Published online Dec 21, 2019. doi: 10.3748/wjg.v25.i47.6823
Peer-review started: September 23, 2019
First decision: November 4, 2019
Revised: November 15, 2019
Accepted: November 29, 2019
Article in press: November 29, 2019
Published online: December 21, 2019
Processing time: 88 Days and 7.6 Hours
Gastric adenocarcinoma (GAC) mortality rates have remained relatively changed over the past 30 years, and it continues to be one of the leading causes of cancer-related death.
To search for novel miRNAs related to GAC prognosis and further investigate the effect of miR-96-5p on MGC-803 cells.
The miRNA expression profile data of GAC based on The Cancer Genome Atlas were obtained and used to screen differently expressed miRNAs (DEMs) and DEMs related to GAC prognosis. Then, the expression of DEMs related to GAC prognosis was identified in GAC tumor samples and adjacent normal samples by qRT-PCR. The target gene, ZDHHC5, of miR-96-5p was predicted using TargetScan, miRTarBase, and miRDB databases and confirmed by luciferase reporter assay. Furthermore, MGC-803 cells were transfected with inhibitor NC, miR-96-5p inhibitor, si-ZDHHC5, or miR-96-5p inhibitor + si-ZDHHC5, and then cell apoptosis was detected by flow cytometry. The expression of ZDHHC5, Bcl-2, and COX-2 was detected using western blotting.
A total of 299 DEMs and 35 DEMs related to GAC prognosis were screened based on The Cancer Genome Atlas. Then compared with adjacent normal samples, the levels of miR-96-5p, miR-222-5p, and miR-652-5p were remarkably increased, while miR-125-5p, miR-145-3p, and miR-379-3p levels were reduced in GAC tumor samples (P < 0.01), which were consistent with bioinformatics analysis. Furthermore, ZDHHC5 was defined as a direct target gene of miR-96-5p. miR-96-5p inhibition increased the number of apoptotic cells as well as promoted the expression of ZDHHC5, Bcl-2, and COX-2 in MGC-803 cells (P < 0.01). After ZDHHC5 inhibition, the number of apoptotic cells and the expression of ZDHHC5, Bcl-2, and COX-2 were reduced. The addition of an miR-96-5p inhibitor partly reversed these effects (P < 0.01).
Our findings identified six miRNAs related to GAC prognosis and suggested that downregulated miR-96-5p might induce cell apoptosis via upregulating ZDHHC5 expression in MGC-803 cells.
Core tip: Gastric adenocarcinoma (GAC) is the most common malignant tumor. It is important to further reveal novel diagnostic and therapeutic methods as well as the underlying molecular mechanism of GAC. This study aimed to search for novel miRNAs related to GAC prognosis. Six miRNAs related to prognosis, including miR-96-5p, miR-125-5p, miR-145-3p, miR-222-5p, miR-379-3p, and miR-652-5p, were identified in GAC samples. Furthermore, downregulated miR-96-5p markedly induced cell apoptosis through targeting ZDHHC5. Current findings provide a potential molecular mechanism of miR-96-5p in GAC.
- Citation: Zhou HY, Wu CQ, Bi EX. MiR-96-5p inhibition induces cell apoptosis in gastric adenocarcinoma. World J Gastroenterol 2019; 25(47): 6823-6834
- URL: https://www.wjgnet.com/1007-9327/full/v25/i47/6823.htm
- DOI: https://dx.doi.org/10.3748/wjg.v25.i47.6823
Gastric adenocarcinoma (GAC) is the most common malignant tumor originating in the stomach and is counted as one of the top ten common cancers worldwide, with approximately 951000 diagnosed cases and 723000 deaths in 2012[1,2]. Currently, the common and effective therapeutic method is the combination of surgery and adjuvant radiation therapy or chemotherapy, which have improved the 5-year survival rate of GAC[3]. However, delayed diagnosis occurs in most patients with proximal or distant metastasis due to the nontypical symptoms of early GAC, which results in poor treatment and prognosis[4]. Therefore, it is important to further reveal novel diagnostic and therapeutic methods as well as the underlying molecular mechanism of GAC.
It is widely known that a major challenge for the treatment of GAC is poor prognosis, and environmental exposure and gene mutation have been identified to be associated with this outcome[5]. Plenty of evidence indicates that the poor prognosis of GAC is significantly related to many molecular biomarkers, such as microRNAs (miRNA)[6,7]. miRNAs, as endogenous noncoding small-molecule RNAs, widely exist in severe conditions[8]. It is known to function in post-transcriptional regulation of gene expression through binding 3’-untranslated region of their target mRNA, accordingly modulating various key cell biological processes, such as embryonic development, tumor cell proliferation, differentiation, and apoptosis[8,9].
Previous studies have demonstrated that miRNA dysregulation significantly influences the prognosis of gastric cancer patients (e.g., miRNA-203[10], miR-21[11], and miR-25[12]. Imaoka et al[10] reported that a low serum miR-203 expression is associated with poor prognosis and may be a noninvasive biomarker for prognosis of gastric cancer patients. Simonian et al[11] observed that circulating miR-21 may be considered as a diagnostic and prognostic biomarker in gastric cancer. In addition, Li et al[12] revealed that miR-25 is associated with the prognosis of gastric cancer and can induce cell migration and proliferation by targeting transducer of ERBB2.1. Thus, it is essential to search for more novel miRNAs related to GAC prognosis, which may contribute to the development of GAC diagnosis.
In the current study, the miRNA expression profile data of GAC based on The Cancer Genome Atlas (TCGA) were analyzed to screen differently expressed miRNAs (DEMs) and DEMs related to GAC prognosis. Furthermore, DEMs were identified in clinical samples, and the mechanism of DEM was investigated in vitro. According to this, we aimed to search for new therapeutic targets for GAC and provide some useful insights in improving the prognosis of GAC patients.
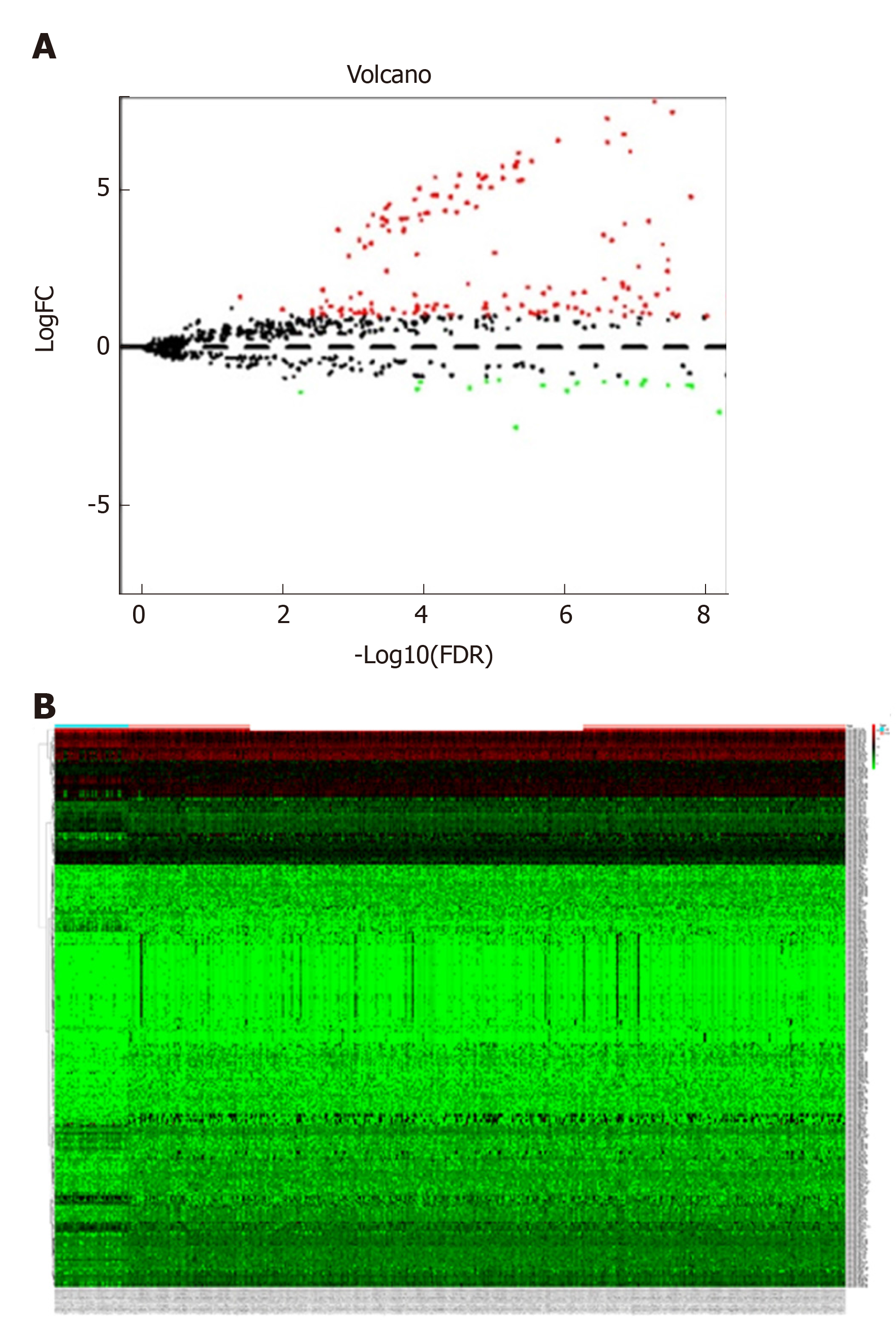
The miRNA expression profile data (level 3, processed and standardized data) and the corresponding clinical information of GAC were downloaded from TCGA (https://portal.gdc.cancer.gov/) on February 11, 2019 based on the platform of Illumina HiSeq 2000 RNA Sequencing platform. A total of 452 samples were obtained from this dataset, including 410 GAC tumor samples and 42 normal control samples. The edgeR package in R was utilized to screen DEMs between GAC samples and normal samples. The thresholds were defined as false discovery rate < 0.05 and |log fold change| > 1. Meanwhile, volcano plots and heat maps were generated based on the obtained DEMs.
The overall survival time was individually extracted from clinical information. Then, combined with the overall survival times and the expression levels of DEMs, DEMs related to prognosis were screened using KMsurv package of R, with the threshold of log-rank P < 0.05.
This study obtained ethical approval from the ethics committee of Jinan Seventh People's Hospital, and the study was performed according to the Helsinki Declaration. A total of 20 paired tumor tissues and adjacent normal tissues (distance of 3-4 cm from the tumor tissue) were collected from GAC patients who underwent surgery in Jinan Seventh People's Hospital between September 2018 to September 2019. The specimens were confirmed by hematoxylin eosin staining and stored in RNA later. In addition, 5 mL peripheral blood was obtained from these 20 GAC patients. Meanwhile, the same amount of peripheral blood was extracted from 20 paired healthy subjects. Written informed consent from all participants was obtained, and the clinical information, including age, weight, gender, distant metastasis, lymph node metastasis, depth of invasion, and TNM stage are shown in Table 1.
| Variables | Patients, n = 20 | Controls, n = 20 | P value1 | ||
| n | % | n | % | ||
| Age in yr, mean ± SD | 62.1 ± 5.2 | 60.1 ± 10.2 | 0.73 | ||
| Weight in kg, mean ± SD | 64.0 ± 7.1 | 69.1 ± 6.6 | 0.88 | ||
| Gender | |||||
| Male | 13 | 65.0 | 1 | 55.0 | 0.64 |
| Female | 7 | 35.0 | 9 | 45.0 | |
| Depth of invasion | |||||
| T1/T2 | 7 | 35.0 | |||
| T3/T4 | 13 | 65.0 | |||
| Lymph node metastasis | |||||
| N0 | 6 | 30.0 | |||
| N1/N2/N3 | 14 | 70.0 | |||
| Distant metastasis | |||||
| M0 | 8 | 40.0 | |||
| M1 | 12 | 60.0 | |||
| TNM stage | |||||
| I/II | 7 | 35.0 | |||
| III/IV | 13 | 65.0 | |||
Target genes of DEMs related to prognosis were predicted using the three online analysis databases, including miRDB, miRTarBase, and TargetScan. Overlapping target genes among the three tools were selected to make the bioinformatic analysis more reliable. Then, the intersection of the predicting target genes among the three databases was obtained using a Venn diagram online tool, and the target genes that overlapped in the three databases were considered as a potential target gene of DEM.
Human gastric carcinoma cell line MGC-803 was obtained from Shanghai Obio Technology Co., Ltd. The cells were maintained in Dulbecco's Modified Eagle Media (DMEM, Gibco, Carlsbad, CA, United States) supplemented with 10% fetal bovine serum (FBS, Gibco, Carlsbad, CA, United States). The construction of ZDHHC5 silence vector (si-ZDHHC5) was performed by GenePharma (Shanghai, China). The miR-96-5p inhibitor and inhibitor NC were purchased from Thermo (Waltham, MA, United States). MGC-803 cells were inoculated in six-well plates for 24 h with approximately 5 × 105 cells in each well, and then inhibitor NC, miR-96-5p inhibitor, si-ZDHHC5, or miR-96-5p inhibitor + si-ZDHHC5 was transfected into MGC-803 cells by Lipofectamine 2000 (Invitrogen, Carlsbad, CA, United States) per the manufacturer's instructions. Meanwhile, MGC-803 cells without transfection served as the control group. Cells were harvested after 48 h of transfection to perform follow-up experiments.
Total RNA from tissues and peripheral blood was obtained by Trizol (Invitrogen) and RNeasy Plus Mini Kit (Qiagen, Valencia, CA, United States) per the manufacturer's instructions. The concentration of RNA was detected using NanoDrop ND-2000 (Invitrogen). To generate cDNA, Mir-X™ miRNA FirstStrand Synthesis Kit (Takara, Dalian, China) was used. The quantitative polymerase chain reaction (qPCR) was carried out using the SYBR Premix ExTaq TM II (Takara) by ABI 7900 qRT-PCR System (Applied Biosystems, Foster City, CA, United States). The primer sequences are listed in Table 2. U6 and glyceraldehyde-phosphate dehydrogenase were used as the internal control of measuring miR-19a and ADIPOR2 expression. Data were analyzed by the 2-ΔΔCt method.
| Gene | Primer sequence |
| miR-96-5p | F:5’-TCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGCAAAAA-3’ |
| R: 5’- ACACTCCAGCTGGGTTTGGCACTAGCACATT-3’ | |
| miR-125a-5p | F: 5’- CCCTGAGACCCTTTAACCT-3’ |
| R: 5’- GTCCAGTTTTTTTTTTTTTTTCACAG-3’ | |
| miR-145-3p | F: 5’- GGTCCAGTTTTCCCAGGA-3’ |
| R: 5’- CCAGTTTTTTTTTTTTTTTAGGGATTC-3’ | |
| miR-222-5p | F: 5’- GCTCAGTAGCCAGTGTAGA-3’ |
| R: 5’- GTCCAGTTTTTTTTTTTTTTTAGGATCT-3’ | |
| miR-379-3p | F: 5’- GCAGTGGTAGACTATGGAAC-3’ |
| R: 5’- GGTCCAGTTTTTTTTTTTTTTTCCT-3’ | |
| miR-652-5p | F: 5’- CCTAGGAGAGGGTGCCA-3’ |
| R: 5’- GTCCAGTTTTTTTTTTTTTTTGAATGG-3’ | |
| miR-708-3p | F: 5’- GCAACTAGACTGTGAGCTTC-3’ |
| R: 5’- GGTCCAGTTTTTTTTTTTTTTTCTAGA-3’ | |
| GAPDH | F: 5’-AGAAGGCTGGGGCTCATTTG-3’ |
| R: 5’-AGGGGCCATCCACAGTCTTC-3’ | |
| U6 | F: 5’-AGGGGCCATCCACAGTCTTC-3’ |
| R: 5’-AACGCTTCACGAATTTGCGT-3’ |
The target gene of miR-96-5p was verified using the luciferase assay. The 3’-untranslated region of ZDHHC5 was cloned into a pGL3-basic vector, named as Luc-ZDHHC5. Luc-ZDHHC5 and phRL-TK plasmid were co-transfected with miR-96-5p mimic, miR-96-5p NC (negative control), or siZDHHC5 (positive control) (synthesized by Biosyntech, Suzhou, China) into 293T cells. After 48 h of transfection, the relative luciferase activity was measured by the Dual-Glo Luciferase Assay System (Promega) in accordance to the manufacturer's introductions. Renilla luciferase activity was used to normalize luciferase activity.
Total proteins were isolated by RIPA Lysis Buffer (Beyotime, Shanghai, China). Proteins concentrations were tested by bicinchoninic acid kit (Beyotime). The protein sample was separated on SDS-PAGE gel, transferred to polyvinylidene fluoride membranes, and followed by the blockage with 5% nonfat milk for 1 h. Next, the membranes were probed with primary antibodies of ZDHHC5 (1:1000, Proteintech), Bcl-2 (1:1000, Abcam), COX-2 (1:1000, Abcam), and GAPDH (1:1000, Beyotime) overnight at 4 °C. Then, membranes were incubated with secondary antibody (1:1000, Beyotime) for 2 h in a dark room at room temperature. GAPDH was used as the control protein. Enhanced chemiluminescence Plus reagent (Beyotime) was used to image blots. The band quantification was performed using Image J software.
Annexin V-FITC Apoptosis Detection kit was used to evaluate cell apoptosis. MGC-803 cells were grown in 6-well plates for 24 h and then transfected with inhibitor NC, miR-96-5p inhibitor, si-ZDHHC5, or miR-96-5p inhibitor + si-ZDHHC5 for 48 h. Next, cells were digested with trypsin and washed with PBS, followed by resuspending in 1 × Binding Buffer, and stained with propidium iodide and FITC-Annexin V for 15 min at 25 °C in the dark. Cells were finally detected using a flow cytometer (Beckman Coulter, Fullerton, CA, United States).
Statistical analysis was conducted using SPSS Statistics software 22.0 (Chicago, IL, United States). Continuous variables were expressed as mean ± standard deviation and analyzed by independent-samples t test. Categorical variables were expressed as percentages and assessed by two-sided chi-square test. The differences of multiple groups were performed by one-way ANOVA following with post-hoc of Dunnett t test. P < 0.05 was considered to be statistically significant.
Based on the selective criteria, a total of 299 DEMs were identified between GAC and normal control samples, including 225 upregulated and 74 downregulated miRNAs. As shown in Figure 1A and 1B, volcano plots and heat maps were conducted for these 299 DEMs.
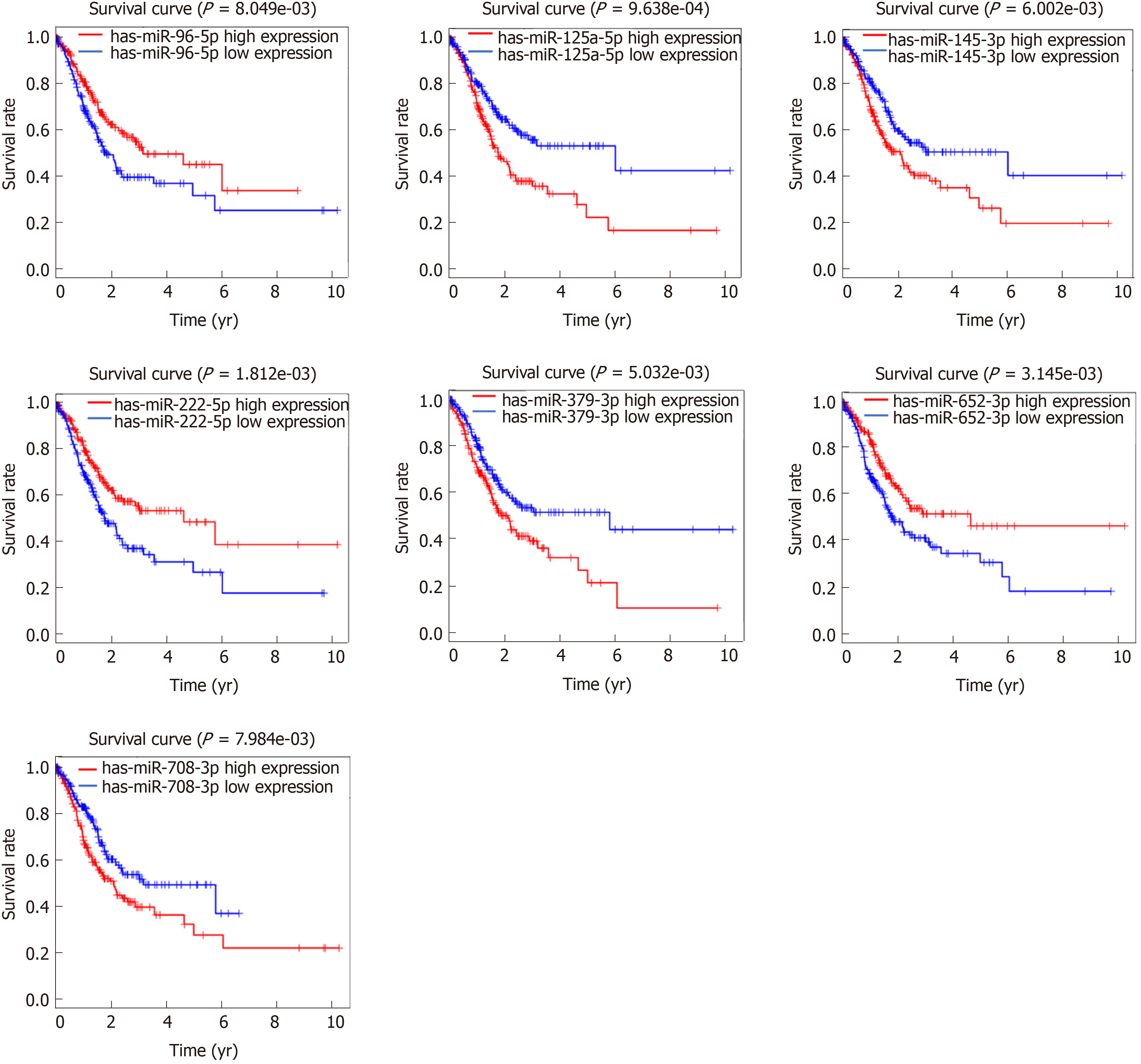
Based on these 299 DEMs, the relationships between patient overall survival and miRNA expression were evaluated, and the results showed that 35 DEMs were significantly related to the prognosis of GAC patients (P < 0.05). Among these DEMs, seven miRNAs had a higher association with GAC prognosis (P < 0.01), including miR-96-5p (P = 8.049 × 10-3), miR-125-5p (P = 9.638 × 10-4), miR-145-3p (P = 6.002 × 10-3), miR-222-5p (P = 1.812 × 10-3), miR-379-3p (P = 5.032 × 10-3), miR-652-5p (P = 3.145 × 10-3), and miR-708-3p (P = 7.984 × 10-3) (Figure 2).
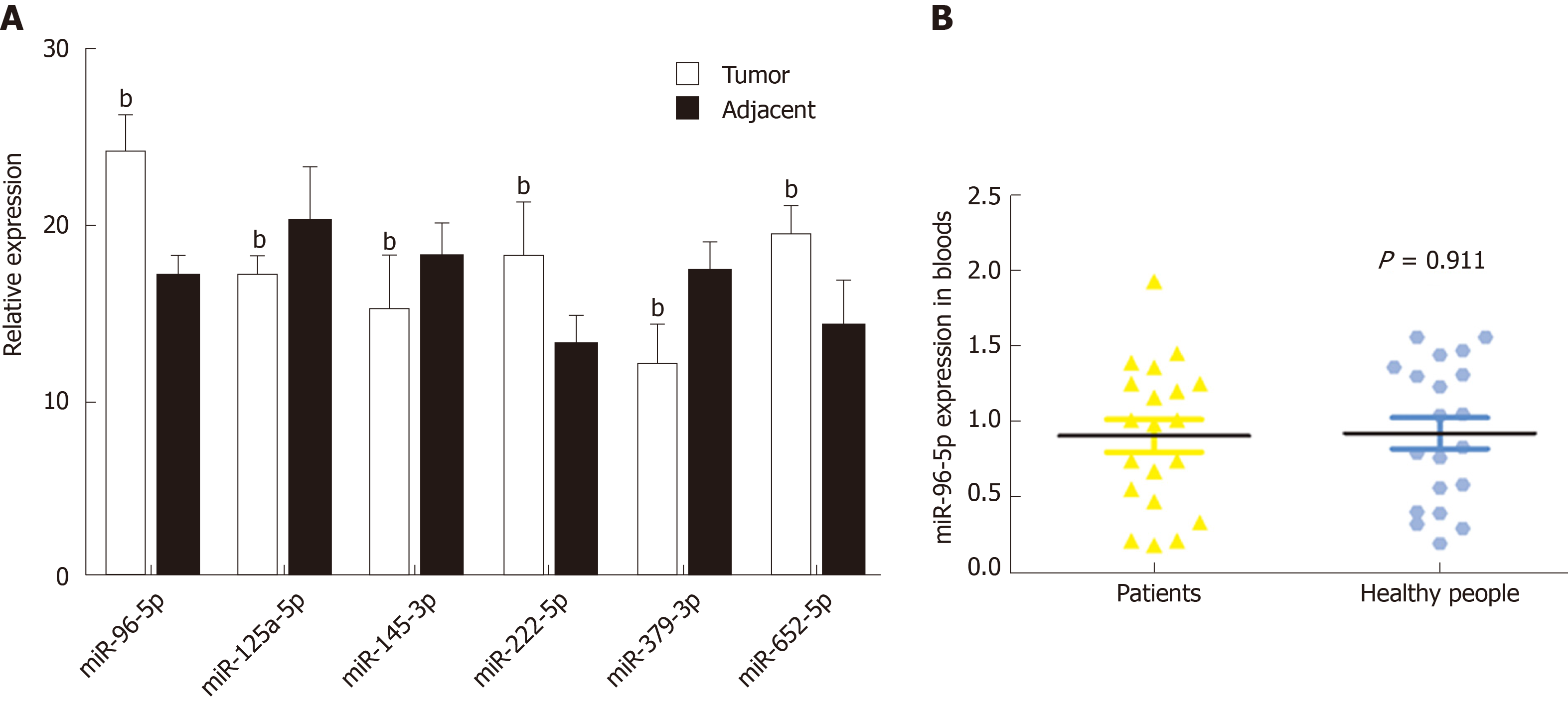
A total of 20 GAC patients and 20 healthy subjects were included in this study. No significant difference was found in age, weight, and gender between GAC patients and healthy subjects (Table 1). Based on the above survival analysis, six miRNAs were selected for identification in GAC tumor samples and adjacent normal samples. qRT-PCR revealed that compared with adjacent normal samples, the levels of miR-96-5p, miR-222-5p, and miR-652-5p were remarkably increased, while miR-125-5p, miR-145-3p, and miR-379-3p levels were obviously reduced in GAC tumor samples (P < 0.01, Figure 3A), which was consistent with bioinformatics analysis results by TCGA. Moreover, miR-96-5p level was detected in the blood of GAC patients and healthy subjects, but no significant difference was found (Figure 3B).
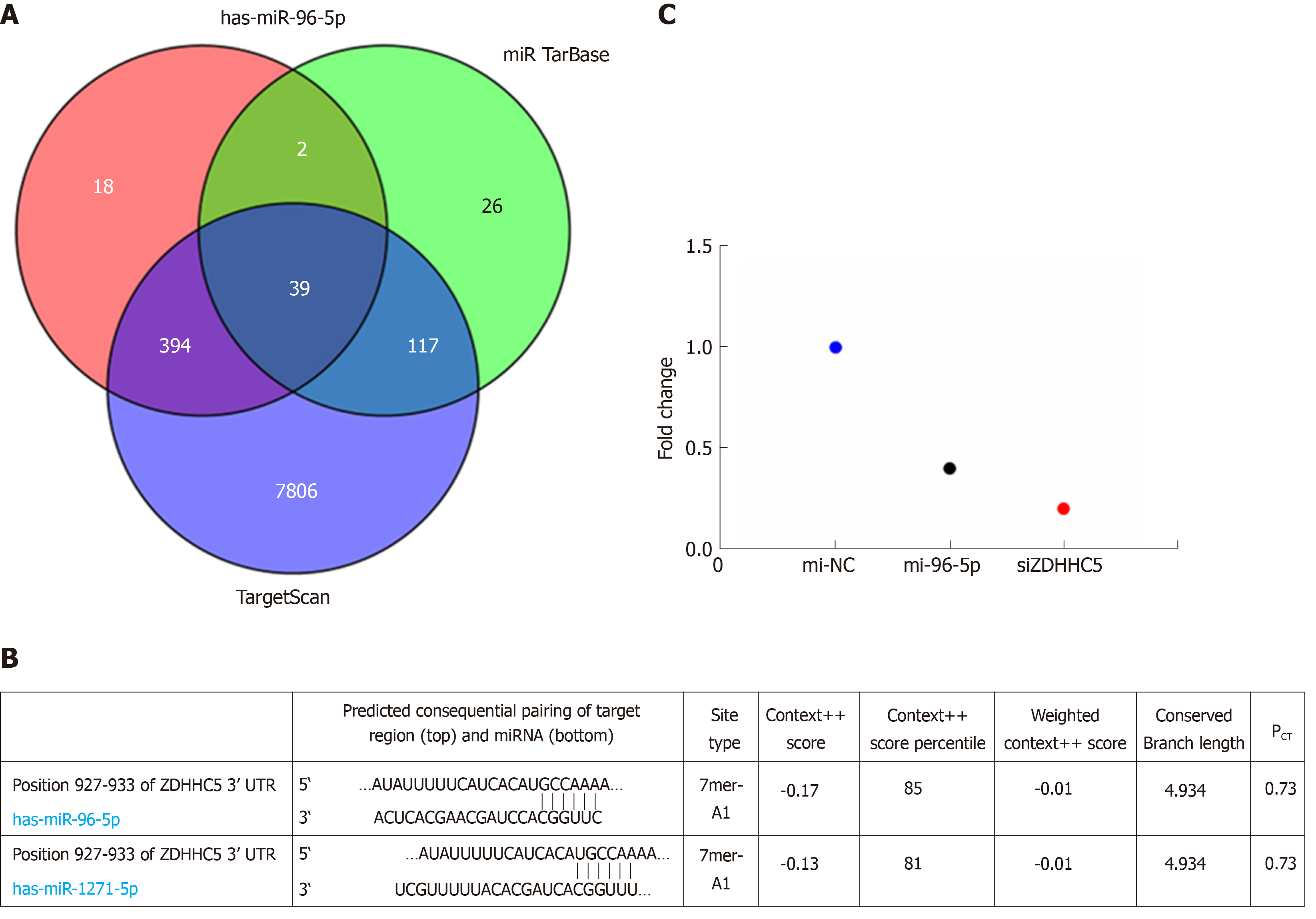
Considering miR-96-5p had the highest association with GAC prognosis, the function of miR-96-5p was investigated in the following experiments. The results found that a total of 39 overlapped target genes existed in TargetScan, miRTarBase, and miRDB databases (Figure 4A). Based on this bioinformatics analysis, ZDHHC5 was considered as a potential target gene of miR-96-5p (Figure 4B). Luciferase receptor assay showed that the relative luciferase activity was reduced after co-transfection with miR-96-5p mimic or siZDHHC5 compared with co-transfection with miR-96-5p NC (Figure 4C), which suggested ZDHHC5 was a direct target gene of miR-96-5p.
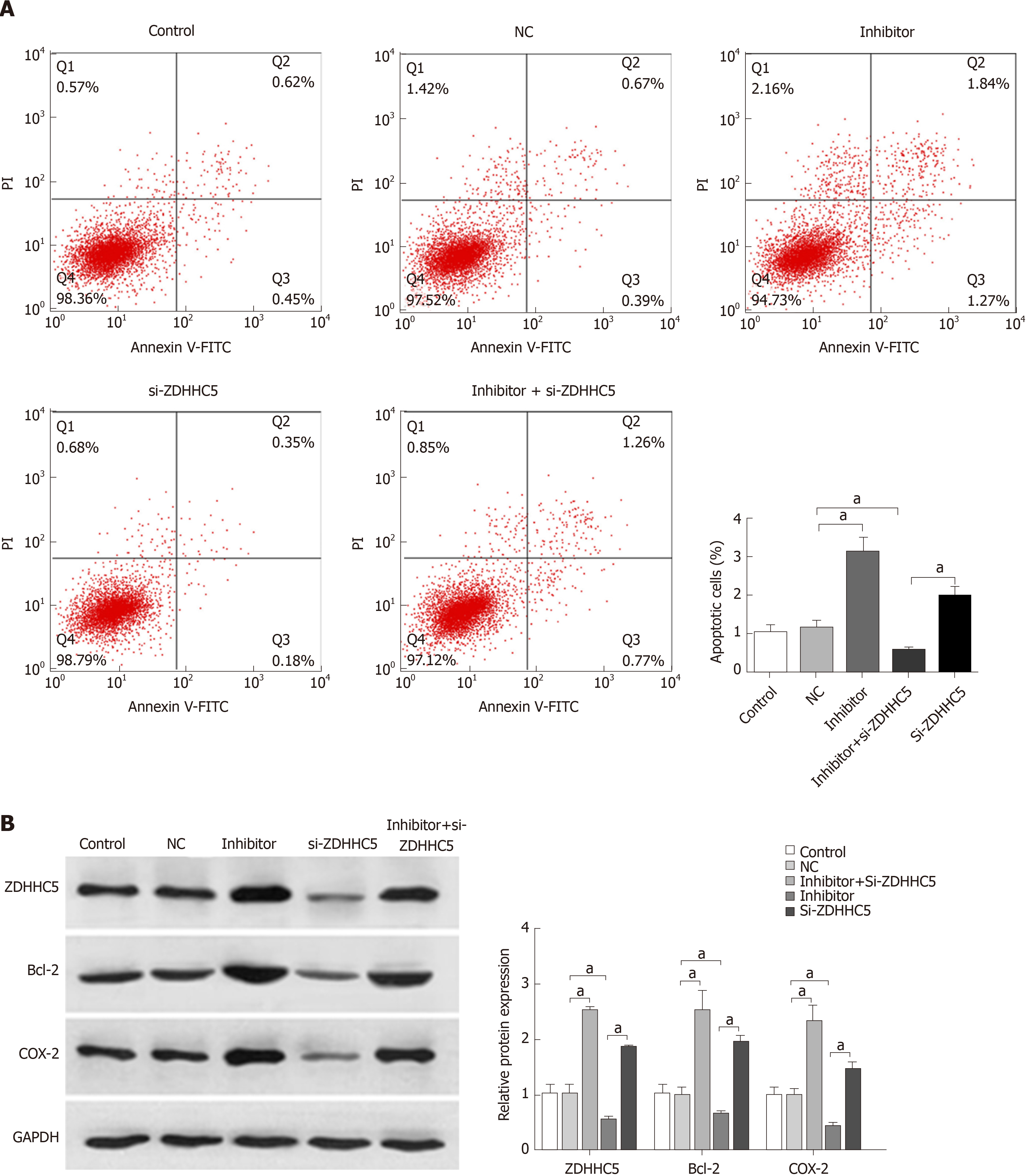
To further investigate the effects of miR-96-5p on GAC, the miR-96-5p inhibitor was used to inhibit miR-96-5p in MGC-803 cells. Flow cytometry assay showed that the number of apoptotic cells increased in MGC-803 cells transfected with the miR-96-5p inhibitor, while inhibition of ZDHHC5 decreased cell apoptosis compared with cells with NC. Notably, co-transfection of the miR-96-5p inhibitor and si-ZDHHC5 partly reversed the effect of inhibiting ZDHHC5 on cell apoptosis (P < 0.01, Figure 5A). In addition, western blotting revealed that compared with MGC-803 cells without treatment, miR-96-5p inhibition promoted the expression of ZDHHC5, Bcl-2, and COX-2 (apoptosis proteins) in MGC-803 cells (P < 0.01, Figure 5B). However, inhibiting ZDHHC5 decreased the expression of ZDHHC5, Bcl-2, and COX-2. The addition of the miR-96-5p inhibitor increased the expression of these three proteins (P < 0.01, Figure 5B).
In the present study, a total of 299 DEMs and 35 DEMs related to GAC prognosis were screened based on the miRNA expression profile data from TCGA. Then, six miRNAs were selected for identification in GAC tumor samples and adjacent normal samples. The results were consistent with bioinformatics analysis. Furthermore, miR-96-5p was considered as an important biomarker and investigated in the in vitro experiments. Our results revealed that ZDHHC5 was a direct target gene of miR-96-5p, and miR-96-5p inhibition increased the expression of Bcl-2 and COX-2.
Six miRNAs were identified in this study, and the results showed that the levels of miR-96-5p, miR-222-5p, and miR-652-5p were overexpressed, while miR-125-5p, miR-145-3p, and miR-379-3p levels were downregulated in GAC sample. Several studies have demonstrated that miR-96-5p is overexpressed in various cancers, including colorectal cancer[13], pancreatic carcinoma[14], prostate cancer[15], hepatocellular carcinoma[16], and breast cancer[17], and it is an oncogene by promoting cell proliferation. miR-652-5p was reported to be associated with non-small cell lung cancer[18], esophageal adenocarcinoma[19], and breast cancer[20], while the mechanism of miR-652-5p was unknown. Current studies of miR-222-5p are focused on the role of angiogenesis in endothelium[21,22], and few studies investigated the effect of miR-222-5p in cancers. miR-125-5p was identified as a tumor suppressor in glioblastoma[23], cervical cancer[24], and renal cell carcinoma[25], and it was involved in proliferation, migration, and apoptosis. Many have investigated the role of miR-145-3p in cancers, such as bladder cancer[26], lung squamous cell carcinoma[27], gallbladder cancer[28], and head and neck squamous cell carcinoma[29]. It is also considered a tumor suppressor. Few studies have investigated the role of miRNA-379-3p; only one recent study reported that miRNA-379-5p exerted an antitumor effect by regulating tumor invasion and metastasis in hepatocellular carcinoma[30]. Unfortunately, the effects of these miRNAs on GAC have not been reported. Therefore, it is essential to further reveal the mechanism and prognostic significance of these miRNAs in GAC.
Due to the highest association of miR-96-5p with GAC prognosis, the effects of miR-96-5p on MGC-803 cells were investigated in this study. Notably, a previous study has shown that miR-96-5p exerts an inhibiting role in cell proliferation and migration by downregulation of FoxQ1 in gastric cancer cells[31]. Contradictorily, a recent study has demonstrated that miR-96-5p exerts a promoting effect on cell progression by directly targeting FOXO3 in gastric cancer. Consistent with this recent study[32], this study found that the miR-96-5p inhibitor induced cell apoptosis in MGC-803 cells.
It is generally acknowledged that miRNAs develop biological functions by impeding translation of target mRNAs. In agreement with the bioinformatics prediction, our study revealed that ZDHHC5 was identified as a target gene of miR-96-5p. ZDHHC5, encoding zinc finger DHHC-type containing 5, is one member of the family of ZDHHC proteins and was identified as a putative palmitoyl S-acyltransferases[33]. It has been suggested that S-palmitoylation is closely associated with cancer development, and ZDHHC enzymes are the key enzymes responsible for palmitoylation[34].
Individual ZDHHC enzymes exert different effects on various cancers, either as tumor suppressors or oncoproteins[34]. A previous study documented that high expression of ZDHHC5 is associated with a poor prognosis in glioma[35]. In addition, the report of Tian et al[36] has suggested that DHHC5 knockdown can dramatically inhibit cell proliferation and invasion in non–small cell lung cancer. The present study revealed that miR-96-5p inhibition increased the number of apoptotic cells as well as promoted the expression of ZDHHC5, Bcl-2, and COX-2 in MGC-803 cells, while inhibiting ZDHHC5 decreased cell apoptosis. Co-transfection of the miR-96-5p inhibitor and si-ZDHHC5 partly reversed the effect of ZDHHC5 inhibition on cell apoptosis. These results indicated that miR-96-5p inhibition induced cell apoptosis by upregulating ZDHHC5 expression. This result was inconsistent with previous studies, which may be due to ZDHHC5 having different functions in different cancer types[37].
Gastric adenocarcinoma (GAC) is one of the leading causes of cancer-related death. However, delayed diagnosis is found in most patients with proximal or distal metastasis due to the nontypical symptoms of early GAC, which results in poor treatment and prognosis. Therefore, it is important to further reveal novel diagnostic and therapeutic methods as well as the underlying molecular mechanism of GAC.
Plenty of evidence indicates that the poor prognosis of GAC is significantly related to many molecular biomarkers, such as microRNA (miRNA). Previous studies have demonstrated that miRNA dysregulation significantly influences the prognosis of gastric cancer patients (e.g., miRNA-203, miR-21, and miR-25). Thus, it is essential to search for novel miRNAs related to GAC prognosis, which may contribute to the development of GAC diagnosis.
This study aimed to search for new miRNA therapeutic targets for GAC and investigate the mechanism of differentially expressed miRNA (DEM) in vitro, which might provide some useful insights in improving the prognosis of GAC patients.
First, the miRNA expression profile data of GAC based on The Cancer Genome Atlas were obtained and used to screen DEMs and DEMs related to GAC prognosis by bioinformatics methods. Then, the expression of DEMs related to GAC prognosis was identified in GAC tumor samples and adjacent normal samples by qRT-PCR. ZDHHC5, a target gene of miR-96-5p, was predicted and confirmed by the luciferase reporter assay. Furthermore, MGC-803 cells were transfected with inhibitor NC, miR-96-5p inhibitor, si-ZDHHC5, or miR-96-5p inhibitor + si-ZDHHC5. Cell apoptosis was detected by flow cytometry. The expression of ZDHHC5, Bcl-2, and COX-2 was detected using western blotting.
A total of 299 DEMs and 35 DEMs related to GAC prognosis were screened based on the miRNA expression profile data from The Cancer Genome Atlas. Six miRNAs, including miR-96-5p, miR-222-5p, miR-652-5p, miR-125-5p, miR-145-3p, and miR-379-3p, were selected for identification in GAC tumor samples and adjacent normal samples. The results were consistent with bioinformatics analysis. Furthermore, miR-96-5p was considered as an important biomarker and investigated in in vitro experiments. Our results revealed that ZDHHC5 was a direct target gene of miR-96-5p, and miR-96-5p inhibition induced cell apoptosis and increased the expression of Bcl-2 and COX-2.
In conclusion, this work identified six miRNAs related to GAC prognosis, including miR-96-5p, miR-125-5p, miR-145-3p, miR-222-5p, miR-379-3p, and miR-652-5p. Furthermore, downregulated miR-96-5p markedly induced cell apoptosis through targeting ZDHHC5.
Current findings provide a potential molecular mechanism of miR-96-5p in GAC. However, further studies are needed to investigate the mechanism and prognostic significance of these miRNAs in GAC.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report classification
Grade A (Excellent): A
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Amiri M, Ulaşoğlu C S-Editor: Wang J L-Editor: Filipodia E-Editor: Ma YJ
| 1. | Ajani JA, Lee J, Sano T, Janjigian YY, Fan D, Song S. Gastric adenocarcinoma. Nat Rev Dis Primers. 2017;3:17036. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 292] [Cited by in RCA: 412] [Article Influence: 51.5] [Reference Citation Analysis (0)] |
| 2. | Torre LA, Siegel RL, Ward EM, Jemal A. Global Cancer Incidence and Mortality Rates and Trends--An Update. Cancer Epidemiol Biomarkers Prev. 2016;25:16-27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2004] [Cited by in RCA: 2486] [Article Influence: 248.6] [Reference Citation Analysis (0)] |
| 3. | Sitarz R, Skierucha M, Mielko J, Offerhaus GJA, Maciejewski R, Polkowski WP. Gastric cancer: epidemiology, prevention, classification, and treatment. Cancer Manag Res. 2018;10:239-248. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 743] [Cited by in RCA: 725] [Article Influence: 103.6] [Reference Citation Analysis (0)] |
| 4. | Kemi N, Eskuri M, Ikäläinen J, Karttunen TJ, Kauppila JH. Tumor Budding and Prognosis in Gastric Adenocarcinoma. Am J Surg Pathol. 2019;43:229-234. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 65] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 5. | Tan P, Yeoh KG. Genetics and Molecular Pathogenesis of Gastric Adenocarcinoma. Gastroenterology. 2015;149:1153-1162.e3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 269] [Cited by in RCA: 363] [Article Influence: 36.3] [Reference Citation Analysis (0)] |
| 6. | Tsai MM, Wang CS, Tsai CY, Huang HW, Chi HC, Lin YH, Lu PH, Lin KH. Potential Diagnostic, Prognostic and Therapeutic Targets of MicroRNAs in Human Gastric Cancer. Int J Mol Sci. 2016;17:pii: E945. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 101] [Cited by in RCA: 104] [Article Influence: 11.6] [Reference Citation Analysis (1)] |
| 7. | Jiang C, Chen X, Alattar M, Wei J, Liu H. MicroRNAs in tumorigenesis, metastasis, diagnosis and prognosis of gastric cancer. Cancer Gene Ther. 2015;22:291-301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 95] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 8. | Acunzo M, Romano G, Wernicke D, Croce CM. MicroRNA and cancer--a brief overview. Adv Biol Regul. 2015;57:1-9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 495] [Cited by in RCA: 495] [Article Influence: 49.5] [Reference Citation Analysis (0)] |
| 9. | Saliminejad K, Khorram Khorshid HR, Soleymani Fard S, Ghaffari SH. An overview of microRNAs: Biology, functions, therapeutics, and analysis methods. J Cell Physiol. 2019;234:5451-5465. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1431] [Cited by in RCA: 1325] [Article Influence: 220.8] [Reference Citation Analysis (0)] |
| 10. | Imaoka H, Toiyama Y, Okigami M, Yasuda H, Saigusa S, Ohi M, Tanaka K, Inoue Y, Mohri Y, Kusunoki M. Circulating microRNA-203 predicts metastases, early recurrence, and poor prognosis in human gastric cancer. Gastric Cancer. 2016;19:744-753. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 69] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 11. | Simonian M, Mosallayi M, Mirzaei H. Circulating miR-21 as novel biomarker in gastric cancer: Diagnostic and prognostic biomarker. J Cancer Res Ther. 2018;14:475. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 35] [Reference Citation Analysis (0)] |
| 12. | Li BS, Zuo QF, Zhao YL, Xiao B, Zhuang Y, Mao XH, Wu C, Yang SM, Zeng H, Zou QM, Guo G. MicroRNA-25 promotes gastric cancer migration, invasion and proliferation by directly targeting transducer of ERBB2, 1 and correlates with poor survival. Oncogene. 2015;34:2556-2565. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 99] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 13. | Ress AL, Stiegelbauer V, Winter E, Schwarzenbacher D, Kiesslich T, Lax S, Jahn S, Deutsch A, Bauernhofer T, Ling H, Samonigg H, Gerger A, Hoefler G, Pichler M. MiR-96-5p influences cellular growth and is associated with poor survival in colorectal cancer patients. Mol Carcinog. 2015;54:1442-1450. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 65] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 14. | Li C, Du X, Tai S, Zhong X, Wang Z, Hu Z, Zhang L, Kang P, Ji D, Jiang X, Zhou Q, Wan M, Jiang G, Cui Y. GPC1 regulated by miR-96-5p, rather than miR-182-5p, in inhibition of pancreatic carcinoma cell proliferation. Int J Mol Sci. 2014;15:6314-6327. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 49] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 15. | Yu JJ, Wu YX, Zhao FJ, Xia SJ. miR-96 promotes cell proliferation and clonogenicity by down-regulating of FOXO1 in prostate cancer cells. Med Oncol. 2014;31:910. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 36] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 16. | Wang TH, Yeh CT, Ho JY, Ng KF, Chen TC. OncomiR miR-96 and miR-182 promote cell proliferation and invasion through targeting ephrinA5 in hepatocellular carcinoma. Mol Carcinog. 2016;55:366-375. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 55] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 17. | Shi Y, Zhao Y, Shao N, Ye R, Lin Y, Zhang N, Li W, Zhang Y, Wang S. Overexpression of microRNA-96-5p inhibits autophagy and apoptosis and enhances the proliferation, migration and invasiveness of human breast cancer cells. Oncol Lett. 2017;13:4402-4412. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 42] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 18. | Wang B, Lv F, Zhao L, Yang K, Gao YS, Du MJ, Zhang YJ. MicroRNA-652 inhibits proliferation and induces apoptosis of non-small cell lung cancer A549 cells. Int J Clin Exp Pathol. 2017;10:6719-6726. |
| 19. | Matsui D, Zaidi AH, Martin SA, Omstead AN, Kosovec JE, Huleihel L, Saldin LT, DiCarlo C, Silverman JF, Hoppo T, Finley GG, Badylak SF, Kelly RJ, Jobe BA. Primary tumor microRNA signature predicts recurrence and survival in patients with locally advanced esophageal adenocarcinoma. Oncotarget. 2016;7:81281-81291. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 25] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 20. | Lagendijk M, Sadaatmand S, Koppert LB, Tilanus-Linthorst MMA, de Weerd V, Ramírez-Moreno R, Smid M, Sieuwerts AM, Martens JWM. MicroRNA expression in pre-treatment plasma of patients with benign breast diseases and breast cancer. Oncotarget. 2018;9:24335-24346. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 11] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 21. | Chistiakov DA, Sobenin IA, Orekhov AN, Bobryshev YV. Human miR-221/222 in Physiological and Atherosclerotic Vascular Remodeling. Biomed Res Int. 2015;2015:354517. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 127] [Cited by in RCA: 141] [Article Influence: 14.1] [Reference Citation Analysis (0)] |
| 22. | Celic T, Metzinger-Le Meuth V, Six I, Massy ZA, Metzinger L. The mir-221/222 Cluster is a Key Player in Vascular Biology via the Fine-Tuning of Endothelial Cell Physiology. Curr Vasc Pharmacol. 2017;15:40-46. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 39] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 23. | Yuan J, Xiao G, Peng G, Liu D, Wang Z, Liao Y, Liu Q, Wu M, Yuan X. MiRNA-125a-5p inhibits glioblastoma cell proliferation and promotes cell differentiation by targeting TAZ. Biochem Biophys Res Commun. 2015;457:171-176. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 67] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 24. | Natalia MA, Alejandro GT, Virginia TJ, Alvarez-Salas LM. MARK1 is a Novel Target for miR-125a-5p: Implications for Cell Migration in Cervical Tumor Cells. Microrna. 2018;7:54-61. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 25] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 25. | Chen D, Li Y, Su Z, Yu Z, Yu W, Li Y, Gui Y, Yang S, Lai Y. Identification of miR‑125a‑5p as a tumor suppressor of renal cell carcinoma, regulating cellular proliferation, migration and apoptosis. Mol Med Rep. 2015;11:1278-1283. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 20] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 26. | Matsushita R, Yoshino H, Enokida H, Goto Y, Miyamoto K, Yonemori M, Inoguchi S, Nakagawa M, Seki N. Regulation of UHRF1 by dual-strand tumor-suppressor microRNA-145 (miR-145-5p and miR-145-3p): Inhibition of bladder cancer cell aggressiveness. Oncotarget. 2016;7:28460-28487. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 83] [Cited by in RCA: 96] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 27. | Mataki H, Seki N, Mizuno K, Nohata N, Kamikawaji K, Kumamoto T, Koshizuka K, Goto Y, Inoue H. Dual-strand tumor-suppressor microRNA-145 (miR-145-5p and miR-145-3p) coordinately targeted MTDH in lung squamous cell carcinoma. Oncotarget. 2016;7:72084-72098. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 67] [Cited by in RCA: 69] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 28. | Letelier P, García P, Leal P, Álvarez H, Ili C, López J, Castillo J, Brebi P, Roa JC. miR-1 and miR-145 act as tumor suppressor microRNAs in gallbladder cancer. Int J Clin Exp Pathol. 2014;7:1849-1867. [PubMed] |
| 29. | Yamada Y, Koshizuka K, Hanazawa T, Kikkawa N, Okato A, Idichi T, Arai T, Sugawara S, Katada K, Okamoto Y, Seki N. Passenger strand of miR-145-3p acts as a tumor-suppressor by targeting MYO1B in head and neck squamous cell carcinoma. Int J Oncol. 2018;52:166-178. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 43] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 30. | Chen JS, Li HS, Huang JQ, Dong SH, Huang ZJ, Yi W, Zhan GF, Feng JT, Sun JC, Huang XH. MicroRNA-379-5p inhibits tumor invasion and metastasis by targeting FAK/AKT signaling in hepatocellular carcinoma. Cancer Lett. 2016;375:73-83. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 85] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 31. | Yang XY, Li N, Deng WY, Ma YJ, Han XL, Zhang ZY, Xie JL, Luo SX. [miRNA-96-5p inhibits the proliferation and migration of gastric cancer cells by targeting FoxQ1]. Zhonghua Zhong Liu Za Zhi. 2019;41:193-199. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 32. | He X, Zou K. MiRNA-96-5p contributed to the proliferation of gastric cancer cells by targeting FOXO3. J Biochem. 2019;pii:mvz080. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 40] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 33. | Linder ME, Deschenes RJ. Palmitoylation: policing protein stability and traffic. Nat Rev Mol Cell Biol. 2007;8:74-84. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 906] [Cited by in RCA: 889] [Article Influence: 49.4] [Reference Citation Analysis (0)] |
| 34. | Yeste-Velasco M, Linder ME, Lu YJ. Protein S-palmitoylation and cancer. Biochim Biophys Acta. 2015;1856:107-120. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 62] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 35. | Chen X, Ma H, Wang Z, Zhang S, Yang H, Fang Z. EZH2 Palmitoylation Mediated by ZDHHC5 in p53-Mutant Glioma Drives Malignant Development and Progression. Cancer Res. 2017;77:4998-5010. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 90] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 36. | Tian H, Lu JY, Shao C, Huffman KE, Carstens RM, Larsen JE, Girard L, Liu H, Rodriguez-Canales J, Frenkel EP, Wistuba II, Minna JD, Hofmann SL. Systematic siRNA Screen Unmasks NSCLC Growth Dependence by Palmitoyltransferase DHHC5. Mol Cancer Res. 2015;13:784-794. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 37] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 37. | Ko PJ, Dixon SJ. Protein palmitoylation and cancer. EMBO Rep. 2018;19. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 262] [Article Influence: 37.4] [Reference Citation Analysis (0)] |