Published online Jul 15, 2019. doi: 10.4239/wjd.v10.i7.376
Peer-review started: February 22, 2019
First decision: May 8, 2019
Revised: May 23, 2019
Accepted: June 11, 2019
Article in press: June 11, 2019
Published online: July 15, 2019
Processing time: 144 Days and 16.1 Hours
Type 2 diabetes (T2D) mellitus is a common complex disease that currently affects more than 400 million people worldwide and has become a global health problem. High-throughput sequencing technologies such as whole-genome and whole-exome sequencing approaches have provided numerous new insights into the molecular bases of T2D. Recent advances in the application of sequencing technologies to T2D research include, but are not limited to: (1) Fine mapping of causal rare and common genetic variants; (2) Identification of confident gene-level associations; (3) Identification of novel candidate genes by specific scoring approaches; (4) Interrogation of disease-relevant genes and pathways by transcriptional profiling and epigenome mapping techniques; and (5) Investigation of microbial community alterations in patients with T2D. In this work we review these advances in application of next-generation sequencing methods for elucidation of T2D pathogenesis, as well as progress and challenges in implementation of this new knowledge about T2D genetics in diagnosis, prevention, and treatment of the disease.
Core tip: Next-generation sequencing (NGS) technologies have a broad range of applications in studying the genetic causes of type 2 diabetes (T2D), such as: (1) Identification of rare and common genetic variants, associated with disease; (2) Functional studies for describing role of genes in disease pathogenesis; and (3) Evaluation of environmental contribution to the disease by using microbiome profiling methods. This review of NGS application to the genetic research of T2D presents the advances and challenges related with sequencing analysis-based studies and implementation of this knowledge in clinical practice.
- Citation: Nasykhova YA, Barbitoff YA, Serebryakova EA, Katserov DS, Glotov AS. Recent advances and perspectives in next generation sequencing application to the genetic research of type 2 diabetes. World J Diabetes 2019; 10(7): 376-395
- URL: https://www.wjgnet.com/1948-9358/full/v10/i7/376.htm
- DOI: https://dx.doi.org/10.4239/wjd.v10.i7.376
Type 2 diabetes (T2D) mellitus is a common complex disease that currently affects more than 400 million people throughout the world, and it is projected 552 million cases of T2D by the year 2030[1]. The disease is characterized by insulin resistance and beta-cell dysfunction and can seriously impair overall quality of life[2]. T2D may lead to increased risk of cardiovascular disease, stroke, kidney failure and can result in lower life expectancy by 5-10 years[3-5]. T2D etiology is known to have a significant genetic component that is confirmed by family- and twin-based studies. The risk of the disease developing is approximately 70% when both parents have T2D and approximately 40% when one parent has disease[6]. Twin studies have shown that the heritability of T2D ranges from 26% to 73%, and the concordance rate for T2D in monozygotic twins can reach 76%[7]. Early identification of individuals at high T2D risk enables delay or prevention of T2D onset through effective lifestyle and/or pharma-cological interventions and has been shown to reduce costs of healthcare that causes continuing strong interest in revealing risk markers of T2D[8,9].
The development of high-throughput and affordable genotyping technologies, statistical tools and computational software has allowed remarkable progress over the past decade in the search for genetic associations. Since the first genome-wide association study (GWAS) for T2D identified novel susceptibility loci in 2007, more than 100 T2D susceptibility loci have been discovered[10]. Next-generation sequencing (NGS) technologies have a broad range of applications in studying the genetic causes of T2D, such as: (1) Identification of rare and common genetic variants, associated with disease; (2) Functional studies for describing role of genes in disease patho-genesis; and (3) Evaluation of environmental contribution to the disease by using microbiome profiling methods. However, it remains uncertain if and to what extent our increasing knowledge of genetic and epigenetic T2D risk factors gained by NGS methods will translate into clinical practice.
The aim of this article is to summarize recent progress and discoveries for T2D genetics focusing on the sequencing analysis-based studies and review the challenges in studying the genetic basis of T2D in order to improve diagnosis, prevention, and treatment.
The earliest genetic studies of T2D susceptibility focused on family-based linkage analysis and analysis of candidate genes in small-size groups of patients. This approach was successful in identifying familial genetic variants with large effects such as those involved in monogenic forms of the disease. In the past two decades, numerous candidate gene studies have been performed to identify genetic variants for T2D. However only 4 genetic markers identified in these studies have been confirmed later by GWAS. The first genetic variant for T2D was the P12A polymorphism (rs1801282) in peroxisomal proliferator activated-receptor gamma gene (PPARG)[11]. Then, in 2003, in a large-scale association study the previously identified association between the E23K (rs5219) polymorphism in a gene encoding inwardly rectifying potassium channel subfamily J, member 11 (KCNJ11) and T2D was replicated[12]. E23K can alter function by inducing spontaneous over-activity of pancreatic β-cells, thus increasing the threshold ATP concentration for insulin release[13]. In previous studies a polymorphism in this genes (KCNJ11 E23K) has been reported to be associated with T2D in several populations, although the data was inconsistent[14-18]. Transcription factor 7-like 2 (T-cell specific, HMG-box) (TCF7L2) was shown to be associated with T2D[19]. TCF7L2 gene product is a member of the high mobility group box family of transcription factors, activated by the WNT signaling pathway and may play a master role in regulating insulin biosynthesis, secretion, and processing. Subsequently, two single nucleotide polymorphisms (SNPs) within intron 3 of TCF7L2, rs7903146 and rs12255372, were confirmed to be strongly associated with T2D risk[20-22]. Wolfram syndrome 1 gene (wolframin) (WFS1) was reported to be associated with T2D on the basis of in-depth studies of candidate genes[23]. The WFS1 gene encodes wolframin, endoplasmic reticulum (ER) membrane protein with a role in ER calcium homeostasis. Mutations in WFS1 are known to be associated with Wolfram syndrome[24].
Advances in technology of SNP genotyping, implementation of recent genetic know-ledge gained from the Human Genome Project, and development of robust statistical methods have allowed GWAS to become the basic method for identification of common genetic variants associated with complex diseases such as T2D. Since the application of GWAS technology the discovery of genetic variants associated with T2D has developed dramatically.
In 2007, the first GWAS performed for T2D has identified three novel susceptibility loci related to pancreatic β-cells: (1) Solute carrier family 30 (zinc transporter), member 8 (SLC30A8), which is expressed exclusively in insulin-producing β-cells; (2) Insulin-degrading enzyme (IDE)–kinesin-interacting factor 11 (KIF11)–hemato-poietically expressed homeobox (HHEX); and (3) Exostosin glycosyltransferase 2 (EXT2)–ALX homeobox 4 (ALX4)[25]. Subsequent GWAS revealed four additional loci associated with T2D, namely CDK5 regulatory subunit associated protein 1-like 1 (CDKAL1), cyclin-dependent kinase inhibitor 2A (CDKN2A/B), insulin-like growth factor 2 mRNA binding protein 2 (IGF2BP2), and fat mass and obesity associated (FTO)[26-30]. In addition, HNF1 homeobox B (HNF1B/ TCF2), a gene related to maturity-onset diabetes of the young type 5 (MODY5), was shown to be associated with T2D[31]. One important finding from the initial GWAS results was that effect sizes for common variants involved in T2D were likely to be modest. The statistical power to detect associations between genetic variants and a trait depends on the sample size, the distribution of effect sizes of (unknown) causal genetic variants, the frequency of those variants, and the linkage disequilibrium (LD) between observed genotyped DNA variants and the unknown causal variants[32]. This led to an innovative data merging strategy now known as GWAS meta-analysis and resulted in multiple waves of GWAS studies for T2D.
In 2008, six new T2D loci including JAZF1, CDC123/calcium/CAMK1D, TSPAN8/ LGR5, THADA, ADAMTS9, and NOTCH2 were reported by a meta-analysis combining three previous GWAS [Diabetes Genetic Initiative (DGI), Finland–United States Investigation of NIDDM Genetics (FUSION), and Wellcome Trust Case Control Consortium (WTCCC)][33]. In 2009, two loci, namely insulin receptor substrate 1 (IRS1) and melatonin receptor 1B (MTNR1B) were identified to be associated with T2D by GWAS[34-36]. The IRS1 gene is related to insulin resistance and hyperinsulinemia, whereas MTNR1B is involved in impaired early insulin response to glucose[35].
In 2010 the second wave of the GWAS identified 17 new loci associated with T2D which was made possible because of improved efficiency of GWAS genotyping technology, enabling interrogation of larger numbers of SNPs that better cover common genetic variation across populations in increased sample sizes, as well as because of methodological innovations, such as imputation (described below), which allows prediction of genotypes at SNPs not typed on GWAS arrays[37].
In the past year a leap forward has occurred from smaller, cumulative advances to the description of up to around 250 genome-wide significant loci of T2D[10]. In this work, a large meta-analysis of GWAS in sample of T2D including 62892 cases and 596424 controls was performed by combining 3 GWAS data sets of European ancestry: DIAbetes Genetics Replication and Meta-analysis (DIAGRAM), Genetic Epidemiology Research on Aging, and the full cohort release of the UK Biobank 39 previously unknown loci have been identified[38]. This study highlighted the benefits of integrating multiple omics data to identify functional genes and putative regulatory mechanisms caused by genetic variation. Future applications of integrative omics data analyses are expected to improve our understanding of the biological mechanisms underlying common diseases such as T2D[38].
While conventional genome-wide association studies allow to identify associated loci, GWAS alone cannot be used to map causal variants (many of which are expectedly rare in population), as the method strictly focuses on pre-selected common variants identified by the HapMap project in the beginning of the century[39]. On the other hand, NGS presents a reasonable alternative to the chip-based methods. For genotyp-ing purposes, NGS reads are aligned to a reference genome, and a set of statistical procedures is performed to identify variant sites[40]. Thus, NGS directly identifies most of the genetic variants present in an individual’s genome irrespective of their frequency, which enables testing of all variants’ association. In this section, we will focus on how NGS datasets might be used for identification of novel causal variants for T2D, and which loci have been identified by these methods.
Large genome and exome sequencing and aggregation consortia, such as the 1000 Genomes project or UK10K provide valuable insights into linkage disequilibrium, i.e., co-occurrence rates, between different variants, enabling probabilistic reconstruction of individual genome sequences from fixed number of genotyped loci (such as in traditional GWAS). This in turn enables testing for the role of rare variants without sequencing per se[37,41,42]. Large reference panels for such genotype imputation have been constructed from sequencing data[43]. Genotype imputation has been widely used in the studies of the genetic architecture of T2D[44]. An interesting example is a 2014 study of Icelandic population[45]. In this work, whole-genome sequencing study of a cohort of 2630 Icelanders was performed; and the identified SNPs and indels were imputed into 98721 controls and T2D patients genotyped with Illumina SNP chips. As a result of this study a rare variant in HNF1A gene, encoding for a transcription factor required for the expression of several liver-specific genes was identified. Moreover, a new signal with association P < 1 × 10−8 at rs76895963, located within the first intron of cyclin D2 (CCND2) was observed[45]. Two of the most recent and comprehensive research efforts aimed at fine mapping of association signal using imputation and islet-specific epigenome maps identified multiple previously unreported loci for T2D, including PNPLA3, LPL, TPCN2, DENND2C, and KIF2B[46,47]. Apart from using NGS datasets for rare variant imputation, different approaches based on combined SNP and exome chip methods have been developed, enhancing the power of imputation-based analyses[48].
As previously stated, many new genetic associations relevant to T2D have been revealed by GWASs, but these findings represent common and mid-frequency genetic variants with small effect sizes and explain only a small proportion of heritability of the disease. Sequencing approach enables more complete assessments of low-frequency and rare genetic variants that can be promising in investigation of complex traits.
Many published studies have focused on identification of T2D susceptibility loci from NGS data. In Danish study, the exomes of 1974 Danes were sequenced to a depth of 8 × and subsequently a two-stage follow-up in 15989 Danes and in a further 63896 Europeans were performed. A low-frequency coding variant in CD300LG associated with fasting HDL-cholesterol and two common coding variants in COBLL1 and MACF1 have been shown to be associated with T2D[49]. CD300LG encodes a protein proposed to serve multiple functions, including endocytosis of various immunoglobulins and mediation of L-selectin-dependent lymphocyte rolling[50,51]. Non-coding SNPs in COBLL1 and MACF1 have previously been associated with other metabolic phenotypes[52-54].
To investigate the hypothesis of “missing heritability”, the Genetics of Type 2 Diabetes and Type 2 Diabetes Genetic Exploration by Next-generation sequencing in multi-Ethnic Samples Consortium (GoT2D/T2D-GENES Consortium) undertook whole genome sequencing in 2657 Europeans with and without diabetes, and exome sequencing in a total of 12940 subjects from five ancestral groups. Results of this study showed that the variants associated with T2D were overwhelmingly common and most located within regions previously identified by GWAS. A few coding variant associations outside established common variant GWAS regions have been identified (rs41278853 in MTMR3 gene; rs11549795, rs28265, rs36571 in ASCC2 gene). A coding variant reached genome-wide significance that was common in East Asian ancestry population (PAX4 Arg192His, rs2233580)[55]. PAX4 gene encodes a transcription factor involved in islet differentiation and function. Some PAX4 variants have been associ-ated with early-onset monogenic diabetes[56,57].
Despite decreasing costs of NGS-based analyses, there still remain certain notable limitations of such studies. The most evident limitation of all the rare variant-based tests on both whole-genome and imputed SNP array datasets is the difficulty of obtaining enough observations to make confident statistical inference. For example, if a causal variant occurs at a rate of 10-4 in a population, one would require many hundreds of thousands of individuals to test its association with the disease. To allow testing for the association of rare variants, especially in smaller samples, a group of techniques were developed, called Rare Variant Association tests. Most of rare-variant tests are designed to identify candidate disease genes through aggregation of all rare variants inside the coding sequence of each gene. Numerous strategies for gene-level testing of rare-variant association have been developed[58]. The two main groups of such methods test either the imbalance of rare allele counts between cases and controls (burden tests) or the proportion of phenotypic variance explained by rare variant genotypes (variance-component tests). However, for T2D almost few significant gene-level associations have been found even in the largest NGS-based population cohorts[55,59]. Only the largest study performed by whole-exome sequencing to date, which included 20791 T2D cases and 24440 controls of multiple ancestries (Hispanic/Latino, European, African-American, East-Asian, South-Asian), identified several gene-level associations: in 3 genes at exome-wide significance, including a T2D protective series of > 30 SLC30A8 alleles, and within 12 gene sets, including those corresponding to T2D drug targets and candidate genes from knockout mice. The strongest T2D rare variant gene-level signals was shown to explain at most 25% of the heritability of the strongest common single variant signals[60].
Several alternative techniques have been developed to overcome the limitations of rare variant testing. In samples of limited size based on exome sequencing or targeted resequencing, contribution of rare variants might be assessed using tests for case-specificity conditioned on true population minor allele frequency[61]. Such strategy may help to identify variants that serve as the candidate causal markers for the pathology. In a recent study by our group, we identified potential association for the VAV3, ADAMTS13, HBQ1, and DBH genes with T2D and obesity. While these genes have not been previously implicated in the disease, they are reasonable targets for further clinical investigation.
Another approach to counteract statistical power limitation of rare-variant based tests in small NGS-based datasets is the usage of pedigrees. The biggest advantage of familial studies is that cohorts of related individuals would have higher frequency of alleles that are rare in the general population. One recent example of pedigree-based analysis is a study of 20 Mexican-American families comprising 1034 highly related individuals[62]. While this study still did not identify any significant associations for individual rare variants, it has shown gene-level association for the CYP3A4 and OR2T11 genes with glycemic traits, such as fasting glucose levels and 2h insulin levels.
Overall, there are several ways in which NGS might be used to assist identification of causal genes and variants for T2D pathology. These associations are of ultimate relevance for genomic risk prediction of T2D and clinical decision making[63]. Some of the inherent limitation of the technology, however, still do not allow thorough analysis of chromosome- and genome-level genetic variation and/or complex genome regions that are poorly accessible to short read sequencing[64-66]. The spread of third-generation sequencing technologies, such as the Oxford Nanopore Technologies single-molecule sequencing, as well as modifications to the existing laboratory and/or bioinformatic practices would shed light on the roles of higher-level genetic variants in T2D pathology.
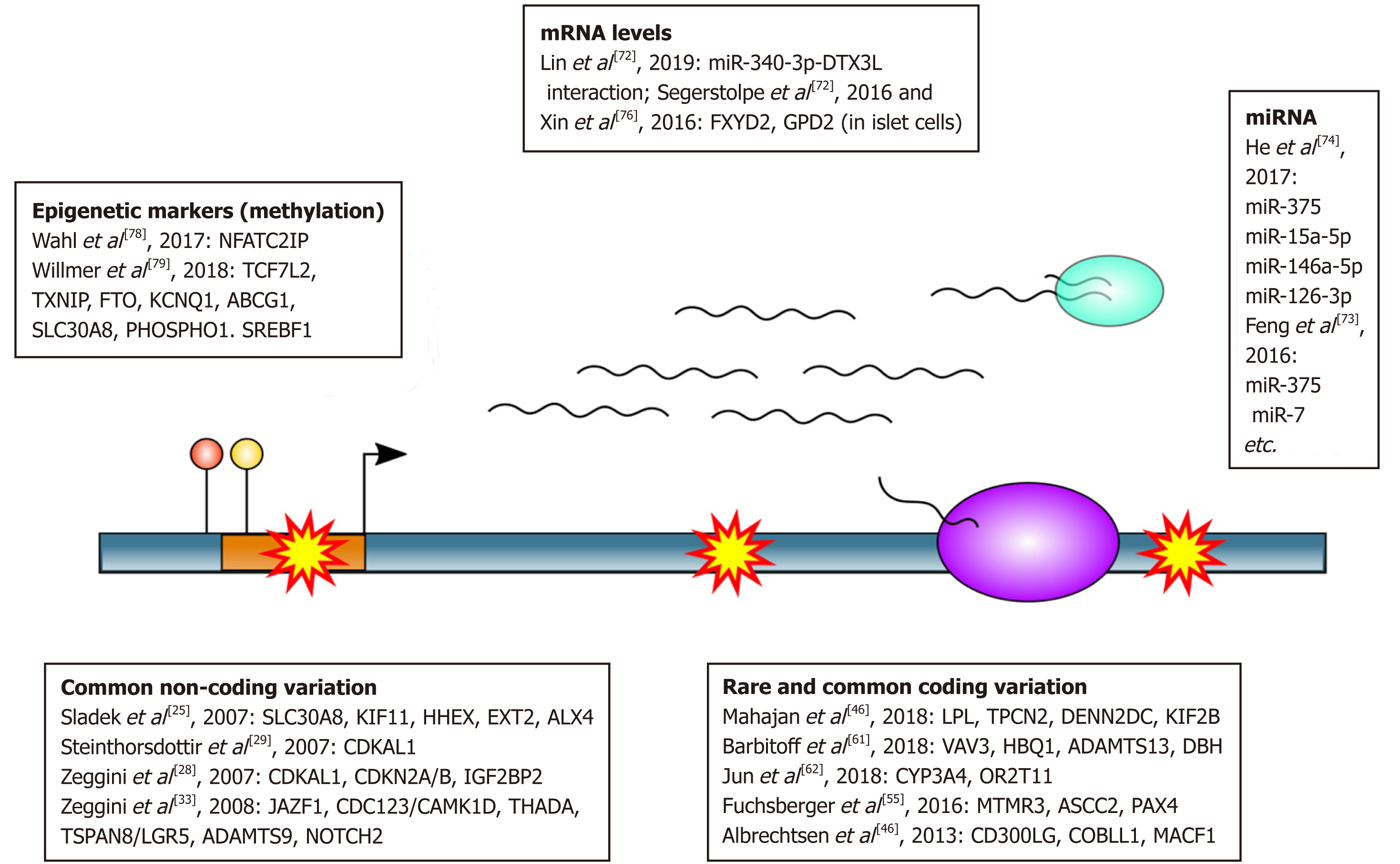
Apart from methods aimed at genotyping, NGS can also be used to dissect functional genome elements rather than sequence variants. NGS techniques for these purposes include transcriptional profiling approaches (RNA-Seq), epigenome mapping techniques (positional methods), and other[67,68]. These methods are commonly used to both identify candidate disease genes and understand pathological mechanisms behind the observed phenotype. Below, we will provide several recent examples of application of these methods to the research of T2D (Figure 1).
Transcriptional profiling methods, such as RNA-Seq, are used to study activity patterns of genes. In the recent decade, transcriptomic technologies were frequently used to decipher the molecular pathology behind human disease[69]. T2D, being one of the most common pathologies, has also been extensively studied by transcriptional profiling techniques in the recent decade[70]. Traditional way to analyze RNA-Seq data is to align the reads to a reference genome and count the numbers of reads or fragm-ents mapped to each gene or transcript. These counts are then used to search for genes which significantly change their expression in case vs controls (differentially expressed genes, DEGs) using conventional statistical tests or linear regression models, and identify biological processes which are dysregulated in one of the conditions. The latter task is solved by a family of gene set enrichment tests that analyze overrepresentation of genes from a certain pathway among the identified DEGs. Multiple downstream analyses can be performed to identify disease genes and pathways from both bulk and single-cell RNA-Seq data[71]. Below, we will focus on several notable examples of how both bulk and single-cell technologies can be used to identify genes involved in pathological mechanisms of T2D.
One example of a conventional bulk RNA-Seq approach used to identify disease-relevant pathways can be found in a recent work that studied transcriptional profiles of diabetic keratinocytes[72]. This study showed extensive dysregulation of immunity-related genes in these cells compared to controls, with as many as 420 differentially expressed genes identified in total. Moreover, this study has suggested a causal role of miR-340-3p-DTX3L interaction in the pathological processes occurring in diabetic skin.
Multiple studies have also focused on the roles of microRNA (miRNA) in the pathology of T2D[73]. microRNAs are a separate class of RNA molecules which play an important role in gene regulation via post-transcriptional gene silencing. One of the most recent studies aiming at systematic analysis of microRNA involvement in T2D by aggregation of published data identified as many 158 microRNAs reported to be differentially expressed in T2D. One example of an important microRNA identified in this study is the miR-375 RNA which affects expression of several disease-relevant genes in islets and other tissues.
Many studies suggest that the alterations in miRNA levels are associated with T2D development and its complications. miRNA may play a key role in regulation of the processes of carbohydrate and lipid metabolisms, adipocytokine and insulin signaling pathways involved in T2D development. It was shown that the dysregulated in the islets miR-7-5p, -129-3p, -136-5p,-187-3p, -224-5p, -369-5p, -375 -495-3p, -589-5p, -655-3p affect the expression of important genes involved in insulin signaling pathway. The altered level of miRNA miR-17-5p, -155-5p, -125b-5p, -30e-5p, -27a-5p, -221-3p, -199a-5p, -130b-3p, -181a-5p, -29a, -29b can cause the dysregulation of lipid and glucose metabolisms. For miR-130b-3p, -140-5p, -147a, -199a-5p, -27b, -221-3p and -30e-5p) their involvement in the regulation of adipogenesis was identified[74]. Stability of miRNAs, their presence in various body fluids and significant changes of specific circulating miRNAs’ concentrations associated with diseases allow studying them as potential reliable biomarkers for complex diseases such as T2D and related complica-tions. However, there are some obstacles for straightforward clinical application of circulating miRNAs. The biggest difficulty is due to the composition of circulating miRNA that are sum of many different tissues and cell types in the body. At the same time, it is well known that the expression of miRNAs varies considerably between different tissues.
Another important branch of NGS-based transcriptional profiling techniques is the single-cell RNA sequencing (scRNA-Seq) which allows researchers to study transcriptional responses of individual cells and cell-types. scRNA-Seq techniques are also being extensively used to identify key disease genes for T2D in pancreas cells. For example, scRNA-Seq of pancreatic islets suggested a role of FXYD2 and GPD2 genes in pathological processes behind T2D in certain islet cell types, with as many as 245 dysregulated genes in total[75,76].
Another widely used group of NGS methods is aimed at understanding the language of epigenetic marks, i.e., non-DNA based units of genetic information. NGS technolo-gies for epigenome studies include but are not limited to: (1) Methods for detection of specific DNA-protein interaction (e.g., Chromatin Immuno Precipitation followed by Sequencing); (2) Methods for identification of DNA methylation sites (such as reduced representation bisulfite sequencing); and (3) Open chromatin mapping technologies (e.g., DNAse-Seq or ATAC-Seq)[67]. All of these methods provide valuable insights into dysregulation of cellular processes, which is of ultimate importance for T2D pathology[77]. Epigenetic marks, as the dynamic features of the cell, are frequently considered as convenient biomarkers for disease risk prediction and prognosis in the clinic. A large-scale survey on the adverse outcome of adiposity showed that methyla-tion pattern at certain loci predicts development of T2D in overweight people[78]. A recent analysis of published data identified 8 differentially methylated genes as potential blood biomarkers of T2D (TCF7L2, KCNQ1, ABCG1, TXNIP, PHOSPHO1, SREBF1, SLC30A8, and FTO)[79]. Epigenome profiles might also be used to enhance identification of causal variants at complex GWAS loci[80].
Overall, RNA-Seq and positional NGS techniques provide a very useful framework to investigate cellular processes that are affected during disease pathogenesis. These data may in turn be used for both prediction of diabetes risk and for designing clinical treatment of the disease; furthermore, simultaneous consideration of genotype, expression profile and epigenetic factors might assist efficient personalized treatment of T2D. Further integration of multiple omics datasets would allow researchers and clinicians to have a comprehensive look into the molecular pathology behind T2D.
Rapid progress of NGS technologies and bioinformatic data processing methods led to the advent of metagenome studies, i.e., investigation of the microbial composition of natural inhabitants. A decade of advances in the field of intestinal microbiome analysis demonstrated that alterations of gut bacteria composition is implicated in a few medical conditions, including diabetes and obesity[81-83]. Such progress can be attributed to a number of factors, for example, stable decrease of price per single run for NGS platforms, continuous development of bioinformatic tool/pipelines[84-87], creation of specialized gut microbiome 16s rRNA databases and use of metaproteo-mics, metabolomics and metatranscriptomics in conjuncture with genetic profiling[88-92]. Still, there is no consensus concerning optimal conditions for conducting microbiome research. Choice between 16S RNA profiling/shotgun sequencing methods , differences in effective coverage between V1-V9 hypervariable regions , more precise quantitative analysis for microbiota constituents[93], and generalized protocols for sample acquisition are still in discussion, with main emphasise often being put on low reproducibility of results, partly due to the unstable nature of samples’ bacterial composition[81,86,87,92,94]. Overall, intestinal microbiome genetic profiling may find use in clinical practice with development of presently elusive “golden standard” for this research field, leading to better understanding of gut microbiota’s role in human homeostasis and associations with diseases[95].
As of 2014, microbial community of human gut was estimated to contain at least 957 bacterial genera with phyla Actinobacteria, Bacteroides, Firmicutes, Proteobacteria and Verrucomicrobia demonstrating most diversity and abundance[96]. While both types of diabetes mellitus are known to cause significant changes in gastrointestinal microbial composition, underlying mechanisms for dysbiosis and roles of all microbiome constituents, including bacteria, archaea, eukaryota and fungi, are still not fully understood. Roseburia intestinalis, Faecalibacterium prausnitzii, and families Ruminococcaceae/Lachnospiraceae, all known as butyrate producers, were detected to be lower during T2D[97,98]. Abundance of Akkermansia muciniphila, a mucin-degrading primarily mucosal bacteria, had been connected to lower insulin resistance, while their low concentrations were associated with obesity, diabetes, IBD, ulcerative colitis and appendicitis, suggesting future use of this bacteria as a biomarker[99]. However, such broad spectrum of diseases makes effective clinical usage questionable. Prevotellacopri and Bacteroides vulgatus were mentioned as possible promoters for insulin resistance due to active branched chain amino acids (BCAA) production[100]. Data on general Firmicutes/Bacteroidetes ratio changes during prediabetes and T2D are contradicting, which may be explained by differences in sequencing methods and bioinformatics approaches[100,101]. Recent 16S/18S/ITS microbiome profiling study of T2D with 49 adult participants in India showed interesting correlation for archaea, where concentration of Methanobrevibacter increased in direction from healthy subjects to fully developed T2D, while Methanosphaera concentration gradually decreased. Fungal component demonstrated overall abundance growth with inclusion of pathogenic Aspergillus and Candida phyla[98]. Most of aforementioned microorganisms were proposed as possible indicators for prediabetes, T1D (type 1 diabetes) and T2D, but their use in clinical practice is not recommended at the moment due to low amou-nt of data and contradictory nature of results between studies, which may be solved in the future[86].
Both T2D and obesity demonstrate a growing trend across the globe, with subjects suffering from the latter being often viewed as possible T2D risk group[102,103]. Recent findings in the field of microbiome variation during diabetes and obesity had reaffirmed earlier theories concerning microbiota’s participation in adipose tissue function and insulin resistance. Network-based gene expression association studies of host’s genome underline digestive metabolism, immunization, and signal transdu-ction as the most prominent mechanisms in development of obesity/T2D[104], while the data on gastrointestional microbiome role is yet to be unified in coherent system. Gut microbiota had been shown to regulate body mass in a set of fecal transplantation experiments conducted on lean, obese and germ-free mice. Transplantation of gastrointestional microbiota from lean to obese mice led to lower insulin resistance, while transfer of microbiota from obese to lean mice led to body mass increase by 60% and higher insulin resistance[83,104,105]. Low grade inflammation, acquired through activation of TLR4/MyD88/NF-κB pathway by lipopolysaccharides from gram-negative bacterial walls, had been connected to insulin resistance through insulin receptor substrate serine phosphorylation by participants of inflammatory cascade[106]. Inhibition of NF-κB led to increase of Akkermansia/Lactobacillus, reduced body mass and lower insulin tolerance[100,107]. Short chain fatty acids (SCFA), obtained by bacteria through fermentation of non-digestible fibers, serve as signaling molecules in a broad list of processes, including proliferation of pancreatic β cells and insulin biosynthesis. This partially explains prebiotic treatment effectiveness and changes in abundance of Roseburia intestinalis and Faecalibacterium prausnitzii, but further research is required, as results from different studies often contradict each other[100,108,109]. High serum levels of BCAAare attributed to both obesity and T2D with steady increase of Prevotella copri and Bacteroides vulgatus during the onset of the diseases[110]. Both probiotics and prebiotics tend to increase insulin sensitivity and lower body mass, although studies have small sample sizes and require longitudinal research[111,112].
Recent findings demonstrate that effectiveness of metformin, most prescribed antidiabetic drug whose pharmacodynamics mainly involve activation of hepatic AMP-activated protein kinase in liver, may be partially attributed to mediation of diabetic dysbiosis. Increase of Akkermansia muciniphila abundance after metformin treatment was detected in both human and animal studies, while in vitro conditions in gut simulator demonstrated metformin as a growth factor for both Akkermansia muciniphila and Bifidobacterium adolescentis[113,114]. Metformin therapy was found to promote growth of SCFA-producing bacteria in rats (Allobaculum, Bacteroides, Blautia, Butyricoccus, Lactobacillus, Akkermansia and Phascolarctobacterium) and humans (Akkermansia, Lactobacillus, Bifidobacterium, Prevotella, Megasphaera, Shewanella, Blautia or Butyrivibrio)[113].
The identification of multiple loci by GWAS and sequencing technologies has given a considerable impetus to the disclosure of pathogenesis of T2D and provides a tempting opportunity to translate genetic information to clinical practice. This knowledge may have potential role in disease risk prediction including identification of subjects at risk of developing disease at an early-stage, and in clinical management of individuals to modify treatment regimens so that affected individuals would benefit most by their therapy and avoid the occurrence of complications[63]. The emerging availability of genomic and electronic health data in large populations is a powerful tool for research that has drawn interest in bringing precision medicine to diabetes[115].
According to the latest polls people are interested in genetic testing for T2D risk since this allows them to evaluate the individual feature of pathology state[116]. However, several studies have shown that some factors contribute to the failure of individuals to conduct a genetic test. The main factors that influence refusal include distrust of medical researchers, religious prejudices and lower levels of education[117,118]. Some have argued that the clinical significance of genetic markers of T2D have only a minor role in predicting the risk with careful clinical risk assessment, the predictive value increases[116,119].
Until recently, it has been assumed that genetic predisposition awareness can motivate healthy behavior[120]. According to some authors, it is considered that the patient does not appear motivated to a healthy lifestyle after identifying his genetic predisposition[121-123]. At the same time, research on the molecular basis of the development of T2D is absolutely necessary when making a diagnosis, since young individuals with T1D can also be obese[124,125]. Misdiagnosis of diabetes can lead to misuse of medical treatment[126].
Many studies have analyzed the utility of genetic variants in T2D risk prediction for undiagnosed individuals with T2D using cross-sectional studies and incident T2D using longitudinal studies. Early studies provided much optimism and showed that common variants at the TCF7L2 locus predict the progression to diabetes in subjects with impaired glucose tolerance[63,127]. Unfortunately, diabetes mellitus is diagnosed on the basis of its biochemical effects (increased glucose), and the absence of detection of the main defect, which indicates the absence of the disease[128]. However, at present, aggregated available data do not provide robust evidence to support the utility of genetic testing for T2D predictions and indicate a modest contribution of genetic variants[129-131]. Several large population-based follow-up studies have been published aiming to investigate the predictive power of common genetic variants on the risk of incident T2D. The results of these studies were similar to those from cross-sectional case-control studies. It was shown that risk variants did not essentially increase the AUC to predict T2D when combined with clinical risk factors[132]. However, it seems possible to improve T2D risk prediction and overcome factors limiting predictive power, such as: (1) Modest effect sizes of common variants, (2) Insufficient knowledge of rare and coding variants missed by GWAS; (3) Heterogeneous nature of the dise-ase; and (4) Genetic diversity between ethnic groups (detailed below).
The limitations related with modest effect sizes of common alleles and necessity of further investigation aimed to identify rare and coding variants involved in T2D pathogenesis have been reviewed above. T2D seemingly encompasses a group of several subtypes of diseases, which makes it rather difficult to distinguish it from other types, as it may be the result of defects in various metabolic pathways. The accuracy of prediction models may be affected by the fact that latent autoimmune diabetes in adults has been identified and the number of monogenic forms of diabetes is increasing, which can also indicate the level of misclassification[133].
In different populations, heterogeneity in association of genetic variants with the disease was demonstrated, apparently related to the design of the study, in particular the results of a large meta-analysis that combines cases of T2D with different origins or signs and evaluates them with a generalized intermediate hyperglycemia phenoty-pe, despite the fact that the phenotype may differ due to a multitude of unrelated causes within the physiology of the body or the environment[134]. In recent years, a large number of projects have been carried out to study the causes of diabetes, large-scale studies have been created and huge biobanks of samples of these patients have been collected. In addition, some variants were found that are important in the prevention and treatment of T2D, found in individual population isolates, demons-trating the value of studying genetically isolated populations[128]. Because of genetic drift, deleterious variants with large phenotypic effects could rise randomly to higher allele frequencies. Which makes investigation of such variants’ association easier in isolated populations compared to the admixed ones, in which these variants might not be present or might be very rare[10].
Circulating miRNAs in plasma or serum have several features that make them ideal candidate biomarkers of complex diseases such as T2D[135]. Hundreds of miRNAs are actively or passively released to the blood circulation to regulate specific gene func-tion[136]. Current studies demonstrate that changes in expression miRNAs involve in dysfunction of insulin and progression of T2D. Many studies confirmed that some miRNAs have been identified and found to be associated with T2D[137]. miR-21, miR-126 and miR146a have been shown to have potential to be biomarkers of early diagnosis of T2D disease[138-140]. Thus, the above mentioned miRNAs and a number of other miRNAs may be candidates for testing the effectiveness of therapy but further studies are needed to identify them[137].
T2D commonly develops with insulin resistance, a disorder in which cells located primarily within the muscles, liver, and fat tissue do not use insulin properly, and progresses to pancreatic beta-cell failure. T2D trigger are insulin resistance and inadequate insulin secretion[141].
Selection of drug therapy based on the genetic features of the individual can be a huge breakthrough because there are individual drug idiosyncrasy and many patients eventually fail to achieve recommended levels of glycemic control due to their genetic characteristics[142,143]. Currently, only half of patients initiating therapy with metformin or sulfonylurea, reached a level of hemoglobin A1c in 7%[144]. It should be emphasized that sulfonylureas and metformin are the most studied classes of drugs used to treat T2D[115].
Sulphonylureas (SUs) are widely used drugs in the clinical practice however, diffe-rent side effects, such as weight gain and increased risk of hypoglycemia, have been frequently[145]. Studies have shown that these drugs can act effectively in response to a defect induced by variants in KCNJ11 (rs5219, rs5215) and ABCC8 (rs757110) in patients with T2D[146,147]. Also important in the selection of SUs play role CYP2C9 (rs1799853, rs1057910), TCF7L2 (rs12255372, rs7903146), IRS1 (rs2943641, rs1801278) and CAPN10 (rs3842570, rs3792267, rs5030952)[148-151]. It should also be noted rs7754840 in the gene CDKAL1, which is significantly associated with the response to treatment with sulfonylurea and in combination with other clinical and pathological data will help move to individual therapy of patients with T2D[152].
Metformin is the most commonly used drug in the treatment of T2D, which is not metabolized in the liver, therefore, the effect of reducing the level of metformin is not affected by genetic variants in the genes encoding metabolizing enzymes[153]. SLC22A1 (rs12208357, rs34130495, rs35167514, rs34059508) is the most studied gene that is involved in the response to metformin[154]. However, other genes involved in the metabolism of metformin have been identified, for example, SLC22A2 (rs316019), PPARG (rs1801282)[145,155]. It should also be noted that the T2D–associated variant rs7903146 in TCF7L2 influences the acute response to both glipizide and metformin in persons free of overt diabetes[156].
The growing power and reducing cost sparked an enormous range of applications of NGS technology that gave us the excellent instrument for solving various problems in molecular biology. Rational usage of this instrument, taking into account all of its benefits and limitations, is the next step on the way to elucidation of pathogenesis of complex diseases such as T2D. Results obtained in sequencing-based studies combined with earlier findings from GWAS and candidate genes studies allow ordering and improving our knowledge about T2D and give us an opportunity to translate genetic information to clinical practice. The increasing knowledge provides a fascinating opportunity to use this information to predict the occurrence of disease and to identify subgroups of patients for whom therapies will have the greatest efficacy or the least adverse effect. However, this new knowledge should be treated with caution. Unfortunately, the accuracy of risk prediction models based on genetic information of T2D is not remarkable to date. Hence, further research and techno-logical improvement is needed in studying the individual and aggregate contribution of genetic markers for the development of diabetes for widespread use in clinical practice.
Manuscript source: Invited manuscript
Specialty type: Endocrinology and metabolism
Country of origin: Russia
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Al-Gayyar M, Ramos S S-Editor: Ji FF L-Editor: A E-Editor: Wang J
| 1. | Whiting DR, Guariguata L, Weil C, Shaw J. IDF diabetes atlas: global estimates of the prevalence of diabetes for 2011 and 2030. Diabetes Res Clin Pract. 2011;94:311-321. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2650] [Cited by in RCA: 2678] [Article Influence: 191.3] [Reference Citation Analysis (2)] |
| 2. | Kahn SE, Cooper ME, Del Prato S. Pathophysiology and treatment of type 2 diabetes: perspectives on the past, present, and future. Lancet. 2014;383:1068-1083. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1110] [Cited by in RCA: 1111] [Article Influence: 101.0] [Reference Citation Analysis (0)] |
| 3. | Fowler MJ. Microvascular and macrovascular complications of diabetes. Clin Diabetes. 2008;26:77-82. [RCA] [DOI] [Full Text] [Cited by in Crossref: 947] [Cited by in RCA: 1057] [Article Influence: 62.2] [Reference Citation Analysis (0)] |
| 4. | Rao Kondapally Seshasai S, Kaptoge S, Thompson A, Di Angelantonio E, Gao P, Sarwar N, Whincup PH, Mukamal KJ, Gillum RF, Holme I, Njølstad I, Fletcher A, Nilsson P, Lewington S, Collins R, Gudnason V, Thompson SG, Sattar N, Selvin E, Hu FB, Danesh J; Emerging Risk Factors Collaboration. Diabetes mellitus, fasting glucose, and risk of cause-specific death. N Engl J Med. 2011;364:829-841. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2142] [Cited by in RCA: 2022] [Article Influence: 144.4] [Reference Citation Analysis (0)] |
| 5. | Tancredi M, Rosengren A, Svensson AM, Kosiborod M, Pivodic A, Gudbjörnsdottir S, Wedel H, Clements M, Dahlqvist S, Lind M. Excess Mortality among Persons with Type 2 Diabetes. N Engl J Med. 2015;373:1720-1732. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 656] [Cited by in RCA: 734] [Article Influence: 73.4] [Reference Citation Analysis (0)] |
| 6. | Rich SS. Mapping genes in diabetes. Genetic epidemiological perspective. Diabetes. 1990;39:1315-1319. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 60] [Cited by in RCA: 50] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 7. | Medici F, Hawa M, Ianari A, Pyke DA, Leslie RD. Concordance rate for type II diabetes mellitus in monozygotic twins: actuarial analysis. Diabetologia. 1999;42:146-150. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 119] [Cited by in RCA: 102] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 8. | Li G, Zhang P, Wang J, Gregg EW, Yang W, Gong Q, Li H, Li H, Jiang Y, An Y, Shuai Y, Zhang B, Zhang J, Thompson TJ, Gerzoff RB, Roglic G, Hu Y, Bennett PH. The long-term effect of lifestyle interventions to prevent diabetes in the China Da Qing Diabetes Prevention Study: a 20-year follow-up study. Lancet. 2008;371:1783-1789. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1074] [Cited by in RCA: 1059] [Article Influence: 62.3] [Reference Citation Analysis (0)] |
| 9. | Lindström J, Ilanne-Parikka P, Peltonen M, Aunola S, Eriksson JG, Hemiö K, Hämäläinen H, Härkönen P, Keinänen-Kiukaanniemi S, Laakso M, Louheranta A, Mannelin M, Paturi M, Sundvall J, Valle TT, Uusitupa M, Tuomilehto J; Finnish Diabetes Prevention Study Group. Sustained reduction in the incidence of type 2 diabetes by lifestyle intervention: follow-up of the Finnish Diabetes Prevention Study. Lancet. 2006;368:1673-1679. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1224] [Cited by in RCA: 1142] [Article Influence: 60.1] [Reference Citation Analysis (0)] |
| 10. | Langenberg C, Lotta LA. Genomic insights into the causes of type 2 diabetes. Lancet. 2018;391:2463-2474. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 90] [Cited by in RCA: 111] [Article Influence: 15.9] [Reference Citation Analysis (0)] |
| 11. | Altshuler D, Hirschhorn JN, Klannemark M, Lindgren CM, Vohl MC, Nemesh J, Lane CR, Schaffner SF, Bolk S, Brewer C, Tuomi T, Gaudet D, Hudson TJ, Daly M, Groop L, Lander ES. The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat Genet. 2000;26:76-80. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1044] [Cited by in RCA: 1038] [Article Influence: 41.5] [Reference Citation Analysis (0)] |
| 12. | Gloyn AL, Weedon MN, Owen KR, Turner MJ, Knight BA, Hitman G, Walker M, Levy JC, Sampson M, Halford S, McCarthy MI, Hattersley AT, Frayling TM. Large-scale association studies of variants in genes encoding the pancreatic beta-cell KATP channel subunits Kir6.2 (KCNJ11) and SUR1 (ABCC8) confirm that the KCNJ11 E23K variant is associated with type 2 diabetes. Diabetes. 2003;52:568-572. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 557] [Cited by in RCA: 511] [Article Influence: 23.2] [Reference Citation Analysis (0)] |
| 13. | Schwanstecher C, Meyer U, Schwanstecher M. K(IR)6.2 polymorphism predisposes to type 2 diabetes by inducing overactivity of pancreatic beta-cell ATP-sensitive K(+) channels. Diabetes. 2002;51:875-879. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 168] [Cited by in RCA: 166] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 14. | Gloyn AL, Hashim Y, Ashcroft SJ, Ashfield R, Wiltshire S, Turner RC; UK Prospective Diabetes Study (UKPDS 53). Association studies of variants in promoter and coding regions of beta-cell ATP-sensitive K-channel genes SUR1 and Kir6.2 with Type 2 diabetes mellitus (UKPDS 53). Diabet Med. 2001;18:206-212. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 129] [Cited by in RCA: 124] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 15. | Hani EH, Boutin P, Durand E, Inoue H, Permutt MA, Velho G, Froguel P. Missense mutations in the pancreatic islet beta cell inwardly rectifying K+ channel gene (KIR6.2/BIR): a meta-analysis suggests a role in the polygenic basis of Type II diabetes mellitus in Caucasians. Diabetologia. 1998;41:1511-1515. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 191] [Cited by in RCA: 170] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 16. | Hansen L, Echwald SM, Hansen T, Urhammer SA, Clausen JO, Pedersen O. Amino acid polymorphisms in the ATP-regulatable inward rectifier Kir6.2 and their relationships to glucose- and tolbutamide-induced insulin secretion, the insulin sensitivity index, and NIDDM. Diabetes. 1997;46:508-512. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 33] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 17. | Inoue H, Ferrer J, Warren-Perry M, Zhang Y, Millns H, Turner RC, Elbein SC, Hampe CL, Suarez BK, Inagaki N, Seino S, Permutt MA. Sequence variants in the pancreatic islet beta-cell inwardly rectifying K+ channel Kir6.2 (Bir) gene: identification and lack of role in Caucasian patients with NIDDM. Diabetes. 1997;46:502-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 57] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 18. | Sakura H, Wat N, Horton V, Millns H, Turner RC, Ashcroft FM. Sequence variations in the human Kir6.2 gene, a subunit of the beta-cell ATP-sensitive K-channel: no association with NIDDM in while Caucasian subjects or evidence of abnormal function when expressed in vitro. Diabetologia. 1996;39:1233-1236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 86] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 19. | Grant SF, Thorleifsson G, Reynisdottir I, Benediktsson R, Manolescu A, Sainz J, Helgason A, Stefansson H, Emilsson V, Helgadottir A, Styrkarsdottir U, Magnusson KP, Walters GB, Palsdottir E, Jonsdottir T, Gudmundsdottir T, Gylfason A, Saemundsdottir J, Wilensky RL, Reilly MP, Rader DJ, Bagger Y, Christiansen C, Gudnason V, Sigurdsson G, Thorsteinsdottir U, Gulcher JR, Kong A, Stefansson K. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat Genet. 2006;38:320-323. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1578] [Cited by in RCA: 1580] [Article Influence: 83.2] [Reference Citation Analysis (0)] |
| 20. | Humphries SE, Gable D, Cooper JA, Ireland H, Stephens JW, Hurel SJ, Li KW, Palmen J, Miller MA, Cappuccio FP, Elkeles R, Godsland I, Miller GJ, Talmud PJ. Common variants in the TCF7L2 gene and predisposition to type 2 diabetes in UK European Whites, Indian Asians and Afro-Caribbean men and women. J Mol Med (Berl). 2006;84:1005-1014. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 113] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 21. | Miyake K, Horikawa Y, Hara K, Yasuda K, Osawa H, Furuta H, Hirota Y, Yamagata K, Hinokio Y, Oka Y, Iwasaki N, Iwamoto Y, Yamada Y, Seino Y, Maegawa H, Kashiwagi A, Yamamoto K, Tokunaga K, Takeda J, Makino H, Nanjo K, Kadowaki T, Kasuga M. Association of TCF7L2 polymorphisms with susceptibility to type 2 diabetes in 4,087 Japanese subjects. J Hum Genet. 2008;53:174-180. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 74] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 22. | Scott LJ, Bonnycastle LL, Willer CJ, Sprau AG, Jackson AU, Narisu N, Duren WL, Chines PS, Stringham HM, Erdos MR, Valle TT, Tuomilehto J, Bergman RN, Mohlke KL, Collins FS, Boehnke M. Association of transcription factor 7-like 2 (TCF7L2) variants with type 2 diabetes in a Finnish sample. Diabetes. 2006;55:2649-2653. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 172] [Cited by in RCA: 176] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 23. | Sandhu MS, Weedon MN, Fawcett KA, Wasson J, Debenham SL, Daly A, Lango H, Frayling TM, Neumann RJ, Sherva R, Blech I, Pharoah PD, Palmer CN, Kimber C, Tavendale R, Morris AD, McCarthy MI, Walker M, Hitman G, Glaser B, Permutt MA, Hattersley AT, Wareham NJ, Barroso I. Common variants in WFS1 confer risk of type 2 diabetes. Nat Genet. 2007;39:951-953. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 299] [Cited by in RCA: 264] [Article Influence: 14.7] [Reference Citation Analysis (0)] |
| 24. | Inoue H, Tanizawa Y, Wasson J, Behn P, Kalidas K, Bernal-Mizrachi E, Mueckler M, Marshall H, Donis-Keller H, Crock P, Rogers D, Mikuni M, Kumashiro H, Higashi K, Sobue G, Oka Y, Permutt MA. A gene encoding a transmembrane protein is mutated in patients with diabetes mellitus and optic atrophy (Wolfram syndrome). Nat Genet. 1998;20:143-148. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 528] [Cited by in RCA: 546] [Article Influence: 20.2] [Reference Citation Analysis (0)] |
| 25. | Sladek R, Rocheleau G, Rung J, Dina C, Shen L, Serre D, Boutin P, Vincent D, Belisle A, Hadjadj S, Balkau B, Heude B, Charpentier G, Hudson TJ, Montpetit A, Pshezhetsky AV, Prentki M, Posner BI, Balding DJ, Meyre D, Polychronakos C, Froguel P. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature. 2007;445:881-885. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2162] [Cited by in RCA: 2103] [Article Influence: 116.8] [Reference Citation Analysis (1)] |
| 26. | Diabetes Genetics Initiative of Broad Institute of Harvard and MIT; Lund University; Novartis Institutes of BioMedical Research; Saxena R, Voight BF, Lyssenko V, Burtt NP, de Bakker PI, Chen H, Roix JJ, Kathiresan S, Hirschhorn JN, Daly MJ, Hughes TE, Groop L, Altshuler D, Almgren P, Florez JC, Meyer J, Ardlie K, Bengtsson Boström K, Isomaa B, Lettre G, Lindblad U, Lyon HN, Melander O, Newton-Cheh C, Nilsson P, Orho-Melander M, Råstam L, Speliotes EK, Taskinen MR, Tuomi T, Guiducci C, Berglund A, Carlson J, Gianniny L, Hackett R, Hall L, Holmkvist J, Laurila E, Sjögren M, Sterner M, Surti A, Svensson M, Svensson M, Tewhey R, Blumenstiel B, Parkin M, Defelice M, Barry R, Brodeur W, Camarata J, Chia N, Fava M, Gibbons J, Handsaker B, Healy C, Nguyen K, Gates C, Sougnez C, Gage D, Nizzari M, Gabriel SB, Chirn GW, Ma Q, Parikh H, Richardson D, Ricke D, Purcell S. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science. 2007;316:1331-1336. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2243] [Cited by in RCA: 2126] [Article Influence: 118.1] [Reference Citation Analysis (0)] |
| 27. | Scott LJ, Mohlke KL, Bonnycastle LL, Willer CJ, Li Y, Duren WL, Erdos MR, Stringham HM, Chines PS, Jackson AU, Prokunina-Olsson L, Ding CJ, Swift AJ, Narisu N, Hu T, Pruim R, Xiao R, Li XY, Conneely KN, Riebow NL, Sprau AG, Tong M, White PP, Hetrick KN, Barnhart MW, Bark CW, Goldstein JL, Watkins L, Xiang F, Saramies J, Buchanan TA, Watanabe RM, Valle TT, Kinnunen L, Abecasis GR, Pugh EW, Doheny KF, Bergman RN, Tuomilehto J, Collins FS, Boehnke M. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science. 2007;316:1341-1345. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2180] [Cited by in RCA: 2055] [Article Influence: 114.2] [Reference Citation Analysis (0)] |
| 28. | Zeggini E, Weedon MN, Lindgren CM, Frayling TM, Elliott KS, Lango H, Timpson NJ, Perry JR, Rayner NW, Freathy RM, Barrett JC, Shields B, Morris AP, Ellard S, Groves CJ, Harries LW, Marchini JL, Owen KR, Knight B, Cardon LR, Walker M, Hitman GA, Morris AD, Doney AS; Wellcome Trust Case Control Consortium (WTCCC), McCarthy MI, Hattersley AT. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science. 2007;316:1336-1341. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1762] [Cited by in RCA: 1653] [Article Influence: 91.8] [Reference Citation Analysis (0)] |
| 29. | Steinthorsdottir V, Thorleifsson G, Reynisdottir I, Benediktsson R, Jonsdottir T, Walters GB, Styrkarsdottir U, Gretarsdottir S, Emilsson V, Ghosh S, Baker A, Snorradottir S, Bjarnason H, Ng MC, Hansen T, Bagger Y, Wilensky RL, Reilly MP, Adeyemo A, Chen Y, Zhou J, Gudnason V, Chen G, Huang H, Lashley K, Doumatey A, So WY, Ma RC, Andersen G, Borch-Johnsen K, Jorgensen T, van Vliet-Ostaptchouk JV, Hofker MH, Wijmenga C, Christiansen C, Rader DJ, Rotimi C, Gurney M, Chan JC, Pedersen O, Sigurdsson G, Gulcher JR, Thorsteinsdottir U, Kong A, Stefansson K. A variant in CDKAL1 influences insulin response and risk of type 2 diabetes. Nat Genet. 2007;39:770-775. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 781] [Cited by in RCA: 777] [Article Influence: 43.2] [Reference Citation Analysis (0)] |
| 30. | Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447:661-678. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7741] [Cited by in RCA: 7160] [Article Influence: 397.8] [Reference Citation Analysis (0)] |
| 31. | Gudmundsson J, Sulem P, Steinthorsdottir V, Bergthorsson JT, Thorleifsson G, Manolescu A, Rafnar T, Gudbjartsson D, Agnarsson BA, Baker A, Sigurdsson A, Benediktsdottir KR, Jakobsdottir M, Blondal T, Stacey SN, Helgason A, Gunnarsdottir S, Olafsdottir A, Kristinsson KT, Birgisdottir B, Ghosh S, Thorlacius S, Magnusdottir D, Stefansdottir G, Kristjansson K, Bagger Y, Wilensky RL, Reilly MP, Morris AD, Kimber CH, Adeyemo A, Chen Y, Zhou J, So WY, Tong PC, Ng MC, Hansen T, Andersen G, Borch-Johnsen K, Jorgensen T, Tres A, Fuertes F, Ruiz-Echarri M, Asin L, Saez B, van Boven E, Klaver S, Swinkels DW, Aben KK, Graif T, Cashy J, Suarez BK, van Vierssen Trip O, Frigge ML, Ober C, Hofker MH, Wijmenga C, Christiansen C, Rader DJ, Palmer CN, Rotimi C, Chan JC, Pedersen O, Sigurdsson G, Benediktsson R, Jonsson E, Einarsson GV, Mayordomo JI, Catalona WJ, Kiemeney LA, Barkardottir RB, Gulcher JR, Thorsteinsdottir U, Kong A, Stefansson K. Two variants on chromosome 17 confer prostate cancer risk, and the one in TCF2 protects against type 2 diabetes. Nat Genet. 2007;39:977-983. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 567] [Cited by in RCA: 545] [Article Influence: 30.3] [Reference Citation Analysis (0)] |
| 32. | Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, Yang J. 10 Years of GWAS Discovery: Biology, Function, and Translation. Am J Hum Genet. 2017;101:5-22. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2025] [Cited by in RCA: 2161] [Article Influence: 270.1] [Reference Citation Analysis (0)] |
| 33. | Zeggini E, Scott LJ, Saxena R, Voight BF, Marchini JL, Hu T, de Bakker PI, Abecasis GR, Almgren P, Andersen G, Ardlie K, Boström KB, Bergman RN, Bonnycastle LL, Borch-Johnsen K, Burtt NP, Chen H, Chines PS, Daly MJ, Deodhar P, Ding CJ, Doney AS, Duren WL, Elliott KS, Erdos MR, Frayling TM, Freathy RM, Gianniny L, Grallert H, Grarup N, Groves CJ, Guiducci C, Hansen T, Herder C, Hitman GA, Hughes TE, Isomaa B, Jackson AU, Jørgensen T, Kong A, Kubalanza K, Kuruvilla FG, Kuusisto J, Langenberg C, Lango H, Lauritzen T, Li Y, Lindgren CM, Lyssenko V, Marvelle AF, Meisinger C, Midthjell K, Mohlke KL, Morken MA, Morris AD, Narisu N, Nilsson P, Owen KR, Palmer CN, Payne F, Perry JR, Pettersen E, Platou C, Prokopenko I, Qi L, Qin L, Rayner NW, Rees M, Roix JJ, Sandbaek A, Shields B, Sjögren M, Steinthorsdottir V, Stringham HM, Swift AJ, Thorleifsson G, Thorsteinsdottir U, Timpson NJ, Tuomi T, Tuomilehto J, Walker M, Watanabe RM, Weedon MN, Willer CJ; Wellcome Trust Case Control Consortium, Illig T, Hveem K, Hu FB, Laakso M, Stefansson K, Pedersen O, Wareham NJ, Barroso I, Hattersley AT, Collins FS, Groop L, McCarthy MI, Boehnke M, Altshuler D. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat Genet. 2008;40:638-645. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1460] [Cited by in RCA: 1395] [Article Influence: 82.1] [Reference Citation Analysis (0)] |
| 34. | Rung J, Cauchi S, Albrechtsen A, Shen L, Rocheleau G, Cavalcanti-Proença C, Bacot F, Balkau B, Belisle A, Borch-Johnsen K, Charpentier G, Dina C, Durand E, Elliott P, Hadjadj S, Järvelin MR, Laitinen J, Lauritzen T, Marre M, Mazur A, Meyre D, Montpetit A, Pisinger C, Posner B, Poulsen P, Pouta A, Prentki M, Ribel-Madsen R, Ruokonen A, Sandbaek A, Serre D, Tichet J, Vaxillaire M, Wojtaszewski JF, Vaag A, Hansen T, Polychronakos C, Pedersen O, Froguel P, Sladek R. Genetic variant near IRS1 is associated with type 2 diabetes, insulin resistance and hyperinsulinemia. Nat Genet. 2009;41:1110-1115. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 369] [Cited by in RCA: 325] [Article Influence: 20.3] [Reference Citation Analysis (0)] |
| 35. | Lyssenko V, Nagorny CL, Erdos MR, Wierup N, Jonsson A, Spégel P, Bugliani M, Saxena R, Fex M, Pulizzi N, Isomaa B, Tuomi T, Nilsson P, Kuusisto J, Tuomilehto J, Boehnke M, Altshuler D, Sundler F, Eriksson JG, Jackson AU, Laakso M, Marchetti P, Watanabe RM, Mulder H, Groop L. Common variant in MTNR1B associated with increased risk of type 2 diabetes and impaired early insulin secretion. Nat Genet. 2009;41:82-88. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 608] [Cited by in RCA: 547] [Article Influence: 34.2] [Reference Citation Analysis (0)] |
| 36. | Bouatia-Naji N, Bonnefond A, Cavalcanti-Proença C, Sparsø T, Holmkvist J, Marchand M, Delplanque J, Lobbens S, Rocheleau G, Durand E, De Graeve F, Chèvre JC, Borch-Johnsen K, Hartikainen AL, Ruokonen A, Tichet J, Marre M, Weill J, Heude B, Tauber M, Lemaire K, Schuit F, Elliott P, Jørgensen T, Charpentier G, Hadjadj S, Cauchi S, Vaxillaire M, Sladek R, Visvikis-Siest S, Balkau B, Lévy-Marchal C, Pattou F, Meyre D, Blakemore AI, Jarvelin MR, Walley AJ, Hansen T, Dina C, Pedersen O, Froguel P. A variant near MTNR1B is associated with increased fasting plasma glucose levels and type 2 diabetes risk. Nat Genet. 2009;41:89-94. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 498] [Cited by in RCA: 445] [Article Influence: 27.8] [Reference Citation Analysis (0)] |
| 37. | Marchini J, Howie B. Genotype imputation for genome-wide association studies. Nat Rev Genet. 2010;11:499-511. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1366] [Cited by in RCA: 1140] [Article Influence: 76.0] [Reference Citation Analysis (0)] |
| 38. | Xue A, Wu Y, Zhu Z, Zhang F, Kemper KE, Zheng Z, Yengo L, Lloyd-Jones LR, Sidorenko J, Wu Y; eQTLGen Consortium, McRae AF, Visscher PM, Zeng J, Yang J. Genome-wide association analyses identify 143 risk variants and putative regulatory mechanisms for type 2 diabetes. Nat Commun. 2018;9:2941. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 589] [Cited by in RCA: 561] [Article Influence: 80.1] [Reference Citation Analysis (0)] |
| 39. | International HapMap Consortium. A haplotype map of the human genome. Nature. 2005;437:1299-1320. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4621] [Cited by in RCA: 4243] [Article Influence: 212.2] [Reference Citation Analysis (0)] |
| 40. | Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G, Levy-Moonshine A, Jordan T, Shakir K, Roazen D, Thibault J, Banks E, Garimella KV, Altshuler D, Gabriel S, DePristo MA. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics. 2013;43:11.10.1-11.1033. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1565] [Cited by in RCA: 3592] [Article Influence: 399.1] [Reference Citation Analysis (0)] |
| 41. | 1000 Genomes Project Consortium; Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, Marchini JL, McCarthy S, McVean GA, Abecasis GR. A global reference for human genetic variation. Nature. 2015;526:68-74. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10217] [Cited by in RCA: 12042] [Article Influence: 1204.2] [Reference Citation Analysis (0)] |
| 42. | UK10K Consortium; Walter K, Min JL, Huang J, Crooks L, Memari Y, McCarthy S, Perry JR, Xu C, Futema M, Lawson D, Iotchkova V, Schiffels S, Hendricks AE, Danecek P, Li R, Floyd J, Wain LV, Barroso I, Humphries SE, Hurles ME, Zeggini E, Barrett JC, Plagnol V, Richards JB, Greenwood CM, Timpson NJ, Durbin R, Soranzo N, . The UK10K project identifies rare variants in health and disease. Nature. 2015;526:82-90. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 823] [Cited by in RCA: 854] [Article Influence: 85.4] [Reference Citation Analysis (0)] |
| 43. | McCarthy S, Das S, Kretzschmar W, Delaneau O, Wood AR, Teumer A, Kang HM, Fuchsberger C, Danecek P, Sharp K, Luo Y, Sidore C, Kwong A, Timpson N, Koskinen S, Vrieze S, Scott LJ, Zhang H, Mahajan A, Veldink J, Peters U, Pato C, van Duijn CM, Gillies CE, Gandin I, Mezzavilla M, Gilly A, Cocca M, Traglia M, Angius A, Barrett JC, Boomsma D, Branham K, Breen G, Brummett CM, Busonero F, Campbell H, Chan A, Chen S, Chew E, Collins FS, Corbin LJ, Smith GD, Dedoussis G, Dorr M, Farmaki AE, Ferrucci L, Forer L, Fraser RM, Gabriel S, Levy S, Groop L, Harrison T, Hattersley A, Holmen OL, Hveem K, Kretzler M, Lee JC, McGue M, Meitinger T, Melzer D, Min JL, Mohlke KL, Vincent JB, Nauck M, Nickerson D, Palotie A, Pato M, Pirastu N, McInnis M, Richards JB, Sala C, Salomaa V, Schlessinger D, Schoenherr S, Slagboom PE, Small K, Spector T, Stambolian D, Tuke M, Tuomilehto J, Van den Berg LH, Van Rheenen W, Volker U, Wijmenga C, Toniolo D, Zeggini E, Gasparini P, Sampson MG, Wilson JF, Frayling T, de Bakker PI, Swertz MA, McCarroll S, Kooperberg C, Dekker A, Altshuler D, Willer C, Iacono W, Ripatti S, Soranzo N, Walter K, Swaroop A, Cucca F, Anderson CA, Myers RM, Boehnke M, McCarthy MI, Durbin R; Haplotype Reference Consortium. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet. 2016;48:1279-1283. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2511] [Cited by in RCA: 2030] [Article Influence: 225.6] [Reference Citation Analysis (0)] |
| 44. | Gaulton KJ, Ferreira T, Lee Y, Raimondo A, Mägi R, Reschen ME, Mahajan A, Locke A, Rayner NW, Robertson N, Scott RA, Prokopenko I, Scott LJ, Green T, Sparso T, Thuillier D, Yengo L, Grallert H, Wahl S, Frånberg M, Strawbridge RJ, Kestler H, Chheda H, Eisele L, Gustafsson S, Steinthorsdottir V, Thorleifsson G, Qi L, Karssen LC, van Leeuwen EM, Willems SM, Li M, Chen H, Fuchsberger C, Kwan P, Ma C, Linderman M, Lu Y, Thomsen SK, Rundle JK, Beer NL, van de Bunt M, Chalisey A, Kang HM, Voight BF, Abecasis GR, Almgren P, Baldassarre D, Balkau B, Benediktsson R, Blüher M, Boeing H, Bonnycastle LL, Bottinger EP, Burtt NP, Carey J, Charpentier G, Chines PS, Cornelis MC, Couper DJ, Crenshaw AT, van Dam RM, Doney AS, Dorkhan M, Edkins S, Eriksson JG, Esko T, Eury E, Fadista J, Flannick J, Fontanillas P, Fox C, Franks PW, Gertow K, Gieger C, Gigante B, Gottesman O, Grant GB, Grarup N, Groves CJ, Hassinen M, Have CT, Herder C, Holmen OL, Hreidarsson AB, Humphries SE, Hunter DJ, Jackson AU, Jonsson A, Jørgensen ME, Jørgensen T, Kao WH, Kerrison ND, Kinnunen L, Klopp N, Kong A, Kovacs P, Kraft P, Kravic J, Langford C, Leander K, Liang L, Lichtner P, Lindgren CM, Lindholm E, Linneberg A, Liu CT, Lobbens S, Luan J, Lyssenko V, Männistö S, McLeod O, Meyer J, Mihailov E, Mirza G, Mühleisen TW, Müller-Nurasyid M, Navarro C, Nöthen MM, Oskolkov NN, Owen KR, Palli D, Pechlivanis S, Peltonen L, Perry JR, Platou CG, Roden M, Ruderfer D, Rybin D, van der Schouw YT, Sennblad B, Sigurðsson G, Stančáková A, Steinbach G, Storm P, Strauch K, Stringham HM, Sun Q, Thorand B, Tikkanen E, Tonjes A, Trakalo J, Tremoli E, Tuomi T, Wennauer R, Wiltshire S, Wood AR, Zeggini E, Dunham I, Birney E, Pasquali L, Ferrer J, Loos RJ, Dupuis J, Florez JC, Boerwinkle E, Pankow JS, van Duijn C, Sijbrands E, Meigs JB, Hu FB, Thorsteinsdottir U, Stefansson K, Lakka TA, Rauramaa R, Stumvoll M, Pedersen NL, Lind L, Keinanen-Kiukaanniemi SM, Korpi-Hyövälti E, Saaristo TE, Saltevo J, Kuusisto J, Laakso M, Metspalu A, Erbel R, Jöcke KH, Moebus S, Ripatti S, Salomaa V, Ingelsson E, Boehm BO, Bergman RN, Collins FS, Mohlke KL, Koistinen H, Tuomilehto J, Hveem K, Njølstad I, Deloukas P, Donnelly PJ, Frayling TM, Hattersley AT, de Faire U, Hamsten A, Illig T, Peters A, Cauchi S, Sladek R, Froguel P, Hansen T, Pedersen O, Morris AD, Palmer CN, Kathiresan S, Melander O, Nilsson PM, Groop LC, Barroso I, Langenberg C, Wareham NJ, O'Callaghan CA, Gloyn AL, Altshuler D, Boehnke M, Teslovich TM, McCarthy MI, Morris AP; DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat Genet. 2015;47:1415-1425. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 372] [Cited by in RCA: 305] [Article Influence: 30.5] [Reference Citation Analysis (0)] |
| 45. | Steinthorsdottir V, Thorleifsson G, Sulem P, Helgason H, Grarup N, Sigurdsson A, Helgadottir HT, Johannsdottir H, Magnusson OT, Gudjonsson SA, Justesen JM, Harder MN, Jørgensen ME, Christensen C, Brandslund I, Sandbæk A, Lauritzen T, Vestergaard H, Linneberg A, Jørgensen T, Hansen T, Daneshpour MS, Fallah MS, Hreidarsson AB, Sigurdsson G, Azizi F, Benediktsson R, Masson G, Helgason A, Kong A, Gudbjartsson DF, Pedersen O, Thorsteinsdottir U, Stefansson K. Identification of low-frequency and rare sequence variants associated with elevated or reduced risk of type 2 diabetes. Nat Genet. 2014;46:294-298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 249] [Cited by in RCA: 229] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 46. | Mahajan A, Taliun D, Thurner M, Robertson NR, Torres JM, Rayner NW, Payne AJ, Steinthorsdottir V, Scott RA, Grarup N, Cook JP, Schmidt EM, Wuttke M, Sarnowski C, Mägi R, Nano J, Gieger C, Trompet S, Lecoeur C, Preuss MH, Prins BP, Guo X, Bielak LF, Below JE, Bowden DW, Chambers JC, Kim YJ, Ng MCY, Petty LE, Sim X, Zhang W, Bennett AJ, Bork-Jensen J, Brummett CM, Canouil M, Ec Kardt KU, Fischer K, Kardia SLR, Kronenberg F, Läll K, Liu CT, Locke AE, Luan J, Ntalla I, Nylander V, Schönherr S, Schurmann C, Yengo L, Bottinger EP, Brandslund I, Christensen C, Dedoussis G, Florez JC, Ford I, Franco OH, Frayling TM, Giedraitis V, Hackinger S, Hattersley AT, Herder C, Ikram MA, Ingelsson M, Jørgensen ME, Jørgensen T, Kriebel J, Kuusisto J, Ligthart S, Lindgren CM, Linneberg A, Lyssenko V, Mamakou V, Meitinger T, Mohlke KL, Morris AD, Nadkarni G, Pankow JS, Peters A, Sattar N, Stančáková A, Strauch K, Taylor KD, Thorand B, Thorleifsson G, Thorsteinsdottir U, Tuomilehto J, Witte DR, Dupuis J, Peyser PA, Zeggini E, Loos RJF, Froguel P, Ingelsson E, Lind L, Groop L, Laakso M, Collins FS, Jukema JW, Palmer CNA, Grallert H, Metspalu A, Dehghan A, Köttgen A, Abecasis GR, Meigs JB, Rotter JI, Marchini J, Pedersen O, Hansen T, Langenberg C, Wareham NJ, Stefansson K, Gloyn AL, Morris AP, Boehnke M, McCarthy MI. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat Genet. 2018;50:1505-1513. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1280] [Cited by in RCA: 1221] [Article Influence: 174.4] [Reference Citation Analysis (1)] |
| 47. | Mahajan A, Wessel J, Willems SM, Zhao W, Robertson NR, Chu AY, Gan W, Kitajima H, Taliun D, Rayner NW, Guo X, Lu Y, Li M, Jensen RA, Hu Y, Huo S, Lohman KK, Zhang W, Cook JP, Prins BP, Flannick J, Grarup N, Trubetskoy VV, Kravic J, Kim YJ, Rybin DV, Yaghootkar H, Müller-Nurasyid M, Meidtner K, Li-Gao R, Varga TV, Marten J, Li J, Smith AV, An P, Ligthart S, Gustafsson S, Malerba G, Demirkan A, Tajes JF, Steinthorsdottir V, Wuttke M, Lecoeur C, Preuss M, Bielak LF, Graff M, Highland HM, Justice AE, Liu DJ, Marouli E, Peloso GM, Warren HR; ExomeBP Consortium; MAGIC Consortium; GIANT Consortium, Afaq S, Afzal S, Ahlqvist E, Almgren P, Amin N, Bang LB, Bertoni AG, Bombieri C, Bork-Jensen J, Brandslund I, Brody JA, Burtt NP, Canouil M, Chen YI, Cho YS, Christensen C, Eastwood SV, Eckardt KU, Fischer K, Gambaro G, Giedraitis V, Grove ML, de Haan HG, Hackinger S, Hai Y, Han S, Tybjærg-Hansen A, Hivert MF, Isomaa B, Jäger S, Jørgensen ME, Jørgensen T, Käräjämäki A, Kim BJ, Kim SS, Koistinen HA, Kovacs P, Kriebel J, Kronenberg F, Läll K, Lange LA, Lee JJ, Lehne B, Li H, Lin KH, Linneberg A, Liu CT, Liu J, Loh M, Mägi R, Mamakou V, McKean-Cowdin R, Nadkarni G, Neville M, Nielsen SF, Ntalla I, Peyser PA, Rathmann W, Rice K, Rich SS, Rode L, Rolandsson O, Schönherr S, Selvin E, Small KS, Stančáková A, Surendran P, Taylor KD, Teslovich TM, Thorand B, Thorleifsson G, Tin A, Tönjes A, Varbo A, Witte DR, Wood AR, Yajnik P, Yao J, Yengo L, Young R, Amouyel P, Boeing H, Boerwinkle E, Bottinger EP, Chowdhury R, Collins FS, Dedoussis G, Dehghan A, Deloukas P, Ferrario MM, Ferrières J, Florez JC, Frossard P, Gudnason V, Harris TB, Heckbert SR, Howson JMM, Ingelsson M, Kathiresan S, Kee F, Kuusisto J, Langenberg C, Launer LJ, Lindgren CM, Männistö S, Meitinger T, Melander O, Mohlke KL, Moitry M, Morris AD, Murray AD, de Mutsert R, Orho-Melander M, Owen KR, Perola M, Peters A, Province MA, Rasheed A, Ridker PM, Rivadineira F, Rosendaal FR, Rosengren AH, Salomaa V, Sheu WH, Sladek R, Smith BH, Strauch K, Uitterlinden AG, Varma R, Willer CJ, Blüher M, Butterworth AS, Chambers JC, Chasman DI, Danesh J, van Duijn C, Dupuis J, Franco OH, Franks PW, Froguel P, Grallert H, Groop L, Han BG, Hansen T, Hattersley AT, Hayward C, Ingelsson E, Kardia SLR, Karpe F, Kooner JS, Köttgen A, Kuulasmaa K, Laakso M, Lin X, Lind L, Liu Y, Loos RJF, Marchini J, Metspalu A, Mook-Kanamori D, Nordestgaard BG, Palmer CNA, Pankow JS, Pedersen O, Psaty BM, Rauramaa R, Sattar N, Schulze MB, Soranzo N, Spector TD, Stefansson K, Stumvoll M, Thorsteinsdottir U, Tuomi T, Tuomilehto J, Wareham NJ, Wilson JG, Zeggini E, Scott RA, Barroso I, Frayling TM, Goodarzi MO, Meigs JB, Boehnke M, Saleheen D, Morris AP, Rotter JI, McCarthy MI. Refining the accuracy of validated target identification through coding variant fine-mapping in type 2 diabetes. Nat Genet. 2018;50:559-571. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 330] [Cited by in RCA: 313] [Article Influence: 44.7] [Reference Citation Analysis (0)] |
| 48. | Kim YJ, Lee J, Kim BJ; T2D-Genes Consortium, Park T. A new strategy for enhancing imputation quality of rare variants from next-generation sequencing data via combining SNP and exome chip data. BMC Genomics. 2015;16:1109. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 49. | Albrechtsen A, Grarup N, Li Y, Sparsø T, Tian G, Cao H, Jiang T, Kim SY, Korneliussen T, Li Q, Nie C, Wu R, Skotte L, Morris AP, Ladenvall C, Cauchi S, Stančáková A, Andersen G, Astrup A, Banasik K, Bennett AJ, Bolund L, Charpentier G, Chen Y, Dekker JM, Doney AS, Dorkhan M, Forsen T, Frayling TM, Groves CJ, Gui Y, Hallmans G, Hattersley AT, He K, Hitman GA, Holmkvist J, Huang S, Jiang H, Jin X, Justesen JM, Kristiansen K, Kuusisto J, Lajer M, Lantieri O, Li W, Liang H, Liao Q, Liu X, Ma T, Ma X, Manijak MP, Marre M, Mokrosiński J, Morris AD, Mu B, Nielsen AA, Nijpels G, Nilsson P, Palmer CN, Rayner NW, Renström F, Ribel-Madsen R, Robertson N, Rolandsson O, Rossing P, Schwartz TW; D. E.S.I.R. Study Group, Slagboom PE, Sterner M; DIAGRAM Consortium, Tang M, Tarnow L, Tuomi T, van't Riet E, van Leeuwen N, Varga TV, Vestmar MA, Walker M, Wang B, Wang Y, Wu H, Xi F, Yengo L, Yu C, Zhang X, Zhang J, Zhang Q, Zhang W, Zheng H, Zhou Y, Altshuler D, 't Hart LM, Franks PW, Balkau B, Froguel P, McCarthy MI, Laakso M, Groop L, Christensen C, Brandslund I, Lauritzen T, Witte DR, Linneberg A, Jørgensen T, Hansen T, Wang J, Nielsen R, Pedersen O. Exome sequencing-driven discovery of coding polymorphisms associated with common metabolic phenotypes. Diabetologia. 2013;56:298-310. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 111] [Cited by in RCA: 97] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 50. | Takatsu H, Hase K, Ohmae M, Ohshima S, Hashimoto K, Taniura N, Yamamoto A, Ohno H. CD300 antigen like family member G: A novel Ig receptor like protein exclusively expressed on capillary endothelium. Biochem Biophys Res Commun. 2006;348:183-191. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 31] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 51. | Umemoto E, Tanaka T, Kanda H, Jin S, Tohya K, Otani K, Matsutani T, Matsumoto M, Ebisuno Y, Jang MH, Fukuda M, Hirata T, Miyasaka M. Nepmucin, a novel HEV sialomucin, mediates L-selectin-dependent lymphocyte rolling and promotes lymphocyte adhesion under flow. J Exp Med. 2006;203:1603-1614. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 50] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 52. | Kooner JS, Saleheen D, Sim X, Sehmi J, Zhang W, Frossard P, Been LF, Chia KS, Dimas AS, Hassanali N, Jafar T, Jowett JB, Li X, Radha V, Rees SD, Takeuchi F, Young R, Aung T, Basit A, Chidambaram M, Das D, Grundberg E, Hedman AK, Hydrie ZI, Islam M, Khor CC, Kowlessur S, Kristensen MM, Liju S, Lim WY, Matthews DR, Liu J, Morris AP, Nica AC, Pinidiyapathirage JM, Prokopenko I, Rasheed A, Samuel M, Shah N, Shera AS, Small KS, Suo C, Wickremasinghe AR, Wong TY, Yang M, Zhang F, DIAGRAM, MuTHER, Abecasis GR, Barnett AH, Caulfield M, Deloukas P, Frayling TM, Froguel P, Kato N, Katulanda P, Kelly MA, Liang J, Mohan V, Sanghera DK, Scott J, Seielstad M, Zimmet PZ, Elliott P, Teo YY, McCarthy MI, Danesh J, Tai ES, Chambers JC. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat Genet. 2011;43:984-989. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 431] [Cited by in RCA: 409] [Article Influence: 29.2] [Reference Citation Analysis (0)] |
| 53. | Teslovich TM, Musunuru K, Smith AV, Edmondson AC, Stylianou IM, Koseki M, Pirruccello JP, Ripatti S, Chasman DI, Willer CJ, Johansen CT, Fouchier SW, Isaacs A, Peloso GM, Barbalic M, Ricketts SL, Bis JC, Aulchenko YS, Thorleifsson G, Feitosa MF, Chambers J, Orho-Melander M, Melander O, Johnson T, Li X, Guo X, Li M, Shin Cho Y, Jin Go M, Jin Kim Y, Lee JY, Park T, Kim K, Sim X, Twee-Hee Ong R, Croteau-Chonka DC, Lange LA, Smith JD, Song K, Hua Zhao J, Yuan X, Luan J, Lamina C, Ziegler A, Zhang W, Zee RY, Wright AF, Witteman JC, Wilson JF, Willemsen G, Wichmann HE, Whitfield JB, Waterworth DM, Wareham NJ, Waeber G, Vollenweider P, Voight BF, Vitart V, Uitterlinden AG, Uda M, Tuomilehto J, Thompson JR, Tanaka T, Surakka I, Stringham HM, Spector TD, Soranzo N, Smit JH, Sinisalo J, Silander K, Sijbrands EJ, Scuteri A, Scott J, Schlessinger D, Sanna S, Salomaa V, Saharinen J, Sabatti C, Ruokonen A, Rudan I, Rose LM, Roberts R, Rieder M, Psaty BM, Pramstaller PP, Pichler I, Perola M, Penninx BW, Pedersen NL, Pattaro C, Parker AN, Pare G, Oostra BA, O'Donnell CJ, Nieminen MS, Nickerson DA, Montgomery GW, Meitinger T, McPherson R, McCarthy MI, McArdle W, Masson D, Martin NG, Marroni F, Mangino M, Magnusson PK, Lucas G, Luben R, Loos RJ, Lokki ML, Lettre G, Langenberg C, Launer LJ, Lakatta EG, Laaksonen R, Kyvik KO, Kronenberg F, König IR, Khaw KT, Kaprio J, Kaplan LM, Johansson A, Jarvelin MR, Janssens AC, Ingelsson E, Igl W, Kees Hovingh G, Hottenga JJ, Hofman A, Hicks AA, Hengstenberg C, Heid IM, Hayward C, Havulinna AS, Hastie ND, Harris TB, Haritunians T, Hall AS, Gyllensten U, Guiducci C, Groop LC, Gonzalez E, Gieger C, Freimer NB, Ferrucci L, Erdmann J, Elliott P, Ejebe KG, Döring A, Dominiczak AF, Demissie S, Deloukas P, de Geus EJ, de Faire U, Crawford G, Collins FS, Chen YD, Caulfield MJ, Campbell H, Burtt NP, Bonnycastle LL, Boomsma DI, Boekholdt SM, Bergman RN, Barroso I, Bandinelli S, Ballantyne CM, Assimes TL, Quertermous T, Altshuler D, Seielstad M, Wong TY, Tai ES, Feranil AB, Kuzawa CW, Adair LS, Taylor HA, Borecki IB, Gabriel SB, Wilson JG, Holm H, Thorsteinsdottir U, Gudnason V, Krauss RM, Mohlke KL, Ordovas JM, Munroe PB, Kooner JS, Tall AR, Hegele RA, Kastelein JJ, Schadt EE, Rotter JI, Boerwinkle E, Strachan DP, Mooser V, Stefansson K, Reilly MP, Samani NJ, Schunkert H, Cupples LA, Sandhu MS, Ridker PM, Rader DJ, van Duijn CM, Peltonen L, Abecasis GR, Boehnke M, Kathiresan S. Biological, clinical and population relevance of 95 loci for blood lipids. Nature. 2010;466:707-713. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3060] [Cited by in RCA: 2841] [Article Influence: 189.4] [Reference Citation Analysis (0)] |
| 54. | Dehghan A, Dupuis J, Barbalic M, Bis JC, Eiriksdottir G, Lu C, Pellikka N, Wallaschofski H, Kettunen J, Henneman P, Baumert J, Strachan DP, Fuchsberger C, Vitart V, Wilson JF, Paré G, Naitza S, Rudock ME, Surakka I, de Geus EJ, Alizadeh BZ, Guralnik J, Shuldiner A, Tanaka T, Zee RY, Schnabel RB, Nambi V, Kavousi M, Ripatti S, Nauck M, Smith NL, Smith AV, Sundvall J, Scheet P, Liu Y, Ruokonen A, Rose LM, Larson MG, Hoogeveen RC, Freimer NB, Teumer A, Tracy RP, Launer LJ, Buring JE, Yamamoto JF, Folsom AR, Sijbrands EJ, Pankow J, Elliott P, Keaney JF, Sun W, Sarin AP, Fontes JD, Badola S, Astor BC, Hofman A, Pouta A, Werdan K, Greiser KH, Kuss O, Meyer zu Schwabedissen HE, Thiery J, Jamshidi Y, Nolte IM, Soranzo N, Spector TD, Völzke H, Parker AN, Aspelund T, Bates D, Young L, Tsui K, Siscovick DS, Guo X, Rotter JI, Uda M, Schlessinger D, Rudan I, Hicks AA, Penninx BW, Thorand B, Gieger C, Coresh J, Willemsen G, Harris TB, Uitterlinden AG, Järvelin MR, Rice K, Radke D, Salomaa V, Willems van Dijk K, Boerwinkle E, Vasan RS, Ferrucci L, Gibson QD, Bandinelli S, Snieder H, Boomsma DI, Xiao X, Campbell H, Hayward C, Pramstaller PP, van Duijn CM, Peltonen L, Psaty BM, Gudnason V, Ridker PM, Homuth G, Koenig W, Ballantyne CM, Witteman JC, Benjamin EJ, Perola M, Chasman DI. Meta-analysis of genome-wide association studies in >80 000 subjects identifies multiple loci for C-reactive protein levels. Circulation. 2011;123:731-738. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 436] [Cited by in RCA: 408] [Article Influence: 29.1] [Reference Citation Analysis (0)] |
| 55. | Fuchsberger C, Flannick J, Teslovich TM, Mahajan A, Agarwala V, Gaulton KJ, Ma C, Fontanillas P, Moutsianas L, McCarthy DJ, Rivas MA, Perry JRB, Sim X, Blackwell TW, Robertson NR, Rayner NW, Cingolani P, Locke AE, Tajes JF, Highland HM, Dupuis J, Chines PS, Lindgren CM, Hartl C, Jackson AU, Chen H, Huyghe JR, van de Bunt M, Pearson RD, Kumar A, Müller-Nurasyid M, Grarup N, Stringham HM, Gamazon ER, Lee J, Chen Y, Scott RA, Below JE, Chen P, Huang J, Go MJ, Stitzel ML, Pasko D, Parker SCJ, Varga TV, Green T, Beer NL, Day-Williams AG, Ferreira T, Fingerlin T, Horikoshi M, Hu C, Huh I, Ikram MK, Kim BJ, Kim Y, Kim YJ, Kwon MS, Lee J, Lee S, Lin KH, Maxwell TJ, Nagai Y, Wang X, Welch RP, Yoon J, Zhang W, Barzilai N, Voight BF, Han BG, Jenkinson CP, Kuulasmaa T, Kuusisto J, Manning A, Ng MCY, Palmer ND, Balkau B, Stančáková A, Abboud HE, Boeing H, Giedraitis V, Prabhakaran D, Gottesman O, Scott J, Carey J, Kwan P, Grant G, Smith JD, Neale BM, Purcell S, Butterworth AS, Howson JMM, Lee HM, Lu Y, Kwak SH, Zhao W, Danesh J, Lam VKL, Park KS, Saleheen D, So WY, Tam CHT, Afzal U, Aguilar D, Arya R, Aung T, Chan E, Navarro C, Cheng CY, Palli D, Correa A, Curran JE, Rybin D, Farook VS, Fowler SP, Freedman BI, Griswold M, Hale DE, Hicks PJ, Khor CC, Kumar S, Lehne B, Thuillier D, Lim WY, Liu J, van der Schouw YT, Loh M, Musani SK, Puppala S, Scott WR, Yengo L, Tan ST, Taylor HA Jr, Thameem F, Wilson G Sr, Wong TY, Njølstad PR, Levy JC, Mangino M, Bonnycastle LL, Schwarzmayr T, Fadista J, Surdulescu GL, Herder C, Groves CJ, Wieland T, Bork-Jensen J, Brandslund I, Christensen C, Koistinen HA, Doney ASF, Kinnunen L, Esko T, Farmer AJ, Hakaste L, Hodgkiss D, Kravic J, Lyssenko V, Hollensted M, Jørgensen ME, Jørgensen T, Ladenvall C, Justesen JM, Käräjämäki A, Kriebel J, Rathmann W, Lannfelt L, Lauritzen T, Narisu N, Linneberg A, Melander O, Milani L, Neville M, Orho-Melander M, Qi L, Qi Q, Roden M, Rolandsson O, Swift A, Rosengren AH, Stirrups K, Wood AR, Mihailov E, Blancher C, Carneiro MO, Maguire J, Poplin R, Shakir K, Fennell T, DePristo M, de Angelis MH, Deloukas P, Gjesing AP, Jun G, Nilsson P, Murphy J, Onofrio R, Thorand B, Hansen T, Meisinger C, Hu FB, Isomaa B, Karpe F, Liang L, Peters A, Huth C, O'Rahilly SP, Palmer CNA, Pedersen O, Rauramaa R, Tuomilehto J, Salomaa V, Watanabe RM, Syvänen AC, Bergman RN, Bharadwaj D, Bottinger EP, Cho YS, Chandak GR, Chan JCN, Chia KS, Daly MJ, Ebrahim SB, Langenberg C, Elliott P, Jablonski KA, Lehman DM, Jia W, Ma RCW, Pollin TI, Sandhu M, Tandon N, Froguel P, Barroso I, Teo YY, Zeggini E, Loos RJF, Small KS, Ried JS, DeFronzo RA, Grallert H, Glaser B, Metspalu A, Wareham NJ, Walker M, Banks E, Gieger C, Ingelsson E, Im HK, Illig T, Franks PW, Buck G, Trakalo J, Buck D, Prokopenko I, Mägi R, Lind L, Farjoun Y, Owen KR, Gloyn AL, Strauch K, Tuomi T, Kooner JS, Lee JY, Park T, Donnelly P, Morris AD, Hattersley AT, Bowden DW, Collins FS, Atzmon G, Chambers JC, Spector TD, Laakso M, Strom TM, Bell GI, Blangero J, Duggirala R, Tai ES, McVean G, Hanis CL, Wilson JG, Seielstad M, Frayling TM, Meigs JB, Cox NJ, Sladek R, Lander ES, Gabriel S, Burtt NP, Mohlke KL, Meitinger T, Groop L, Abecasis G, Florez JC, Scott LJ, Morris AP, Kang HM, Boehnke M, Altshuler D, McCarthy MI. The genetic architecture of type 2 diabetes. Nature. 2016;536:41-47. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 809] [Cited by in RCA: 780] [Article Influence: 86.7] [Reference Citation Analysis (0)] |
| 56. | Collombat P, Mansouri A, Hecksher-Sorensen J, Serup P, Krull J, Gradwohl G, Gruss P. Opposing actions of Arx and Pax4 in endocrine pancreas development. Genes Dev. 2003;17:2591-2603. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 416] [Cited by in RCA: 416] [Article Influence: 18.9] [Reference Citation Analysis (0)] |
| 57. | Kooptiwut S, Plengvidhya N, Chukijrungroat T, Sujjitjoon J, Semprasert N, Furuta H, Yenchitsomanus PT. Defective PAX4 R192H transcriptional repressor activities associated with maturity onset diabetes of the young and early onset-age of type 2 diabetes. J Diabetes Complications. 2012;26:343-347. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 24] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 58. | Nicolae DL. Association Tests for Rare Variants. Annu Rev Genomics Hum Genet. 2016;17:117-130. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 38] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 59. | Flannick J, Fuchsberger C, Mahajan A, Teslovich TM, Agarwala V, Gaulton KJ, Caulkins L, Koesterer R, Ma C, Moutsianas L, McCarthy DJ, Rivas MA, Perry JRB, Sim X, Blackwell TW, Robertson NR, Rayner NW, Cingolani P, Locke AE, Tajes JF, Highland HM, Dupuis J, Chines PS, Lindgren CM, Hartl C, Jackson AU, Chen H, Huyghe JR, van de Bunt M, Pearson RD, Kumar A, Müller-Nurasyid M, Grarup N, Stringham HM, Gamazon ER, Lee J, Chen Y, Scott RA, Below JE, Chen P, Huang J, Go MJ, Stitzel ML, Pasko D, Parker SCJ, Varga TV, Green T, Beer NL, Day-Williams AG, Ferreira T, Fingerlin T, Horikoshi M, Hu C, Huh I, Ikram MK, Kim BJ, Kim Y, Kim YJ, Kwon MS, Lee J, Lee S, Lin KH, Maxwell TJ, Nagai Y, Wang X, Welch RP, Yoon J, Zhang W, Barzilai N, Voight BF, Han BG, Jenkinson CP, Kuulasmaa T, Kuusisto J, Manning A, Ng MCY, Palmer ND, Balkau B, Stančáková A, Abboud HE, Boeing H, Giedraitis V, Prabhakaran D, Gottesman O, Scott J, Carey J, Kwan P, Grant G, Smith JD, Neale BM, Purcell S, Butterworth AS, Howson JMM, Lee HM, Lu Y, Kwak SH, Zhao W, Danesh J, Lam VKL, Park KS, Saleheen D, So WY, Tam CHT, Afzal U, Aguilar D, Arya R, Aung T, Chan E, Navarro C, Cheng CY, Palli D, Correa A, Curran JE, Rybin D, Farook VS, Fowler SP, Freedman BI, Griswold M, Hale DE, Hicks PJ, Khor CC, Kumar S, Lehne B, Thuillier D, Lim WY, Liu J, Loh M, Musani SK, Puppala S, Scott WR, Yengo L, Tan ST, Taylor HA, Thameem F, Wilson G, Wong TY, Njølstad PR, Levy JC, Mangino M, Bonnycastle LL, Schwarzmayr T, Fadista J, Surdulescu GL, Herder C, Groves CJ, Wieland T, Bork-Jensen J, Brandslund I, Christensen C, Koistinen HA, Doney ASF, Kinnunen L, Esko T, Farmer AJ, Hakaste L, Hodgkiss D, Kravic J, Lyssenko V, Hollensted M, Jørgensen ME, Jørgensen T, Ladenvall C, Justesen JM, Käräjämäki A, Kriebel J, Rathmann W, Lannfelt L, Lauritzen T, Narisu N, Linneberg A, Melander O, Milani L, Neville M, Orho-Melander M, Qi L, Qi Q, Roden M, Rolandsson O, Swift A, Rosengren AH, Stirrups K, Wood AR, Mihailov E, Blancher C, Carneiro MO, Maguire J, Poplin R, Shakir K, Fennell T, DePristo M, de Angelis MH, Deloukas P, Gjesing AP, Jun G, Nilsson P, Murphy J, Onofrio R, Thorand B, Hansen T, Meisinger C, Hu FB, Isomaa B, Karpe F, Liang L, Peters A, Huth C, O'Rahilly SP, Palmer CNA, Pedersen O, Rauramaa R, Tuomilehto J, Salomaa V, Watanabe RM, Syvänen AC, Bergman RN, Bharadwaj D, Bottinger EP, Cho YS, Chandak GR, Chan JC, Chia KS, Daly MJ, Ebrahim SB, Langenberg C, Elliott P, Jablonski KA, Lehman DM, Jia W, Ma RCW, Pollin TI, Sandhu M, Tandon N, Froguel P, Barroso I, Teo YY, Zeggini E, Loos RJF, Small KS, Ried JS, DeFronzo RA, Grallert H, Glaser B, Metspalu A, Wareham NJ, Walker M, Banks E, Gieger C, Ingelsson E, Im HK, Illig T, Franks PW, Buck G, Trakalo J, Buck D, Prokopenko I, Mägi R, Lind L, Farjoun Y, Owen KR, Gloyn AL, Strauch K, Tuomi T, Kooner JS, Lee JY, Park T, Donnelly P, Morris AD, Hattersley AT, Bowden DW, Collins FS, Atzmon G, Chambers JC, Spector TD, Laakso M, Strom TM, Bell GI, Blangero J, Duggirala R, Tai ES, McVean G, Hanis CL, Wilson JG, Seielstad M, Frayling TM, Meigs JB, Cox NJ, Sladek R, Lander ES, Gabriel S, Mohlke KL, Meitinger T, Groop L, Abecasis G, Scott LJ, Morris AP, Kang HM, Altshuler D, Burtt NP, Florez JC, Boehnke M, McCarthy MI. Sequence data and association statistics from 12,940 type 2 diabetes cases and controls. Sci Data. 2017;4:170179. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 24] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 60. | Flannick JA, Mercader JM, Fuchsberger C, Udler MS, Mahajan A, Wessel J, Altshuler D, Burtt NP, Scott L, Morris A, Florez J. Genetic discovery and translational decision support from exome sequencing of 20,791 type 2 diabetes cases and 24,440 controls from five ancestries. 2018 Preprint. Available from: BioRxiv:371450. . [DOI] [Full Text] |
| 61. | Barbitoff YA, Serebryakova EA, Nasykhova YA, Predeus AV, Polev DE, Shuvalova AR, Vasiliev EV, Urazov SP, Sarana AM, Scherbak SG, Gladyshev DV, Pokrovskaya MS, Sivakova OV, Meshkov AN, Drapkina OM, Glotov OS, Glotov AS. Identification of Novel Candidate Markers of Type 2 Diabetes and Obesity in Russia by Exome Sequencing with a Limited Sample Size. Genes (Basel). 2018;9:pii: E415. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 19] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 62. | Jun G, Manning A, Almeida M, Zawistowski M, Wood AR, Teslovich TM, Fuchsberger C, Feng S, Cingolani P, Gaulton KJ, Dyer T, Blackwell TW, Chen H, Chines PS, Choi S, Churchhouse C, Fontanillas P, King R, Lee S, Lincoln SE, Trubetskoy V, DePristo M, Fingerlin T, Grossman R, Grundstad J, Heath A, Kim J, Kim YJ, Laramie J, Lee J, Li H, Liu X, Livne O, Locke AE, Maller J, Mazur A, Morris AP, Pollin TI, Ragona D, Reich D, Rivas MA, Scott LJ, Sim X, Tearle RG, Teo YY, Williams AL, Zöllner S, Curran JE, Peralta J, Akolkar B, Bell GI, Burtt NP, Cox NJ, Florez JC, Hanis CL, McKeon C, Mohlke KL, Seielstad M, Wilson JG, Atzmon G, Below JE, Dupuis J, Nicolae DL, Lehman D, Park T, Won S, Sladek R, Altshuler D, McCarthy MI, Duggirala R, Boehnke M, Frayling TM, Abecasis GR, Blangero J. Evaluating the contribution of rare variants to type 2 diabetes and related traits using pedigrees. Proc Natl Acad Sci U S A. 2018;115:379-384. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 25] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 63. | Dorajoo R, Liu J, Boehm BO. Genetics of Type 2 Diabetes and Clinical Utility. Genes (Basel). 2015;6:372-384. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 23] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 64. | Barbitoff YA, Polev DE, Glotov AS, Serebryakova EA, Shcherbakova IV, Kiselev AM, Kostareva AA, Glotov OS, Predeus AV. Systematic dissection of biases in whole-exome and whole-genome sequencing reveals major determinants of coding sequence coverage. 2018 Preprint. Available from: BioRxiv:387639. . [DOI] [Full Text] |
| 65. | Ebbert MT, Jensen TD, Jansen-West K, Sens JP, Reddy JS, Ridge PG, Kauwe JS, Belzil V, Pregent L, Carrasquillo MM, Keene D. Systematic analysis of dark and camouflaged genes: disease-relevant genes hiding in plain sight. Genome Biol. 2019;20:97. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 131] [Cited by in RCA: 121] [Article Influence: 20.2] [Reference Citation Analysis (0)] |
| 66. | Sedlazeck FJ, Lee H, Darby CA, Schatz MC. Piercing the dark matter: bioinformatics of long-range sequencing and mapping. Nat Rev Genet. 2018;19:329-346. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 289] [Cited by in RCA: 322] [Article Influence: 53.7] [Reference Citation Analysis (0)] |
| 67. | Furey TS. ChIP-seq and beyond: new and improved methodologies to detect and characterize protein-DNA interactions. Nat Rev Genet. 2012;13:840-852. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 542] [Cited by in RCA: 532] [Article Influence: 40.9] [Reference Citation Analysis (0)] |
| 68. | van Dijk EL, Auger H, Jaszczyszyn Y, Thermes C. Ten years of next-generation sequencing technology. Trends Genet. 2014;30:418-426. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 932] [Cited by in RCA: 908] [Article Influence: 82.5] [Reference Citation Analysis (0)] |
| 69. | Costa V, Aprile M, Esposito R, Ciccodicola A. RNA-Seq and human complex diseases: recent accomplishments and future perspectives. Eur J Hum Genet. 2013;21:134-142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 168] [Cited by in RCA: 168] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 70. | Jenkinson CP, Göring HH, Arya R, Blangero J, Duggirala R, DeFronzo RA. Transcriptomics in type 2 diabetes: Bridging the gap between genotype and phenotype. Genom Data. 2015;8:25-36. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 27] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 71. | Hung JH, Yang TH, Hu Z, Weng Z, DeLisi C. Gene set enrichment analysis: performance evaluation and usage guidelines. Brief Bioinform. 2012;13:281-291. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 152] [Cited by in RCA: 182] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 72. | Lin ES, Chang WA, Chen YY, Wu LY, Chen YJ, Kuo PL. Deduction of Novel Genes Potentially Involved in Keratinocytes of Type 2 Diabetes Using Next-Generation Sequencing and Bioinformatics Approaches. J Clin Med. 2019;8:pii: E73. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 73. | Feng J, Xing W, Xie L. Regulatory Roles of MicroRNAs in Diabetes. Int J Mol Sci. 2016;17:pii: E1729. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 101] [Cited by in RCA: 102] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 74. | He Y, Ding Y, Liang B, Lin J, Kim TK, Yu H, Hang H, Wang K. A Systematic Study of Dysregulated MicroRNA in Type 2 Diabetes Mellitus. Int J Mol Sci. 2017;18:pii: E456. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 98] [Article Influence: 12.3] [Reference Citation Analysis (0)] |
| 75. | Segerstolpe Å, Palasantza A, Eliasson P, Andersson EM, Andréasson AC, Sun X, Picelli S, Sabirsh A, Clausen M, Bjursell MK, Smith DM, Kasper M, Ämmälä C, Sandberg R. Single-Cell Transcriptome Profiling of Human Pancreatic Islets in Health and Type 2 Diabetes. Cell Metab. 2016;24:593-607. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 961] [Cited by in RCA: 1060] [Article Influence: 117.8] [Reference Citation Analysis (0)] |
| 76. | Xin Y, Kim J, Okamoto H, Ni M, Wei Y, Adler C, Murphy AJ, Yancopoulos GD, Lin C, Gromada J. RNA Sequencing of Single Human Islet Cells Reveals Type 2 Diabetes Genes. Cell Metab. 2016;24:608-615. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 404] [Cited by in RCA: 436] [Article Influence: 48.4] [Reference Citation Analysis (0)] |
| 77. | Dziewulska A, Dobosz AM, Dobrzyn A. High-Throughput Approaches onto Uncover (Epi)Genomic Architecture of Type 2 Diabetes. Genes (Basel). 2018;9:pii: E374. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 17] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 78. | Wahl S, Drong A, Lehne B, Loh M, Scott WR, Kunze S, Tsai PC, Ried JS, Zhang W, Yang Y, Tan S, Fiorito G, Franke L, Guarrera S, Kasela S, Kriebel J, Richmond RC, Adamo M, Afzal U, Ala-Korpela M, Albetti B, Ammerpohl O, Apperley JF, Beekman M, Bertazzi PA, Black SL, Blancher C, Bonder MJ, Brosch M, Carstensen-Kirberg M, de Craen AJ, de Lusignan S, Dehghan A, Elkalaawy M, Fischer K, Franco OH, Gaunt TR, Hampe J, Hashemi M, Isaacs A, Jenkinson A, Jha S, Kato N, Krogh V, Laffan M, Meisinger C, Meitinger T, Mok ZY, Motta V, Ng HK, Nikolakopoulou Z, Nteliopoulos G, Panico S, Pervjakova N, Prokisch H, Rathmann W, Roden M, Rota F, Rozario MA, Sandling JK, Schafmayer C, Schramm K, Siebert R, Slagboom PE, Soininen P, Stolk L, Strauch K, Tai ES, Tarantini L, Thorand B, Tigchelaar EF, Tumino R, Uitterlinden AG, van Duijn C, van Meurs JB, Vineis P, Wickremasinghe AR, Wijmenga C, Yang TP, Yuan W, Zhernakova A, Batterham RL, Smith GD, Deloukas P, Heijmans BT, Herder C, Hofman A, Lindgren CM, Milani L, van der Harst P, Peters A, Illig T, Relton CL, Waldenberger M, Järvelin MR, Bollati V, Soong R, Spector TD, Scott J, McCarthy MI, Elliott P, Bell JT, Matullo G, Gieger C, Kooner JS, Grallert H, Chambers JC. Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity. Nature. 2017;541:81-86. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 578] [Cited by in RCA: 637] [Article Influence: 70.8] [Reference Citation Analysis (0)] |
| 79. | Willmer T, Johnson R, Louw J, Pheiffer C. Blood-Based DNA Methylation Biomarkers for Type 2 Diabetes: Potential for Clinical Applications. Front Endocrinol (Lausanne). 2018;9:744. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 54] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 80. | Cannon ME, Mohlke KL. Deciphering the Emerging Complexities of Molecular Mechanisms at GWAS Loci. Am J Hum Genet. 2018;103:637-653. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 83] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 81. | Feng XW, Ding WP, Xiong LY, Guo L, Sun JM, Xiao P. Recent Advancements in Intestinal Microbiota Analyses: A Review for Non-Microbiologists. Curr Med Sci. 2018;38:949-961. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 4] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 82. | Raygan F, Rezavandi Z, Bahmani F, Ostadmohammadi V, Mansournia MA, Tajabadi-Ebrahimi M, Borzabadi S, Asemi Z. The effects of probiotic supplementation on metabolic status in type 2 diabetic patients with coronary heart disease. Diabetol Metab Syndr. 2018;10:51. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 48] [Cited by in RCA: 69] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 83. | Chen X, Devaraj S. Gut Microbiome in Obesity, Metabolic Syndrome, and Diabetes. Curr Diab Rep. 2018;18:129. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 102] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 84. | Cickovski T, Narasimhan G. Constructing lightweight and flexible pipelines using Plugin-Based Microbiome Analysis (PluMA). Bioinformatics. 2018;34:2881-2888. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 85. | Concolino P, Rizza R, Mignone F, Costella A, Guarino D, Carboni I, Capoluongo E, Santonocito C, Urbani A, Minucci A. A comprehensive BRCA1/2 NGS pipeline for an immediate Copy Number Variation (CNV) detection in breast and ovarian cancer molecular diagnosis. Clin Chim Acta. 2018;480:173-179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 32] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 86. | Allaband C, McDonald D, Vázquez-Baeza Y, Minich JJ, Tripathi A, Brenner DA, Loomba R, Smarr L, Sandborn WJ, Schnabl B, Dorrestein P, Zarrinpar A, Knight R. Microbiome 101: Studying, Analyzing, and Interpreting Gut Microbiome Data for Clinicians. Clin Gastroenterol Hepatol. 2019;17:218-230. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 189] [Article Influence: 31.5] [Reference Citation Analysis (0)] |
| 87. | Jovel J, Patterson J, Wang W, Hotte N, O'Keefe S, Mitchel T, Perry T, Kao D, Mason AL, Madsen KL, Wong GK. Characterization of the Gut Microbiome Using 16S or Shotgun Metagenomics. Front Microbiol. 2016;7:459. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 563] [Cited by in RCA: 548] [Article Influence: 60.9] [Reference Citation Analysis (0)] |
| 88. | Ritari J, Salojärvi J, Lahti L, de Vos WM. Improved taxonomic assignment of human intestinal 16S rRNA sequences by a dedicated reference database. BMC Genomics. 2015;16:1056. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 101] [Cited by in RCA: 103] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 89. | Forster SC, Browne HP, Kumar N, Hunt M, Denise H, Mitchell A, Finn RD, Lawley TD. HPMCD: the database of human microbial communities from metagenomic datasets and microbial reference genomes. Nucleic Acids Res. 2016;44:D604-D609. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 50] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 90. | Laudadio I, Fulci V, Palone F, Stronati L, Cucchiara S, Carissimi C. Quantitative Assessment of Shotgun Metagenomics and 16S rDNA Amplicon Sequencing in the Study of Human Gut Microbiome. OMICS. 2018;22:248-254. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 161] [Article Influence: 26.8] [Reference Citation Analysis (0)] |
| 91. | Ranjan R, Rani A, Metwally A, McGee HS, Perkins DL. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem Biophys Res Commun. 2016;469:967-977. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 449] [Cited by in RCA: 584] [Article Influence: 58.4] [Reference Citation Analysis (0)] |
| 92. | Zhang J, Ding X, Guan R, Zhu C, Xu C, Zhu B, Zhang H, Xiong Z, Xue Y, Tu J, Lu Z. Evaluation of different 16S rRNA gene V regions for exploring bacterial diversity in a eutrophic freshwater lake. Sci Total Environ. 2018;618:1254-1267. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 76] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 93. | Jian C, Luukkonen P, Yki-Jarvinen H, Salonen A, Korpela K. Quantitative PCR provides a simple and accessible method for quantitative microbiome profiling. 2018 Preprint. Available from: BioRxiv:478685 . . [DOI] [Full Text] |
| 94. | Couch RD, Navarro K, Sikaroodi M, Gillevet P, Forsyth CB, Mutlu E, Engen PA, Keshavarzian A. The approach to sample acquisition and its impact on the derived human fecal microbiome and VOC metabolome. PLoS One. 2013;8:e81163. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 35] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 95. | Malla MA, Dubey A, Kumar A, Yadav S, Hashem A, Abd Allah EF. Exploring the Human Microbiome: The Potential Future Role of Next-Generation Sequencing in Disease Diagnosis and Treatment. Front Immunol. 2019;9:2868. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 175] [Cited by in RCA: 178] [Article Influence: 29.7] [Reference Citation Analysis (0)] |
| 96. | Rajilić-Stojanović M, de Vos WM. The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol Rev. 2014;38:996-1047. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 765] [Cited by in RCA: 787] [Article Influence: 71.5] [Reference Citation Analysis (0)] |
| 97. | Aydin Ö, Nieuwdorp M, Gerdes V. The Gut Microbiome as a Target for the Treatment of Type 2 Diabetes. Curr Diab Rep. 2018;18:55. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 82] [Cited by in RCA: 77] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 98. | Bhute SS, Suryavanshi MV, Joshi SM, Yajnik CS, Shouche YS, Ghaskadbi SS. Gut Microbial Diversity Assessment of Indian Type-2-Diabetics Reveals Alterations in Eubacteria, Archaea, and Eukaryotes. Front Microbiol. 2017;8:214. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 96] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 99. | Belzer C, de Vos WM. Microbes inside--from diversity to function: the case of Akkermansia. ISME J. 2012;6:1449-1458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 418] [Cited by in RCA: 520] [Article Influence: 40.0] [Reference Citation Analysis (0)] |
| 100. | Sircana A, Framarin L, Leone N, Berrutti M, Castellino F, Parente R, De Michieli F, Paschetta E, Musso G. Altered Gut Microbiota in Type 2 Diabetes: Just a Coincidence? Curr Diab Rep. 2018;18:98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 138] [Article Influence: 19.7] [Reference Citation Analysis (0)] |
| 101. | Moreno-Indias I, Cardona F, Tinahones FJ, Queipo-Ortuño MI. Impact of the gut microbiota on the development of obesity and type 2 diabetes mellitus. Front Microbiol. 2014;5:190. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 183] [Cited by in RCA: 218] [Article Influence: 19.8] [Reference Citation Analysis (0)] |
| 102. | NCD Risk Factor Collaboration (NCD-RisC). Worldwide trends in body-mass index, underweight, overweight, and obesity from 1975 to 2016: a pooled analysis of 2416 population-based measurement studies in 128·9 million children, adolescents, and adults. Lancet. 2017;390:2627-2642. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4848] [Cited by in RCA: 4666] [Article Influence: 583.3] [Reference Citation Analysis (2)] |
| 103. | Harsch IA, Konturek PC. The Role of Gut Microbiota in Obesity and Type 2 and Type 1 Diabetes Mellitus: New Insights into "Old" Diseases. Med Sci (Basel). 2018;6:pii: E3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 89] [Cited by in RCA: 94] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 104. | Zhang S, Wang B, Shi J, Li J. Network-Based Association Study of Obesity and Type 2 Diabetes with Gene Expression Profiles. Biomed Res Int. 2015;2015:619730. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 4] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 105. | Ridaura VK, Faith JJ, Rey FE, Cheng J, Duncan AE, Kau AL, Griffin NW, Lombard V, Henrissat B, Bain JR, Muehlbauer MJ, Ilkayeva O, Semenkovich CF, Funai K, Hayashi DK, Lyle BJ, Martini MC, Ursell LK, Clemente JC, Van Treuren W, Walters WA, Knight R, Newgard CB, Heath AC, Gordon JI. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science. 2013;341:1241214. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2415] [Cited by in RCA: 2725] [Article Influence: 227.1] [Reference Citation Analysis (0)] |
| 106. | Saad MJ, Santos A, Prada PO. Linking Gut Microbiota and Inflammation to Obesity and Insulin Resistance. Physiology (Bethesda). 2016;31:283-293. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 276] [Cited by in RCA: 471] [Article Influence: 58.9] [Reference Citation Analysis (0)] |
| 107. | Zheng J, Yuan X, Zhang C, Jia P, Jiao S, Zhao X, Yin H, Du Y, Liu H. N-Acetylcysteine alleviates gut dysbiosis and glucose metabolic disorder in high-fat diet-fed mice. J Diabetes. 2019;11:32-45. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 40] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 108. | Teixeira TF, Grześkowiak Ł, Franceschini SC, Bressan J, Ferreira CL, Peluzio MC. Higher level of faecal SCFA in women correlates with metabolic syndrome risk factors. Br J Nutr. 2013;109:914-919. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 96] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 109. | Grasset E, Puel A, Charpentier J, Collet X, Christensen JE, Tercé F, Burcelin R. A Specific Gut Microbiota Dysbiosis of Type 2 Diabetic Mice Induces GLP-1 Resistance through an Enteric NO-Dependent and Gut-Brain Axis Mechanism. Cell Metab. 2017;25:1075-1090.e5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 141] [Cited by in RCA: 186] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 110. | Pedersen HK, Gudmundsdottir V, Nielsen HB, Hyotylainen T, Nielsen T, Jensen BA, Forslund K, Hildebrand F, Prifti E, Falony G, Le Chatelier E, Levenez F, Doré J, Mattila I, Plichta DR, Pöhö P, Hellgren LI, Arumugam M, Sunagawa S, Vieira-Silva S, Jørgensen T, Holm JB, Trošt K; MetaHIT Consortium, Kristiansen K, Brix S, Raes J, Wang J, Hansen T, Bork P, Brunak S, Oresic M, Ehrlich SD, Pedersen O. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature. 2016;535:376-381. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1072] [Cited by in RCA: 1456] [Article Influence: 161.8] [Reference Citation Analysis (0)] |
| 111. | Kobyliak N, Conte C, Cammarota G, Haley AP, Styriak I, Gaspar L, Fusek J, Rodrigo L, Kruzliak P. Probiotics in prevention and treatment of obesity: a critical view. Nutr Metab (Lond). 2016;13:14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 175] [Cited by in RCA: 208] [Article Influence: 23.1] [Reference Citation Analysis (0)] |
| 112. | Sun Z, Sun X, Li J, Li Z, Hu Q, Li L, Hao X, Song M, Li C. Using probiotics for type 2 diabetes mellitus intervention: Advances, questions, and potential. Crit Rev Food Sci Nutr. 2019;1-14. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 73] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 113. | Wu H, Esteve E, Tremaroli V, Khan MT, Caesar R, Mannerås-Holm L, Ståhlman M, Olsson LM, Serino M, Planas-Fèlix M, Xifra G, Mercader JM, Torrents D, Burcelin R, Ricart W, Perkins R, Fernàndez-Real JM, Bäckhed F. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat Med. 2017;23:850-858. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 824] [Cited by in RCA: 1137] [Article Influence: 142.1] [Reference Citation Analysis (0)] |
| 114. | Rodriguez J, Hiel S, Delzenne NM. Metformin: old friend, new ways of action-implication of the gut microbiome? Curr Opin Clin Nutr Metab Care. 2018;21:294-301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 82] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 115. | Floyd JS, Psaty BM. The Application of Genomics in Diabetes: Barriers to Discovery and Implementation. Diabetes Care. 2016;39:1858-1869. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 15] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 116. | Wessel J, Gupta J, de Groot M. Factors Motivating Individuals to Consider Genetic Testing for Type 2 Diabetes Risk Prediction. PLoS One. 2016;11:e0147071. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 117. | Henderson G, Garrett J, Bussey-Jones J, Moloney ME, Blumenthal C, Corbie-Smith G. Great expectations: views of genetic research participants regarding current and future genetic studies. Genet Med. 2008;10:193-200. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 42] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 118. | Kaphingst KA, McBride CM, Wade C, Alford SH, Brody LC, Baxevanis AD. Consumers' use of web-based information and their decisions about multiplex genetic susceptibility testing. J Med Internet Res. 2010;12:e41. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 119. | Meigs JB, Shrader P, Sullivan LM, McAteer JB, Fox CS, Dupuis J, Manning AK, Florez JC, Wilson PW, D'Agostino RB Sr, Cupples LA. Genotype score in addition to common risk factors for prediction of type 2 diabetes. N Engl J Med. 2008;359:2208-2219. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 610] [Cited by in RCA: 561] [Article Influence: 33.0] [Reference Citation Analysis (0)] |
| 120. | Vassy JL, Meigs JB. Is genetic testing useful to predict type 2 diabetes? Best Pract Res Clin Endocrinol Metab. 2012;26:189-201. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 40] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 121. | Grant RW, O'Brien KE, Waxler JL, Vassy JL, Delahanty LM, Bissett LG, Green RC, Stember KG, Guiducci C, Park ER, Florez JC, Meigs JB. Personalized genetic risk counseling to motivate diabetes prevention: a randomized trial. Diabetes Care. 2013;36:13-19. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 101] [Cited by in RCA: 112] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 122. | Hivert MF, Jablonski KA, Perreault L, Saxena R, McAteer JB, Franks PW, Hamman RF, Kahn SE, Haffner S; DIAGRAM Consortium, Meigs JB, Altshuler D, Knowler WC, Florez JC; Diabetes Prevention Program Research Group. Updated genetic score based on 34 confirmed type 2 diabetes Loci is associated with diabetes incidence and regression to normoglycemia in the diabetes prevention program. Diabetes. 2011;60:1340-1348. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 162] [Cited by in RCA: 144] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 123. | Vassy JL. Can genetic information change patient behavior to reduce Type 2 diabetes risk? Per Med. 2013;10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 6] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 124. | Ogden CL, Carroll MD, Kit BK, Flegal KM. Prevalence of childhood and adult obesity in the United States, 2011-2012. JAMA. 2014;311:806-814. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6227] [Cited by in RCA: 5926] [Article Influence: 538.7] [Reference Citation Analysis (1)] |
| 125. | Farmer A, Fox R. Diagnosis, classification, and treatment of diabetes. BMJ. 2011;342:d3319. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 30] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 126. | Oram RA, Patel K, Hill A, Shields B, McDonald TJ, Jones A, Hattersley AT, Weedon MN. A Type 1 Diabetes Genetic Risk Score Can Aid Discrimination Between Type 1 and Type 2 Diabetes in Young Adults. Diabetes Care. 2016;39:337-344. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 175] [Cited by in RCA: 238] [Article Influence: 26.4] [Reference Citation Analysis (0)] |
| 127. | Florez JC, Jablonski KA, Bayley N, Pollin TI, de Bakker PI, Shuldiner AR, Knowler WC, Nathan DM, Altshuler D; Diabetes Prevention Program Research Group. TCF7L2 polymorphisms and progression to diabetes in the Diabetes Prevention Program. N Engl J Med. 2006;355:241-250. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 645] [Cited by in RCA: 621] [Article Influence: 32.7] [Reference Citation Analysis (0)] |
| 128. | Fitipaldi H, McCarthy MI, Florez JC, Franks PW. A Global Overview of Precision Medicine in Type 2 Diabetes. Diabetes. 2018;67:1911-1922. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 74] [Cited by in RCA: 62] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 129. | Lin X, Song K, Lim N, Yuan X, Johnson T, Abderrahmani A, Vollenweider P, Stirnadel H, Sundseth SS, Lai E, Burns DK, Middleton LT, Roses AD, Matthews PM, Waeber G, Cardon L, Waterworth DM, Mooser V. Risk prediction of prevalent diabetes in a Swiss population using a weighted genetic score--the CoLaus Study. Diabetologia. 2009;52:600-608. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 98] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 130. | Sparsø T, Grarup N, Andreasen C, Albrechtsen A, Holmkvist J, Andersen G, Jørgensen T, Borch-Johnsen K, Sandbaek A, Lauritzen T, Madsbad S, Hansen T, Pedersen O. Combined analysis of 19 common validated type 2 diabetes susceptibility gene variants shows moderate discriminative value and no evidence of gene-gene interaction. Diabetologia. 2009;52:1308-1314. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 45] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 131. | Wang J, Stancáková A, Kuusisto J, Laakso M. Identification of undiagnosed type 2 diabetic individuals by the finnish diabetes risk score and biochemical and genetic markers: a population-based study of 7232 Finnish men. J Clin Endocrinol Metab. 2010;95:3858-3862. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 39] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 132. | Lyssenko V, Laakso M. Genetic screening for the risk of type 2 diabetes: worthless or valuable? Diabetes Care. 2013;36 Suppl 2:S120-S126. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 110] [Cited by in RCA: 108] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 133. | Stone MA, Camosso-Stefinovic J, Wilkinson J, de Lusignan S, Hattersley AT, Khunti K. Incorrect and incomplete coding and classification of diabetes: a systematic review. Diabet Med. 2010;27:491-497. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 134. | Ali S, Nafis S, Kalaiarasan P, Rai E, Sharma S, Bamezai RN. Understanding Genetic Heterogeneity in Type 2 Diabetes by Delineating Physiological Phenotypes: SIRT1 and its Gene Network in Impaired Insulin Secretion. Rev Diabet Stud. 2016;13:17-34. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 135. | Kroh EM, Parkin RK, Mitchell PS, Tewari M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods. 2010;50:298-301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 867] [Cited by in RCA: 933] [Article Influence: 62.2] [Reference Citation Analysis (0)] |
| 136. | Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215-233. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14460] [Cited by in RCA: 16089] [Article Influence: 1005.6] [Reference Citation Analysis (2)] |
| 137. | Demirsoy İH, Ertural DY, Balci Ş, Çınkır Ü, Sezer K, Tamer L, Aras N. Profiles of Circulating MiRNAs Following Metformin Treatment in Patients with Type 2 Diabetes. J Med Biochem. 2018;37:499-506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 79] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 138. | Zhu H, Leung SW. Identification of microRNA biomarkers in type 2 diabetes: a meta-analysis of controlled profiling studies. Diabetologia. 2015;58:900-911. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 176] [Cited by in RCA: 207] [Article Influence: 20.7] [Reference Citation Analysis (0)] |
| 139. | Prattichizzo F, Giuliani A, Ceka A, Rippo MR, Bonfigli AR, Testa R, Procopio AD, Olivieri F. Epigenetic mechanisms of endothelial dysfunction in type 2 diabetes. Clin Epigenetics. 2015;7:56. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 67] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 140. | Nath D, Heemels M T, Anson L. Obesity and diabetes. Nature. 2006;444:839–888. [RCA] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 16] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 142. | Walford GA, Colomo N, Todd JN, Billings LK, Fernandez M, Chamarthi B, Warner AS, Davis J, Littleton KR, Hernandez AM, Fanelli RR, Lanier A, Barbato C, Ackerman RJ, Khan SQ, Bui R, Garber L, Stolerman ES, Moore AF, Huang C, Kaur V, Harden M, Taylor A, Chen L, Manning AK, Huang P, Wexler D, McCarthy RM, Lo J, Thomas MK, Grant RW, Goldfine A, Hudson MS, Florez JC. The study to understand the genetics of the acute response to metformin and glipizide in humans (SUGAR-MGH): design of a pharmacogenetic resource for type 2 diabetes. PLoS One. 2015;10:e0121553. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 19] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 143. | TODAY Study Group; Zeitler P, Hirst K, Pyle L, Linder B, Copeland K, Arslanian S, Cuttler L, Nathan DM, Tollefsen S, Wilfley D, Kaufman F. A clinical trial to maintain glycemic control in youth with type 2 diabetes. N Engl J Med. 2012;366:2247-2256. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 662] [Cited by in RCA: 727] [Article Influence: 55.9] [Reference Citation Analysis (2)] |
| 144. | Cook MN, Girman CJ, Stein PP, Alexander CM. Initial monotherapy with either metformin or sulphonylureas often fails to achieve or maintain current glycaemic goals in patients with Type 2 diabetes in UK primary care. Diabet Med. 2007;24:350-358. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 130] [Cited by in RCA: 141] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 145. | Pollastro C, Ziviello C, Costa V, Ciccodicola A. Pharmacogenomics of Drug Response in Type 2 Diabetes: Toward the Definition of Tailored Therapies? PPAR Res. 2015;2015:415149. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 14] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 146. | Pearson ER, Flechtner I, Njølstad PR, Malecki MT, Flanagan SE, Larkin B, Ashcroft FM, Klimes I, Codner E, Iotova V, Slingerland AS, Shield J, Robert JJ, Holst JJ, Clark PM, Ellard S, Søvik O, Polak M, Hattersley AT; Neonatal Diabetes International Collaborative Group. Switching from insulin to oral sulfonylureas in patients with diabetes due to Kir6.2 mutations. N Engl J Med. 2006;355:467-477. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 708] [Cited by in RCA: 671] [Article Influence: 35.3] [Reference Citation Analysis (0)] |
| 147. | Rafiq M, Flanagan SE, Patch AM, Shields BM, Ellard S, Hattersley AT; Neonatal Diabetes International Collaborative Group. Effective treatment with oral sulfonylureas in patients with diabetes due to sulfonylurea receptor 1 (SUR1) mutations. Diabetes Care. 2008;31:204-209. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 208] [Cited by in RCA: 187] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 148. | Kirchheiner J, Brockmöller J, Meineke I, Bauer S, Rohde W, Meisel C, Roots I. Impact of CYP2C9 amino acid polymorphisms on glyburide kinetics and on the insulin and glucose response in healthy volunteers. Clin Pharmacol Ther. 2002;71:286-296. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 152] [Cited by in RCA: 139] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 149. | Tong Y, Lin Y, Zhang Y, Yang J, Zhang Y, Liu H, Zhang B. Association between TCF7L2 gene polymorphisms and susceptibility to type 2 diabetes mellitus: a large Human Genome Epidemiology (HuGE) review and meta-analysis. BMC Med Genet. 2009;10:15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 168] [Cited by in RCA: 176] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 150. | Aquilante CL. Sulfonylurea pharmacogenomics in Type 2 diabetes: the influence of drug target and diabetes risk polymorphisms. Expert Rev Cardiovasc Ther. 2010;8:359-372. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 77] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 151. | Sun X, Yu W, Hu C. Genetics of type 2 diabetes: insights into the pathogenesis and its clinical application. Biomed Res Int. 2014;2014:926713. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 57] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 152. | Soltani G, Hatefi Z, Salehi AR, Khosravi S, Ghiasi MR, Teke K, Aminorroaya A, Salehi R. Pharmacogenomics of Sulfonylureas Response in Relation to rs7754840 Polymorphisms in Cyclin-Dependent Kinase 5 Regulatory Subunit-associated Protein 1-like (CDKAL1) Gene in Iranian Type 2 Diabetes Patients. Adv Biomed Res. 2018;7:96. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 5] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 153. | Graham GG, Punt J, Arora M, Day RO, Doogue MP, Duong JK, Furlong TJ, Greenfield JR, Greenup LC, Kirkpatrick CM, Ray JE, Timmins P, Williams KM. Clinical pharmacokinetics of metformin. Clin Pharmacokinet. 2011;50:81-98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 711] [Cited by in RCA: 849] [Article Influence: 60.6] [Reference Citation Analysis (0)] |
| 154. | Shu Y, Brown C, Castro RA, Shi RJ, Lin ET, Owen RP, Sheardown SA, Yue L, Burchard EG, Brett CM, Giacomini KM. Effect of genetic variation in the organic cation transporter 1, OCT1, on metformin pharmacokinetics. Clin Pharmacol Ther. 2008;83:273-280. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 392] [Cited by in RCA: 353] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 155. | Aprile M, Ambrosio MR, D'Esposito V, Beguinot F, Formisano P, Costa V, Ciccodicola A. PPARG in Human Adipogenesis: Differential Contribution of Canonical Transcripts and Dominant Negative Isoforms. PPAR Res. 2014;2014:537865. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 63] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 156. | Srinivasan S, Kaur V, Chamarthi B, Littleton KR, Chen L, Manning AK, Merino J, Thomas MK, Hudson M, Goldfine A, Florez JC. <i>TCF7L2</i> Genetic Variation Augments Incretin Resistance and Influences Response to a Sulfonylurea and Metformin: The Study to Understand the Genetics of the Acute Response to Metformin and Glipizide in Humans (SUGAR-MGH). Diabetes Care. 2018;41:554-561. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 40] [Article Influence: 5.7] [Reference Citation Analysis (0)] |