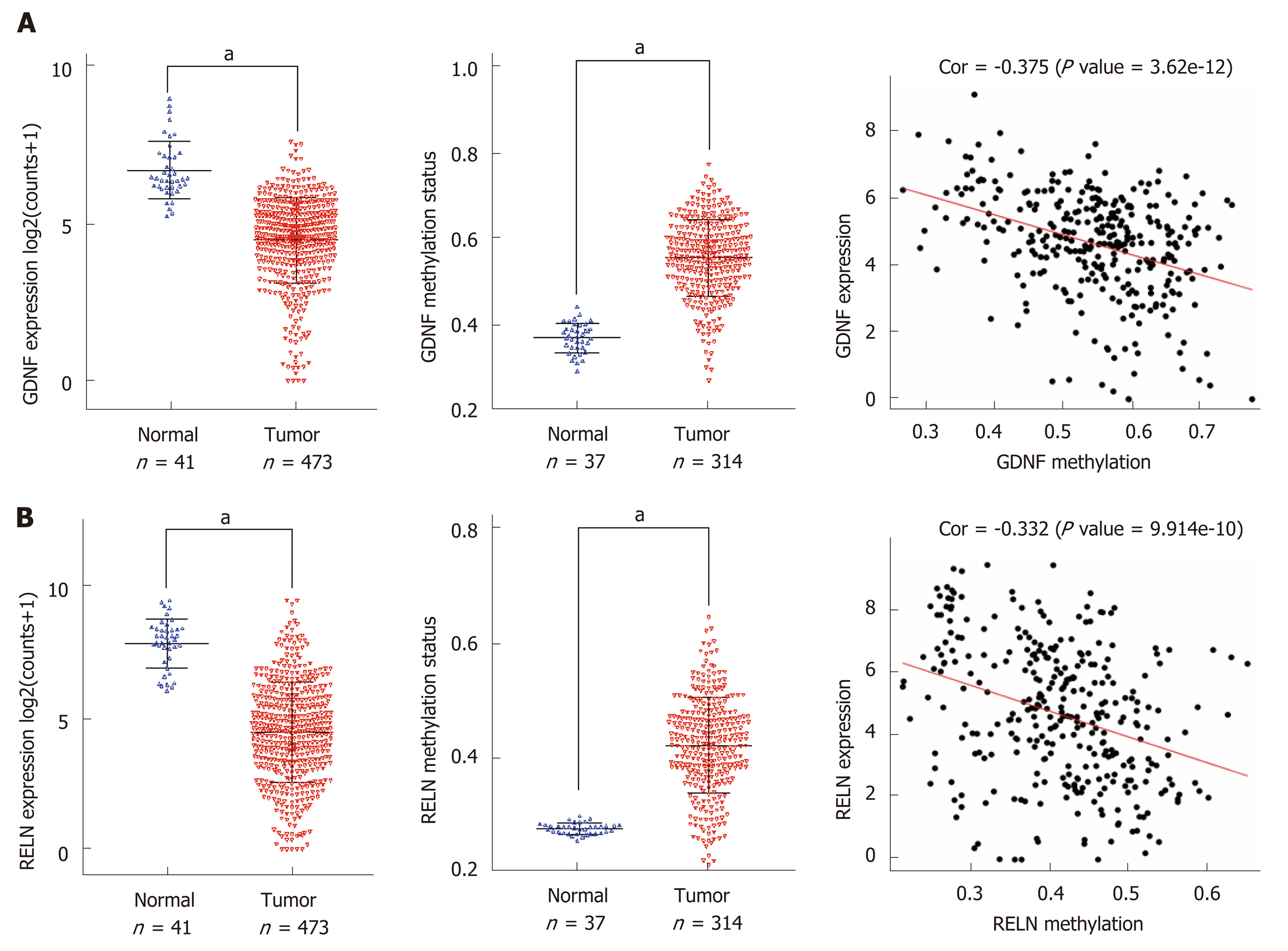
We performed a Kaplan–Meier curve analysis to identify the MeDEGs related to overall survival of colon cancer patients. First, we analyzed the relationship between MeDEG methylation value and prognosis. The Kaplan–Meier curves showed that hypermethylation of glial cell-derived neurotrophic factor (GDNF) and reelin (RELN) were negatively correlated with overall survival (Figure 4A and B). Next, we further compared the prognosis of the above two MeDEGs between the Hyper-LG group and the Hypo-HG group. The results showed that Hyper-LGs were significantly related to poor survival of patients (Figure 4C and D). The expression, methylation data, and Spearman’s correlation analyses of GDNF and RELN are shown in Figure 5.
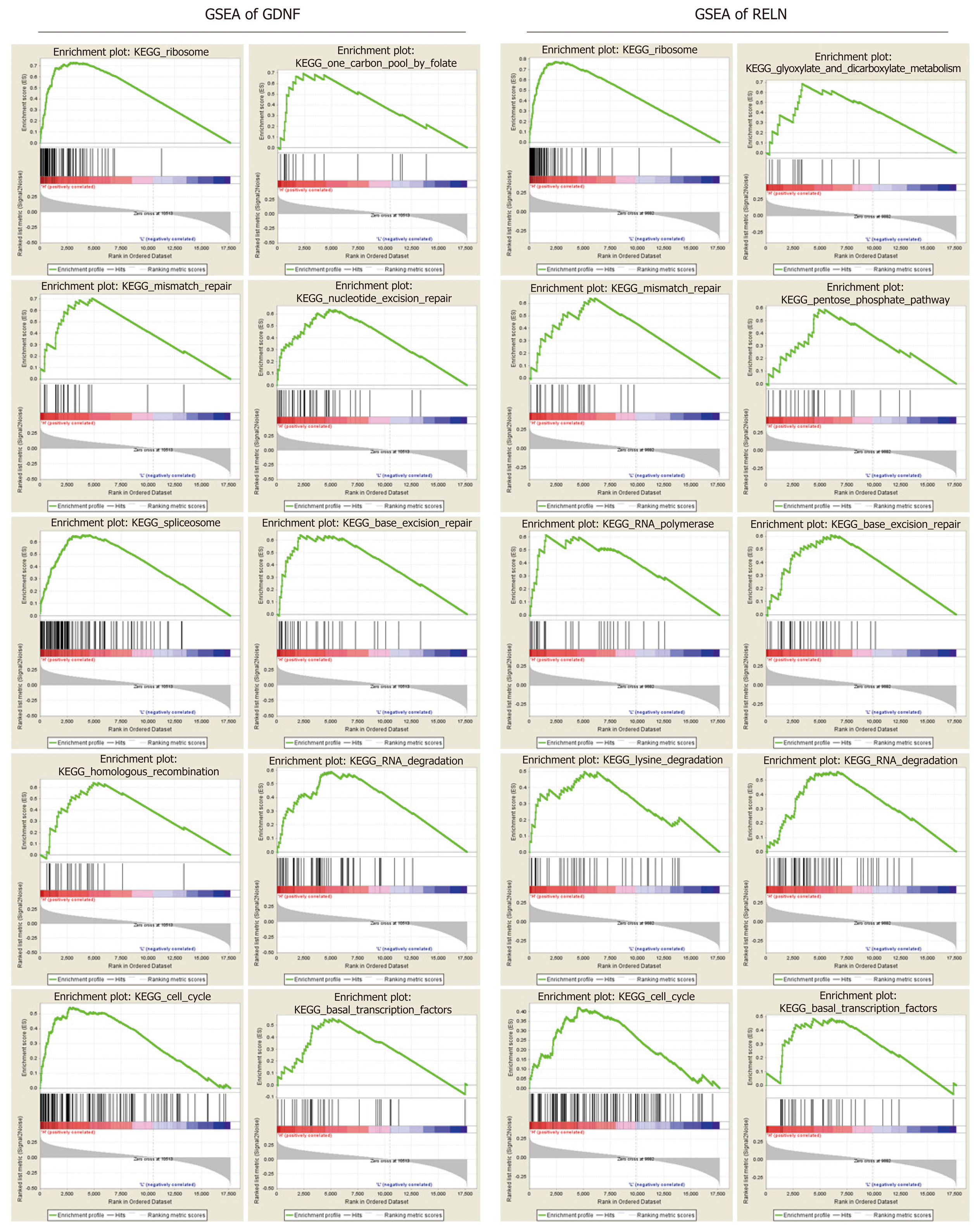
To clarify the biological function of GDNF and RELN methylation status, the effects of GDNF and RELN methylation status on KEGG pathways were analyzed using GSEA 3.0 software. The top ten GSEA results of GDNF and RELN are shown in Figure 6. Ultimately, hypermethylation of GDNF and RELN was both significantly associated with cancer-related pathways including “ribosome”, “RNA degradation”, “mismatch repair”, “cell cycle”, and “base excision repair”. More interestingly, the “ribosome” pathway was identified as the most significantly associated with GDNF and RELN methylation data.
Furthermore, we constructed PPI networks of GDNF and RELN using the STRING protein database. The PPI enrichment P-value was 1.74 × 10-5 (Figure 7). The results revealed that GDNF and RELN interact with neural cell adhesion molecule 1. Moreover, GDNF can interact with several proteins including GDNF family receptor alpha (GFRA)-1, -2, and -3 and the proto-oncogene tyrosine-protein kinase receptor (RET). Additionally, RELN can interact with low-density lipoprotein receptor-related protein 8, glutamate receptor ionotropic, NMDA 2B, disabled homolog 1, very low-density lipoprotein receptor, and platelet-activating factor acetylhydrolase IB subunit alpha.
Molecular pathological epidemiology (MPE), integrating molecular pathology and data science, can use tumor markers as surrogate of disease pathologies and help precision medicine. MPE deeply studies environmental exposures, intermediate variables, and molecular changes in cancer[13-15]. DNA methylation has been widely recognized as an important cancer-related biomarker and potential therapeutic target[16,17] in MPE. Previous studies mainly focused on the correlation between individual genes and methylation, so there was a lack of systematic analysis of DNA methylation in colon cancer. Thus, identification and analysis of MeDEGs in large cohorts of colon cancer patients are urgently required, providing potential directions and targets for future research.
In the present research, we identified a total of 5 up-regulated and 81 down-regulated MeDEGs by integrating DEGs, DMGs, and the results of Spearman’s correlation analysis. Among the 86 MeDEGs we report in this article, some of them have been confirmed to be regulated by methylation in previous research. For instance, the promoter of forkhead box D3 (FOXD3) has been reported to be hypermethylated in colon cancer and FOXD3 expression could be restored by treatment with 5-azacytidine[18,19]. The promoter methylation status of protocadherin 10 was related to disease-free survival and overall survival of colorectal patients, suggesting that it could be utilized as a biomarker for patients and to facilitate treatment decisions in colorectal cancer[20-22]. However, the methylation regulation mechanisms of the majority of MeDEGs have not been revealed and need to be further studied.
We further performed functional enrichment analysis to clarify the role of methylation in colon cancer. In biological process enrichment of GO, as many as 12 terms were related to the regulation of transcription. Therefore, we speculated that transcription factors regulated by methylation can further regulate the transcription of cancer-related genes and thus affect the occurrence and development of cancer. Aberrant cell proliferation is regarded as the distinguishing feature of cancer cells as opposed to normal cells. The term “negative regulation of cell proliferation” was enriched, meaning that multiple cancer suppressor genes were down-regulated by promoter hypermethylation. As shown in Table 1, MeDEGs including FEZ family zinc finger protein 1 (FEZF1), T-box transcription factor 5 (TBX5), secreted frizzled-related protein 5 (SFRP5), RAS-like estrogen regulated growth inhibitor (RERG), ripply transcriptional repressor 3 (RIPPLY3), and pancreatic and duodenal homeobox 1 (PDX1) were involved. Among them, TBX5[23], SFRP5[24,25], RERG[26-28], and PDX1[29] have been reported to be regulated by methylation in colon cancer, while FEZF1 and RIPPLY3 have not been reported. Bone morphogenetic proteins (BMPs), which form part of the transforming growth factor β signaling pathway, can promote colon cancer cell migration and invasion[30,31]. Moreover, methylation of BMP3[32,33] and NK2 homeobox 5[34] promoters has been revealed as an independent biomarker for colon cancer. However, apart from colon cancer, there have been no reports on the role of CHRDL1 promoter region methylation in tumors[35]. KEGG pathway analysis has further clarified the function of MeDEGs. The “cAMP signaling pathway” has been indicated as one of the most critical mechanisms in regulating colon cancer cell apoptosis[36-38]. The term “transcriptional misregulation in cancer” is consistent with the GO enrichment results. Babykutty et al[39] indicated that “cGMP-PKG signaling pathway” could promote colon cancer cell migration and invasion by upregulating matrix metalloprotein 2/9.
Moreover, we identified the methylation status of GDNF and RELN which was associated with clinical prognosis. GDNF is a neurotrophic factor which could guarantee the function of neuron in nervous systems[40]. According to TCGA database, GDNF is downregulated in colon cancer tissues. Interestingly, recent research indicated that GDNF promotes colon cancer cell proliferation and migration[41,42]. Luo et al[43] reported that GDNF was only detected in normal colon mucosa. Moreover, GDNF blocked apoptosis and suppression of anchorage-independent growth effects mediated by RET (rearranged during transfection). However, there has been no study on the correlation between GDNF methylation and the prognosis of colon cancer. RELN has been reported to play a critical role in maintaining epithelial cell ho-meostasis and protecting the colon from pathological processes[44]. Furthermore, RELN is epigenetically regulated by methylation in gastric cancer[45], pancreatic cancer[46], hepatocellular cancer[47], and myeloma[48]. However, the methylation state and function of RELN in colon cancer are still controversial.
Next, we performed GSEA to clarify the function of GDNF and RELN in KEGG pathways. The “ribosome” pathway is the most significantly enriched, which has been confirmed as an important mechanism in colon cancer development. For instance, precursor 45s ribosomal RNA (pre-45s rRNA) is up-regulated in colon cancer tissues and cell lines, and is associated with the prognosis of tumor patients. Pre-45s rRNA could promote cancer cell proliferation by inhibiting P53 by interfering in the interaction between murine double minute 2 (MDM2) and ribosomal protein L11 (RPL11)[49]. P73 is a p53 family tumor suppressor, which is promoted by RPL26 through direct binding to the 3’-UTR region. Furthermore, RPL26 maintains the protein stability of P73 by interacting with MDM2[50]. The “mismatch repair” and “base excision repair” pathways are primary mechanisms in DNA lesion repair and microsatellite instability in colon cancer[51]. Moreover, microsatellite-instability has been demonstrated to frequently occur in the hypermethylated cases. For instance, MLH1 (DNA mismatch repair protein Mlh1) methylation is a frequent molecular event and is closely linked to tumor invasion in veins and microsatellite-instability[52,53]. Moreover, “base excision repair” capacity[54] and “cell cycle”[55] pathway are also the critical mechanism in colon cancer progression. From the above, the methylation state and function of GDNF and RELN should be better elucidated and replicated in a larger validation cohort.
In conclusion, we identified MeDEGs by analyzing the expression profiles and methylation data of colon cancer samples from TCGA database. Functional enrichment analyses further confirmed the role of MeDEGs in colon cancer. Moreover, we revealed that GDNF and RELN are related to overall survival. GSEA and PPI networks further clarified the function of prognosis-related MeDEGs. Our study deepens the understanding of methylation and provides novel therapeutic targets and prognosis-related biomarkers for further research.
Accumulating evidence has indicated that DNA methylation modification is a reversible process of gene regulation in epigenetics. However, studies of methylation in the individual genes and pathways are still insufficient. In the present research, we conducted a conjoint analysis of correlation between methylation and gene expression and patient prognosis in large cohorts based on the Illumina Methylation 450K BeadChip.
DNA methylation modification has been considered as a potential therapeutic target and biomarker that may improve the prognosis of colon cancer. Therefore, identification and analysis of methylation-regulated differentially expressed genes (MeDEGs) will be of great significant.
In our study, we aimed to conduct bioinformatics analysis to identify MeDEGs and prognosis-related MeDEGs in colon cancer. Functional enrichment analysis was performed to clarify the function of MeDEGs. Furthermore, our study elucidated the potential mechanisms of prognosis-related MeDEGs.
We downloaded RNA expression profiles, Illumina Human Methylation 450K BeadChip data, and clinical data of colon cancer from The Cancer Genome Atlas project. Differentially expressed genes and differentially methylated genes were identified using with the “edgeR” package and the “limma” package in R software. Then, we performed Spearman’s correlation analysis to clarify the relationship between methylation and expression. The in silico function of MeDEGs was further analyzed in the DAVID database and Kyoto Encyclopedia of Genes and Genomes (KEGG) Orthology-Based Annotation System 3.0, respectively. The relationship between methylation and expression and overall survival was revealed through a Kaplan–Meier curve test. Gene set enrichment analysis (GSEA) and investigation of protein-protein interactions were performed to clarify the function of prognosis-related genes.
We identified a total of 5 up-regulated and 81 down-regulated MeDEGs that satisfied the conditions. Gene ontology analysis indicated that the enrichment terms are mainly associated with transcription regulation. According to KEGG pathway analysis, three cancer-related pathways were involved by MeDEGs. Hypermethylation of glial cell-derived neurotrophic factor (GDNF) and reelin (RELN) was negatively correlated with overall survival. Based on GSEA, hypermethylation of GDNF and RELN was both significantly associated with pathways including “RNA degradation,” “ribosome,” “mismatch repair,” “cell cycle”, and “base excision repair.”
In conclusion, we provide a new and reliable pathway to identify MeDEGs based on Illumina Human Methylation 450K BeadChip. Methylation plays a critical role in regulating cancer-related gene expression, especially tumor-suppressor genes. Our study provides an in-depth understanding of methylation. Furthermore, the prognosis-related MeDEGs may be potential biomarkers and therapeutic targets in colon cancer.
The application of high-throughput platform will provide great support for the precision medicine in cancer. We will conduct more studies to reveal the function of the prognosis-related MeDEGs and establish a valid and reliable high-throughput analysis system in the future.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report classification
Grade A (Excellent): A, A
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Bordonaro M, Kim TI, Linnebacher M, Ogino S S-Editor: Yan JP L-Editor: Wang TQ E-Editor: Zhang YL
| 1. | Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394-424. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53206] [Cited by in RCA: 55820] [Article Influence: 7974.3] [Reference Citation Analysis (132)] |
| 2. | Fearon ER. Molecular genetics of colorectal cancer. Annu Rev Pathol. 2011;6:479-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1163] [Cited by in RCA: 1271] [Article Influence: 90.8] [Reference Citation Analysis (1)] |
| 3. | Chen Y, Xie H, Gao Q, Zhan H, Xiao H, Zou Y, Zhang F, Liu Y, Li J. Colon cancer associated transcripts in human cancers. Biomed Pharmacother. 2017;94:531-540. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 24] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 4. | Hong SN. Genetic and epigenetic alterations of colorectal cancer. Intest Res. 2018;16:327-337. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 102] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 5. | Hervouet E, Peixoto P, Delage-Mourroux R, Boyer-Guittaut M, Cartron PF. Specific or not specific recruitment of DNMTs for DNA methylation, an epigenetic dilemma. Clin Epigenetics. 2018;10:17. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 115] [Cited by in RCA: 163] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 6. | Tanaka H, Kuwano Y, Nishikawa T, Rokutan K, Nishida K. ZNF350 promoter methylation accelerates colon cancer cell migration. Oncotarget. 2018;9:36750-36769. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 12] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 7. | Naumov VA, Generozov EV, Zaharjevskaya NB, Matushkina DS, Larin AK, Chernyshov SV, Alekseev MV, Shelygin YA, Govorun VM. Genome-scale analysis of DNA methylation in colorectal cancer using Infinium HumanMethylation450 BeadChips. Epigenetics. 2013;8:921-934. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 115] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 8. | Dumenil TD, Wockner LF, Bettington M, McKeone DM, Klein K, Bowdler LM, Montgomery GW, Leggett BA, Whitehall VL. Genome-wide DNA methylation analysis of formalin-fixed paraffin embedded colorectal cancer tissue. Genes Chromosomes Cancer. 2014;53:537-548. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 35] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 9. | Sandoval J, Heyn H, Moran S, Serra-Musach J, Pujana MA, Bibikova M, Esteller M. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics. 2011;6:692-702. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 763] [Cited by in RCA: 755] [Article Influence: 53.9] [Reference Citation Analysis (0)] |
| 10. | Wang Z, Jensen MA, Zenklusen JC. A Practical Guide to The Cancer Genome Atlas (TCGA). Methods Mol Biol. 2016;1418:111-141. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 219] [Cited by in RCA: 434] [Article Influence: 48.2] [Reference Citation Analysis (0)] |
| 11. | Gene Ontology Consortium. The Gene Ontology (GO) project in 2006. Nucleic Acids Res. 2006;34:D322-D326. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 671] [Cited by in RCA: 821] [Article Influence: 43.2] [Reference Citation Analysis (0)] |
| 12. | Draghici S, Khatri P, Tarca AL, Amin K, Done A, Voichita C, Georgescu C, Romero R. A systems biology approach for pathway level analysis. Genome Res. 2007;17:1537-1545. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 806] [Cited by in RCA: 939] [Article Influence: 52.2] [Reference Citation Analysis (0)] |
| 13. | Ogino S, Chan AT, Fuchs CS, Giovannucci E. Molecular pathological epidemiology of colorectal neoplasia: An emerging transdisciplinary and interdisciplinary field. Gut. 2011;60:397-411. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 430] [Cited by in RCA: 452] [Article Influence: 32.3] [Reference Citation Analysis (0)] |
| 14. | Ogino S, Lochhead P, Chan AT, Nishihara R, Cho E, Wolpin BM, Meyerhardt JA, Meissner A, Schernhammer ES, Fuchs CS, Giovannucci E. Molecular pathological epidemiology of epigenetics: Emerging integrative science to analyze environment, host, and disease. Mod Pathol. 2013;26:465-484. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 168] [Cited by in RCA: 167] [Article Influence: 13.9] [Reference Citation Analysis (0)] |
| 15. | Ogino S, Nowak JA, Hamada T, Milner DA, Nishihara R. Insights into Pathogenic Interactions Among Environment, Host, and Tumor at the Crossroads of Molecular Pathology and Epidemiology. Annu Rev Pathol. 2019;14:83-103. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 147] [Cited by in RCA: 176] [Article Influence: 29.3] [Reference Citation Analysis (0)] |
| 16. | Liang G, Weisenberger DJ. DNA methylation aberrancies as a guide for surveillance and treatment of human cancers. Epigenetics. 2017;12:416-432. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 90] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 17. | Weisenberger DJ, Liang G, Lenz HJ. DNA methylation aberrancies delineate clinically distinct subsets of colorectal cancer and provide novel targets for epigenetic therapies. Oncogene. 2018;37:566-577. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 62] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 18. | van Roon EH, Boot A, Dihal AA, Ernst RF, van Wezel T, Morreau H, Boer JM. BRAF mutation-specific promoter methylation of FOX genes in colorectal cancer. Clin Epigenetics. 2013;5:2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 29] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 19. | Sarkar S, O'Connell MR, Okugawa Y, Lee BS, Toiyama Y, Kusunoki M, Daboval RD, Goel A, Singh P. FOXD3 Regulates CSC Marker, DCLK1-S, and Invasive Potential: Prognostic Implications in Colon Cancer. Mol Cancer Res. 2017;15:1678-1691. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 30] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 20. | Zhong X, Zhu Y, Mao J, Zhang J, Zheng S. Frequent epigenetic silencing of PCDH10 by methylation in human colorectal cancer. J Cancer Res Clin Oncol. 2013;139:485-490. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 34] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 21. | Heitzer E, Artl M, Filipits M, Resel M, Graf R, Weißenbacher B, Lax S, Gnant M, Wrba F, Greil R, Dietze O, Hofbauer F, Böhm G, Höfler G, Samonigg H, Schaberl-Moser R, Balic M, Dandachi N. Differential survival trends of stage II colorectal cancer patients relate to promoter methylation status of PCDH10, SPARC, and UCHL1. Mod Pathol. 2014;27:906-915. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 22. | Zhong X, Shen H, Mao J, Zhang J, Han W. Epigenetic silencing of protocadherin 10 in colorectal cancer. Oncol Lett. 2017;13:2449-2453. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 11] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 23. | Yu J, Ma X, Cheung KF, Li X, Tian L, Wang S, Wu CW, Wu WK, He M, Wang M, Ng SS, Sung JJ. Epigenetic inactivation of T-box transcription factor 5, a novel tumor suppressor gene, is associated with colon cancer. Oncogene. 2010;29:6464-6474. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 75] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 24. | Belshaw NJ, Pal N, Tapp HS, Dainty JR, Lewis MP, Williams MR, Lund EK, Johnson IT. Patterns of DNA methylation in individual colonic crypts reveal aging and cancer-related field defects in the morphologically normal mucosa. Carcinogenesis. 2010;31:1158-1163. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 59] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 25. | Silva TD, Vidigal VM, Felipe AV, DE Lima JM, Neto RA, Saad SS, Forones NM. DNA methylation as an epigenetic biomarker in colorectal cancer. Oncol Lett. 2013;6:1687-1692. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 56] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 26. | Oster B, Thorsen K, Lamy P, Wojdacz TK, Hansen LL, Birkenkamp-Demtröder K, Sørensen KD, Laurberg S, Orntoft TF, Andersen CL. Identification and validation of highly frequent CpG island hypermethylation in colorectal adenomas and carcinomas. Int J Cancer. 2011;129:2855-2866. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 134] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 27. | Luo X, Huang R, Sun H, Liu Y, Bi H, Li J, Yu H, Sun J, Lin S, Cui B, Zhao Y. Methylation of a panel of genes in peripheral blood leukocytes is associated with colorectal cancer. Sci Rep. 2016;6:29922. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 22] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 28. | Andrew AS, Baron JA, Butterly LF, Suriawinata AA, Tsongalis GJ, Robinson CM, Amos CI. Hyper-Methylated Loci Persisting from Sessile Serrated Polyps to Serrated Cancers. Int J Mol Sci. 2017;18:pii: E535. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 14] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 29. | Wu J, Xiao Y, Xia C, Yang F, Li H, Shao Z, Lin Z, Zhao X. Identification of Biomarkers for Predicting Lymph Node Metastasis of Stomach Cancer Using Clinical DNA Methylation Data. Dis Markers. 2017;2017:5745724. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 25] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 30. | Deng H, Ravikumar TS, Yang WL. Overexpression of bone morphogenetic protein 4 enhances the invasiveness of Smad4-deficient human colorectal cancer cells. Cancer Lett. 2009;281:220-231. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 41] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 31. | Slattery ML, Lundgreen A, Herrick JS, Kadlubar S, Caan BJ, Potter JD, Wolff RK. Genetic variation in bone morphogenetic protein and colon and rectal cancer. Int J Cancer. 2012;130:653-664. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 24] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 32. | Koga Y, Yamazaki N, Matsumura Y. [Fecal Biomarker for Colorectal Cancer Diagnosis]. Rinsho Byori. 2015;63:361-368. [PubMed] |
| 33. | Park SK, Song CS, Yang HJ, Jung YS, Choi KY, Koo DH, Kim KE, Jeong KU, Kim HO, Kim H, Chun HK, Park DI. Field Cancerization in Sporadic Colon Cancer. Gut Liver. 2016;10:773-780. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 31] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 34. | Chung W, Kwabi-Addo B, Ittmann M, Jelinek J, Shen L, Yu Y, Issa JP. Identification of novel tumor markers in prostate, colon and breast cancer by unbiased methylation profiling. PLoS One. 2008;3:e2079. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 86] [Cited by in RCA: 98] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 35. | Pei YF, Zhang YJ, Lei Y, Wu DW, Ma TH, Liu XQ. Hypermethylation of the CHRDL1 promoter induces proliferation and metastasis by activating Akt and Erk in gastric cancer. Oncotarget. 2017;8:23155-23166. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 39] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 36. | Insel PA, Zhang L, Murray F, Yokouchi H, Zambon AC. Cyclic AMP is both a pro-apoptotic and anti-apoptotic second messenger. Acta Physiol (Oxf). 2012;204:277-287. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 163] [Cited by in RCA: 171] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 37. | Leiphrakpam PD, Brattain MG, Black JD, Wang J. TGFβ and IGF1R signaling activates protein kinase A through differential regulation of ezrin phosphorylation in colon cancer cells. J Biol Chem. 2018;293:8242-8254. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 19] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 38. | Hara M, Takeba Y, Iiri T, Ohta Y, Ootaki M, Watanabe M, Watanabe D, Koizumi S, Otsubo T, Matsumoto N. Vasoactive intestinal peptide increases apoptosis of hepatocellular carcinoma by inhibiting the cAMP/Bcl-xL pathway. Cancer Sci. 2019;110:235-244. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 25] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 39. | Babykutty S, Suboj P, Srinivas P, Nair AS, Chandramohan K, Gopala S. Insidious role of nitric oxide in migration/invasion of colon cancer cells by upregulating MMP-2/9 via activation of cGMP-PKG-ERK signaling pathways. Clin Exp Metastasis. 2012;29:471-492. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 77] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 40. | Henderson CE, Phillips HS, Pollock RA, Davies AM, Lemeulle C, Armanini M, Simmons L, Moffet B, Vandlen RA, Simpson LC corrected to Simmons L, Koliatsos VE, Rosenthal A. GDNF: A potent survival factor for motoneurons present in peripheral nerve and muscle. Science. 1994;266:1062-1064. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 918] [Cited by in RCA: 972] [Article Influence: 31.4] [Reference Citation Analysis (0)] |
| 41. | Huang SM, Chen TS, Chiu CM, Chang LK, Liao KF, Tan HM, Yeh WL, Chang GR, Wang MY, Lu DY. GDNF increases cell motility in human colon cancer through VEGF-VEGFR1 interaction. Endocr Relat Cancer. 2013;21:73-84. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 36] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 42. | Zhong F, Zhang W, Cao Y, Wen Q, Cao Y, Lou B, Li J, Shi W, Liu Y, Luo R, Chen C. LncRNA NEAT1 promotes colorectal cancer cell proliferation and migration via regulating glial cell-derived neurotrophic factor by sponging miR-196a-5p. Acta Biochim Biophys Sin (Shanghai). 2018;50:1190-1199. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 32] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 43. | Luo Y, Tsuchiya KD, Il Park D, Fausel R, Kanngurn S, Welcsh P, Dzieciatkowski S, Wang J, Grady WM. RET is a potential tumor suppressor gene in colorectal cancer. Oncogene. 2013;32:2037-2047. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 58] [Cited by in RCA: 72] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 44. | Carvajal AE, Serrano-Morales JM, Vázquez-Carretero MD, García-Miranda P, Calonge ML, Peral MJ, Ilundain AA. Reelin protects from colon pathology by maintaining the intestinal barrier integrity and repressing tumorigenic genes. Biochim Biophys Acta Mol Basis Dis. 2017;1863:2126-2134. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 20] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 45. | Dohi O, Takada H, Wakabayashi N, Yasui K, Sakakura C, Mitsufuji S, Naito Y, Taniwaki M, Yoshikawa T. Epigenetic silencing of RELN in gastric cancer. Int J Oncol. 2010;36:85-92. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 46. | Sato N, Fukushima N, Chang R, Matsubayashi H, Goggins M. Differential and epigenetic gene expression profiling identifies frequent disruption of the RELN pathway in pancreatic cancers. Gastroenterology. 2006;130:548-565. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 108] [Cited by in RCA: 120] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 47. | Okamura Y, Nomoto S, Kanda M, Hayashi M, Nishikawa Y, Fujii T, Sugimoto H, Takeda S, Nakao A. Reduced expression of reelin (RELN) gene is associated with high recurrence rate of hepatocellular carcinoma. Ann Surg Oncol. 2011;18:572-579. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 48. | Lin L, Wang P, Liu X, Zhao D, Zhang Y, Hao J, Liang X, Huang X, Lu J, Ge Q. Epigenetic regulation of reelin expression in multiple myeloma. Hematol Oncol. 2017;35:685-692. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 49. | Tsoi H, Lam KC, Dong Y, Zhang X, Lee CK, Zhang J, Ng SC, Ng SSM, Zheng S, Chen Y, Fang J, Yu J. Pre-45s rRNA promotes colon cancer and is associated with poor survival of CRC patients. Oncogene. 2017;36:6109-6118. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 36] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 50. | Zhang M, Zhang J, Yan W, Chen X. p73 expression is regulated by ribosomal protein RPL26 through mRNA translation and protein stability. Oncotarget. 2016;7:78255-78268. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 15] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 51. | Koi M, Okita Y, Carethers JM. Fusobacterium nucleatum Infection in Colorectal Cancer: Linking Inflammation, DNA Mismatch Repair and Genetic and Epigenetic Alterations. J Anus Rectum Colon. 2018;2:37-46. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 54] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 52. | Ma Y, Chen Y, Petersen I. Expression and promoter DNA methylation of MLH1 in colorectal cancer and lung cancer. Pathol Res Pract. 2017;213:333-338. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 30] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 53. | Li X, Yao X, Wang Y, Hu F, Wang F, Jiang L, Liu Y, Wang D, Sun G, Zhao Y. MLH1 promoter methylation frequency in colorectal cancer patients and related clinicopathological and molecular features. PLoS One. 2013;8:e59064. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 62] [Cited by in RCA: 78] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 54. | Vodenkova S, Jiraskova K, Urbanova M, Kroupa M, Slyskova J, Schneiderova M, Levy M, Buchler T, Liska V, Vodickova L, Vymetalkova V, Collins A, Opattova A, Vodicka P. Base excision repair capacity as a determinant of prognosis and therapy response in colon cancer patients. DNA Repair (Amst). 2018;72:77-85. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 25] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 55. | Eyvani H, Moghaddaskho F, Kabuli M, Zekri A, Momeny M, Tavakkoly-Bazzaz J, Alimoghaddam K, Ghavamzadeh A, Ghaffari SH. Arsenic trioxide induces cell cycle arrest and alters DNA methylation patterns of cell cycle regulatory genes in colorectal cancer cells. Life Sci. 2016;167:67-77. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 30] [Article Influence: 3.3] [Reference Citation Analysis (0)] |