Published online Nov 14, 2017. doi: 10.3748/wjg.v23.i42.7505
Peer-review started: May 23, 2017
First decision: June 23, 2017
Revised: July 31, 2017
Accepted: August 15, 2017
Article in press: August 15, 2017
Published online: November 14, 2017
Processing time: 173 Days and 20 Hours
Celiac disease (CD) is a chronic immune-mediated disorder triggered by the ingestion of gluten in genetically predisposed individuals. Before activating the immune system, gluten peptides are transferred by the epithelial barrier to the mucosal lamina propria, where they are deamidated by intestinal tissue transglutaminase 2. As a result, they strongly bind to human leucocyte antigens (HLAs), especially HLA-DQ2 and HLA-DQ8, expressed on antigen-presenting cells. This induces an inflammatory response, which results in small bowel enteropathy. Although gluten is the main external trigger activating both innate and adaptive (specific) immunity, its presence in the intestinal lumen does not fully explain CD pathogenesis. It has been hypothesized that an early disruption of the gut barrier in genetically susceptible individuals, which would result in an increased intestinal permeability, could precede the onset of gluten-induced immune events. The intestinal barrier is a complex functional structure, whose functioning is dependent on intestinal microbiota homeostasis, epithelial layer integrity, and the gut-associated lymphoid tissue with its intraepithelial lymphocytes (IELs). The aim of this paper was to review the current literature and summarize the role of the gut microbiota, epithelial cells and their intercellular junctions, and IELs in CD development.
Core tip: There is evidence that the host-microbiota homeostasis is disrupted in celiac disease (CD) patients. Dysbiosis, meaning an imbalance in the gut microbiota and its metabolome, may activate innate immunity leading to pro-inflammatory changes, which induces intraepithelial lymphocyte infiltration and epithelial barrier damage, ultimately resulting in increased transfer of gluten peptides and inflammatory activation leading to CD development. The intestinal microbiota also has a direct effect on the breakdown of gluten and formation of immunogenic peptides. As colonization of the gut with microorganisms may be dependent on genetic factors, future prophylactic strategies may focus on gut microbiota modulation in genetically predisposed infants.
- Citation: Cukrowska B, Sowińska A, Bierła JB, Czarnowska E, Rybak A, Grzybowska-Chlebowczyk U. Intestinal epithelium, intraepithelial lymphocytes and the gut microbiota - Key players in the pathogenesis of celiac disease. World J Gastroenterol 2017; 23(42): 7505-7518
- URL: https://www.wjgnet.com/1007-9327/full/v23/i42/7505.htm
- DOI: https://dx.doi.org/10.3748/wjg.v23.i42.7505
Celiac disease (CD) is a chronic immune-mediated disorder triggered by the ingestion of gluten in genetically predisposed individuals[1]. Gluten is a storage protein that consists of alcohol-insoluble glutenins and soluble prolamines, such as gliadin in wheat, secalin in rye, and hordein in barley. CD development requires the presence of gluten, the intestinal enzyme tissue transglutaminase 2 (TTG2), which modifies gluten peptides, and the genes encoding human leucocyte antigen (HLA)-DQ2 or HLA-DQ8[2]. Gluten from food products is degraded by gastrointestinal tract enzymes into peptides, which then are transferred through the epithelial barrier into the mucosal lamina propria.
In CD individuals, some of these peptides can bind to HLA-DQ2 or HLA-DQ8 heterodimers expressed on the surface of antigen-presenting cells (e.g., macrophages, lymphocytes or dendritic cells) and, after triggering T-cell responses, lead to local tissue damage[3]. TTG2 converts glutamine residues present in gluten peptides into glutamic acid, and this conversion generates deamidated gluten peptides (DGP), which strongly bind to HLA-DQ2/-DQ8 molecules. Consequently, increased gluten antigenicity amplifies a gluten-specific T-cell response.
Gluten-activated T cells release pro-inflammatory cytokines [(mainly interferon-gamma (IFN-γ), interleukin (IL)-21 and IL-17)], which induce mucosal inflammation and have a direct cytotoxic effect on the epithelium, all of which finally leads to villous atrophy in the small intestine. Moreover, specific T cells induce B cells to produce antibodies directed against DGP and TTG2[4]. Thus, this adaptive (specific) T-cell response is a requirement for CD development. Nonetheless, innate immunity also plays an important role in CD development. The increased transfer of gluten peptides through the epithelial barrier could be a consequence of earlier activation of innate (non-specific) immunity, dependent on the function of both the epithelium and the lymphocytes located between epithelial cells, i.e. intraepithelial lymphocytes (IELs)[5].
Some of the gluten peptides can directly react with epithelial cells and activate production of pro-inflammatory cytokines, especially IL-15. IL-15 plays a key role in enhanced cytolytic activity of IELs via increasing the expression of both intestinal epithelial cell surface ligands (such as MICA and MICB, i.e. major histocompatibility complex class I chain-related molecules), which are targeted by cytotoxic, natural killer (NK)-like IELs, and NK receptors, such as NKG2D and CD94/NKG2C, on the surface of IELs. Finally, IL-15 activation leads to innate cytotoxic disruption of epithelial cells, resulting in increased intestinal permeability to different luminal macromolecules, including immunogenic gluten peptides[6].
Although gluten is the main external trigger of CD, gluten ingestion does not fully explain CD pathogenesis. Introduction of gluten into the diet starts in early childhood, but CD can develop at any point during a person’s lifetime. The role of both breastfeeding and the time when gluten is first introduced into the diet in the risk of CD has long been debated. Retrospective data from Sweden indicated that introducing gluten in small amounts to breastfed infants at the age between 4 mo and 6 mo reduced the risk of CD compared with introducing gluten in larger amounts at older ages[7,8]. However, a recently published systematic review with meta-analysis of studies that assessed the effect of gluten consumption on CD development showed that for infants at high genetic risk of CD, gluten introduction at the age of 4 mo, 6 mo or 12 mo, resulted in similar rates of CD diagnosis in childhood, and neither breastfeeding as such (at any time during an infant’s life) nor breastfeeding during gluten introduction were shown to reduce the risk of CD[9]. Also, the recently published prospective PreventCD cohort study showed that neither the gluten consumption pattern nor the amount of gluten consumed at the age of 11 mo to 36 mo influenced CD development in children with a genetic risk[10].
Thus, the time of gluten introduction into the diet seems not to play a key role in CD development. In addition, gluten-free diet (GFD) has been reported to improve mucosal lesions and decrease specific antibody levels, but not to correct the increased activation of pro-inflammatory mediators, which is characteristic for CD[11]. That is why it has been hypothesized that an early disruption of the gut barrier in genetically susceptible individuals, which is not associated with gluten peptides and results in an increased intestinal permeability, could precede the onset of gluten-induced immune events.
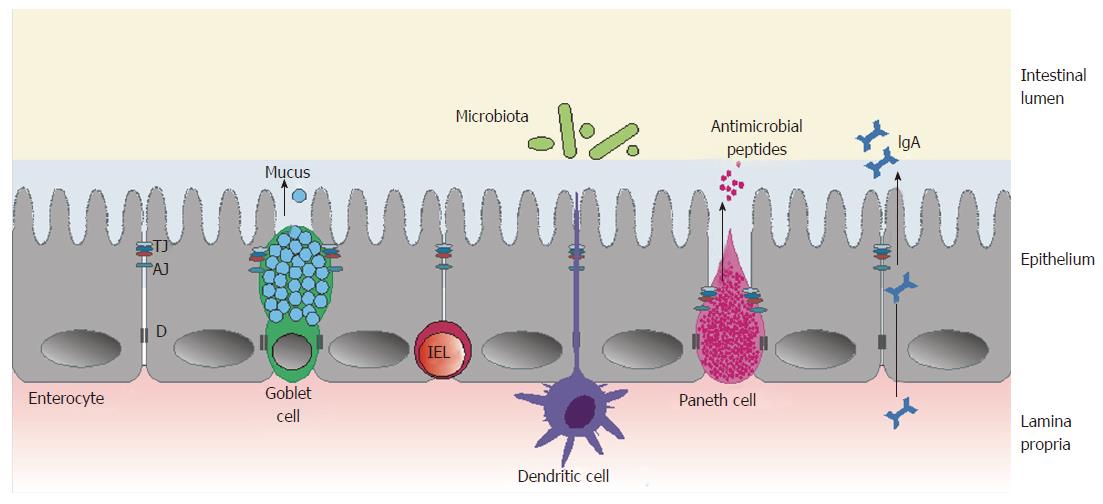
The intestinal barrier is a complex structure that separates the internal milieu from the luminal environment[12]. It consists of three main functional components: the microbiota that colonize the intestines; the epithelium, with its specialized mucus-producing cells and cells producing antimicrobial peptides; and gut-associated lymphoid tissue, composed of various immune cells (including IELs, which come in direct contact with gut luminal antigens, and lamina propria cells, producing secretory IgA) (Figure 1).
This review summarizes the role of epithelial cells and their intercellular junctions as well as IELs and the gut microbiota in the activation of early processes leading to the pathomechanisms associated with CD.
The small intestinal epithelium is organized into a monolayer of specialized cells: enterocytes (constituting approximately 80%), goblet cells (secreting mucus), Paneth cells (synthesizing defensins and other antimicrobial agents), endocrine cells (secreting hormones), and intestinal stem cells (responsible for epithelial cell homeostasis and regeneration)[13,14]. Epithelial cells form a continuous layer thanks to being sealed together by intercellular junctions, including tight junctions (TJs), adherens junctions (AJs), desmosomes, and gap junctions[15].
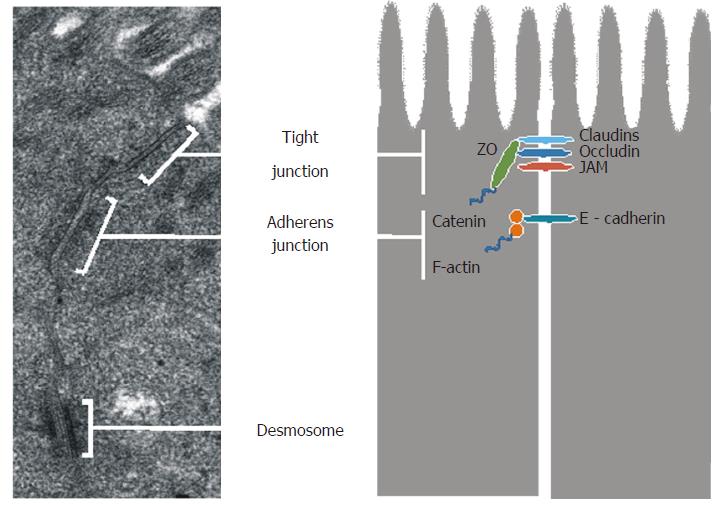
The ultrastructure of epithelial junctions is presented in Figure 2. TJs and AJs are supported by a dense perijunctional ring of actin and myosin, and they form the apical junctional complex and regulate epithelial paracellular permeability[16,17]. TJs are located near the apical surface of enterocytes and they act as a gate in the paracellular transport of ions, solutes, water, and cells. TJs are highly dynamic structures, whose degree of sealing varies in response to external stimuli as well as physiological and pathological conditions. TJs are composed of transmembrane proteins: occludin, claudins, junction adhesion molecules, tricellulin, and scaffold proteins - zonula occludens (ZO-1, ZO-2 and ZO-3)[17].
Occludin is an integral membrane protein with two extracellular loops, a short cytoplasmic N-terminal region, and a long cytoplasmic C-terminal region, which interacts with a ZO-1 protein that links occludin to the actin cytoskeleton[18,19]. Occludin plays a role in TJ maintenance and assembly, which are regulated by phosphorylation of serine (Ser), threonine (Thr), and tyrosine (Tyr) residues[20]. In an intact epithelium, occludin is highly phosphorylated on Ser and Thr residues[21,22] and poorly phosphorylated on Tyr residues[23]. Dephosphorylation of Ser/Thr residues and increased phosphorylation of Tyr residues reduces occludin’s interaction with ZO-1, leading to its separation from the junctional complex and TJ disruption[24,25].
The claudin family can be divided into sealing proteins (claudins 1, 3, 4, 5 and 8), which reduce permeability, and pore-forming proteins (claudins 2, 7, 10 and 12), which increase permeability[26]. Thus, claudins 1, 3, 4, 5 and 8 strengthen the intestinal barrier, whereas claudins 2, 7, 10 and 12 weaken it. The extracellular loops of claudins are involved in the formation of ion-selective channels[27], while the intracellular C-terminal domain is connected to the cytoskeleton via a domain containing ZO-1, ZO-2 and ZO-3[28,29]. ZO-1, ZO-2 and ZO-3 are multidomain bridging proteins that function as cross-linkers, anchoring the TJ strand proteins to the actin cytoskeleton[30].
Recently, tricellulin has been identified as a component maintaining TJ structure and regulating the passage of macromolecules through the junctions[31]. TJ development may be dependent on AJ formation, since the ability of ZO-1 proteins to migrate apically to join occludin was observed only after AJ assembly[32]. The main component of AJ is E-cadherin, a transmembrane protein that forms homodimers with other cadherin molecules on adjacent cells. This protein is connected to the actin cytoskeleton by a complex of cytoplasmic proteins: α-, β- and γ-catenins[33].
Despite the major progress in knowledge on TJ structure and function, the mechanisms regulating TJs are still incompletely understood. The discovery of the Vibrio cholerae-derived Zonula occludens toxin, which reversibly regulates TJ permeability, helped identify its intestinal mammalian analogue - a human protein named zonulin[34,35]. Zonulin was identified as pre-haptoglobin 2. Structural analysis of this protein revealed similarities with several growth factors, such as hepatocyte growth factor or epidermal growth factor, which affect intercellular TJ integrity[36,37].
Zonulin was shown to induce TJ disassembly and a subsequent increase in intestinal permeability. Zonulin transactivates the epidermal growth factor receptor through proteinase-activated receptor 2, and then activates phospholipase C, which hydrolyzes phosphatidylinositol to release inositol 1, 4, 5-tris phosphate and diacylglycerol[38,39]. Protein kinase Cα is then activated, either directly (via diacylglycerol) or through the release of intracellular calcium ions (via inositol 1, 4, 5-tris phosphate). Membrane-associated, activated protein kinase Cα catalyzes the phosphorylation of target proteins, including ZO-1 and myosin 1C, as well as polymerization of soluble G-actin in F-actin. This polymerization results in actin filament rearrangement and subsequent displacement of proteins (including ZO-1) from the junctional complex. As result, intestinal TJs become looser, which increases the paracellular transport of luminal molecules[35].
Zonulin is over-expressed in tissues and sera of subjects affected by autoimmune diseases, including CD[35]. In vitro studies showed that increased zonulin release in the small intestine can be triggered by both gluten peptides[38,39] and enteric bacteria[40]. Zonulin secretion has been demonstrated to be independent of either the species or the virulence of the microorganisms tested[40]. However, recently an association of low serum zonulin levels with lower quantities of Bacteroidaceae and Veillonellaceae and higher quantities of Faecalibacterium has been found in overweight pregnant women[41]. Thus, this in vivo study suggests that zonulin release could be affected by changes in gut microbiota composition.
Recently, epithelial polarity regulators, especially the Par-3 protein, have been reported to be likely involved in regulating TJ permeability[42]. Par-3 and other proteins regulating cell polarity, such as Par-6 and atypical protein kinase C, form the apical polarity complex that orchestrates the formation of apical junctional complex. In addition, Par-3 located in the junctional complex together with ZO-1 and catenins is able to affect TJs by rearranging the actin cytoskeleton. Schumann et al[43] in 2012 found a reduced level of Par-3 and a defect in performing lateral exclusion of Par-3 in the epithelial cells of CD patients. In this context, genetic studies on non-HLA gene candidates associated with CD seem to be very interesting. Wapenaar et al[44] in 2008 found two candidate genes: Par-3 and Magi2, encoding the proteins regulating of epithelial polarity. However, this study involved a homogenous Dutch population, and further genome-wide association studies did not confirm this association[45].
One of the first studies on the structure of epithelial junctions using freeze-fracture electron micrographs presented severely altered TJs with strand discontinuities and a reduced number of strands in children with active CD[46]. GFD improved these abnormalities, but only partially - strand numbers were restored to normal at the surface, but remained low in the crypts. The recent transmission electron microscopy analyses on duodenal biopsies of CD patients also showed changes in TJ ultrastructure: dilatation (saccular or fusiform) and destruction of pentalaminar structures[47]. Interestingly, ultrastructural abnormalities of TJs were also found in asymptomatic and serologically negative first-degree relatives of CD patients[48].
Furthermore, over-expression of occludin and the pore-forming protein claudin-2 was demonstrated in CD patients, as well as an under-expression of pore-sealing proteins claudin-3 and 4, and scaffold protein ZO-1[47,49,50]. After introduction of a GFD, normalization of claudin expression was observed. No improvement after GFD introduction was reported in about 3% of the patients with refractory CD, whose mucosa undergoes a constant inflammatory process[51]. Other studies indicated a subcellular localization and downregulation of claudin 4 and claudin 5 in refractory CD patients[52].
Alterations in AJ structure were also reported. The expression of E-cadherin and β catenin - proteins required for TJ formation - was shown to be reduced in the duodenal epithelium of children with CD. Ciccocioppo et al[50] in 2006 showed that a lack of ZO-1 phosphorylation in active CD led to TJ disruption. The authors suggested that non-phosphorylated ZO-1 was unable to detach from β-catenin and to connect with occludin. It was also found that a higher phosphorylation of β-catenin was responsible for the absence of membranous E-cadherin. On the other hand, highly phosphorylated β-catenin was unable to connect with E-cadherin, which, in turn, could bind to the αEβ7-integrin of IELs. However, the levels of both E-cadherin and β-catenin returned to normal following GFD introduction[53,54]. Interestingly, a recent study by Mishra et al[48] in 2015 indicated the presence of altered ZO-1 and occludin expression not only in active CD patients but also in asymptomatic and serologically negative first-degree relatives of CD patients.
Fasano et al[35] in 2000 tried to explain the increased expression of zonulin found in CD patients. Some studies suggested that gliadin, by binding to the proinflammatory chemokine CXCR3 receptor on the intestinal epithelium, initiates the release of zonulin, which induces cytoskeleton rearrangement, ZO-1 and occludin down-regulation, leading to disruption of TJ integrity and finally to an increase in epithelial permeability[38,55]. Thus, the receptor CXCR3 could be involved in early TJ dysfunction, preceding the immune cascade of events observed in CD patients. Recently, Bondar et al[55] in 2014 showed that CXCL10 - a ligand for CXCR - is over-expressed in the small intestine of CD patients and strongly activated by poly I:C (an experimental model of viral infections) and IL-15 in non-CD controls. Thus, it cannot be excluded that the CXCR3/CXCL10 axis activated by infectious agents may play a role in initiating gluten-induced inflammatory processes in the small intestinal mucosa.
Overall, the presented results show that epithelial barrier impairment occurring in CD patients can play an important role in CD development. Because epithelial function is regulated by microorganisms colonizing the intestines[56], there is a hypothesis that dysbiosis, i.e. disturbances in both the quantity and composition of the gut microbiota, is a critical factor for the activation of innate immunity, leading to epithelial barrier dysfunctions.
The microbiota colonizing the gut after birth reaches the pattern found in adults within 2-3 years of life. Eventually, the human intestine is colonized with more than 1000 species categorized into subgroups of phyla, classes, orders, families and genera, with Firmicutes and Bacteroidetes constituting the most abundant phyla[57]. The number of bacteria in the gut microbiota is similar to the number of cells making up the human body[58], and microbiota genes (microbiome) outnumber those in the human genome by approximately 100-fold. This complex microbial community adjusts the immune system, protects the body against pathogens, harvests nutrients and energy from the diet, and ferments non-digestible carbohydrates.
Extensive studies in germ-free (GF) animals, i.e. animals deprived of the gut microbiota, have demonstrated an indispensable role of microbiota in shaping the local mucosal gut-associated lymphoid tissue as well as systemic immunity[59,60]. In contrast to conventionally raised (CV) mice, GF mice have hypoplastic Peyer’s patches and decreased number of both IgA-secreting plasma cells and lymphocytes located in the lamina propria. Colonization of GF animals with components of the gut microbiota induces production of secretory immunoglobulins A (sIgA). sIgAs are natural antibodies that constitute the first line of defense by reacting with a wide spectrum of microorganisms and toxic molecules, which directly affects the composition of the gut microbiota[61]. Experimental data have shown that sIgAs cooperate with innate defense factors to reinforce the epithelial barrier[62].
Epithelial barrier integrity also depends on homeostatic regulatory mechanisms, including mucosal induction of regulatory T (Treg) cells, and the gut microbiota plays a decisive role in this process[63]. According to some reports, gut-colonizing commensals are responsible for differentiation of effector T helper (Th) 1, Th17, and Treg cells responsible for Th1/Th2/Th17 homeostasis[64]. Colonization of GF mice with components of conventional microbiota also induced the recruitment and activation of IELs, some of which (especially γδ IELs) were reported to be involved in epithelial cell generation and differentiation[65,66]. Thus, the gut microbiota seems capable of protecting the epithelium and strengthening its barrier function[59].
Recently, using transmission electron microscopy, we found ultrastructural differences of enterocytes and epithelial junctions in GF mice, CV or specific pathogen-free (SPF) mice, and mice inoculated with a mixture of Lactobacillus strains obtained from stools of healthy children[61]. Brush borders of GF-mouse enterocytes were irregularly arranged and exhibited decreased numbers of cytoskeletal microfilaments and a lack of elongation into the terminal web. The AJ region was significantly broader and shorter in GF animals compared both with that in CV mice and in mice colonized with Lactobacillus strains. Consistent with other reports[67,68], we observed that the gut microbiota and Lactobacillus strains significantly increased the expression of TJ proteins: occludin and ZO-1[61]. On the other hand, there is experimental evidence that certain components of the gut microbiota, such as Escherichia coli, Klebsiella pneumoniae and Streptococcus viridans, are able to increase gut permeability[69].
The gut microbiota interacts with the host via pattern recognition receptors (PRRs), including Toll-like receptors (TLRs) expressed on the surface of epithelial and dendritic cells. Recognition of specific microbial structures, called microorganism-associated molecular patterns, by PRRs induces signaling cascades that eventually result in immune response activation and the production of cytokines responsible for intestinal barrier strengthening (e.g., TGF-β and IL-10) or weakening (e.g., IL-15, TNF-α and IFN-γ)[70]. Alterations in TLR4 and TLR2 expression, as well as functional single-nucleotide polymorphisms in the genes expressed upon TLR4 activation, have also been associated with CD[45,71,72].
Interestingly, epithelial barrier function may be controlled indirectly by the intestinal metabolome, e.g., gut microbiota metabolites in the form of low-molecular weight chemical intermediates[73]. Soluble dietary fibers (such as fructans, pectin, inulin and xylans) and resistant starches can be actively fermented by commensal microbiota in the human colon, producing biologically active short-chain fatty acids (SCFAs), such as acetate, propionate and butyrate. These SCFAs are the main metabolites produced by gut-colonizing bacteria and a major source of energy for intestinal epithelial cells[74]. Acetate and propionate are predominantly produced by bacterial species of the phylum Bacteroidetes, whereas butyrate is primarily produced by those of the phylum Firmicutes. SCFAs serve as specific activators of orphan G-protein-coupled receptors, such as GPR43 and GPR41, predominantly expressed in intestinal epithelial cells[75,76]. GPR43 deficiency leads to expansion of Firmicutes in the gut microbiota and consequently raises fecal SCFAs and plasma acetate levels. Indoles, produced from tryptophan by various Gram-positive and Gram-negative intestinal bacteria, and acetate, produced by Bifidobacterium strains, enhance epithelial defense functions and suppress intestinal inflammation[77-79]. Microbe-derived SCFAs also have an impact on terminal differentiation of CD4+ Th cells[80].
The gut microbiota is responsible not only for immune homeostasis and epithelial barrier function, but also can have direct impact on gluten digestion in the intestinal tract. There is evidence that certain bacterial strains isolated from feces, e.g., Bifidobacterium and Bacteroides fragilis, are capable of digesting immunogenic gliadin peptides, which are rich in proline residues but resistant to human enzymes[81,82].
Several studies addressed the phenomenon of gut dysbiosis in CD patients with active untreated disease and those on a GFD. Fecal analyses in untreated CD patients showed an imbalance in the composition of intestinal microbiota characterized by an increase in the number of Bacteroides species and reduced numbers of Bifidobacterium species[83-86]. In addition, CD patients, both untreated and treated with a GFD, demonstrated a lesser diversity of Bacteroides species in biopsy samples of the duodenal microbiota in comparison with controls[87]. The numbers of Escherichia coli and Staphylococcus bacteria were also higher in fecal and biopsy specimens of untreated CD children than in controls[88]. Escherichia coli strains from CD children carried a higher number of virulence genes than those from healthy children. Nadal et al[89] in 2007 reported a significantly lower ratio of harmless Gram-positive bacteria (Lactobacillus and Bifidobacterium) to potentially harmful Gram-negative bacteria (Bacteroides/Prevotella and Escherichia coli) in CD patients compared to controls, with no distinction between active and inactive CD. The numbers of bacteria of Streptococcus and Prevotella genera were found to be lower both in adults and children with untreated CD in comparison with healthy controls.
The disturbances in intestinal microbiota composition found in CD patients have been associated with changes in the metabolome[90]. Metabolic profiles of serum, urine and feces in celiac patients revealed a significantly altered profile of volatile organic compounds (e.g., phenols and ketones), SCFAs and amino acids (e.g., proline, methionine, histidine and tryptophan)[91,92]. CD patients were also characterized by higher urine levels of certain gut microbiota-derived metabolites, such as indoxyl sulfate, meta-[hydroxyphenyl] propionic acid and phenylacetylglycine, which were associated with untreated CD[92]. Interestingly, metabolic abnormalities found in celiac patients and “potential” celiac patients (i.e. individuals with a positive antibody test but no evidence of intestinal damage) were similar, indicating that CD-related dysmetabolism/dysbiosis precedes the intestinal damage[93].
Only a few serum metabolites can help differentiate between potential and overt CD; none of these metabolites are related to energy metabolism. Glycolysis appears to be somehow impaired in potential CD patients, just as is the case in overt CD patients. This is consistent with the hypothesis that the gut microbiota of CD patients is altered or contains specific species with their distinctive microbial metabolome. Schirmer et al[94] in 2016 reported that TNF-α and IFN-γ production was associated with specific microbial metabolic pathways: palmitoleic acid metabolism and tryptophan degradation to tryptophol.
Low doses of pro-inflammatory cytokines, such as IFN-γ, were shown not to affect TJ protein expression but to activate bacterial endocytosis by epithelial cells[95]. This process is dependent on extracellular signal-regulated kinase Cζ and ADP-ribosylation factor-6 signaling[96]. Thus, some commensal bacteria might interact with certain intracellular PRRs, namely, nucleotide-binding oligomerization domain (NOD)-like receptors, and activate epithelium-derived pro-inflammatory cytokines and free radicals that may cause secondary TJ damage[96,97]. An increased activity and expression of inducible nitric oxide synthase in human duodenal enterocytes has been reported in CD patients[97].
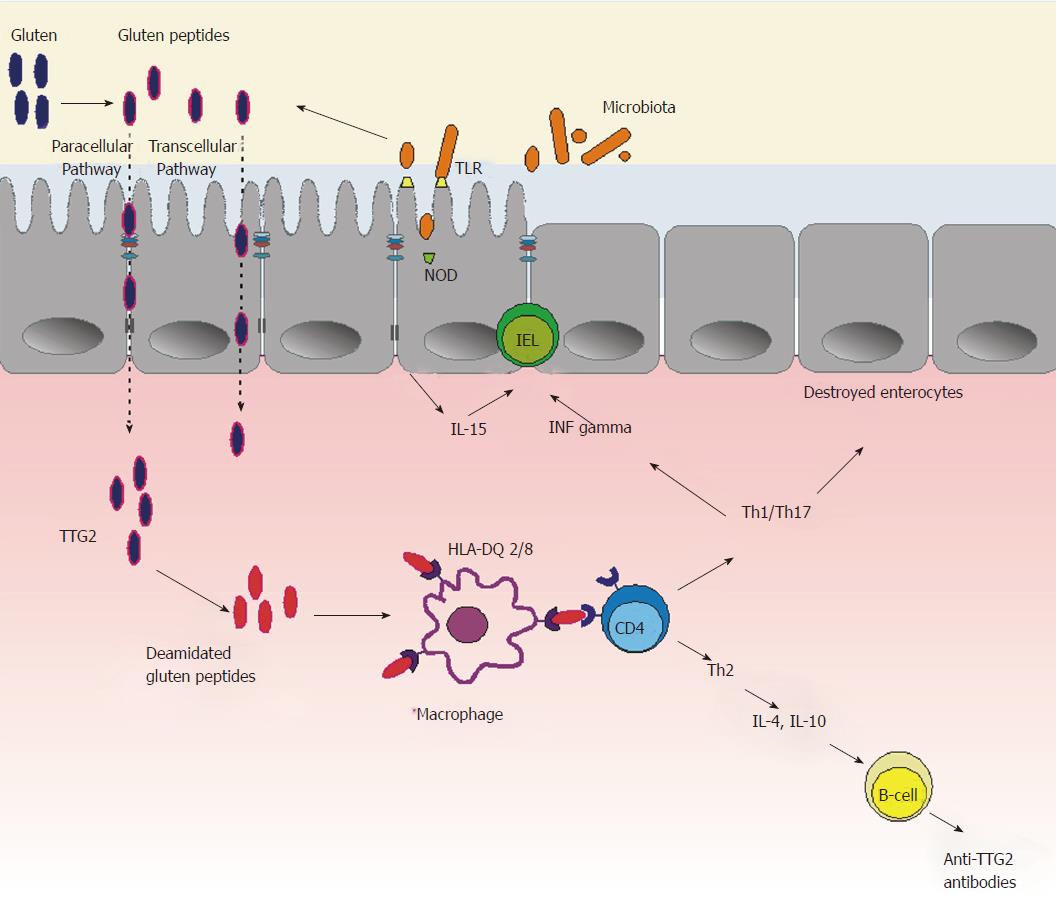
Thus, dysbiosis, which can follow viral or bacterial infections or antibiotic therapy, may activate innate immunity leading to pro-inflammatory changes, with the resulting IEL infiltration, epithelial barrier disruption, and increased transfer of immunogenic gluten peptides, which in turn activate inflammation leading to CD development (Figure 3).
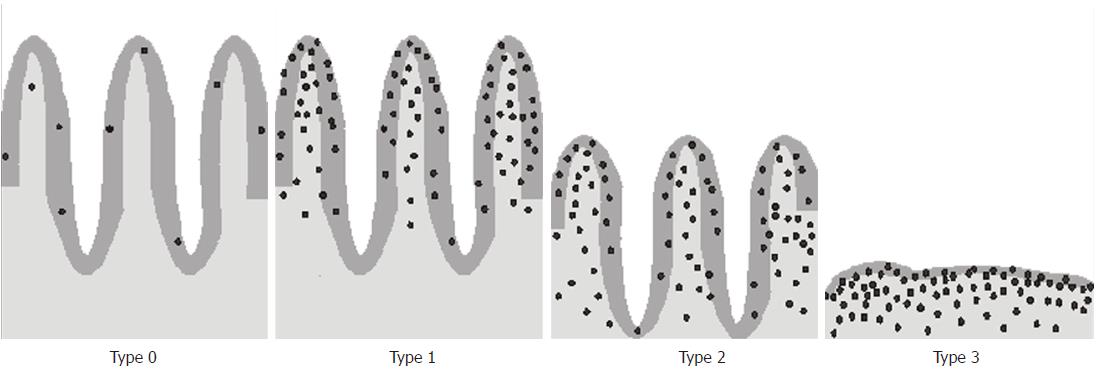
The typical histopathological presentation of CD is small intestinal enteropathy characterized by an increase in IELs, crypt hyperplasia, and villous atrophy. The changes develop gradually over time. The increased number of IELs is one of the earliest signs of CD[98] and may herald the impending disease[99]. Histological changes in the small intestine can be graded using the Marsh classification[100] modified by Oberhuber[101] (Figure 4). The Marsh-Oberhuber classification includes four categories of CD-associated lesions: infiltrative (type 1), infiltrative-hyperplastic (type 2), flat-destructive (type 3) and atrophic-hypoplastic (type 4)[101].
Irrespective of the type of changes found in CD patients, an increase in the number of IELs is considered to be the most sensitive histopathological marker of CD. The upper limit of normal for IELs in duodenal or jejunal mucosa is 25 IELs per 100 enterocytes. An IEL count between 25 and 29 is considered to be “borderline intraepithelial lymphocytosis”, and 30 or more means “pathological lymphocytosis” in the duodenum[103,104]. IELs are classified into two major subgroups based on their phenotypical and functional characteristics: one bears the αβ T-cell receptor (αβ-IEL), while the other bears the γδ T-cell receptor (γδ-IEL). When it comes to the typical composition of the small-intestinal IEL population, approximately 75% of it consists of CD8-positive αβ T cells, 10% constitute CD4-positive αβ T cells, and 15% constitute γδ T cells which are CD4- and either CD8- or CD8+[5].
In CD sensitive patients, gluten exposure causes rapid activation of both αβ-IELs and γδ-IELs[105], while a GFD lowers both αβ-IEL and γδ-IEL counts; however, lowering of the latter IEL subtype takes months or even years[106]. It is believed that CD8+ αβ-IELs represent the effector T cell subset that mediates epithelial cell destruction (after IL-15 up-regulation) and, ultimately, induces villous atrophy in CD. The role of γδ-IELs in CD pathogenesis remains unclear.
A recent study showed that IEL expansion can be modulated by the host microbiota. Mice deficient in NOD2 (receptors recognizing bacterial molecules)[107] exhibited a significant reduction in IEL counts and IL-15 expression in the epithelium, with the residual IELs displaying reduced proliferation and increased apoptosis. Moreover, Lactobacillus strains were able to decrease the number of IELs activated by TLR3 after an experimentally induced viral infection (poly I:C). They also significantly reduced the levels of pro-inflammatory cytokines, such as TNF-α and IL-15, and increased serum and intestinal regulatory IL-10 levels[108]. Finally, the immunomodulatory capacity of Lactobacilli helped significantly reduce intestinal tissue damage.
The data above indicate that IEL homeostasis is controlled by commensal microbiota, which affects cytokine production by epithelial cells via PRR activation. Moreover, increased IEL counts in CD patients, which lead to epithelial barrier disturbances, may be primarily induced by microbiota dysbiosis.
Recent research has shown that early bacterial colonization may affect the risk of developing CD later in life. This phenomenon is called microbial programming[109]. There is evidence indicating that the pioneer microbiota of the neonatal gut is essential for gut maturation as well as for metabolic and immunologic programming[109,110]. Establishment of the human gut microbiota is a complex, stepwise process. The composition of microbiota within the 1st year of life is characterized by low diversity, high instability, and high inter-individual variation[111]. By the age of 2-3 years, the microbiota becomes stable, more diverse, and resembles that found in adults, with Firmicutes and Bacteroidetes as the predominant phyla. Gut microbiota formation after the birth is dependent on different environmental factors, such as the mode of delivery, breast or formula feeding, or antibiotic therapy[111].
Although the evidence that the perinatal environment influences CD development is still only circumstantial[112], there have been studies showing that cesarean sections and antibiotic treatment in infancy increased the risk of CD[113-115]. There is also evidence that colonization of the gut with microorganisms may be dependent on genetic factors[116,117]. The hypothesis that gut microbiota composition is affected by host genes has been confirmed by studies in twins, showing that fecal microbiota of monozygotic twins was much more similar than that of dizygotic twins[118]. Recent microbiome analyses performed on 22 infants demonstrated that certain HLA genes predisposing to CD could affect microbiota composition[119]. The infants at high genetic risk of CD, i.e. those with an HLA-DQ2 genotype, showed a higher proportion of Firmicutes and Proteobacteria and lower proportion of Actinobacteria than those at low genetic risk. At the genus level, the gut microbiota of high-risk infants had a significantly lower proportion of Bifidobacterium and unclassified Bifidobacteriaceae and a higher proportion of Corynebacterium, Gemella, Clostridium, unclassified Clostridiaceae, unclassified Enterobacteriaceae and Raoultella. Sellitto et al[120] in 2012 reported an overall lack of bacteria of the phylum Bacteriodetes, with a high abundance of Firmicutes, in infants genetically predisposed to CD compared with microbiota composition of low-risk infants. Those differences were stable until 2 years of age.
As CD is strongly associated with HLA genes - almost 100% individuals with CD are carriers of alleles encoding HLA-DQ2/DQ8 molecules - these findings suggest that children with the CD risk genotype have a different microbiota profile than those without genetic predisposition. However, it must be emphasized that about 25%-30% of the general population exhibits the same HLA genotypes as CD patients[121]. In addition, there are also non-HLA genes associated with CD.
Although gluten is necessary in order to activate the processes leading to CD, there is evidence that an imbalance in the gut microbiota and intestinal epithelium can precede the specific gluten-dependent immune response. Under certain conditions affecting the intestinal microbiota, e.g., after infections or antibiotic therapy, an increased translocation of dietary macromolecules (including gluten peptides) via the opening of epithelial junctions triggers a cascade of events in genetically susceptible individuals, leading to overt CD. Microbiota disturbances are observed not only in untreated CD patients, but also in potential CD patients and those following a GFD as well as in infants at high genetic risk of CD. The microbial fingerprint associated with CD is likely dependent on specific genetic factors, including (but not exclusively) the HLA-DQ2/-DQ8 genotype. Future strategies should include prospective, birth cohort studies involving comprehensive genome, microbiome and metabolome analyses. Such an approach could help identify a “CD-specific“ microbial/metabolic fingerprint, which would become the target for both primary prevention and management of CD.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: Poland
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: De Re V Romano M, Schwarz SM, S- Editor: Qi Y L- Editor: Filipodia E- Editor: Ma YJ
| 1. | Troncone R, Discepolo V. Celiac disease and autoimmunity. J Pediatr Gastroenterol Nutr. 2014;59 Suppl 1:S9-S11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 2. | Koning F. Celiac disease: quantity matters. Semin Immunopathol. 2012;34:541-549. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 51] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 3. | Sollid LM, Qiao SW, Anderson RP, Gianfrani C, Koning F. Nomenclature and listing of celiac disease relevant gluten T-cell epitopes restricted by HLA-DQ molecules. Immunogenetics. 2012;64:455-460. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 355] [Cited by in RCA: 339] [Article Influence: 26.1] [Reference Citation Analysis (0)] |
| 4. | van Bergen J, Mulder CJ, Mearin ML, Koning F. Local communication among mucosal immune cells in patients with celiac disease. Gastroenterology. 2015;148:1187-1194. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 26] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 5. | Jabri B, Abadie V. IL-15 functions as a danger signal to regulate tissue-resident T cells and tissue destruction. Nat Rev Immunol. 2015;15:771-783. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 218] [Cited by in RCA: 238] [Article Influence: 23.8] [Reference Citation Analysis (0)] |
| 6. | Abadie V, Jabri B. IL-15: a central regulator of celiac disease immunopathology. Immunol Rev. 2014;260:221-234. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 145] [Cited by in RCA: 170] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 7. | Ivarsson A, Persson LA, Nyström L, Ascher H, Cavell B, Danielsson L, Dannaeus A, Lindberg T, Lindquist B, Stenhammar L. Epidemic of coeliac disease in Swedish children. Acta Paediatr. 2000;89:165-171. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 268] [Cited by in RCA: 251] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 8. | Ivarsson A, Hernell O, Stenlund H, Persson LA. Breast-feeding protects against celiac disease. Am J Clin Nutr. 2002;75:914-921. [PubMed] |
| 9. | Szajewska H, Shamir R, Chmielewska A, Pieścik-Lech M, Auricchio R, Ivarsson A, Kolacek S, Koletzko S, Korponay-Szabo I, Mearin ML. Systematic review with meta-analysis: early infant feeding and coeliac disease--update 2015. Aliment Pharmacol Ther. 2015;41:1038-1054. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 84] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 10. | Crespo-Escobar P, Mearin ML, Hervás D, Auricchio R, Castillejo G, Gyimesi J, Martinez-Ojinaga E, Werkstetter K, Vriezinga SL, Korponay-Szabo IR. The role of gluten consumption at an early age in celiac disease development: a further analysis of the prospective PreventCD cohort study. Am J Clin Nutr. 2017;105:890-896. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 41] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 11. | Lahdenperä A, Ludvigsson J, Fälth-Magnusson K, Högberg L, Vaarala O. The effect of gluten-free diet on Th1-Th2-Th3-associated intestinal immune responses in celiac disease. Scand J Gastroenterol. 2011;46:538-549. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 25] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 12. | Bischoff SC, Barbara G, Buurman W, Ockhuizen T, Schulzke JD, Serino M, Tilg H, Watson A, Wells JM. Intestinal permeability--a new target for disease prevention and therapy. BMC Gastroenterol. 2014;14:189. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 893] [Cited by in RCA: 1218] [Article Influence: 110.7] [Reference Citation Analysis (0)] |
| 13. | Moens E, Veldhoen M. Epithelial barrier biology: good fences make good neighbours. Immunology. 2012;135:1-8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 110] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 14. | Antoni L, Nuding S, Wehkamp J, Stange EF. Intestinal barrier in inflammatory bowel disease. World J Gastroenterol. 2014;20:1165-1179. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 248] [Cited by in RCA: 302] [Article Influence: 27.5] [Reference Citation Analysis (0)] |
| 15. | Gumbiner BM. Cell adhesion: the molecular basis of tissue architecture and morphogenesis. Cell. 1996;84:345-357. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2618] [Cited by in RCA: 2557] [Article Influence: 88.2] [Reference Citation Analysis (0)] |
| 16. | Tsukita S, Furuse M, Itoh M. Multifunctional strands in tight junctions. Nat Rev Mol Cell Biol. 2001;2:285-293. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1851] [Cited by in RCA: 1903] [Article Influence: 79.3] [Reference Citation Analysis (0)] |
| 17. | Van Itallie CM, Anderson JM. Architecture of tight junctions and principles of molecular composition. Semin Cell Dev Biol. 2014;36:157-165. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 316] [Cited by in RCA: 383] [Article Influence: 34.8] [Reference Citation Analysis (0)] |
| 18. | Furuse M, Itoh M, Hirase T, Nagafuchi A, Yonemura S, Tsukita S, Tsukita S. Direct association of occludin with ZO-1 and its possible involvement in the localization of occludin at tight junctions. J Cell Biol. 1994;127:1617-1626. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 717] [Cited by in RCA: 726] [Article Influence: 23.4] [Reference Citation Analysis (0)] |
| 19. | Saitou M, Furuse M, Sasaki H, Schulzke JD, Fromm M, Takano H, Noda T, Tsukita S. Complex phenotype of mice lacking occludin, a component of tight junction strands. Mol Biol Cell. 2000;11:4131-4142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 875] [Cited by in RCA: 894] [Article Influence: 35.8] [Reference Citation Analysis (0)] |
| 20. | Rao R. Occludin phosphorylation in regulation of epithelial tight junctions. Ann N Y Acad Sci. 2009;1165:62-68. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 170] [Cited by in RCA: 175] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 21. | Sakakibara A, Furuse M, Saitou M, Ando-Akatsuka Y, Tsukita S. Possible involvement of phosphorylation of occludin in tight junction formation. J Cell Biol. 1997;137:1393-1401. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 450] [Cited by in RCA: 436] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
| 22. | Seth A, Sheth P, Elias BC, Rao R. Protein phosphatases 2A and 1 interact with occludin and negatively regulate the assembly of tight junctions in the CACO-2 cell monolayer. J Biol Chem. 2007;282:11487-11498. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 136] [Cited by in RCA: 145] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 23. | Rao RK, Basuroy S, Rao VU, Karnaky Jr KJ, Gupta A. Tyrosine phosphorylation and dissociation of occludin-ZO-1 and E-cadherin-beta-catenin complexes from the cytoskeleton by oxidative stress. Biochem J. 2002;368:471-481. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 303] [Cited by in RCA: 334] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 24. | Elias BC, Suzuki T, Seth A, Giorgianni F, Kale G, Shen L, Turner JR, Naren A, Desiderio DM, Rao R. Phosphorylation of Tyr-398 and Tyr-402 in occludin prevents its interaction with ZO-1 and destabilizes its assembly at the tight junctions. J Biol Chem. 2009;284:1559-1569. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 161] [Cited by in RCA: 161] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 25. | Lee SH. Intestinal permeability regulation by tight junction: implication on inflammatory bowel diseases. Intest Res. 2015;13:11-18. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 408] [Cited by in RCA: 579] [Article Influence: 57.9] [Reference Citation Analysis (0)] |
| 26. | Angelow S, Ahlstrom R, Yu AS. Biology of claudins. Am J Physiol Renal Physiol. 2008;295:F867-F876. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 268] [Cited by in RCA: 274] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 27. | Van Itallie CM, Anderson JM. Claudins and epithelial paracellular transport. Annu Rev Physiol. 2006;68:403-429. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 837] [Cited by in RCA: 865] [Article Influence: 45.5] [Reference Citation Analysis (0)] |
| 28. | Fanning AS, Van Itallie CM, Anderson JM. Zonula occludens-1 and -2 regulate apical cell structure and the zonula adherens cytoskeleton in polarized epithelia. Mol Biol Cell. 2012;23:577-590. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 181] [Cited by in RCA: 190] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 29. | Itoh M, Furuse M, Morita K, Kubota K, Saitou M, Tsukita S. Direct binding of three tight junction-associated MAGUKs, ZO-1, ZO-2, and ZO-3, with the COOH termini of claudins. J Cell Biol. 1999;147:1351-1363. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 829] [Cited by in RCA: 854] [Article Influence: 32.8] [Reference Citation Analysis (0)] |
| 30. | Fanning AS, Ma TY, Anderson JM. Isolation and functional characterization of the actin binding region in the tight junction protein ZO-1. FASEB J. 2002;16:1835-1837. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 216] [Cited by in RCA: 215] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 31. | Mariano C, Sasaki H, Brites D, Brito MA. A look at tricellulin and its role in tight junction formation and maintenance. Eur J Cell Biol. 2011;90:787-796. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 59] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 32. | Takeichi M. Cadherin cell adhesion receptors as a morphogenetic regulator. Science. 1991;251:1451-1455. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2396] [Cited by in RCA: 2422] [Article Influence: 71.2] [Reference Citation Analysis (0)] |
| 33. | Laukoetter MG, Bruewer M, Nusrat A. Regulation of the intestinal epithelial barrier by the apical junctional complex. Curr Opin Gastroenterol. 2006;22:85-89. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 175] [Cited by in RCA: 195] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 34. | Fasano A, Uzzau S, Fiore C, Margaretten K. The enterotoxic effect of zonula occludens toxin on rabbit small intestine involves the paracellular pathway. Gastroenterology. 1997;112:839-846. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 103] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 35. | Fasano A, Not T, Wang W, Uzzau S, Berti I, Tommasini A, Goldblum SE. Zonulin, a newly discovered modulator of intestinal permeability, and its expression in coeliac disease. Lancet. 2000;355:1518-1519. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 397] [Cited by in RCA: 436] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 36. | Tripathi A, Lammers KM, Goldblum S, Shea-Donohue T, Netzel-Arnett S, Buzza MS, Antalis TM, Vogel SN, Zhao A, Yang S. Identification of human zonulin, a physiological modulator of tight junctions, as prehaptoglobin-2. Proc Natl Acad Sci USA. 2009;106:16799-16804. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 261] [Cited by in RCA: 319] [Article Influence: 19.9] [Reference Citation Analysis (0)] |
| 37. | Hollande F, Blanc EM, Bali JP, Whitehead RH, Pelegrin A, Baldwin GS, Choquet A. HGF regulates tight junctions in new nontumorigenic gastric epithelial cell line. Am J Physiol Gastrointest Liver Physiol. 2001;280:G910-G921. [PubMed] |
| 38. | Lammers KM, Lu R, Brownley J, Lu B, Gerard C, Thomas K, Rallabhandi P, Shea-Donohue T, Tamiz A, Alkan S. Gliadin induces an increase in intestinal permeability and zonulin release by binding to the chemokine receptor CXCR3. Gastroenterology. 2008;135:194-204.e3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 384] [Cited by in RCA: 370] [Article Influence: 21.8] [Reference Citation Analysis (0)] |
| 39. | Clemente MG, De Virgiliis S, Kang JS, Macatagney R, Musu MP, Di Pierro MR, Drago S, Congia M, Fasano A. Early effects of gliadin on enterocyte intracellular signalling involved in intestinal barrier function. Gut. 2003;52:218-223. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 248] [Cited by in RCA: 255] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 40. | El Asmar R, Panigrahi P, Bamford P, Berti I, Not T, Coppa GV, Catassi C, Fasano A. Host-dependent zonulin secretion causes the impairment of the small intestine barrier function after bacterial exposure. Gastroenterology. 2002;123:1607-1615. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 225] [Cited by in RCA: 256] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 41. | Mokkala K, Röytiö H, Munukka E, Pietilä S, Ekblad U, Rönnemaa T, Eerola E, Laiho A, Laitinen K. Gut Microbiota Richness and Composition and Dietary Intake of Overweight Pregnant Women Are Related to Serum Zonulin Concentration, a Marker for Intestinal Permeability. J Nutr. 2016;146:1694-1700. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 99] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 42. | Schumann M, Siegmund B, Schulzke JD, Fromm M. Celiac Disease: Role of the Epithelial Barrier. Cell Mol Gastroenterol Hepatol. 2017;3:150-162. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 106] [Cited by in RCA: 105] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 43. | Schumann M, Günzel D, Buergel N, Richter JF, Troeger H, May C, Fromm A, Sorgenfrei D, Daum S, Bojarski C. Cell polarity-determining proteins Par-3 and PP-1 are involved in epithelial tight junction defects in coeliac disease. Gut. 2012;61:220-228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 79] [Cited by in RCA: 101] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 44. | Wapenaar MC, Monsuur AJ, van Bodegraven AA, Weersma RK, Bevova MR, Linskens RK, Howdle P, Holmes G, Mulder CJ, Dijkstra G. Associations with tight junction genes PARD3 and MAGI2 in Dutch patients point to a common barrier defect for coeliac disease and ulcerative colitis. Gut. 2008;57:463-467. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 122] [Cited by in RCA: 127] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 45. | Dubois PC, Trynka G, Franke L, Hunt KA, Romanos J, Curtotti A, Zhernakova A, Heap GA, Adány R, Aromaa A. Multiple common variants for celiac disease influencing immune gene expression. Nat Genet. 2010;42:295-302. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 843] [Cited by in RCA: 771] [Article Influence: 51.4] [Reference Citation Analysis (0)] |
| 46. | Schulzke JD, Bentzel CJ, Schulzke I, Riecken EO, Fromm M. Epithelial tight junction structure in the jejunum of children with acute and treated celiac sprue. Pediatr Res. 1998;43:435-441. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 132] [Cited by in RCA: 129] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 47. | Goswami P, Das P, Verma AK, Prakash S, Das TK, Nag TC, Ahuja V, Gupta SD, Makharia GK. Are alterations of tight junctions at molecular and ultrastructural level different in duodenal biopsies of patients with celiac disease and Crohn’s disease? Virchows Arch. 2014;465:521-530. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 35] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 48. | Mishra A, Prakash S, Sreenivas V, Das TK, Ahuja V, Gupta SD, Makharia GK. Structural and Functional Changes in the Tight Junctions of Asymptomatic and Serology-negative First-degree Relatives of Patients With Celiac Disease. J Clin Gastroenterol. 2016;50:551-560. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 20] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 49. | Szakál DN, Gyorffy H, Arató A, Cseh A, Molnár K, Papp M, Dezsofi A, Veres G. Mucosal expression of claudins 2, 3 and 4 in proximal and distal part of duodenum in children with coeliac disease. Virchows Arch. 2010;456:245-250. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 63] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 50. | Ciccocioppo R, Finamore A, Ara C, Di Sabatino A, Mengheri E, Corazza GR. Altered expression, localization, and phosphorylation of epithelial junctional proteins in celiac disease. Am J Clin Pathol. 2006;125:502-511. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 80] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 51. | Tjon JM, van Bergen J, Koning F. Celiac disease: how complicated can it get? Immunogenetics. 2010;62:641-651. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 112] [Cited by in RCA: 115] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 52. | Schumann M, Kamel S, Pahlitzsch ML, Lebenheim L, May C, Krauss M, Hummel M, Daum S, Fromm M, Schulzke JD. Defective tight junctions in refractory celiac disease. Ann N Y Acad Sci. 2012;1258:43-51. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 41] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 53. | Perry I, Tselepis C, Hoyland J, Iqbal TH, Scott D, Sanders SA, Cooper BT, Jankowski JA. Reduced cadherin/catenin complex expression in celiac disease can be reproduced in vitro by cytokine stimulation. Lab Invest. 1999;79:1489-1499. [PubMed] |
| 54. | Dolfini E, Roncoroni L, Elli L, Fumagalli C, Colombo R, Ramponi S, Forlani F, Bardella MT. Cytoskeleton reorganization and ultrastructural damage induced by gliadin in a three-dimensional in vitro model. World J Gastroenterol. 2005;11:7597-7601. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 30] [Cited by in RCA: 29] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 55. | Bondar C, Araya RE, Guzman L, Rua EC, Chopita N, Chirdo FG. Role of CXCR3/CXCL10 axis in immune cell recruitment into the small intestine in celiac disease. PLoS One. 2014;9:e89068. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 60] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 56. | Tlaskalová-Hogenová H, Stepánková R, Hudcovic T, Tucková L, Cukrowska B, Lodinová-Zádníková R, Kozáková H, Rossmann P, Bártová J, Sokol D. Commensal bacteria (normal microflora), mucosal immunity and chronic inflammatory and autoimmune diseases. Immunol Lett. 2004;93:97-108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 470] [Cited by in RCA: 479] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 57. | Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, Gill SR, Nelson KE, Relman DA. Diversity of the human intestinal microbial flora. Science. 2005;308:1635-1638. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5700] [Cited by in RCA: 5592] [Article Influence: 279.6] [Reference Citation Analysis (2)] |
| 58. | Sender R, Fuchs S, Milo R. Revised Estimates for the Number of Human and Bacteria Cells in the Body. PLoS Biol. 2016;14:e1002533. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2328] [Cited by in RCA: 2927] [Article Influence: 325.2] [Reference Citation Analysis (0)] |
| 59. | Tlaskalová-Hogenová H, Stěpánková R, Kozáková H, Hudcovic T, Vannucci L, Tučková L, Rossmann P, Hrnčíř T, Kverka M, Zákostelská Z. The role of gut microbiota (commensal bacteria) and the mucosal barrier in the pathogenesis of inflammatory and autoimmune diseases and cancer: contribution of germ-free and gnotobiotic animal models of human diseases. Cell Mol Immunol. 2011;8:110-120. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 528] [Cited by in RCA: 500] [Article Influence: 35.7] [Reference Citation Analysis (0)] |
| 60. | Cukrowska B, Kozáková H, Reháková Z, Sinkora J, Tlaskalová-Hogenová H. Specific antibody and immunoglobulin responses after intestinal colonization of germ-free piglets with non-pathogenic Escherichia coli O86. Immunobiology. 2001;204:425-433. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 34] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 61. | Kozakova H, Schwarzer M, Tuckova L, Srutkova D, Czarnowska E, Rosiak I, Hudcovic T, Schabussova I, Hermanova P, Zakostelska Z. Colonization of germ-free mice with a mixture of three lactobacillus strains enhances the integrity of gut mucosa and ameliorates allergic sensitization. Cell Mol Immunol. 2016;13:251-262. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 126] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 62. | Pabst O. New concepts in the generation and functions of IgA. Nat Rev Immunol. 2012;12:821-832. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 416] [Cited by in RCA: 484] [Article Influence: 37.2] [Reference Citation Analysis (0)] |
| 63. | Feng T, Elson CO, Cong Y. Treg cell-IgA axis in maintenance of host immune homeostasis with microbiota. Int Immunopharmacol. 2011;11:589-592. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 40] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 64. | Mikulic J, Longet S, Favre L, Benyacoub J, Corthesy B. Secretory IgA in complex with Lactobacillus rhamnosus potentiates mucosal dendritic cell-mediated Treg cell differentiation via TLR regulatory proteins, RALDH2 and secretion of IL-10 and TGF-β. Cell Mol Immunol. 2017;14:546-556. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 45] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 65. | Ismail AS, Severson KM, Vaishnava S, Behrendt CL, Yu X, Benjamin JL, Ruhn KA, Hou B, DeFranco AL, Yarovinsky F. Gammadelta intraepithelial lymphocytes are essential mediators of host-microbial homeostasis at the intestinal mucosal surface. Proc Natl Acad Sci USA. 2011;108:8743-8748. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 263] [Cited by in RCA: 241] [Article Influence: 17.2] [Reference Citation Analysis (0)] |
| 66. | Ismail AS, Behrendt CL, Hooper LV. Reciprocal interactions between commensal bacteria and gamma delta intraepithelial lymphocytes during mucosal injury. J Immunol. 2009;182:3047-3054. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 174] [Cited by in RCA: 163] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 67. | Ukena SN, Singh A, Dringenberg U, Engelhardt R, Seidler U, Hansen W, Bleich A, Bruder D, Franzke A, Rogler G. Probiotic Escherichia coli Nissle 1917 inhibits leaky gut by enhancing mucosal integrity. PLoS One. 2007;2:e1308. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 310] [Cited by in RCA: 349] [Article Influence: 19.4] [Reference Citation Analysis (0)] |
| 68. | Ewaschuk JB, Diaz H, Meddings L, Diederichs B, Dmytrash A, Backer J, Looijer-van Langen M, Madsen KL. Secreted bioactive factors from Bifidobacterium infantis enhance epithelial cell barrier function. Am J Physiol Gastrointest Liver Physiol. 2008;295:G1025-G1034. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 376] [Cited by in RCA: 413] [Article Influence: 24.3] [Reference Citation Analysis (0)] |
| 69. | García-Lafuente A, Antolín M, Guarner F, Crespo E, Malagelada JR. Modulation of colonic barrier function by the composition of the commensal flora in the rat. Gut. 2001;48:503-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 99] [Cited by in RCA: 94] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 70. | Rhee SH. Basic and translational understandings of microbial recognition by toll-like receptors in the intestine. J Neurogastroenterol Motil. 2011;17:28-34. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 26] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 71. | Szebeni B, Veres G, Dezsofi A, Rusai K, Vannay A, Bokodi G, Vásárhelyi B, Korponay-Szabó IR, Tulassay T, Arató A. Increased mucosal expression of Toll-like receptor (TLR)2 and TLR4 in coeliac disease. J Pediatr Gastroenterol Nutr. 2007;45:187-193. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 70] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 72. | Kalliomäki M, Satokari R, Lähteenoja H, Vähämiko S, Grönlund J, Routi T, Salminen S. Expression of microbiota, Toll-like receptors, and their regulators in the small intestinal mucosa in celiac disease. J Pediatr Gastroenterol Nutr. 2012;54:727-732. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 79] [Cited by in RCA: 82] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 73. | Gilbert JA, Quinn RA, Debelius J, Xu ZZ, Morton J, Garg N, Jansson JK, Dorrestein PC, Knight R. Microbiome-wide association studies link dynamic microbial consortia to disease. Nature. 2016;535:94-103. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 457] [Cited by in RCA: 479] [Article Influence: 53.2] [Reference Citation Analysis (0)] |
| 74. | Tan J, McKenzie C, Potamitis M, Thorburn AN, Mackay CR, Macia L. The role of short-chain fatty acids in health and disease. Adv Immunol. 2014;121:91-119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1076] [Cited by in RCA: 1616] [Article Influence: 146.9] [Reference Citation Analysis (0)] |
| 75. | Maslowski KM, Vieira AT, Ng A, Kranich J, Sierro F, Yu D, Schilter HC, Rolph MS, Mackay F, Artis D. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature. 2009;461:1282-1286. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2007] [Cited by in RCA: 2399] [Article Influence: 149.9] [Reference Citation Analysis (0)] |
| 76. | Kim MH, Kang SG, Park JH, Yanagisawa M, Kim CH. Short-chain fatty acids activate GPR41 and GPR43 on intestinal epithelial cells to promote inflammatory responses in mice. Gastroenterology. 2013;145:396-406.e1-10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 576] [Cited by in RCA: 770] [Article Influence: 64.2] [Reference Citation Analysis (0)] |
| 77. | Shimada Y, Kinoshita M, Harada K, Mizutani M, Masahata K, Kayama H, Takeda K. Commensal bacteria-dependent indole production enhances epithelial barrier function in the colon. PLoS One. 2013;8:e80604. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 204] [Cited by in RCA: 284] [Article Influence: 23.7] [Reference Citation Analysis (0)] |
| 78. | Bansal T, Alaniz RC, Wood TK, Jayaraman A. The bacterial signal indole increases epithelial-cell tight-junction resistance and attenuates indicators of inflammation. Proc Natl Acad Sci USA. 2010;107:228-233. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 485] [Cited by in RCA: 639] [Article Influence: 39.9] [Reference Citation Analysis (0)] |
| 79. | Inan MS, Rasoulpour RJ, Yin L, Hubbard AK, Rosenberg DW, Giardina C. The luminal short-chain fatty acid butyrate modulates NF-kappaB activity in a human colonic epithelial cell line. Gastroenterology. 2000;118:724-734. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 330] [Cited by in RCA: 356] [Article Influence: 14.2] [Reference Citation Analysis (0)] |
| 80. | Geuking MB, McCoy KD, Macpherson AJ. Metabolites from intestinal microbes shape Treg. Cell Res. 2013;23:1339-1340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 37] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 81. | Laparra JM, Sanz Y. Bifidobacteria inhibit the inflammatory response induced by gliadins in intestinal epithelial cells via modifications of toxic peptide generation during digestion. J Cell Biochem. 2010;109:801-807. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 63] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 82. | Sánchez E, Laparra JM, Sanz Y. Discerning the role of Bacteroides fragilis in celiac disease pathogenesis. Appl Environ Microbiol. 2012;78:6507-6515. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 66] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 83. | Sánchez E, De Palma G, Capilla A, Nova E, Pozo T, Castillejo G, Varea V, Marcos A, Garrote JA, Polanco I. Influence of environmental and genetic factors linked to celiac disease risk on infant gut colonization by Bacteroides species. Appl Environ Microbiol. 2011;77:5316-5323. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 94] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 84. | Sanz Y, Sánchez E, Marzotto M, Calabuig M, Torriani S, Dellaglio F. Differences in faecal bacterial communities in coeliac and healthy children as detected by PCR and denaturing gradient gel electrophoresis. FEMS Immunol Med Microbiol. 2007;51:562-568. [PubMed] |
| 85. | Wacklin P, Kaukinen K, Tuovinen E, Collin P, Lindfors K, Partanen J, Mäki M, Mättö J. The duodenal microbiota composition of adult celiac disease patients is associated with the clinical manifestation of the disease. Inflamm Bowel Dis. 2013;19:934-941. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 126] [Cited by in RCA: 144] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 86. | Cheng J, Kalliomäki M, Heilig HG, Palva A, Lähteenoja H, de Vos WM, Salojärvi J, Satokari R. Duodenal microbiota composition and mucosal homeostasis in pediatric celiac disease. BMC Gastroenterol. 2013;13:113. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 98] [Cited by in RCA: 111] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 87. | Sánchez E, Donat E, Ribes-Koninckx C, Fernández-Murga ML, Sanz Y. Duodenal-mucosal bacteria associated with celiac disease in children. Appl Environ Microbiol. 2013;79:5472-5479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 121] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 88. | Collado MC, Donat E, Ribes-Koninckx C, Calabuig M, Sanz Y. Imbalances in faecal and duodenal Bifidobacterium species composition in active and non-active coeliac disease. BMC Microbiol. 2008;8:232. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 144] [Cited by in RCA: 151] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 89. | Nadal I, Donat E, Ribes-Koninckx C, Calabuig M, Sanz Y. Imbalance in the composition of the duodenal microbiota of children with coeliac disease. J Med Microbiol. 2007;56:1669-1674. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 271] [Cited by in RCA: 288] [Article Influence: 16.9] [Reference Citation Analysis (0)] |
| 90. | Bertini I, Calabrò A, De Carli V, Luchinat C, Nepi S, Porfirio B, Renzi D, Saccenti E, Tenori L. The metabonomic signature of celiac disease. J Proteome Res. 2009;8:170-177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 122] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 91. | Calabrò A, Gralka E, Luchinat C, Saccenti E, Tenori L. A metabolomic perspective on coeliac disease. Autoimmune Dis. 2014;2014:756138. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 20] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 92. | Duran M, Ketting D, de Bree PK, van der Heiden C, Wadman SK. Gas chromatographic analysis of urinary volatile phenols in patients with gastro-intestinal disorders and normals. Clin Chim Acta. 1973;45:341-347. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 20] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 93. | Bernini P, Bertini I, Calabrò A, la Marca G, Lami G, Luchinat C, Renzi D, Tenori L. Are patients with potential celiac disease really potential? The answer of metabonomics. J Proteome Res. 2011;10:714-721. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 94. | Schirmer M, Smeekens SP, Vlamakis H, Jaeger M, Oosting M, Franzosa EA, Jansen T, Jacobs L, Bonder MJ, Kurilshikov A. Linking the Human Gut Microbiome to Inflammatory Cytokine Production Capacity. Cell. 2016;167:1125-1136.e8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 495] [Cited by in RCA: 723] [Article Influence: 90.4] [Reference Citation Analysis (0)] |
| 95. | Yu LC. Commensal bacterial internalization by epithelial cells: An alternative portal for gut leakiness. Tissue Barriers. 2015;3:e1008895. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 12] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 96. | Wu LL, Chiu HD, Peng WH, Lin BR, Lu KS, Lu YZ, Yu LC. Epithelial inducible nitric oxide synthase causes bacterial translocation by impairment of enterocytic tight junctions via intracellular signals of Rho-associated kinase and protein kinase C zeta. Crit Care Med. 2011;39:2087-2098. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 52] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 97. | Murray IA, Daniels I, Coupland K, Smith JA, Long RG. Increased activity and expression of iNOS in human duodenal enterocytes from patients with celiac disease. Am J Physiol Gastrointest Liver Physiol. 2002;283:G319-G326. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 36] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 98. | Sergi C, Shen F, Bouma G. Intraepithelial lymphocytes, scores, mimickers and challenges in diagnosing gluten-sensitive enteropathy (celiac disease). World J Gastroenterol. 2017;23:573-589. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 26] [Cited by in RCA: 23] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 99. | Järvinen TT, Collin P, Rasmussen M, Kyrönpalo S, Mäki M, Partanen J, Reunala T, Kaukinen K. Villous tip intraepithelial lymphocytes as markers of early-stage coeliac disease. Scand J Gastroenterol. 2004;39:428-433. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 83] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 100. | Marsh MN. Gluten, major histocompatibility complex, and the small intestine. A molecular and immunobiologic approach to the spectrum of gluten sensitivity (‘celiac sprue’). Gastroenterology. 1992;102:330-354. [PubMed] |
| 101. | Oberhuber G, Granditsch G, Vogelsang H. The histopathology of coeliac disease: time for a standardized report scheme for pathologists. Eur J Gastroenterol Hepatol. 1999;11:1185-1194. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1142] [Cited by in RCA: 1205] [Article Influence: 46.3] [Reference Citation Analysis (0)] |
| 102. | Hayat M, Cairns A, Dixon MF, O’Mahony S. Quantitation of intraepithelial lymphocytes in human duodenum: what is normal? J Clin Pathol. 2002;55:393-394. [PubMed] |
| 103. | Mahadeva S, Wyatt JI, Howdle PD. Is a raised intraepithelial lymphocyte count with normal duodenal villous architecture clinically relevant? J Clin Pathol. 2002;55:424-428. [PubMed] |
| 104. | Veress B, Franzén L, Bodin L, Borch K. Duodenal intraepithelial lymphocyte-count revisited. Scand J Gastroenterol. 2004;39:138-144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 111] [Cited by in RCA: 105] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 105. | Iltanen S, Holm K, Ashorn M, Ruuska T, Laippala P, Mäki M. Changing jejunal gamma delta T cell receptor (TCR)-bearing intraepithelial lymphocyte density in coeliac disease. Clin Exp Immunol. 1999;117:51-55. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 40] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 106. | Collin P, Wahab PJ, Murray JA. Intraepithelial lymphocytes and coeliac disease. Best Pract Res Clin Gastroenterol. 2005;19:341-350. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 30] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 107. | Jiang W, Wang X, Zeng B, Liu L, Tardivel A, Wei H, Han J, MacDonald HR, Tschopp J, Tian Z. Recognition of gut microbiota by NOD2 is essential for the homeostasis of intestinal intraepithelial lymphocytes. J Exp Med. 2013;210:2465-2476. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 126] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 108. | Tada A, Zelaya H, Clua P, Salva S, Alvarez S, Kitazawa H, Villena J. Immunobiotic Lactobacillus strains reduce small intestinal injury induced by intraepithelial lymphocytes after Toll-like receptor 3 activation. Inflamm Res. 2016;771-783. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 33] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 109. | Kaplan JL, Shi HN, Walker WA. The role of microbes in developmental immunologic programming. Pediatr Res. 2011;69:465-472. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 132] [Cited by in RCA: 132] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 110. | Bäckhed F, Roswall J, Peng Y, Feng Q, Jia H, Kovatcheva-Datchary P, Li Y, Xia Y, Xie H, Zhong H. Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell Host Microbe. 2015;17:690-703. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1477] [Cited by in RCA: 1978] [Article Influence: 197.8] [Reference Citation Analysis (0)] |
| 111. | Penders J, Thijs C, Vink C, Stelma FF, Snijders B, Kummeling I, van den Brandt PA, Stobberingh EE. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics. 2006;118:511-521. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1513] [Cited by in RCA: 1528] [Article Influence: 80.4] [Reference Citation Analysis (0)] |
| 112. | Mårild K, Ludvigsson JF, Størdal K. Current evidence on whether perinatal risk factors influence coeliac disease is circumstantial. Acta Paediatr. 2016;105:366-375. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 16] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 113. | Adlercreutz EH, Wingren CJ, Vincente RP, Merlo J, Agardh D. Perinatal risk factors increase the risk of being affected by both type 1 diabetes and coeliac disease. Acta Paediatr. 2015;104:178-184. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 41] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 114. | Canova C, Zabeo V, Pitter G, Romor P, Baldovin T, Zanotti R, Simonato L. Association of maternal education, early infections, and antibiotic use with celiac disease: a population-based birth cohort study in northeastern Italy. Am J Epidemiol. 2014;180:76-85. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 82] [Cited by in RCA: 107] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 115. | Decker E, Hornef M, Stockinger S. Cesarean delivery is associated with celiac disease but not inflammatory bowel disease in children. Gut Microbes. 2011;2:91-98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 48] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 116. | Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, Blekhman R, Beaumont M, Van Treuren W, Knight R, Bell JT. Human genetics shape the gut microbiome. Cell. 2014;159:789-799. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1911] [Cited by in RCA: 2150] [Article Influence: 215.0] [Reference Citation Analysis (0)] |
| 117. | Murphy K, O’ Shea CA, Ryan CA, Dempsey EM, O’ Toole PW, Stanton C, Ross RP. The gut microbiota composition in dichorionic triplet sets suggests a role for host genetic factors. PLoS One. 2015;10:e0122561. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 24] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 118. | Stewart JA, Chadwick VS, Murray A. Investigations into the influence of host genetics on the predominant eubacteria in the faecal microflora of children. J Med Microbiol. 2005;54:1239-1242. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 130] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 119. | Olivares M, Neef A, Castillejo G, Palma GD, Varea V, Capilla A, Palau F, Nova E, Marcos A, Polanco I. The HLA-DQ2 genotype selects for early intestinal microbiota composition in infants at high risk of developing coeliac disease. Gut. 2015;64:406-417. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 203] [Cited by in RCA: 222] [Article Influence: 22.2] [Reference Citation Analysis (1)] |
| 120. | Sellitto M, Bai G, Serena G, Fricke WF, Sturgeon C, Gajer P, White JR, Koenig SS, Sakamoto J, Boothe D. Proof of concept of microbiome-metabolome analysis and delayed gluten exposure on celiac disease autoimmunity in genetically at-risk infants. PLoS One. 2012;7:e33387. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 192] [Cited by in RCA: 184] [Article Influence: 14.2] [Reference Citation Analysis (0)] |
| 121. | Monsuur AJ, de Bakker PI, Alizadeh BZ, Zhernakova A, Bevova MR, Strengman E, Franke L, van’t Slot R, van Belzen MJ, Lavrijsen IC. Myosin IXB variant increases the risk of celiac disease and points toward a primary intestinal barrier defect. Nat Genet. 2005;37:1341-1344. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 170] [Article Influence: 8.5] [Reference Citation Analysis (0)] |