Published online Mar 15, 2021. doi: 10.4239/wjd.v12.i3.261
Peer-review started: July 27, 2020
First decision: November 4, 2020
Revised: November 10, 2020
Accepted: December 23, 2020
Article in press: December 23, 2020
Published online: March 15, 2021
Processing time: 217 Days and 23.1 Hours
The causality between education and type 2 diabetes (T2DM) remains unclear.
To identify the causality between education and T2DM and the potential metabolic risk factors [coronary heart disease (CHD), total cholesterol, low-density lipoprotein, triglycerides (TG), body mass index (BMI), waist circumference (WC), waist-to-hip ratio (WHR), fasting insulin, fasting glucose, and glycated hemoglobin] from summarized genome-wide association study (GWAS) data used a network Mendelian randomization (MR).
Two-sample MR and network MR were performed to obtain the causality between education-T2DM, education-mediator, and mediator-T2DM. Summary statistics from the Social Science Genetic Association Consortium (discovery data) and Neale Lab consortium (replication data) were used for education and DIAGRAMplusMetabochip for T2DM.
The odds ratio for T2DM was 0.392 (95%CI: 0.263-0.583) per standard deviation increase (3.6 years) in education by the inverse variance weighted method, without heterogeneity or horizontal pleiotropy. Education was genetically associated with CHD, TG, BMI, WC, and WHR in the discovery phase, yet only the results for CHD, BMI, and WC were replicated in the replication data. Moreover, BMI was genetically associated with T2DM.
Short education was found to be associated with an increased T2DM risk. BMI might serve as a potential mediator between them.
Core Tip: Genetically predicted education was negatively causally associated with type 2 diabetes (T2DM). The odds ratio for T2DM was 0.392 (95%CI: 0.263-0.583) per standard deviation increase (3.6 years) in education. Body mass index might serve as a potential mediator.
- Citation: Liao LZ, Chen ZC, Li WD, Zhuang XD, Liao XX. Causal effect of education on type 2 diabetes: A network Mendelian randomization study. World J Diabetes 2021; 12(3): 261-277
- URL: https://www.wjgnet.com/1948-9358/full/v12/i3/261.htm
- DOI: https://dx.doi.org/10.4239/wjd.v12.i3.261
Type 2 diabetes mellitus (T2DM), affecting approximately 415 million people, has become a major global health problem. This number will rapidly increase to 642 million by 2040[1]. Environmental, genetic, and metabolic risk factors all contribute to T2DM[2]. Among these, controlling the modifiable risk factors to decrease the T2DM morbidity, turns into a key public health priority[3]. A previous systematic review and meta-analysis reported that diabetes self-management education can reduce all-cause mortality risk in T2DM patients[4], indicating that education may help T2DM management. However, the causality between primary education (years of schooling) and T2DM remains unclear. If the causal association exists, the potential pathways involved in the association from education to T2DM have not yet been studied.
Mendelian randomization (MR) serves as a strategy for assessing the causality between disease and common risk factors[5]. Since the genetically determined risk factors occur before the disease onset, MR can avoid the potential bias of reverse causation in retrospective studies. The risk of confounding is also reduced[5]. In this study, the causality between education and T2DM was analyzed and potential metabolic risk factors were explored from summarized genome-wide association study (GWAS) data. The potential metabolic risk factors included coronary heart disease (CHD), total cholesterol (TC), low-density lipoprotein (LDL), triglycerides (TG), body mass index (BMI), waist circumference (WC), waist-to-hip ratio (WHR), fasting insulin, fasting glucose, and glycated hemoglobin (HbA1c).
Summary data from array-based analysis for single nucleotide polymorphism (SNP) was included. We selected all SNPs associated with education at genome-wide significance (P < 5 × 10-8) in the available GWAS (Supplementary Table 1). The metrics of SNP quality were requested as follows. There was strong evidence of between-study heterogeneity in the SNP-trait association (P ≤ 0.005). The imputation quality metric (info or r2) was ≤ 0.90, and there was Hardy-Weinberg disequilibrium (P ≤ 0.001). Summary statistics from the Social Science Genetic Association Consortium (SSGAC)[6] and Neale Lab consortium (http://www.nealelab.is/blog/2017/9/11/details-and-considerations-of-the-uk-biobank-gwas) were used for education. Because several large summary statistics from different consortia were available for the same biomarker, we used the earliest and largest summary data (SSGAC) as the discovery data set and the most recent data (Neale Lab) as the replication data set. DIAGRAM plus Metabochip consortium data were used for T2DM[7], CARDIoGRAM plus C4D consortium data were utilized for CHD[8], GLGC consortia data were used for TC[9], LDL[9], and TG[9], Genetic Investigation of Anthropometric Traits consortium data were used for BMI[10], WC[11], and WHR[12], and Mapping and Geographic Information Consortium data were used for fasting insulin[13], fasting glucose[14], and HbA1c[15]. The datasets are presented in Table 1.
| Exposure/Outcome | Cases, n | Controls, n | Sample size | PubMed ID | First author | Consortium | Year | Units |
| Education (discovery data) | NA | NA | 293723 | 27225129 | Okbay | SSGAC | 2016 | SD (yr) |
| Education 2 (replication data) | NA | NA | 226899 | UKB-a:505 | Neale | Neale Lab | 2017 | NA |
| T2DM | 34840 | 114981 | 77312 | 22885922 | Morris | DIAGRAM plus Metabochip | 2012 | Log odds |
| CHD | 60801 | 123504 | 184305 | 26343387 | Nikpay | CARDIoGRAM plus C4D | 2015 | Log odds |
| TG | NA | NA | 108514 | 24097068 | Willer | GLGC | 2013 | SD (mg/dL) |
| LDL | NA | NA | 99073 | 24097068 | Willer | GLGC | 2013 | SD (mg/dL) |
| TC | NA | NA | 108363 | 24097068 | Willer | GLGC | 2013 | SD (mg/dL) |
| BMI | NA | NA | 152893 | 25673413 | Locke | GIANT | 2015 | SD (kg/m2) |
| WC | NA | NA | 232101 | 25673412 | Shungin | GIANT | 2015 | SD (cm) |
| WHR | NA | NA | 60586 | 23754948 | Randall | GIANT | 2013 | SD (ratio) |
| Fasting insulin | NA | NA | 38238 | 20081858 | Dupuis | MAGIC | 2010 | Log pmol/L |
| Fasting glucose | NA | NA | 58074 | 22581228 | Manning | MAGIC | 2012 | mmol/L |
| HbA1c | NA | NA | 46368 | 20858683 | Soranzo | MAGIC | 2010 | % |
Our MR study was conducted on the MR-Base platform online (http://www.mrbase.org)[16]. We explored the associations as follows: (1) Causality: The conventional MR approach [inverse variance weighted (IVW)] method, MR Egger method, and weighted median method were used: (a) Causality between genetically determined education and T2DM; (b) Causality between education and metabolic risk factors (CHD, TG, LDL, TC, BMI, WC, WHR, fasting insulin, fasting glucose, and HbA1c); and (c) Causality between metabolic risk factors and T2DM; (2) Heterogeneity: We conducted heterogeneity tests in MR analyses using IVW and MR Egger; (3) Horizontal pleiotropy: The MR egger intercept was assessed; (4) Leave-one-out analysis; and (5) Funnel plots.
Two-sample MR and network MR were performed to obtain the causality between education-T2DM, education-mediator, and mediator-T2DM[18]. A network MR analysis consisted of three two-sample MR tests: (1) The causality between education and T2DM; (2) The causality between education and the potential mediators; and (3) The causality between the potential mediators and T2DM.
We could conclude that the specific metabolic risk factor might serve as a mediator between education and T2DM if the causality was estimated in all three steps.
The statistical tests were two-sided. The statistical test for the MR analyses was considered statistically significant at P < 0.05. All of the calculations were conducted using Stata (College Station, TX, United States) and R language.
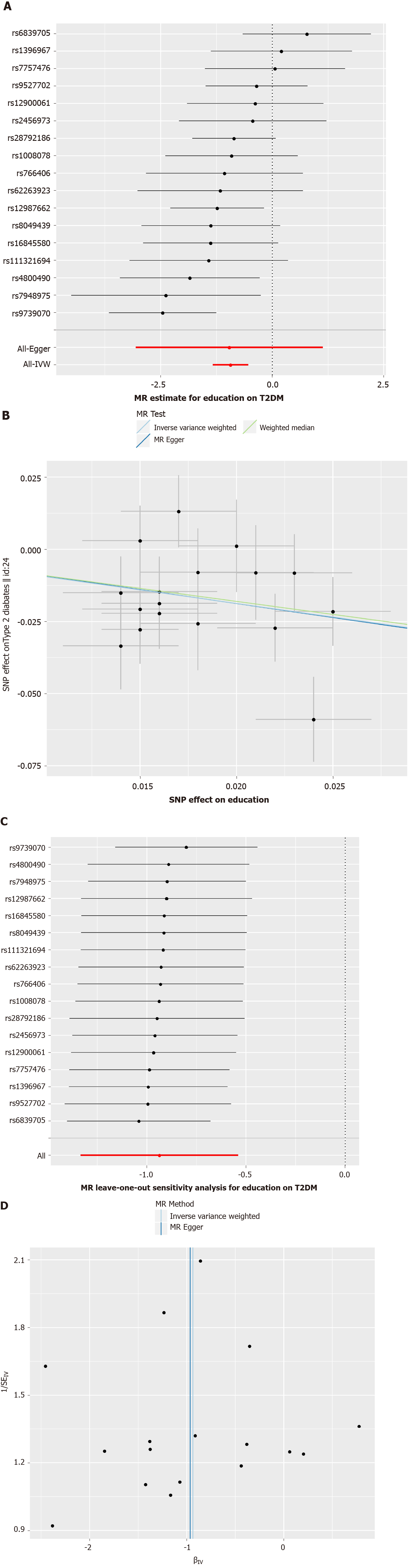
Characteristics of the SNPs are shown in Supplementary Table 1. We used the SSGAC consortia for education to explore the causal associations between education and T2DM. In the IVW method, the odds ratio [OR (95%CI)] for T2DM was 0.392 (0.263-0.583) per standard deviation increase (3.6 years) in education (Table 2 and Figure 1A). Results were consistent in weighted median method (OR: 0.406, 95%CI: 0.246-0.672; P = 0.000) (Table 2). Both IVW and MR Egger estimates indicated no heterogeneity amongst these 17 SNPs (P = 0.163 and P = 0.124, respectively) (Table 2). There was no directional horizontal pleiotropy (MR egger intercept P = 0.979) (Table 2). In a leave-one-out analysis, no single instrument was strongly driving the overall effect of education on T2DM (Figure 1C). Besides, there was no funnel plot asymmetry (Figure 1D). Both the leave-one-out analysis and funnel plot further suggested that no SNPs exhibited horizontal pleiotropy.
| Trait | Method | nSNP | OR | 95%CI | P value | Heterogeneity P | MR Egger intercept P | |
| Education-T2DM | MR Egger | 17 | 0.381 | 0.047 | 3.093 | 0.381 | 0.124 | 0.979 |
| Weighted median | 17 | 0.406 | 0.246 | 0.672 | 0.000 | |||
| Inverse variance weighted | 17 | 0.392 | 0.263 | 0.583 | 0.000 | 0.163 | ||
To sum up, the genetically predicted education was negatively causally associated with T2DM.
The causality between education and metabolic biomarkers, including CHD, TG, LDL, TC, BMI, WC, WHR, fasting insulin, fasting glucose, and HbA1c, is shown in Table 3. Education was causally associated with CHD, TG, BMI, WC, and WHR in the discovery phase. Yet in the replication data, only the results for CHD, BMI, and WC were duplicated.
| Trait | Method | nSNP | Beta | SE | P value | Heterogeneity P | MR Egger intercept P |
| Discovery (education-metabolic risk factors) | |||||||
| CHD | MR Egger | 71 | 0.145 | 0.417 | 0.729 | 0.123 | 0.263 |
| Weighted median | 71 | -0.238 | 0.107 | 0.026 | |||
| Inverse variance weighted | 71 | -0.318 | 0.079 | 0.000 | 0.117 | ||
| TG | MR Egger | 57 | -0.183 | 0.263 | 0.489 | 0.112 | 0.937 |
| Weighted median | 57 | -0.192 | 0.057 | 0.001 | |||
| Inverse variance weighted | 57 | -0.204 | 0.043 | 0.000 | 0.130 | ||
| LDL | MR Egger | 57 | -0.274 | 0.275 | 0.324 | 0.316 | 0.415 |
| Weighted median | 57 | 0.007 | 0.063 | 0.909 | |||
| Inverse variance weighted | 57 | -0.051 | 0.045 | 0.261 | 0.326 | ||
| TC | MR Egger | 57 | -0.087 | 0.277 | 0.755 | 0.197 | 0.838 |
| Weighted median | 57 | -0.018 | 0.064 | 0.775 | |||
| Inverse variance weighted | 57 | -0.031 | 0.045 | 0.500 | 0.223 | ||
| BMI | MR Egger | 59 | 0.013 | 0.353 | 0.970 | 0.000 | 0.415 |
| Weighted median | 59 | -0.209 | 0.048 | 0.000 | |||
| Inverse variance weighted | 59 | -0.272 | 0.058 | 0.000 | 0.000 | ||
| WC | MR Egger | 59 | -0.002 | 0.404 | 0.996 | 0.000 | 0.429 |
| Weighted median | 59 | -0.269 | 0.056 | 0.000 | |||
| Inverse variance weighted | 59 | -0.319 | 0.066 | 0.000 | 0.000 | ||
| WHR | MR Egger | 58 | -0.743 | 0.461 | 0.113 | 0.028 | 0.346 |
| Weighted median | 58 | -0.298 | 0.097 | 0.002 | |||
| Inverse variance weighted | 58 | -0.311 | 0.076 | 0.000 | 0.027 | ||
| Fasting insulin | MR Egger | 58 | -0.269 | 0.215 | 0.215 | 0.010 | 0.312 |
| Weighted median | 58 | -0.039 | 0.048 | 0.419 | |||
| Inverse variance weighted | 58 | -0.053 | 0.035 | 0.129 | 0.014 | ||
| Fasting glucose | MR Egger | 58 | -0.185 | 0.172 | 0.287 | 0.167 | 0.398 |
| Weighted median | 58 | -0.044 | 0.039 | 0.257 | |||
| Inverse variance weighted | 58 | -0.041 | 0.028 | 0.152 | 0.172 | ||
| HbA1c | MR Egger | 58 | 0.004077 | 0.1779 | 0.9818 | 0.5674 | 0.889 |
| Weighted median | 58 | -0.009323 | 0.04094 | 0.8199 | |||
| Inverse variance weighted | 58 | -0.02056 | 0.02908 | 0.4796 | 0.6038 | ||
| Replication (education-metabolic risk factors) | |||||||
| CHD | MR Egger | 20 | -0.985 | 0.772 | 0.218 | 0.196 | 0.537 |
| Weighted median | 20 | -0.474 | 0.209 | 0.024 | |||
| Inverse variance weighted | 20 | -0.511 | 0.164 | 0.002 | 0.222 | ||
| TG | MR Egger | 13 | 0.099 | 1.076 | 0.928 | 0.000 | 0.702 |
| Weighted median | 13 | -0.161 | 0.134 | 0.229 | |||
| Inverse variance weighted | 13 | -0.319 | 0.163 | 0.051 | 0.000 | ||
| BMI | MR Egger | 14 | 0.830 | 0.582 | 0.179 | 0.079 | 0.071 |
| Weighted median | 14 | -0.381 | 0.106 | 0.000 | |||
| Inverse variance weighted | 14 | -0.308 | 0.095 | 0.001 | 0.018 | ||
| WC | MR Egger | 14 | 0.669 | 0.637 | 0.314 | 0.179 | 0.169 |
| Weighted median | 14 | -0.246 | 0.113 | 0.030 | |||
| Inverse variance weighted | 14 | -0.252 | 0.096 | 0.009 | 0.118 | ||
| WHR | MR Egger | 15 | 0.235 | 1.375 | 0.867 | 0.010 | 0.661 |
| Weighted median | 15 | -0.511 | 0.222 | 0.021 | |||
| Inverse variance weighted | 15 | -0.375 | 0.213 | 0.079 | 0.014 | ||
Based on the above results, CHD, BMI, and WC might serve as potential mediators between education and T2DM. Thus, we further evaluated whether these three potential mediators were associated with T2DM using MR analyses. Only BMI (but not CHD or WC) was positively associated with T2DM (Table 4). Hence, BMI might serve as a potential mediator between education and T2DM.
| Trait | Method | nSNP | OR | 95%CI | P value | Heterogeneity P | MR Egger intercept P | |
| CHD-T2DM | MR Egger | 28 | 0.947 | 0.697 | 1.288 | 0.733 | 0.000 | 0.815 |
| Weighted median | 28 | 1.111 | 0.996 | 1.239 | 0.059 | |||
| Inverse variance weighted | 28 | 1.073 | 0.954 | 1.207 | 0.243 | 0.000 | ||
| BMI-T2DM | MR Egger | 72 | 3.370 | 1.328 | 8.556 | 0.013 | 0.000 | 0.240 |
| Weighted median | 72 | 2.622 | 2.164 | 3.178 | 0.000 | |||
| Inverse variance weighted | 72 | 2.046 | 1.374 | 3.048 | 0.000 | 0.000 | ||
| WC-T2DM | MR Egger | 39 | 11.670 | 1.590 | 85.654 | 0.021 | 0.000 | 0.049 |
| Weighted median | 39 | 2.371 | 1.788 | 3.145 | 0.000 | |||
| Inverse variance weighted | 39 | 1.607 | 0.879 | 2.938 | 0.123 | 0.000 | ||
Diabetes has rapidly become a global epidemic and a significant public health concern. Identifying the high-risk populations and addressing the risk factors for diabetes might prove to be an effective diabetes prevention strategy. Traditional risk factors for T2DM include obesity, diet, physical activity, socioeconomic status, etc.[19], some of which are modifiable through behavioral or pharmacological intervention (e.g., obesity, diet, and physical activity). It is of great importance to determine whether traditional risk factors have a causal role in T2DM or are merely bystanders.
Previous studies have reported that education might help T2DM management. A prospective, randomized, single-center study revealed that pharmacotherapeutic education of patients with T2DM could significantly improve 30-d post-discharge medication adherence without a significant reduction in adverse clinical outcomes[20]. A systematic review and meta-analysis also demonstrated that educational interventions improved medication adherence among adult patients diagnosed with diabetes[20]. Moreover, structured education had a positive impact on glucose control and hypoglycemia in T2DM[22]. However, there has been no direct study exploring the causality between original education and T2DM.
Interestingly, previous findings indicated a causal association between low educational attainment and increased risk of smoking[23]. Also, observational studies suggested an association between smoking and risk of T2DM[24,25]. Based on the above reviews, we made a bold assumption that education might be negatively causally associated with T2DM. As we expected, our MR results revealed that the OR (95%CI) for T2DM was 0.392 (0.263-0.583) per standard deviation increase (3.6 years) in education, indicating that the genetically predicted education was negatively causally associated with T2DM. Since education was shown to be a protective factor against T2DM, and it was modifiable, longer education years among the population are recommended for T2DM prevention.
As there was no full description of the underlying mechanisms possibly connecting education to T2DM, we investigated this relationship. As for the possible mediators from education to T2DM, ten representative modifiable metabolic risk factors were chosen for further MR analysis. Our results indicated that education was causally associated with CHD, TG, BMI, WC, and WHR in the discovery phase. Yet, in the replication data set, only the results for CHD, BMI, and WC were duplicated. Therefore, there were negative causal associations between genetically determined education and CHD, BMI, and WC, which were considered to be the potential mediators between education and T2DM.
A previous study revealed that genetic predisposition towards 3.6 years of additional education was associated with a one-third lower risk of CHD, indicating that low education was a causal risk factor in the development of CHD[26], which was in accord with our finding. Another MR analysis also suggested that there might be a negative causal effect of education on BMI[27]. Our results provide more evidence supporting that more education years might help the public better control BMI, which is beneficial for blood glucose regulation and T2DM prevention. There was no direct causal study exploring the effect of education on WC. A randomized clinical trial revealed that nutrition therapy and a multimedia diabetes education program positively impacted achieving metabolic control goals in T2DM, including HbA1C, glucose decrease, TG, and weight loss. Yet, the WC change was still not statistically different[28]. Another study reported that diet-related and lifestyle-related school-based education could reduce central adiposity in pre-teenagers[29]. We analyzed whether the different effects of education on WC might be due to various educational programs or diverse populations used in the studies.
Since our MR investigation indicated that CHD, BMI, and WC might be potential mediators from less education years to increased risk of T2DM, we further evaluated whether these three potential mediators were associated with T2DM by MR analysis. Only BMI was positively associated with T2DM. In summary, BMI might serve as a mediator between education and T2DM.
BMI is a well-described risk factor for T2DM[30], and several large-scale MR studies have addressed its positive causal association with T2DM[31-33]. Our results also demonstrated that higher BMI could lead to higher risk of T2MD. Combined with the effects of education on BMI and T2MD, for most developing countries where the majority of population receives short education years, we recommend longer education time and a BMI control program for public T2DM prevention.
Short education was found to be associated with an increased T2DM risk. BMI might serve as a potential mediator between them.
The causality between education and type 2 diabetes mellitus (T2DM) remains unclear.
In this study, a network Mendelian randomization (MR) framework was applied to determine the causality between education and T2DM from summarized genome-wide association study data.
We used a network MR to identify the causality between education and T2DM and the potential metabolic risk factors [coronary heart disease (CHD), total cholesterol, low-density lipoprotein, triglycerides, body mass index (BMI), waist circumference (WC), waist-to-hip ratio, fasting insulin, fasting glucose, and glycated hemoglobin] from summarized genome-wide association study data.
Two-sample MR and network MR were performed to obtain the causality between education-T2DM, education-mediator, and mediator-T2DM. Summary statistics from the Social Science Genetic Association Consortium (discovery data) and Neale Lab consortium (replication data) were used for education. DIAGRAM plus Metabochip consortium data were utilized for T2DM.
In the IVW method, the odds ratio (95%CI) for T2DM was 0.392 (0.263-0.583) per standard deviation increase (3.6 years) in education, without heterogeneity or horizontal pleiotropy. Education was genetically associated with CHD, triglycerides, BMI, WC, and waist-to-hip ratio in the discovery phase, yet only the results for CHD, BMI, and WC were confirmed in the replication data. Moreover, BMI was positively associated with T2DM.
Short education was found to be associated with increased T2DM risk. BMI might serve as a potential mediator between them.
For most developing countries, the majority of the population receive short education years. Longer education time is recommended as is a BMI control program, for public T2DM prevention.
Manuscript source: Unsolicited manuscript
Specialty type: Genetics and heredity
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Karatza AA S-Editor: Zhang H L-Editor: Wang TQ P-Editor: Yuan YY
| 1. | Dall TM, Narayan KM, Gillespie KB, Gallo PD, Blanchard TD, Solcan M, O'Grady M, Quick WW. Detecting type 2 diabetes and prediabetes among asymptomatic adults in the United States: modeling American Diabetes Association versus US Preventive Services Task Force diabetes screening guidelines. Popul Health Metr. 2014;12:12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 53] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 2. | Fletcher B, Gulanick M, Lamendola C. Risk factors for type 2 diabetes mellitus. J Cardiovasc Nurs. 2002;16:17-23. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 172] [Cited by in RCA: 225] [Article Influence: 9.8] [Reference Citation Analysis (1)] |
| 3. | Vitale M, Masulli M, Calabrese I, Rivellese AA, Bonora E, Signorini S, Perriello G, Squatrito S, Buzzetti R, Sartore G, Babini AC, Gregori G, Giordano C, Clemente G, Grioni S, Dolce P, Riccardi G, Vaccaro O; TOSCA. IT Study Group. Impact of a Mediterranean Dietary Pattern and Its Components on Cardiovascular Risk Factors, Glucose Control, and Body Weight in People with Type 2 Diabetes: A Real-Life Study. Nutrients. 2018;10. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 97] [Article Influence: 13.9] [Reference Citation Analysis (0)] |
| 4. | He X, Li J, Wang B, Yao Q, Li L, Song R, Shi X, Zhang JA. Diabetes self-management education reduces risk of all-cause mortality in type 2 diabetes patients: a systematic review and meta-analysis. Endocrine. 2017;55:712-731. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 136] [Cited by in RCA: 157] [Article Influence: 19.6] [Reference Citation Analysis (0)] |
| 5. | Emdin CA, Khera AV, Kathiresan S. Mendelian Randomization. JAMA. 2017;318:1925-1926. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 500] [Cited by in RCA: 2190] [Article Influence: 273.8] [Reference Citation Analysis (0)] |
| 6. | Okbay A, Beauchamp JP, Fontana MA, Lee JJ, Pers TH, Rietveld CA, Turley P, Chen GB, Emilsson V, Meddens SF, Oskarsson S, Pickrell JK, Thom K, Timshel P, de Vlaming R, Abdellaoui A, Ahluwalia TS, Bacelis J, Baumbach C, Bjornsdottir G, Brandsma JH, Pina Concas M, Derringer J, Furlotte NA, Galesloot TE, Girotto G, Gupta R, Hall LM, Harris SE, Hofer E, Horikoshi M, Huffman JE, Kaasik K, Kalafati IP, Karlsson R, Kong A, Lahti J, van der Lee SJ, deLeeuw C, Lind PA, Lindgren KO, Liu T, Mangino M, Marten J, Mihailov E, Miller MB, van der Most PJ, Oldmeadow C, Payton A, Pervjakova N, Peyrot WJ, Qian Y, Raitakari O, Rueedi R, Salvi E, Schmidt B, Schraut KE, Shi J, Smith AV, Poot RA, St Pourcain B, Teumer A, Thorleifsson G, Verweij N, Vuckovic D, Wellmann J, Westra HJ, Yang J, Zhao W, Zhu Z, Alizadeh BZ, Amin N, Bakshi A, Baumeister SE, Biino G, Bønnelykke K, Boyle PA, Campbell H, Cappuccio FP, Davies G, De Neve JE, Deloukas P, Demuth I, Ding J, Eibich P, Eisele L, Eklund N, Evans DM, Faul JD, Feitosa MF, Forstner AJ, Gandin I, Gunnarsson B, Halldórsson BV, Harris TB, Heath AC, Hocking LJ, Holliday EG, Homuth G, Horan MA, Hottenga JJ, de Jager PL, Joshi PK, Jugessur A, Kaakinen MA, Kähönen M, Kanoni S, Keltigangas-Järvinen L, Kiemeney LA, Kolcic I, Koskinen S, Kraja AT, Kroh M, Kutalik Z, Latvala A, Launer LJ, Lebreton MP, Levinson DF, Lichtenstein P, Lichtner P, Liewald DC; LifeLines Cohort Study, Loukola A, Madden PA, Mägi R, Mäki-Opas T, Marioni RE, Marques-Vidal P, Meddens GA, McMahon G, Meisinger C, Meitinger T, Milaneschi Y, Milani L, Montgomery GW, Myhre R, Nelson CP, Nyholt DR, Ollier WE, Palotie A, Paternoster L, Pedersen NL, Petrovic KE, Porteous DJ, Räikkönen K, Ring SM, Robino A, Rostapshova O, Rudan I, Rustichini A, Salomaa V, Sanders AR, Sarin AP, Schmidt H, Scott RJ, Smith BH, Smith JA, Staessen JA, Steinhagen-Thiessen E, Strauch K, Terracciano A, Tobin MD, Ulivi S, Vaccargiu S, Quaye L, van Rooij FJ, Venturini C, Vinkhuyzen AA, Völker U, Völzke H, Vonk JM, Vozzi D, Waage J, Ware EB, Willemsen G, Attia JR, Bennett DA, Berger K, Bertram L, Bisgaard H, Boomsma DI, Borecki IB, Bültmann U, Chabris CF, Cucca F, Cusi D, Deary IJ, Dedoussis GV, van Duijn CM, Eriksson JG, Franke B, Franke L, Gasparini P, Gejman PV, Gieger C, Grabe HJ, Gratten J, Groenen PJ, Gudnason V, van der Harst P, Hayward C, Hinds DA, Hoffmann W, Hyppönen E, Iacono WG, Jacobsson B, Järvelin MR, Jöckel KH, Kaprio J, Kardia SL, Lehtimäki T, Lehrer SF, Magnusson PK, Martin NG, McGue M, Metspalu A, Pendleton N, Penninx BW, Perola M, Pirastu N, Pirastu M, Polasek O, Posthuma D, Power C, Province MA, Samani NJ, Schlessinger D, Schmidt R, Sørensen TI, Spector TD, Stefansson K, Thorsteinsdottir U, Thurik AR, Timpson NJ, Tiemeier H, Tung JY, Uitterlinden AG, Vitart V, Vollenweider P, Weir DR, Wilson JF, Wright AF, Conley DC, Krueger RF, Davey Smith G, Hofman A, Laibson DI, Medland SE, Meyer MN, Yang J, Johannesson M, Visscher PM, Esko T, Koellinger PD, Cesarini D, Benjamin DJ. Genome-wide association study identifies 74 loci associated with educational attainment. Nature. 2016;533:539-542. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 947] [Cited by in RCA: 805] [Article Influence: 89.4] [Reference Citation Analysis (0)] |
| 7. | Morris AP, Voight BF, Teslovich TM, Ferreira T, Segrè AV, Steinthorsdottir V, Strawbridge RJ, Khan H, Grallert H, Mahajan A, Prokopenko I, Kang HM, Dina C, Esko T, Fraser RM, Kanoni S, Kumar A, Lagou V, Langenberg C, Luan J, Lindgren CM, Müller-Nurasyid M, Pechlivanis S, Rayner NW, Scott LJ, Wiltshire S, Yengo L, Kinnunen L, Rossin EJ, Raychaudhuri S, Johnson AD, Dimas AS, Loos RJ, Vedantam S, Chen H, Florez JC, Fox C, Liu CT, Rybin D, Couper DJ, Kao WH, Li M, Cornelis MC, Kraft P, Sun Q, van Dam RM, Stringham HM, Chines PS, Fischer K, Fontanillas P, Holmen OL, Hunt SE, Jackson AU, Kong A, Lawrence R, Meyer J, Perry JR, Platou CG, Potter S, Rehnberg E, Robertson N, Sivapalaratnam S, Stančáková A, Stirrups K, Thorleifsson G, Tikkanen E, Wood AR, Almgren P, Atalay M, Benediktsson R, Bonnycastle LL, Burtt N, Carey J, Charpentier G, Crenshaw AT, Doney AS, Dorkhan M, Edkins S, Emilsson V, Eury E, Forsen T, Gertow K, Gigante B, Grant GB, Groves CJ, Guiducci C, Herder C, Hreidarsson AB, Hui J, James A, Jonsson A, Rathmann W, Klopp N, Kravic J, Krjutškov K, Langford C, Leander K, Lindholm E, Lobbens S, Männistö S, Mirza G, Mühleisen TW, Musk B, Parkin M, Rallidis L, Saramies J, Sennblad B, Shah S, Sigurðsson G, Silveira A, Steinbach G, Thorand B, Trakalo J, Veglia F, Wennauer R, Winckler W, Zabaneh D, Campbell H, van Duijn C, Uitterlinden AG, Hofman A, Sijbrands E, Abecasis GR, Owen KR, Zeggini E, Trip MD, Forouhi NG, Syvänen AC, Eriksson JG, Peltonen L, Nöthen MM, Balkau B, Palmer CN, Lyssenko V, Tuomi T, Isomaa B, Hunter DJ, Qi L; Wellcome Trust Case Control Consortium; Meta-Analyses of Glucose and Insulin-related traits Consortium (MAGIC) Investigators; Genetic Investigation of ANthropometric Traits (GIANT) Consortium; Asian Genetic Epidemiology Network–Type 2 Diabetes (AGEN-T2D) Consortium; South Asian Type 2 Diabetes (SAT2D) Consortium, Shuldiner AR, Roden M, Barroso I, Wilsgaard T, Beilby J, Hovingh K, Price JF, Wilson JF, Rauramaa R, Lakka TA, Lind L, Dedoussis G, Njølstad I, Pedersen NL, Khaw KT, Wareham NJ, Keinanen-Kiukaanniemi SM, Saaristo TE, Korpi-Hyövälti E, Saltevo J, Laakso M, Kuusisto J, Metspalu A, Collins FS, Mohlke KL, Bergman RN, Tuomilehto J, Boehm BO, Gieger C, Hveem K, Cauchi S, Froguel P, Baldassarre D, Tremoli E, Humphries SE, Saleheen D, Danesh J, Ingelsson E, Ripatti S, Salomaa V, Erbel R, Jöckel KH, Moebus S, Peters A, Illig T, de Faire U, Hamsten A, Morris AD, Donnelly PJ, Frayling TM, Hattersley AT, Boerwinkle E, Melander O, Kathiresan S, Nilsson PM, Deloukas P, Thorsteinsdottir U, Groop LC, Stefansson K, Hu F, Pankow JS, Dupuis J, Meigs JB, Altshuler D, Boehnke M, McCarthy MI; DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat Genet. 2012;44:981-990. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1706] [Cited by in RCA: 1477] [Article Influence: 113.6] [Reference Citation Analysis (0)] |
| 8. | Nikpay M, Goel A, Won HH, Hall LM, Willenborg C, Kanoni S, Saleheen D, Kyriakou T, Nelson CP, Hopewell JC, Webb TR, Zeng L, Dehghan A, Alver M, Armasu SM, Auro K, Bjonnes A, Chasman DI, Chen S, Ford I, Franceschini N, Gieger C, Grace C, Gustafsson S, Huang J, Hwang SJ, Kim YK, Kleber ME, Lau KW, Lu X, Lu Y, Lyytikäinen LP, Mihailov E, Morrison AC, Pervjakova N, Qu L, Rose LM, Salfati E, Saxena R, Scholz M, Smith AV, Tikkanen E, Uitterlinden A, Yang X, Zhang W, Zhao W, de Andrade M, de Vries PS, van Zuydam NR, Anand SS, Bertram L, Beutner F, Dedoussis G, Frossard P, Gauguier D, Goodall AH, Gottesman O, Haber M, Han BG, Huang J, Jalilzadeh S, Kessler T, König IR, Lannfelt L, Lieb W, Lind L, Lindgren CM, Lokki ML, Magnusson PK, Mallick NH, Mehra N, Meitinger T, Memon FU, Morris AP, Nieminen MS, Pedersen NL, Peters A, Rallidis LS, Rasheed A, Samuel M, Shah SH, Sinisalo J, Stirrups KE, Trompet S, Wang L, Zaman KS, Ardissino D, Boerwinkle E, Borecki IB, Bottinger EP, Buring JE, Chambers JC, Collins R, Cupples LA, Danesh J, Demuth I, Elosua R, Epstein SE, Esko T, Feitosa MF, Franco OH, Franzosi MG, Granger CB, Gu D, Gudnason V, Hall AS, Hamsten A, Harris TB, Hazen SL, Hengstenberg C, Hofman A, Ingelsson E, Iribarren C, Jukema JW, Karhunen PJ, Kim BJ, Kooner JS, Kullo IJ, Lehtimäki T, Loos RJF, Melander O, Metspalu A, März W, Palmer CN, Perola M, Quertermous T, Rader DJ, Ridker PM, Ripatti S, Roberts R, Salomaa V, Sanghera DK, Schwartz SM, Seedorf U, Stewart AF, Stott DJ, Thiery J, Zalloua PA, O'Donnell CJ, Reilly MP, Assimes TL, Thompson JR, Erdmann J, Clarke R, Watkins H, Kathiresan S, McPherson R, Deloukas P, Schunkert H, Samani NJ, Farrall M. A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat Genet. 2015;47:1121-1130. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1530] [Cited by in RCA: 1810] [Article Influence: 181.0] [Reference Citation Analysis (0)] |
| 9. | Willer CJ, Schmidt EM, Sengupta S, Peloso GM, Gustafsson S, Kanoni S, Ganna A, Chen J, Buchkovich ML, Mora S, Beckmann JS, Bragg-Gresham JL, Chang HY, Demirkan A, Den Hertog HM, Do R, Donnelly LA, Ehret GB, Esko T, Feitosa MF, Ferreira T, Fischer K, Fontanillas P, Fraser RM, Freitag DF, Gurdasani D, Heikkilä K, Hyppönen E, Isaacs A, Jackson AU, Johansson Å, Johnson T, Kaakinen M, Kettunen J, Kleber ME, Li X, Luan J, Lyytikäinen LP, Magnusson PKE, Mangino M, Mihailov E, Montasser ME, Müller-Nurasyid M, Nolte IM, O'Connell JR, Palmer CD, Perola M, Petersen AK, Sanna S, Saxena R, Service SK, Shah S, Shungin D, Sidore C, Song C, Strawbridge RJ, Surakka I, Tanaka T, Teslovich TM, Thorleifsson G, Van den Herik EG, Voight BF, Volcik KA, Waite LL, Wong A, Wu Y, Zhang W, Absher D, Asiki G, Barroso I, Been LF, Bolton JL, Bonnycastle LL, Brambilla P, Burnett MS, Cesana G, Dimitriou M, Doney ASF, Döring A, Elliott P, Epstein SE, Ingi Eyjolfsson G, Gigante B, Goodarzi MO, Grallert H, Gravito ML, Groves CJ, Hallmans G, Hartikainen AL, Hayward C, Hernandez D, Hicks AA, Holm H, Hung YJ, Illig T, Jones MR, Kaleebu P, Kastelein JJP, Khaw KT, Kim E, Klopp N, Komulainen P, Kumari M, Langenberg C, Lehtimäki T, Lin SY, Lindström J, Loos RJF, Mach F, McArdle WL, Meisinger C, Mitchell BD, Müller G, Nagaraja R, Narisu N, Nieminen TVM, Nsubuga RN, Olafsson I, Ong KK, Palotie A, Papamarkou T, Pomilla C, Pouta A, Rader DJ, Reilly MP, Ridker PM, Rivadeneira F, Rudan I, Ruokonen A, Samani N, Scharnagl H, Seeley J, Silander K, Stančáková A, Stirrups K, Swift AJ, Tiret L, Uitterlinden AG, van Pelt LJ, Vedantam S, Wainwright N, Wijmenga C, Wild SH, Willemsen G, Wilsgaard T, Wilson JF, Young EH, Zhao JH, Adair LS, Arveiler D, Assimes TL, Bandinelli S, Bennett F, Bochud M, Boehm BO, Boomsma DI, Borecki IB, Bornstein SR, Bovet P, Burnier M, Campbell H, Chakravarti A, Chambers JC, Chen YI, Collins FS, Cooper RS, Danesh J, Dedoussis G, de Faire U, Feranil AB, Ferrières J, Ferrucci L, Freimer NB, Gieger C, Groop LC, Gudnason V, Gyllensten U, Hamsten A, Harris TB, Hingorani A, Hirschhorn JN, Hofman A, Hovingh GK, Hsiung CA, Humphries SE, Hunt SC, Hveem K, Iribarren C, Järvelin MR, Jula A, Kähönen M, Kaprio J, Kesäniemi A, Kivimaki M, Kooner JS, Koudstaal PJ, Krauss RM, Kuh D, Kuusisto J, Kyvik KO, Laakso M, Lakka TA, Lind L, Lindgren CM, Martin NG, März W, McCarthy MI, McKenzie CA, Meneton P, Metspalu A, Moilanen L, Morris AD, Munroe PB, Njølstad I, Pedersen NL, Power C, Pramstaller PP, Price JF, Psaty BM, Quertermous T, Rauramaa R, Saleheen D, Salomaa V, Sanghera DK, Saramies J, Schwarz PEH, Sheu WH, Shuldiner AR, Siegbahn A, Spector TD, Stefansson K, Strachan DP, Tayo BO, Tremoli E, Tuomilehto J, Uusitupa M, van Duijn CM, Vollenweider P, Wallentin L, Wareham NJ, Whitfield JB, Wolffenbuttel BHR, Ordovas JM, Boerwinkle E, Palmer CNA, Thorsteinsdottir U, Chasman DI, Rotter JI, Franks PW, Ripatti S, Cupples LA, Sandhu MS, Rich SS, Boehnke M, Deloukas P, Kathiresan S, Mohlke KL, Ingelsson E, Abecasis GR; Global Lipids Genetics Consortium. Discovery and refinement of loci associated with lipid levels. Nat Genet. 2013;45:1274-1283. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2340] [Cited by in RCA: 2324] [Article Influence: 193.7] [Reference Citation Analysis (0)] |
| 10. | Locke AE, Kahali B, Berndt SI, Justice AE, Pers TH, Day FR, Powell C, Vedantam S, Buchkovich ML, Yang J, Croteau-Chonka DC, Esko T, Fall T, Ferreira T, Gustafsson S, Kutalik Z, Luan J, Mägi R, Randall JC, Winkler TW, Wood AR, Workalemahu T, Faul JD, Smith JA, Zhao JH, Zhao W, Chen J, Fehrmann R, Hedman ÅK, Karjalainen J, Schmidt EM, Absher D, Amin N, Anderson D, Beekman M, Bolton JL, Bragg-Gresham JL, Buyske S, Demirkan A, Deng G, Ehret GB, Feenstra B, Feitosa MF, Fischer K, Goel A, Gong J, Jackson AU, Kanoni S, Kleber ME, Kristiansson K, Lim U, Lotay V, Mangino M, Leach IM, Medina-Gomez C, Medland SE, Nalls MA, Palmer CD, Pasko D, Pechlivanis S, Peters MJ, Prokopenko I, Shungin D, Stančáková A, Strawbridge RJ, Sung YJ, Tanaka T, Teumer A, Trompet S, van der Laan SW, van Setten J, Van Vliet-Ostaptchouk JV, Wang Z, Yengo L, Zhang W, Isaacs A, Albrecht E, Ärnlöv J, Arscott GM, Attwood AP, Bandinelli S, Barrett A, Bas IN, Bellis C, Bennett AJ, Berne C, Blagieva R, Blüher M, Böhringer S, Bonnycastle LL, Böttcher Y, Boyd HA, Bruinenberg M, Caspersen IH, Chen YI, Clarke R, Daw EW, de Craen AJM, Delgado G, Dimitriou M, Doney ASF, Eklund N, Estrada K, Eury E, Folkersen L, Fraser RM, Garcia ME, Geller F, Giedraitis V, Gigante B, Go AS, Golay A, Goodall AH, Gordon SD, Gorski M, Grabe HJ, Grallert H, Grammer TB, Gräßler J, Grönberg H, Groves CJ, Gusto G, Haessler J, Hall P, Haller T, Hallmans G, Hartman CA, Hassinen M, Hayward C, Heard-Costa NL, Helmer Q, Hengstenberg C, Holmen O, Hottenga JJ, James AL, Jeff JM, Johansson Å, Jolley J, Juliusdottir T, Kinnunen L, Koenig W, Koskenvuo M, Kratzer W, Laitinen J, Lamina C, Leander K, Lee NR, Lichtner P, Lind L, Lindström J, Lo KS, Lobbens S, Lorbeer R, Lu Y, Mach F, Magnusson PKE, Mahajan A, McArdle WL, McLachlan S, Menni C, Merger S, Mihailov E, Milani L, Moayyeri A, Monda KL, Morken MA, Mulas A, Müller G, Müller-Nurasyid M, Musk AW, Nagaraja R, Nöthen MM, Nolte IM, Pilz S, Rayner NW, Renstrom F, Rettig R, Ried JS, Ripke S, Robertson NR, Rose LM, Sanna S, Scharnagl H, Scholtens S, Schumacher FR, Scott WR, Seufferlein T, Shi J, Smith AV, Smolonska J, Stanton AV, Steinthorsdottir V, Stirrups K, Stringham HM, Sundström J, Swertz MA, Swift AJ, Syvänen AC, Tan ST, Tayo BO, Thorand B, Thorleifsson G, Tyrer JP, Uh HW, Vandenput L, Verhulst FC, Vermeulen SH, Verweij N, Vonk JM, Waite LL, Warren HR, Waterworth D, Weedon MN, Wilkens LR, Willenborg C, Wilsgaard T, Wojczynski MK, Wong A, Wright AF, Zhang Q; LifeLines Cohort Study, Brennan EP, Choi M, Dastani Z, Drong AW, Eriksson P, Franco-Cereceda A, Gådin JR, Gharavi AG, Goddard ME, Handsaker RE, Huang J, Karpe F, Kathiresan S, Keildson S, Kiryluk K, Kubo M, Lee JY, Liang L, Lifton RP, Ma B, McCarroll SA, McKnight AJ, Min JL, Moffatt MF, Montgomery GW, Murabito JM, Nicholson G, Nyholt DR, Okada Y, Perry JRB, Dorajoo R, Reinmaa E, Salem RM, Sandholm N, Scott RA, Stolk L, Takahashi A, Tanaka T, van 't Hooft FM, Vinkhuyzen AAE, Westra HJ, Zheng W, Zondervan KT; ADIPOGen Consortium; AGEN-BMI Working Group; CARDIOGRAMplusC4D Consortium; CKDGen Consortium; GLGC; ICBP; MAGIC Investigators; MuTHER Consortium; MIGen Consortium; PAGE Consortium; ReproGen Consortium; GENIE Consortium; International Endogene Consortium, Heath AC, Arveiler D, Bakker SJL, Beilby J, Bergman RN, Blangero J, Bovet P, Campbell H, Caulfield MJ, Cesana G, Chakravarti A, Chasman DI, Chines PS, Collins FS, Crawford DC, Cupples LA, Cusi D, Danesh J, de Faire U, den Ruijter HM, Dominiczak AF, Erbel R, Erdmann J, Eriksson JG, Farrall M, Felix SB, Ferrannini E, Ferrières J, Ford I, Forouhi NG, Forrester T, Franco OH, Gansevoort RT, Gejman PV, Gieger C, Gottesman O, Gudnason V, Gyllensten U, Hall AS, Harris TB, Hattersley AT, Hicks AA, Hindorff LA, Hingorani AD, Hofman A, Homuth G, Hovingh GK, Humphries SE, Hunt SC, Hyppönen E, Illig T, Jacobs KB, Jarvelin MR, Jöckel KH, Johansen B, Jousilahti P, Jukema JW, Jula AM, Kaprio J, Kastelein JJP, Keinanen-Kiukaanniemi SM, Kiemeney LA, Knekt P, Kooner JS, Kooperberg C, Kovacs P, Kraja AT, Kumari M, Kuusisto J, Lakka TA, Langenberg C, Marchand LL, Lehtimäki T, Lyssenko V, Männistö S, Marette A, Matise TC, McKenzie CA, McKnight B, Moll FL, Morris AD, Morris AP, Murray JC, Nelis M, Ohlsson C, Oldehinkel AJ, Ong KK, Madden PAF, Pasterkamp G, Peden JF, Peters A, Postma DS, Pramstaller PP, Price JF, Qi L, Raitakari OT, Rankinen T, Rao DC, Rice TK, Ridker PM, Rioux JD, Ritchie MD, Rudan I, Salomaa V, Samani NJ, Saramies J, Sarzynski MA, Schunkert H, Schwarz PEH, Sever P, Shuldiner AR, Sinisalo J, Stolk RP, Strauch K, Tönjes A, Trégouët DA, Tremblay A, Tremoli E, Virtamo J, Vohl MC, Völker U, Waeber G, Willemsen G, Witteman JC, Zillikens MC, Adair LS, Amouyel P, Asselbergs FW, Assimes TL, Bochud M, Boehm BO, Boerwinkle E, Bornstein SR, Bottinger EP, Bouchard C, Cauchi S, Chambers JC, Chanock SJ, Cooper RS, de Bakker PIW, Dedoussis G, Ferrucci L, Franks PW, Froguel P, Groop LC, Haiman CA, Hamsten A, Hui J, Hunter DJ, Hveem K, Kaplan RC, Kivimaki M, Kuh D, Laakso M, Liu Y, Martin NG, März W, Melbye M, Metspalu A, Moebus S, Munroe PB, Njølstad I, Oostra BA, Palmer CNA, Pedersen NL, Perola M, Pérusse L, Peters U, Power C, Quertermous T, Rauramaa R, Rivadeneira F, Saaristo TE, Saleheen D, Sattar N, Schadt EE, Schlessinger D, Slagboom PE, Snieder H, Spector TD, Thorsteinsdottir U, Stumvoll M, Tuomilehto J, Uitterlinden AG, Uusitupa M, van der Harst P, Walker M, Wallaschofski H, Wareham NJ, Watkins H, Weir DR, Wichmann HE, Wilson JF, Zanen P, Borecki IB, Deloukas P, Fox CS, Heid IM, O'Connell JR, Strachan DP, Stefansson K, van Duijn CM, Abecasis GR, Franke L, Frayling TM, McCarthy MI, Visscher PM, Scherag A, Willer CJ, Boehnke M, Mohlke KL, Lindgren CM, Beckmann JS, Barroso I, North KE, Ingelsson E, Hirschhorn JN, Loos RJF, Speliotes EK. Genetic studies of body mass index yield new insights for obesity biology. Nature. 2015;518:197-206. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2987] [Cited by in RCA: 3253] [Article Influence: 325.3] [Reference Citation Analysis (0)] |
| 11. | Shungin D, Winkler TW, Croteau-Chonka DC, Ferreira T, Locke AE, Mägi R, Strawbridge RJ, Pers TH, Fischer K, Justice AE, Workalemahu T, Wu JMW, Buchkovich ML, Heard-Costa NL, Roman TS, Drong AW, Song C, Gustafsson S, Day FR, Esko T, Fall T, Kutalik Z, Luan J, Randall JC, Scherag A, Vedantam S, Wood AR, Chen J, Fehrmann R, Karjalainen J, Kahali B, Liu CT, Schmidt EM, Absher D, Amin N, Anderson D, Beekman M, Bragg-Gresham JL, Buyske S, Demirkan A, Ehret GB, Feitosa MF, Goel A, Jackson AU, Johnson T, Kleber ME, Kristiansson K, Mangino M, Leach IM, Medina-Gomez C, Palmer CD, Pasko D, Pechlivanis S, Peters MJ, Prokopenko I, Stančáková A, Sung YJ, Tanaka T, Teumer A, Van Vliet-Ostaptchouk JV, Yengo L, Zhang W, Albrecht E, Ärnlöv J, Arscott GM, Bandinelli S, Barrett A, Bellis C, Bennett AJ, Berne C, Blüher M, Böhringer S, Bonnet F, Böttcher Y, Bruinenberg M, Carba DB, Caspersen IH, Clarke R, Daw EW, Deelen J, Deelman E, Delgado G, Doney AS, Eklund N, Erdos MR, Estrada K, Eury E, Friedrich N, Garcia ME, Giedraitis V, Gigante B, Go AS, Golay A, Grallert H, Grammer TB, Gräßler J, Grewal J, Groves CJ, Haller T, Hallmans G, Hartman CA, Hassinen M, Hayward C, Heikkilä K, Herzig KH, Helmer Q, Hillege HL, Holmen O, Hunt SC, Isaacs A, Ittermann T, James AL, Johansson I, Juliusdottir T, Kalafati IP, Kinnunen L, Koenig W, Kooner IK, Kratzer W, Lamina C, Leander K, Lee NR, Lichtner P, Lind L, Lindström J, Lobbens S, Lorentzon M, Mach F, Magnusson PK, Mahajan A, McArdle WL, Menni C, Merger S, Mihailov E, Milani L, Mills R, Moayyeri A, Monda KL, Mooijaart SP, Mühleisen TW, Mulas A, Müller G, Müller-Nurasyid M, Nagaraja R, Nalls MA, Narisu N, Glorioso N, Nolte IM, Olden M, Rayner NW, Renstrom F, Ried JS, Robertson NR, Rose LM, Sanna S, Scharnagl H, Scholtens S, Sennblad B, Seufferlein T, Sitlani CM, Smith AV, Stirrups K, Stringham HM, Sundström J, Swertz MA, Swift AJ, Syvänen AC, Tayo BO, Thorand B, Thorleifsson G, Tomaschitz A, Troffa C, van Oort FV, Verweij N, Vonk JM, Waite LL, Wennauer R, Wilsgaard T, Wojczynski MK, Wong A, Zhang Q, Zhao JH, Brennan EP, Choi M, Eriksson P, Folkersen L, Franco-Cereceda A, Gharavi AG, Hedman ÅK, Hivert MF, Huang J, Kanoni S, Karpe F, Keildson S, Kiryluk K, Liang L, Lifton RP, Ma B, McKnight AJ, McPherson R, Metspalu A, Min JL, Moffatt MF, Montgomery GW, Murabito JM, Nicholson G, Nyholt DR, Olsson C, Perry JR, Reinmaa E, Salem RM, Sandholm N, Schadt EE, Scott RA, Stolk L, Vallejo EE, Westra HJ, Zondervan KT; ADIPOGen Consortium; CARDIOGRAMplusC4D Consortium; CKDGen Consortium; GEFOS Consortium; GENIE Consortium; GLGC; ICBP; International Endogene Consortium; LifeLines Cohort Study; MAGIC Investigators; MuTHER Consortium; PAGE Consortium; ReproGen Consortium, Amouyel P, Arveiler D, Bakker SJ, Beilby J, Bergman RN, Blangero J, Brown MJ, Burnier M, Campbell H, Chakravarti A, Chines PS, Claudi-Boehm S, Collins FS, Crawford DC, Danesh J, de Faire U, de Geus EJ, Dörr M, Erbel R, Eriksson JG, Farrall M, Ferrannini E, Ferrières J, Forouhi NG, Forrester T, Franco OH, Gansevoort RT, Gieger C, Gudnason V, Haiman CA, Harris TB, Hattersley AT, Heliövaara M, Hicks AA, Hingorani AD, Hoffmann W, Hofman A, Homuth G, Humphries SE, Hyppönen E, Illig T, Jarvelin MR, Johansen B, Jousilahti P, Jula AM, Kaprio J, Kee F, Keinanen-Kiukaanniemi SM, Kooner JS, Kooperberg C, Kovacs P, Kraja AT, Kumari M, Kuulasmaa K, Kuusisto J, Lakka TA, Langenberg C, Le Marchand L, Lehtimäki T, Lyssenko V, Männistö S, Marette A, Matise TC, McKenzie CA, McKnight B, Musk AW, Möhlenkamp S, Morris AD, Nelis M, Ohlsson C, Oldehinkel AJ, Ong KK, Palmer LJ, Penninx BW, Peters A, Pramstaller PP, Raitakari OT, Rankinen T, Rao DC, Rice TK, Ridker PM, Ritchie MD, Rudan I, Salomaa V, Samani NJ, Saramies J, Sarzynski MA, Schwarz PE, Shuldiner AR, Staessen JA, Steinthorsdottir V, Stolk RP, Strauch K, Tönjes A, Tremblay A, Tremoli E, Vohl MC, Völker U, Vollenweider P, Wilson JF, Witteman JC, Adair LS, Bochud M, Boehm BO, Bornstein SR, Bouchard C, Cauchi S, Caulfield MJ, Chambers JC, Chasman DI, Cooper RS, Dedoussis G, Ferrucci L, Froguel P, Grabe HJ, Hamsten A, Hui J, Hveem K, Jöckel KH, Kivimaki M, Kuh D, Laakso M, Liu Y, März W, Munroe PB, Njølstad I, Oostra BA, Palmer CN, Pedersen NL, Perola M, Pérusse L, Peters U, Power C, Quertermous T, Rauramaa R, Rivadeneira F, Saaristo TE, Saleheen D, Sinisalo J, Slagboom PE, Snieder H, Spector TD, Stefansson K, Stumvoll M, Tuomilehto J, Uitterlinden AG, Uusitupa M, van der Harst P, Veronesi G, Walker M, Wareham NJ, Watkins H, Wichmann HE, Abecasis GR, Assimes TL, Berndt SI, Boehnke M, Borecki IB, Deloukas P, Franke L, Frayling TM, Groop LC, Hunter DJ, Kaplan RC, O'Connell JR, Qi L, Schlessinger D, Strachan DP, Thorsteinsdottir U, van Duijn CM, Willer CJ, Visscher PM, Yang J, Hirschhorn JN, Zillikens MC, McCarthy MI, Speliotes EK, North KE, Fox CS, Barroso I, Franks PW, Ingelsson E, Heid IM, Loos RJ, Cupples LA, Morris AP, Lindgren CM, Mohlke KL. New genetic loci link adipose and insulin biology to body fat distribution. Nature. 2015;518:187-196. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1401] [Cited by in RCA: 1181] [Article Influence: 118.1] [Reference Citation Analysis (0)] |
| 12. | Randall JC, Winkler TW, Kutalik Z, Berndt SI, Jackson AU, Monda KL, Kilpeläinen TO, Esko T, Mägi R, Li S, Workalemahu T, Feitosa MF, Croteau-Chonka DC, Day FR, Fall T, Ferreira T, Gustafsson S, Locke AE, Mathieson I, Scherag A, Vedantam S, Wood AR, Liang L, Steinthorsdottir V, Thorleifsson G, Dermitzakis ET, Dimas AS, Karpe F, Min JL, Nicholson G, Clegg DJ, Person T, Krohn JP, Bauer S, Buechler C, Eisinger K; DIAGRAM Consortium, Bonnefond A, Froguel P; MAGIC Investigators, Hottenga JJ, Prokopenko I, Waite LL, Harris TB, Smith AV, Shuldiner AR, McArdle WL, Caulfield MJ, Munroe PB, Grönberg H, Chen YD, Li G, Beckmann JS, Johnson T, Thorsteinsdottir U, Teder-Laving M, Khaw KT, Wareham NJ, Zhao JH, Amin N, Oostra BA, Kraja AT, Province MA, Cupples LA, Heard-Costa NL, Kaprio J, Ripatti S, Surakka I, Collins FS, Saramies J, Tuomilehto J, Jula A, Salomaa V, Erdmann J, Hengstenberg C, Loley C, Schunkert H, Lamina C, Wichmann HE, Albrecht E, Gieger C, Hicks AA, Johansson A, Pramstaller PP, Kathiresan S, Speliotes EK, Penninx B, Hartikainen AL, Jarvelin MR, Gyllensten U, Boomsma DI, Campbell H, Wilson JF, Chanock SJ, Farrall M, Goel A, Medina-Gomez C, Rivadeneira F, Estrada K, Uitterlinden AG, Hofman A, Zillikens MC, den Heijer M, Kiemeney LA, Maschio A, Hall P, Tyrer J, Teumer A, Völzke H, Kovacs P, Tönjes A, Mangino M, Spector TD, Hayward C, Rudan I, Hall AS, Samani NJ, Attwood AP, Sambrook JG, Hung J, Palmer LJ, Lokki ML, Sinisalo J, Boucher G, Huikuri H, Lorentzon M, Ohlsson C, Eklund N, Eriksson JG, Barlassina C, Rivolta C, Nolte IM, Snieder H, Van der Klauw MM, Van Vliet-Ostaptchouk JV, Gejman PV, Shi J, Jacobs KB, Wang Z, Bakker SJ, Mateo Leach I, Navis G, van der Harst P, Martin NG, Medland SE, Montgomery GW, Yang J, Chasman DI, Ridker PM, Rose LM, Lehtimäki T, Raitakari O, Absher D, Iribarren C, Basart H, Hovingh KG, Hyppönen E, Power C, Anderson D, Beilby JP, Hui J, Jolley J, Sager H, Bornstein SR, Schwarz PE, Kristiansson K, Perola M, Lindström J, Swift AJ, Uusitupa M, Atalay M, Lakka TA, Rauramaa R, Bolton JL, Fowkes G, Fraser RM, Price JF, Fischer K, Krjutå Kov K, Metspalu A, Mihailov E, Langenberg C, Luan J, Ong KK, Chines PS, Keinanen-Kiukaanniemi SM, Saaristo TE, Edkins S, Franks PW, Hallmans G, Shungin D, Morris AD, Palmer CN, Erbel R, Moebus S, Nöthen MM, Pechlivanis S, Hveem K, Narisu N, Hamsten A, Humphries SE, Strawbridge RJ, Tremoli E, Grallert H, Thorand B, Illig T, Koenig W, Müller-Nurasyid M, Peters A, Boehm BO, Kleber ME, März W, Winkelmann BR, Kuusisto J, Laakso M, Arveiler D, Cesana G, Kuulasmaa K, Virtamo J, Yarnell JW, Kuh D, Wong A, Lind L, de Faire U, Gigante B, Magnusson PK, Pedersen NL, Dedoussis G, Dimitriou M, Kolovou G, Kanoni S, Stirrups K, Bonnycastle LL, Njølstad I, Wilsgaard T, Ganna A, Rehnberg E, Hingorani A, Kivimaki M, Kumari M, Assimes TL, Barroso I, Boehnke M, Borecki IB, Deloukas P, Fox CS, Frayling T, Groop LC, Haritunians T, Hunter D, Ingelsson E, Kaplan R, Mohlke KL, O'Connell JR, Schlessinger D, Strachan DP, Stefansson K, van Duijn CM, Abecasis GR, McCarthy MI, Hirschhorn JN, Qi L, Loos RJ, Lindgren CM, North KE, Heid IM. Sex-stratified genome-wide association studies including 270,000 individuals show sexual dimorphism in genetic loci for anthropometric traits. PLoS Genet. 2013;9:e1003500. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 345] [Cited by in RCA: 325] [Article Influence: 27.1] [Reference Citation Analysis (0)] |
| 13. | Dupuis J, Langenberg C, Prokopenko I, Saxena R, Soranzo N, Jackson AU, Wheeler E, Glazer NL, Bouatia-Naji N, Gloyn AL, Lindgren CM, Mägi R, Morris AP, Randall J, Johnson T, Elliott P, Rybin D, Thorleifsson G, Steinthorsdottir V, Henneman P, Grallert H, Dehghan A, Hottenga JJ, Franklin CS, Navarro P, Song K, Goel A, Perry JR, Egan JM, Lajunen T, Grarup N, Sparsø T, Doney A, Voight BF, Stringham HM, Li M, Kanoni S, Shrader P, Cavalcanti-Proença C, Kumari M, Qi L, Timpson NJ, Gieger C, Zabena C, Rocheleau G, Ingelsson E, An P, O'Connell J, Luan J, Elliott A, McCarroll SA, Payne F, Roccasecca RM, Pattou F, Sethupathy P, Ardlie K, Ariyurek Y, Balkau B, Barter P, Beilby JP, Ben-Shlomo Y, Benediktsson R, Bennett AJ, Bergmann S, Bochud M, Boerwinkle E, Bonnefond A, Bonnycastle LL, Borch-Johnsen K, Böttcher Y, Brunner E, Bumpstead SJ, Charpentier G, Chen YD, Chines P, Clarke R, Coin LJ, Cooper MN, Cornelis M, Crawford G, Crisponi L, Day IN, de Geus EJ, Delplanque J, Dina C, Erdos MR, Fedson AC, Fischer-Rosinsky A, Forouhi NG, Fox CS, Frants R, Franzosi MG, Galan P, Goodarzi MO, Graessler J, Groves CJ, Grundy S, Gwilliam R, Gyllensten U, Hadjadj S, Hallmans G, Hammond N, Han X, Hartikainen AL, Hassanali N, Hayward C, Heath SC, Hercberg S, Herder C, Hicks AA, Hillman DR, Hingorani AD, Hofman A, Hui J, Hung J, Isomaa B, Johnson PR, Jørgensen T, Jula A, Kaakinen M, Kaprio J, Kesaniemi YA, Kivimaki M, Knight B, Koskinen S, Kovacs P, Kyvik KO, Lathrop GM, Lawlor DA, Le Bacquer O, Lecoeur C, Li Y, Lyssenko V, Mahley R, Mangino M, Manning AK, Martínez-Larrad MT, McAteer JB, McCulloch LJ, McPherson R, Meisinger C, Melzer D, Meyre D, Mitchell BD, Morken MA, Mukherjee S, Naitza S, Narisu N, Neville MJ, Oostra BA, Orrù M, Pakyz R, Palmer CN, Paolisso G, Pattaro C, Pearson D, Peden JF, Pedersen NL, Perola M, Pfeiffer AF, Pichler I, Polasek O, Posthuma D, Potter SC, Pouta A, Province MA, Psaty BM, Rathmann W, Rayner NW, Rice K, Ripatti S, Rivadeneira F, Roden M, Rolandsson O, Sandbaek A, Sandhu M, Sanna S, Sayer AA, Scheet P, Scott LJ, Seedorf U, Sharp SJ, Shields B, Sigurethsson G, Sijbrands EJ, Silveira A, Simpson L, Singleton A, Smith NL, Sovio U, Swift A, Syddall H, Syvänen AC, Tanaka T, Thorand B, Tichet J, Tönjes A, Tuomi T, Uitterlinden AG, van Dijk KW, van Hoek M, Varma D, Visvikis-Siest S, Vitart V, Vogelzangs N, Waeber G, Wagner PJ, Walley A, Walters GB, Ward KL, Watkins H, Weedon MN, Wild SH, Willemsen G, Witteman JC, Yarnell JW, Zeggini E, Zelenika D, Zethelius B, Zhai G, Zhao JH, Zillikens MC; DIAGRAM Consortium; GIANT Consortium; Global BPgen Consortium, Borecki IB, Loos RJ, Meneton P, Magnusson PK, Nathan DM, Williams GH, Hattersley AT, Silander K, Salomaa V, Smith GD, Bornstein SR, Schwarz P, Spranger J, Karpe F, Shuldiner AR, Cooper C, Dedoussis GV, Serrano-Ríos M, Morris AD, Lind L, Palmer LJ, Hu FB, Franks PW, Ebrahim S, Marmot M, Kao WH, Pankow JS, Sampson MJ, Kuusisto J, Laakso M, Hansen T, Pedersen O, Pramstaller PP, Wichmann HE, Illig T, Rudan I, Wright AF, Stumvoll M, Campbell H, Wilson JF; Anders Hamsten on behalf of Procardis Consortium; MAGIC investigators, Bergman RN, Buchanan TA, Collins FS, Mohlke KL, Tuomilehto J, Valle TT, Altshuler D, Rotter JI, Siscovick DS, Penninx BW, Boomsma DI, Deloukas P, Spector TD, Frayling TM, Ferrucci L, Kong A, Thorsteinsdottir U, Stefansson K, van Duijn CM, Aulchenko YS, Cao A, Scuteri A, Schlessinger D, Uda M, Ruokonen A, Jarvelin MR, Waterworth DM, Vollenweider P, Peltonen L, Mooser V, Abecasis GR, Wareham NJ, Sladek R, Froguel P, Watanabe RM, Meigs JB, Groop L, Boehnke M, McCarthy MI, Florez JC, Barroso I. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet. 2010;42:105-116. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1722] [Cited by in RCA: 1721] [Article Influence: 114.7] [Reference Citation Analysis (0)] |
| 14. | Manning AK, Hivert MF, Scott RA, Grimsby JL, Bouatia-Naji N, Chen H, Rybin D, Liu CT, Bielak LF, Prokopenko I, Amin N, Barnes D, Cadby G, Hottenga JJ, Ingelsson E, Jackson AU, Johnson T, Kanoni S, Ladenvall C, Lagou V, Lahti J, Lecoeur C, Liu Y, Martinez-Larrad MT, Montasser ME, Navarro P, Perry JR, Rasmussen-Torvik LJ, Salo P, Sattar N, Shungin D, Strawbridge RJ, Tanaka T, van Duijn CM, An P, de Andrade M, Andrews JS, Aspelund T, Atalay M, Aulchenko Y, Balkau B, Bandinelli S, Beckmann JS, Beilby JP, Bellis C, Bergman RN, Blangero J, Boban M, Boehnke M, Boerwinkle E, Bonnycastle LL, Boomsma DI, Borecki IB, Böttcher Y, Bouchard C, Brunner E, Budimir D, Campbell H, Carlson O, Chines PS, Clarke R, Collins FS, Corbatón-Anchuelo A, Couper D, de Faire U, Dedoussis GV, Deloukas P, Dimitriou M, Egan JM, Eiriksdottir G, Erdos MR, Eriksson JG, Eury E, Ferrucci L, Ford I, Forouhi NG, Fox CS, Franzosi MG, Franks PW, Frayling TM, Froguel P, Galan P, de Geus E, Gigante B, Glazer NL, Goel A, Groop L, Gudnason V, Hallmans G, Hamsten A, Hansson O, Harris TB, Hayward C, Heath S, Hercberg S, Hicks AA, Hingorani A, Hofman A, Hui J, Hung J, Jarvelin MR, Jhun MA, Johnson PC, Jukema JW, Jula A, Kao WH, Kaprio J, Kardia SL, Keinanen-Kiukaanniemi S, Kivimaki M, Kolcic I, Kovacs P, Kumari M, Kuusisto J, Kyvik KO, Laakso M, Lakka T, Lannfelt L, Lathrop GM, Launer LJ, Leander K, Li G, Lind L, Lindstrom J, Lobbens S, Loos RJ, Luan J, Lyssenko V, Mägi R, Magnusson PK, Marmot M, Meneton P, Mohlke KL, Mooser V, Morken MA, Miljkovic I, Narisu N, O'Connell J, Ong KK, Oostra BA, Palmer LJ, Palotie A, Pankow JS, Peden JF, Pedersen NL, Pehlic M, Peltonen L, Penninx B, Pericic M, Perola M, Perusse L, Peyser PA, Polasek O, Pramstaller PP, Province MA, Räikkönen K, Rauramaa R, Rehnberg E, Rice K, Rotter JI, Rudan I, Ruokonen A, Saaristo T, Sabater-Lleal M, Salomaa V, Savage DB, Saxena R, Schwarz P, Seedorf U, Sennblad B, Serrano-Rios M, Shuldiner AR, Sijbrands EJ, Siscovick DS, Smit JH, Small KS, Smith NL, Smith AV, Stančáková A, Stirrups K, Stumvoll M, Sun YV, Swift AJ, Tönjes A, Tuomilehto J, Trompet S, Uitterlinden AG, Uusitupa M, Vikström M, Vitart V, Vohl MC, Voight BF, Vollenweider P, Waeber G, Waterworth DM, Watkins H, Wheeler E, Widen E, Wild SH, Willems SM, Willemsen G, Wilson JF, Witteman JC, Wright AF, Yaghootkar H, Zelenika D, Zemunik T, Zgaga L; DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium; Multiple Tissue Human Expression Resource (MUTHER) Consortium, Wareham NJ, McCarthy MI, Barroso I, Watanabe RM, Florez JC, Dupuis J, Meigs JB, Langenberg C. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat Genet. 2012;44:659-669. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 690] [Cited by in RCA: 635] [Article Influence: 48.8] [Reference Citation Analysis (0)] |
| 15. | Soranzo N, Sanna S, Wheeler E, Gieger C, Radke D, Dupuis J, Bouatia-Naji N, Langenberg C, Prokopenko I, Stolerman E, Sandhu MS, Heeney MM, Devaney JM, Reilly MP, Ricketts SL, Stewart AF, Voight BF, Willenborg C, Wright B, Altshuler D, Arking D, Balkau B, Barnes D, Boerwinkle E, Böhm B, Bonnefond A, Bonnycastle LL, Boomsma DI, Bornstein SR, Böttcher Y, Bumpstead S, Burnett-Miller MS, Campbell H, Cao A, Chambers J, Clark R, Collins FS, Coresh J, de Geus EJ, Dei M, Deloukas P, Döring A, Egan JM, Elosua R, Ferrucci L, Forouhi N, Fox CS, Franklin C, Franzosi MG, Gallina S, Goel A, Graessler J, Grallert H, Greinacher A, Hadley D, Hall A, Hamsten A, Hayward C, Heath S, Herder C, Homuth G, Hottenga JJ, Hunter-Merrill R, Illig T, Jackson AU, Jula A, Kleber M, Knouff CW, Kong A, Kooner J, Köttgen A, Kovacs P, Krohn K, Kühnel B, Kuusisto J, Laakso M, Lathrop M, Lecoeur C, Li M, Li M, Loos RJ, Luan J, Lyssenko V, Mägi R, Magnusson PK, Mälarstig A, Mangino M, Martínez-Larrad MT, März W, McArdle WL, McPherson R, Meisinger C, Meitinger T, Melander O, Mohlke KL, Mooser VE, Morken MA, Narisu N, Nathan DM, Nauck M, O'Donnell C, Oexle K, Olla N, Pankow JS, Payne F, Peden JF, Pedersen NL, Peltonen L, Perola M, Polasek O, Porcu E, Rader DJ, Rathmann W, Ripatti S, Rocheleau G, Roden M, Rudan I, Salomaa V, Saxena R, Schlessinger D, Schunkert H, Schwarz P, Seedorf U, Selvin E, Serrano-Ríos M, Shrader P, Silveira A, Siscovick D, Song K, Spector TD, Stefansson K, Steinthorsdottir V, Strachan DP, Strawbridge R, Stumvoll M, Surakka I, Swift AJ, Tanaka T, Teumer A, Thorleifsson G, Thorsteinsdottir U, Tönjes A, Usala G, Vitart V, Völzke H, Wallaschofski H, Waterworth DM, Watkins H, Wichmann HE, Wild SH, Willemsen G, Williams GH, Wilson JF, Winkelmann J, Wright AF; WTCCC, Zabena C, Zhao JH, Epstein SE, Erdmann J, Hakonarson HH, Kathiresan S, Khaw KT, Roberts R, Samani NJ, Fleming MD, Sladek R, Abecasis G, Boehnke M, Froguel P, Groop L, McCarthy MI, Kao WH, Florez JC, Uda M, Wareham NJ, Barroso I, Meigs JB. Common variants at 10 genomic loci influence hemoglobin A₁(C) levels via glycemic and nonglycemic pathways. Diabetes. 2010;59:3229-3239. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 377] [Cited by in RCA: 340] [Article Influence: 22.7] [Reference Citation Analysis (0)] |
| 16. | Hartwig FP, Davies NM, Hemani G, Davey Smith G. Two-sample Mendelian randomization: avoiding the downsides of a powerful, widely applicable but potentially fallible technique. Int J Epidemiol. 2016;45:1717-1726. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 243] [Cited by in RCA: 556] [Article Influence: 79.4] [Reference Citation Analysis (0)] |
| 17. | Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, Laurin C, Burgess S, Bowden J, Langdon R, Tan VY, Yarmolinsky J, Shihab HA, Timpson NJ, Evans DM, Relton C, Martin RM, Davey Smith G, Gaunt TR, Haycock PC. The MR-Base platform supports systematic causal inference across the human phenome. Elife. 2018;7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4747] [Cited by in RCA: 4859] [Article Influence: 694.1] [Reference Citation Analysis (0)] |
| 18. | Morgan RG. Network Mendelian Randomization Study Design to Assess Factors Mediating the Causal Link Between Telomere Length and Heart Disease. Circ Res. 2017;121:200-202. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 19. | Weisman A, Fazli GS, Johns A, Booth GL. Evolving Trends in the Epidemiology, Risk Factors, and Prevention of Type 2 Diabetes: A Review. Can J Cardiol. 2018;34:552-564. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 107] [Article Influence: 15.3] [Reference Citation Analysis (0)] |
| 20. | Marušić S, Meliš P, Lucijanić M, Grgurević I, Turčić P, Neto PRO, Bilić-Ćurčić I. Impact of pharmacotherapeutic education on medication adherence and adverse outcomes in patients with type 2 diabetes mellitus: a prospective, randomized study. Croat Med J. 2018;59:290-297. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 9] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 21. | Tan JP, Cheng KKF, Siah RC. A systematic review and meta-analysis on the effectiveness of education on medication adherence for patients with hypertension, hyperlipidaemia and diabetes. J Adv Nurs. 2019;75:2478-2494. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 69] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 22. | Yorke E, Atiase Y. Impact of structured education on glucose control and hypoglycaemia in Type-2 diabetes: a systematic review of randomized controlled trials. Ghana Med J. 2018;52:41-60. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 23. | Gage SH, Bowden J, Davey Smith G, Munafò MR. Investigating causality in associations between education and smoking: a two-sample Mendelian randomization study. Int J Epidemiol. 2018;47:1131-1140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 51] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 24. | Akter S, Goto A, Mizoue T. Smoking and the risk of type 2 diabetes in Japan: A systematic review and meta-analysis. J Epidemiol. 2017;27:553-561. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 82] [Cited by in RCA: 106] [Article Influence: 13.3] [Reference Citation Analysis (0)] |
| 25. | Xu H, Wang Q, Sun Q, Qin Y, Han A, Cao Y, Yang Q, Yang P, Lu J, Liu Q, Xiang Q. In type 2 diabetes induced by cigarette smoking, activation of p38 MAPK is involved in pancreatic β-cell apoptosis. Environ Sci Pollut Res Int. 2018;25:9817-9827. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 14] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 26. | Tillmann T, Vaucher J, Okbay A, Pikhart H, Peasey A, Kubinova R, Pajak A, Tamosiunas A, Malyutina S, Hartwig FP, Fischer K, Veronesi G, Palmer T, Bowden J, Davey Smith G, Bobak M, Holmes MV. Education and coronary heart disease: mendelian randomisation study. BMJ. 2017;358:j3542. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 121] [Article Influence: 15.1] [Reference Citation Analysis (0)] |
| 27. | Böckerman P, Viinikainen J, Pulkki-Råback L, Hakulinen C, Pitkänen N, Lehtimäki T, Pehkonen J, Raitakari OT. Does higher education protect against obesity? Prev Med. 2017;101:195-198. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 37] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 28. | Velázquez-López L, Muñoz-Torres AV, Medina-Bravo P, Vilchis-Gil J, Klϋnder-Klϋnder M, Escobedo-de la Peña J. Multimedia education program and nutrition therapy improves HbA1c, weight, and lipid profile of patients with type 2 diabetes: a randomized clinical trial. Endocrine. 2017;58:236-245. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 13] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 29. | Wadolowska L, Hamulka J, Kowalkowska J, Ulewicz N, Hoffmann M, Gornicka M, Bronkowska M, Leszczynska T, Glibowski P, Korzeniowska-Ginter R. Changes in Sedentary and Active Lifestyle, Diet Quality and Body Composition Nine Months after an Education Program in Polish Students Aged 11⁻12 Years: Report from the ABC of Healthy Eating Study. Nutrients. 2019;11. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 28] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 30. | Prospective Studies Collaboration. , Whitlock G, Lewington S, Sherliker P, Clarke R, Emberson J, Halsey J, Qizilbash N, Collins R, Peto R. Body-mass index and cause-specific mortality in 900 000 adults: collaborative analyses of 57 prospective studies. Lancet. 2009;373:1083-1096. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3582] [Cited by in RCA: 3265] [Article Influence: 204.1] [Reference Citation Analysis (0)] |
| 31. | Lyall DM, Celis-Morales C, Ward J, Iliodromiti S, Anderson JJ, Gill JMR, Smith DJ, Ntuk UE, Mackay DF, Holmes MV, Sattar N, Pell JP. Association of Body Mass Index With Cardiometabolic Disease in the UK Biobank: A Mendelian Randomization Study. JAMA Cardiol. 2017;2:882-889. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 128] [Cited by in RCA: 183] [Article Influence: 30.5] [Reference Citation Analysis (0)] |
| 32. | Zhu Z, Zheng Z, Zhang F, Wu Y, Trzaskowski M, Maier R, Robinson MR, McGrath JJ, Visscher PM, Wray NR, Yang J. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat Commun. 2018;9:224. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 402] [Cited by in RCA: 599] [Article Influence: 85.6] [Reference Citation Analysis (0)] |
| 33. | Cheng L, Zhuang H, Ju H, Yang S, Han J, Tan R, Hu Y. Exposing the Causal Effect of Body Mass Index on the Risk of Type 2 Diabetes Mellitus: A Mendelian Randomization Study. Front Genet. 2019;10:94. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 48] [Article Influence: 8.0] [Reference Citation Analysis (0)] |