Published online Sep 28, 2019. doi: 10.3748/wjg.v25.i36.5530
Peer-review started: April 30, 2019
First decision: May 30, 2019
Revised: July 16, 2019
Accepted: August 19, 2019
Article in press: Auguet 19, 2019
Published online: September 28, 2019
Processing time: 151 Days and 18.1 Hours
DNA mutational analysis of pancreatic cystic fluid (CF) is a useful adjunct to the evaluation of pancreatic cysts. KRAS/GNAS or RAF/PTPRD/CTNNB1/RNF43 mutations are highly specific to precancerous or advanced neoplasia. Several studies recently demonstrated the ability of next-generation sequencing (NGS) analysis to detect DNA mutations in pancreatic CF, but few studies have performed a systematic comparative analysis between pancreatic CF and neoplastic surgical tissue (NT). The value of CF-NGS analysis indicators for determining surgical resection necessitates evaluation.
To confirm whether CF genomic profiles are a reliable malignancy predictor by comparing NGS mutational analyses of CF and NT.
Patients requiring surgery for high-risk pancreatic cysts were included in a multicenter prospective pilot study. DNA from CF (collected by endoscopic ultrasound-guided fine needle aspiration (known as EUS-FNA)) and NT (collected by surgery) were analyzed by NGS. The primary objective was to compare the mutation profiles of paired DNA samples. The secondary objective was to correlate the presence of specific mutations (KRAS/GNAS, RAF/ PTPRD/CTNNB1/RNF43/POLD1/TP53) with a final cancer diagnosis. Sensitivity and specificity were also evaluated.
Between December 2016 and October 2017, 20 patients were included in this pilot study. Surgery was delayed for 3 patients. Concordant CF-NT genotypes were found in 15/17 paired DNA, with a higher proportion of mutated alleles in CF than in NT. NGS was possible for all pancreatic CF collected by EUS-FNA. In 2 cases, the presence of a KRAS/GNAS mutation was discordant between CF and NT. No mutations were found in 3 patients with NT or pancreatic cysts with high-grade dysplasia. The sensitivity and specificity of KRAS/GNAS mutations in CF to predict an appropriate indication for surgical resection were 0.78 and 0.62, respectively. The sensitivity and specificity of RAF/PTPRD/CTNNB1 /RNF43/POLD1/TP53 mutations in CF were 0.55 and 1.0, respectively.
Mutational analyses of CF and NT were highly concordant, confirming the value of NGS analysis of CF in the preoperative malignancy assessment. However, these results need to be confirmed on a larger scale.
Core tip: This pilot study (20 patients) was a comparative next-generation sequencing mutational analysis between pancreatic cyst fluid (CF) and neoplastic surgical tissue (NT) to confirm whether the CF genomic profile was a reliable malignancy predictor. Concordant genotypes were found in 15 of 17 paired DNA samples. The sensitivity and specificity of CF mutations to predict an appropriate indication for surgical resection were 0.78 and 0.62 for the KRAS/GNAS mutations and 0.55 and 1.0 for the RAF/PTPRD/CTNNB1/RNF43/POLD1/TP53 mutations, respectively. Mutational analyses of CF and NT were highly concordant, confirming the interest of CF next-generation sequencing analysis in the preoperative malignancy assessment.
- Citation: Laquière A, Lagarde A, Napoléon B, Bourdariat R, Atkinson A, Donatelli G, Pol B, Lecomte L, Curel L, Urena-Campos R, Helbert T, Valantin V, Mithieux F, Buono JP, Grandval P, Olschwang S. Genomic profile concordance between pancreatic cyst fluid and neoplastic tissue. World J Gastroenterol 2019; 25(36): 5530-5542
- URL: https://www.wjgnet.com/1007-9327/full/v25/i36/5530.htm
- DOI: https://dx.doi.org/10.3748/wjg.v25.i36.5530
Cystic neoplasms of the pancreas are frequent in the general population, with an estimated prevalence ranging from 5%-15% in those over the age of 70 years[1,2]. Pancreatic cysts can be divided into mucinous and non-mucinous. Their distinction is important because mucinous cysts, which comprise mucinous cystic neoplasms (MCNs) and intraductal papillary mucinous neoplasms (IPMNs), are considered premalignant. Indeed, the risk of pancreatic adenocarcinoma developing from main-duct IPMN is high, at 40%-90% within 5 years of diagnosis[3,4]. For branch-duct IPMN, the risk is moderate, ranging between 6%-46% within 5 years[5-8]. This risk considerably decreases for mucinous cystadenoma (an MCN) to under 15%[8], and is quite rare for serous cystadenoma (SCA)[2]. The clinical management of patients with pancreatic cysts is unfortunately imperfect, and distinguishing between different cystic tumors and their risks of malignant evolution can be challenging.
A few guidelines based on clinical features and cystic tumor morphology were published to help set intervals of follow-up, and to define criteria for surgical resection[9-13]. However, in a meta-analysis, the pooled sensitivity and specificity for malignancy in Sendai-positive lesions were 56% and 74%, respectively, and under the Fukuoka criteria, they were 83% and 53%[14]. Worrisome features in those guidelines are not sufficient to correctly select patients for surgery[10,12]. Currently, 75% of resected IPMNs harbor only low- or intermediate-grade dysplasia, which could have been safely observed[15]. Similarly, in a retrospective multicentric study, 96.5% of patients presenting an IPMN with worrisome features who did not undergo surgery were still alive 5 years later without developing pancreatic cancer[16].
Other tests that can help in cystic lesion diagnosis and management include cytology and biochemical tests of cystic fluid (CF), such as carcinoembryonic antigen (CEA). However, those tests are also limited; cytology had a sensitivity for malignancy of 42% in a meta-analysis of 12 articles[17], and there are varying cut-off values for CEA. This is why molecular analysis was developed in the last decade, with new techniques of advanced sequencing becoming available to help in pancreatic cyst differential diagnosis. DNA mutational analysis of the different types of pancreatic cysts involves several specific alterations, with each cyst type having different mutational profiles. IPMN and mucinous cysts harbor KRAS mutations at diagnosis, in addition to GNAS mutations in IPMN, while PTEN, CDKN2A and TP53 mutations are mostly found in cancerous lesions[18]. In contrast, SCA contains a mutation in the Von Hippel Lindau gene, and a mutation in the CTNNB1 gene is described in solid pseudopapillary neoplasms.
Several studies recently demonstrated the ability of next-generation sequencing (NGS) analysis to detect DNA mutations in pancreatic CF, which contains “circulating” DNA from surrounding epithelial cells. Nevertheless, few studies have yet to perform a systematic comparative analysis of pancreatic CF and neoplastic surgical tissue (NT). Therefore, molecular analysis of DNA mutations in CF would potentially be a very good prognostic marker[19,20]. The current 50% diagnostic accuracy to indicate a surgical treatment could reach 90% with molecular analysis of CF-DNA mutations[21].
Although the molecular results for CF-DNA analysis seem very promising, we have little data on the feasibility of this analysis, especially when the fluid is collected after puncturing the cyst under endoscopic ultrasound (EUS) control. In addition, these few preliminary results must be confirmed with other studies. The value of NGS analysis in CF as an indicator for therapeutic decision needs to be evaluated. This pilot prospective study was conducted to evaluate the feasibility, sensitivity and specificity of NGS analysis of CF in comparison with the "gold-standard" anatomopathological analysis of the surgical specimen. The main objective was to evaluate and compare the genomic profiles of pancreatic CF and pancreatic tumors to confirm whether the genomic profile of pancreatic CF was a reliable predictor of malignancy, and could thus help in clinical management.
This prospective multicenter study promoted by Saint Joseph Hospital was approved by the French Ethical Committee (ID RCB: 2016-A01399-42/ref 16-107 available upon request) and registered in the ClinicalTrial.gov database (NCT03305146). Six centers, all expert centers in digestive oncology and surgery, participated in the study. The primary objective of this pilot prospective study was to determine the mutational concordance in the molecular biology analysis of paired DNA samples from CF and pancreatic tumor NT. The secondary objective was to analyze specific mutations (KRAS, GNAS, RAF, PTPRD, CTNNB1, RNF43, POLD1, and TP53) to correlate their presence with a final cancer diagnosis. The sensitivity and specificity of the DNA mutational analysis in CF was also evaluated for pancreatic cysts requiring surgery.
Patients diagnosed with pancreatic cystic lesions with worrisome features or high-risk stigmata and scheduled for surgery in the participating centers were enrolled in the study. Worrisome features were defined as a cyst > 3 cm, thickened/enhancing cyst walls, main pancreatic duct size 5-9 mm, non-enhancing mural nodules, and an abrupt change in the caliber of the pancreatic duct with distal pancreatic atrophy. High-risk stigmata were defined by obstructive jaundice in a patient with a cystic lesion on the head of the pancreas, and an enhancing solid component within the cyst or main pancreatic duct ≥ 10 mm in size. Malignant cytopathology was defined as being either suspicious for adenocarcinoma or positive for adenocarcinoma. Pathology slides were reviewed for each surgical specimen, and diagnoses for all pancreatic cysts were rendered based on standard histo-morphological criteria. Pathological staging was performed as outlined by the American Joint Committee in the Cancer Staging Manual (8th edition). Inclusion and exclusion criteria are described in Table 1. Relevant collected data are presented in Table 2.
| Inclusion criteria | Exclusion criteria |
| Patients over the age of 18 yr, male or female | Contraindications to perform digestive endoscopy |
| Cystic tumor of the pancreas > 15 mm to benefit from a puncture under EUS control to determine the etiological and gravity diagnosis. | Hemorrhagic disease, hemostasis or coagulation disorder (PR < 60%, APTT > 40 s or platelets < 60000/mm3). |
| No transient shutdown of AVK, AOD or AAP | |
| Complete clinical examination | |
| Patients able to follow and understand the study and answer the questionnaire | |
| Pregnant or lactating women (a urine pregnancy test was carried out in women of childbearing age) | |
| Informed and written consent | Vulnerable persons: deprived of liberty or guardianship |
| Affiliation to a social security |
| Final diagn-osis | Pat. code | Gender | Age | Status | Sample | Proportion of mutated allele | Patholo-gical feature-s | |||||||
| KRAS | GNAS | TP53 | RAF1 | PTPRD | CTNNB1 | RNF43 | POLD1 | |||||||
| Cancer | 1 | F | 73 | No mutatio-n | CF | - | - | - | - | - | - | - | - | MCN with invasive ADK |
| NT | - | - | - | - | - | - | - | - | ||||||
| Cancer | 9 | M | 72 | Noncon-cordant mutatio-ns | CF | 0 | - | - | 0 | 0 | - | - | - | IPMN with invasive adenoca-rcinoma pT2N1 |
| NT | 100 | - | - | 26 | 100 | - | - | - | ||||||
| Cancer | 10 | F | 60 | Conc-ordant mutatio-ns | CF | 5 | - | - | - | - | - | - | 50 | IPMN with invasive adenoca-rcinoma pT3N1 |
| NT | 0 | - | - | - | - | - | - | 24 | ||||||
| Cancer | 13 | M | 66 | Concor-dant mutatio-ns | CF | 3 | - | 1 | - | - | - | - | - | IPMN with invasive adenoca-rcinoma pT3N1 |
| NT | 35 | - | 22 | - | - | - | - | - | ||||||
| Cancer | 14 | M | 63 | Concor-dant mutatio-ns | CF | - | 37 | - | - | - | - | - | - | MCN with invasive ADK |
| NT | - | 19 | - | - | - | - | - | - | ||||||
| Cancer | 15 | M | 83 | Concor-dant mutatio-ns | CF | 32 | 50 | - | - | - | - | - | - | IPMN with focal adenoca-rcinoma pTisN0 |
| NT | 9 | 19 | - | - | - | - | - | - | ||||||
| Cancer | 17 | F | 71 | Concor-dant mutatio-ns | CF | 25 | 48 | - | - | - | 56 | - | - | IPMN with focal adenoca-rcinoma pTisN0 |
| NT | 21 | 42 | - | - | - | 37 | - | - | ||||||
| Cancer | 18 | M | 82 | Concor-dant mutatio-ns | CF | 41 | - | - | - | - | - | 39 | - | IPMN with focal adenoca-rcinoma pTisN0 |
| NT | 32 | - | - | - | - | - | 35 | - | ||||||
| Cancer | 19 | F | 67 | Concor-dant mutatio-ns | CF | 23 | - | 31 | - | - | - | - | - | MCN with invasive ADK |
| NT | 0 | - | 18 | - | - | - | - | - | ||||||
| NO Cancer | 2 | M | 86 | No mutatio-n | CF | - | - | - | - | - | - | - | - | IPMN with low-grade dysplasi-a |
| NT | - | - | - | - | - | - | - | - | ||||||
| NO Cancer | 3 | M | 75 | No mutatio-n | CF | - | - | - | - | - | - | - | - | Serous cystaden-oma |
| NT | - | - | - | - | - | - | - | - | ||||||
| NO Cancer | 4 | F | 66 | No mutatio-n | CF | - | - | - | - | - | - | - | - | IPMN with low-grade dysplasi-a |
| NT | - | - | - | - | - | - | - | - | ||||||
| NO Cancer | 8 | F | 81 | No mutatio-n | CF | - | - | - | - | - | - | - | - | Mucinou-s cystaden-oma |
| NT | - | - | - | - | - | - | - | - | ||||||
| NO Cancer | 16 | M | 63 | No mutatio-n | CF | - | - | - | - | - | - | - | - | Cystic lymphan-gioma |
| NT | - | - | - | - | - | - | - | - | ||||||
| NO Cancer | 6 | M | 71 | Nonconc-ordant mutatio-ns | CF | 12 | 31 | - | - | - | - | - | - | IPMN with low-grade dysplasi-a |
| NT | 0 | 0 | - | - | - | - | - | - | ||||||
| NO Cancer | 5 | M | 74 | Concor-dant mutatio-ns | CF | 41 | 12 | - | - | - | - | - | - | IPMN with low-grade dysplasi-a |
| NT | 31 | 0 | - | - | - | - | - | - | ||||||
| NO Cancer | 20 | M | 51 | Concor-dant mutatio-ns | CF | - | 26 | - | - | - | - | - | - | IPMN with low-grade dysplasi-a |
| NT | - | 7 | - | - | - | - | - | - | ||||||
A sample size of 20 patients for this pilot study was determined according to various parameters: The frequency of the disease, the inclusion capacity of the centers, and a 12-mo maximum inclusion duration.
Pancreatic CF samples were collected at the time of EUS. CF samples were aspirated under EUS control using 22-gauge needles (Boston Scientific, Marlborough, MA, United States), and studied for biochemical markers in the systematic preoperative evaluation; 1.5 mL was preserved for molecular analysis in 2 mL of ATL Buffer (Qiagen, Hilden, Germany). Cytological analysis of CF was not systematically performed because of its poor diagnostic performance in the presence of poor cellularity. NT samples were withdrawn from the cystic wall on the pancreatic surgical resection by the pathologist before formalin fixation. A fragment of approximately 25 mg was placed into 2 mL of Allprotect Tissue Reagent (Qiagen). Blood samples (10 mL) were collected to eliminate germ line mutations in the case of an ambiguous sequencing profile. Samples were stored at 4 °C for no more than a month before treatment.
Sample processing, NGS sequencing, and sequence analyses were performed in the INSERM-MMG laboratory. DNA was extracted from cyst fluids and tissues using the QIAamp DNA Micro Kit (Qiagen) following the manufacturer’s recommendations, with the exception of three points (Supplement 1).
A panel of 526 genes was created using SureDesign for the HaloPlex Target Enrichment System (Agilent Technologies Inc., Santa Clara, CA, United States) adapted for NextSeq Illumina NGS (Illumina, San Diego, CA, United States) (Supplement 2/genes list). Designed probes for the total target covered more than 99% of coding regions, as well as 20 base-pair regions flanking exon-intron boundaries, corresponding to 101,803 amplicons and 1.48 megabase pairs. Library preparation for sequencing was carried out using the HaloPlex Target Enrichment Kit 0.25-2.5 Mbp (Agilent Technologies Inc.). The protocol is briefly described in Supplement 2.
Sequencing run outputs were loaded onto a server in the Bioinformatics Department. Raw data generated by the NextSeq 500 sequencer were demultiplexed and converted into FASTQ files using bcl2fastq 2.18. Read quality was assessed using FastQC 0.11.5[22].
The obtained sequences were mapped to the human reference assembly genome hg19 using the Maximum Exact Matches algorithm in Burrows–Wheeler Aligner 0.7.15[23]. On average, 98% of targeted sequences were successfully covered with a depth > 50× (Supplement 3). Alignment quality was evaluated using Qualimap 2.2.1[24].
First, the Genome Analysis Software Kit 3.7 (GATK) pipeline was applied following the germline version of GATK best-practice recommendations to perform variant calling[25]. The GATK base quality score recalibrator was applied to correct sequencing artifacts, and variants were called using HaplotypeCaller. At the time of data processing, HaplotypeCaller was the variant analysis software embedded into the GATK best-practice pipeline. It is best suited to germline variant calling, but also detects allele frequencies outside of an expected 50:50 ratio. MuTect2 beta release was used to perform somatic variant calling. MuTect2 detects a range of allele frequencies, thus making it more suitable for somatic genotyping in heterogeneous, often aneuploid, tumor tissue compared to HaplotypeCaller, which was designed more for germline variant calling. Normal samples were briefly combined to generate a panel of normal (referred to as PoN) variations. Variant calling was run by normal/tumor comparison for each patient, filtered using the dbSNP 138 database, COSMIC and the pregenerated PoN. Variant annotation, mining and manual review were performed using VarAFT (http://varaft.eu) and ALAMUT software (Interactive Biosoftware, Rouen, France). Variants of interest were then verified using the Integrative Genomics Viewer.
We considered that a pancreatic cyst postoperatively characterized and associated with cancer (in situ or invasive carcinoma) was the most appropriate pancreatic cyst for surgical excision. Both KRAS/GNAS and RAF/PTPRD/CTNNB1/RNF43 /POLD1/TP53 mutations were analyzed in the CF and NT in each cancerous pancreatic cyst. Sensitivity was defined as the probability of a positive mutation in in situ or invasive carcinoma. Specificity was defined as the probability of a negative mutation (no mutation) in a healthy person (no cancer).
This pilot study did not assess statistical significance. Numerical data are expressed as the mean ± standard deviation (SD) or as percent.
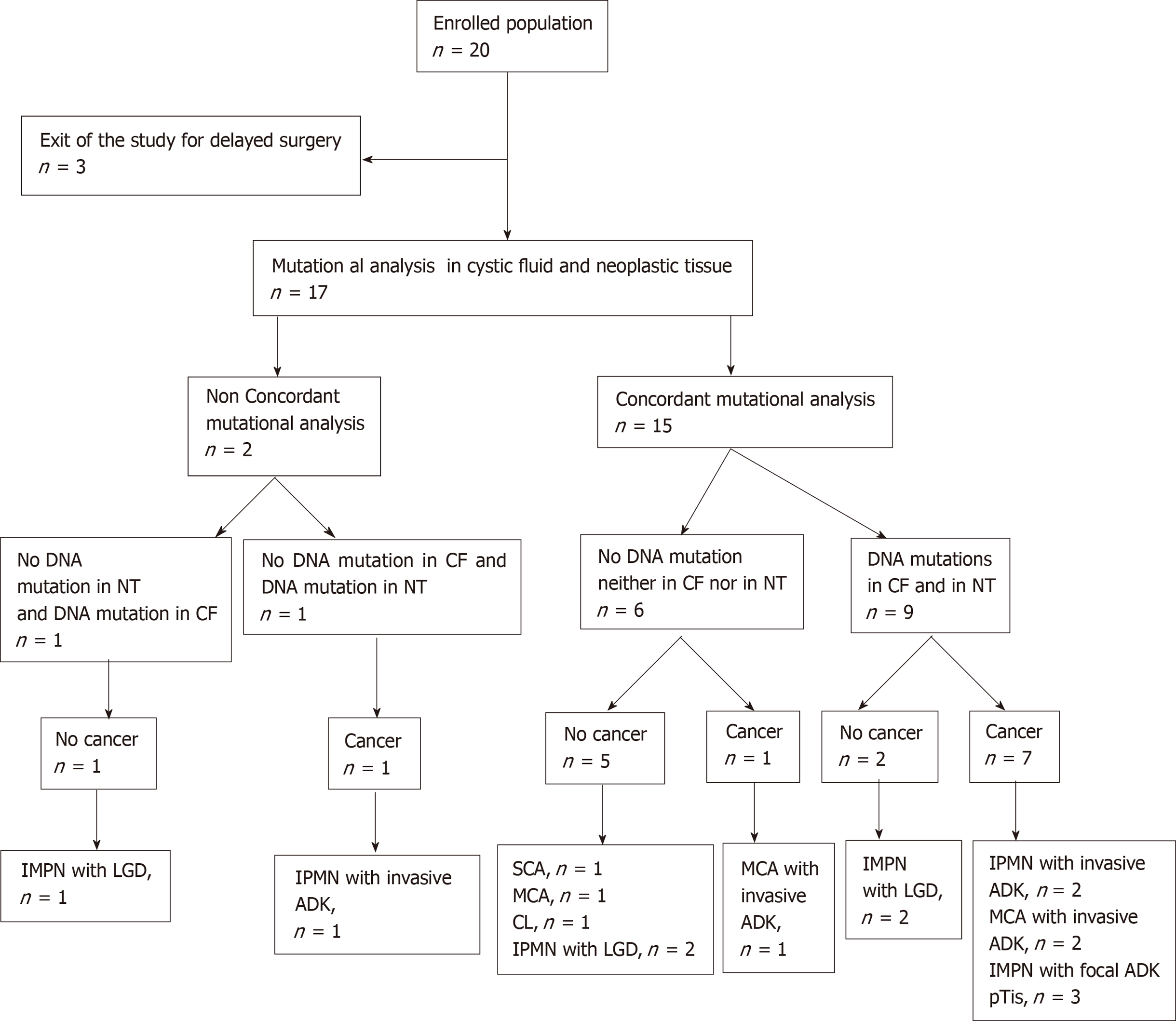
From December 2016 to October 2017, 20 patients (13 males, 7 females), aged from 51 years to 91 years (mean age: 71.5 ± 10.3 years), diagnosed with pancreatic cystic lesions with worrisome features or high-risk stigmata and scheduled for surgery, were enrolled in the study. Surgery was delayed for 3 patients because of postoperative high mortality risk. The results of DNA analysis are reported in Table 2, together with pathological features of the surgical cyst resections. Raw data corresponding to each DNA mutation are presented in Supplement 4.
NGS was carried out in 100% of the pancreatic CF samples collected by ultrasound-guided fine needle aspiration (EUS-FNA), and in 100% of the surgically resected lesion samples.
Concordant genotypes were found in 15 of 17 paired DNA samples, with various proportions of mutated alleles including a higher proportion in CF-DNA than in NT-DNA, as shown in the raw data detailed in Supplement 4. In one case (patient 9, Table 2), no mutation was found in CF-DNA. This patient had invasive cancer harboring heterozygous mutations in three different genes. In another case (patient 6, Table 2), no mutation was found in NT-DNA, while two mutations were abundantly detected in CF-DNA.
All 17 patients undergoing partial pancreatectomy had one or more worrisome features or “high-risk stigmata” according to international guidelines. For 8/17 patients histopathologically (i.e. postoperatively) diagnosed with a benign pancreatic cyst, 1 patient had a SCA, 1 patient had a cystic lymphangioma (CL), 1 patient had a benign MCN, and 5 patients had IPMN with low-grade dysplasia. In the rest of the population (9/17 patients), 3/9 patients had IPMN with invasive adenocarcinoma, 3/9 had IPMN with focal adenocarcinoma, and 3/9 patients had MCN with invasive adenocarcinoma (Table 2, Figure 1).
For 6/17 patients, no mutation was found in either CF or paired tissue. Pathological reports mentioned the absence of (high-grade) dysplastic cells in 5 of them (1 SCA, 1 MCN, 1 CL, and 2 IPMN), and a true cancer in one of them (MCN with invasive adenocarcinoma; patient 1) (Table 2). For the 11 other patients, only 8/526 of the sequenced genes exhibited mutations (KRAS/GNAS/RAF/PTPRD/CTNNB1 /RNF43/POLD1/TP53), with one to three mutations for each patient (Table 2). Three patients were confirmed to have IPMN with low-grade dysplasia, 3 patients had in situ cancer, and 5 patients had invasive carcinoma. No mutation was present in the germ line.
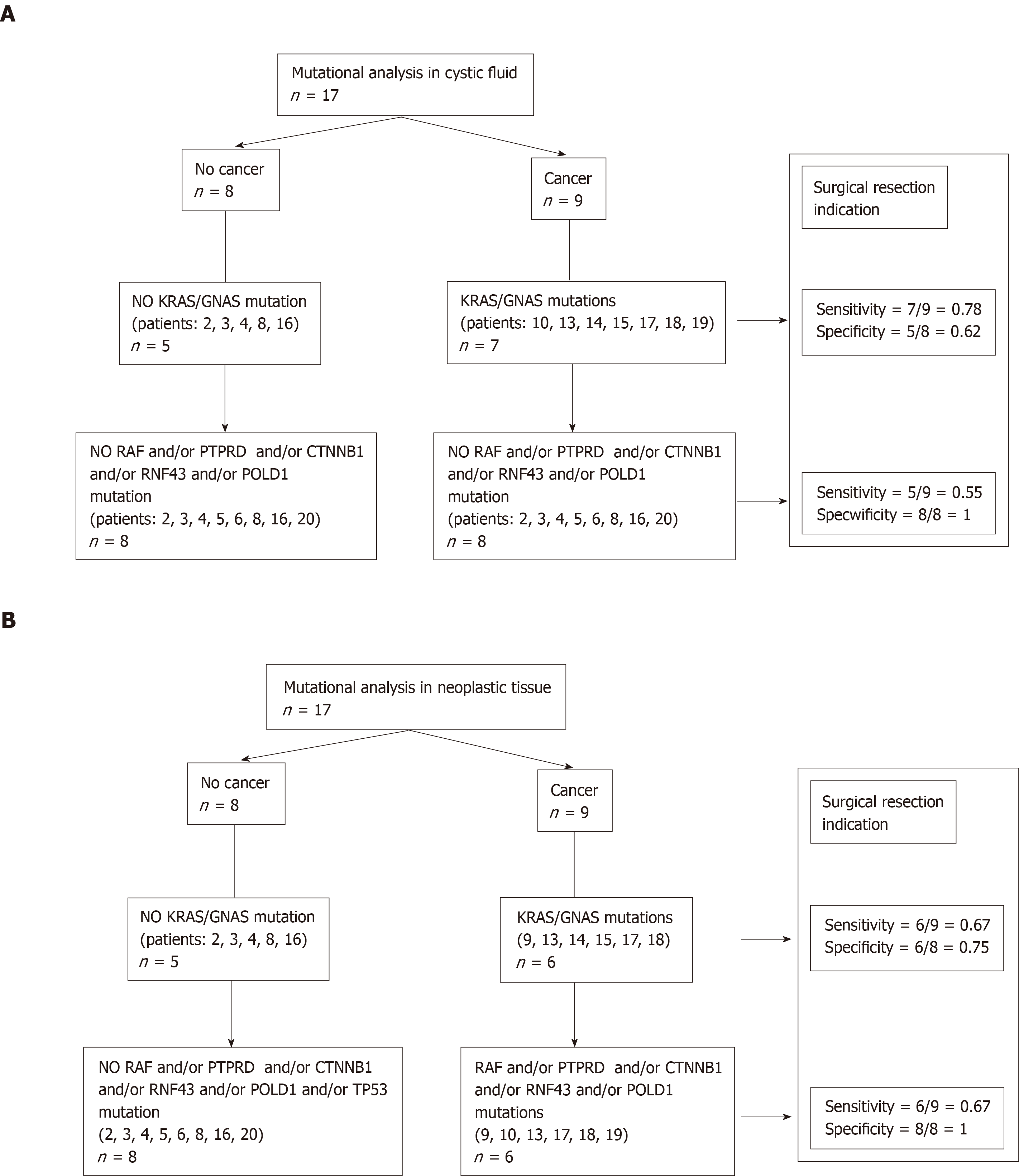
CF: Of the 9 patients who had a cancerous cyst, CF mutations were found in 7 patients (patients: 10-13-14-15-17-18-19). Regarding the 2 patients who did not have CF mutations, one had no NT mutation (patient 1), whereas the second had a NT mutation (patient 9) (see Table 3). Finally, the sensitivity and specificity of the KRAS/GNAS mutations in the CF collected by EUS-FNA to indicate surgical resection (or to predict the risk of cancer) were 0.78 (7/9) and 0.62 (5/8), respectively (Figure 2A).
| KRAS/GNAS mutations analysis in cystic fluid, n = 17 | Cancer | No cancer |
| Positive mutations (+) | ||
| Population size, n | 7 | 3 |
| Patient codes | True positive: 10, 13, 14, 15, 17, 18, 19 | False positive: 5, 6, 20 |
| Negative mutations (-) | ||
| Population size, n | 2 | 5 |
| Patient codes | False negative: 1, 9 | True negative: 2, 3, 4, 8, 16 |
| Total population, n | 9 | 8 |
NT: Of the 9 patients who had a cancerous cyst, only 6 patients had NT mutations (patients: 9-13-14-15-17-18). Of the 8 patients without cancer, 2 patients had mutations (patients: 5, 20). These 2 patients had IPMN with low-grade dysplasia. Finally, the sensitivity and specificity of the KRAS/GNAS mutations in the CF collected by surgery or EUS-FNA to predict the risk of cancer (or to indicate surgical resection) were 0.66 (6/9) and 0.75 (6/8), respectively (Figure 2B).
CF: RAF and/or PTPRD and/or CTNNB1 and/or RNF43 and/or POLD1 and/or TP53 mutations were present only in 5 patients with in situ or invasive cancer (patients: 10-13-17-18-19). The sensitivity and specificity of these mutations in the CF collected by EUS-FNA to predict the risk of cancer were 0.55 (5/9) and 1.0 (8/8), respectively. No mutation was present in the germ line (Figure 2A, Table 4).
| RAF/PTPRD/CTNNB1/RNF43/POLD1/TP53 mutations in cystic fluid, n = 17 | Cancer | No cancer |
| Positive (+) | ||
| Population size, n | 5 | 0 |
| Patient codes | True positive: 10, 13, 17, 18, 19 | False positive |
| Negative mutations (-) | ||
| Population size, n | 4 | 8 |
| Patient codes | False negative: 1, 9, 14, 15 | True negative: 2, 3, 4, 5, 6, 8, 16, 20 |
| Total population, n | 9 | 8 |
NT: RAF and/or PTPRD and/or CTNNB1 and/or RNF43 and/or POLD1 and/or TP53 mutations were present only in 6 patients with in situ or invasive cancer (patients: 9-10-13-17-18-19). The sensitivity and specificity of these mutations in the NT collected by surgery or EUS-FNA to predict the risk of cancer were 0.67 (6/9) and 1.0 (8/8), respectively (Figure 2B).
Several studies confirming the presence of KRAS, GNAS, RAF, PTPRD, CTNNB1, RNF43, or TP53 mutations in CF[19-21] did not compare CF with surgically resected NT mutations. To our knowledge, this study is the only one that compares CF with NT mutations as its main objective. The unique feature of this study lies in the fact that, for all patients, the profile of the genetic mutations in CF and NT was systematically compared.
Concordant genotypes were found in 15 of 17 paired DNA, with various proportions of mutated alleles that were generally higher in CF-DNA than in NT-DNA (Table 2). The sequencing technology HaloPlexHS enables detection of alleles present in a proportion of 1%. This technical approach, combining high sensitivity and specificity, is compatible with degraded DNA, even if there is less than 50 ng. It therefore appears to be an appropriate technique for these samples with an expected high level of degraded DNA and cell heterogeneity.
As for the 2 discordant cases, patient 9 harbored multiple cystic lesions, one of which was a suspected adenocarcinoma that was not observed during EUS exploration. We presume that CF and tissue were withdrawn from different benign cysts, because a KRAS mutation was reported in the cancerous lesions of both cases by the molecular pathologist. Therefore, in the case of multiple lesions, several fluid samples might be needed for mutational analysis. This patient was the only one without mutations detected in CF and with cancer as the final diagnosis. To avoid this specificity problem, further studies will be carried out with additional CF mutational analyses. For patient 6, the proportion of mutated alleles was not high in CF, but sufficient to exclude sample contamination. Tumor heterogeneity might explain this observation by itself, but it could also be interesting to systematically calculate the proportion of neoplastic cells. The mutated profile was not detected in the corresponding NT, emphasizing the importance of detecting CF rather than performing an NT biopsy.
For the 3 patients who presented a pancreatic cyst that required surgical resection (patients: 1, 2 and 3), no mutations were found. The possibility of false-negative results is always a concern with any sequencing technique. This study was designed to detect all known mutations in oncogenes and most mutations in tumor suppressor genes that occur in cysts based on genome-wide sequencing. Furthermore, the molecular panel consisted of commonly altered genes in pancreatic cysts that were present in invasive adenocarcinoma, but did not include CDKN2A and SMAD4. Deletions in CDKN2A and SMAD4 are associated with IPMNs harboring high-grade dysplasia and invasive adenocarcinoma. Another source of false-negative results could be tumor suppressor genes. Large deletions or insertions, as well as translocations, will not be detected upon sequencing. These facts could explain the low sensitivity and specificity of mutational analysis to predict cyst cancer in this study compared to other studies[19,20].
Our study showed that there was a good correspondence between the mutations found in the liquid part of the degenerate cyst and the tumor itself. However, when analyzing mutations in the NT of the tumor, the sensitivity and specificity of KRAS/GNAS mutations to estimate the risk of cancer were not excellent, with values of 0.67 and 0.75, respectively (Figure 2). Regarding the other RAF/PTPRD/ CTNNB1/RNF43/POLD1/TP53 mutations, which are more specific to cancerous lesions, the sensitivity was insufficient: 0.55 in CF vs 0.67 in NT. This means that additional mutations sensitive to degeneration should be studied. From a diagnostic point of view, KRAS and GNAS mutations were not found either in CF or in NT in the 2 cases of IPMN. The sensitivity and diagnostic specificity of the GNAS/KRAS mutations were inferior to the results published in the literature (the sensitivity was 96% in the study by Singhi et al[20]).
Regarding KRAS/GNAS mutations, we observed a higher sensitivity and lower specificity of CT vs NT. This could be explained by the generally higher proportions of mutated alleles in CF-DNA than in NT-DNA (Table 2). Despite quite low rates, these preliminary results are encouraging, confirming the hypothesis of CF genomic profile reliability (at least for RAF PTPRD/CTNNB1/RNF43/POLD1/TP53 mutations). Further analyses are needed to confirm these results and avoid bias. In this series, all patients had precancerous or cancerous cysts. No patient had a pseudocyst or dilatation of the main pancreatic duct in relation to chronic calcific pancreatitis, because morphological criteria are now well-defined in imaging[26]. The main limitation of this study lies in the small number of patients. As a result, the final diagnosis, as well as the resulting sensitivity and specificity calculations, may lack precision. This pilot study was a first step, and a multicenter study on a larger scale is ongoing. It was reassuring in this respect that our analysis of paired fluids obtained from EUS-guided cyst aspiration and subsequent surgery revealed very similar molecular genetic alterations. In the future, the optimal study design should incorporate examination of the CF collected by routine EUS sessions over time, then comparing the results with those obtained by surgery.
Our analysis of paired fluids obtained from EUS-guided cyst aspiration and subsequent surgery revealed very similar genetic alterations. In the future, the optimal study design should incorporate examination of the CF collected by routine EUS sessions over time, and the results should then be compared with those obtained by surgery. Although the aim of this study was to compare fluid with tissue, the most important issue was to determine whether the genomic profiles from fluid analysis could predict a malignant diagnosis and thus indicate surgical resection. Despite the fact that 15/17 patients presented the same genomic profile between CF and NT, the specificity and sensitivity observed in this study were not satisfactory because the mutations were not systematically present in all patients with cancer. In our study, 3 patients with cancer had mutations in neither CF nor NT. The predictive effect of CF mutations on malignity needs to be further analyzed with some additional mutations. This analysis has been planned in a future larger-scale study.
In conclusion, this study reinforces the value of including CF-DNA profiling before entering patients into a surgical program, in addition to using new imaging approaches. As circulating tumor cells harbor molecular alterations now used as cancer diagnostic or recurrence markers, tumor cells are present in the CF and can be collected for molecular investigations, validating this pilot approach as relevant to assess the diagnostic and prognostic criteria in a larger series.
DNA mutational analysis of pancreatic cystic fluid (CF) is a useful adjunct to the evaluation of pancreatic cysts. KRAS/GNAS or RAF/PTPRD/CTNNB1/RNF43 mutations are highly specific to precancerous or advanced neoplasia. Several studies recently demonstrated the ability of next-generation sequencing (NGS) analysis to detect DNA mutations in pancreatic CF, but few studies have yet to perform a systematic comparative analysis between pancreatic CF and neoplastic surgical tissue (NT). The value of CF-NGS analysis indicators for determining surgical resection requires evaluation.
The main subject of this study is to evaluate the diagnostic performance of some specific mutations in CF, and to establish a concordance of detected mutations in CF and NT. The key problem to be solved was to appropriately select patients for preventive pancreatic surgery. It is very important to solve this problem in the future to avoid unnecessary procedures. Future research will be targeted to assess and validate this new technique of non-cancerous cyst prognosis.
The primary objective of this pilot prospective study was to determine the mutational concordance in the molecular biology analysis of paired DNA samples from CF and pancreatic tumor NT. The secondary objective was to analyze specific mutations (KRAS, GNAS, RAF, PTPRD, CTNNB1, RNF43, POLD1, and TP53) to correlate their presence with a final cancer diagnosis. The sensitivity and specificity of the DNA mutational analysis in CF was also evaluated for pancreatic cysts requiring surgery.
Patients requiring surgery for high-risk pancreatic cysts were included in a multicenter prospective pilot study. DNA from CF (collected by Endoscopic Ultrasound-Guided Fine Needle Aspiration Biopsy (EUS-FNA)) and NT (collected by surgery) were analyzed by NGS. The primary objective was to compare the mutation profiles of paired DNA samples. The secondary objective was to correlate the presence of specific mutations (KRAS/GNAS, RAF/PTPRD/CTNNB1/RNF43/POLD1/TP53) with a final cancer diagnosis. Sensitivity and specificity were also evaluated.
Between December 2016 and October 2017, 20 patients were included in this pilot study. Surgery was delayed for three patients. Concordant CF-NT genotypes were found in 15/17 paired DNA, with a higher proportion of mutated alleles in CF than in NT. NGS was possible for all pancreatic CF collected by EUS-FNA. In two cases, the presence of a KRAS/GNAS mutation was discordant between CF and NT. In three patients with NT or pancreatic cysts with high-grade dysplasia, no mutations were found. The sensitivity and specificity of KRAS/GNAS mutations in CF to predict an appropriate indication for surgical resection were 0.78 and 0.62, respectively. The sensitivity and specificity of RAF/PTPRD/CTNNB1/RNF43/POLD1/TP53 mutations in CF were 0.55 and 1.0, respectively.
The main goal of this study was to confirm the concordance between CF and NT mutations. The prognosis of pancreatic cysts can be evaluated by analyzing CF mutations. KRAS, GNAS, RAF, PTPRD, CTNNB1, RNF43, POLD1, and TP53 mutations are good indicators of cyst malignant degeneration risks. This study offers the opportunity to accurately assess pancreatic cyst prognosis using a minimally invasive technique. The new hypothesis assessed in this study is that CF mutations are good indicators of malignant degeneration risks. The new methods proposed in this study involve DNA extraction from a body fluid other than blood. The POLD1 gene might be of interest in the evaluation of the malignant degeneration risks of pancreatic cysts. We confirmed that CF mutations are good indicators of malignant risk. In the future, all patients with pancreatic cysts could benefit from CF analysis collected by EUS-FNA.
All of the collected CF samples were able to be genetically analyzed. The mutations selected for evaluation (KRAS, GNAS, RAF, PTPRD, CTNNB1, RNF43, POLD1, and TP53) were not sufficient and did not provide excellent diagnostic performance. Additional mutations should be identified in the future to improve this diagnostic performance. Future national multicentric research is being implemented, with the collection of all pancreatic cysts of 12 mm-diameter and over. Future research should use the same methodology with comparative CF and NT analyses, but conducted on a larger scale and including the analysis of a higher number of mutations.
We thank the patients and nurses for their important contributions to this study.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: France
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C, C, C
Grade D (Fair): D
Grade E (Poor): 0
P-Reviewer: Abd El-Razek A, Sperti C, Yang ZH, Zhang XB, Wada R S-Editor: Yan JP L-Editor: Filipodia E-Editor: Liu MY
| 1. | Lee KS, Sekhar A, Rofsky NM, Pedrosa I. Prevalence of incidental pancreatic cysts in the adult population on MR imaging. Am J Gastroenterol. 2010;105:2079-2084. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 420] [Cited by in RCA: 436] [Article Influence: 29.1] [Reference Citation Analysis (0)] |
| 2. | Farrell JJ, Fernández-del Castillo C. Pancreatic cystic neoplasms: Management and unanswered questions. Gastroenterology. 2013;144:1303-1315. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 174] [Cited by in RCA: 144] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 3. | Crippa S, Fernández-Del Castillo C, Salvia R, Finkelstein D, Bassi C, Domínguez I, Muzikansky A, Thayer SP, Falconi M, Mino-Kenudson M, Capelli P, Lauwers GY, Partelli S, Pederzoli P, Warshaw AL. Mucin-producing neoplasms of the pancreas: An analysis of distinguishing clinical and epidemiologic characteristics. Clin Gastroenterol Hepatol. 2010;8:213-219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 236] [Cited by in RCA: 233] [Article Influence: 15.5] [Reference Citation Analysis (2)] |
| 4. | Marchegiani G, Mino-Kenudson M, Sahora K, Morales-Oyarvide V, Thayer S, Ferrone C, Warshaw AL, Lillemoe KD, Fernández-Del Castillo C. IPMN involving the main pancreatic duct: Biology, epidemiology, and long-term outcomes following resection. Ann Surg. 2015;261:976-983. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 100] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 5. | Bournet B, Kirzin S, Carrère N, Portier G, Otal P, Selves J, Musso C, Suc B, Moreau J, Fourtanier G, Pradère B, Lazorthes F, Escourrou J, Buscail L. Clinical fate of branch duct and mixed forms of intraductal papillary mucinous neoplasia of the pancreas. J Gastroenterol Hepatol. 2009;24:1211-1217. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 31] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 6. | Sugiyama M, Izumisato Y, Abe N, Masaki T, Mori T, Atomi Y. Predictive factors for malignancy in intraductal papillary-mucinous tumours of the pancreas. Br J Surg. 2003;90:1244-1249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 351] [Cited by in RCA: 308] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 7. | Kanno A, Satoh K, Hirota M, Hamada S, Umino J, Itoh H, Masamune A, Asakura T, Shimosegawa T. Prediction of invasive carcinoma in branch type intraductal papillary mucinous neoplasms of the pancreas. J Gastroenterol. 2010;45:952-959. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 74] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 8. | Crippa S, Salvia R, Warshaw AL, Domínguez I, Bassi C, Falconi M, Thayer SP, Zamboni G, Lauwers GY, Mino-Kenudson M, Capelli P, Pederzoli P, Castillo CF. Mucinous cystic neoplasm of the pancreas is not an aggressive entity: Lessons from 163 resected patients. Ann Surg. 2008;247:571-579. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 340] [Cited by in RCA: 274] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 9. | Tanaka M, Chari S, Adsay V, Fernandez-del Castillo C, Falconi M, Shimizu M, Yamaguchi K, Yamao K, Matsuno S; International Association of Pancreatology. International consensus guidelines for management of intraductal papillary mucinous neoplasms and mucinous cystic neoplasms of the pancreas. Pancreatology. 2006;6:17-32. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1539] [Cited by in RCA: 1441] [Article Influence: 75.8] [Reference Citation Analysis (0)] |
| 10. | Tanaka M, Fernández-del Castillo C, Adsay V, Chari S, Falconi M, Jang JY, Kimura W, Levy P, Pitman MB, Schmidt CM, Shimizu M, Wolfgang CL, Yamaguchi K, Yamao K; International Association of Pancreatology. International consensus guidelines 2012 for the management of IPMN and MCN of the pancreas. Pancreatology. 2012;12:183-197. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1714] [Cited by in RCA: 1614] [Article Influence: 124.2] [Reference Citation Analysis (0)] |
| 11. | Vege SS, Ziring B, Jain R, Moayyedi P; Clinical Guidelines Committee; American Gastroenterology Association. American gastroenterological association institute guideline on the diagnosis and management of asymptomatic neoplastic pancreatic cysts. Gastroenterology. 2015;148:819-22; quize12-3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 629] [Cited by in RCA: 759] [Article Influence: 75.9] [Reference Citation Analysis (1)] |
| 12. | Tanaka M, Fernández-Del Castillo C, Kamisawa T, Jang JY, Levy P, Ohtsuka T, Salvia R, Shimizu Y, Tada M, Wolfgang CL. Revisions of international consensus Fukuoka guidelines for the management of IPMN of the pancreas. Pancreatology. 2017;17:738-753. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 868] [Cited by in RCA: 1156] [Article Influence: 144.5] [Reference Citation Analysis (1)] |
| 13. | Elta GH, Enestvedt BK, Sauer BG, Lennon AM. ACG Clinical Guideline: Diagnosis and Management of Pancreatic Cysts. Am J Gastroenterol. 2018;113:464-479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 282] [Cited by in RCA: 423] [Article Influence: 60.4] [Reference Citation Analysis (1)] |
| 14. | Heckler M, Michalski CW, Schaefle S, Kaiser J, Büchler MW, Hackert T. The Sendai and Fukuoka consensus criteria for the management of branch duct IPMN - A meta-analysis on their accuracy. Pancreatology. 2017;17:255-262. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 54] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 15. | Sahora K, Mino-Kenudson M, Brugge W, Thayer SP, Ferrone CR, Sahani D, Pitman MB, Warshaw AL, Lillemoe KD, Fernandez-del Castillo CF. Branch duct intraductal papillary mucinous neoplasms: Does cyst size change the tip of the scale? A critical analysis of the revised international consensus guidelines in a large single-institutional series. Ann Surg. 2013;258:466-475. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 207] [Cited by in RCA: 227] [Article Influence: 18.9] [Reference Citation Analysis (0)] |
| 16. | Crippa S, Bassi C, Salvia R, Malleo G, Marchegiani G, Rebours V, Levy P, Partelli S, Suleiman SL, Banks PA, Ahmed N, Chari ST, Fernández-Del Castillo C, Falconi M. Low progression of intraductal papillary mucinous neoplasms with worrisome features and high-risk stigmata undergoing non-operative management: A mid-term follow-up analysis. Gut. 2017;66:495-506. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 147] [Cited by in RCA: 152] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 17. | Gillis A, Cipollone I, Cousins G, Conlon K. Does EUS-FNA molecular analysis carry additional value when compared to cytology in the diagnosis of pancreatic cystic neoplasm? A systematic review. HPB (Oxford). 2015;17:377-386. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 50] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 18. | Huang ES, Turner BG, Fernandez-Del-Castillo C, Brugge WR, Hur C. Pancreatic cystic lesions: Clinical predictors of malignancy in patients undergoing surgery. Aliment Pharmacol Ther. 2010;31:285-294. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 12] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 19. | Springer S, Wang Y, Dal Molin M, Masica DL, Jiao Y, Kinde I, Blackford A, Raman SP, Wolfgang CL, Tomita T, Niknafs N, Douville C, Ptak J, Dobbyn L, Allen PJ, Klimstra DS, Schattner MA, Schmidt CM, Yip-Schneider M, Cummings OW, Brand RE, Zeh HJ, Singhi AD, Scarpa A, Salvia R, Malleo G, Zamboni G, Falconi M, Jang JY, Kim SW, Kwon W, Hong SM, Song KB, Kim SC, Swan N, Murphy J, Geoghegan J, Brugge W, Fernandez-Del Castillo C, Mino-Kenudson M, Schulick R, Edil BH, Adsay V, Paulino J, van Hooft J, Yachida S, Nara S, Hiraoka N, Yamao K, Hijioka S, van der Merwe S, Goggins M, Canto MI, Ahuja N, Hirose K, Makary M, Weiss MJ, Cameron J, Pittman M, Eshleman JR, Diaz LA, Papadopoulos N, Kinzler KW, Karchin R, Hruban RH, Vogelstein B, Lennon AM. A combination of molecular markers and clinical features improve the classification of pancreatic cysts. Gastroenterology. 2015;149:1501-1510. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 378] [Cited by in RCA: 326] [Article Influence: 32.6] [Reference Citation Analysis (0)] |
| 20. | Singhi AD, McGrath K, Brand RE, Khalid A, Zeh HJ, Chennat JS, Fasanella KE, Papachristou GI, Slivka A, Bartlett DL, Dasyam AK, Hogg M, Lee KK, Marsh JW, Monaco SE, Ohori NP, Pingpank JF, Tsung A, Zureikat AH, Wald AI, Nikiforova MN. Preoperative next-generation sequencing of pancreatic cyst fluid is highly accurate in cyst classification and detection of advanced neoplasia. Gut. 2018;67:2131-2141. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 210] [Cited by in RCA: 264] [Article Influence: 37.7] [Reference Citation Analysis (0)] |
| 21. | Al-Haddad MA, Kowalski T, Siddiqui A, Mertz HR, Mallat D, Haddad N, Malhotra N, Sadowski B, Lybik MJ, Patel SN, Okoh E, Rosenkranz L, Karasik M, Golioto M, Linder J, Catalano MF. Integrated molecular pathology accurately determines the malignant potential of pancreatic cysts. Endoscopy. 2015;47:136-142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 23] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 22. | Andrews S. FastQC A Quality Control tool for High Throughput Sequence Data [cited 28 Jan 2019]. Babraham Bioinformatics. Available from: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/. |
| 23. | Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754-1760. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29052] [Cited by in RCA: 34409] [Article Influence: 2150.6] [Reference Citation Analysis (0)] |
| 24. | Okonechnikov K, Conesa A, García-Alcalde F. Qualimap 2: Advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics. 2016;32:292-294. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 440] [Cited by in RCA: 1101] [Article Influence: 110.1] [Reference Citation Analysis (0)] |
| 25. | Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G, Levy-Moonshine A, Jordan T, Shakir K, Roazen D, Thibault J, Banks E, Garimella KV, Altshuler D, Gabriel S, DePristo MA. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics. 2013;43:11.10.1-11.1033. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1565] [Cited by in RCA: 3583] [Article Influence: 398.1] [Reference Citation Analysis (0)] |
| 26. | Razek AAKA, Elfar E, Abubacker S. Interobserver agreement of computed tomography reporting standards for chronic pancreatitis. Abdom Radiol (NY). 2019;44:2459-2465. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 15] [Article Influence: 2.5] [Reference Citation Analysis (0)] |