Published online Mar 28, 2019. doi: 10.3748/wjg.v25.i12.1492
Peer-review started: November 27, 2018
First decision: January 11, 2019
Revised: January 29, 2019
Accepted: January 30, 2019
Article in press: January 30, 2019
Published online: March 28, 2019
Processing time: 121 Days and 22.7 Hours
Nonalcoholic fatty liver disease (NAFLD), the most common chronic liver disease, can progress into nonalcoholic steatohepatitis (NASH), cirrhosis, and even hepatocellular carcinoma. Bile acids such as ursodeoxycholic acid (UDCA) play an essential role in the pathogenesis of NAFLD by regulating the level of sterol regulatory element-binding protein (SREBP) 1c, but the underlying regulatory mechanism remains elusive. Increased evidence indicates that the AKT/mTOR/SREBP-1 signaling pathway is a key pathway to regulate hepatic cellular lipid metabolism. UDCA may regulate the AKT/mTOR/SREBP-1 signaling pathway to ameliorate hepatic lipid metabolism.
To investigate the functional mechanism of UDCA in an oleic acid (OA)-induced cellular model of NAFLD.
The cellular model of NAFLD was established using OA and treated with UDCA. First, the best concentration of UDCA was selected. For the best time-dependent assay, cells were stimulated with OA only or co-treated with OA and 2 mmol/L UDCA for 24 h, 48 h, and 72 h. Oil red O staining was used to observe the accumulation of intracellular lipids, while the intracellular contents of triglyceride, alanine aminotransferase (ALT), gamma-glutamyl transpeptidase (GGT), and aspartate aminotransferase (AST) were detected by enzymatic methods. Meanwhile, the expression levels of AKT/mTOR/SREBP-1 signaling pathway-related proteins were detected by real-time PCR and Western blot.
In the NAFLD cell model established with LO2 cells induced using OA, lipid accumulation was obvious. UDCA significantly inhibited lipid accumulation at different concentrations (especially 2 mmol/L) and decreased cell growth ability at different time points. The biochemical parameters like ALT, AST, and GGT were significant improved by UDCA. UDCA treatment vividly repressed the activation of AKT, mTOR, and CRTC2 and the expression of nSREBP-1 in LO2 cells induced with OA.
Our findings demonstrate the effect of UDCA in improving NAFLD. UDCA attenuates OA-induced hepatic steatosis mainly by regulation of AKT/mTOR/SREBP-1 signal transduction.
Core tip: Nonalcoholic fatty liver disease (NAFLD) is the most common form of liver disease. Many studies show that the disorder of hepatic lipid metabolism is the major pathogenesis. Increased evidence indicates that the AKT/mTOR/SREBP-1 signaling pathway is a key pathway to regulate hepatocellular lipid metabolism. At present, there are few studies on the mechanism of NAFLD with regard to hepatic lipid metabolism. We aimed to investigate the functional mechanism of ursodeoxycholic acid (UDCA) in the oleic acid-induced cellular model of NAFLD. The possible molecular mechanism and related targets of regulating hepatic lipid metabolism were explored, and the correlation between the occurrence of NAFLD and the AKT/mTOR/SREBP-1 signaling pathway was explored. We provided more sufficient experimental basis for clinical application of UDCA in the treatment of NAFLD.
- Citation: Hu J, Hong W, Yao KN, Zhu XH, Chen ZY, Ye L. Ursodeoxycholic acid ameliorates hepatic lipid metabolism in LO2 cells by regulating the AKT/mTOR/SREBP-1 signaling pathway. World J Gastroenterol 2019; 25(12): 1492-1501
- URL: https://www.wjgnet.com/1007-9327/full/v25/i12/1492.htm
- DOI: https://dx.doi.org/10.3748/wjg.v25.i12.1492
Nonalcoholic fatty liver disease (NAFLD) is characterized by macrovesicular fat accumulation of more than 5% in the hepatocytes of patients, due to the deposition of fat for reasons other than excessive alcohol use[1]. The prevalence of NAFLD ranges from 9%to 36.9% in different regions of the world, and is growing rapidly worldwide[2,3]. In some cases, the progression of NAFLD may lead to the formation of nonalcoholic steatohepatitis (NASH) or other types of liver diseases[4]. Hepatic lipid metabolism dysbiosis-induced hepatic steatosis is regarded as the major cause of NAFLD[5]. Thus, investigation of the proposed mechanisms of hepatic lipid metabolism in the pathogenesis of NAFLD is potentially contributing to improving the pathological symptoms of NAFLD and NAFLD-related liver disease.
As amphipathic molecules, bile acids (BAs) are generated from cholesterol oxidation in the liver and play an essential role in the progression of NAFLD[6,7]. At the right concentration, BAs function as a master regulator in the digestion and absorption of lipids. However, excessive BAs exert detrimental effects on hepatocytes[8]. BAs can maintain triglyceride (TG) homeostasis and act as a metabolic regulator of glucose uptake and lipid metabolism[9]. It is confirmed that BAs binding to activated farnesoid X receptor (FXR) in the liver inhibit sterol regulatory element-binding protein 1c (SREBP1c)-mediated lipogenesis[10,11]. Conversely, activation of FXR also suppresses synthesis of BAs when BAs are excessive[12]. The production of BAs is the main route of cholesterol catabolism and disturbance of the enterohepatic circulation of bile acid leads to the decrease of endogenous cholesterol[3]. Downregulation of endogenous cholesterol induces the generation of mature SREBP-1 which can initiate lipogenesis in turn. Thus, the complicated correlation of BAs and hepatic lipid metabolism is essential for the maintenance of liver homeostasis.
Ursodeoxycholic acid (UDCA) is one of the secondary BAs and has been widely used for the treatment of cholestatic liver diseases, such as gallstones, and non-surgical treatment of primary biliary cirrhosis, through its cytoprotective effect and anti-apoptotic activities in the liver[13,14]. UDCA-mediated glutathione synthesis through the activation of the PI3K/AKT/Nrf2 pathway exerts antioxidative action and is regarded as a potential treatment for chronic hepatitis[15]. Additionally, it has been reported that UDCA administration promotes the progression of lipogenesis in the livers of obese people by the antagonism of FXR[16]. In contrast, recent studies have indicated that UDCA and taurine-conjugated UDCA facilitate the improvement of abnormal glucose metabolism and insulin resistance[17]. UDCA also acts as a protection factor for hepatic steatosis and hepatitis in mice[18]. Although application studies of UDCA in NAFLD have been widely used and have achieved good results, the underlying mechanism of UDCA in regulating the pathogenesis of NAFLD remains largely unknown.
Herein, we discovered that UDCA administration reduced oleic acid (OA)-induced production of lipid droplets and disturbance of hepatic lipid metabolism by regulating the AKT/mTOR/SREBP-1 signaling pathway. These findings could possibly provide a great reference for clinical application of UDCA in NAFLD.
Human LO2 cells were purchased from Bogoo Biotechnology (Shanghai, China). Cells were cultured in RPMI 1640 medium (HyClone) containing 10% fetal bovine serum (FBS; Wisent) and incubated at 37 °C in a humidified atmosphere containing 50 mL/L CO2. An in vitro cell model of NAFLD was established by treating cells with 20 μg/mL OA for 48 h. In cell viability experiments, cells were treated with OA only or co-treated with OA and a concentration gradient of UDCA for 48 h. For time-dependent assays, cells were stimulated with OA only or co-treated with OA and 2 mmol/L UDCA for 24 h, 48 h, and 72 h. For the detection of the lipid profile and the involved signaling pathways, cells were incubated with OA only or co-treated with OA and 2 mmol/L UDCA for 72 h.
After treatment with OA or UDCA, cells were washed twice with PBS and fixed using 10% paraformaldehyde overnight. Then, cells were washed twice again and stained in oil red O solution (isopropanol: 0.5% oil red O, 3:2) for 20 min at room temperature. Oil red O solution was removed, and the cells were stained with haematoxylin for 30 s. Next, cells were differentiated using hydrochloric acid alcohol (1:1) for 5-10 s. After washing again, oil red O staining results were observed under an optical microscope.
LO2 cells were seeded in 100 μL in a 96-well plate at a density of 5 × 104 cells/mL and incubated at 37 °C in a humidified atmosphere containing 50 mL/L CO2 overnight. Then, cells were treated with OA and UDCA according to the method we described above. At different time points, 100 μL MTT (1 mg/mL) was added into the wells after removing the medium. After incubation for another 4 h, 150 μL DMSO was used to dissolve cells stained with MTT, and OD values were measured using a microplate reader at the wavelength of 570 nm. Relative cell viability was calculated according to the formula: OD value of experimental group/OD value of control group × 100%.
Levels of TG, alanine aminotransferase (ALT), aspartate aminotransferase (AST), and gamma-glutamyl transpeptidase (GGT) were measured using commercial kits according to the manufacturers’ instructions. ELISA kits for TG (SBJ-H0240), AST (SBJ-H0659), and GGT (SBJ-H0669) were obtained from Nanjing Senbeijia Biotechnology (Nanjing, China). An ELISA kit for ALT (E-EL-H0312c) was purchased from Elab science (Wuhan, China).
Total protein was extracted after cells were treated in the presence of OA and UDCA. Then, 30 μg of protein was used to perform immunoblotting. Primary antibodies against mSREBP-1 (AF6283), p-CRTC2 (AF8328), p-AKT (AF0016), and p-mTOR (AF3308) were purchased from Affinity Biosciences (OH, United States). Relative quantitative analysis of protein bands was performed using Quantity One analysis software (Bio-Rad).
Statistical analyses were performed using SPSS 21.0 (Chicago, IL, United States) software. All results were from three independent experiments, and are shown as the mean ± standard error of the mean. The results were analyzed with GraphPad Prism 5.0 (GraphPad Software Inc, United States) using one-way ANOVA. P < 0.05 was regarded as significantly different.
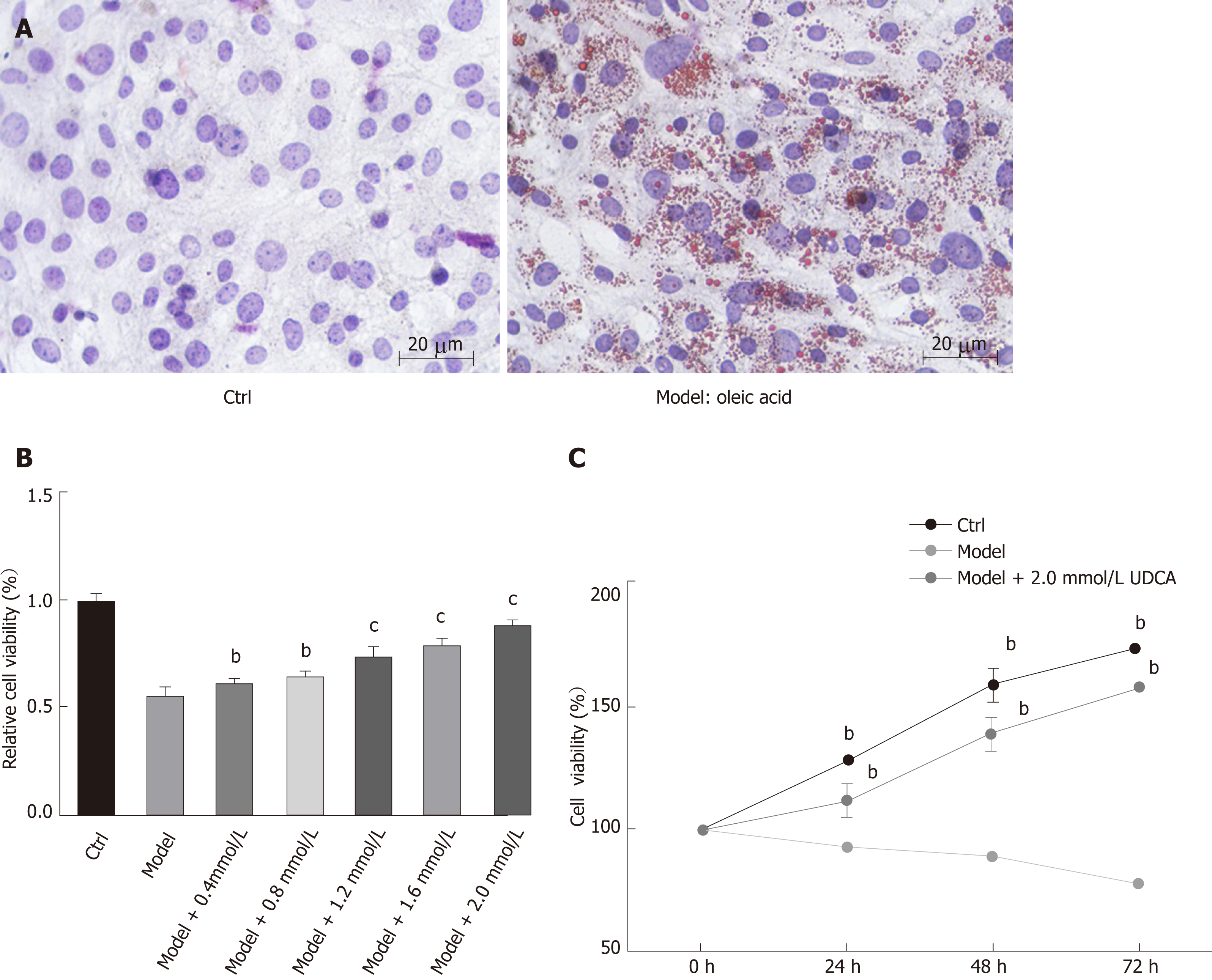
To determine the effect of UDCA on NAFLD, we first established a cell model of NAFLD. The oil red O staining analysis showed that 20 μg/mL OA significantly promoted the production of lipid droplets in LO2 cells, which suggested that we successfully developed the NAFLD cell model (Figure 1A, the right image). Given that OA treatment inhibited cell growth ability, we further evaluated the proliferation ability of the OA-induced LO2 cells treated with different concentrations of UDCA using MTT assay. When exposed to 20 μg/mL OA for 48 h, LO2 cell proliferation was inhibited significantly. This inhibition of proliferation by OA can be overcome to varying degrees by the addition of different levels of UDCA, especially under the condition of 2 mmol/L UDCA (cP < 0.001, Figure 1B). Additionally, relative cell viability of LO2 cells prominently declined in the presence of OA at 24 h, 48 h, and 72 h in comparison with the control cells. However, when treated with 2 mmol/L UDCA for 24 h, 48 h, and 72 h, OA-mediated reduction of cell viability at different time points was notably upregulated; cell viability was close to the normal level at 72 h (bP < 0.01, Figure 1C). These data indicate that 2 mmol/L UDCA was effective for improving OA-induced cell growth inhibition.
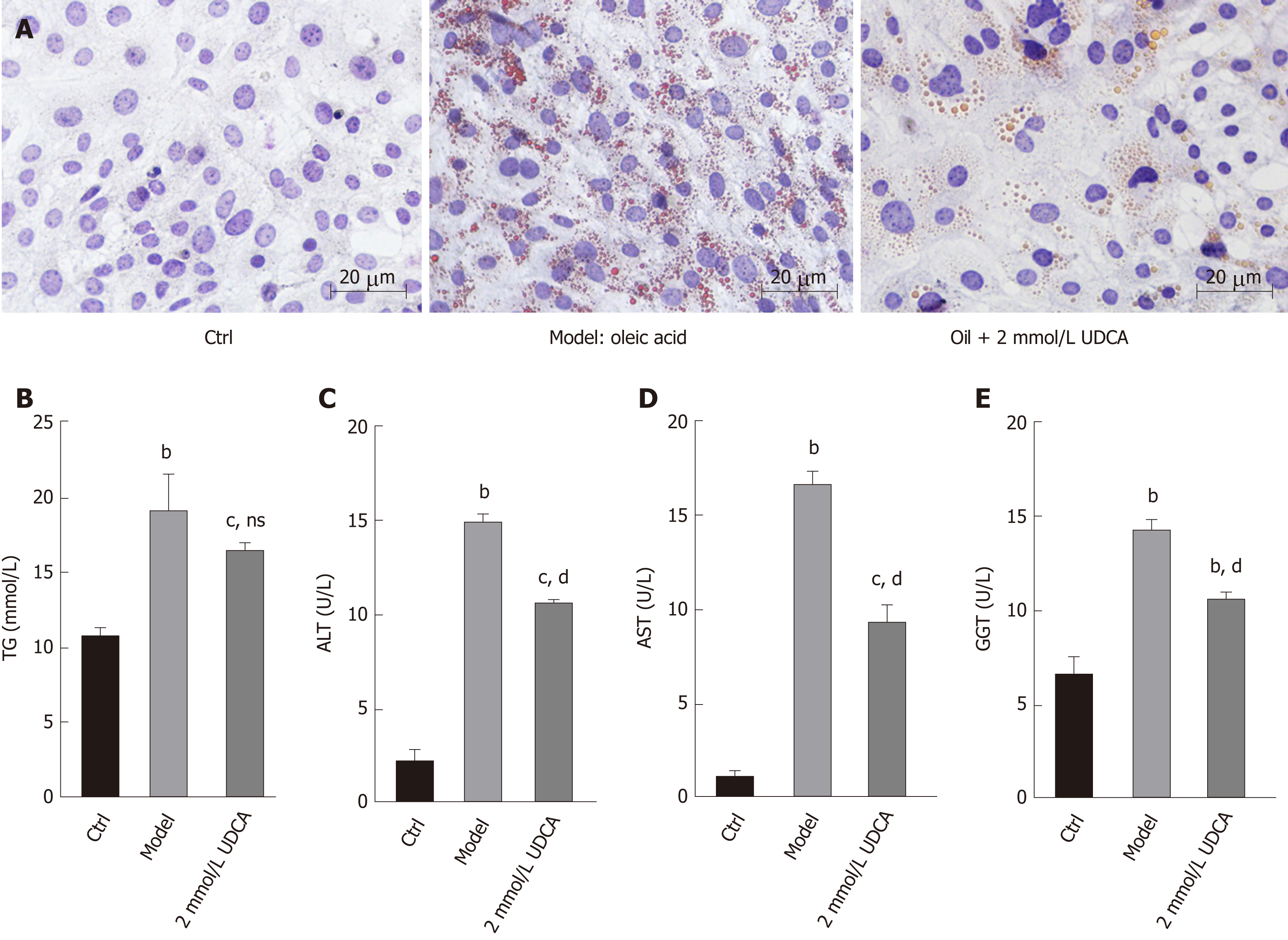
The production of lipid droplets is closely related to various biochemical parameters. The levels of lipid metabolic indexes such as TG, ALT, AST, and GGT can accurately reflect the state of hepatic lipid metabolism. As shown in Figure 2A, OA treatment-induced massive lipid accumulation of LO2 cells was dramatically suppressed under the treatment with UDCA for 72 h (Figure 2A, the middle image vs the right image). In addition, OA-induced LO2 cells showed a prominent increase of TG (1.5-fold), ALT (6-fold), AST (11-fold), and GGT (2-fold) levels compared to the control group. Upon the treatment with UDCA, the upregulation of ALT, AST, and GGT was significantly blocked (bP < 0.01, Figure 2C-E), whereas UDCA treatment had no significant inhibitory effect on TG level in OA-exposed cells (Figure 2B). Although UDCA administration partly attenuated hepatic steatosis, there was a high level of lipid profile (TG, ALT, AST, and GGT) compared with the control group, which implied that adjuvant therapy was needed to better improve the disorder of hepatic lipid metabolism.
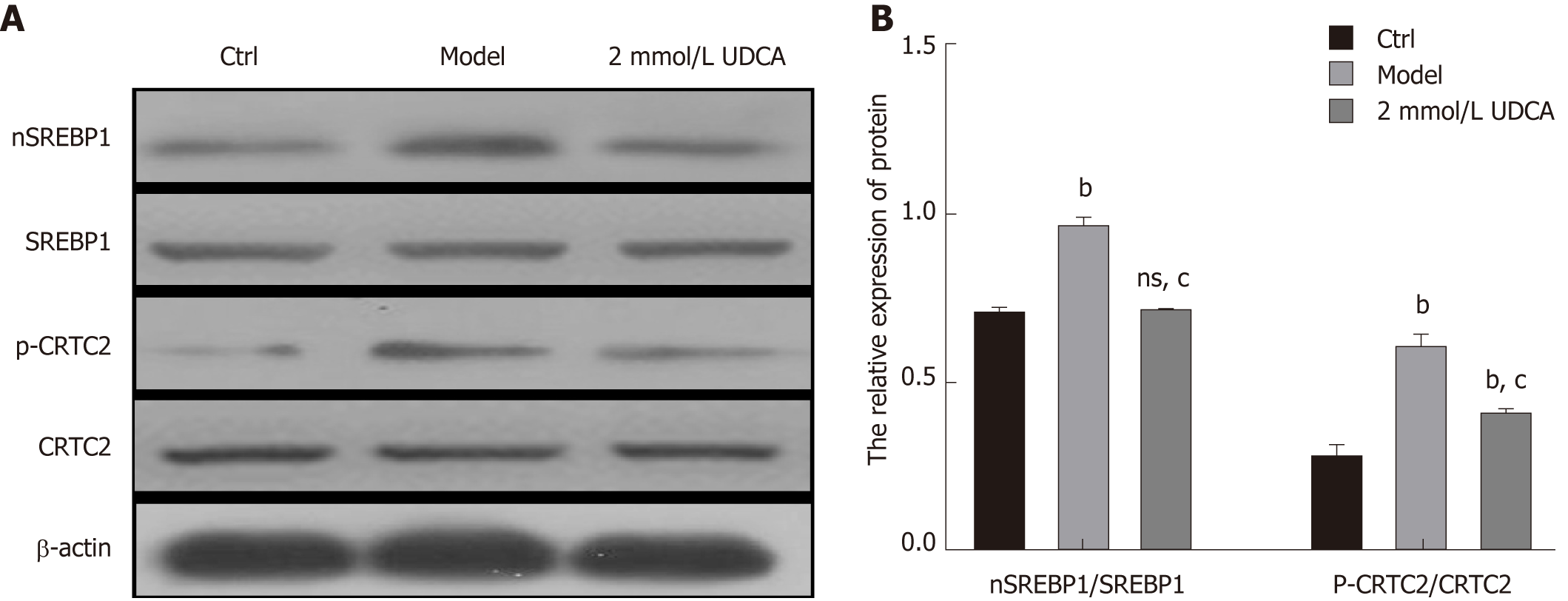
In the course of hepatic lipid metabolism, the cyclic adenosine monophosphate (cAMP) response element‐binding protein (CREB)‐regulated transcription coactivator 2 (CRTC2) mediates the activation of mature SREBP-1 (nSREBP-1) which subsequently upregulates lipogenic gene expression and controls the biosynthesis of fatty acids and TG in hepatocytes[19]. To further confirm the protective effect of UDCA on excessive lipid synthesis, we determined the expression pattern of nSREBP-1 and activated CTRC2 in OA-exposed LO2 cells. Once treated with OA for 72 h, phosphorylated CRTC2 and mature SREBP-1 (nSREBP-1) showed a 1.5-fold and 2.5-fold increase, respectively (bP < 0.01, Figure 3A and B, the second band vs the first band, the second box vs the first box, and the fifth box vs the fourth box). When exposed to UDCA, the enhancement of activated CRTC2 and SREBP-1 robustly declined compared with OA-exposed cells (cP < 0.01, Figure 3A and B, the second band vs the third band, the second box vs the third box, and the fifth box vs the sixth box). These results suggest that CRTC2/SREBP-1 signaling transduction is implicated in the UDCA-mediated protective effect on lipid accumulation.
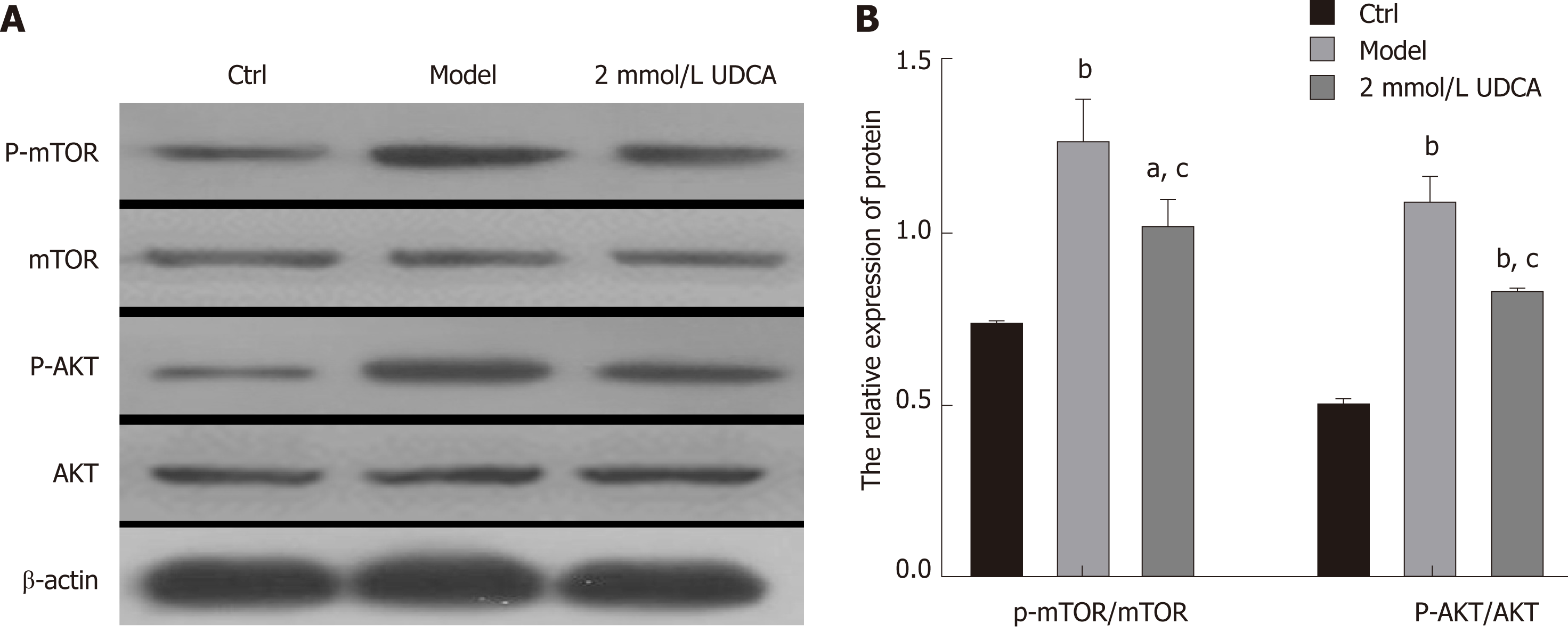
It is known that AKT activation increases the activity of its downstream effector mTOR, which then promotes the activation of mature SREBP-1 by integrating CRTC2[20,21]. Thus, we performed an immunoblotting assay to detect the changes of AKT/mTOR signaling and found that the activation of AKT and mTOR presented a 3-fold and 1.5-fold increase, respectively, under the condition of OA treatment for 72 h (Figure 4A and B, the second band vs the first band, the second box vs the first box, the fifth box vs the fourth box, and the eighth box vs the seventh box). UDCA administration recovered the levels of phosphorylated AKT and mTOR in OA-induced LO2 cells (aP < 0.05, bP < 0.01, Figure 4A and B, the second band vs the third band, the second box vs the third box, the fifth box vs the sixth box, and the eighth box vs the ninth box). Based on these results, we believe that UDCA ameliorates lipid metabolic disorder partly by regulating the AKT/mTOR signaling pathway.
There is increasing evidence that disorder of hepatic lipid metabolism is not only a hallmark of NAFLD, but also a biologic marker of cancer cells[22,23]. Numerous key enzymes involved in the progression of de novo fatty acid synthesis are enhanced and correlated with poor clinical outcomes in hepatic carcinoma (HCC)[24]. That means the importance of homeostasis of hepatic lipid metabolism is imaginable in liver diseases. In the present study, we found that the hydrophilic BA, UDCA, prevented lipogenesis by regulating the expression of mature SREBP-1 in LO2 cells, which suggested that UDCA administration might improve lipid metabolic disorder in NAFLD patients and slow the progression of metabolic disturbance-derived liver cancer.
Previous studies have pointed that OA-induced hepatic steatosis is always accompanied by oxide stress, cell apoptosis, and cell viability reduction in hepatocytes[25,26]. Oxidative stress acting as an apoptotic mediator of liver cells can cause damage of membrane lipid peroxidation and cell degeneration in the progression of NAFLD[27,28]. Here, we observed that 2 mmol/L UDCA significantly relieved the growth inhibition of OA-exposed LO2 cells, implying that 2 mmol/L was a safe and effective dosage for in vitro study. UDCA exerts a protective effect on NAFLD in rats by reducing the lipid profile including total cholesterol (TC) and TG[29]. Treatment with UDCA suppressed activated liver X receptor α (LXRα)-mediated hepatic lipogenesis in high-fat diet (HFD)-fed mice[30]. These reports indicate that UDCA can serve as a regulator of lipid metabolic indexes in abnormal hepatic lipid metabolism-related liver diseases. Our findings indicated that administration with UDCA ameliorated OA-induced disturbance of liver dysfunction such as the increase of ALT, AST, and GGT levels in LO2 cells. UDCA treatment cannot result in a significant reduction of TG level in OA-induced cells, but had a certain regulatory role. Therefore, our data further supported the protective effect of UDCA in improving hepatocyte fatty degeneration and relieving the inflammatory reaction.
SREBP-1, a member of basic helix–loop–helix-leucine zipper transcription factor family, is mainly responsible for regulating fatty acid and cholesterol synthesis[31,32]. SREBP-1 is synthesized as a precursor located on the nuclear membrane and endoplasmic reticulum[33]. Upon stimulus or sterol depletion, the precursor (full-length SREBP-1) is cleaved into mature SREBP-1 which subsequently translocates to the nucleus (nSREBP-1) and activates transcription by binding to the promoter region of genes containing sterol regulatory element-1 (SREBP-1)[34,35]. Once SREBP-1 is activated, nSREBP-1 enhances the expression of cholesterol and fatty acid synthesis related genes[36]. In the course of lipid metabolism, activated AKT subsequently induces the activation of mTOR, which leads to the upregulation of its downstream CRTC2[37,38]. mTOR is an essential regulator of lipogenic metabolism by activating SREBP-1 cleavage[39]. CRTC2 as a mediator of the mTOR molecule, can modulate hepatic lipid metabolism by affecting COPII-dependent SREBP1 processing[21], which indicates that AKT-induced activation of mTOR can recruit CRTC2 as a complex to promote nSREBP-1 activity and the following lipogenesis. The present research demonstrated that CRTC2 activation, the AKT/MTOR signaling pathway, and nSREBP-1 expression were enhanced in the presence of OA. UDCA stimulation notably mitigated OA-initiated lipid metabolic disorder by suppressing this signaling pathway transduction. It is possible that UDCA attenuates OA-induced hepatic steatosis mainly by regulation of the AKT/mTOR/SREBP-1 signaling transduction.
In conclusion, we illustrated the protective role of UDCA in NAFLD. At a proper concentration of UDCA, OA-evoked lipid accumulation was nearly blocked by affecting SREBP-1 cleavage, implying that UDCA may contribute to developing more effective BA-based therapies for NAFLD. At present, the effects of UDCA on NAFLD in the clinic have seldom been reported, and especially the research of UDCA on liver histology is also limited. We carried out research at the cellular level to explore the AKT/mTOR/SREBP-1 signaling pathway as a new target for NAFLD treatment. Our research is expected to enrich the academic research content in this field and provide scientific basis for clinical application.
Ursodeoxycholic acid (UDCA) plays an essential role in the pathogenesis of nonalcoholic fatty liver disease (NAFLD), but the underlying regulatory mechanism remains elusive. Increased evidence indicates that the AKT/mTOR/SREBP-1 signaling pathway is a key pathway to regulate cellular hepatic lipid metabolism. UDCA may regulate the AKT/mTOR/SREBP-1 signaling pathway to ameliorate hepatic lipid metabolism. However, no report has discussed the regulatory effects of UDCA on lipid metabolism in vitro.
Although UDCA has been widely used in the treatment of NAFLD, the mechanism remains unclear. Understanding and regulating the AKT/mTOR/SREBP-1 signaling pathway using UDCA will be an important area for future research.
This study aimed to explore the protective effects of UDCA on NAFLD by regulating the AKT/mTOR/SREBP-1 signaling pathway. This will provide a therapeutic target for future clinical applications of UDCA to prevent and improve hepatic lipid metabolism disorders related diseases.
First, we established a cellular model of NAFLD using LO2 cells induced with oleic acid (OA). Second, we used a MTT assay to determine the optimal concentration of OA for inducing hepatocyte steatosis and the best treatment duration of UDCA. Third, oil red O staining was used to observe the formation of intracellular lipid droplets, and subsequently we analyzed the intracellular levels of biochemical indexes with UDCA treatment by enzymatic methods. Finally, we examined the effects of UDCA on the expression levels of the AKT/mTOR/SREBP-1 signaling pathway related proteins by real-time PCR and Western blot.
UDCA inhibited the lipid accumulation of LO2 cells induced with OA. UDCA decreased cell growth ability under the treatment of OA in a dose- and time-dependent manner. UDCA treatment significantly improved the biochemical parameters like alanine aminotransferase, gamma-glutamyl transpeptidase, and aspartate aminotransferase. UDCA suppressed the OA-induced upregulation of AKT, mTOR, CRTC2, and nSREBP-1 expression in LO2 cells.
UDCA administration improves OA induced growth inhibition of LO2 cells and relieves OA induced lipid accumulation. UDCA has a protective effect in improving hepatic steatosis and amending liver function in our in vitro experiments. We may provide the first evidence of the AKT/mTOR/SREBP-1 signaling pathway regulated by UDCA in OA-induced LO2 cells as an in vitro model of NAFLD. Our hypothesis was confirmed using LO2 cells treated with OA. We present a novel theory here that UDCA can ameliorate hepatic lipid metabolism via the AKT/mTOR/SREBP-1 signaling pathway. Thus, UDCA and other bile acids may have novel clinical applications to ameliorate hepatic lipid metabolism for NAFLD in future.
Our findings provide evidence that UDCA has the effect of ameliorating hepatic lipid metabolism. This study has some limitations and further studies using animals and further in vitro research with agonists and inhibitors of mTOR as tools should be performed.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): D
Grade E (Poor): 0
P-Reviewer: Colak Y, Joven J, Lee MK S-Editor: Ma RY L-Editor: Wang TQ E-Editor: Ma YJ
| 1. | Yuan L, Bambha K. Bile acid receptors and nonalcoholic fatty liver disease. World J Hepatol. 2015;7:2811-2818. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 69] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 2. | Shen L, Fan JG, Shao Y, Zeng MD, Wang JR, Luo GH, Li JQ, Chen SY. Prevalence of nonalcoholic fatty liver among administrative officers in Shanghai: an epidemiological survey. World J Gastroenterol. 2003;9:1106-1110. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 130] [Cited by in RCA: 123] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 3. | Chow MD, Lee YH, Guo GL. The role of bile acids in nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Mol Aspects Med. 2017;56:34-44. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 112] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 4. | McCullough AJ. The clinical features, diagnosis and natural history of nonalcoholic fatty liver disease. Clin Liver Dis. 2004;8:521-533, viii. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 382] [Cited by in RCA: 397] [Article Influence: 18.9] [Reference Citation Analysis (0)] |
| 5. | Almeda-Valdés P, Cuevas-Ramos D, Aguilar-Salinas CA. Metabolic syndrome and non-alcoholic fatty liver disease. Ann Hepatol. 2009;8 Suppl 1:S18-S24. [PubMed] |
| 6. | Arab JP, Karpen SJ, Dawson PA, Arrese M, Trauner M. Bile acids and nonalcoholic fatty liver disease: Molecular insights and therapeutic perspectives. Hepatology. 2017;65:350-362. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 349] [Cited by in RCA: 457] [Article Influence: 57.1] [Reference Citation Analysis (1)] |
| 7. | Merlen G, Ursic-Bedoya J, Jourdainne V, Kahale N, Glenisson M, Doignon I, Rainteau D, Tordjmann T. Bile acids and their receptors during liver regeneration: "Dangerous protectors". Mol Aspects Med. 2017;56:25-33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 37] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 8. | Carulli L, Gabbi C, Bertolotti M. Bile acids and nonalcoholic fatty liver disease: An intriguing relationship. Hepatology. 2016;63:1739-1740. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 9. | Kobayashi Y, Hara N, Sugimoto R, Mifuji-Moroka R, Tanaka H, Eguchi A, Iwasa M, Hasegawa H, Iwata K, Takei Y, Taguchi O. The Associations between Circulating Bile Acids and the Muscle Volume in Patients with Non-alcoholic Fatty Liver Disease (NAFLD). Intern Med. 2017;56:755-762. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 28] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 10. | Han B, Kim BK, Kim K, Fang SS. Essential roles of bile acids and their nuclear receptors, FXR and PXR, in the cholestatic liver disease. Anim Cells Syst. 2016;20:175-178. [RCA] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 11. | Zhu Y, Liu H, Zhang M, Guo GL. Fatty liver diseases, bile acids, and FXR. Acta Pharm Sin B. 2016;6:409-412. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 67] [Cited by in RCA: 93] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 12. | Manley S, Ding W. Role of farnesoid X receptor and bile acids in alcoholic liver disease. Acta Pharm Sin B. 2015;5:158-167. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 83] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 13. | Lim SC, Duong HQ, Parajuli KR, Han SI. Pro-apoptotic role of the MEK/ERK pathway in ursodeoxycholic acid-induced apoptosis in SNU601 gastric cancer cells. Oncol Rep. 2012;28:1429-1434. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 25] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 14. | Tonin F, Arends IWCE. Latest development in the synthesis of ursodeoxycholic acid (UDCA): a critical review. Beilstein J Org Chem. 2018;14:470-483. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 71] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 15. | Arisawa S, Ishida K, Kameyama N, Ueyama J, Hattori A, Tatsumi Y, Hayashi H, Yano M, Hayashi K, Katano Y, Goto H, Takagi K, Wakusawa S. Ursodeoxycholic acid induces glutathione synthesis through activation of PI3K/Akt pathway in HepG2 cells. Biochem Pharmacol. 2009;77:858-866. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 50] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 16. | Mueller M, Thorell A, Claudel T, Jha P, Koefeler H, Lackner C, Hoesel B, Fauler G, Stojakovic T, Einarsson C, Marschall HU, Trauner M. Ursodeoxycholic acid exerts farnesoid X receptor-antagonistic effects on bile acid and lipid metabolism in morbid obesity. J Hepatol. 2015;62:1398-1404. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 210] [Cited by in RCA: 234] [Article Influence: 23.4] [Reference Citation Analysis (0)] |
| 17. | Ratziu V, de Ledinghen V, Oberti F, Mathurin P, Wartelle-Bladou C, Renou C, Sogni P, Maynard M, Larrey D, Serfaty L, Bonnefont-Rousselot D, Bastard JP, Rivière M, Spénard J; FRESGUN. A randomized controlled trial of high-dose ursodesoxycholic acid for nonalcoholic steatohepatitis. J Hepatol. 2011;54:1011-1019. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 208] [Cited by in RCA: 236] [Article Influence: 16.9] [Reference Citation Analysis (0)] |
| 18. | Ozcan U, Yilmaz E, Ozcan L, Furuhashi M, Vaillancourt E, Smith RO, Görgün CZ, Hotamisligil GS. Chemical chaperones reduce ER stress and restore glucose homeostasis in a mouse model of type 2 diabetes. Science. 2006;313:1137-1140. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1994] [Cited by in RCA: 1959] [Article Influence: 103.1] [Reference Citation Analysis (0)] |
| 19. | Matsuzaka T, Shimano H. Novel role for the CRTC2 in lipid homeostasis. J Diabetes Investig. 2016;7:677-679. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 20. | Lim W, Yang C, Bazer FW, Song G. Luteolin Inhibits Proliferation and Induces Apoptosis of Human Placental Choriocarcinoma Cells by Blocking the PI3K/AKT Pathway and Regulating Sterol Regulatory Element Binding Protein Activity. Biol Reprod. 2016;95:82. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 47] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 21. | Han J, Li E, Chen L, Zhang Y, Wei F, Liu J, Deng H, Wang Y. The CREB coactivator CRTC2 controls hepatic lipid metabolism by regulating SREBP1. Nature. 2015;524:243-246. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 153] [Cited by in RCA: 237] [Article Influence: 23.7] [Reference Citation Analysis (0)] |
| 22. | Piscaglia F, Svegliati-Baroni G, Barchetti A, Pecorelli A, Marinelli S, Tiribelli C, Bellentani S; HCC-NAFLD Italian Study Group. Clinical patterns of hepatocellular carcinoma in nonalcoholic fatty liver disease: A multicenter prospective study. Hepatology. 2016;63:827-838. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 466] [Cited by in RCA: 465] [Article Influence: 51.7] [Reference Citation Analysis (0)] |
| 23. | Glaysher J. Lipid metabolism and cancer. Curr Opin Lipidol. 2013;24:530-531. [RCA] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 15] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 24. | Menendez JA, Lupu R. Fatty acid synthase and the lipogenic phenotype in cancer pathogenesis. Nat Rev Cancer. 2007;7:763-777. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1925] [Cited by in RCA: 2205] [Article Influence: 122.5] [Reference Citation Analysis (0)] |
| 25. | Patel TP, Rawal K, Soni S, Gupta S. Swertiamarin ameliorates oleic acid induced lipid accumulation and oxidative stress by attenuating gluconeogenesis and lipogenesis in hepatic steatosis. Biomed Pharmacother. 2016;83:785-791. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 39] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 26. | Xie C, Chen Z, Zhang C, Xu X, Jin J, Zhan W, Han T, Wang J. Dihydromyricetin ameliorates oleic acid-induced lipid accumulation in L02 and HepG2 cells by inhibiting lipogenesis and oxidative stress. Life Sci. 2016;157:131-139. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 70] [Article Influence: 7.8] [Reference Citation Analysis (1)] |
| 27. | Berardo C, Palladini G, Pasqua LG, Rizzo V, Perlini S, Richelmi P, Vairetti M, Ferrigno A. Oxidative stress, mitochondria damage and matrix metalloprotease activation in the pathogenesis of NAFLD. Digest Liver Dis. 2015;7 Suppl 1:e60-e61. [RCA] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 28. | Sumida Y, Niki E, Naito Y, Yoshikawa T. Involvement of free radicals and oxidative stress in NAFLD/NASH. Free Radic Res. 2013;47:869-880. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 181] [Cited by in RCA: 226] [Article Influence: 18.8] [Reference Citation Analysis (0)] |
| 29. | Ali MH, Messiha BA, Abdel-Latif HA. Protective effect of ursodeoxycholic acid, resveratrol, and N-acetylcysteine on nonalcoholic fatty liver disease in rats. Pharm Biol. 2016;54:1198-1208. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 18] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 30. | Lee JM, Gang GT, Kim DK, Kim YD, Koo SH, Lee CH, Choi HS. Ursodeoxycholic acid inhibits liver X receptor α-mediated hepatic lipogenesis via induction of the nuclear corepressor SMILE. J Biol Chem. 2014;289:1079-1091. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 39] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 31. | Brown MS, Goldstein JL. The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell. 1997;89:331-340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2786] [Cited by in RCA: 2901] [Article Influence: 103.6] [Reference Citation Analysis (0)] |
| 32. | Peng CH, Yang MY, Yang YS, Yu CC, Wang CJ. Antrodia cinnamomea Prevents Obesity, Dyslipidemia, and the Derived Fatty Liver via Regulating AMPK and SREBP Signaling. Am J Chin Med. 2017;45:67-83. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 33] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 33. | Yokoyama C, Wang X, Briggs MR, Admon A, Wu J, Hua X, Goldstein JL, Brown MS. SREBP-1, a basic-helix-loop-helix-leucine zipper protein that controls transcription of the low density lipoprotein receptor gene. Cell. 1993;75:187-197. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 595] [Cited by in RCA: 607] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 34. | Horton JD, Goldstein JL, Brown MS. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest. 2002;109:1125-1131. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 99] [Cited by in RCA: 1747] [Article Influence: 76.0] [Reference Citation Analysis (0)] |
| 35. | Xu H, Luo J, Ma G, Zhang X, Yao D, Li M, Loor JJ. Acyl-CoA synthetase short-chain family member 2 (ACSS2) is regulated by SREBP-1 and plays a role in fatty acid synthesis in caprine mammary epithelial cells. J Cell Physiol. 2018;233:1005-1016. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 60] [Cited by in RCA: 85] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 36. | Eberlé D, Hegarty B, Bossard P, Ferré P, Foufelle F. SREBP transcription factors: master regulators of lipid homeostasis. Biochimie. 2004;86:839-848. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 929] [Cited by in RCA: 1121] [Article Influence: 53.4] [Reference Citation Analysis (0)] |
| 37. | Han C, Wei S, Song Q, He F, Xiong X, Wan H, Liu D, Ye F, Liu H, Li L, Xu H, Du X, Kang B, Zeng X. Insulin Stimulates Goose Liver Cell Growth by Activating PI3K-AKT-mTOR Signal Pathway. Cell Physiol Biochem. 2016;38:558-570. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 19] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 38. | Yecies JL, Zhang HH, Menon S, Liu S, Yecies D, Lipovsky AI, Gorgun C, Kwiatkowski DJ, Hotamisligil GS, Lee CH, Manning BD. Akt stimulates hepatic SREBP1c and lipogenesis through parallel mTORC1-dependent and independent pathways. Cell Metab. 2011;14:21-32. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 532] [Cited by in RCA: 509] [Article Influence: 36.4] [Reference Citation Analysis (0)] |
| 39. | Bakan I, Laplante M. Connecting mTORC1 signaling to SREBP-1 activation. Curr Opin Lipidol. 2012;23:226-234. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 201] [Article Influence: 15.5] [Reference Citation Analysis (0)] |