Published online May 7, 2007. doi: 10.3748/wjg.v13.i17.2496
Revised: January 14, 2007
Accepted: March 23, 2007
Published online: May 7, 2007
AIM: To study the relationship between quantitative structure and pharmacokinetics (QSPkR) of fluoroquinolone antibacterials.
METHODS: The pharmacokinetic (PK) parameters of oral fluoroquinolones were collected from the litera-ture. These pharmacokinetic data were averaged, 19 compounds were used as the training set, and 3 served as the test set. Genetic function approximation (GFA) module of Cerius2 software was used in QSPkR analysis.
RESULTS: A small volume and large polarizability and surface area of substituents at C-7 contribute to a large area under the curve (AUC) for fluoroquinolones. Large polarizability and small volume of substituents at N-1 contribute to a long half life elimination.
CONCLUSION: QSPkR models can contribute to some fluoroquinolones antibacterials with excellent pharmacokinetic properties.
- Citation: Cheng D, Xu WR, Liu CX. Relationship of quantitative structure and pharmacokinetics in fluoroquinolone antibacterials. World J Gastroenterol 2007; 13(17): 2496-2503
- URL: https://www.wjgnet.com/1007-9327/full/v13/i17/2496.htm
- DOI: https://dx.doi.org/10.3748/wjg.v13.i17.2496
H pylori is generally considered to be the most important cause of peptic ulcer diseases, gastric adenocarcinoma and mucosa-associated lymphoid tissue (MALT) lymphoma of the stomach[1]. The widespread use of antibacterial therapy is suggested to be the cause for the decline in the prevalence of H pylori infection[2]. Among the different types of antibacterial agents, the effects of fluoroquinolones are better and have attracted much attention. Unfortunately, complete eradication of H pylori is still in the initial stage, especially in South East Asia and Southern Europe, where resistance to antibiotics has become more prevalent[3]. It is therefore important to search for better antibacterial agents against resistant H pylori strains[4].
Successful drugs must have suitable properties in toxicity, bioavailability and pharmacokinetic parameters. Screening of a large number of compounds with excellent absorption, distribution, metabolism, and excretion (ADME) properties is time-consuming and expensive[5]. So the extension of the idea of quantitative structure-activity relationship to the pharmacokinetics has led to the emergence of a new tool called the quantitative structure pharmacokinetic relationship (QSPkR) studies. QSPkR studies can be utilized at early stages of drug design. Both one- and two-dimensional topological indices have been used extensively to numerically relate molecular structure with activity[6]. These descriptors rely only on the molecular graph for their calculation. In contrast, three-dimensional descriptors require the absolute conformation of a molecule, and have been successfully used to develop QSPkR analysis[7].
The QSPkR models integrated properties of che-mical structures (e.g. LogP) and their pharmacokinetic parameters (total clearance, distribution volume, etc.) of fluoroquinolones have been reported[8]. But these existing models cannot demonstrate the influence of the substituents to pharmacokinetic parameters. That is to say, these models can only predict pharmacokinetic parameters of the existing chemicals.
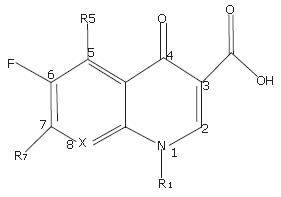
After examining the structures of all marketed fluoro-quinolones, we found that their diversities in structures were mainly within R1 and R7 (Figure 1). Considering the connections between the groups (R1 and R7) and matrix were single bonds, the conjugations between groups and matrix were limited, and the groups had relatively independent properties. To simplify the design for high efficiency in practice, the properties of fragments were applied as the descriptors of calculation. In this study, a two-step process was used to develop QSPkR models clinically using fluoroquinolone antibacterials. The first step was to calculate properties related to chemical structures and their conformation, especially constituent structures. These properties include 2D descriptors representing physical properties (logP), 3D descriptors (volume), and quantum chemical parameters (polarizability). After calculating these properties, the QSPkR models were developed by multivariate linear regression based on genetic algorithms.
Using these QSPkR models, we can illustrate how the changes at N-1 and C-7 of the fluoroquinolones affect their pharmacokinetic parameters. Hopefully, these QSPkR models can contribute to some fluoroquinolones with excellent pharmacokinetic properties.
All 22 compounds used in this study are analogues of the fluoroquinolone antibacterials which are widely used clinically except DW116 (No.5). The matrix of the compounds is shown in Figure 1, and their detailed substituents are listed in Figure 2.
The PK parameters of these fluoroquinolones were collected from literature[9-68]. Data were taken from the studies of oral fluoroquinolones. These pharmacokinetic data were averaged after AUC and Cmax data were normalized by 100 mg of drugs (Table 1). t1/2 in this paper is elimination half life, it is also known as t1/2(®). Nineteen compounds were used as the training set, and the others served as test set.
| Compounds | PK parameters | References | ||||||
| 1AUC0-∞ (μg·h/mL) | 1t1/2(h) | 1Cmax (mg/L) | ||||||
| No. | Name | Range | Average | Range | Average | Range | Average | |
| Training set | ||||||||
| 1 | Amifloxacin | 5.5-5.62 | 5.57 | 3.58-4.83 | 4.14 | 0.9-1.26 | 1.14 | 9, 10 |
| 2 | Balofloxacin | 8.55 | 8.55 | 7.8 | 7.8 | 1.08 | 1.08 | 11 |
| 3 | Ciprofloxacin | 2.12-3.53 | 2.56 | 3.01-4.7 | 4.16 | 0.4-0.69 | 0.56 | 12-15 |
| 4 | Clinafloxacin | 4.63-5.93 | 5.34 | 5.09-6.13 | 5.74 | 0.6-0.84 | 0.72 | 16-18 |
| 5 | DW116 | 18.54-23.3 | 21.86 | 14.53-18.7 | 15.82 | 1.1-1.22 | 1.17 | 19 |
| 6 | Enoxacin | 2.9-5.47 | 4.36 | 2.35-4.98 | 3.54 | 0.62-0.81 | 0.66 | 20-22 |
| 7 | Gatifloxacin | 6.5-8.92 | 7.87 | 6.52-8.6 | 7.46 | 0.84-1.03 | 0.9 | 23-26 |
| 8 | Gemifloxacin | 2.79-3.43 | 3.02 | 5.87-8.2 | 6.65 | 0.46-0.73 | 0.56 | 27-29 |
| 9 | Grepafloxacin | 2.83-4.05 | 3.43 | 9.2-12.7 | 11.53 | 0.24-0.41 | 0.32 | 11, 12, 30, 31 |
| 10 | Levofloxacin | 8.96-9.5 | 9.33 | 6-7.4 | 6.78 | 0.16-0.3 | 0.24 | 32-34 |
| 11 | Lomefloxacin | 8.05-13.53 | 9.84 | 5.5-12.7 | 7.73 | 0.95-1.18 | 1.06 | 35-37 |
| 12 | Norfloxacin | 1.7-1.85 | 1.77 | 3.5-4.02 | 3.7 | 0.32-0.36 | 0.33 | 38-40 |
| 13 | Ofloxacin | 6.68-11.64 | 7.67 | 4.6-6.7 | 5.32 | 0.71-1.33 | 0.87 | 41-45 |
| 14 | Pefloxacin | 24.4-40.78 | 29.97 | 10.9-15.06 | 14.63 | 1.03-1.68 | 1.44 | 46-48 |
| 15 | Rufloxacin | 35.8-44.03 | 39.43 | 28.2-40 | 34.25 | 0.68-1.13 | 0.99 | 49-52 |
| 16 | Sitafloxacin | 5.62-6.02 | 5.88 | 4.6-7 | 5.4 | 0.9-0.93 | 0.92 | 53-54 |
| 17 | Sparfloxacin | 8.08-11.96 | 8.35 | 16.5-25.56 | 20.06 | 0.23-0.4 | 0.34 | 55-57 |
| 18 | Temafloxacin | 7.42-10.63 | 8.45 | 7.8-10.6 | 8.55 | 0.61-0.9 | 0.74 | 58-60 |
| 19 | Trovafloxacin | 9.75-14.47 | 11.91 | 7.8-10.8 | 9.66 | 0.97-1.5 | 1.23 | 61-63 |
| Test set | ||||||||
| 20 | Difloxacin | 26.6-28.3 | 27.8 | 20.6-28.8 | 25.7 | 1.02-1.1 | 1.04 | 64 |
| 21 | Fleroxacin | 16.3-20.65 | 18.13 | 7.9-13 | 11.02 | 1.19-1.58 | 1.4 | 32, 65, 66 |
| 22 | Tosufloxacin | 1.49-3.3 | 2.62 | 3.6-4.85 | 4.02 | 0.21-0.4 | 0.34 | 67-69 |
The 3D structure of each compound was constructed by HyperChem 7.0 (Hypercube Inc., USA) and then optimized with MM+ force field. All molecules were aligned by minimizing the rms distance of their matrix by SYBYL 7.0 (Tripos Inc., 2004). The alignment of molecules is displayed in Figure 3. The descriptors were calculated for substituents R1 and R7 by HyperChem 7.0. The definitions of all descriptors are shown in Table 2.
| Descriptors | Physicochemical meaning |
| SA7 | Surface area (grid) of R7 |
| V7 | Volume of R7 |
| HE7 | Hydration energy of R7 |
| LP7 | Logp of R7 |
| RF7 | Refractivity of R7 |
| P7 | Polarizability of R7 |
| MW7 | Molecular weight of R7 |
| SA1 | Surface area (grid) of R1 |
| V1 | Volume of R1 |
| HE1 | Hydration energy of R1 |
| LP1 | Logp of R1 |
| RF1 | Refractivity of R1 |
| P1 | Polarizability of R1 |
| MW1 | Molecular weight of R1 |
The logarithmic values of the PK parameters were used as the dependent variables. All the descriptors were scaled by the mean values of data from the training set .
The models related to three dependent variables [ln(AUC), ln(t1/2) and ln(Cmax)] and 14 independent variables were built respectively according to the data of the training set. To obtain a high quality of QSPkR models, genetic algorithms (GA) and partial least squares analysis (PLS) were used in calculation. The calculation was conducted with the QSAR module of Cerius2 (Accelrys Software Inc.) molecular modeling software.
We selected three and four independent variables to search their best models. QSPkR analysis based on GA began with a population of random models. These models were generated by randomly selecting three or four features from the data file. Product of multiple linear regression coefficient and leave-one-out cross-validation coefficient was used as a fitness function to generate the fitness scores of these models. For this data set 200 populations were used, and the number of elite populations was 100. The genetic operator was applied until the total fitness score of the elite populations could not be improved over a period of 30 crossover operations. The convergence criteria was met after 430 operations for four features and 280 operations for three features.
The parameters like correlation coefficient (R), variance ratio (F), lack of fit (LOF) scores and leave-one-out cross-validation coefficient (S) were also computed for the suitability of fitness.
The data of the left test set were then predicted by these models.
The descriptors were calculated for substituents R1 and R7 by HyperChem 7.0. And their values are displayed in Tables 3 and 4.
| No. | R7 | ||||||
| SA7 | V7 | HE7 | LP7 | RF7 | P7 | MW7 | |
| 1 | 271.21 | 398.4 | 5.48 | -0.36 | 27.44 | 11.56 | 99.16 |
| 2 | 299.04 | 445.86 | 5.04 | -0.15 | 31.33 | 13.39 | 113.18 |
| 3 | 247.54 | 347.95 | 1.46 | -0.72 | 22.15 | 9.72 | 85.13 |
| 4 | 248.59 | 344.96 | 0.9 | -1 | 21.91 | 9.72 | 85.13 |
| 5 | 266.41 | 382.26 | 5.33 | -0.36 | 27.44 | 11.56 | 99.16 |
| 6 | 251.07 | 350.09 | 1.58 | -0.72 | 22.15 | 9.72 | 85.13 |
| 7 | 265.94 | 383.08 | 3.5 | -0.31 | 26.57 | 11.56 | 99.16 |
| 8 | 300 | 442.55 | -2.21 | -0.28 | 35.48 | 14.96 | 142.18 |
| 9 | 261.99 | 381 | 3.58 | -0.31 | 26.57 | 11.56 | 99.16 |
| 10 | 264.36 | 391.96 | 5.45 | -0.36 | 27.44 | 11.56 | 99.16 |
| 11 | 260.22 | 378.35 | 3.68 | -0.31 | 26.57 | 11.56 | 99.16 |
| 12 | 250.06 | 350.92 | 1.37 | -0.72 | 22.15 | 9.72 | 85.13 |
| 13 | 269.89 | 397.69 | 5.39 | -0.36 | 27.44 | 11.56 | 99.16 |
| 14 | 258.05 | 374.48 | 5.68 | -0.36 | 27.44 | 11.56 | 99.16 |
| 15 | 258.5 | 378.79 | 5.73 | -0.36 | 27.44 | 11.56 | 99.16 |
| 16 | 281.66 | 422.58 | 3 | -0.4 | 28.87 | 12.62 | 111.17 |
| 17 | 298.38 | 442.76 | 5.55 | 0.11 | 30.99 | 13.39 | 113.18 |
| 18 | 266.8 | 385.12 | 3.43 | -0.31 | 26.57 | 11.56 | 99.16 |
| 19 | 244.47 | 346.87 | 4.31 | -1.24 | 24.6 | 10.78 | 97.14 |
| 20 | 260.91 | 378.28 | 5.31 | -0.36 | 27.44 | 11.56 | 99.16 |
| 21 | 264 | 387.29 | 5.61 | -0.36 | 27.44 | 11.56 | 99.16 |
| 22 | 231.53 | 320.13 | 2.75 | -1 | 21.91 | 9.72 | 85.13 |
| No. | R1 | ||||||
| SA1 | V1 | HE1 | LP1 | RF1 | P1 | MW1 | |
| 1 | 162.65 | 191.65 | -4.39 | -0.41 | 6.5 | 3.64 | 30.05 |
| 2 | 184.12 | 230.99 | 2.6 | 1.13 | 10.1 | 5.41 | 41.07 |
| 3 | 183.13 | 229.07 | 2.59 | 1.13 | 10.1 | 5.41 | 41.07 |
| 4 | 181.83 | 225.71 | 2.61 | 1.13 | 10.1 | 5.41 | 41.07 |
| 5 | 232.61 | 320.17 | -3.57 | 1.43 | 24.71 | 9.18 | 96.08 |
| 6 | 171.57 | 206.82 | 0.73 | 1.32 | 7.29 | 4.35 | 29.06 |
| 7 | 192.14 | 241.82 | 2.56 | 1.13 | 10.1 | 5.41 | 41.07 |
| 8 | 180.91 | 225.56 | 2.61 | 1.13 | 10.1 | 5.41 | 41.07 |
| 9 | 185.87 | 233.6 | 2.57 | 1.13 | 10.1 | 5.41 | 41.07 |
| 10 | 209.76 | 274.58 | 0.58 | 2.4 | 13.05 | 6.37 | 58.08 |
| 11 | 173.79 | 211.56 | 0.71 | 1.32 | 7.29 | 4.35 | 29.06 |
| 12 | 172.99 | 213.28 | 0.7 | 1.32 | 7.29 | 4.35 | 29.06 |
| 13 | 213.93 | 278.93 | 0.71 | 2.4 | 13.05 | 6.37 | 58.08 |
| 14 | 172.31 | 210.04 | 0.72 | 1.32 | 7.29 | 4.35 | 29.06 |
| 15 | 187.74 | 241.74 | -1.24 | 0.8 | 15.93 | 7.69 | 60.11 |
| 16 | 194.84 | 249.45 | 2.6 | 0.82 | 9.92 | 5.32 | 59.06 |
| 17 | 193.09 | 243.16 | 2.55 | 1.13 | 10.1 | 5.41 | 41.07 |
| 18 | 246.11 | 343.09 | -3.4 | 2.14 | 26.43 | 9.8 | 113.09 |
| 19 | 246.34 | 343.09 | -3.4 | 2.14 | 26.43 | 9.8 | 113.09 |
| 20 | 239.88 | 334.78 | -2.47 | 2 | 26.21 | 9.89 | 95.1 |
| 21 | 168.2 | 205.76 | 0.8 | 0.92 | 7.37 | 4.26 | 47.05 |
| 22 | 248.12 | 343.14 | -3.4 | 2.14 | 26.43 | 9.8 | 113.09 |
The GA calculation gave 100 models for each pharmacokinetic parameter. The models with the best fitness are listed in Table 5. Results showed that GA was a powerful tool to find the best models. Maximum R2 of models based on ln(Cmax) was only 0.327. Therefore, these models might not be significant. That is to say these 14 descriptors were not correlated with Cmax.
| No. | Model | R2 | R | F | S | LOF |
| AUC | ln(AUC) = 2.27895 + 1.22614 (HE7) + 9.96141 (P7) - 20.5953 (V7) + 9.13637 (SA7) | 0.737 | 0.858 | 9.801 | 0.550 | 0.472 |
| t1/2 | ln(t1/2) = 1.49842 + 1.80503 (P7) + 0.492241 (HE7) - 5.26324 (V1) + 3.53476 (P1) | 0.729 | 0.854 | 9.400 | 0.555 | 0.280 |
| Cmax | ln(Cmax) = 2.96161 - 5.92537 (V1) + 2.15698 (MW1) + 0.206369 (P7) + 0.26873 (HE7) | 0.327 | 0.572 | 1.697 | -0.093 | 0.523 |
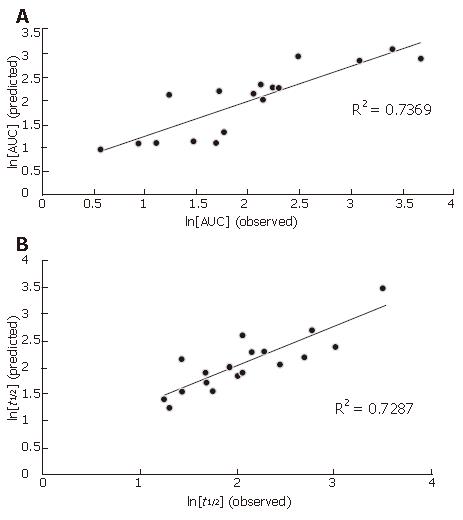
All the predicted and observed data of ln(AUC) and ln(t1/2) from the training set are displayed in Figure 4.
We normalized the data of all the descriptors before model construction, making the coefficient of all the descriptors comparable in the same model. In the model based on AUC, coefficient of V7 was the largest and negative, and that of HE7 was quite small, suggesting that the substituents at position 7 are very significant to AUC, and small volume, large polarizability and large surface area substituents at C-7 are preferred, while hydration energy has little influence on AUC.
In fact, compounds with relatively small volume and large polarizability and surface area of substituents at C-7 (Table 4) all had relatively large AUC (Table 1). Although compounds 3, 4, 12 and 22 (Table 4) had substituents at C-7 with very small volume, their AUCs were all small(Table 1) because of extremely small polarizability and surface area (Table 4), suggesting that coefficient of V7 is not the definitive factor to affect AUC. Volume, polarizability and surface area of R7 determined AUC, and small volume, large polarizability and large surface area of substituents at C-7 were of benefit to large AUC. It is coincident with the results of coefficients in the AUC-based model.
In the t1/2-based model, coefficient of V1 was the largest and negative, but that of P7 and HE7 was quite small, suggesting that the substituents at position 1 are significant to t1/2, large polarizability and small volume of substituents at N-1 are therefore preferred.
In fact, compounds with relatively small volume and large polarizability of substituents at N-1 (Table 4) all had relatively large t1/2 (Table 1). Compounds 1, 6, 11 and 12 with very small volume of substituents at N-1(Table 4) had small t1/2 (Table 1) because of extremely small polarizability (Table 4), and compounds 18, 19 and 22 with extremely large polarizability of substituents at N-1 (Table 4) had relatively small t1/2 (Table 1) because of too large volumes. Therefore, volume and polarizability of R1 determine t1/2 and small volume and large polarizability of substituents are beneficial to large t1/2. It is coincident with the coefficients in the t1/2-based model.
The AUC and t1/2 data of test set (Table 6) were predicted by models displayed in Table 5.
| Compounds | Observed | Predicted | ||
| AUC | t1/2 | AUC | t1/2 | |
| 20 | 27.8 | 25.7 | 17.431 | 16.395 |
| 21 | 18.13 | 11.02 | 13.269 | 9.388 |
| 22 | 2.62 | 4.02 | 12.034 | 6.852 |
The ln(AUC) values predicted by the model correlated well with the observed ln(AUC) values for the training data set with correlation coefficient (R2) equal to 0.7369 (Figure 4A). In addition, application of the model to an external test data set consisting of 3 compounds demonstrated that the model-predicted AUC values were approximate to the observed AUC values (Table 6), indicating that the constructed model is valid for AUC.
The ln(t1/2) values predicted by the model also correlated well with the observed ln(t1/2) values for the training data set with correlation coefficient (R2) equal to 0.7287 (Figure 4B). In addition, the model-predicted t1/2 values were approximate to the observed t1/2 values (Table 6), indicating that the constructed model is also valid for t1/2.
These models may be used to predict the pharmacokinetic parameters (AUC and t1/2) of untried fluoroquinolones. But residual values between predicted and observed data of the test set are slightly larger especially for AUC. It is mainly due to non-precise pharmacokinetic data. Although all the pharmacokinetic data obtained from the literature were averaged, they were not precise enough to get excellent models. The other reason is that we only considered diversities within R1 and R7 to simplify the models. These models however, are very useful as in-silicon prefilters of fluoroquinolone compounds in virtual high throughput screening. And qualitative analysis of substituents at N-1 and C-7 may contribute to guide design of novel fluoroquinolones with excellent pharmacokinetic properties
In conclusion, this model can contribute to a series of fluoroquinolone antibacterial drugs with excellent pharmacokinetic properties for complete eradication of H pylori.
We thank Shandong University (Shandong, China) for allowing us to use the Cerius2 molecular modeling software (Accelrys) and Hypercube Inc. for providing us HyperChem 7.0 Evaluation for Windows (download from http://www.hyper.com/).
Successful drugs must have suitable properties in toxicity, bioavailability and pharmacokinetic parameters. Screening for a large number of compounds with excellent absorption, distribution, metabolism, and excretion (ADME) properties is time-consuming and expensive. So the extension of the idea of quantitative structure-activity relationship (QSAR) to pharmacokinetic data has led to emergence of new tool called quantitative structure pharmacokinetic relationship (QSPkR) study. QSPkR study can be utilized in drug design.
Both one- and two-dimensional topological indices have been used extensively to numerically relate molecular structure with activity and/or property. (These descriptors rely only on the molecular graph for their calculation. In contrast, three-dimensional descriptors require the absolute conformation of a molecule. They, too, have been successfully used to develop QSPkRs.
In this study the authors have developed and demonstrated novel computational approaches for the efficient and accurate prediction of AUC and t1/2 of fluoroquinolones. They constructed simple models which can directly correlate physical and chemical properties to pharmacokinetic data. These models can be used not only to predict pharmacokinetic parameters but also to guide the design of novel fluoroquinolones.
Using these QSPkR models, the authors can illustrate how the changes at N-1 and C-7 of the fluoroquinolones affect their pharmacokinetic parameters. Such computational models may be useful as in-silico prefilters of fluoroquinolones compounds in a virtual high throughput screening environment and as a research tool for identifying and improving the pharmacokinetic profiles of fluoroquinolones candidates.
In the present study, the authors have tried to develop computational approaches for the prediction of the pharmacokinetics of fluoroquinolones. Quantitative structure-pharmacokinetics relationship analysis can be an important tool at the early stage of drug design. The authors demonstrated that small volume and large polarizability of substitutents of R-1 are beneficial to large t1/2 and small volume, large polarizability and surface area of substitutents at C-7 are of benefit to large AUC in fluoroquinolones.
S- Editor Liu Y L- Editor Ma JY E- Editor Wang HF
| 1. | NIH Consensus Conference. Helicobacter pylori in peptic ulcer disease. NIH Consensus Development Panel on Helicobacter pylori in Peptic Ulcer Disease. JAMA. 1994;272:65-69. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 524] [Cited by in RCA: 483] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
| 2. | Leung WK, Hung LC, Kwok CK, Leong RW, Ng DK, Sung JJ. Follow up of serial urea breath test results in patients after consumption of antibiotics for non-gastric infections. World J Gastroenterol. 2002;8:703-706. [PubMed] |
| 3. | Ibrahim M, Khan AA, Tiwari SK, Habeeb MA, Khaja MN, Habibullah CM. Antimicrobial activity of Sapindus mukorossi and Rheum emodi extracts against H pylori: In vitro and in vivo studies. World J Gastroenterol. 2006;12:7136-7142. [PubMed] |
| 4. | Klopman G, Fercu D, Renau TE, Jacobs MR. N-1-tert-butyl-substituted quinolones: in vitro anti-Mycobacterium avium activities and structure-activity relationship studies. Antimicrob Agents Chemother. 1996;40:2637-2643. [PubMed] |
| 5. | Norris DA, Leesman GD, Sinko PJ, Grass GM. Development of predictive pharmacokinetic simulation models for drug discovery. J Control Release. 2000;65:55-62. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 71] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 6. | Ghafourian T, Fooladi S. The effect of structural QSAR parameters on skin penetration. Int J Pharm. 2001;217:1-11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 19] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 7. | Feher M, Sourial E, Schmidt JM. A simple model for the prediction of blood-brain partitioning. Int J Pharm. 2000;201:239-247. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 76] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 8. | Wajima T, Fukumura K, Yano Y, Oguma T. Prediction of human pharmacokinetics from animal data and molecular structural parameters using multivariate regression analysis: oral clearance. J Pharm Sci. 2003;92:2427-2440. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 25] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 9. | Cook JA, Silverman MH, Schelling DJ, Nix DE, Schentag JJ, Brown RR, Stroshane RM. Multiple-dose pharmacokinetics and safety of oral amifloxacin in healthy volunteers. Antimicrob Agents Chemother. 1990;34:974-979. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 4] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 10. | Stroshane RM, Silverman MH, Sauerschell R, Brown RR, Boddy AW, Cook JA. Preliminary study of the pharmacokinetics of oral amifloxacin in elderly subjects. Antimicrob Agents Chemother. 1990;34:751-754. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 11. | Kozawa O, Uematsu T, Matsuno H, Niwa M, Nagashima S, Kanamaru M. Comparative study of pharmacokinetics of two new fluoroquinolones, balofloxacin and grepafloxacin, in elderly subjects. Antimicrob Agents Chemother. 1996;40:2824-2828. [PubMed] |
| 12. | Lubasch A, Keller I, Borner K, Koeppe P, Lode H. Comparative pharmacokinetics of ciprofloxacin, gatifloxacin, grepafloxacin, levofloxacin, trovafloxacin, and moxifloxacin after single oral administration in healthy volunteers. Antimicrob Agents Chemother. 2000;44:2600-2603. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 129] [Cited by in RCA: 130] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 13. | Boy D, Well M, Kinzig-Schippers M, Sörgel F, Ankel-Fuchs D, Naber KG. Urinary bactericidal activity, urinary excretion and plasma concentrations of gatifloxacin (400 mg) versus ciprofloxacin (500 mg) in healthy volunteers after a single oral dose. Int J Antimicrob Agents. 2004;23 Suppl 1:S6-16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 25] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 14. | Zhang J, Zhang YY, Shi YG, Yu JC, Wang L. Clinical pharmacokinetics study on ciprofloxacin. Zhongguo Yaolixue Tongbao. 1996;12:430-433. |
| 15. | Xiao YH, Wang QN, Qian YS, Du JZ, Zheng XP, Liu HY. Pharmacokinetics and in vitro antibacterial activity of ciprofloxacin. Zhongguo Kangshengsu Zazhi. 1992;17:293-296. |
| 16. | Randinitis EJ, Koup JR, Rausch G, Abel R, Bron NJ, Hounslow NJ, Vassos AB, Sedman AJ. Clinafloxacin pharmacokinetics in subjects with various degrees of renal function. Antimicrob Agents Chemother. 2001;45:2536-2542. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 8] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 17. | Randinitis EJ, Brodfuehrer JI, Eiseman I, Vassos AB. Pharmacokinetics of clinafloxacin after single and multiple doses. Antimicrob Agents Chemother. 2001;45:2529-2535. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 18. | Wise R, Jones S, Das I, Andrews JM. Pharmacokinetics and inflammatory fluid penetration of clinafloxacin. Antimicrob Agents Chemother. 1998;42:428-430. [PubMed] |
| 19. | Meyerhoff C, Dilger C, Yoon SJ, Chung YH, Lee DK, Lee CW, Ryu JM, Choi MS, Pabst G, Reh C. Safety, tolerability and pharmacokinetics of the new long-acting quinolone DW-116 after single and multiple dosing in healthy subjects. J Antimicrob Chemother. 1998;42:349-361. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 20. | Qian YS, Wang QN, Xiao YH, Du JZ, Zheng XP, Li CZ, Liu HY, Qiu J, Jiang FY. Pharmacokinetics study of enoxacin in human and in vitro antibactial activity. Zhongguo Kangshengsu Zazhi. 1992;17:215-218. |
| 21. | Zeng JZ, Liang DR, Qin YP, Liang MZ, Liu DY, Huang Y, Zhang HM, Yu Q, Zou YG. Zhang HL. Study on the pharmacokinetics of oral enoxacin and bioavailabilities of preparations in healthy volunteers. Zhongguo Kangshengsu Zazhi. 1993;18:341-346. |
| 22. | Xiao YH, Wang QN, Qian YS, Du JZ, He HX. Pharmacokinetics of enoxacin, ciprofloxacin, ofloxacin and pefloxacin. Chongqing Yikedaxue Xuebao. 1994;19:177-179. |
| 23. | Trampuz A, Laifer G, Wenk M, Rajacic Z, Zimmerli W. Pharmacokinetics and pharmacodynamics of gatifloxacin against Streptococcus pneumoniae and Staphylococcus aureus in a granulocyte-rich exudate. Antimicrob Agents Chemother. 2002;46:3630-3633. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 8] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 24. | Lober S, Ziege S, Rau M, Schreiber G, Mignot A, Koeppe P, Lode H. Pharmacokinetics of gatifloxacin and interaction with an antacid containing aluminum and magnesium. Antimicrob Agents Chemother. 1999;43:1067-1071. [PubMed] |
| 25. | Liu XD, Xie L, Wang J, Liang Y, Li L, Lu L. Comparison of pharmacokinetics of gatifloxacin in rats, dogs and humans. Asian J drug Metab Pharmacokinet. 2005;5:71-76. |
| 26. | Wise R, Andrews JM, Ashby JP, Marshall J. A study to determine the pharmacokinetics and inflammatory fluid penetration of gatifloxacin following a single oral dose. J Antimicrob Chemother. 1999;44:701-704. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 20] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 27. | Allen A, Bygate E, Oliver S, Johnson M, Ward C, Cheon AJ, Choo YS, Kim IC. Pharmacokinetics and tolerability of gemifloxacin (SB-265805) after administration of single oral doses to healthy volunteers. Antimicrob Agents Chemother. 2000;44:1604-1608. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 50] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 28. | Gee T, Andrews JM, Ashby JP, Marshall G, Wise R. Pharmacokinetics and tissue penetration of gemifloxacin following a single oral dose. J Antimicrob Chemother. 2001;47:431-434. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 25] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 29. | Allen A, Bygate E, Vousden M, Oliver S, Johnson M, Ward C, Cheon A, Choo YS, Kim I. Multiple-dose pharmacokinetics and tolerability of gemifloxacin administered orally to healthy volunteers. Antimicrob Agents Chemother. 2001;45:540-545. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 25] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 30. | Child J, Andrews JM, Wise R. Pharmacokinetics and tissue penetration of the new fluoroquinolone grepafloxacin. Antimicrob Agents Chemother. 1995;39:513-515. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 31] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 31. | Efthymiopoulos C. Pharmacokinetics of grepafloxacin. J Antimicrob Chemother. 1997;40 Suppl A:35-43. [PubMed] |
| 32. | Chien SC, Rogge MC, Gisclon LG, Curtin C, Wong F, Natarajan J, Williams RR, Fowler CL, Cheung WK, Chow AT. Pharmacokinetic profile of levofloxacin following once-daily 500-milligram oral or intravenous doses. Antimicrob Agents Chemother. 1997;41:2256-2260. [PubMed] |
| 33. | Chien SC, Chow AT, Natarajan J, Williams RR, Wong FA, Rogge MC, Nayak RK. Absence of age and gender effects on the pharmacokinetics of a single 500-milligram oral dose of levofloxacin in healthy subjects. Antimicrob Agents Chemother. 1997;41:1562-1565. [PubMed] |
| 34. | Stone JW, Andrews JM, Ashby JP, Griggs D, Wise R. Pharmacokinetics and tissue penetration of orally administered lomefloxacin. Antimicrob Agents Chemother. 1988;32:1508-1510. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 50] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 35. | Kovarik JM, Hoepelman AI, Smit JM, Sips PA, Rozenberg-Arska M, Glerum JH, Verhoef J. Steady-state pharmacokinetics and sputum penetration of lomefloxacin in patients with chronic obstructive pulmonary disease and acute respiratory tract infections. Antimicrob Agents Chemother. 1992;36:2458-2461. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 3] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 36. | Tan L, Xu DK, Diao Y, Yuan YS. Highperformance liquid chromatographic assay for lomefloxacin in plasma and its pharmacokinetics in healthy volunteers. YaoXue XueBao. 1993;28:286-289. [PubMed] |
| 37. | Tang X, Cai YM. Effects of probenicid on the pharmacokinetics of norfloxacin. Shenyang Yaoxueyuan Xuebao. 1991;8:1-4. |
| 38. | Adhami ZN, Wise R, Weston D, Crump B. The pharmacokinetics and tissue penetration of norfloxacin. J Antimicrob Chemother. 1984;13:87-92. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 58] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 39. | Lehto P, Kivistö KT. Effect of sucralfate on absorption of norfloxacin and ofloxacin. Antimicrob Agents Chemother. 1994;38:248-251. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 40] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 40. | Yuk JH, Nightingale CH, Quintiliani R, Sweeney KR. Bioavailability and pharmacokinetics of ofloxacin in healthy volunteers. Antimicrob Agents Chemother. 1991;35:384-386. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 35] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 41. | Immanuel C, Hemanthkumar AK, Gurumurthy P, Venkatesan P. Dose related pharmacokinetics of ofloxacin in healthy volunteers. Int J Tuberc Lung Dis. 2002;6:1017-1022. [PubMed] |
| 42. | Hameed A, Iqbal T, Nawaz M, Batool F, Baig TT, Ali S, Nurjis F. Pharmacokinetics of ofloxacin in male volunteers following oral administration. Pak J Biol Sci. 2002;5:1098-1100. [DOI] [Full Text] |
| 43. | Israel D, Gillum JG, Turik M, Harvey K, Ford J, Dalton H, Towle M, Echols R, Heller AH, Polk R. Pharmacokinetics and serum bactericidal titers of ciprofloxacin and ofloxacin following multiple oral doses in healthy volunteers. Antimicrob Agents Chemother. 1993;37:2193-2199. [RCA] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 28] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 44. | Lode H, Höffken G, Olschewski P, Sievers B, Kirch A, Borner K, Koeppe P. Pharmacokinetics of ofloxacin after parenteral and oral administration. Antimicrob Agents Chemother. 1987;31:1338-1342. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 111] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 45. | De Lepeleire I, Van Hecken A, Verbesselt R, Tjandra-Maga TB, De Schepper PJ. Comparative oral pharmacokinetics of fleroxacin and pefloxacin. J Antimicrob Chemother. 1988;22:197-202. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 46. | Wang WL, Lu H, Wang T, Hou F, Li JT. Pharmacokinetics of pefloxacin in normal volunteers. Zhongguo Linchuang Yaolixue Zazhi. 1990;6:235-241. |
| 47. | Xiao YH, Wang QN, Qian YS, Du JZ, Zheng XP, Liu HY. Pharmacokinetics and in vitro antibacterial activity of pefloxacin. Zhongguo Kangshengsu Zazhi. 1992;17:302-305. |
| 48. | Imbimbo BP, Broccali G, Cesana M, Crema F, Attardo-Parrinello G. Inter- and intrasubject variabilities in the pharmacokinetics of rufloxacin after single oral administration to healthy volunteers. Antimicrob Agents Chemother. 1991;35:390-393. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 36] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 49. | Wise R, Johnson J, O'Sullivan N, Andrews JM, Imbimbo BP. Pharmacokinetics and tissue penetration of rufloxacin, a long acting quinolone antimicrobial agent. J Antimicrob Chemother. 1991;28:905-909. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 27] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 50. | Perry G, Mant TG, Morrison PJ, Sacks S, Woodcook J, Wise R, Imbimbo BP. Pharmacokinetics of rufloxacin in patients with impaired renal function. Antimicrob Agents Chemother. 1993;37:637-641. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 51. | Kisicki JC, Griess RS, Ott CL, Cohen GM, McCormack RJ, Troetel WM, Imbimbo BP. Multiple-dose pharmacokinetics and safety of rufloxacin in normal volunteers. Antimicrob Agents Chemother. 1992;36:1296-1301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 24] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 52. | O'Grady J, Briggs A, Atarashi S, Kobayashi H, Smith RL, Ward J, Ward C, Milatovic D. Pharmacokinetics and absolute bioavailability of sitafloxacin, a new fluoroquinolone antibiotic, in healthy male and female Caucasian subjects. Xenobiotica. 2001;31:811-822. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 24] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 53. | Nakashima M, Uematsu T, Kosuge K, Umemura K, Hakusui H, Tanaka M. Pharmacokinetics and tolerance of DU-6859a, a new fluoroquinolone, after single and multiple oral doses in healthy volunteers. Antimicrob Agents Chemother. 1995;39:170-174. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 54. | Johnson JH, Cooper MA, Andrews JM, Wise R. Pharmacokinetics and inflammatory fluid penetration of sparfloxacin. Antimicrob Agents Chemother. 1992;36:2444-2446. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 40] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 55. | Zix JA, Geerdes-Fenge HF, Rau M, Vöckler J, Borner K, Koeppe P, Lode H. Pharmacokinetics of sparfloxacin and interaction with cisapride and sucralfate. Antimicrob Agents Chemother. 1997;41:1668-1672. [PubMed] |
| 56. | Mody VD, Pandya KK, Satia MC, Modi IA, Modi RI, Gandhi TP. High performance thin-layer chromatographic method for the determination of sparfloxacin in human plasma and its use in pharmacokinetic studies. J Pharm Biomed Anal. 1998;16:1289-1294. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 21] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 57. | Matsuoka M, Banno K, Sato T. Analytical chiral separation of a new quinolone compound in biological fluids by high-performance liquid chromatography. J Chromatogr B Biomed Appl. 1996;676:117-124. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 15] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 58. | Granneman GR, Carpentier P, Morrison PJ, Pernet AG. Pharmacokinetics of temafloxacin in humans after single oral doses. Antimicrob Agents Chemother. 1991;35:436-441. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 43] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 59. | Granneman GR, Braeckman R, Kraut J, Shupien S, Craft JC. Temafloxacin pharmacokinetics in subjects with normal and impaired renal function. Antimicrob Agents Chemother. 1991;35:2345-2351. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 9] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 60. | Teng R, Liston TE, Harris SC. Multiple-dose pharmacokinetics and safety of trovafloxacin in healthy volunteers. J Antimicrob Chemother. 1996;37:955-963. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 54] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 61. | Vincent J, Venitz J, Teng R, Baris BA, Willavize SA, Polzer RJ, Friedman HL. Pharmacokinetics and safety of trovafloxacin in healthy male volunteers following administration of single intravenous doses of the prodrug, alatrofloxacin. J Antimicrob Chemother. 1997;39 Suppl B:75-80. [PubMed] |
| 62. | Wise R, Mortiboy D, Child J, Andrews JM. Pharmacokinetics and penetration into inflammatory fluid of trovafloxacin (CP-99,219). Antimicrob Agents Chemother. 1996;40:47-49. [PubMed] |
| 63. | Granneman GR, Snyder KM, Shu VS. Difloxacin metabolism and pharmacokinetics in humans after single oral doses. Antimicrob Agents Chemother. 1986;30:689-693. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 43] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 64. | Stass H, Dalhoff A, Kubitza D, Schühly U. Pharmacokinetics, safety, and tolerability of ascending single doses of moxifloxacin, a new 8-methoxy quinolone, administered to healthy subjects. Antimicrob Agents Chemother. 1998;42:2060-2065. [PubMed] |
| 65. | Siefert HM, Domdey-Bette A, Henninger K, Hucke F, Kohlsdorfer C, Stass HH. Pharmacokinetics of the 8-methoxyquinolone, moxifloxacin: a comparison in humans and other mammalian species. J Antimicrob Chemother. 1999;43 Suppl B:69-76. [PubMed] |
| 66. | Minami R, Inotsume N, Nakamura C, Nakano M. Stereoselective analysis of the disposition of tosufloxacin enantiomers in man. Eur J Clin Pharmacol. 1993;45:489-491. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 67. | Niki Y. Pharmacokinetics and safety assessment of tosufloxacin tosilate. J Infect Chemother. 2002;8:1-18. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 17] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 68. | Minami R, Nakamura C, Inotsume N, Nakano M. Effects of aluminum hydroxide and famotidine on bioavailability of tosufloxacin in healthy volunteers. Antimicrob Agents Chemother. 1998;42:453-455. [PubMed] |