Published online Oct 15, 2015. doi: 10.4251/wjgo.v7.i10.233
Peer-review started: May 12, 2015
First decision: July 6, 2015
Revised: August 2, 2015
Accepted: September 7, 2015
Article in press: September 8, 2015
Published online: October 15, 2015
Processing time: 173 Days and 7.4 Hours
The human gastrointestinal tract hosts a complex and vast microbial community with up to 1011-1012 microorganisms colonizing the colon. The gut microbiota has a serious effect on homeostasis and pathogenesis through a number of mechanisms. In recent years, the relationship between the intestinal microbiota and sporadic colorectal cancer has attracted much scientific interest. Mechanisms underlying colonic carcinogenesis include the conversion of procarcinogenic diet-related factors to carcinogens and the stimulation of procarcinogenic signaling pathways in luminal epithelial cells. Understanding each of these mechanisms will facilitate future studies, leading to the development of novel strategies for the diagnosis, treatment, and prevention of colorectal cancer. In this review, we discuss the relationship between colorectal cancer and the intestinal microbiota.
Core tip: Microbiota’s role in providing intestinal homeostasis is not as an audience, but it is active. Both the composition of microbiota and its metabolic activity impact the sensitivity of the host and can cause many pathologies including colorectal cancer.
- Citation: Cipe G, Idiz UO, Firat D, Bektasoglu H. Relationship between intestinal microbiota and colorectal cancer. World J Gastrointest Oncol 2015; 7(10): 233-240
- URL: https://www.wjgnet.com/1948-5204/full/v7/i10/233.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v7.i10.233
Colorectal cancer is the third commonest cancer type worldwide and causes 600000 deaths every year[1]. Because colorectal cancer patients are frequently asymptomatic in the early phase of the disease, diagnosis at this stage presents a significant clinical challenge. Detection of early stage cancers (stages 1-2) allows curative surgery with a 5-year survival rate of 80%. However, survival rates decrease to approximately 10% for metastatic and late stage tumors[2]. Although there are currently methods for the early diagnosis methods, including computed tomography, colonoscopy, and blood tests, it is expected that evaluation of the intestinal microbiota will prove to be a valuable method allowing earlier diagnosis of colorectal cancer.
In humans, a relationship between cancer and microorganisms has been demonstrated in a number of organs, with the most well-known example being the relationship between Helicobacter pylori and gastric cancer and mucosa-associated lymphoid tissue lymphoma[3].
In adults, while the bacterial population in the stomach and small intestine is smaller (103-104 CFU/g contents), increased concentrations of microorganisms are found in the colon (1011-1012 CFU/g contents) compared with the upper gastrointestinal tract. The majority of these microorganisms exist in a favorable symbiotic relationship with humans[3,4]. The intestinal microbiota develops specific to individual variation and environmental conditions beginning at birth[5].
Recently, etiology of colorectal cancer has been shown to be related to genetic mutations, diet, inflammatory processes, lifestyle, and the gut microbiota, with up to 95% of colorectal cancer thought to sporadically develop in individuals with no genetic predisposition[6].
The colonic microbiota is thought to contribute to the development of colorectal cancer by controlling the epithelial cell proliferation and differentiation, synthesizing essential nutrients and bioactive products, preventing the reproduction of pathogenic organisms, and stimulating the immune system[7]. In this review, studies investigating the role of the intestinal microbiota in the development of colorectal cancer development are discussed.
There are 100 billion bacteria in the human intestine with an approximate weight equivalent to 1.5-2 kg. Bacteroidetes and Firmicutes are the major species of the adult intestinal microbiota with the next most frequent species being Actinobacteria, Proteobacteria, and Verrucomicrobia[8].
Normally, colonic bacteria exist in a mutually beneficial symbiotic relationship with humans without adverse effects on the host cells. In situations where this balance is deregulated because of a number of possible causes, the numbers and species of harmful bacteria increase, providing a basis for the development of inflammatory and chronic disease. Changes in the intestinal microbiota have been shown to be associated with obesity, fatty liver, type 1 and 2 diabetes, kidney disease, arthritis, inflammatory bowel disease, and colorectal cancer[9-13]. However, the precise relationship between changes in the microbiota and colorectal cancer has yet to be fully elucidated.
The intestinal microbiota is affected by a number of factors, such as antibiotics, diet, and inflammation[4-18]. A number of studies have reported a high degree of similarity in the intestinal microbiota between members of the same family but a low degree of similarity between heterozygous mice despite being housed in the same cage[9,14,19].
The intestinal microbiota of mice fed standard low-in-fat nutrients has been shown to change within a few weeks with particularly great changes in the composition of Bacteroidetes and Firmicutes species. After mice returned to a low-fat diet, a particularly significant reduction in Mollicutes, a species of Firmicutes, was observed[9,20]. Similar changes have observed with diets high in fat, particularly in obese people, genetically obese mice, and obesity-resistant mice[9,14,21]. Transfer of colon microbiota from mice fed a high-fat diet to mice fed a low-in-fat diet has been shown to accelerate tumor growth suggesting diet-induced changes in the colon microbiota may have a synergistic effect with genetic factors on tumor development[22]. Diet-related changes in intestinal microbiota have also been shown to be associated with colorectal cancer[23].
The relationship between the intestinal microbiota and disease has drawn increased attention in recent years. In particular, recent studies have demonstrated strong associations between the development of colorectal cancer and intestinal bacteria. In these studies, DNA damage caused by superoxide radicals, genotoxin formation, increased T-cell proliferation, and activation of procarcinogenic pathways through a number of receptors have all been shown to contribute to cancer development[24-27].
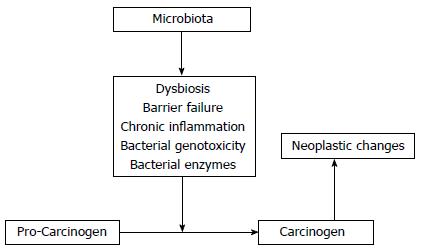
The enzymatic activation or detoxification of carcinogens, and therefore modulation of their tumorigenic activity, has been shown to be influenced by the intestinal microbiota[24,28-35]. In the 1960s, it was observed that germ-free rats exposed to the glycoside, cyasin, did not develop intestinal tumors. Conversely, germ-free rats directly exposed to methylazoximethanol, a sub-active metabolite of cyasin, did develop intestinal tumors[36]. As the formation of methylazoximethanol depends on bacterial β-glucosidase enzyme activity[36], this study was a potent demonstration of the effect of the intestinal microbiota on bioactive carcinogenic compounds. Subsequent research has revealed that the intestinal microbiota converts latent carcinogens to bioactive forms through a number of enzymes, including β-glucuronidase, β-glucosidase, azoreductase, and nitroreductase[37]. Azoxymethane (AOM) is the most frequently used experimental colon carcinogen. AOM is first hydrolyzed in the liver to methylazoximethanol and conjugated to glucuronic acid before bilious excretion into the intestine where it is converted into a highly reactive methyl carbon ion by bacterial β-glucuronidase[34,37,38]. Interestingly, it has been reported that inhibition of β-glucuronidase activity significantly decreases the tumor-inducing potential of AOM in rats[39]. Furthermore, probiotic bacteria, such as Lactobacillus and Bifidobacterium species, have been shown to have anti-carcinogenic effects through the inactivation of microbial enzymes involved in procarcinogenic activation[40]. For example, Lactobacillales, such as L. Casei and L. Acidophilus suppress β-glucuronidase, azoreductase, and nitroreductase activity[41,42]. This balance between the activation and detoxification of potential carcinogens underlies the activation of host oncogenes and tumor suppressors (Figure 1).
In the study by Boleij et al[43] investigating the expression of the Bacteroides fragilis gene (BFT) in colonoscopic samples from 49 healthy individuals and 49 colorectal cancer patients, BFT gene expression was detected more frequently in samples from colorectal cancer patients. When comparing early and late stage cancer patients, BFT gene expression was more frequently detected in late stage cancer patients.
DNA damage and chromosomal instability are early genetic events in the development of colorectal cancer. As with aneuploidy, chromosomal instability is associated with long-term inflammatory bowel disease (IBD) and frequently a precedent event in the subsequent development of colorectal cancer[44-46]. Enterococcus faecalis (E. faecalis), an intestinal bacteria, has been repeatedly found to induce aneuploidy in colonic epithelial cells in monoassociated interleukin (IL)-10 -/- rats and cause aggressive colitis[47,48]. Inhibitors of reactive oxygen and nitrogen species can prevent aneuploidy induced by E. faecalis[49]. These findings demonstrate that intestinal microbiota (particularly specific species) can induce RONS and lead to carcinogenesis.
In intestinal hemostasis, the protective role of the microbiota is thought to be through an effect on epithelial cell proliferation and apoptosis. The main mechanism underlying this effect has been proposed as the conversion of dietary fiber into short chain fatty acids (SCFA), such as acetate, propionate, and butyrate, through microbial fermentation. These SCFAs, particularly butyrate, are readily absorbed easily by the colon and are used as a primary energy source. In addition to significant anti-inflammatory effects[50,51], SCFAs stimulate cell proliferation and differentiation in non-neoplastic normal colon, promote intestinal hemostasis, and the resolution of intestinal injury[51,52]. In addition, SCFAs demonstrate a trans-effect on cancer cells. In particular, butyrate induces apoptosis in colorectal cancer cell lines through a number of mechanisms but predominantly via inhibition of histone deacetylase and activation of intrinsic/mitochondrial apoptosis[53-57].
However, SLC5A and GPR109A, the two major receptors of butyrate, provide protection in the early phases of tumorigenesis as they are frequently inactivated in human cancers[58-60]. It is believed that regulation of microbiota species responsible for the production of butyrate will have efficacy in the treatment of gastrointestinal diseases[61,62]. Therefore, probiotics and in-absorbable food are thought to alter the intestinal microbiota leading to a beneficial increase in the production of short chain fatty acids[63].
Although the development of colorectal cancer has not been attributed to any specific microorganism, a number of cancer-promoting bacteria have been identified (Table 1).
| Bacteria | Subject of study | Evidence | Ref. |
| Helicobacterhepaticus | Animal | Augments azoxymethane induced, and spontaneous colorectal cancer in mice | [64-69] |
| H. hepaticus + H.bilis | Animal | Dual infection induces colorectal cancer in mice | [70,71] |
| H. typhlonius + H. rodentium | Animal | Dual infection in neonates induces colorectal cancer in mice | [72,73] |
| Streptococcus bovis | Human | S.bovis bacteremia and endocarditis associated with human colorectal cancer | [74-77] |
| Animal | Augments azoxymethane induced colorectal cancer in rats | [78] | |
| Human | Increased humoral immune response to S.bovis antigenRpL7/L12, sassociated with increased risk for colorectal cancer | [79] | |
| Bacteroides fragilis | Animal | Enterotoxigenic B.fragilis augments spontaneous colorectal cancer in mice | [26] |
| Human | Increased prevalence of enterotoxigenic B.fragilis in human colorectal cancer | [80] | |
| Human | Increased prevalence in tumor vs normal colonic tissue by quantative PCR analysis | [81] | |
| Human | Increased prevalence in tumor vs normal colonic tissue by quantative PCR analysis | [43] | |
| B. vulgatus | Animal | Induces azoxymethane induced, colorectal cancer in mice | [82] |
| Escherichia coli | Human | Increased mucosa-associated Escherichia coli in human colorectal cancer | [83] |
| Citrobacter rodentium and C. freundii | Animal | Etiologic agent of transmissible murine colonic hyperplasia | [84] |
| Animal | Augments spontaneous and 1,2 dimethylhydrazine induced colorectal cancer in mice | [85,86] | |
| Fusobacterium nucleatum | Human | Increased prevalence in tumor vs normal colonic tissue by quantative PCR analysis | [87] |
| Human | Increased prevalence in tumor vs normal colonic tissue by quantative PCR analysis and 16S ribosomal RNA Gene V3 pyrosequencing analysis | [88] | |
| Human | Increased prevalence in tumor vs normal colonic tissue by quantative PCR analysis | [89] | |
| Animal | 16S ribosomal RNA Gene V3 pyrosequencing analysis | [90] | |
| Enterococcus faecalis | Human | Increased in the feces of colorectal cancer patients by quantative PCR analysis | [91] |
| Furmicutes | Animal | 16S ribosomal RNA Gene V3 pyrosequencing analysis | [90] |
| Akkermansia muciniphila | Human | 16S ribosomal RNA Gene V4 pyrosequencing analysis and Gas Chromatography-Mass Spectrometry | [92] |
| Methanobrevibacterium | Human | Increased prevalence in tumor vs normal colonic tissue by quantative PCR analysis and 16S ribosomal RNA Gene V3 pyrosequencing analysis in fecal samples | [89] |
In rats, Helicobacter hepaticus increases the development of colorectal cancer related to experimental colitis and spontaneous colorectal cancer[65,67]. Bacteroides fragilis is a widespread intestinal bacteria and a potential cause of spontaneous colon tumorigenesis in rats as an enterotoxigenic variant[26].
Exclusion of opportunist pathogens by colonic bacteria may represent a natural defense against colorectal cancer. Similarly, food containing species of Lactobacillus and Bifidobacteria, used as probiotics, provide a number of protective benefits against inflammatory bowel diseases[93-95]. Upon colonizing the host and on the condition of the formation of an additional biofilm, probiotic bacteria have been shown to prevent the adhesion and invasion of pathogen types, maintain host tight junction protein structure, decrease host cytokine production, modulate inflammation and immunity, and neutralize carcinogens and toxins[96-100].
Intestinal microbiota have been shown to cause the release of host antibacterial lectins, stimulate antimicrobial host epithelial responses, and deplete subsets of potentially pathogenic bacteria providing a protective role against abnormal immune responses.
In a study by Sobhani et al[81] of 179 individuals undergoing colonoscopy (60 colorectal cancer, 119 normal), significantly greater levels of Bacteroides/Prevotella bacterial DNA were found in patients with colorectal cancer. Further, it was shown that a greater proportion of IL-17 immunomodulatory cells were isolated from patients with colorectal cancer.
In a study by Gao et al[88] in 2015 examining colon samples from 30 healthy and 31 cancer patients, distal and proximal colon microbiota from both healthy individuals and cancer patients were evaluated using the 16S RNA V3 sequence. No significant difference was observed between proximal and distal colon microbiota; however, in patients with colorectal cancer, Firmicutes and Fusobacteria were over-represented and Proteobacteria were under-represented. Further, Lactococcus and Fusobacterium were identified more often, and Pseudomonas and Escherichia–Shigella less often, in tissues from patients with colorectal cancer compared to those without cancer[88].
In a study by Zhu et al[90] using the 1,2-dimethylhydrazine cancer model, V3 sequences of 16S ribosomal RNA isolated from intestinal microbiota samples from rats with cancer and healthy rats were determined. While Firmucutesin was more frequently observed in rats with colorectal cancer, Bacteroidetes and Spirochetes were less commonly observed. There was no significant difference in the Proteobacteria types between the two groups; however, Prevotella, Lactobacillus, and Treponema were more frequently detected in healthy rats. Furthermore, while Fusobacterium was not observed in healthy rats, it could be identified specifically in cancer rats[90]. In a study of feces samples from healthy individuals and colorectal cancer patients, Akkermansia muciniphila was identified 4 times as often in colorectal cancer patients than healthy individuals[92].
As emphasized in many studies discussed above, intestinal microbiota have a substantial impact on intestinal health through controlling the immune and inflammatory response to individual species of intestinal microbiota, the activation or detoxification of carcinogens, the stimulation of DNA damage and chromosomal instability, dysregulation of the balance between proliferation and apoptosis, and prevention of invasion by pathogens.
Although colorectal cancer development is a complex process, recent studies have shown that the microbiota is actively involved.
Recently, we have developed a greater understanding of the effect of the microbiota on bowel health and diseases, including esophagitis/Barrett’s esophagus, stomach cancer, IBD, and colorectal cancer. However, while a strong relationship between gastrointestinal diseases and the microbiota content is evident, many questions remain unanswered. One of the most clinically challenging issues is to understand how a change in intestinal microbiota will likely impact on the course of disease. Knowledge obtained from dysbiotic microbiota research in germ-free animals and clinical studies involving a variety of intestinal diseases will help provide answers to these important questions. Further, there is currently a lack of data regarding which microorganisms in the microbiota cause disease and are protective.
Continuous improvements in the development of increasingly cost-effective research methods, gene sequencing technology, and high productivity techniques are expected to provide substantial information regarding the healthy and dysbiotic microbiota composition. This information will facilitate functional experiments utilizing cause and effect animal models.
Understanding the relationship between pathology and the microbiota is important; however, the role of microbiota in pathogenesis has yet to be fully elucidated. Therapeutic microbial transplantation has been trialed in metabolic syndrome and also has utility in the treatment of colorectal cancer; however, this technique has many limitations including infection and the promotion of autoimmune disease. Despite this, there is hope that treatments targeting the human microbiota may provide therapies for the prevention and treatment of colorectal cancer in the future.
In summary, the microbiota plays an active role in intestinal homeostasis. Both the composition of microbiota and its metabolic activity have an impact on the host susceptibility to disease and can directly contribute to a number of varied pathologies, including colorectal cancer.
P- Reviewer: Das S S- Editor: Song XX L- Editor: A E- Editor: Wu HL
| 1. | Bonnet M, Buc E, Sauvanet P, Darcha C, Dubois D, Pereira B, Déchelotte P, Bonnet R, Pezet D, Darfeuille-Michaud A. Colonization of the human gut by E. coli and colorectal cancer risk. Clin Cancer Res. 2014;20:859-867. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 364] [Cited by in RCA: 330] [Article Influence: 30.0] [Reference Citation Analysis (0)] |
| 2. | O’Connell JB, Maggard MA, Liu JH, Etzioni DA, Ko CY. Are survival rates different for young and older patients with rectal cancer? Dis Colon Rectum. 2004;47:2064-2069. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 102] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 3. | Gueimonde M, Ouwehand A, Huhtinen H, Salminen E, Salminen S. Qualitative and quantitative analyses of the bifidobacterial microbiota in the colonic mucosa of patients with colorectal cancer, diverticulitis and inflammatory bowel disease. World J Gastroenterol. 2007;13:3985-3989. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 83] [Cited by in RCA: 91] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 4. | Savage DC. Microbial ecology of the gastrointestinal tract. Annu Rev Microbiol. 1977;31:107-133. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1659] [Cited by in RCA: 1467] [Article Influence: 30.6] [Reference Citation Analysis (0)] |
| 5. | Salminen S, Isolauri E. Intestinal colonization, microbiota and probiotics. J Pediatr. 2006;149:115-120. [RCA] [DOI] [Full Text] [Cited by in Crossref: 79] [Cited by in RCA: 66] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 6. | Watson AJ, Collins PD. Colon cancer: a civilization disorder. Dig Dis. 2011;29:222-228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 166] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 7. | Tappenden KA, Deutsch AS. The physiological relevance of the intestinal microbiota--contributions to human health. J Am Coll Nutr. 2007;26:679S-683S. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 73] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 8. | Bäckhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI. Host-bacterial mutualism in the human intestine. Science. 2005;307:1915-1920. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3394] [Cited by in RCA: 3540] [Article Influence: 177.0] [Reference Citation Analysis (5)] |
| 9. | Larsen N, Vogensen FK, van den Berg FW, Nielsen DS, Andreasen AS, Pedersen BK, Al-Soud WA, Sørensen SJ, Hansen LH, Jakobsen M. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS One. 2010;5:e9085. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1783] [Cited by in RCA: 2077] [Article Influence: 138.5] [Reference Citation Analysis (0)] |
| 10. | Abdollahi-Roodsaz S, Joosten LA, Koenders MI, Devesa I, Roelofs MF, Radstake TR, Heuvelmans-Jacobs M, Akira S, Nicklin MJ, Ribeiro-Dias F. Stimulation of TLR2 and TLR4 differentially skews the balance of T cells in a mouse model of arthritis. J Clin Invest. 2008;118:205-216. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 365] [Cited by in RCA: 409] [Article Influence: 24.1] [Reference Citation Analysis (0)] |
| 11. | Ley RE, Turnbaugh PJ, Klein S, Gordon JI. Microbial ecology: human gut microbes associated with obesity. Nature. 2006;444:1022-1023. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7222] [Cited by in RCA: 6403] [Article Influence: 337.0] [Reference Citation Analysis (0)] |
| 12. | Sidhu H, Allison MJ, Chow JM, Clark A, Peck AB. Rapid reversal of hyperoxaluria in a rat model after probiotic administration of Oxalobacter formigenes. J Urol. 2001;166:1487-1491. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 110] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 13. | Tannock GW. Molecular analysis of the intestinal microflora in IBD. Mucosal Immunol. 2008;1 Suppl 1:S15-S18. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 29] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 14. | Turnbaugh PJ, Bäckhed F, Fulton L, Gordon JI. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe. 2008;3:213-223. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2421] [Cited by in RCA: 2189] [Article Influence: 128.8] [Reference Citation Analysis (0)] |
| 15. | Hoffmann C, Hill DA, Minkah N, Kirn T, Troy A, Artis D, Bushman F. Community-wide response of the gut microbiota to enteropathogenic Citrobacter rodentium infection revealed by deep sequencing. Infect Immun. 2009;77:4668-4678. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 113] [Cited by in RCA: 114] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 16. | Preidis GA, Versalovic J. Targeting the human microbiome with antibiotics, probiotics, and prebiotics: gastroenterology enters the metagenomics era. Gastroenterology. 2009;136:2015-2031. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 364] [Cited by in RCA: 327] [Article Influence: 20.4] [Reference Citation Analysis (0)] |
| 17. | Wlodarska M, Finlay BB. Host immune response to antibiotic perturbation of the microbiota. Mucosal Immunol. 2010;3:100-103. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 18. | Lupp C, Robertson ML, Wickham ME, Sekirov I, Champion OL, Gaynor EC, Finlay BB. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe. 2007;2:204. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 311] [Cited by in RCA: 370] [Article Influence: 20.6] [Reference Citation Analysis (0)] |
| 19. | Zhang C, Zhang M, Wang S, Han R, Cao Y, Hua W, Mao Y, Zhang X, Pang X, Wei C. Interactions between gut microbiota, host genetics and diet relevant to development of metabolic syndromes in mice. ISME J. 2010;4:232-241. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 651] [Cited by in RCA: 830] [Article Influence: 51.9] [Reference Citation Analysis (1)] |
| 20. | Wen L, Ley RE, Volchkov PY, Stranges PB, Avanesyan L, Stonebraker AC, Hu C, Wong FS, Szot GL, Bluestone JA. Innate immunity and intestinal microbiota in the development of Type 1 diabetes. Nature. 2008;455:1109-1113. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1623] [Cited by in RCA: 1495] [Article Influence: 87.9] [Reference Citation Analysis (1)] |
| 21. | Turnbaugh PJ, Hamady M, Yatsunenko T, Cantarel BL, Duncan A, Ley RE, Sogin ML, Jones WJ, Roe BA, Affourtit JP. A core gut microbiome in obese and lean twins. Nature. 2009;457:480-484. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6397] [Cited by in RCA: 5660] [Article Influence: 353.8] [Reference Citation Analysis (1)] |
| 22. | Schulz MD, Atay C, Heringer J, Romrig FK, Schwitalla S, Aydin B, Ziegler PK, Varga J, Reindl W, Pommerenke C. High-fat-diet-mediated dysbiosis promotes intestinal carcinogenesis independently of obesity. Nature. 2014;514:508-512. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 284] [Cited by in RCA: 352] [Article Influence: 32.0] [Reference Citation Analysis (0)] |
| 23. | Bingham S, Riboli E. Diet and cancer--the European Prospective Investigation into Cancer and Nutrition. Nat Rev Cancer. 2004;4:206-215. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 276] [Cited by in RCA: 265] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 24. | Toprak NU, Yagci A, Gulluoglu BM, Akin ML, Demirkalem P, Celenk T, Soyletir G. A possible role of Bacteroides fragilis enterotoxin in the aetiology of colorectal cancer. Clin Microbiol Infect. 2006;12:782-786. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 286] [Cited by in RCA: 332] [Article Influence: 17.5] [Reference Citation Analysis (0)] |
| 25. | Wang X, Allen TD, May RJ, Lightfoot S, Houchen CW, Huycke MM. Enterococcus faecalis induces aneuploidy and tetraploidy in colonic epithelial cells through a bystander effect. Cancer Res. 2008;68:9909-9917. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 159] [Cited by in RCA: 152] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 26. | Wu S, Rhee KJ, Albesiano E, Rabizadeh S, Wu X, Yen HR, Huso DL, Brancati FL, Wick E, McAllister F. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med. 2009;15:1016-1022. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1113] [Cited by in RCA: 1303] [Article Influence: 81.4] [Reference Citation Analysis (0)] |
| 27. | Cuevas-Ramos G, Petit CR, Marcq I, Boury M, Oswald E, Nougayrède JP. Escherichia coli induces DNA damage in vivo and triggers genomic instability in mammalian cells. Proc Natl Acad Sci USA. 2010;107:11537-11542. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 495] [Cited by in RCA: 611] [Article Influence: 40.7] [Reference Citation Analysis (0)] |
| 28. | Ge Z, Rogers AB, Feng Y, Lee A, Xu S, Taylor NS, Fox JG. Bacterial cytolethal distending toxin promotes the development of dysplasia in a model of microbially induced hepatocarcinogenesis. Cell Microbiol. 2007;9:2070-2080. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 119] [Cited by in RCA: 122] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 29. | Kim DH, Jin YH. Intestinal bacterial beta-glucuronidase activity of patients with colon cancer. Arch Pharm Res. 2001;24:564-567. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 135] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 30. | Jubelin G, Chavez CV, Taieb F, Banfield MJ, Samba-Louaka A, Nobe R, Nougayrède JP, Zumbihl R, Givaudan A, Escoubas JM. Cycle inhibiting factors (CIFs) are a growing family of functional cyclomodulins present in invertebrate and mammal bacterial pathogens. PLoS One. 2009;4:e4855. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 49] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 31. | Bagnoli F, Buti L, Tompkins L, Covacci A, Amieva MR. Helicobacter pylori CagA induces a transition from polarized to invasive phenotypes in MDCK cells. Proc Natl Acad Sci USA. 2005;102:16339-16344. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 203] [Cited by in RCA: 208] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 32. | Knasmüller S, Steinkellner H, Hirschl AM, Rabot S, Nobis EC, Kassie F. Impact of bacteria in dairy products and of the intestinal microflora on the genotoxic and carcinogenic effects of heterocyclic aromatic amines. Mutat Res. 2001;480-481:129-138. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 90] [Cited by in RCA: 87] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 33. | Femia AP, Dolara P, Giannini A, Salvadori M, Biggeri A, Caderni G. Frequent mutation of Apc gene in rat colon tumors and mucin-depleted foci, preneoplastic lesions in experimental colon carcinogenesis. Cancer Res. 2007;67:445-449. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 51] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 34. | Fiala ES. Investigations into the metabolism and mode of action of the colon carcinogens 1,2-dimethylhydrazine and azoxymethane. Cancer. 1977;40:2436-2445. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 35. | Rowland IR, Rumney CJ, Coutts JT, Lievense LC. Effect of Bifidobacterium longum and inulin on gut bacterial metabolism and carcinogen-induced aberrant crypt foci in rats. Carcinogenesis. 1998;19:281-285. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 324] [Cited by in RCA: 258] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 36. | Laqueur GL, McDaniel EG, Matsumoto H. Tumor induction in germfree rats with methylazoxymethanol (MAM) and synthetic MAM acetate. J Natl Cancer Inst. 1967;39:355-371. [PubMed] |
| 37. | Rowland IR. The role of the gastrointestinal microbiota in colorectal cancer. Curr Pharm Des. 2009;15:1524-1527. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 97] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 38. | Neufert C, Becker C, Neurath MF. An inducible mouse model of colon carcinogenesis for the analysis of sporadic and inflammation-driven tumor progression. Nat Protoc. 2007;2:1998-2004. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 460] [Cited by in RCA: 556] [Article Influence: 30.9] [Reference Citation Analysis (0)] |
| 39. | Takada H, Hirooka T, Hiramatsu Y, Yamamoto M. Effect of beta-glucuronidase inhibitor on azoxymethane-induced colonic carcinogenesis in rats. Cancer Res. 1982;42:331-334. [PubMed] |
| 40. | Geier MS, Butler RN, Howarth GS. Probiotics, prebiotics and synbiotics: a role in chemoprevention for colorectal cancer? Cancer Biol Ther. 2006;5:1265-1269. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 122] [Cited by in RCA: 97] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 41. | Goldin BR, Gorbach SL. Alterations of the intestinal microflora by diet, oral antibiotics, and Lactobacillus: decreased production of free amines from aromatic nitro compounds, azo dyes, and glucuronides. J Natl Cancer Inst. 1984;73:689-695. [PubMed] |
| 42. | Goldin BR, Swenson L, Dwyer J, Sexton M, Gorbach SL. Effect of diet and Lactobacillus acidophilus supplements on human fecal bacterial enzymes. J Natl Cancer Inst. 1980;64:255-261. [PubMed] |
| 43. | Boleij A, Hechenbleikner EM, Goodwin AC, Badani R, Stein EM, Lazarev MG, Ellis B, Carroll KC, Albesiano E, Wick EC. The Bacteroides fragilis toxin gene is prevalent in the colon mucosa of colorectal cancer patients. Clin Infect Dis. 2015;60:208-215. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 313] [Cited by in RCA: 449] [Article Influence: 40.8] [Reference Citation Analysis (0)] |
| 44. | Rubin CE, Haggitt RC, Burmer GC, Brentnall TA, Stevens AC, Levine DS, Dean PJ, Kimmey M, Perera DR, Rabinovitch PS. DNA aneuploidy in colonic biopsies predicts future development of dysplasia in ulcerative colitis. Gastroenterology. 1992;103:1611-1620. [PubMed] |
| 45. | Porschen R, Robin U, Schumacher A, Schauseil S, Borchard F, Hengels KJ, Strohmeyer G. DNA aneuploidy in Crohn’s disease and ulcerative colitis: results of a comparative flow cytometric study. Gut. 1992;33:663-667. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 28] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 46. | Sjöqvist U, Befrits R, Söderlund S, Ost A, Karlén P, Tribukait B, Rubio C, Rutgeerts P, Geboes K, Löfberg R. Colorectal cancer in colonic Crohn’s disease--high frequency of DNA-aneuploidy. Anticancer Res. 2005;25:4393-4397. [PubMed] |
| 47. | Balish E, Warner T. Enterococcus faecalis induces inflammatory bowel disease in interleukin-10 knockout mice. Am J Pathol. 2002;160:2253-2257. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 241] [Cited by in RCA: 246] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 48. | Kim SC, Tonkonogy SL, Albright CA, Tsang J, Balish EJ, Braun J, Huycke MM, Sartor RB. Variable phenotypes of enterocolitis in interleukin 10-deficient mice monoassociated with two different commensal bacteria. Gastroenterology. 2005;128:891-906. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 331] [Cited by in RCA: 339] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 49. | Wang X, Huycke MM. Extracellular superoxide production by Enterococcus faecalis promotes chromosomal instability in mammalian cells. Gastroenterology. 2007;132:551-561. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 152] [Cited by in RCA: 179] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 50. | Segain JP, Raingeard de la Blétière D, Bourreille A, Leray V, Gervois N, Rosales C, Ferrier L, Bonnet C, Blottière HM, Galmiche JP. Butyrate inhibits inflammatory responses through NFkappaB inhibition: implications for Crohn’s disease. Gut. 2000;47:397-403. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 886] [Cited by in RCA: 972] [Article Influence: 38.9] [Reference Citation Analysis (0)] |
| 51. | Maslowski KM, Vieira AT, Ng A, Kranich J, Sierro F, Yu D, Schilter HC, Rolph MS, Mackay F, Artis D. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature. 2009;461:1282-1286. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2007] [Cited by in RCA: 2396] [Article Influence: 149.8] [Reference Citation Analysis (0)] |
| 52. | Scheppach W. Effects of short chain fatty acids on gut morphology and function. Gut. 1994;35:S35-S38. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 517] [Cited by in RCA: 506] [Article Influence: 16.3] [Reference Citation Analysis (0)] |
| 53. | Bolden JE, Peart MJ, Johnstone RW. Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov. 2006;5:769-784. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2201] [Cited by in RCA: 2297] [Article Influence: 120.9] [Reference Citation Analysis (0)] |
| 54. | Heerdt BG, Houston MA, Augenlicht LH. Short-chain fatty acid-initiated cell cycle arrest and apoptosis of colonic epithelial cells is linked to mitochondrial function. Cell Growth Differ. 1997;8:523-532. [PubMed] |
| 55. | Bonnotte B, Favre N, Reveneau S, Micheau O, Droin N, Garrido C, Fontana A, Chauffert B, Solary E, Martin F. Cancer cell sensitization to fas-mediated apoptosis by sodium butyrate. Cell Death Differ. 1998;5:480-487. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 70] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 56. | Hague A, Elder DJ, Hicks DJ, Paraskeva C. Apoptosis in colorectal tumour cells: induction by the short chain fatty acids butyrate, propionate and acetate and by the bile salt deoxycholate. Int J Cancer. 1995;60:400-406. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 291] [Cited by in RCA: 277] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 57. | Ruemmele FM, Schwartz S, Seidman EG, Dionne S, Levy E, Lentze MJ. Butyrate induced Caco-2 cell apoptosis is mediated via the mitochondrial pathway. Gut. 2003;52:94-100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 108] [Cited by in RCA: 119] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 58. | Thangaraju M, Cresci GA, Liu K, Ananth S, Gnanaprakasam JP, Browning DD, Mellinger JD, Smith SB, Digby GJ, Lambert NA. GPR109A is a G-protein-coupled receptor for the bacterial fermentation product butyrate and functions as a tumor suppressor in colon. Cancer Res. 2009;69:2826-2832. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 604] [Cited by in RCA: 568] [Article Influence: 35.5] [Reference Citation Analysis (0)] |
| 59. | Park JY, Helm JF, Zheng W, Ly QP, Hodul PJ, Centeno BA, Malafa MP. Silencing of the candidate tumor suppressor gene solute carrier family 5 member 8 (SLC5A8) in human pancreatic cancer. Pancreas. 2008;36:e32-e39. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 52] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 60. | Whitman SP, Hackanson B, Liyanarachchi S, Liu S, Rush LJ, Maharry K, Margeson D, Davuluri R, Wen J, Witte T. DNA hypermethylation and epigenetic silencing of the tumor suppressor gene, SLC5A8, in acute myeloid leukemia with the MLL partial tandem duplication. Blood. 2008;112:2013-2016. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 58] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 61. | Donohoe DR, Collins LB, Wali A, Bigler R, Sun W, Bultman SJ. The Warburg effect dictates the mechanism of butyrate-mediated histone acetylation and cell proliferation. Mol Cell. 2012;48:612-626. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 512] [Cited by in RCA: 660] [Article Influence: 50.8] [Reference Citation Analysis (0)] |
| 62. | Belcheva A, Irrazabal T, Martin A. Gut microbial metabolism and colon cancer: can manipulations of the microbiota be useful in the management of gastrointestinal health? Bioessays. 2015;37:403-412. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 37] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 63. | Yang T, Owen JL, Lightfoot YL, Kladde MP, Mohamadzadeh M. Microbiota impact on the epigenetic regulation of colorectal cancer. Trends Mol Med. 2013;19:714-725. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 69] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 64. | Erdman SE, Poutahidis T, Tomczak M, Rogers AB, Cormier K, Plank B, Horwitz BH, Fox JG. CD4+ CD25+ regulatory T lymphocytes inhibit microbially induced colon cancer in Rag2-deficient mice. Am J Pathol. 2003;162:691-702. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 243] [Cited by in RCA: 245] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 65. | Nagamine CM, Rogers AB, Fox JG, Schauer DB. Helicobacter hepaticus promotes azoxymethane-initiated colon tumorigenesis in BALB/c-IL10-deficient mice. Int J Cancer. 2008;122:832-838. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 49] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 66. | Maggio-Price L, Treuting P, Zeng W, Tsang M, Bielefeldt-Ohmann H, Iritani BM. Helicobacter infection is required for inflammation and colon cancer in SMAD3-deficient mice. Cancer Res. 2006;66:828-838. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 164] [Cited by in RCA: 177] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 67. | Nagamine CM, Sohn JJ, Rickman BH, Rogers AB, Fox JG, Schauer DB. Helicobacter hepaticus infection promotes colon tumorigenesis in the BALB/c-Rag2(-/-) Apc(Min/+) mouse. Infect Immun. 2008;76:2758-2766. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 50] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 68. | Rao VP, Poutahidis T, Ge Z, Nambiar PR, Boussahmain C, Wang YY, Horwitz BH, Fox JG, Erdman SE. Innate immune inflammatory response against enteric bacteria Helicobacter hepaticus induces mammary adenocarcinoma in mice. Cancer Res. 2006;66:7395-7400. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 155] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 69. | Erdman SE, Rao VP, Poutahidis T, Ihrig MM, Ge Z, Feng Y, Tomczak M, Rogers AB, Horwitz BH, Fox JG. CD4(+)CD25(+) regulatory lymphocytes require interleukin 10 to interrupt colon carcinogenesis in mice. Cancer Res. 2003;63:6042-6050. [PubMed] |
| 70. | Maggio-Price L, Bielefeldt-Ohmann H, Treuting P, Iritani BM, Zeng W, Nicks A, Tsang M, Shows D, Morrissey P, Viney JL. Dual infection with Helicobacter bilis and Helicobacter hepaticus in p-glycoprotein-deficient mdr1a-/- mice results in colitis that progresses to dysplasia. Am J Pathol. 2005;166:1793-1806. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 83] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 71. | Maggio-Price L, Shows D, Waggie K, Burich A, Zeng W, Escobar S, Morrissey P, Viney JL. Helicobacter bilis infection accelerates and H. hepaticus infection delays the development of colitis in multiple drug resistance-deficient (mdr1a-/-) mice. Am J Pathol. 2002;160:739-751. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 110] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 72. | Chichlowski M, Sharp JM, Vanderford DA, Myles MH, Hale LP. Helicobacter typhlonius and Helicobacter rodentium differentially affect the severity of colon inflammation and inflammation-associated neoplasia in IL10-deficient mice. Comp Med. 2008;58:534-541. [PubMed] |
| 73. | Hale LP, Perera D, Gottfried MR, Maggio-Price L, Srinivasan S, Marchuk D. Neonatal co-infection with helicobacter species markedly accelerates the development of inflammation-associated colonic neoplasia in IL-10(-/-) mice. Helicobacter. 2007;12:598-604. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 22] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 74. | Zarkin BA, Lillemoe KD, Cameron JL, Effron PN, Magnuson TH, Pitt HA. The triad of Streptococcus bovis bacteremia, colonic pathology, and liver disease. Ann Surg. 1990;211:786-791; discussion 791-792. [PubMed] |
| 75. | Gold JS, Bayar S, Salem RR. Association of Streptococcus bovis bacteremia with colonic neoplasia and extracolonic malignancy. Arch Surg. 2004;139:760-765. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 141] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 76. | Klein RS, Recco RA, Catalano MT, Edberg SC, Casey JI, Steigbigel NH. Association of Streptococcus bovis with carcinoma of the colon. N Engl J Med. 1977;297:800-802. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 403] [Cited by in RCA: 380] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 77. | Ruoff KL, Miller SI, Garner CV, Ferraro MJ, Calderwood SB. Bacteremia with Streptococcus bovis and Streptococcus salivarius: clinical correlates of more accurate identification of isolates. J Clin Microbiol. 1989;27:305-308. [PubMed] |
| 78. | Ellmerich S, Schöller M, Duranton B, Gossé F, Galluser M, Klein JP, Raul F. Promotion of intestinal carcinogenesis by Streptococcus bovis. Carcinogenesis. 2000;21:753-756. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 171] [Cited by in RCA: 177] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 79. | Boleij A, Roelofs R, Schaeps RM, Schülin T, Glaser P, Swinkels DW, Kato I, Tjalsma H. Increased exposure to bacterial antigen RpL7/L12 in early stage colorectal cancer patients. Cancer. 2010;116:4014-4022. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 40] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 80. | Huang JQ, Zheng GF, Sumanac K, Irvine EJ, Hunt RH. Meta-analysis of the relationship between cagA seropositivity and gastric cancer. Gastroenterology. 2003;125:1636-1644. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 365] [Cited by in RCA: 380] [Article Influence: 17.3] [Reference Citation Analysis (0)] |
| 81. | Sobhani I, Tap J, Roudot-Thoraval F, Roperch JP, Letulle S, Langella P, Corthier G, Tran Van Nhieu J, Furet JP. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS One. 2011;6:e16393. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 630] [Cited by in RCA: 620] [Article Influence: 44.3] [Reference Citation Analysis (0)] |
| 82. | Uronis JM, Mühlbauer M, Herfarth HH, Rubinas TC, Jones GS, Jobin C. Modulation of the intestinal microbiota alters colitis-associated colorectal cancer susceptibility. PLoS One. 2009;4:e6026. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 315] [Cited by in RCA: 336] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 83. | Martin HM, Campbell BJ, Hart CA, Mpofu C, Nayar M, Singh R, Englyst H, Williams HF, Rhodes JM. Enhanced Escherichia coli adherence and invasion in Crohn’s disease and colon cancer. Gastroenterology. 2004;127:80-93. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 508] [Cited by in RCA: 558] [Article Influence: 26.6] [Reference Citation Analysis (0)] |
| 84. | Luperchio SA, Schauer DB. Molecular pathogenesis of Citrobacter rodentium and transmissible murine colonic hyperplasia. Microbes Infect. 2001;3:333-340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 206] [Cited by in RCA: 218] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 85. | Barthold SW, Jonas AM. Morphogenesis of early 1, 2-dimethylhydrazine-induced lesions and latent period reduction of colon carcinogenesis in mice by a variant of Citrobacter freundii. Cancer Res. 1977;37:4352-4360. [PubMed] |
| 86. | Newman JV, Kosaka T, Sheppard BJ, Fox JG, Schauer DB. Bacterial infection promotes colon tumorigenesis in Apc(Min/+) mice. J Infect Dis. 2001;184:227-230. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 115] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 87. | Castellarin M, Warren RL, Freeman JD, Dreolini L, Krzywinski M, Strauss J, Barnes R, Watson P, Allen-Vercoe E, Moore RA. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 2012;22:299-306. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1164] [Cited by in RCA: 1493] [Article Influence: 106.6] [Reference Citation Analysis (0)] |
| 88. | Gao Z, Guo B, Gao R, Zhu Q, Qin H. Microbiota disbiosis is associated with colorectal cancer. Front Microbiol. 2015;6:20. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 242] [Cited by in RCA: 385] [Article Influence: 38.5] [Reference Citation Analysis (0)] |
| 89. | Mira-Pascual L, Cabrera-Rubio R, Ocon S, Costales P, Parra A, Suarez A, Moris F, Rodrigo L, Mira A, Collado MC. Microbial mucosal colonic shifts associated with the development of colorectal cancer reveal the presence of different bacterial and archaeal biomarkers. J Gastroenterol. 2015;50:167-179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 161] [Cited by in RCA: 196] [Article Influence: 19.6] [Reference Citation Analysis (0)] |
| 90. | Zhu Q, Jin Z, Wu W, Gao R, Guo B, Gao Z, Yang Y, Qin H. Analysis of the intestinal lumen microbiota in an animal model of colorectal cancer. PLoS One. 2014;9:e90849. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 141] [Cited by in RCA: 149] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 91. | Balamurugan R, Rajendiran E, George S, Samuel GV, Ramakrishna BS. Real-time polymerase chain reaction quantification of specific butyrate-producing bacteria, Desulfovibrio and Enterococcus faecalis in the feces of patients with colorectal cancer. J Gastroenterol Hepatol. 2008;23:1298-1303. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 208] [Cited by in RCA: 227] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 92. | Weir TL, Manter DK, Sheflin AM, Barnett BA, Heuberger AL, Ryan EP. Stool microbiome and metabolome differences between colorectal cancer patients and healthy adults. PLoS One. 2013;8:e70803. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 414] [Cited by in RCA: 491] [Article Influence: 40.9] [Reference Citation Analysis (0)] |
| 93. | Lightfoot YL, Yang T, Sahay B, Mohamadzadeh M. Targeting aberrant colon cancer-specific DNA methylation with lipoteichoic acid-deficient Lactobacillus acidophilus. Gut Microbes. 2013;4:84-88. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 35] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 94. | Mohamadzadeh M, Pfeiler EA, Brown JB, Zadeh M, Gramarossa M, Managlia E, Bere P, Sarraj B, Khan MW, Pakanati KC. Regulation of induced colonic inflammation by Lactobacillus acidophilus deficient in lipoteichoic acid. Proc Natl Acad Sci USA. 2011;108 Suppl 1:4623-4630. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 224] [Cited by in RCA: 216] [Article Influence: 15.4] [Reference Citation Analysis (0)] |
| 95. | Licciardi PV, Wong SS, Tang ML, Karagiannis TC. Epigenome targeting by probiotic metabolites. Gut Pathog. 2010;2:24. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 50] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 96. | Jones SE, Versalovic J. Probiotic Lactobacillus reuteri biofilms produce antimicrobial and anti-inflammatory factors. BMC Microbiol. 2009;9:35. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 229] [Cited by in RCA: 281] [Article Influence: 17.6] [Reference Citation Analysis (0)] |
| 97. | Resta-Lenert S, Barrett KE. Live probiotics protect intestinal epithelial cells from the effects of infection with enteroinvasive Escherichia coli (EIEC). Gut. 2003;52:988-997. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 468] [Cited by in RCA: 428] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 98. | Qin H, Zhang Z, Hang X, Jiang Y. L. plantarum prevents enteroinvasive Escherichia coli-induced tight junction proteins changes in intestinal epithelial cells. BMC Microbiol. 2009;9:63. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 122] [Cited by in RCA: 139] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 99. | Pagnini C, Saeed R, Bamias G, Arseneau KO, Pizarro TT, Cominelli F. Probiotics promote gut health through stimulation of epithelial innate immunity. Proc Natl Acad Sci USA. 2010;107:454-459. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 278] [Cited by in RCA: 265] [Article Influence: 17.7] [Reference Citation Analysis (0)] |
| 100. | Liévin-Le Moal V, Amsellem R, Servin AL, Coconnier MH. Lactobacillus acidophilus (strain LB) from the resident adult human gastrointestinal microflora exerts activity against brush border damage promoted by a diarrhoeagenic Escherichia coli in human enterocyte-like cells. Gut. 2002;50:803-811. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 106] [Article Influence: 4.6] [Reference Citation Analysis (0)] |