Copyright
©The Author(s) 2015.
World J Nephrol. May 6, 2015; 4(2): 295-306
Published online May 6, 2015. doi: 10.5527/wjn.v4.i2.295
Published online May 6, 2015. doi: 10.5527/wjn.v4.i2.295
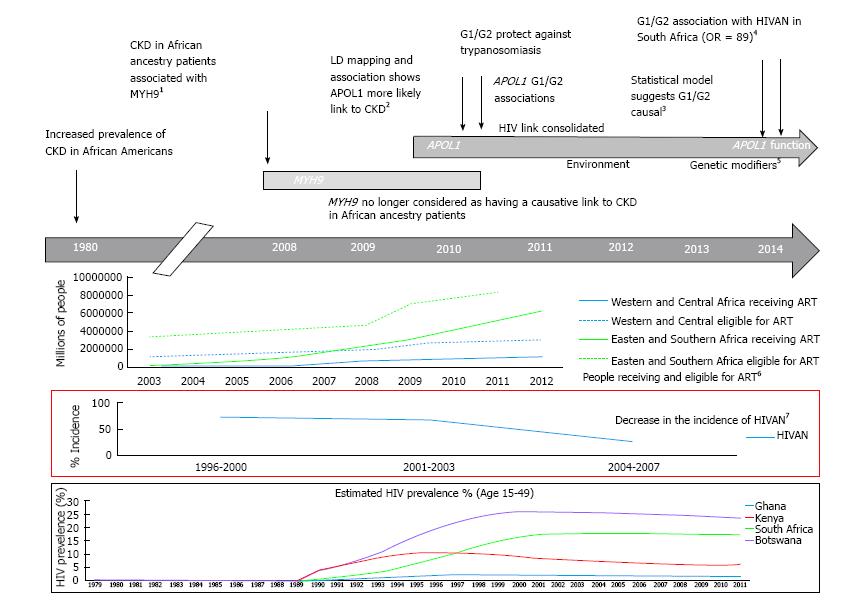
Figure 1 Historical timeline reflecting the discovery of genetic association to chronic kidney disease in populations with African ancestry.
1Adapted from Kopp et al[31] and Kao et al[30]; 2Adapted from Freedman et al[49], Genovese et al[13], Tzur et al[29]; 3Adapted from Genovese et al[84]; 4Adapted from Kasembeli et al (2014 unpublished observations); 5Adapted from Freedman et al[64]; 6Adapted from UNAIDS report on global AIDS epidemic[18]; 7Adapted from USRDS 2012 Annual Data Report. APOL1: Apolipoprotein L1; MYH9: Non-muscle myosin heavy chain 9; ART: Antiretroviral therapy; CKD: Chronic kidney disease.
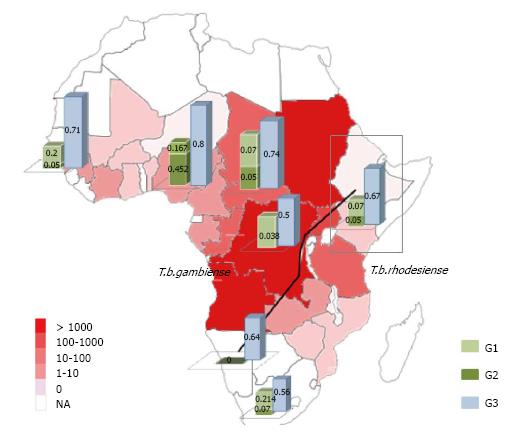
Figure 2 Distribution of Human African Trypanosomiasis (T.
b gambiense and T.b rhodesiense), MYH9 E1 and APOL1 G1 and G2 risk alleles in Africa[12,42]. The frequency of distribution of APOL1 risk variants in Africa are represented by bar charts and overlap the areas distribution of Human African Trypanosomiasis. The numbers reflect the reported cases of Trypanosomiasis from the WHO, 2010. T.b: Trypanosoma brucei.
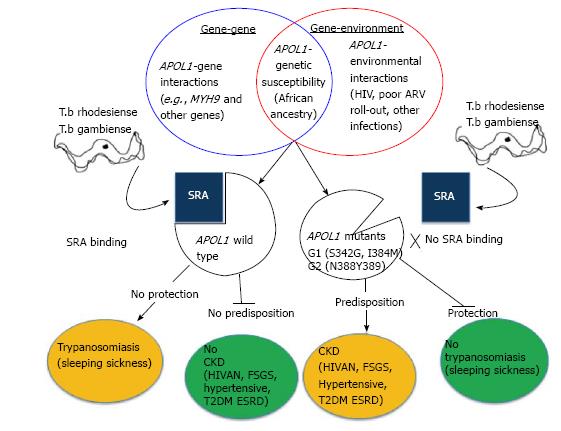
Figure 3 Gene-Gene, Gene-Environment steering contribution to APOL1 associated CKD and the positive selection of APOL1 associated CKD variants as a result of Trypanosomiasis.
SRA: Serum resistant associated protein; HIV: Human immunodeficiency virus; T.b: Trypanosoma brucei; APOL1: Apolipoprotein L1; MYH9: Non-muscle myosin heavy chain 9; HIVAN: Human immunodeficiency virus-associated nephropathy; FSGS: Focal segmental glomerulosclerosis; T2DM: Type 2 diabetes mellitus; ESRD: End stage renal disease; CKD: Chronic kidney disease.
- Citation: Kasembeli AN, Duarte R, Ramsay M, Naicker S. African origins and chronic kidney disease susceptibility in the human immunodeficiency virus era. World J Nephrol 2015; 4(2): 295-306
- URL: https://www.wjgnet.com/2220-6124/full/v4/i2/295.htm
- DOI: https://dx.doi.org/10.5527/wjn.v4.i2.295