Published online Aug 21, 2013. doi: 10.3748/wjg.v19.i31.5144
Revised: March 28, 2013
Accepted: July 17, 2013
Published online: August 21, 2013
Processing time: 209 Days and 3.9 Hours
AIM: To characterize high mobility group box chromosomal protein 1 (HMGB1) polymorphisms in patients infected with hepatitis B virus (HBV) and determine the different patterns in patient subgroups.
METHODS: A total of 1495 unrelated Han Chinese HBV carriers were recruited in this hospital-based case-control study. The HMGB1 1176 G/C polymorphism was genotyped by polymerase chain reaction-restriction fragment length polymorphism assay.
RESULTS: A significant association was observed between HMGB1 1176 G/C polymorphism and outcome of HBV infection. The subjects bearing 1176G/G genotype had an increased risk of susceptibility to chronic hepatitis B, liver cirrhosis and severe hepatitis B when compared with those bearing at least one 1176C allele.
CONCLUSION: Patients with 1176G/G genotype of HMGB1 gene are more likely to have a progressive status in HBV infection.
Core tip: We analyzed the relationship between the high mobility group box chromosomal protein 1 (HMGB1) 1176 G/C polymorphism and the susceptibility and outcome to hepatitis B virus (HBV) infection in a large hospital-based case-control study. Our results indicated that patients with 1176G/G genotype of HMGB1 gene are more likely to have a progressive status in HBV infection. Our study emphasizes the importance of HMGB1 in the pathophysiology of HBV-related diseases on the population level and will provide researchers new clue for the further basic research in pathogenesis of chronic HBV infection.
-
Citation: Deng CQ, Deng GH, Wang YM.
HMGB1 gene polymorphisms in patients with chronic hepatitis B virus infection. World J Gastroenterol 2013; 19(31): 5144-5149 - URL: https://www.wjgnet.com/1007-9327/full/v19/i31/5144.htm
- DOI: https://dx.doi.org/10.3748/wjg.v19.i31.5144
Hepatitis B virus (HBV) infection is associated with a variety of diseases, including asymptomatic carrier (AsC), fulminant hepatitis, chronic hepatitis (CHB), liver cirrhosis (LC), and hepatocellular carcinoma (HCC). Persistent HBV infection has been considered as a multifactorial and polygenic disorder with viral, environmental and genetic components. HBV genomic variability and a number of conventional risk factors, including age, gender, concurrent infection with hepatitis C virus, hepatitis D virus and human immune deficiency virus, are clearly the important factors contributing to the incidence of persistent HBV infection[1-4]. However, segregation analysis and twin studies strongly support the role of host genetic components in determining the chronicity of HBV infection[5,6]. A known and unknown number of identified or unidentified genes are likely to modify the susceptibility to persistent HBV infection[7-10]. Single nucleotide polymorphism (SNP) is currently believed to be a powerful tool for identifying genetic susceptibilities to common complex diseases[11,12].
The intranuclear architectural protein termed high mobility group box chromosomal protein 1 (HMGB1) has recently been identified as a potent proinflammatory mediator when passively released to extracellular by necrotic cells, as opposed to apoptotic cells that will induce inflammation[13,14]. Furthermore, HMGB1 can also be actively secreted by stimulated macrophages or monocytes[15-17]. Active secretion from living inflammatory cells and passive release from necrotic cells implicate that HMGB1 may play a central role in proinflammatory reactions. It is well known that HBV infection is closely related with cytokines. Polymorphisms of cytokine gene, such as human leukocyte antigen, estrogen receptor alpha (ESR1), have been reported to be associated with HBV infection[18-21]. However, so far there has been no report on the association between HMGB1 gene and HBV infection. We conducted a hospital-based case-control study including more than one thousand subjects with HBV infection to characterize the relationship between HMGB1 gene polymorphism and HBV infection.
Patients with HBV infection were randomly selected from the outpatient and inpatient referral center affiliated to the Institute for Infectious Diseases of Southwest Hospital treated between February 2002 and February 2012. Informed consent was obtained from all the patients to participate in the study. Participants finally included in the current study were from a subset of unrelated individuals from the referral center. The diagnostic criteria for chronic HBV infection were as follows: persistent presence of hepatitis B surface antigen (HBsAg), absence of anti-hepatitis B surface antibodies (anti-HBs), presence of anti-core IgG antibodies (anti-HBc), and presence of hepatitis B early antigen (HBeAg) or anti-hepatitis B e antibodies (anti-HBe) for 6 mo or longer despite of virus replication. Asymptomatic carriers had no fluctuation of serum alanine aminotransferase (ALT) levels and no obvious clinical symptoms. Chronic hepatitis B had a serum ALT fluctuation, 1 × the upper limit of normal (ULN) < ALT < 5 × ULN, with or without other abnormal hepatic functions. Severe hepatitis B (SHB), which is currently equal to acute-on-chronic liver failure, presents the following symptoms: (1) fatigue with striking gastrointestinal tract symptoms; (2) rapidly worsening jaundice, with serum total bilirubin (TBIL) 10 times higher than ULN, or with a daily increase ≥ 17.1 μmol/L; (3) hemorrhagic tendency with international normalised ratio ≥ 1.5 or prothrombin activity ≤ 40% where other causes have been excluded; (4) progressive reduction in liver size; and (5) occurrence of hepatic encephalopathy. Liver cirrhosis and HCC were confirmed by liver biopsy, ultrasound, and/or computerized tomography scan. Healthy control individuals were recruited from Red Cross blood donor centers with or without anti-HBs, but HBsAg, anti-HBc, HBeAg, and anti-HBe were negative.
The leukocytes genomic DNA from 5 mL whole blood was isolated using Miller’s method[22]. DNA samples were diluted to 8 ng/μL and distributed into 96-well plates (DNA panels), with 94 samples and 2 controls (DNA-free water) in each plate.
We used the current recommendations of human genome SNP described at http://www.ncbi.nlm.gov/SNP under accession number NT024524. The higher allele variation frequency selected in position 1176 G/C, the intron 4 of HMGB1 gene, was studied to determine whether any association identified was specific to HBV infection. The SNP was named in a same way to HMGB1 (1176G/C).
The genotyping was analyzed by polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) assay. Appropriate primer pairs (sense 5’-3’GTCTCCTTTGCCCAGTGTATCTC and anti-sense 5’-3’GTACACAGCCTTTGTCTGAGTCTG) were designed by Primer Premier 5.0 software (Premier Biosoft International, Palo Alto, CA, United States). PCR condition was as follows: one cycle of predenature 3 min at 95 °C, 30 cycles of denature 30 s at 94 °C, hybridization for 30 s at 54 °C, an extension cycle of 50 s at 72 °C, and a last cycle of delay 5 min at 72 °C. Restriction enzyme BcLI (recognition site T/GATCA) was obtained from NEB; the fragments were separated by electrophoresis on 3% agarose gel and stained with ethidium bromide for visualization under ultraviolet light. The observed genotypes were also identified by direct sequencing before large-scale test was started.
An allele frequency was directly calculated by its genotype. The observed genotype frequencies and allele frequency were compared using χ2 test between the variables to determine if they were in Hardy-Weinberg equilibrium. An observed P > 0.01 was considered in Hardy-Weinberg equilibrium, and P < 0.05 was considered significantly different between the variables. All the analyses were performed with SPSS11.0 statistical software (SPSS Inc., Chicago, IL, United States).
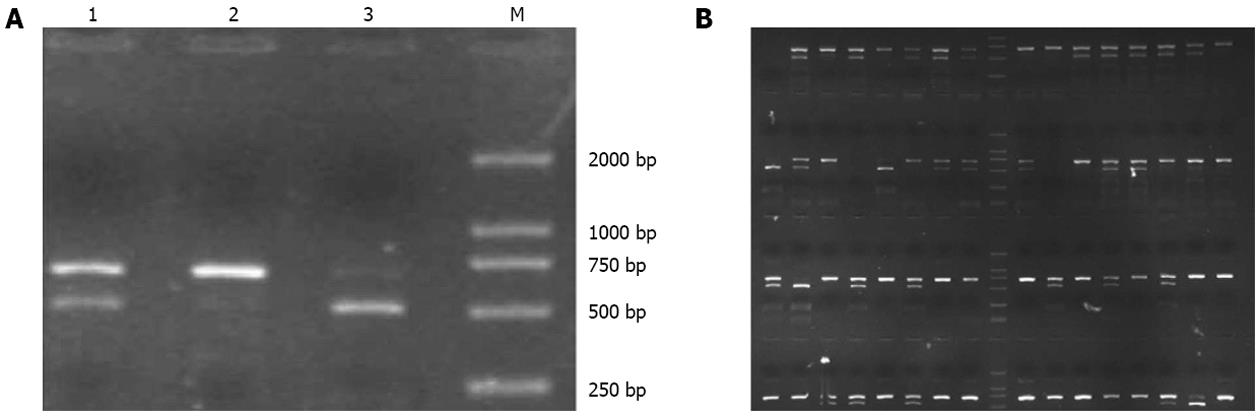
HMGB1 1176G/C polymorphism was genotyped by PCR-RFLP assay (Figure 1). A total of 1495 clearly diagnosed and genotyped patients were enrolled. The clinical characteristics, such as age and sex, are listed in Table 1. Apparently, age or sex difference existed in the studied subgroups. Hardy-Weinberg equilibrium by χ2 test showed P = 0.494 > 0.01 (Table 1), which confirmed that the studied group was in Hardy-Weinberg equilibrium.
| Sex | Age (yr) | Genotype | Allele frequency | ||||
| (M/F) | (mean ± SD) | GG | CC | GC | G | C | |
| AsC (n = 199) | 116/83 | 34.762 ± 11.282 | 107 | 9 | 83 | 0.7462 | 0.2538 |
| AHB (n = 15) | 11/4 | 30.201 ± 10.221 | 9 | 1 | 5 | ||
| CHB (n = 929) | 730/199 | 34.312 ± 11.549 | 572 | 33 | 324 | 0.6530 | 0.3470 |
| SHB (n = 157) | 129/28 | 39.989 ± 11.792 | 91 | 6 | 60 | 0.7707 | 0.2293 |
| LC (n = 175) | 142/33 | 41.950 ± 11.437 | 104 | 13 | 58 | 0.7600 | 0.2300 |
| HCC (n = 20) | 14/6 | 49.256 ± 12.232 | 10 | 1 | 9 | ||
| LC + CHB (n = 1104) | 872/232 | 35.461 ± 11.642 | 676 | 46 | 382 | 0.7853 | 0.2147 |
| LC + CHB + SHB (n = 1261) | 1001/260 | 36.001 ± 11.852 | 767 | 52 | 442 | 0.7835 | 0.2165 |
Because age or sex difference existed in the studied subgroups, it is essential to detect the association between observed SNP and HBV infected subgroups, and age and sex factors were considered by logistic regression (Table 2). A statistically significant difference in the distribution of HMGB1 polymorphism (1176G/C) was observed between subgroups of AsC and LC (OR = 1.571, 95%CI: 1.108-2.227, P = 0.011, codominant model); AsC and CHB (OR = 1.354, 95%CI: 1.085-1.689, P = 0.007, codominant model); AsC and CHB + SHB + LC (OR = 1.401, 95%CI: 1.010-1.944, P = 0.044, recessive model); OR = 1.329, 95%CI: 1.070-1.651, P = 0.010, codominant model, AsC and CHB + LC (OR = 1.406, 95%CI: 1.011-1.956, P = 0.043, recessive model; OR = 1.355, 95%CI: 1.088-1.687, P = 0.007, codominant model).
| Subgroup | Dominant model | Recessive model | Codominant model | |||
| P | OR (95%CI) | P | OR (95%CI) | P | OR (95%CI) | |
| LC/CHB | 0.075 | 0.533 (0.267-1.065) | 0.945 | 0.988 (0.701-1.392) | 0.120 | 0.844 (0.682-1.045) |
| LC/SHB | 0.183 | 0.508 (0.188-1.376) | 0.747 | 1.075 (0.693-1.669) | 0.326 | 0.872 (0.664-1.146) |
| LC/AsC | 0.168 | 2.320 (0.702-7.672) | 0.108 | 1.572 (0.906-2.728) | 0.011 | 1.571 (1.108-2.227) |
| AsC/SHB | 0.473 | 1.616 (0.436-5.999) | 0.489 | 1.204 (0.711-2.040) | 0.206 | 1.254 (0.883-1.782) |
| AsC/CHB | 0.238 | 1.660 (0.716-3.851) | 0.052 | 1.389 (0.997-1.935) | 0.007 | 1.354 (1.085-1.689) |
| SHB/CHB | 0.916 | 1.050 (0.425-2.592) | 0.474 | 1.137 (0.800-1.614) | 0.462 | 1.090 (0.866-1.371) |
| SHB + LC/CHB + AsC | 0.362 | 0.760 (0.421-1.371) | 0.824 | 0.971 (0.747-1.262) | 0.316 | 0.918 (0.777-1.085) |
| AsC/CHB + SHB + LC | 0.330 | 1.505 (0.662-3.421) | 0.044 | 1.401 (1.010-1.944) | 0.010 | 1.329 (1.070-1.651) |
| AsC/CHB + LC | 0.256 | 1.619 (0.704-3.721) | 0.043 | 1.406 (1.011-1.956) | 0.007 | 1.355 (1.088-1.687) |
HMGB1 is a nuclear DNA-binding protein, which also functions as a pleiotropic cytokine, implicated in the pathology of several different immune-mediated diseases. The human HMGB1 gene is located on chromosome 13. Kornblit et al[23] firstly elaborated six polymorphisms and four mutations identified in the HMGB1 gene, located in -1615A/G, 982C/T, 3814C/G, 1779T/G, -196C/A, 1808C/G, 4519_4521delGAT, -1377delA,1747delT, 1888insT, respectively. In other studies, several associations have been observed, revealing the importance of the genetic variation in the HMGB1 gene. In their report, the -1377delAA/- genotype or the -1377delA-/- genotype showed a significant association with delayed mortality, independent of age and number of the systemic inflammatory response syndrome (SIRS) criteria[24]. Subsequent estimation revealed that several polymorphisms have a potential regulatory impact on HMGB1 transcription. Genetically determined risk factors associated with early and late mortality and death due to infection have been identified, explaining some of the inherited risks in this heterogeneous patient population. Associations between genetic variation and disease severity parameters are also established. Studies of association between HMGB1 polymorphisms and disease have been also reported with allogeneic hematopoietic cell transplantation (HCT)[25]. Patient homozygosity or heterozygosity for the-1377delA minor allele is associated with increased risk of relapse and increased relapse-related mortality. Furthermore, patient homozygosity for the 3814C/G minor allele is associated with increased overall survival and progression-free survival. Patient carriage of the 2351insT minor allele can reduce the risk from grade II to IV acute graft-versus-host disease whereas donor homozygosity is associated with chronic acute graft-versus-host disease. These findings suggest that the inherited variation in HMGB1 is associated with outcome after allogeneic HCT following myeloablative conditioning regimens. Zeng et al[26] found that three SNPs act as tag SNPs for the entire HMGB1 gene in multiple organ dysfunction syndromes. The rs2249825 and the haplotype TCG can be used as relevant risk estimate for the development of sepsis in patients with major trauma.
As is well known, the susceptibility of HBV infection is closely related to the variation of some important genes. Deng et al[27] have demonstrated that polymorphisms at the ESR1 gene locus are associated with persistent HBV infection. Subsequently, Yan et al[18] have also demonstrated an association between cis-acting regulatory variation of the ESR1 gene and hepatitis B virus-related liver cirrhosis. Some important variations of cytokine gene also influenced the susceptibility to HBV infection. Deng et al[28] have found that the novel regulatory polymorphism G-201A in the promoter of interferon gamma-inducible protein of 10 kilodaltons (IP-10, CXCL10) gene might be a part of the genetic variation underlying the susceptibility of individuals to disease progression of chronic HBV infection. In another study, Yan et al[29] demonstrated that the -592C allele and the -1082A-819C-592C haplotype in the IL-10 gene promoter were associated with an increased susceptibility to acute liver failure in HBV carriers.
Nevertheless, there are few reports about the association between HMGB1 gene and HBV infection, especially reports about the association between HMGB1 polymorphisms and HBV infection. In this study, we used the current recommendations of human genome SNPs described at http://www.ncbi.nlm.gov/SNP under accession number NT024524. The higher allele variation frequency was selected in position 1176 G/C, the intron 4 of HMGB1 gene. There has been no report about this SNP up to date. We genotyped the polymorphisms of 1495 cases, including AsC, CHB, SHB, LC and HCC. The distribution of HMGB1 1176G/C genotypes in studied sample of unrelated men and women from the referral center were in Hardy-Weinberg equilibrium (P > 0.01), so it is important to consider whether our studied sample could be representative. Yan et al[18] and Deng et al[28] had scanned the polymorphisms on the same cohort. Because differences in age or sex existed in the studied subgroups, we detected the association by logistic regression between observed SNP and HBV infected subgroups, and the age and sex factors were considered. As a result, there was statistically significant evidence of association. The fraction calculated by relative risk indicated that HMGB1 1176G/G genotype was more susceptible to CHB, LC and SHB than 1176C/C and 1176G/C genotype. In other words, the patients with 1176G/G genotype of HMGB1 gene are more likely to have a progressive status in HBV infection. The results suggest that allele 1176G is closely related to the ponderance of disease. These findings underscore a potentially important role of HMGB1 in influencing the development of HBV infection.
In another study, we had successfully cloned and analyzed 154 bp nucleotides in intron 4 near the fourth exon-intron boundary, and found that the region contained sequences 1176 G/C polymorphism characteristic of an enhancer using PGL3 reporter gene systems. We demonstrated that the SNP 1176 G/C could affect the function. Furthermore, this activity was enhanced by the SNP: G→C change in position 1176,providing the basis for molecular investigations of the HMGB1 gene in HBV infection. Subsequent reports would focus on this investigation.
In conclusion, our results showed that the HMGB1 1176G/G genotype was related to the outcomes of hepatitis B infection, and patients with 1176G/G genotype of HMGB1 gene are more likely to have a progressive status in HBV infection. The subjects bearing 1176G/G genotype have an increased risk of susceptibility to CHB, LC and SHB compared with those bearing at least one 1176C allele. However, further work is needed to validate our results, and clarify more potential functions of human HMGB1 gene.
Chronic hepatitis B virus (HBV) infection is a serious public health problem worldwide. Host genetic factors play a role in determining to the outcome and progression of the infection. A large number of studies on the association between cytokine gene polymorphisms and the risk of chronic hepatitis B (CHB) have been conducted. High mobility group box 1 (HMGB1) functioned as a pleiotropic cytokine and implicated in the pathology of several different immune-mediated diseases. However, there has been no report about the association between HMGB1 gene and HBV infection.
HMGB1 has recently been identified as a potent proinflammatory mediator when passively released extracellularly by necrotic cells, as opposed to apoptotic cells that will induce inflammation. Furthermore, HMGB1 can also be actively secreted by stimulated macrophages or monocytes. Active secretion from living inflammatory cells and passive release from necrotic cells implicate that HMGB1 may play a central role in proinflammatory reactions.
This study characterize the relationship between HMGB1 gene polymorphism and HBV infection, and concluded that the HMGB1 1176G/G genotype was related to the outcomes of hepatitis B infection, and patients with 1176G/G genotype of HMGB1 gene are more likely to have a progressive status in HBV infection.
The study results suggest that the subjects bearing HMGB1 1176G/G genotype have an increased risk of susceptibility to CHB, liver cirrhosis and severe hepatitis B compared with those bearing at least one 1176C allele, which will provide new clue for the further basic research in pathogenesis of chronic HBV infection.
The authors have done a good job and found an association between the 1176G/C polymorphism of HMGB1, a proinflammatory mediator, and hepatitis B virus infection. The results are interesting and suggest that HMGB1 is a mediator of the immune response to HBV infection.
P- Reviewer Zhang L S- Editor Huang XZ L- Editor A E- Editor Zhang DN
| 1. | Kacprzak-Bergman I, Nowakowska B. Influence of genetic factors on the susceptibility to HBV infection, its clinical pictures, and responsiveness to HBV vaccination. Arch Immunol Ther Exp (Warsz). 2005;53:139-142. [PubMed] |
| 2. | Zhang YY, Theele DP, Summers J. Age-related differences in amplification of covalently closed circular DNA at early times after duck hepatitis B virus infection of ducks. J Virol. 2005;79:9896-9903. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 17] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 3. | Song BC, Cui XJ, Kim H. Hepatitis B virus genotypes in Korea: an endemic area of hepatitis B virus infection. Intervirology. 2005;48:133-137. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 70] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 4. | Michitaka K, Horiike N, Chen Y, Yatsuhashi H, Yano M, Kojima N, Ohkubo K, Tanaka Y, Yamamoto K, Ohno N. Infectious source factors affecting the severity of sexually transmitted acute hepatitis due to hepatitis B virus genotype C. Intervirology. 2005;48:112-119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 11] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 5. | Song le H, Binh VQ, Duy DN, Jüliger S, Bock TC, Luty AJ, Kremsner PG, Kun JF. Mannose-binding lectin gene polymorphisms and hepatitis B virus infection in Vietnamese patients. Mutat Res. 2003;522:119-125. [PubMed] |
| 6. | Höhler T, Reuss E, Evers N, Dietrich E, Rittner C, Freitag CM, Vollmar J, Schneider PM, Fimmers R. Differential genetic determination of immune responsiveness to hepatitis B surface antigen and to hepatitis A virus: a vaccination study in twins. Lancet. 2002;360:991-995. [PubMed] |
| 7. | Wang C, Tang J, Song W, Lobashevsky E, Wilson CM, Kaslow RA. HLA and cytokine gene polymorphisms are independently associated with responses to hepatitis B vaccination. Hepatology. 2004;39:978-988. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 152] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 8. | Miyazoe S, Hamasaki K, Nakata K, Kajiya Y, Kitajima K, Nakao K, Daikoku M, Yatsuhashi H, Koga M, Yano M. Influence of interleukin-10 gene promoter polymorphisms on disease progression in patients chronically infected with hepatitis B virus. Am J Gastroenterol. 2002;97:2086-2092. [PubMed] |
| 9. | Höhler T, Kruger A, Gerken G, Schneider PM, Meyer zum Büschenefelde KH, Rittner C. A tumor necrosis factor-alpha (TNF-alpha) promoter polymorphism is associated with chronic hepatitis B infection. Clin Exp Immunol. 1998;111:579-582. [PubMed] |
| 10. | Sobao Y, Sugi K, Tomiyama H, Saito S, Fujiyama S, Morimoto M, Hasuike S, Tsubouchi H, Tanaka K, Takiguch M. Identification of hepatitis B virus-specific CTL epitopes presented by HLA-A*2402, the most common HLA class I allele in East Asia. J Hepatol. 2001;34:922-929. [PubMed] |
| 11. | Collins FS, Patrinos A, Jordan E, Chakravarti A, Gesteland R, Walters L. New goals for the U.S. Human Genome Project: 1998-2003. Science. 1998;282:682-689. [PubMed] |
| 13. | O’Connor KA, Hansen MK, Rachal Pugh C, Deak MM, Biedenkapp JC, Milligan ED, Johnson JD, Wang H, Maier SF, Tracey KJ. Further characterization of high mobility group box 1 (HMGB1) as a proinflammatory cytokine: central nervous system effects. Cytokine. 2003;24:254-265. [PubMed] |
| 14. | Guazzi S, Strangio A, Franzi AT, Bianchi ME. HMGB1, an architectural chromatin protein and extracellular signalling factor, has a spatially and temporally restricted expression pattern in mouse brain. Gene Expr Patterns. 2003;3:29-33. [PubMed] |
| 15. | Dumitriu IE, Baruah P, Manfredi AA, Bianchi ME, Rovere-Querini P. HMGB1: guiding immunity from within. Trends Immunol. 2005;26:381-387. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 247] [Cited by in RCA: 271] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 16. | DeMarco RA, Fink MP, Lotze MT. Monocytes promote natural killer cell interferon gamma production in response to the endogenous danger signal HMGB1. Mol Immunol. 2005;42:433-444. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 71] [Cited by in RCA: 74] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 17. | Weigand MA, Hörner C, Bardenheuer HJ, Bouchon A. The systemic inflammatory response syndrome. Best Pract Res Clin Anaesthesiol. 2004;18:455-475. [PubMed] |
| 18. | Yan Z, Tan W, Xu B, Dan Y, Zhao W, Deng C, Chen W, Tan S, Mao Q, Wang Y. A cis-acting regulatory variation of the estrogen receptor α (ESR1) gene is associated with hepatitis B virus-related liver cirrhosis. Hum Mutat. 2011;32:1128-1136. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 18] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 19. | Yan Z, Tan S, Dan Y, Sun X, Deng G, Wang Y. Relationship between HLA-DP gene polymorphisms and clearance of chronic hepatitis B virus infections: case-control study and meta-analysis. Infect Genet Evol. 2012;12:1222-1228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 21] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 20. | Deng CQ, Deng GH, Wang YM. Relationship between polymorphisms of 3-hydroxy-3-methylglutaryl coenzyme A reductase gene and hepatitis B virus infection. Shijie Huaren Xiaohua Zazhi. 2005;17:2086-2089. |
| 21. | Deng CQ, Deng GH, Wang YM. eNOS gene 894G/T polymorphisms among patients infected with HBV. Virologica Sinica. 2005;20:476-479. |
| 22. | Miller SA, Dykes DD, Polesky HF. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988;16:1215. [PubMed] |
| 23. | Kornblit B, Munthe-Fog L, Petersen SL, Madsen HO, Vindeløv L, Garred P. The genetic variation of the human HMGB1 gene. Tissue Antigens. 2007;70:151-156. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 28] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 24. | Kornblit B, Munthe-Fog L, Madsen HO, Strøm J, Vindeløv L, Garred P. Association of HMGB1 polymorphisms with outcome in patients with systemic inflammatory response syndrome. Crit Care. 2008;12:R83. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 41] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 25. | Kornblit B, Masmas T, Petersen SL, Madsen HO, Heilmann C, Schejbel L, Sengeløv H, Müller K, Garred P, Vindeløv L. Association of HMGB1 polymorphisms with outcome after allogeneic hematopoietic cell transplantation. Biol Blood Marrow Transplant. 2010;16:239-252. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 44] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 26. | Zeng L, Zhang AQ, Gu W, Chen KH, Jiang DP, Zhang LY, Du DY, Hu P, Huang SN, Wang HY. Clinical relevance of single nucleotide polymorphisms of the high mobility group box 1 protein gene in patients with major trauma in southwest China. Surgery. 2012;151:427-436. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 19] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 27. | Deng G, Zhou G, Zhai Y, Li S, Li X, Li Y, Zhang R, Yao Z, Shen Y, Qiang B. Association of estrogen receptor alpha polymorphisms with susceptibility to chronic hepatitis B virus infection. Hepatology. 2004;40:318-326. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 83] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 28. | Deng G, Zhou G, Zhang R, Zhai Y, Zhao W, Yan Z, Deng C, Yuan X, Xu B, Dong X. Regulatory polymorphisms in the promoter of CXCL10 gene and disease progression in male hepatitis B virus carriers. Gastroenterology. 2008;134:716-726. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 77] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 29. | Yan Z, Tan W, Zhao W, Dan Y, Wang X, Mao Q, Wang Y, Deng G. Regulatory polymorphisms in the IL-10 gene promoter and HBV-related acute liver failure in the Chinese population. J Viral Hepat. 2009;16:775-783. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 30] [Article Influence: 1.9] [Reference Citation Analysis (0)] |