Published online Nov 26, 2014. doi: 10.4252/wjsc.v6.i5.620
Revised: September 5, 2014
Accepted: September 17, 2014
Published online: November 26, 2014
Processing time: 66 Days and 19.4 Hours
Induced pluripotent stem (iPS) cells, somatic cells reprogrammed to the pluripotent state by forced expression of defined factors, represent a uniquely valuable resource for research and regenerative medicine. However, this methodology remains inefficient due to incomplete mechanistic understanding of the reprogramming process. In recent years, various groups have endeavoured to interrogate the cell signalling that governs the reprogramming process, including LIF/STAT3, BMP, PI3K, FGF2, Wnt, TGFβ and MAPK pathways, with the aim of increasing our understanding and identifying new mechanisms of improving safety, reproducibility and efficiency. This has led to a unified model of reprogramming that consists of 3 stages: initiation, maturation and stabilisation. Initiation of reprogramming occurs in almost all cells that receive the reprogramming transgenes; most commonly Oct4, Sox2, Klf4 and cMyc, and involves a phenotypic mesenchymal-to-epithelial transition. The initiation stage is also characterised by increased proliferation and a metabolic switch from oxidative phosphorylation to glycolysis. The maturation stage is considered the major bottleneck within the process, resulting in very few “stabilisation competent” cells progressing to the final stabilisation phase. To reach this stage in both mouse and human cells, pre-iPS cells must activate endogenous expression of the core circuitry of pluripotency, comprising Oct4, Sox2, and Nanog, and thus reach a state of transgene independence. By the stabilisation stage, iPS cells generally use the same signalling networks that govern pluripotency in embryonic stem cells. These pathways differ between mouse and human cells although recent work has demonstrated that this is context dependent. As iPS cell generation technologies move forward, tools are being developed to interrogate the process in more detail, thus allowing a greater understanding of this intriguing biological phenomenon.
Core tip: Induced pluripotent stem (iPS) cells present great promise, both to research and to medicine. However, we know very little regarding the mechanisms that occur throughout the iPS cell reprogramming process and thus the process remains inefficient. In this review, we discuss the 3 stages of reprogramming, initiation, maturation and stabilisation, and clarify the signalling pathways underlying each phase. We draw together the current knowledge to propose a model for the interactions between the key pathways in iPS cell reprogramming with the aim of illuminating this complex yet fascinating process.
- Citation: Hawkins K, Joy S, McKay T. Cell signalling pathways underlying induced pluripotent stem cell reprogramming. World J Stem Cells 2014; 6(5): 620-628
- URL: https://www.wjgnet.com/1948-0210/full/v6/i5/620.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v6.i5.620
Pluripotency, the ability of a single cell to give rise to all cells within an entire living organism, is of great biological interest both in terms of understanding developmental mechanisms as well as the medical potential that pluripotent stem cells possess. However, our understanding of the cell signalling networks underlying this complex process still remains incomplete. The first pluripotent stem cells were isolated from mouse blastocysts simultaneously by 2 groups in 1981[1,2]. This was replicated 17 years later using human blastocysts[3]. Embryonic stem (ES) cells have since been isolated from other species including rhesus monkeys[4] and rats[5,6]. Both human and mouse ES cells have provided and invaluable resource to understand the basic biology of the pluripotent state.
A “core circuitry” of homeodomain transcription factors, Oct4[7], Sox2[8] and Nanog[9], governs pluripotency in both mouse and human ES cells[10]. These transcription factors are expressed both in vivo in the inner cell mass (ICM) of the blastocyst and in vitro, in pluripotent cells. These 3 factors closely interact within the cell; for example Oct4 and Sox2 have been shown to form a heterodimeric transcription complex[11-13] and all 3 factors share target genes[14,15]. This interaction facilitates the precise regulation of the core circuitry necessary to maintain the pluripotent state; for instance Oct4 overexpression leads to endoderm and mesoderm differentiation whereas blockade of Oct4 induces trophoblast differentiation[7]. This may be explained by its biphasic role in Nanog regulation whereby low levels of Oct4 result in upregulation of Nanog whereas higher levels of Oct4 result in downregulation of Nanog[15]. Similarly, small increases in Sox2 expression or ablation of Sox2 expression both induce multilineage differentiation[16]. Blockade of Nanog does not induce differentiation, thus indicating that Nanog’s role in the core circuitry of pluripotency is to stabilise the pluripotent state rather than acting as a housekeeper. However, Nanog knockdown does lead to an increased capacity for differentiation into primitive ectoderm[9].
The core pluripotency circuitry is also autoregulatory since all 3 factors have been shown to regulate the expression of each other as well as themselves[14,15,17]. Interestingly, SOX2 is dispensable for the activation of Oct4/Sox2 target genes since forced expression of Oct4 is able to rescue pluripotency in Sox2-/- cells, however, Sox2 expression is necessary to maintain Oct4 expression[8]. Although it is clear that OCT4, SOX2 and NANOG occupy the top level of the pluripotency hierarchy, these core factors also regulate a wide range of genes associated with pluripotency signalling networks including Stat3, Zic3, Tdgf1, Lefty/Ebaf, Dkk1 and Frat2[14].
With the emergence of this complex molecular inter-play of dosage dependency between hierarchical transcription factors in the maintenance of the somewhat unstable pluripotent ground state, it seems surprising that simply over-expressing these factors in somatic cells can induce the pluripotent state. However, the collective seminal studies of Yamanaka and Thomson show this to be feasible in their descriptions of reprogramming somatic cells to induced Pluripotent Stem (iPS) cells[18-20].
The original iPS cell reprogramming strategy published by Takahashi et al[19] 7 years ago remains robust and largely unaltered to the present day. The “Yamanaka factors”, Oct4, Sox2, Klf4 and cMyc were constitutively expressed using genome integrating retroviruses in both mouse[18] and subsequently human[19] fibroblasts, and under ES cell culture conditions were able to induce pluripotency. To date, this methodology is still widely used, however, various adaptations to the method of vector delivery and reprogramming factors (Table 1) have been made. Advances in vector delivery have generally been made to either improve efficiency or safety, by preventing integration of the transgenes into the genome. For example, iPS cells have now been successfully generated using episomal plasmids[21], Sendai viruses[22] and piggyBac transposons[23] to deliver the reprogramming factors and even proteins[24] or small molecules[25] alone. Many divergent cell-types have been successfully reprogrammed to pluripotency including neural stem cells[26], neural progenitor cells[27], keratinocytes[28], B lymphocytes[29], meningeal membrane cells[30], peripheral blood mononuclear cells[31] and pancreatic β cells[32]. Often the minimal factors necessary to reprogram a cell depend on the endogenous “stemness” of the starting cell, for example, neural stem cells can be reprogrammed using Oct4 alone since they express high levels of the other Yamanaka factors[26].
| Reprogramming factor | Human/mouse | Ref. |
| Oct4 | Both | Takahashi et al[18,19] |
| Sox2 | Both | Takahashi et al[18,19] |
| cMyc | Both | Takahashi et al[18,19] |
| Klf4 | Both | Takahashi et al[18,19] |
| Nanog | Human | Yu et al[20] |
| Esrrb | Mouse | Feng et al[73] |
| Glis1 | Both | Maekawa et al[49] |
| E-cadherin | Mouse | Redmer et al[43] |
| shp53 | Both | Hanna et al[39] |
| Lin28 | Both | Hanna et al[39] |
| UTX | Both | Mansour et al[82] |
The common aspiration is that iPS cells will provide an autologous source of cells for a multitude of regenerative medicine therapies in the future and clinical trials using iPS cells have begun[33]. However, the most immediate utility of iPS cell technologies is the ability to study patient-derived cells in the lab. iPS cells present the opportunity to study a range of diseases in novel ways by isolating and reprogramming patient-specific cells and then differentiating them into the cell type of interest. For example, iPS cells have been generated from patients suffering from a wide range of disorders including Duchenne muscular dystrophy, Parkinson’s disease, Huntingdon’s disease, type I diabetes and Down’s syndrome (reviewed in[34]). In addition, cells such as disease-specific cardiomyocytes, which would be difficult to obtain from patients, can also be generated and used to test specific drugs[35]. In summary, the generation of iPS cells has stimulated the growth of a hugely active new area of research with promise to revolutionise medicine. However, the reprogramming process remains extremely inefficient and the basic molecular understanding of a process that does not appear to readily occur in nature is only just being unravelled. A greater understanding of the basic biology will lead to more efficient methodologies for iPS cell reprogramming in vitro and also potentially lead to strategies to therapeutically manipulate differentiated cells in vivo to become stem cells and repair or regenerate diseased tissues.
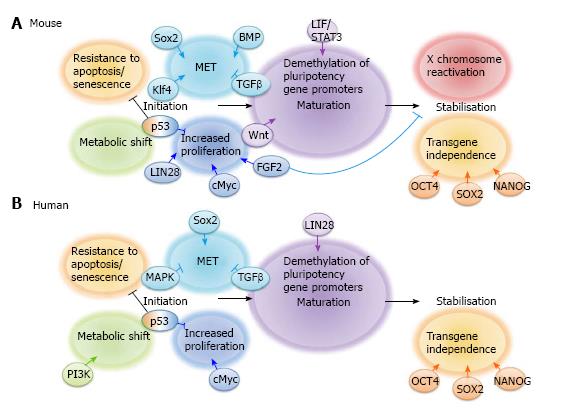
Much progress has been made in recent years to define the molecular mechanisms involved in iPS cell reprogramming. This has led to the general acceptance of the model proposed by Samavarchi-Tehrani et al[36] that reprogramming consists of 3 phases: initiation, maturation and stabilisation (Summarised in Figure 1). Throughout reprogramming various changes occur not only to the cell phenotype but also to gene and non-coding RNA expression, epigenetic status and metabolism. In this review we will focus on cell signalling during the 3 stages of iPS cell reprogramming whilst other aspects are reviewed elsewhere by Papp et al[37] and Jia et al[38].
The initiation phase of reprogramming occurs in virtually all successfully transfected cells[39] and is characterised by somatic genes being switched off by methylation, an increase in cell proliferation, a metabolic switch from oxidative phosphorylation to glycolysis, reactivation of telomerase activity and a mesenchymal-to-epithelial transition (MET)[40]. MET is a feature of both mouse[41] and human[42] somatic cell reprogramming and involves the loss of mesenchymal characteristics such as motility and the acquisition of epithelial characteristics such as cell polarity and expression of the cell adhesion molecule E-CADHERIN, perhaps explaining why E-cadherin can replace Oct4 in the reprogramming process[43]. MET and the opposite transition, epithelial-to-mesenchymal transition (EMT), are key features of embryogenesis[44], tumour metastasis[45] and both mouse[46] and human[47] ES cell differentiation. Interestingly, the MET that marks the initiation of cellular reprogramming is reversible since removal of the reprogramming factors from mouse “pre-iPS” cells after induction of reprogramming has been shown to lead to reversion of the cells to a mesenchymal phenotype[36], thus demonstrating that continued transgene expression is necessary to allow cells to progress to the maturation stage.
Mechanistically, Sox2 suppresses expression of Snail, an EMT inducer[48], and Klf4 induces E-cadherin expression, thus promoting MET[41]. In addition, Maekawa et al[49] have shown that the Glis family zinc finger 1 protein Glis1 can substitute cMyc in the reprogramming cocktail by inducing MET, thus initiating iPS cell reprogramming. MET can also be induced by chemicals, for example, various groups have demonstrated the ability of transforming growth factor (TGF)β inhibition to enhance the initiation stage of both mouse[50,51] and human[42] somatic cell reprogramming. This observation is supported by the finding that addition of recombinant TGFβ abrogates iPS cell formation[42] and is likely due to the EMT-inducing action of TGFβ signalling, which then prevents the MET that is critical to successful iPS cell reprogramming. TGFβ signalling promotes EMT via a wide variety of mechanisms, including mediating the disassembly of junctional complexes, reorganising the cell cytoskeleton, and EMT gene activation[52]. Various TGFβ inhibitors have been used to promote reprogramming, including A-83-01[41,53], E616452[25,50] (also known as RepSox) and SB431542[42] (Table 2). In addition to promoting MET, TGFβ inhibitors promote Nanog expression[50], thus providing 2 potential mechanisms for their ability to enhance reprogramming. Mitogen-activated protein kinase (MAPK) signalling, activated by TGFβ, further induces the expression of mesodermal genes[52]. Inhibitors of MAPK signalling such as PD0325901 have therefore been used in combination with TGFβ inhibitors to promote MET[42].
| Small molecule | Function | Ref. |
| BIX-01294 | Histone methyltransferase inhibitor | Shi et al[51] |
| Bayk8644 | Calcium channel agonist | Shi et al[51] |
| RG108 | DNA methyltransferase inhibitor | Shi et al[51] |
| 5-Aza-2’-Deoxycytidine | DNA methyltransferase inhibitor | Huangfu et al[89] |
| Dexamethasone | Steroid glucocorticoid | Huangfu et al[89] |
| Valproic acid | HDAC inhibitor | Huangfu et al[89] |
| Trichostatin A | HDAC inhibitor | Huangfu et al[89] |
| SAHA | HDAC inhibitor | Huangfu et al[89] |
| PD0325901 + CHIR99021 | MAPK inhibition and GSK3 inhibition | Shi et al[51], Silva et al[77] |
| SB 431542+ PD0325901 | TGFβ inhibitor | Lin et al[42] |
| And MAPK inhibitor | ||
| A-83-01 | TGFβ inhibitor | Li et al[41], Zhu et al[53] |
| E616452 | TGFβ inhibitor | Ichida et al[90] |
| AMI-5 | Protein arginine methyltransferase inhibitor | Yuan et al[13] |
| Kenpaullone | Unknown “novel function” | Lyssiotis et al[91] |
Bone morphogenetic protein (BMP) signalling also plays an important role in the initiation stage of mouse iPS cell reprogramming by promoting MET via upregulation of epithelial genes such as E-cadherin, Occludin and Epithelial cell adhesion molecule[36]. Chen et al[54] have shown that BMPs can replace Klf4 in the reprogramming cocktail, allowing mouse embryonic fibroblasts (MEFs) to be reprogrammed using Oct4 alone. However, constitutive BMP activation prevents human somatic cell reprogramming. This was discovered through the observation that a naturally occurring Alk2 mutation, which causes fibrodysplasia ossificans progressiva in humans, prevents iPS cell reprogramming and that this blockade can be rescued by inhibition of the ALK2 receptor[55].
Increased proliferation has been observed in cells undergoing reprogramming as early as 3 d after induction of reprogramming[56] and is likely to be initiated by cMyc transgene expression[57]. Lin28 expression and p53 knockdown also increase the efficiency of iPS cell reprogramming by stimulating cell proliferation[39]. Specifically, LIN28 has been shown to regulate cell cycle genes such as Cyclin A, Cyclin B and Cdk4[58] whilst p53 induces cell cycle arrest via p21 and thus p53 knockdown promotes proliferation[59].
Fibroblast growth factor (FGF) signalling has also been implicated at the initiation stage[60]. Araki et al[61] show that Fgf4 is upregulated on day 3 after induction of reprogramming in MEFs and Jiao et al[60] show that FGF2 can improve the reprogramming efficiency in the early phases of mouse somatic cell reprogramming, whereas it has adverse effects in the later stages. Mechanistically, this group have shown that FGF2 promotes the early stages of reprogramming through accelerating cell proliferation, facilitating MET and eliminating extracellular collagens. In addition to an increased proliferation rate, the minority of cells that undergo successful reprogramming also exhibit resistance to apoptosis and senescence, by transgene expression[56]. Recent studies have shown that miR-302 expression allows cells to overcome reprogramming-induced senescence[62] and that silencing of the INK4/ARF locus is also likely to be involved, since INK4/ARF blockade improves reprogramming efficiency[63,64]. The INK4/ARF locus encodes tumour suppressor genes that activate the retinoblastoma and p53 pathways. Its inactivation therefore blocks apoptosis and senescence and facilitates reprogramming.
The initiation phase is also characterised by a metabolic switch from oxidative phosphorylation to glycolysis[65] that occurs around 7 d after induction of reprogramming[66] and involves phosphatidylinositol-3-kinase (PI3K)/AKT signalling[53,67]. For example, Chen et al[67] have demonstrated that the PI3K/AKT pathway was activated during reprogramming in parallel with the upregulation of glycolytic gene expression, showing specifically that AKT activated 2 key glycolytic regulators, AS1060 and PFKB2. Zhu et al[53] have also shown that PS48, an activator of the PI3K/AKT pathway, is able to enhance reprogramming by upregulating glycolytic genes. By switching their metabolism from oxidative phosphorylation to anaerobic glycolysis, pre-iPS cells assume an ES cell-like phenotype[68]. ES cells are likely to have developed this form of metabolism as an adaptation to the hypoxic in vivo environment of the early embryo[69]. Interestingly, various groups have shown that iPS cell reprogramming is enhanced by hypoxia[70,71], likely due to the acceleration of this metabolic shift.
Tanabe et al[72] have recently identified the maturation stage of iPS cell reprogramming as being a major bottleneck in the process, which is likely to account for the low efficiency of the process generally. They demonstrate that LIN28, but not NANOG, shp53 or CYCLIN D1, promotes maturation of iPS cells. During maturation, epigenetic changes occur allowing expression of the first pluripotency-associated genes[40]. These genes include Fbxo15, Sall4, Oct4, Nanog and Esrrb. Interestingly, Esrrb has been shown to be sufficient to reprogram MEFs in collaboration with Sox2 and Oct4[73].
LIF/STAT3 signalling is required for the maturation phase of mouse iPS cell reprogramming[74]. Interestingly, pre-iPS cell colony formation has been observed in the absence of LIF, however, beyond day 6 of reprogramming these colonies detach. This is likely due to the requirement that cells undergoing the reprogramming process have for LIF signalling to maintain cMyc expression[75]. In addition, Tang et al[74] demonstrate that LIF/STAT3 activation induces earlier formation of an increased number of pre-iPS cell colonies. Mechanistically, this group demonstrate that LIF/STAT3 signalling is required for demethylation of pluripotency-associated gene promoters. Specifically, STAT3 signalling was shown to directly block the action of the DNA methyltransferase DNMT1 and Histone deacetylases 2, 3 and 8.
Wnt signalling also enhances the maturation phase of mouse somatic cell reprogramming whereby exogenous stimulation of the pathway using Wnt3a between days 6 and 9 after induction of reprogramming enhances the formation of Nanog positive colonies[76]. Various groups have suggested that expression of Nanog is necessary for cells to advance from the maturation phase to the stabilisation stage[39,77] and thus, Samavarchi et al[36] suggest that Nanog expression alone is responsible for mediating the transition from pre-iPS cells to stably reprogrammed cells. This group demonstrate that removal of the reprogramming factors from mouse iPS cells at day 9 after induction of reprogramming did not induce phenotypic reversion. Other groups, however, have reported different time points for the stabilisation stage, including day 11[78,79] and day 16[80], suggesting that this can vary depending on discrete protocols and culture variations. It is clear that there remains substantial information to be learned regarding this critical intermediary step but NANOG appears to play a pivotal role in iPS cell maturation.
Only around 1% of cells that initiate reprogramming make it to the stabilisation stage[72]. This can be explained by the observation made by Golipour et al[81] that not all cells are “stabilisation competent”. This group identify a gene expression signature that distinguishes stabilisation competent and stabilisation incompetent cells and show that stabilisation competent cells require transgene repression to enter this stage. Since the stabilisation stage is characterised by transgene independence, only cells that have activated endogenous pluripotency gene expression are able to maintain pluripotency at this late stage. Endogenous pluripotency gene expression is facilitated by demethylation of pluripotency gene promoters, thus explaining why various DNA and histone methyltransferase inhibitors have been shown to accelerate iPS cell reprogramming, amongst other small molecules (Table 2). This may also explain the ability of the H3K27 demethylase UTX to substitute for some of the original reprogramming factors[82].
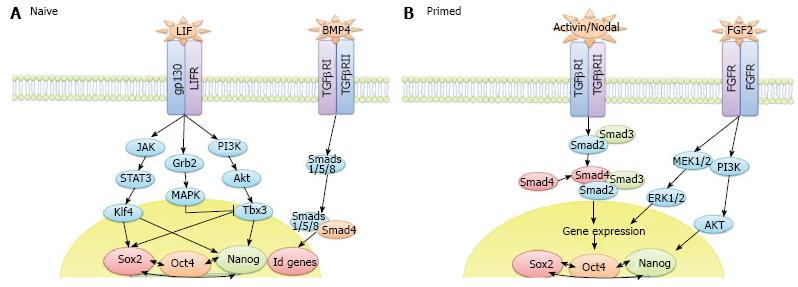
The end-point of iPS cell reprogramming is a matter of some controversy. For example, the stabilisation stage of mouse iPS cell reprogramming involves X chromosome reactivation whereas human iPS cell reprogramming does not[83]. X chromosome inactivation is a process that occurs as female embryonic cells, which have 2 active X chromosomes, commit to differentiation. This feature of human ES and human iPS cells, amongst others (reviewed in[84]), means that they represent the primed pluripotent state. Human iPS cells generated in the presence of ACTIVIN/NODAL and FGF2 ligands are stabilised in this primed state whereas mouse iPS cells reprogrammed in the presence of LIF and BMP4 can be fully reprogrammed to the uncommitted naïve ground state (Figure 2). Interestingly, human dermal fibroblasts (HDFs) have been shown to give rise to naïve human iPS cells when reprogrammed in the presence of LIF, FGF2 and TGFβ1 plus inhibitors of c-Jun NH2-terminal kinase, p38, MAPK and glycogen synthase kinase 3 (3i)[85], thus demonstrating that the cell signalling context is critical to the determination of naïve and primed pluripotency rather than the two states representing a species difference. The derivation of various novel stem cell lines, including intermediate epiblast stem cells which exhibit dual responsiveness to LIF and ACTIVIN/NODAL signalling[86], has challenged the concept of 2 distinct pluripotent states, instead suggesting that a spectrum of pluripotency exists, an idea we develop in Hawkins et al[87]. Thorough investigation into this spectrum of pluripotency, and therefore the transition from pluripotent cells to differentiated cells, should accelerate the delineation of mechanisms occurring throughout the reverse process, from a somatic cell to an iPS cell.
A proposed model for the signalling networks required for the various stages of mouse and human iPS cell reprogramming can be found in Figure 1. However, this knowledge is still vastly incomplete. New technological advances are required to thoroughly interrogate the contribution of a wide range of signalling pathways to somatic cell reprogramming. One of the limitations of many current approaches is the inability to track reprogramming cell signalling in real-time since cells must be sacrificed to obtain data, for example for microarray analysis[36], fluorescence-activated cell sorting or protein extracts[78] at various time points. Some advances have been made to track reprogramming cells in real-time, for example, Smith et al[88] carried out time-lapse imaging with the aim of tracking single cells undergoing the reprogramming process. However, they concluded that this was virtually impossible. We are currently interrogating the role of cell signalling networks in iPS cell reprogramming using a range of GFP reporter HDF lines activated by transcription factors involved in relevant cell signalling pathways. This allows us to monitor signalling pathway activity throughout an entire iPS cell reprogramming experiment in real-time. We anticipate this will enable us to temporally map the contribution of a wide range of signalling pathways to iPS cell reprogramming, thus illuminating this enigmatic biological phenomenon.
P- Reviewer: Imamura M, Niyibizi C, Niu W, Song J S- Editor: Song XX L- Editor: A E- Editor: Lu YJ
| 1. | Evans MJ, Kaufman MH. Establishment in culture of pluripotential cells from mouse embryos. Nature. 1981;292:154-156. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5956] [Cited by in RCA: 5431] [Article Influence: 123.4] [Reference Citation Analysis (0)] |
| 2. | Martin GR. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc Natl Acad Sci USA. 1981;78:7634-7638. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3879] [Cited by in RCA: 3581] [Article Influence: 81.4] [Reference Citation Analysis (0)] |
| 3. | Thomson JA, Itskovitz-Eldor J, Shapiro SS, Waknitz MA, Swiergiel JJ, Marshall VS, Jones JM. Embryonic stem cell lines derived from human blastocysts. Science. 1998;282:1145-1147. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11399] [Cited by in RCA: 10429] [Article Influence: 386.3] [Reference Citation Analysis (0)] |
| 4. | Mitalipov S, Kuo HC, Byrne J, Clepper L, Meisner L, Johnson J, Zeier R, Wolf D. Isolation and characterization of novel rhesus monkey embryonic stem cell lines. Stem Cells. 2006;24:2177-2186. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 82] [Cited by in RCA: 82] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 5. | Buehr M, Meek S, Blair K, Yang J, Ure J, Silva J, McLay R, Hall J, Ying QL, Smith A. Capture of authentic embryonic stem cells from rat blastocysts. Cell. 2008;135:1287-1298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 654] [Cited by in RCA: 584] [Article Influence: 34.4] [Reference Citation Analysis (0)] |
| 6. | Li P, Tong C, Mehrian-Shai R, Jia L, Wu N, Yan Y, Maxson RE, Schulze EN, Song H, Hsieh CL. Germline competent embryonic stem cells derived from rat blastocysts. Cell. 2008;135:1299-1310. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 570] [Cited by in RCA: 511] [Article Influence: 30.1] [Reference Citation Analysis (0)] |
| 7. | Niwa H, Miyazaki J, Smith AG. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat Genet. 2000;24:372-376. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2725] [Cited by in RCA: 2670] [Article Influence: 106.8] [Reference Citation Analysis (0)] |
| 8. | Masui S, Nakatake Y, Toyooka Y, Shimosato D, Yagi R, Takahashi K, Okochi H, Okuda A, Matoba R, Sharov AA. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat Cell Biol. 2007;9:625-635. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 856] [Cited by in RCA: 868] [Article Influence: 48.2] [Reference Citation Analysis (0)] |
| 9. | Chambers I, Silva J, Colby D, Nichols J, Nijmeijer B, Robertson M, Vrana J, Jones K, Grotewold L, Smith A. Nanog safeguards pluripotency and mediates germline development. Nature. 2007;450:1230-1234. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1266] [Cited by in RCA: 1118] [Article Influence: 62.1] [Reference Citation Analysis (0)] |
| 10. | Chambers I, Tomlinson SR. The transcriptional foundation of pluripotency. Development. 2009;136:2311-2322. [PubMed] |
| 11. | Botquin V, Hess H, Fuhrmann G, Anastassiadis C, Gross MK, Vriend G, Schöler HR. New POU dimer configuration mediates antagonistic control of an osteopontin preimplantation enhancer by Oct-4 and Sox-2. Genes Dev. 1998;12:2073-2090. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 242] [Cited by in RCA: 247] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 12. | Nishimoto M, Fukushima A, Okuda A, Muramatsu M. The gene for the embryonic stem cell coactivator UTF1 carries a regulatory element which selectively interacts with a complex composed of Oct-3/4 and Sox-2. Mol Cell Biol. 1999;19:5453-5465. [PubMed] |
| 13. | Yuan H, Corbi N, Basilico C, Dailey L. Developmental-specific activity of the FGF-4 enhancer requires the synergistic action of Sox2 and Oct-3. Genes Dev. 1995;9:2635-2645. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 576] [Cited by in RCA: 572] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 14. | Boyer LA, Lee TI, Cole MF, Johnstone SE, Levine SS, Zucker JP, Guenther MG, Kumar RM, Murray HL, Jenner RG. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell. 2005;122:947-956. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3597] [Cited by in RCA: 3374] [Article Influence: 168.7] [Reference Citation Analysis (0)] |
| 15. | Loh YH, Wu Q, Chew JL, Vega VB, Zhang W, Chen X, Bourque G, George J, Leong B, Liu J. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nat Genet. 2006;38:431-440. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1833] [Cited by in RCA: 1847] [Article Influence: 97.2] [Reference Citation Analysis (0)] |
| 16. | Kopp JL, Ormsbee BD, Desler M, Rizzino A. Small increases in the level of Sox2 trigger the differentiation of mouse embryonic stem cells. Stem Cells. 2008;26:903-911. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 251] [Cited by in RCA: 249] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 17. | Pan G, Li J, Zhou Y, Zheng H, Pei D. A negative feedback loop of transcription factors that controls stem cell pluripotency and self-renewal. FASEB J. 2006;20:1730-1732. [PubMed] |
| 18. | Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663-676. [PubMed] |
| 19. | Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861-872. [PubMed] |
| 20. | Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S, Nie J, Jonsdottir GA, Ruotti V, Stewart R. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917-1920. [PubMed] |
| 21. | Yu J, Hu K, Smuga-Otto K, Tian S, Stewart R, Slukvin II, Thomson JA. Human induced pluripotent stem cells free of vector and transgene sequences. Science. 2009;324:797-801. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1896] [Cited by in RCA: 1692] [Article Influence: 105.8] [Reference Citation Analysis (0)] |
| 22. | Fusaki N, Ban H, Nishiyama A, Saeki K, Hasegawa M. Efficient induction of transgene-free human pluripotent stem cells using a vector based on Sendai virus, an RNA virus that does not integrate into the host genome. Proc Jpn Acad Ser B Phys Biol Sci. 2009;85:348-362. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1024] [Cited by in RCA: 965] [Article Influence: 60.3] [Reference Citation Analysis (0)] |
| 23. | Wang W, Yang J, Liu H, Lu D, Chen X, Zenonos Z, Campos LS, Rad R, Guo G, Zhang S. Rapid and efficient reprogramming of somatic cells to induced pluripotent stem cells by retinoic acid receptor gamma and liver receptor homolog 1. Proc Natl Acad Sci USA. 2011;108:18283-18288. [PubMed] |
| 24. | Zhou H, Wu S, Joo JY, Zhu S, Han DW, Lin T, Trauger S, Bien G, Yao S, Zhu Y. Generation of induced pluripotent stem cells using recombinant proteins. Cell Stem Cell. 2009;4:381-384. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1309] [Cited by in RCA: 1229] [Article Influence: 76.8] [Reference Citation Analysis (0)] |
| 25. | Hou P, Li Y, Zhang X, Liu C, Guan J, Li H, Zhao T, Ye J, Yang W, Liu K. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science. 2013;341:651-654. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 938] [Cited by in RCA: 1024] [Article Influence: 85.3] [Reference Citation Analysis (0)] |
| 26. | Kim JB, Sebastiano V, Wu G, Araúzo-Bravo MJ, Sasse P, Gentile L, Ko K, Ruau D, Ehrich M, van den Boom D. Oct4-induced pluripotency in adult neural stem cells. Cell. 2009;136:411-419. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 712] [Cited by in RCA: 690] [Article Influence: 43.1] [Reference Citation Analysis (0)] |
| 27. | Eminli S, Utikal J, Arnold K, Jaenisch R, Hochedlinger K. Reprogramming of neural progenitor cells into induced pluripotent stem cells in the absence of exogenous Sox2 expression. Stem Cells. 2008;26:2467-2474. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 251] [Cited by in RCA: 246] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 28. | Aasen T, Raya A, Barrero MJ, Garreta E, Consiglio A, Gonzalez F, Vassena R, Bilić J, Pekarik V, Tiscornia G. Efficient and rapid generation of induced pluripotent stem cells from human keratinocytes. Nat Biotechnol. 2008;26:1276-1284. [PubMed] |
| 29. | Hanna J, Markoulaki S, Schorderet P, Carey BW, Beard C, Wernig M, Creyghton MP, Steine EJ, Cassady JP, Foreman R. Direct reprogramming of terminally differentiated mature B lymphocytes to pluripotency. Cell. 2008;133:250-264. [PubMed] |
| 30. | Qin D, Gan Y, Shao K, Wang H, Li W, Wang T, He W, Xu J, Zhang Y, Kou Z. Mouse meningiocytes express Sox2 and yield high efficiency of chimeras after nuclear reprogramming with exogenous factors. J Biol Chem. 2008;283:33730-33735. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 37] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 31. | Okita K, Yamakawa T, Matsumura Y, Sato Y, Amano N, Watanabe A, Goshima N, Yamanaka S. An efficient nonviral method to generate integration-free human-induced pluripotent stem cells from cord blood and peripheral blood cells. Stem Cells. 2013;31:458-466. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 483] [Cited by in RCA: 561] [Article Influence: 46.8] [Reference Citation Analysis (0)] |
| 32. | Stadtfeld M, Brennand K, Hochedlinger K. Reprogramming of pancreatic beta cells into induced pluripotent stem cells. Curr Biol. 2008;18:890-894. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 324] [Cited by in RCA: 279] [Article Influence: 16.4] [Reference Citation Analysis (0)] |
| 33. | Cyranoski D. Stem cells cruise to clinic. Nature. 2013;494:413. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 80] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 34. | Park IH, Arora N, Huo H, Maherali N, Ahfeldt T, Shimamura A, Lensch MW, Cowan C, Hochedlinger K, Daley GQ. Disease-specific induced pluripotent stem cells. Cell. 2008;134:877-886. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1775] [Cited by in RCA: 1608] [Article Influence: 94.6] [Reference Citation Analysis (0)] |
| 35. | Matsa E, Rajamohan D, Dick E, Young L, Mellor I, Staniforth A, Denning C. Drug evaluation in cardiomyocytes derived from human induced pluripotent stem cells carrying a long QT syndrome type 2 mutation. Eur Heart J. 2011;32:952-962. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 328] [Cited by in RCA: 285] [Article Influence: 20.4] [Reference Citation Analysis (0)] |
| 36. | Samavarchi-Tehrani P, Golipour A, David L, Sung HK, Beyer TA, Datti A, Woltjen K, Nagy A, Wrana JL. Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell. 2010;7:64-77. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 792] [Cited by in RCA: 801] [Article Influence: 53.4] [Reference Citation Analysis (0)] |
| 37. | Papp B, Plath K. Epigenetics of reprogramming to induced pluripotency. Cell. 2013;152:1324-1343. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 241] [Article Influence: 20.1] [Reference Citation Analysis (0)] |
| 38. | Jia W, Chen W, Kang J. The functions of microRNAs and long non-coding RNAs in embryonic and induced pluripotent stem cells. Genomics Proteomics Bioinformatics. 2013;11:275-283. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 33] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 39. | Hanna J, Saha K, Pando B, van Zon J, Lengner CJ, Creyghton MP, van Oudenaarden A, Jaenisch R. Direct cell reprogramming is a stochastic process amenable to acceleration. Nature. 2009;462:595-601. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 797] [Cited by in RCA: 763] [Article Influence: 47.7] [Reference Citation Analysis (0)] |
| 40. | David L, Polo JM. Phases of reprogramming. Stem Cell Res. 2014;12:754-761. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 97] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 41. | Li R, Liang J, Ni S, Zhou T, Qing X, Li H, He W, Chen J, Li F, Zhuang Q. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell. 2010;7:51-63. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 895] [Cited by in RCA: 924] [Article Influence: 61.6] [Reference Citation Analysis (0)] |
| 42. | Lin T, Ambasudhan R, Yuan X, Li W, Hilcove S, Abujarour R, Lin X, Hahm HS, Hao E, Hayek A. A chemical platform for improved induction of human iPSCs. Nat Methods. 2009;6:805-808. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 460] [Cited by in RCA: 432] [Article Influence: 27.0] [Reference Citation Analysis (0)] |
| 43. | Redmer T, Diecke S, Grigoryan T, Quiroga-Negreira A, Birchmeier W, Besser D. E-cadherin is crucial for embryonic stem cell pluripotency and can replace OCT4 during somatic cell reprogramming. EMBO Rep. 2011;12:720-726. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 229] [Cited by in RCA: 238] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 44. | Yang J, Weinberg RA. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell. 2008;14:818-829. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2157] [Cited by in RCA: 2335] [Article Influence: 137.4] [Reference Citation Analysis (0)] |
| 45. | Yao D, Dai C, Peng S. Mechanism of the mesenchymal-epithelial transition and its relationship with metastatic tumor formation. Mol Cancer Res. 2011;9:1608-1620. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 282] [Cited by in RCA: 338] [Article Influence: 24.1] [Reference Citation Analysis (0)] |
| 46. | Spencer HL, Eastham AM, Merry CL, Southgate TD, Perez-Campo F, Soncin F, Ritson S, Kemler R, Stern PL, Ward CM. E-cadherin inhibits cell surface localization of the pro-migratory 5T4 oncofetal antigen in mouse embryonic stem cells. Mol Biol Cell. 2007;18:2838-2851. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 87] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 47. | Eastham AM, Spencer H, Soncin F, Ritson S, Merry CL, Stern PL, Ward CM. Epithelial-mesenchymal transition events during human embryonic stem cell differentiation. Cancer Res. 2007;67:11254-11262. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 215] [Cited by in RCA: 232] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 48. | Liu X, Sun H, Qi J, Wang L, He S, Liu J, Feng C, Chen C, Li W, Guo Y. Sequential introduction of reprogramming factors reveals a time-sensitive requirement for individual factors and a sequential EMT-MET mechanism for optimal reprogramming. Nat Cell Biol. 2013;15:829-838. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 190] [Article Influence: 15.8] [Reference Citation Analysis (0)] |
| 49. | Maekawa M, Yamaguchi K, Nakamura T, Shibukawa R, Kodanaka I, Ichisaka T, Kawamura Y, Mochizuki H, Goshima N, Yamanaka S. Direct reprogramming of somatic cells is promoted by maternal transcription factor Glis1. Nature. 2011;474:225-229. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 297] [Cited by in RCA: 302] [Article Influence: 21.6] [Reference Citation Analysis (0)] |
| 50. | Maherali N, Hochedlinger K. Tgfbeta signal inhibition cooperates in the induction of iPSCs and replaces Sox2 and cMyc. Curr Biol. 2009;19:1718-1723. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 273] [Cited by in RCA: 279] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 51. | Shi Y, Do JT, Desponts C, Hahm HS, Schöler HR, Ding S. A combined chemical and genetic approach for the generation of induced pluripotent stem cells. Cell Stem Cell. 2008;2:525-528. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 544] [Cited by in RCA: 501] [Article Influence: 29.5] [Reference Citation Analysis (0)] |
| 52. | Thiery JP, Sleeman JP. Complex networks orchestrate epithelial-mesenchymal transitions. Nat Rev Mol Cell Biol. 2006;7:131-142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2893] [Cited by in RCA: 3087] [Article Influence: 162.5] [Reference Citation Analysis (0)] |
| 53. | Zhu S, Li W, Zhou H, Wei W, Ambasudhan R, Lin T, Kim J, Zhang K, Ding S. Reprogramming of human primary somatic cells by OCT4 and chemical compounds. Cell Stem Cell. 2010;7:651-655. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 506] [Cited by in RCA: 516] [Article Influence: 36.9] [Reference Citation Analysis (0)] |
| 54. | Chen J, Liu J, Yang J, Chen Y, Chen J, Ni S, Song H, Zeng L, Ding K, Pei D. BMPs functionally replace Klf4 and support efficient reprogramming of mouse fibroblasts by Oct4 alone. Cell Res. 2011;21:205-212. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 105] [Cited by in RCA: 113] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 55. | Hamasaki M, Hashizume Y, Yamada Y, Katayama T, Hohjoh H, Fusaki N, Nakashima Y, Furuya H, Haga N, Takami Y. Pathogenic mutation of ALK2 inhibits induced pluripotent stem cell reprogramming and maintenance: mechanisms of reprogramming and strategy for drug identification. Stem Cells. 2012;30:2437-2449. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 53] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 56. | Papp B, Plath K. Reprogramming to pluripotency: stepwise resetting of the epigenetic landscape. Cell Res. 2011;21:486-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 135] [Cited by in RCA: 132] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 57. | Apostolou E, Hochedlinger K. Chromatin dynamics during cellular reprogramming. Nature. 2013;502:462-471. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 327] [Cited by in RCA: 303] [Article Influence: 25.3] [Reference Citation Analysis (0)] |
| 58. | Xu B, Zhang K, Huang Y. Lin28 modulates cell growth and associates with a subset of cell cycle regulator mRNAs in mouse embryonic stem cells. RNA. 2009;15:357-361. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 156] [Cited by in RCA: 163] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 59. | el-Deiry WS, Tokino T, Velculescu VE, Levy DB, Parsons R, Trent JM, Lin D, Mercer WE, Kinzler KW, Vogelstein B. WAF1, a potential mediator of p53 tumor suppression. Cell. 1993;75:817-825. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6088] [Cited by in RCA: 6305] [Article Influence: 197.0] [Reference Citation Analysis (0)] |
| 60. | Jiao J, Dang Y, Yang Y, Gao R, Zhang Y, Kou Z, Sun XF, Gao S. Promoting reprogramming by FGF2 reveals that the extracellular matrix is a barrier for reprogramming fibroblasts to pluripotency. Stem Cells. 2013;31:729-740. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 41] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 61. | Araki R, Jincho Y, Hoki Y, Nakamura M, Tamura C, Ando S, Kasama Y, Abe M. Conversion of ancestral fibroblasts to induced pluripotent stem cells. Stem Cells. 2010;28:213-220. [PubMed] |
| 62. | Banito A, Rashid ST, Acosta JC, Li S, Pereira CF, Geti I, Pinho S, Silva JC, Azuara V, Walsh M. Senescence impairs successful reprogramming to pluripotent stem cells. Genes Dev. 2009;23:2134-2139. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 473] [Cited by in RCA: 492] [Article Influence: 30.8] [Reference Citation Analysis (0)] |
| 63. | Li H, Collado M, Villasante A, Strati K, Ortega S, Cañamero M, Blasco MA, Serrano M. The Ink4/Arf locus is a barrier for iPS cell reprogramming. Nature. 2009;460:1136-1139. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 793] [Cited by in RCA: 766] [Article Influence: 47.9] [Reference Citation Analysis (0)] |
| 64. | Utikal J, Polo JM, Stadtfeld M, Maherali N, Kulalert W, Walsh RM, Khalil A, Rheinwald JG, Hochedlinger K. Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature. 2009;460:1145-1148. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 654] [Cited by in RCA: 658] [Article Influence: 41.1] [Reference Citation Analysis (0)] |
| 65. | Panopoulos AD, Yanes O, Ruiz S, Kida YS, Diep D, Tautenhahn R, Herrerías A, Batchelder EM, Plongthongkum N, Lutz M. The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Res. 2012;22:168-177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 383] [Cited by in RCA: 402] [Article Influence: 28.7] [Reference Citation Analysis (0)] |
| 66. | Park SJ, Yeo HC, Kang NY, Kim H, Lin J, Ha HH, Vendrell M, Lee JS, Chandran Y, Lee DY. Mechanistic elements and critical factors of cellular reprogramming revealed by stepwise global gene expression analyses. Stem Cell Res. 2014;12:730-741. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 50] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 67. | Chen M, Zhang H, Wu J, Xu L, Xu D, Sun J, He Y, Zhou X, Wang Z, Wu L. Promotion of the induction of cell pluripotency through metabolic remodeling by thyroid hormone triiodothyronine-activated PI3K/AKT signal pathway. Biomaterials. 2012;33:5514-5523. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 36] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 68. | Kondoh H, Lleonart ME, Nakashima Y, Yokode M, Tanaka M, Bernard D, Gil J, Beach D. A high glycolytic flux supports the proliferative potential of murine embryonic stem cells. Antioxid Redox Signal. 2007;9:293-299. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 289] [Cited by in RCA: 265] [Article Influence: 14.7] [Reference Citation Analysis (0)] |
| 69. | Ottosen LD, Hindkaer J, Husth M, Petersen DE, Kirk J, Ingerslev HJ. Observations on intrauterine oxygen tension measured by fibre-optic microsensors. Reprod Biomed Online. 2006;13:380-385. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 70] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 70. | Shimada H, Hashimoto Y, Nakada A, Shigeno K, Nakamura T. Accelerated generation of human induced pluripotent stem cells with retroviral transduction and chemical inhibitors under physiological hypoxia. Biochem Biophys Res Commun. 2012;417:659-664. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 28] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 71. | Yoshida Y, Takahashi K, Okita K, Ichisaka T, Yamanaka S. Hypoxia enhances the generation of induced pluripotent stem cells. Cell Stem Cell. 2009;5:237-241. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 574] [Cited by in RCA: 567] [Article Influence: 35.4] [Reference Citation Analysis (0)] |
| 72. | Tanabe K, Nakamura M, Narita M, Takahashi K, Yamanaka S. Maturation, not initiation, is the major roadblock during reprogramming toward pluripotency from human fibroblasts. Proc Natl Acad Sci USA. 2013;110:12172-12179. [PubMed] |
| 73. | Feng B, Jiang J, Kraus P, Ng JH, Heng JC, Chan YS, Yaw LP, Zhang W, Loh YH, Han J. Reprogramming of fibroblasts into induced pluripotent stem cells with orphan nuclear receptor Esrrb. Nat Cell Biol. 2009;11:197-203. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 358] [Cited by in RCA: 353] [Article Influence: 22.1] [Reference Citation Analysis (0)] |
| 74. | Tang Y, Tian XC. JAK-STAT3 and somatic cell reprogramming. JAKSTAT. 2013;2:e24935. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 28] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 75. | Yang J, van Oosten AL, Theunissen TW, Guo G, Silva JC, Smith A. Stat3 activation is limiting for reprogramming to ground state pluripotency. Cell Stem Cell. 2010;7:319-328. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 209] [Cited by in RCA: 194] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 76. | Ho R, Papp B, Hoffman JA, Merrill BJ, Plath K. Stage-specific regulation of reprogramming to induced pluripotent stem cells by Wnt signaling and T cell factor proteins. Cell Rep. 2013;3:2113-2126. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 73] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 77. | Silva J, Nichols J, Theunissen TW, Guo G, van Oosten AL, Barrandon O, Wray J, Yamanaka S, Chambers I, Smith A. Nanog is the gateway to the pluripotent ground state. Cell. 2009;138:722-737. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 859] [Cited by in RCA: 782] [Article Influence: 48.9] [Reference Citation Analysis (0)] |
| 78. | Hansson J, Rafiee MR, Reiland S, Polo JM, Gehring J, Okawa S, Huber W, Hochedlinger K, Krijgsveld J. Highly coordinated proteome dynamics during reprogramming of somatic cells to pluripotency. Cell Rep. 2012;2:1579-1592. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 194] [Cited by in RCA: 203] [Article Influence: 16.9] [Reference Citation Analysis (0)] |
| 79. | Polo JM, Anderssen E, Walsh RM, Schwarz BA, Nefzger CM, Lim SM, Borkent M, Apostolou E, Alaei S, Cloutier J. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell. 2012;151:1617-1632. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 729] [Cited by in RCA: 671] [Article Influence: 51.6] [Reference Citation Analysis (0)] |
| 80. | Brambrink T, Foreman R, Welstead GG, Lengner CJ, Wernig M, Suh H, Jaenisch R. Sequential expression of pluripotency markers during direct reprogramming of mouse somatic cells. Cell Stem Cell. 2008;2:151-159. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 672] [Cited by in RCA: 608] [Article Influence: 35.8] [Reference Citation Analysis (0)] |
| 81. | Golipour A, David L, Liu Y, Jayakumaran G, Hirsch CL, Trcka D, Wrana JL. A late transition in somatic cell reprogramming requires regulators distinct from the pluripotency network. Cell Stem Cell. 2012;11:769-782. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 126] [Cited by in RCA: 124] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 82. | Mansour AA, Gafni O, Weinberger L, Zviran A, Ayyash M, Rais Y, Krupalnik V, Zerbib M, Amann-Zalcenstein D, Maza I. The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature. 2012;488:409-413. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 273] [Cited by in RCA: 279] [Article Influence: 21.5] [Reference Citation Analysis (0)] |
| 83. | Plath K, Lowry WE. Progress in understanding reprogramming to the induced pluripotent state. Nat Rev Genet. 2011;12:253-265. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 241] [Cited by in RCA: 215] [Article Influence: 15.4] [Reference Citation Analysis (0)] |
| 84. | Nichols J, Smith A. Naive and primed pluripotent states. Cell Stem Cell. 2009;4:487-492. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1599] [Cited by in RCA: 1377] [Article Influence: 86.1] [Reference Citation Analysis (0)] |
| 85. | Gafni O, Weinberger L, Mansour AA, Manor YS, Chomsky E, Ben-Yosef D, Kalma Y, Viukov S, Maza I, Zviran A. Derivation of novel human ground state naive pluripotent stem cells. Nature. 2013;504:282-286. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 764] [Cited by in RCA: 834] [Article Influence: 69.5] [Reference Citation Analysis (0)] |
| 86. | Chang KH, Li M. Clonal isolation of an intermediate pluripotent stem cell state. Stem Cells. 2013;31:918-927. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 23] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 87. | Hawkins K, Keramari M, Soncin F, Segal JM, Mohamet L, Miazga N, Ritson S, Bobola N, Merry CLR, Ward CM. Novel cell lines isolated from mES cells exhibiting de novo methylation of the E-cadherin promoter. CMB. 2014;In press. |
| 88. | Smith AG. Embryo-derived stem cells: of mice and men. Annu Rev Cell Dev Biol. 2001;17:435-462. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 937] [Cited by in RCA: 855] [Article Influence: 37.2] [Reference Citation Analysis (0)] |
| 89. | Huangfu D, Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nat Biotechnol. 2008;26:79-797. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1320] [Cited by in RCA: 1199] [Article Influence: 70.5] [Reference Citation Analysis (0)] |
| 90. | Ichida JK, A small-molecule inhibitor of Tgf-β signaling replaces Sox2 in reprogramming by inducing Nanog. Cell Stem Cell. 2009;5:491-503. [PubMed] [DOI] [Full Text] |
| 91. | Lyssiotis CA, Reprogramming of murine fibroblasts to induced pluripotent stem cells with chemical complementation of Klf4. Proc Natl Acad Sci. 2009;106:8912-8917. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 303] [Cited by in RCA: 282] [Article Influence: 17.6] [Reference Citation Analysis (0)] |