Published online Dec 15, 2003. doi: 10.3748/wjg.v9.i12.2839
Revised: May 19, 2003
Accepted: June 2, 2003
Published online: December 15, 2003
AIM: To investigate the mutation of EDNRB gene and EDN-3 gene in sporadic Hirschsprung’s disease (HD) in Chinese population.
METHODS: Genomic DNA was extracted from bowel tissues of 34 unrelated HD patients which were removed by surgery. Exon 3, 4, 6 of EDNRB gene and Exon 1, 2 of EDN-3 gene were amplified by polymerase chain reaction (PCR) and analyzed by single strand conformation polymorphism (SSCP).
RESULTS: EDNRB mutations were detected in 2 of the 13 short-segment HD. One mutant was in the exon 3, the other was in the exon 6. EDN-3 mutation was detected in one of the 13 short-segment HD and in the exon 2. Both EDNRB and EDN-3 mutations were detected in one short-segment HD. No mutations were detected in the ordinary or long-segment HD.
CONCLUSION: The mutations of EDNRB gene and EDN-3 gene are found in the short-segment HD of sporadic Hirschsprung’s disease in Chinese population, which suggests that the EDNRB gene and EDN-3 gene play important roles in the pathogenesis of HD.
- Citation: Duan XL, Zhang XS, Li GW. Clinical relationship between EDN-3 gene, EDNRB gene and Hirschsprung’s disease. World J Gastroenterol 2003; 9(12): 2839-2842
- URL: https://www.wjgnet.com/1007-9327/full/v9/i12/2839.htm
- DOI: https://dx.doi.org/10.3748/wjg.v9.i12.2839
Hirschsprung’s disease (HD) is a congenital malformation with an incidence of one in 5000 live newborns. The absence of intramural intestinal ganglia of Meissner and Auerbach results in poor coordination of propulsive movement, and hence functional intestinal obstruction. Patients were treated surgically with removal of the affected intestine[1-4]. In 1994, two major genes associated with HD were recognized. First, in the RET (receptor tyrosin kinase) gene, there are inactivating mutations in isolated HD. RET accounts for up to 20% of sporadic and 50% of familial cases[5-11]. The second major gene is the EDNRB (endothelin receptor B) gene. EDNRB accounts for 5%-10% of all HD cases. Heterozygous mutations in the EDNRB gene were reported in nonsyndromic HD[12-16]. The preferred ligand for the G-protein coupled transmembranous receptor EDNRB was EDN-3 (endothelin-3). The interaction between EDN-3 and EDNRB was reported to be essential for normal development of enteric ganglia. The importance of the EDN-3-EDNRB interaction in promoting the normal development of neural crest cells has been clearly demonstrated. Human mutations in the EDN-3 gene have been reported recently: heterozygous missense mutations in two cases of sporadic HD[17-19]. However, there are fewer reports about mutations of EDNRB gene and EDN-3 gene in HD in Chinese population. In order to further investigate the pathogenic mechanism of HD, we examined mutations on exon 3, 4, 6 of EDNRB gene and exon 1, 2 of EDN-3 gene in 34 sporadic HD cases with the single strand conformation ploymorphism analysis of polymerase chain reaction products (PCR-SSCP).
Thirty-four specimens of sporadic HD cases were collected after operation in the Second Hospital of Xi’an Jiaotong University between 1999 and 2001, and the pathological statement was approved pathologically. There are four cases of long-segment HD, seventeen cases of common HD, and thirteen cases of short-segment HD based on Romen’s division. At the same time, normal recta and sigmoid flexure tissues were collected, serving as a control group. All specimens were put into liquid nitrogen to freeze quickly after cut off in 15 minutes, and stored at a temperature of -80 °C. DNA was extracted according to the standard protocols.
The designed primers were synthesized by Bioasia Company. The specific primer sequences of exon 3, 4, 6 of EDNRB gene and exon 1, 2 of EDN-3 gene are summarized in Table 1. The PCR mixture (total volume: 30 μL) contained 2 μL of template-DNA, 3 μL of 10×PCR buffer, 3 μL of 2.5 mmol/L MgCl2, 3 μL of 2.5 mmol/L dNTPs, 1 μL each of two fragment-specific primers, 17 μL of triplex distilled water, and 1 unit of Taq DNA polymerase. Thirty-five PCR amplification cycles were performed with the following condition of temperature: 94 °C for 35 seconds, 55 °C for 50 seconds and 72 °C for 1 minute. Amplifications were performed with a final extension for 10 minutes at 72 °C. The amplified fragments were run in 15 g/L agarose gel, and were confirmed to be in existence.
| Gene | Exon | Primer | bp | |
| EDNRB | 3 | Forwoard | ATCTTCAGATATCGAGCTGTT | 223 bp |
| Reverse | TGAAATTTACCTGCATGAAAG | |||
| 4 | Forwoard | ATCCCTATAGTTTTACAAGACAGC | 170 bp | |
| Reverse | ATTTTCTTACCTGCTTTAGGTG | |||
| 6 | Forwoard | ACAGAAGCTACAATGACTAC | 240 bp | |
| Reverse | GAAAGGCTTATATTTGAGCC | |||
| EDN-3 | 1 | Forwoard | CAAGCGGCCGTCCTCCTGGTCCGGT | 180 bp |
| Reverse | CTTCTCCGCGCCTCGGTCC | |||
| 2A | Forwoard | CCCTCCTCAGGTGTTTGGG | 239 bp | |
| Reverse | TCGGCCGCCTGCTCCTGC | |||
| 2B | Forwoard | TGGCGAGGAGACTGTGGCT | 218 bp | |
| Reverse | TGGCGAGGAGACTGTGGCT |
A conventional electrophoresis apparatus (PC-3000 Mini Electrophoresis Unit; Bio-Red Company, USA) was used with a constant temperature of 10 °C for SSCP. For SSCP, the PCR products were heated for 10 min at 94 °C, transferred into an ice-cold water bath for 3 min, and then run on 60 g/L polyacrylamide gel for 3 hours. The gel was stained by ethidium bromide for 10 - 20 min to visualize DNA band patterns.
EDNRB gene PCR products The increment of all DNA samples from HD patients was a single strand with a length of 223 bp, 170 bp and 240 bp and so was the normal control, which indicated that a large fragment insertion and deletion did not exist in the region of EDNRB gene exon 3, 4 and 6 among 34 HD patients.
EDN-3 gene PCR products The increment of all DNA samples from HD patients was a single strand with a length of 180 bp, 239 bp and 219 bp and so was the normal control, which indicated that a large fragment insertion and deletion did not exist in the region of EDN-3 gene exon 1, 2A and 2B among 34 HD patients.
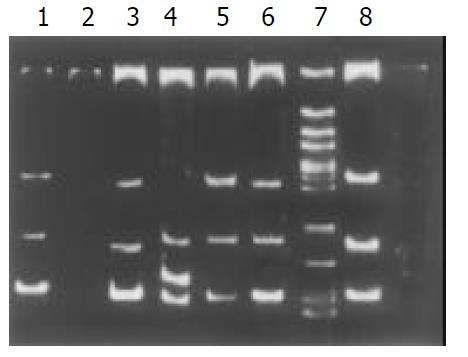
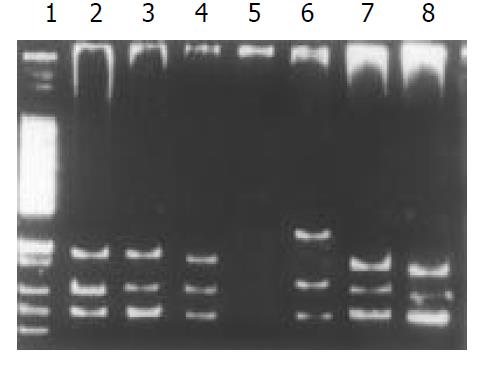
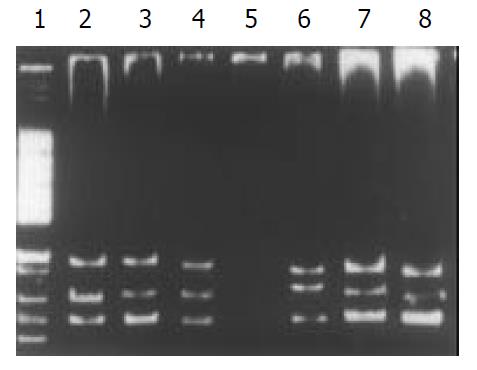
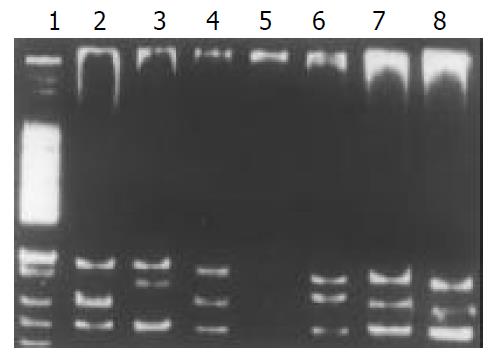
Among the 34 HD patients, abnormal SSCP migration patterns were found in 2 unrelated HD. EDNRB mutations were detected in 2 of the 17 short-segment HD. One mutant was in the exon 3 (Figure 1), the other was in the exon 6 (Figure 2). EDN-3 mutation was detected in 1 of the 17 short-segment HD and in the exon 2 (Figure 3). Both EDNRB mutation and EDN-3 mutation were detected in one short-segment HD (Figure 4). The mutation was absent in the ordinary, long-segment HD and normal control group samples.
The endothelin peptide family of secreted peptides comprises four members to date: EDN-1, EDN-2, EDN-3, and VIP (vasoactive intestinal polypeptide)[20,21]. A diverse set of pharmacologic activities with different potencies are exerted by endothelin family peptides, suggesting the existence of endothelin receptor subtypes.
The EDNRB gene encodes a heptahelical receptor that is involved in the G-protein-mediated intracellular signaling pathway. The human EDNRB gene lies on chromosome band 13q22 and comprises 7 exons, with a length of about 24 kb[22-27]. The predicted protein had 442 amino acids with a transmembrane topology similar to that of other G protein-coupled receptors which, when activated by a ligand, induce a calcium flux into the cells. Activation of EDNRB may result in upregulation of secretion of the endothelins, thereby amplifying their effects.
The EDN-3 gene encodes a large inactive preproendothelin-3 precursor which yields a biologically active 21 amino acid peptide containing four cysteines involved in two disulphide bonds. The human EDN-3 gene lies on chromosome band 20q13.3 and comprises 5 exons[28-30]. The EDN-3 is produced by a two-step proteolytic cleavage of a larger precursor molecule, preproendothelin. This molecule is enzymatically processed to an inactive progenitor (big endothelin) which is subsequently converted to the active peptide by a specific endothelin-converting enzyme 1. The mature peptide mediates its effect through two receptors, one of which is the EDNRB.
We have examined mutations on exon 3, 4, 6 of EDNRB gene and exon 1, 2 of EDN-3 gene in 34 sporadic Chinese HD patients with PCR-SSCP. The PCR result revealed that the increment of all DNA samples from HD patients was a single strand with a length of 223 bp, 170 bp and 240 bp and so was that from normal control, which indicated that a large fragment insertion and deletion did not exist in the region of EDNRB gene exon 3, 4 and 6 among 34 HD patients. And the increment of all DNA samples from HD patients was a single strand with a length of 180 bp, 239 bp and 219 bp and so was that from normal control, which indicated that a large fragment insertion and deletion did not exist in the region of EDN-3 gene exon 1, 2A and 2B among 34 HD patients. Among the 34 HD patients, we found abnormal SSCP migration patterns in 2 unrelated HD. EDNRB mutations were detected in 2 of the 17 short-segment HDs. One mutant was in the exon 3, the other in the exon 6. EDN-3 mutation was detected in 1 of the 17 short-segment HD and in the exon 2. Both EDNRB and EDN-3 mutations were detected in one short-segment HD. The mutation is absent in the ordinary, long-segment HD and normal control group samples. Due to the mutation of EDNRB, there would be no upregulation of secretion of the endothelins and amplification of endothelin effect, and the total amount of endothelin produced would be too small to initiate migration. Alteration of the structure of the preproendothelin by the mutation may conceivably result in a less efficient cleavage, or even a complete failure of cleavage of the preproendothelin, resulting in EDN-3 deficiency during development. This might lead to an incomplete colonization of the bowel by ganglion cells. We should bear in mind that EDNRB is the receptor for EDN-3, so it is reasonable to assume that the mutations of EDNRB and EDN-3 caused the maldevelopment of the enteric nervous system.
Similar to the receptor-ligand relationship between RET and GDNF observed in the etiology of some HD patients, in human fetuses, both EDNRB and EDN-3 have been demonstrated on enteric neurons and gut mesenchyme cells[31], suggesting that EDN-3 and EDNRB may regulate interactions between neural crest and gut mesenchyme cells, necessary for normal migration. There are reports on HSCR patients with GDNF-RET or NTN-RET gene mutation combinations, as well as a case with mutations in both RET and EDNRB[32-39]. So far, there has been no report on an EDN-3 mutation in combination with a mutation in other HSCR genes. In the present study, we found an EDN-3 mutation in combination with an EDNRB mutation in one short-segment HD patient.
To date, at least 16 different mutations or alterations of the EDNRB gene and 4 different mutations or alterations of the EDN-3 gene have been identified in HD patients. A variety of frameshift, nonsense, or missense mutations scattered along EDNRB gene and EDN-3 gene has been identified in HD patients[40,41]. The combined results of our study for mutations in EDN3 and EDNRB may indicate the contributions of these genes to the HD phenotype. EDNRB and EDN3 mutations seem to account for a minority of cases. The majority of HSCR cases cannot be explained by mutations in any of the genes analysed so far, suggesting that other genes or additional factors may contribute to the occurrence of HD phenotype and that HD is a multifactorial disease.
Edited by Ma JY
| 1. | Bates MD. Development of the enteric nervous system. Clin Perinatol. 2002;29:97-114. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 2. | Amiel J, Lyonnet S. Hirschsprung disease, associated syndromes, and genetics: a review. J Med Genet. 2001;38:729-739. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 345] [Cited by in RCA: 286] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 3. | Meier-Ruge WA, Brunner LA. Morphometric assessment of Hirschsprung's disease: associated hypoganglionosis of the colonic myenteric plexus. Pediatr Dev Pathol. 2001;4:53-61. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 18] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 4. | Li JC, Mi KH, Zhou JL, Busch L, Kuhnel W. The development of colon innervation in trisomy 16 mice and Hirschsprung's disease. World J Gastroenterol. 2001;7:16-21. [PubMed] |
| 5. | Borrego S, Wright FA, Fernández RM, Williams N, López-Alonso M, Davuluri R, Antiñolo G, Eng C. A founding locus within the RET proto-oncogene may account for a large proportion of apparently sporadic Hirschsprung disease and a subset of cases of sporadic medullary thyroid carcinoma. Am J Hum Genet. 2003;72:88-100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 74] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 6. | Griseri P, Pesce B, Patrone G, Osinga J, Puppo F, Sancandi M, Hofstra R, Romeo G, Ravazzolo R, Devoto M. A rare haplotype of the RET proto-oncogene is a risk-modifying allele in hirschsprung disease. Am J Hum Genet. 2002;71:969-974. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 7. | Sancandi M, Ceccherini I, Costa M, Fava M, Chen B, Wu Y, Hofstra R, Laurie T, Griffths M, Burge D. Incidence of RET mutations in patients with Hirschsprung's disease. J Pediatr Surg. 2000;35:139-142; discussion 142-143. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 47] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 8. | Fitze G, Cramer J, Ziegler A, Schierz M, Schreiber M, Kuhlisch E, Roesner D, Schackert HK. Association between c135G/A genotype and RET proto-oncogene germline mutations and phenotype of Hirschsprung's disease. Lancet. 2002;359:1200-1205. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 77] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 9. | Pasini B, Rossi R, Ambrosio MR, Zatelli MC, Gullo M, Gobbo M, Collini P, Aiello A, Pansini G, Trasforini G. RET mutation profile and variable clinical manifestations in a family with multiple endocrine neoplasia type 2A and Hirschsprung's disease. Surgery. 2002;131:373-381. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 35] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 10. | Julies MG, Moore SW, Kotze MJ, du Plessis L. Novel RET mutations in Hirschsprung's disease patients from the diverse South African population. Eur J Hum Genet. 2001;9:419-423. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 21] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 11. | Shimotake T, Go S, Inoue K, Tomiyama H, Iwai N. A homozygous missense mutation in the tyrosine E kinase domain of the RET proto-oncogene in an infant with total intestinal aganglionosis. Am J Gastroenterol. 2001;96:1286-1291. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 17] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 12. | Newby DE, Strachan FE, Webb DJ. Abnormal endothelin B receptor vasomotor responses in patients with Hirschsprung's disease. QJM. 2002;95:159-163. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 13. | von Boyen GB, Krammer HJ, Süss A, Dembowski C, Ehrenreich H, Wedel T. Abnormalities of the enteric nervous system in heterozygous endothelin B receptor deficient (spotting lethal) rats resembling intestinal neuronal dysplasia. Gut. 2002;51:414-419. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 33] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 14. | Zaahl MG, du Plessis L, Warnich L, Kotze MJ, Moore SW. Significance of novel endothelin-B receptor gene polymorphisms in Hirschsprung's disease: predominance of a novel variant (561C/T) in patients with co-existing Down's syndrome. Mol Cell Probes. 2003;17:49-54. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 17] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 15. | Fuchs S, Amiel J, Claudel S, Lyonnet S, Corvol P, Pinet F. Functional characterization of three mutations of the endothelin B receptor gene in patients with Hirschsprung's disease: evidence for selective loss of Gi coupling. Mol Med. 2001;7:115-124. [PubMed] |
| 16. | Matsushima Y, Shinkai Y, Kobayashi Y, Sakamoto M, Kunieda T, Tachibana M. A mouse model of Waardenburg syndrome type 4 with a new spontaneous mutation of the endothelin-B receptor gene. Mamm Genome. 2002;13:30-35. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 48] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 17. | Bidaud C, Salomon R, Van Camp G, Pelet A, Attié T, Eng C, Bonduelle M, Amiel J, Nihoul-Fékété C, Willems PJ. Endothelin-3 gene mutations in isolated and syndromic Hirschsprung disease. Eur J Hum Genet. 1997;5:247-251. [PubMed] |
| 18. | Pingault V, Bondurand N, Lemort N, Sancandi M, Ceccherini I, Hugot JP, Jouk PS, Goossens M. A heterozygous endothelin 3 mutation in Waardenburg-Hirschsprung disease: is there a dosage effect of EDN3/EDNRB gene mutations on neurocristopathy phenotypes. J Med Genet. 2001;38:205-209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 39] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 19. | Svensson PJ, Von Tell D, Molander ML, Anvret M, Nordenskjöld A. A heterozygous frameshift mutation in the endothelin-3 (EDN-3) gene in isolated Hirschsprung's disease. Pediatr Res. 1999;45:714-717. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 20] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 20. | Goraca A. New views on the role of endothelin (minireview). Endocr Regul. 2002;36:161-167. [PubMed] |
| 21. | Milla PJ. Endothelins, pseudo-obstruction and Hirschsprung's disease. Gut. 1999;44:148-149. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 22. | Gariepy CE, Williams SC, Richardson JA, Hammer RE, Yanagisawa M. Transgenic expression of the endothelin-B receptor prevents congenital intestinal aganglionosis in a rat model of Hirschsprung disease. J Clin Invest. 1998;102:1092-1101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 140] [Cited by in RCA: 130] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 23. | Shanske A, Ferreira JC, Leonard JC, Fuller P, Marion RW. Hirschsprung disease in an infant with a contiguous gene syndrome of chromosome 13. Am J Med Genet. 2001;102:231-236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 43] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 24. | Shin MK, Levorse JM, Ingram RS, Tilghman SM. The temporal requirement for endothelin receptor-B signalling during neural crest development. Nature. 1999;402:496-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 270] [Cited by in RCA: 257] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 25. | Tanoue A, Koshimizu TA, Tsuchiya M, Ishii K, Osawa M, Saeki M, Tsujimoto G. Two novel transcripts for human endothelin B receptor produced by RNA editing/alternative splicing from a single gene. J Biol Chem. 2002;277:33205-33212. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 11] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 26. | Syrris P, Carter ND, Patton MA. Novel nonsense mutation of the endothelin-B receptor gene in a family with Waardenburg-Hirschsprung disease. Am J Med Genet. 1999;87:69-71. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 27. | Tanaka H, Moroi K, Iwai J, Takahashi H, Ohnuma N, Hori S, Takimoto M, Nishiyama M, Masaki T, Yanagisawa M. Novel mutations of the endothelin B receptor gene in patients with Hirschsprung's disease and their characterization. J Biol Chem. 1998;273:11378-11383. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 62] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 28. | Dupin E, Glavieux C, Vaigot P, Le Douarin NM. Endothelin 3 induces the reversion of melanocytes to glia through a neural crest-derived glial-melanocytic progenitor. Proc Natl Acad Sci USA. 2000;97:7882-7887. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 125] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 29. | Kenny SE, Hofstra RM, Buys CH, Vaillant CR, Lloyd DA, Edgar DH. Reduced endothelin-3 expression in sporadic Hirschsprung disease. Br J Surg. 2000;87:580-585. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 19] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 30. | Woodward MN, Kenny SE, Vaillant C, Lloyd DA, Edgar DH. Time-dependent effects of endothelin-3 on enteric nervous system development in an organ culture model of Hirschsprung's disease. J Pediatr Surg. 2000;35:25-29. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 35] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 31. | McCallion AS, Chakravarti A. EDNRB/EDN3 and Hirschsprung disease type II. Pigment Cell Res. 2001;14:161-169. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 78] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 32. | Angrist M, Bolk S, Halushka M, Lapchak PA, Chakravarti A. Germline mutations in glial cell line-derived neurotrophic factor (GDNF) and RET in a Hirschsprung disease patient. Nat Genet. 1996;14:341-344. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 182] [Cited by in RCA: 174] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 33. | Carrasquillo MM, McCallion AS, Puffenberger EG, Kashuk CS, Nouri N, Chakravarti A. Genome-wide association study and mouse model identify interaction between RET and EDNRB pathways in Hirschsprung disease. Nat Genet. 2002;32:237-244. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 200] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 34. | Auricchio A, Griseri P, Carpentieri ML, Betsos N, Staiano A, Tozzi A, Priolo M, Thompson H, Bocciardi R, Romeo G. Double heterozygosity for a RET substitution interfering with splicing and an EDNRB missense mutation in Hirschsprung disease. Am J Hum Genet. 1999;64:1216-1221. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 57] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 35. | Tomiyama H, Shimotake T, Ono S, Kimura O, Tokiwa K, Iwai N. Relationship between the type of RET/GDNF/NTN or SOX10 gene mutations and long-term results after surgery for total colonic aganglionosis with small bowel involvement. J Pediatr Surg. 2001;36:1685-1688. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 36. | Svensson PJ, Anvret M, Molander ML, Nordenskjöld A. Phenotypic variation in a family with mutations in two Hirschsprung-related genes (RET and endothelin receptor B). Hum Genet. 1998;103:145-148. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 22] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 37. | Inoue K, Shimotake T, Tomiyama H, Iwai N. Mutational analysis of the RET and GDNF gene in children with hypoganglionosis. Eur J Pediatr Surg. 2001;11:120-123. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 21] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 38. | Ivanchuk SM, Myers SM, Eng C, Mulligan LM. De novo mutation of GDNF, ligand for the RET/GDNFR-alpha receptor complex, in Hirschsprung disease. Hum Mol Genet. 1996;5:2023-2026. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 87] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 39. | McCallion AS, Stames E, Conlon RA, Chakravarti A. Phenotype variation in two-locus mouse models of Hirschsprung disease: tissue-specific interaction between Ret and Ednrb. Proc Natl Acad Sci USA. 2003;100:1826-1831. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 115] [Cited by in RCA: 114] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 40. | Parisi MA, Kapur RP. Genetics of Hirschsprung disease. Curr Opin Pediatr. 2000;12:610-617. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 161] [Cited by in RCA: 140] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 41. | Gath R, Goessling A, Keller KM, Koletzko S, Coerdt W, Müntefering H, Wirth S, Hofstra RM, Mulligan L, Eng C. Analysis of the RET, GDNF, EDN3, and EDNRB genes in patients with intestinal neuronal dysplasia and Hirschsprung disease. Gut. 2001;48:671-675. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 50] [Article Influence: 2.1] [Reference Citation Analysis (0)] |