Published online Oct 21, 2015. doi: 10.3748/wjg.v21.i39.11088
Peer-review started: April 24, 2015
First decision: June 2, 2015
Revised: June 11, 2015
Accepted: September 2, 2015
Article in press: September 2, 2015
Published online: October 21, 2015
Processing time: 178 Days and 17.6 Hours
In the Western world, nonalcoholic fatty liver disease (NAFLD) is considered as one of the most significant liver diseases of the twenty-first century. Its development is certainly driven by environmental factors, but it is also regulated by genetic background. The role of heritability has been widely demonstrated by several epidemiological, familial, and twin studies and case series, and likely reflects the wide inter-individual and inter-ethnic genetic variability in systemic metabolism and wound healing response processes. Consistent with this idea, genome-wide association studies have clearly identified Patatin-like phosholipase domain-containing 3 gene variant I148M as a major player in the development and progression of NAFLD. More recently, the transmembrane 6 superfamily member 2 E167K variant emerged as a relevant contributor in both NAFLD pathogenesis and cardiovascular outcomes. Furthermore, numerous case-control studies have been performed to elucidate the potential role of candidate genes in the pathogenesis and progression of fatty liver, although findings are sometimes contradictory. Accordingly, we performed a comprehensive literature search and review on the role of genetics in NAFLD. We emphasize the strengths and weaknesses of the available literature and outline the putative role of each genetic variant in influencing susceptibility and/or progression of the disease.
Core tip: Nonalcoholic fatty liver disease (NAFLD) is regarded as the most significant liver disease from the twenty-first century in the Western world. Although its development is surely driven by environmental factors, it is also regulated by genetic background. The role of heritability has been widely demonstrated by several studies, likely reflecting the diverse genetic variability in systemic metabolism and wound healing response processes. Accordingly, we performed a review of the literature on the role of genetics in NAFLD and outlined here the putative role of each genetic variant in influencing susceptibility and/or progression of the disease.
- Citation: Macaluso FS, Maida M, Petta S. Genetic background in nonalcoholic fatty liver disease: A comprehensive review. World J Gastroenterol 2015; 21(39): 11088-11111
- URL: https://www.wjgnet.com/1007-9327/full/v21/i39/11088.htm
- DOI: https://dx.doi.org/10.3748/wjg.v21.i39.11088
Nonalcoholic fatty liver disease (NAFLD) consists of a spectrum of disorders characterized predominantly by macrovesicular hepatic steatosis in absence of significant alcohol consumption. In this context, it is correct to discriminate between a condition of simple fatty liver, where the only histological finding is the presence of steatosis, and a state of nonalcoholic steatohepatitis (NASH), featured by hepatocellular injury and inflammation, with or without fibrosis[1]. NAFLD is regarded as the most relevant liver disease of the twenty-first century. Indeed, it has been estimated that NAFLD affects approximately 1 billion individuals worldwide[2]. It is the number one cause of altered aminotransferases in the Western world[3], where at least one third of the population is affected[4]. Importantly, a considerable proportion of NAFLD subjects (20%-30%) develop NASH, and this condition, as opposed to simple fatty liver, is a potentially progressive hepatic disorder that can lead to end-stage liver disease and hepatocellular carcinoma (HCC)[5]. In addition, several lines of evidence clearly demonstrated that all NAFLD/NASH patients are at high risk of cardiovascular diseases, type 2 diabetes (T2D), kidney failure, and colorectal cancer[6]. In this complex scenario, NAFLD development is surely driven by environmental factors - particularly dietary habits and a sedentary lifestyle - but it also requires a background of genetic susceptibility. Indeed, the real explanation for the wide inter-individual variability in the occurrence of NAFLD and progression to NASH - after correction for environmental factors - is provided by heritability. Much data has been accumulated over the years about the burden of heritability in NAFLD, as provided by epidemiological, familial, twin studies, and case series[7-10]. Furthermore, racial and ethnic differences have been reported in the prevalence of NAFLD, where it is most common in East Asian Indians, followed by Hispanics, Asians, Caucasians, and less frequently in African Americans[11-13]. In addition to differential exposure to metabolic risk factors, genetic variability in metabolism and wound healing response have surely influenced - at least in part - such differences. Not by chance, a great amount of evidence on the role of genetics in NAFLD/NASH has been produced during the last 10-15 years. Genetic studies can be divided into two categories: candidate gene studies and genome-wide association studies (GWAS). A GWAS is a hypothesis-free method for testing the association between all common variants in the human genome and polymorphic traits, such as diseases, drug response, and others. It is a powerful and statistically poorly biased method. On the other hand, candidate gene studies are generally derived from the results of previous genomic/proteomic and/or animal studies, where then a candidate gene is selected to investigate its putative role in the pathogenesis of a disease through a case-control single nucleotide polymorphism (SNP) study, with all potential methodological limits inherent to such type of study[14].
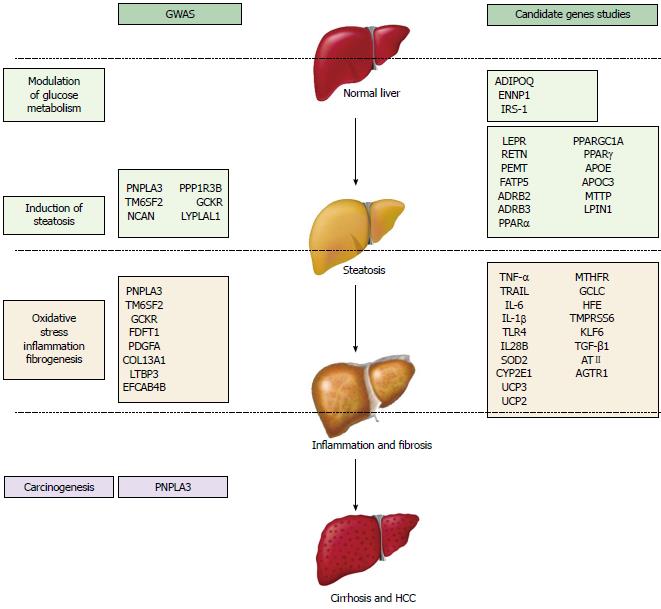
In this review, we have attempted to perform a comprehensive summary of the literature on the role of genetics in NAFLD/NASH, including the most recent evidence on genetic variants identified both by GWAS and candidate gene studies. Furthermore, we emphasize the strengths and weaknesses of the available literature for each variant, trying also to outline their putative role in NAFLD/NASH susceptibility and disease progression (Figure 1). Despite recent progress, several key issues remain to be addressed in the next years, particularly the details about the interaction between genetic background and acquired risk factors in disease pathogenesis and response to current treatments.
Romeo et al[15] was the first to report that the rs738409 C>G SNP in the Patatin-like phospholipase domain-containing 3 (PNPLA3) gene, encoding the isoleucine to methionine variant at protein position 148 (I148M), was strongly associated with increased liver fat content. Since then, several other pieces of evidence have highlighted the role of PNPLA3 in the development and progression of NAFLD. Furthermore, other SNPs have been identified by GWAS (Table 1). Among them, transmembrane 6 superfamily member 2 (TM6SF2) E167K variant is currently emerging as another relevant contributor both for NAFLD pathogenesis and cardiovascular outcomes.
| Gene | SNP | Association with |
| PNPLA3, patatine-like phospholipase domain containing 3 | rs738409 | Steatosis |
| NASH/necroinflammation | ||
| Severity of fibrosis | ||
| HCC development | ||
| TM6SF2, transmembrane 6 superfamily member 2 | rs58542926 | Steatosis |
| NASH/necroinflammation | ||
| Severity of fibrosis | ||
| Reduced cardiovascular risk | ||
| NCAN, neurocan | rs2228603 | Steatosis |
| PPP1R3B, protein phosphatase 1 regulatory subunit 3b | rs4240624 | Steatosis |
| GCKR, glucokinase regulatory protein | rs780094 | Steatosis |
| Severity of fibrosis | ||
| LYPLAL1, lysophospholipase-like 1 | rs12137855 | Steatosis |
| FDFT1, farnesyl diphosphate farnesyl transferase 1 | rs2645424 | NAFLD activity score |
| PDGFA, platelet-derived growth factor alpha | rs343062 | Severity of fibrosis |
| COL13A1, collagen type XIII alpha1 | rs1227756 | Lobular inflammation |
| LTBP3, latent transforming growth factor-beta-protein 3 | rs6591182 | Lobular inflammation |
| EFCAB4B, EF-hand calcium binding domain 4B | rs887304 | Lobular inflammation |
The PNPLA3 (also known as adiponutrin) gene encodes a transmembrane polypeptide chain exhibiting triglyceride hydrolase activity[16], which is highly expressed on the endoplasmic reticulum and lipid membranes of hepatocytes and adipose tissue[17]. PNPLA3 activity is regulated by glucose and insulin[18], mainly via pathways involving the sterol regulatory element binding protein-1c, as demonstrated both in animal models and human hepatocytes[19]. The I148M variant - a SNP with a risk allele frequency of 21%-28% in European populations - impairs the phospholipase activity of the enzyme, thus reducing lipid catabolism, although it might also gain new functions[17] with a resulting increase in the synthesis of phosphatidic acid[20]. In addition, the PNPLA3 variant has been associated with a loss of retinyl-palmitate lipase activity in stellate cells[21]. Taken together, these data support a link between the PNPLA3 variant and the above reported wide spectrum of liver damage. As previously mentioned, the first report on the PNPLA3 I148M variant in NAFLD came from the GWAS by Romeo et al[15]. These authors identified the relationship between this SNP and liver fat content, and this association remained significant after adjusting for metabolic factors, ethanol use, and ancestry. Of great relevance, the link between PNPLA3 I148M variant and NAFLD is not confounded by the presence of metabolic syndrome (MS) and its features; indeed, even if some authors reported an interplay between insulin resistance (IR) and the variant[22,23], most studies did not find such association, as confirmed by a recent meta-analysis[24]. Interestingly, this independent association between the PNPLA3 I148M variant and NAFLD could be more relevant in women than in men, as highlighted by Speliotes et al[25] in a gender specific analysis performed on a histological NASH cohort. Beyond these gender differences, however, the PNPLA3 I148M variant could explain, at least in part, the variations in NAFLD prevalence across different multiple ethnicities. Indeed, the original report by Romeo et al[15] already found that the frequencies of the 148M allele matched the prevalence of NAFLD in the Dallas Heart Study[11], such that Hispanics had the highest frequency of the 148M allele (49%), followed by European Americans (23%) and African Americans (17%). These ethnic differences were subsequently confirmed by other investigators[26]. Over the last few years, several studies not only have further emphasized how the PNPLA3 I148M variant is associated robustly with liver fat content[27,28] but also revealed the link between the variant and the severity of liver injury, in terms of portal and lobular inflammation and Mallory-Denk bodies[29], presence of NASH, and severity of histological liver fibrosis[25,30] or liver stiffness measurement values[31]. This interplay between the PNPLA3 I148M variant and advanced fibrosis in patients with NASH has been further confirmed by a recent meta-analysis[32]. It is noteworthy that the role of PNPLA3 in NAFLD susceptibility and progression has been reported also in pediatric patients. In this line, the 148M allele was associated with higher liver fat content in Hispanic[33] and obese Taiwanese children[34], and with histological hallmarks of severity of liver injury - steatosis, hepatocellular ballooning and lobular inflammation, and presence of NASH and fibrosis - in Caucasian children and adolescents[35]. Interestingly, the PNPLA3 genotype seems to influence steatosis development also in chronic hepatitis C (CHC) patients, and it has been independently associated with the progression of CHC, including fibrosis, cirrhosis, and HCC occurrence[36,37]. Furthermore, it has been associated with susceptibility to steatosis in patients with chronic hepatitis B[38] and with cirrhosis and HCC development in patients with alcohol abuse[39,40]. Recently, the association between the PNPLA3 variant I148M and the risk of HCC development has been robustly validated in patients with NAFLD[41,42], and it has been estimated that the homozygous carriers of the p.148M mutation carry a 12-fold increased HCC risk as compared to p.I148 homozygotes[43]. Considering all the aforementioned effects of PNPLA3 genotype on not only NAFLD, but also on alcoholic liver disease and CHC, some authors have proposed defining a novel clinical entity based on the presence of PNPLA3 risk allele - PNPLA3-associated steatohepatitis (“PASH”) - i.e., patients with fatty liver disease in whom PNPLA3 appears to be a major driver of disease progression in combination with ethanol consumption and Western diet[44]. Furthermore, PNPLA3 genotype has been evaluated as a possible modifier of NAFLD-associated systemic alterations. Our group recently examined the presence of carotid atherosclerosis in a Sicilian NAFLD cohort and its relation with several SNPs, including PNPLA3[45]. We found that the prevalence of carotid plaques and intima media thickness thickening was significantly higher in PNPLA3 GG compared to CC/CG genotype, particularly among patients under 50 years. This finding was also confirmed in a validation cohort from Northern Italy, where PNPLA3 GG genotype was independently associated with intima media thickness progression. Recently, Musso et al[46] associated the PNPLA I148M variant with the presence of chronic kidney disease, a well-known marker of a higher cardiovascular risk in NAFLD. Finally, a recent study by Sevastianova et al[23] evaluated whether weight loss was able to decrease liver fat in homozygous carriers of the G allele of PNPLA3; investigators found that liver fat content decreased significantly more in the 148MM group than in the 148II after a short course of low carbohydrate diet, although 148II and 148MM patients lost similar amounts of body weight. Overall, although the major role of PNPLA3 in susceptibility and progression of fatty liver has been widely elucidated, further research is needed to fully understand the role of PNPLA3 genotype on systemic alterations and treatment outcomes in patients with NAFLD/NASH.
One of the most recently described and intriguing genetic factors in NAFLD scenario is the nonsynonymous variant rs58542926 (c.449 C>T) within a gene of mostly unknown functions called TM6SF2 at the 19p13.11 locus, which encodes an E167K amino acid substitution. This variant is in strong linkage disequilibrium with other variants around the 19p13.11 locus that were previously reported by another GWAS (see further) to be risk factors for NAFLD[47], suggesting that the new and old signals could be the same, even if conditional analyses indicate that TM6SF2 rs58542926 may be the real causal variant underlying the association at this locus. The first evidence on this new SNP originated from three independent groups. Kozlitina et al[48] performed an exome-wide association study in a multiethnic, population-based cohort derived from the Dallas Heart Study, identifying the association between hepatic triglycerides content - evaluated by proton magnetic resonance spectroscopy - and the TM6SF2 variant rs58542926. In addition, the investigators highlighted the association between the TM6SF2 variant with higher serum alanine aminotransferase (ALT) and aspartate aminotransferase (AST) levels - as surrogate for NASH - and with reduced plasma levels of triglycerides and low-density lipoprotein (LDL)-cholesterol. Finally, they performed a functional analysis for the TM6SF2 in mouse models by silencing the gene via adeno-associated viral vectors. Silencing of the gene showed a 3-fold increase in hepatic triglycerides levels and a decrease in plasma levels of triglycerides, LDL- and high-density lipoprotein (HDL)- cholesterols and very low density lipoprotein (VLDL). Overall, their results demonstrated that the TM6SF2 gene regulated hepatic triglyceride secretion and that the functional impairment of TM6SF2 promoted NAFLD. The second study conducted by Mahdessian et al[49] reported a positive correlation between hepatic TM6SF2 mRNA and plasma triglycerides levels and identified the subcellular localization and function of TM6SF2. Indeed, TM6SF2 was mainly localized in the endoplasmic reticulum and endoplasmic reticulum-Golgi intermediate compartment in human hepatoma cells. The TM6SF2 silencing in hepatoma cell lines reduced the expression of genes involved in the synthesis of triglycerides and the secretion of triglycerides-rich lipoprotein, demonstrating that TM6SF2 not only regulated hepatic lipoprotein secretion but also the hepatic synthesis of triglycerides. The third study reported by Liu et al[50] analyzed the relationship between the TM6SF2 rs58542926 SNP and the severity of liver disease in patients with biopsy-proven NAFLD. The authors found that the TM6SF2 rs58542926 SNP was associated with necroinflammation, ballooning, and advanced liver fibrosis. Taken together, these three studies provided evidence that the TM6SF2 variant was associated with the development of NAFLD/NASH via the deregulation of hepatic lipid metabolism. However, not all the authors reported unequivocal findings. Two studies, one from China[51] and one from South America[52], have been unable to replicate the relationship between TM6SF2 and NAFLD. This may be due ethnic differences in the frequency of carriage of the SNP and to the analysis of underpowered cohorts. Conversely, another study from China[53] confirmed, once again, the association between the TM6SF2 167K allele and NAFLD after adjusting for age, sex, body mass index, and presence of T2D. Thus, the reasons for such discrepancies have not yet been elucidated fully. The most interesting aspect about this variant, however, lies in its key role for the elucidation of the mechanistic basis of progressive NAFLD and for the development of a novel point of view on the association between NAFLD and cardiovascular disease. Consistent with this line, Dongiovanni et al[54] found that 188 (13%) out of 1201 subjects who underwent liver biopsy for suspected NASH were carriers of the E167K variant and that they had lower serum lipid levels than noncarriers, more severe steatosis, necroinflammation, ballooning, and fibrosis and were more likely to have NASH and advanced fibrosis after adjusting for metabolic factors and the I148M PNPLA3 risk variant. In addition, E167K carriers had lower risk of developing carotid plaque; in Swedish obese subjects assessed for cardiovascular outcomes, E167K carriers had higher ALT and lower lipid levels but also a lower incidence of cardiovascular events. Consequently, carriers of the TM6SF2 E167K variant seem to be more at risk for progressive NASH, but at the same time they could be protected against cardiovascular diseases. Furthermore, Musso et al[46] found that the TM6SF2 T allele was associated with higher eGFR and with a lower prevalence of albuminuria and chronic kidney disease - another known marker of an increased risk for cardiovascular disease in NAFLD. In other words, TM6SF2 may act as a switch gene able to disconnect the risk of NAFLD/NASH progression from cardiovascular risk.
In 2011, Speliotes et al[47] aimed to discover additional genetic variants influencing NAFLD susceptibility using a genome wide analysis of hepatic steatosis assessed by computed tomography (CT) in large population based samples. First, authors confirmed the prominent role of rs738409 of PNPLA3 as the main genetic risk factor for NAFLD. In addition, they identified four other SNPs. These were localized in or near the genes neurocan (NCAN - rs2228603), protein phosphatase 1, regulatory (inhibitor) subunit 3B (PPP1R3B - rs4240624), glucokinase regulator (GCKR - rs780094), and lysophospholipase-like 1 (LYPLAL1 - rs12137855). NCAN, GCKR, and LYPLAL1, together with PNPLA3, were associated with both increasing CT hepatic steatosis and histological NAFLD, whereas PPP1R3B was associated with CT-assessed steatosis but not histological NAFLD. NCAN is involved in mechanisms of cell adhesion and in lipoprotein metabolism, and its locus was subsequently casually related to the TM6SF2 minor allele (see above). LYPLAL1 likely exerts a complementary function to the PNPLA3 protein in trigliceride catabolism. The protein product of GCKR has been proposed to interfere with glucose and lipid homeostasis via the interaction with hepatic glucokinase and the consequent increased activity of the enzyme[55], ultimately raising the hepatic glycolytic flux, de novo lipogenesis, and triglyceride levels[56]. Several genetic association studies have confirmed the connection between GCKR rs780094 and NAFLD[57-60], including progression of the disease and fibrosis[61]. These findings were further confirmed by a recent meta-analysis[62] that demonstrated a similar effect size of such association in both Asian and non-Asian populations.
Finally, Chalasani et al[63] reported another GWAS in 2010, identifying other variants conferring susceptibility to occurrence of NAFLD and disease progression. On a cohort of patients with biopsy-proven NAFLD, investigators demonstrated an association between severity of histological NAFLD activity score and SNP rs2645424 in the gene encoding farnesyl diphosphate farnesyl transferase 1 - an enzyme involved in cholesterol biosynthesis. Strangely, they did not identify PNPLA3 as a risk factor. However, other associations were reported, including SNP rs343062 on chromosome 7 (near platelet-derived growth factor alpha gene) with the degree of fibrosis; SNP rs1227756 on chromosome 10 in the collagen type XIII alpha1 (COL13A1) gene, rs6591182 on chromosome 11 (near latent transforming growth factor-beta-protein 3 gene), and rs887304 on chromosome 12 in EF-hand calcium binding domain 4B (EFCAB4B) gene with lobular inflammation; and SNP rs2499604 on chromosome 1, rs6487679 on chromosome 12, rs1421201 on chromosome 18, and rs2710833 on chromosome 4 with serum levels of ALT. However, all of them require extensive validation in larger cohorts.
Several genes have been identified as potential candidates in the pathogenesis and progression of fatty liver. In order to give a schematic overview, we roughly divided all candidate genes into two categories: genes influencing glucidic or lipid metabolism - directly or indirectly involved in fatty liver development (Table 2) - and genes involved in mechanisms of liver injury (Table 3).
| Gene | Functions of encoded protein | SNP |
| ENPP1, ectonucleotide pyrophosphatase/phosphodiesterase1 or PC-1 | Interaction with the insulin receptor with consequent reduction of insulin receptor activity | rs1044498 |
| IRS-1, insulin receptor substrate 1 | Part of the machinery involved in insulin pathway as transductor of insulin receptor signaling | rs1801278 |
| ADIPOQ, adiponectin | Relevant adipocytokine associated with insulin resistance, type 2 diabetes, and NAFLD pathogenesis | rs2241766 rs1501299 |
| LEPR, leptin receptor | Receptor of leptin, a hormone synthesized by adipocytes that regulates food intake, insulin action, thermogenesis, and immune system | rs62589000 |
| rs6700986 | ||
| rs1137100 | ||
| rs1137101 | ||
| rs8179183 | ||
| RETN, Resistin | Adipocytokine involved in lipid metabolism, hepatic insulin resistance, inflammatory cascade reactions, and fibrogenesis | rs3745367 |
| PEMT, phosphatidylethanolamine N-methyltransferase | Enzyme involved in the de novo synthesis of phosphatidylcholine in the liver, a biochemical pathway essential for VLDL formation | rs7946 |
| FATP5, Fatty Acid Transport Protein 5 | Transporter involved in the hepatic uptake of fatty acids | rs56225452 |
| ADRB2 and ADRB3, β-adrenergic receptor 2 and 3 | β-adrenergic receptors, with several functions including regulation of basal metabolism and induction of lipolysis | rs4994 |
| rs1042714 | ||
| rs2053044 | ||
| rs11168070 | ||
| rs11959427 | ||
| rs1042711 | ||
| PPARα, peroxisome proliferative activated receptor α | Transcription factor whose activation improves steatosis, inflammation, and fibrosis in pre-clinical models of NAFLD | rs1800206 |
| PPARGC1A, peroxisome proliferator-activated receptor γ coactivator 1-α | PGC-1α, involved in mitochondrial functions, oxidative stress, gluconeogenesis, and lipogenesis | rs8192678 |
| rs2290602 | ||
| PPARγ, peroxisome proliferative activated receptor γ | Transcription factor whose activation improves IR, restores adipose tissue insulin sensitivity, and decreases fatty free acids flux to the liver | rs1801282 |
| APOE, apolipoprotein E | Mediator of remnant lipoprotein binding to LDL receptors to favor the clearance of chylomicrons and VLDL | N/A |
| APOC3, apolipoprotein C-III | A constituent of plasma VLDL, chylomicrons, and HDL-C that inhibits lipoprotein lipase and triglycerides clearance | rs2854116 |
| rs2854117 | ||
| MTTP, microsomal triglyceride transfer protein | Transfer protein involved in apoB-lipoprotein assembly | rs1800591 |
| rs1800804 | ||
| rs1057613 rs3805335 | ||
| LPIN1, lipin 1 | Phosphatase specifically involved in metabolic pathways between adipose tissue and liver | rs13412852 |
| Gene | Functions of encoded protein | SNP |
| TNF-α, tumor necrosis factor-α | Proinflammatory cytokine also involved in the regulation of insulin resistance, release of free fatty acids, and apoptosis in hepatocytes | rs1800629 |
| rs361525 | ||
| rs1799964 | ||
| rs1800630 | ||
| TRAIL, TNF-related apoptosis -inducing ligand | Protein functioning as a ligand that induces celluar apoptosis | rs6763816 rs4491934 |
| IL-6, interleukin-6 | Proinflammatory cytokine produced by adipocytes, hepatocytes, and immune cells also involved in the modulation of insulin resistance | rs1800795 |
| IL-1β, interleukin-1β | Member of IL-1 family cytokine, mainly produced by adipose tissue | rs16944 |
| TLR4, toll-like receptor 4 | Receptor involved in the interaction with bacterial endotoxins capable to favor hepatic injury and a proinflammatory systemic status | rs4986790 |
| IL28B, interleukin-28B | Cytokine belonging to the type III Interferon family | rs12979860 |
| SOD2, superoxide dismutase 2 | Manganese-dependent mitochondrial enzyme involved in protection from cellular injury induced by superoxide radicals | rs4880 |
| CYP2E1, cytochrome P450 2E1 | Part of the cytochrome P450 complex | rs2031920 |
| UCP3, uncoupling protein 3 | Mitochondrial anion carrier involved in the metabolism of superoxide radicals and in the modulation of lipid homeostasis | rs1800849 rs11235972 |
| UCP2, uncoupling protein 2 | Similar to uncoupling protein 3 | rs695366 |
| MTHFR, methylenetetrahydrofolate reductase | Enzyme involved in the methylation of homocysteine to methionine | rs1801133 |
| rs1801131 | ||
| GCLC, Glutamate-cysteine ligase catalytic subunit | Limiting enzyme in the formation of glutathione, a relevant endogen antioxidant | rs17883901 |
| HFE, hemochromatosis | Crucial protein for the regulation of iron homeostasis via the modulation of the expression of hepcidin | rs1800562 |
| rs1799945 | ||
| TMPRSS6, trans-membrane protease serine 6 | Matriptase-2, which cleaves the membrane-bound hemojuvelin, the co-receptor required for hepcidin expression in the liver | rs855791 |
| KLF6, kruppel-like factor 6 | One of the Kruppel-like factors, a family of transcriptional factors that regulate cellular proliferation, differentiation, and apoptosis | rs3750861 |
| TGF-β1, transforming growth factor β1 | In the liver, a promoter of hepatic fibrosis via the activation of hepatic stellate cells | rs1800471 |
| ATII, angiotensin II | Part of the renin-angiotensin system, also advocated as an inducer of TGF-β1 production and accumulation of extracellular matrix in the liver | rs699 |
| AGTR1, Angiotensin II Type 1 Receptor | Type 1 Receptor of Angiotensin II | rs3772622 rs3772633 rs2276736 rs3772630 rs3772627 |
Ectonucleotide pyrophosphatase/phosphodiesterase1 or plasma cell antigen-1 and insulin receptor substrate 1: Insulin resistance - the hallmark of NAFLD pathophysiology - is strongly related to disease progression. Not by chance, SNPs of genes included in the hepatic insulin signalling pathway have consistently been reported to influence IR and to be potential causes of hepatic injury[64]. Among them, the ectonucleotide pyrophosphatase/phosphodiesterase1 (ENPP1)/plasma cell antigen-1 Lys121Gln SNP enhances the interaction between the ENPP1 membrane glycoprotein and the insulin receptor, resulting in inhibition of insulin receptor activity. This SNP has been associated with an increased risk of T2D[65]. Furthermore, the loss-of-function Gly972Arg SNP of IRS-1 - part of the machinery involved in the insulin signaling pathway - decreases activity of IRS-1, thereby inhibiting insulin receptor autophosphorylation and activity[66] and thus increasing the risk of IR and T2D[67]. Dongiovanni et al[68] analyzed the role of these two SNPs in influencing liver damage in 702 patients with biopsy-proven NAFLD from Italy and the United Kingdom, finding that both were independently associated with a marked reduction of insulin signaling activity and with increased the severity of liver fibrosis. Interestingly, the effect of the ENPP1 and IRS-1 SNPs on the severity of liver fibrosis was independent of ethnic background, as it was observed in patients from both Italy and the United Kingdom, thus emphasizing how hepatic IR has a causal role in the progression of liver damage in NASH.
Adiponectin: Adiponectin is a relevant adipocytokine associated with IR and T2D[69]. Several papers have demonstrated a significant decrease in the serum levels of adiponectin in NASH patients[70] and a reduced expression of its receptor in livers with NASH compared to those with simple steatosis[71]. Furthermore, adiponectin has been associated with liver fibrosis and inflammation[72,73], suggesting that it might be directly or indirectly involved in NASH pathogenesis. Variants in adiponectin (ADIPOQ) - the gene encoding adiponectin - have been investigated in order to find potential associations with NAFLD and its severity. Musso et al[74] showed that the at-risk ADIPOQ SNPs 45TT and 276GT were significantly more prevalent in NAFLD than in the general population and that they were associated with the severity of liver disease and with an atherogenic postprandial lipoprotein profile in NASH, independent of fasting adipokine and lipid levels. Consistent with this line, a Japanese study highlighted how such SNPs were associated with IR and progression of liver fibrosis in NAFLD Japanese patients[75]. However, these findings were not replicated in other cohorts. Although hypoadiponectinemia and IR were observed also in Chinese NAFLD patients, the 45TT and 276GT SNPs were not directly associated with NAFLD, even if they might have indirect effects on systemic metabolism and/or NAFLD pathogenesis by influencing serum ALT, body mass index, IR, and plasma adiponectin concentration[76]. It is possible that ethnic differences could explain the discrepancies among these studies.
Leptin receptor: Leptin is a hormone synthesized by adipocytes that regulates food intake, insulin action, thermogenesis, and the immune system[77]. Several studies[78,79] have demonstrated the association between serum leptin levels and risk of NASH, although results have been sometimes conflicting[80]. Accordingly, the leptin receptor has been investigated due to its potential relevance in the modulation of leptin sensitivity: common variants in the human leptin receptor (LEPR) gene have been related with obesity and lipid metabolism[81], IR and T2D[82], and NAFLD[83-86]. The LEPR G3057A variant has been associated with the risk of NAFLD in Chinese diabetic patients[83], whereas Swellam et al[84] showed that NAFLD occurrence was associated with another SNP in LEPR - rs6700986 - in an Egyptian cohort. Furthermore, Zain et al[85] investigated the relationship between polymorphisms in LEPR and NAFLD across different Asiatic ethnic groups (Malayan, Indian, and Chinese). Two SNPs (LEPR rs1137100 and rs1137101) were associated with susceptibility to NAFLD and NASH; and, intriguingly, analysis of gene-gene interaction showed a potential interplay between the LEPR and PNPLA3 genes. Finally, Lys656Asn SNP of LEPR was associated with metabolic factors - namely IR, obesity parameters, and glucose levels - in patients with NAFLD[86]. Thus, LEPR variants may be involved in the occurrence and progression of NAFLD by influencing insulin sensitivity and/or lipid metabolism, even if further evidence should be provided to reinforce such observations.
Resistin: Resistin (RETN) is an adipokine with relevant metabolic actions and a potential role in NAFLD pathogenesis. Indeed, murine models showed that RETN is able to modulate lipid metabolism and hepatic IR[87,88] and may also participate in inflammatory cascade reactions known to be involved in NASH development[89] and in processes of fibrogenesis[90]. Many SNPs of RETN gene have been investigated as potential risk factors for MS and its components[91]. A Chinese study[92] investigated the role of the RETN intronic +299G/A SNP in a NAFLD setting and found that patients with both T2D and NAFLD had the highest plasma RETN levels compared with diabetic patients without evidence of NAFLD and with controls. Furthermore, the AA genotype at the +299 site of the RTEN gene was found to be an independent risk factor for the development of NAFLD in T2D patients at multivariate analysis. However, further studies are needed to confirm this simple association.
Phosphatidylethanolamine N-methyltransferase: Phosphatidylethanolamine N-methyltransferase (PEMT) is a relevant enzyme involved in the de novo synthesis of phosphatidylcholine in the liver[93], a biochemical pathway essential for VLDL formation. Thus, PEMP is involved in the flux of lipid between the liver and plasma, where lack of phosphatidylcholine caused severe steatosis in mice models[94]. A higher frequency of a nonsynonymous sequence variation (V175M) in the PEMT gene, which results in a loss-of-function in the encoded protein, was reported in patients with biopsy-proven NAFLD compared with subjects with normal hepatic triglyceride content assessed by magnetic resonance or by liver biopsy[95]. Similarly, Dong et al[96] found that the occurrence of the V175M variant allele was significantly more frequent in 107 Japanese patients with biopsy-proven NASH than in 150 healthy controls. Conversely, Jun et al[97] did not find any difference in PEMT genotype frequency between NAFLD patients and controls, and Romeo et al[98] demonstrated a lack of any association between the V175M allele and hepatic triglyceride content - assessed by proton magnetic resonance spectroscopy - in their cohort derived from the Dallas Heart Study, a population-based sample from Dallas, Texas[99]. Overall, the available evidence is not enough to firmly consider PEMT as a relevant genetic factor for NAFLD susceptibility and more studies are needed in this setting.
Fatty acid transport proteins: Fatty acid transport proteins (FATPs) are critically involved in the uptake of fatty acids[100], and two different FATP isoforms are expressed in the liver, namely FATP2 and FATP5[101]. Mice models have emphasized the role of FATP5 in increasing the hepatic uptake and trafficking of fatty acids, so that gain-of-function polymorphisms may result in increased steatosis[102]. Auinger et al[103] investigated the consequences of the rs56225452 FATP5 promoter polymorphism on lipid and glucose metabolism and on features of MS in a cohort derived from the Metabolic Intervention Cohort Kiel - a prospective population-based cohort study of the town of Kiel, in Germany, on natural incidence of the MS[104] - and subjects with histologically proven NAFLD. Triglycerides, ALT, and postprandial insulin levels were higher in subjects with the A allele compared with GG homozygotes in the Metabolic Intervention Cohort Kiel cohort, whereas in NAFLD patients, the A allele was associated with higher ALT only. However, the impact of body mass index on the severity of steatosis differed according to FATP5 promoter SNP, suggesting that this polymorphism may be associated with MS and - probably indirectly - with liver damage in NAFLD. Additional independent studies are needed to fully clarify this interesting, even if still unclear, association.
β-adrenergic receptors:β-adrenergic receptors (ADRB) play an important role in regulating basal metabolism, mostly by stimulating lipid mobilization through lipolysis. Several polymorphisms have been detected in ADRB genes that influence IR, hypertriglyceridemia, and features of MS[105-108]. These polymorphisms were evaluated in NAFLD settings, although with conflicting results. A Japanese study involving 63 patients with biopsy-proven NASH analyzed a W64R codon substitution in ADRB3 gene: the R allele frequency in patients with NASH was significantly higher compared with controls[109]. Other authors examined two nonsynonymous polymorphisms involving the ADRB2 gene (Gln27Glu and Arg16Gly): no significant association with fatty liver was observed for the Arg16Gly allele, whereas the Gln27Glu heterozygotes showed a higher prevalence of fatty liver compared with those without the mutation at univariate analysis, even if this association was not confirmed at multivariate analysis[106]. Loomba et al[110] have published the most relevant study on ADRB2 in 2010. The authors evaluated whether common variants at ADRB2 gene in twins were associated with plasma γGT levels - a well-known significant predictor of the MS as well as NAFLD[111,112]. Interestingly, five SNPs in ADRB2 were associated with levels of γGT, and ADRB2 haplotypes displayed pleiotropic effects on γGT and triglyceride levels, suggesting that adrenergic pathways may act as a link between genetic susceptibility to NAFLD and MS.
Peroxisome proliferative activated receptorα, peroxisome proliferative activated receptorγ, and peroxisome proliferator-activated receptorγcoactivator 1-α: Peroxisome proliferative activated receptor (PPAR) α is a transcription factor belonging, together with PPARγ and PPARβ/δ, to the NR1C nuclear receptor subfamily. PPARα activation improves steatosis, inflammation, and fibrosis in pre-clinical models of NAFLD[113], whereas PPARγ improves IR and has been reported to restore adipose tissue insulin sensitivity and decrease fatty free acids flux to the liver[114]. Regarding PPARα SNPs and NAFLD, a Chinese study evaluated the frequency of the val227ala variant on patients with NAFLD compared with control subjects[115]. As the distribution of PPARα val227ala polymorphism was significantly different between the two groups, the authors hypothesized that the Val227 isoform - the one predominant in NAFLD subjects - has lower activity than the Ala227 isoform, thus resulting in a reduced lipid catabolism and an increased risk for NAFLD. Another PPARα variant examined in a setting of NAFLD is the loss-of-function Leu162Val. Dongiovanni et al[116] did not find any association between this SNP and the risk of NAFLD occurrence and histological severity, although it was independently related to IR. The same study also assessed the Pro12Ala loss-of-function SNP in PPARγ2 gene. Even if this polymorphism had been identified as an important mediator for the development of obesity, IR, and T2D[117], no significant association with NAFLD susceptibility and severity was found. Importantly, this SNP was not even associated with IR in this cohort. Similar conclusions were argued by a recent meta-analysis[118] including 1697 cases and 2427 controls derived from eight studies[116,119-125]. No clear evidence of an association between the Pro12Ala polymorphism and susceptibility to NAFLD emerged. The protein PGC-1a is encoded by the peroxisome proliferator-activated receptor γ coactivator 1-α (PPARGC1A) gene and regulates mitochondrial functions, oxidative stress, gluconeogenesis, and lipogenesis[126]. The Gly482Ser SNP in PPARGC1A gene has been repeatedly associated with T2D, hypertension, and obesity in clinical studies[127-129] and also with an impaired capability of PGC-1α to decrease fat deposition in cultured hepatocytes[130]. In this line, it was also associated with the development of NAFLD in Taiwanese obese children after controlling for body mass index, sex, and PNPLA3 genotype[131]. Yoneda et al[132] examined 15 SNPs in PPARGC1A in the Japanese population; they found that rs2290602 SNP was associated with NASH, with an odds ratio (OR) of 2.73 for the T allele. In addition, AST and ALT values of NAFLD patients with the TT genotype were significantly higher than those of patients with the GT or GG allele. However, this association was not further confirmed; a study among the Chinese Han people did not find any association between rs2290602 SNP in PPARGC1A gene and NAFLD[133].
Apolipoprotein E and apolipoprotein C-III: Apolipoprotein E plays a key role in the metabolism of cholesterol and triglycerides. Indeed, it mediates the binding of the remnant lipoproteins to LDL receptors to favor the clearance of chylomicrons and VLDL from the bloodstream. Two SNPs within the apolipoprotein E (APOE) gene have been identified, resulting in three different alleles (e2, e3, e4) and six APOE genotypes with different binding powers[134]. Some association studies investigated the role of APOE genotypes on NAFLD/NASH susceptibility with conflicting results. The APOE 3/3 genotype was associated with an increased risk of NASH in a cohort of Turkish patients[135], whereas the APOE 3/4 genotype had a protective effect[136]. Conversely, Lee et al[137] showed no significant difference in APOE genotypes distribution among 116 Korean NAFLD patients and 50 controls. However, a protective effect of the e4 allele on fatty liver disease was later shown by Yang et al[138] on a large Korean population. Finally, an Italian hospital-based case-control study including 310 NAFLD cases and 422 controls showed that APOE e4 allele carriers had a 2-fold reduction of NAFLD risk compared with e3 homozygotes[139]. The discrepancies between these studies might be attributable to several factors, including different sample sizes, ethnic variability, possible inclusion of alcohol consumers, and lack of clear adjustments for potential metabolic confounders. Apolipoprotein C-III is a major constituent of plasma VLDL, chylomicrons, and HDL-C, which inhibits lipoprotein lipase and triglyceride clearance[140]. Two SNPs in the promoter region of the APOC3 gene - -482C > T and -455T > C, which are in strong linkage disequilibrium with each other - have been repeatedly associated with MS and coronary artery disease[141]. Based on these findings, several studies investigated the association between SNPs of APOC3 gene and NAFLD occurrence, although with conflicting results. Petersen et al[142] firstly reported that ApoC3 T-455C and C-482T promoter SNPs predispose Indian men to liver fat accumulation by altering lipid metabolism and IR. Similar positive results were obtained in Indian[143] and Southern Han Chinese cohorts[144]. However, this association was not further replicated in other studies conducted on Italian[145], British[145], American[146], Finnish[147], German[148], Belgian[149] and Chinese Han[150] subjects. A recent meta-analysis confirmed the absence of a robust association and, therefore, the lack of a causal pathogenetic role of APOC3 gene polymorphisms in patients with NAFLD[151]. These contrasting findings raise doubts about the methodology and quality of some of these studies, particularly about the methods used to diagnose NAFLD and to adjust for confounders.
Microsomal triglyceride transfer protein: Microsomal transfer tryglicerides protein is a transfer protein involved in apoB-lipoprotein assembly[152]. A large number of common genetic polymorphisms in the microsomal triglyceride transfer protein (MTTP) gene have been identified. The G allele of MTTP - 493 G>T polymorphism has been associated with impaired MTTP transcription, and, thus, with a reduced export of triglycerides from hepatocytes and increased susceptibility to NAFLD[153]. Accordingly, the G allele frequency was significantly higher in Japanese patients with NASH, and the severity of NASH was higher in patients with the G/G genotype than in patients with the G/T genotype[154]. Similarly, the -493 G/G genotype was reported to be associated with more severe liver disease and a more atherogenic lipoprotein profile in an Italian cohort[155]. Furthermore, in diabetic French patients, this SNP was associated with elevated ALT as a surrogate marker for NASH[156]. However, other studies did not confirm these reports. Oliveira et al[157] did not find any association between - 493 G>T polymorphism and NAFLD in a Brazilian cohort. Similarly, Peng et al[158] did not find any significant association between the - 493 G>T polymorphism and the risk for NAFLD in a Chinese Han population, even if other SNPs were found to be associated with NAFLD susceptibility. Specifically, in that study, the rs1800804 T/C was associated with an increased risk of NAFLD, while the rs1057613 A/G and rs3805335 C/T SNPs were associated with a decreased risk. Carulli et al[159] found that the distribution of MTTP polymorphisms was not significantly different between NAFLD patients compared with the control group nor associated with clinical or histological characteristics. Finally, a recent meta-analysis including 11 case-control studies with a total of 636 cases and 918 healthy controls revealed that MTP - 493G > T polymorphism was correlated overall with an increased risk of NAFLD among both Caucasian and non-Caucasian populations[160]. However, it should be noted that some of the studies included in the meta-analysis evaluated also featured superimposed NAFLD in HCV-infected patients.
Lipin 1: Lipin 1 is a phosphatase expressed specifically by adipose tissue and liver. It seems to be critically involved in metabolic pathways linking adipose tissue and liver[161]. Several polymorphisms of lipin 1 (LPIN1) have been associated with occurrence of MS and its components[162]. In particular, the LPIN1 rs13412852 T allele was associated with lower body mass index and insulin levels[163]. An Italian study[164] evaluated the LIPIN1 rs13412852 C>T polymorphism in pediatric patients with NAFLD. Investigators demonstrated that the TT genotype, even if underrepresented in pediatric NAFLD patients, was associated with less severe dysplipidemia and a lower prevalence and severity of NASH even after adjustment for genetic - PNPLA3 genotype - and metabolic confounders.
Tumor necrosis factor-αand tumor necrosis factor-related apoptosis-inducing ligand: Tumor necrosis factor-α is an important proinflammatory cytokine involved in the regulation of IR, release of free fatty acids, and induction of apoptosis in hepatocytes under stimuli driven by oxidative stress[165]. Thus, it is not surprising that serum tumor necrosis factor (TNF)-α levels were found to be higher in patients with NASH compared with healthy controls[70] and that elevated levels have been associated with the occurrence of both NAFLD and NASH[166]. Two polymorphisms of the promoter of TNF-α gene have been linked to an increased susceptibility of NAFLD: TNF2 allele (at position -308) and TNFA allele (at position -238)[167,168], both associated with higher TNF-α serum levels[169,170]. However, consistency of this association is still debated[171,172]. Valenti et al[167] found that the prevalence of the -238, but not of the -308, TNF-α polymorphism was higher in Italian patients with NAFLD than in controls and that patients with NAFLD positive for both TNF-α polymorphisms had higher IR but a lower number of associated risk factors for steatosis. Furthermore, Tokushige et al[171] determined the prevalence of six TNF-α promoter region polymorphisms in a group of Japanese patients with NAFLD and in control subjects. Surprisingly, there were no significant differences in the allele frequencies of any of the six polymorphisms between patients and controls. However, they found that two polymorphisms - -1031C and -863A - were significantly higher in the NASH group compared with subjects with simple steatosis only and that they were associated with an increased homeostasis model assessment for IR (HOMA-IR) score. Finally, negative results were also derived from a prospective cohort of Chinese patients with NAFLD, since TNF-α gene polymorphisms were not shown to be associated with NAFLD nor with significant fibrosis[172]. A recent meta-analysis[173] comprising several studies on this topic[167,168,171,172,174-177] concluded that there was a significant difference in TNF-α -238 genotype distribution between NAFLD and control, while there was no clear association between TNF-α -308 genotype and susceptibility for NAFLD. Overall, it is still unclear whether TNF-α polymorphisms are critically involved in NAFLD and/or NASH pathogenesis, probably due to ethnic differences and incomplete control for confounding metabolic factors in most of the studies. Finally, another member of the TNF family, TNF-related apoptosis-inducing ligand (TRAIL), should be mentioned. A Chinese study[178] found that soluble TRAIL levels were significantly higher in NAFLD subjects than in controls and positively correlated with triglyceride concentrations in NAFLD patients and that the AA/TT genotypes of TRAIL at position 1525/1595 conferred a lower risk of NAFLD occurrence and a less severe form of steatosis in NAFLD patients.
Interleukin (IL)-6 and IL-1β: IL-6 is a proinflammatory cytokine produced by adipocytes, hepatocytes, and immune cells, involved in both inflammation and IR[179]. Experimental models have investigated its role in NAFLD pathogenesis and progression, although the results were often contradictory[180-182], whereas certain polymorphisms of the IL-6 gene were associated with NAFLD susceptibility. A small Italian study[159] found that the IL-6 -174C variant C - an allele associated with IR, T2D, and MS in some cohorts[183,184] but not in others[185] - was more prevalent in NAFLD than in healthy subjects, associated with increased insulin levels and HOMA-IR, and an independent predictor of NAFLD and NASH. Intriguingly, this finding is in contrast with other studies reporting that it was the IL-6 -174 G variant that was associated with metabolic abnormalities[186,187]. IL-1 family cytokine members are produced mainly by human adipose tissue; certain IL-1 cytokines - such as IL-1α, IL-1β, IL-18 - have proinflammatory properties, while others - IL-1 receptor antagonist, for example - are anti-inflammatory[188]. Interestingly, IL-1α and IL-1β were shown to have a role in the transition from steatosis to steatohepatitis and liver fibrosis[189]. Based on these findings, Interleukin-1β-511 T/C polymorphism, a functional variant that affects DNA-protein interactions in vitro[190], was determined in 63 Japanese NASH patients and 100 healthy volunteers[109]. The authors found that Interleukin-1β -511 T allele frequency and the T/T genotype frequency were significantly higher in NASH patients than in control subjects.
Toll-like receptor 4: Bacterial overgrowth and endotoxemia have recently emerged as two relevant factors in the pathogenesis of NASH[191]. Indeed, the interplay between toll-like receptor 4 (TLR4) and endotoxins results in the release of several mediators capable of favoring hepatic injury and a proinflammatory systemic status[192]. Variants encoded in the ectodomain of the TLR4 gene, D299G and T399I, have been linked with endotoxin hyporesponsiveness[193] and with possible effects on inflammatory and metabolic disorders like atherosclerosis, IR, MS, and T2D[194,195]. Animal models showed a potential direct link between TLR-4 and Kupffer cells in the pathogenesis of steatohepatitis[192], and, notably, Guo et al[196] demonstrated that the D299G and T399I variants were associated with protection from hepatic fibrosis by reducing TLR4-mediated inflammatory and fibrogenic signalling and lowering the apoptotic threshold of activated hepatic stellate cells. Regarding the interaction between NAFLD and TLR-4 polymorphisms in humans, a recent case-control study[197] revealed that the frequency of the heterozygous mutation at position -299 was significantly lower in patients with NAFLD than in controls. However, further studies are needed to clarify the protective role of such polymorphisms in NAFLD pathogenesis and progression.
IL-28B: Several studies repeatedly showed that genetic variations around the IL-28B gene strongly predict the spontaneous and treatment-induced clearance of hepatitis C viral infection[198,199]. In particular, IL-28B rs12979860 CC and IL-28B rs8099917 TT genotypes were shown to be closely related to the achievement of a sustained virological response following antiviral therapy[200-202]. Furthermore, other studies revealed a link between IL-28B polymorphisms and the severity of CHC in terms of steatosis[203,204], necroinflammatory activity[205], and fibrosis[206-208]. Our group reported on a cohort of 160 patients with histological diagnosis of NAFLD that IL-28B rs12979860 CC genotype was associated with the histological severity of liver disease, independently of HOMA and hyperuricemia - well-known risk factors for liver damage in NAFLD[209]. Interestingly, the at-risk CC rs12979860 variant was associated with severe necroinflammation, particularly in subjects with the PNPLA3 G allele, thus leading to hypothesize a potential interplay between these two genes. Such findings were recently confirmed by Eslam et al[210] on a large cohort, including 3129 patients with CHC, 555 with chronic hepatitis B, and 488 with NAFLD. The authors demonstrated that rs12979860 genotype acted as a strong predictor of tissue inflammation and fibrosis among all these chronic liver diseases, independent of the underlying etiology. However, Garrett et al[211] did not confirm these findings on their North American Caucasian patients with NAFLD, even if they enrolled a cohort of severe obese NAFLD patients evaluated for bariatric surgery, and, therefore, very different from our cohort. Overall, these data suggest an effect of IL-28B CC genotype in patients at lower metabolic risk only, and not in obese patients, where the burden of metabolic alterations on NAFLD severity likely overcomes the role of the genetic background.
Superoxide dismutase 2 and cytochrome P450 2E1: The superoxide dismutase 2 (SOD2) gene encodes for the mitochondrial enzyme manganese-dependent superoxide dismutase, a protein that protects cells from injury induced by superoxide radicals[212]. Interestingly, oxidative stress is regarded as a relevant factor involved into the transition from simple steatosis to steatohepatitis[213]. A common polymorphism in the SOD2 gene - C47T, rs4880 - has been related to relatively efficient protein function by in vitro studies[214,215], and SOD2 variants have been investigated in settings of alcoholic liver disease with inconsistent results[216,217]. Regarding the role of SOD2 C47T polymorphism in NAFLD, a small study performed on 63 Japanese subjects revealed an increased prevalence of the lower activity homozygous T genotype among patients with NASH compared with controls[154]. Similar conclusions were drawn from a cohort of obese Egyptian children with steatosis or NASH[218]. Al-Serri et al[219] performed a two-step analysis of the relevance of this SNP in NAFLD: the preferential transmission of alleles from parents to affected children in 71 family trios and a classical case-control study involving a cohort of 502 European patients with fatty liver. Investigators demonstrated that SOD2 genotype - together with PNPLA3 genotype, T2D, and histological severity of NASH - was associated with an advanced stage of fibrosis. Conversely, a Chinese study did not find any significant difference in the frequencies of the three SOD2 genotypes among patients and controls but highlighted how the frequency of the SOD2 C variant was higher in the NASH group than in subjects with simple steatosis and in controls[220]. The same study evaluated another gene potentially involved in NAFLD pathogenesis: cytochrome P450 2E1 (CYP2E1), encoding for cytochrome P450 2E1 - another enzyme related to superoxide radicals in humans. Indeed, induction of CYP2E1 is a central process involved in generating oxidative stress in both alcoholic and nonalcoholic steatohepatitis[221]. However, evidence about a potential role of CYP2E1 gene SNPs in NAFLD pathogenesis are elusive. On the one hand, the above mentioned study[220] did not report any association between the CYP2E1 -1053C>T variation (*1/*5 - rs2031920) and increased susceptibility to NAFLD or NASH in Chinese subjects; on the other hand, Varela et al[222] found that the CYP2E1 *5 variant was positively associated with liver injury in obese women with NASH, and similar positive results were also found on a Chinese population[223]. It is likely that ethnic differences and the incomplete understanding of the real effect of SOD2 and CYP2E1 genotypes on related enzymatic activities could be the main reasons underlying these conflicting results.
Uncoupling protein 3 and uncoupling protein 2: Uncoupling protein 3 is a mitochondrial anion carrier selectively expressed in skeletal muscle - the major site of thermogenesis in humans - involved in the metabolism of superoxide radicals and in the modulation of energy and lipid homeostasis[224-226]. The rs1800849 -55C/T polymorphism of uncoupling protein (UCP) 3 has been associated with an increased susceptibility to T2D and obesity and with an atherogenic lipid profile[227-229]. Interestingly, the rs1800849 UCP3 -55CT genotype was also associated with IR, increased adiponectin levels, the presence of moderate-severe steatosis, and NASH in a small Spanish study[230]. Furthermore, an interesting Chinese paper aiming to evaluate the frequency of four nonsynonymous SNPs in the UCP3 gene in a pediatric cohort found a higher prevalence of another variant - rs11235972 GG genotype - among patients with NAFLD compared with control subjects[231]. Similar to UCP3, UCP2 is involved in the regulation of mitochondrial lipid efflux and oxidative metabolism. Its increased hepatic expression has been reported both in experimental models and in NASH patients as a protective mechanism against liver injury progression[232]. A promoter region polymorphism of UCP2 - -866 G>A variant - is able to influence the extrahepatic expression of UCP2 and insulin release and sensitivity, although the overall metabolic impact is still controversial[233]. A recent Italian paper investigated the role of this SNP in patients who underwent liver biopsy for suspected NASH[234]. UCP2 -866 A/A genotype was associated with a reduced risk of NASH after adjustment for age, sex, body mass index, impaired fasting glucose or diabetes, and PNPLA3 I148M allele and with a reduced risk of steatosis grade G2-G3 and NASH in patients without, but not in those with, impaired fasting glucose/diabetes. Concerning the metabolic traits, the UCP2 A/A genotype was associated with higher total serum cholesterol levels but not with serum HDL, triglycerides or impaired fasting glucose/diabetes. Overall, SNPs in UCP genes may confer susceptibility or protection to NAFLD/NASH, even if further evidence needs to be provided.
Methylenetetrahydrofolate reductase: Homocysteine is an intermediate amino acid formed during methionine metabolism in the liver. Today, hyperhomocysteinemia is regarded as a risk factor for liver diseases via the promotion of oxidative and endoplasmic reticulum stress, and the activation of proinflammatory factors[235,236]. Methylenetetrahydrofolate reductase (MTHFR) catalyzes the reduction of 5,10-methylenetetrahydrofolate to 5-methyltetrahydrofolate, a metabolic pathway fundamental for the methylation of homocysteine to methionine. Several genetic polymorphisms in the MTHFR gene have been identified, and among them, the C677T polymorphism (rs1801133) and the A1298C (rs1801131) - inducing both a reduction of MTHFR activity - were extensively investigated[237,238] in the setting of NAFLD[239-242]. Sazci et al[239] analyzed the frequency of C677T and A1298C polymorphisms of MTHFR gene in a Turkish cohort comprising 57 NASH patients and 324 healthy controls, showing that the MTHFR 1298C allele in all NASH patients, the C677C/C1298C compound genotype in women, and the C677C/A1298C compound genotype in men were genetic risk factors for NASH. Similarly, Catalano et al[240] recently identified the MTHFR A1298C heterozygous polymorphisms as a weak predictor for NAFLD severity in an Italian cohort. However, the relationship between MTHFR polymorphisms and NAFLD remains controversial. Franco Brochado et al[241] did not find any association between the MTHFR C677T and A1298C polymorphisms and NAFLD and its severity. Similarly, Serin et al[242] showed that the MTHFR C677T polymorphism was not a risk factor for NAFLD in their Turkish cohort. As a consequence, more rigorous work needs to be performed in this field.
Glutamate-cysteine ligase catalytic subunit: The glutamate-cysteine ligase catalytic subunit (GCLC) gene codes the catalytic subunit of the heterodimeric γ-Glutamate-cysteine ligase, the limiting enzyme in the formation of glutathione, a relevant endogen antioxidant. The base T in the position -129, as opposed to base C, determines a sharp decrease in the promoter activity of the GCLC gene and was identified as a significant independent risk factor for myocardial infarction in a Japanese population[243]. In addition, mitochondrial glutathione depletion has been associated with the development of alcoholic steatohepatitis due to the increased sensitivity of hepatocytes to the pro-oxidant effects of cytokines generated by ethanol metabolism[244]. Interestingly, Oliveira et al[157] found that, among 131 biopsy-proven NAFLD patients, the presence of at least one T allele in the -129 C/T polymorphism of the GCLC gene was independently associated with NASH detection, with an OR of 12.14. Thus, such polymorphism could be an important factor in the development of liver injury mediated by oxidative stress.
Hemochromatosis and trans-membrane protease serine 6: Human hemochromatosis protein (HFE) is crucial for the regulation of iron homeostasis via modulation of the expression of hepcidin[245]. Excessive hepatic iron deposition is a frequent histological feature of NASH, and it has been investigated as a potential contributor to oxidative stress in the liver, and thus as a second hit promoter[246]. In this regard, even if the C282Y and H63D mutations of the HFE gene - common in Caucasians and responsible for most cases of hereditary hemochromatosis - are well-known causes of potential iron overload, their prevalence and relevance in patients with NAFLD have been variable, depending on the examined cohorts. The first reports about the association between HFE mutations and NAFLD came in the late 1990s and showed a positive correlation between these two conditions[247,248]. Later, Lee et al[249] identified the presence of H63D mutation as an independent factor associated with NAFLD in the Korean population, and Nelson et al[250] suggested that the presence of the C282Y mutation was a risk factor for the development of advanced hepatic fibrosis among American Caucasian patients with NASH. Nonetheless, other studies have not confirmed such associations. Indeed, even if several reports suggested that increased ferritin levels may be markers of histological damage, the HFE mutations did not consistently contribute to hepatic fibrosis in NAFLD[251] nor to its susceptibility[252]. The poor relevance of HFE mutations in NAFLD have been resumed by a recent meta-analysis including 610 cases and 7298 controls[253]: authors found no associations between iron-overloading HFE mutations and NAFLD susceptibility or severity. However, other genetic variants influencing iron deposition may be involved in NAFLD/NASH pathogenesis. Beta-globin mutations have been identified as a good genetic predictor of parenchymal iron overload in Italian patients with NAFLD and have been associated with a two-fold higher risk of severe fibrosis[254]. More recently, the rs855791 C>T polymorphism of the trans-membrane protease serine 6 (TMPRSS6) gene - encoding for matriptase-2, which cleaves the membrane-bound hemojuvelin, a co-receptor required for hepcidin expression in the liver[255] - has been associated with lower hepatic iron stores, ferritin levels, and ballooning in 216 patients with histological NAFLD[256].
Kruppel-like factor 6: The kruppel-like factor 6 (KLFs) are a family of zinc finger-containing transcriptional factors that regulate cellular processes, such as proliferation, differentiation, and apoptosis[257]. In the liver, injury and/or cytokines are able to induce KLF6 gene expression, which in turn plays an essential role in the transactivation of several genes involved in the development of liver fibrosis, mainly via the activation of hepatic stellate cells[258]. Miele et al[259] reported the association between a functional polymorphism in the KLF6 gene - IVS1-27G>A SNP (rs3750861) - and the severity of NAFLD. In particular, they demonstrated increased levels of total and wild type KLF6 expression in patients with NAFLD and higher steatosis, inflammation, and fibrosis, whereas KLF6 IVS1-27G>A SNP was associated with reduced fibrosis, and thus, acted as a protective factor against NASH progression. Intriguingly, the effects of KLF6 genotype on NAFLD/NASH pathogenesis may also involve the modulation of metabolic pathways: Bechmann et al[260] observed that KLF6 IVS1-27G wild-type allele was associated with increased fasting glucose and insulin levels and with decreased hepatic insulin sensitivity via the reduced expression of glucokinase. KLF6 increased PPARα activity, whereas KLF6 loss led to PPARα repression and attenuation of lipid and glucose abnormalities[261].
Transforming growth factor-β1, angiotensin II, and angiotensin II type 1 receptor: The transforming growth factor (TGF)-β1 is a well-known promoter of hepatic fibrosis that contributes to the activation of hepatic stellate cells[262]. TGF-β1 production can be stimulated by angiotensin II (ATII), part of the renin-angiotensin system that has been advocated as a potential inducer of extracellular matrix accumulation[263]. A higher frequency of a pro-fibrotic TGF-β1 SNP (Arg/Arg at codon 25) has been identified in patients with hypertension compared with controls[264]. Furthermore, this TGF-β1 SNP and an ATII variant in the promoter region of the gene (AT-6 G>A), leading to a higher transcription of AT, were both associated with increased hepatic fibrosis in patients with CHC[265]. Based on these findings, Dixon et al[266] investigated these two polymorphisms in a group of severely obese patients with NASH. The investigators found a positive association between AT-6 A/A polymorphism and severe fibrosis, even if such correlation was lost after correction for gender. However, patients with both high ATII and TGF-β1 producing polymorphisms had a higher risk of advanced fibrosis. In addition, animal models had demonstrated that the Angiotensin II Type 1 Receptor (AGTR1) gene could be implicated in the susceptibility to NAFLD[267]. In this line, none of the five variants of the AGTR1 gene were associated with susceptibility to NAFLD in a multi-ethnic Asiatic cohort composed of Malayan, Indian, and Chinese subjects, with the exception of the Indian subgroup, where the rs2276736, rs3772630, and rs3772627 were found to be protective against NAFLD and NASH[268]. Furthermore, five SNPs of AGTR1 gene (rs3772622, rs3772633, rs2276736, rs3772630, and rs3772627) were significantly associated with NAFLD in a Japanese cohort[269]. All in all, the potential involvement of the renin-angiotensin system in NAFLD/NASH pathogenesis is still unclear, and further research is needed.
In the complex pathogenetic puzzle of NAFLD, genes clearly act as major disease modifiers affecting NAFLD occurrence and severity and sometimes cardiovascular risk as well. To date, the PNPLA3 gene variant is the most validated susceptibility factor for steatosis, NASH, fibrosis, and HCC, despite a number of other genetic variants contributing to liver damage. However, even if the identification of these variants helped us to understand better NAFLD in terms of both clinical phenotypes and pathogenetic mechanisms, their utility in clinical practice and in the individual patients is far from being relevant. Therefore, further efforts should be done to generate a genetic map useful to stratify the hepatic and non-hepatic risk of NAFLD patients and to define better therapeutic approaches in terms of both lifestyle intervention and new pharmacological therapies.
P- Reviewer: de Oliveira CPMS, Gazouli M S- Editor: Ma YJ L- Editor: Filipodia E- Editor: Liu XM
| 1. | Petta S, Muratore C, Craxì A. Non-alcoholic fatty liver disease pathogenesis: the present and the future. Dig Liver Dis. 2009;41:615-625. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 188] [Cited by in RCA: 196] [Article Influence: 12.3] [Reference Citation Analysis (1)] |
| 2. | Loomba R, Sanyal AJ. The global NAFLD epidemic. Nat Rev Gastroenterol Hepatol. 2013;10:686-690. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1127] [Cited by in RCA: 1330] [Article Influence: 110.8] [Reference Citation Analysis (0)] |
| 3. | Clark JM, Brancati FL, Diehl AM. The prevalence and etiology of elevated aminotransferase levels in the United States. Am J Gastroenterol. 2003;98:960-967. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 4. | Vernon G, Baranova A, Younossi ZM. Systematic review: the epidemiology and natural history of non-alcoholic fatty liver disease and non-alcoholic steatohepatitis in adults. Aliment Pharmacol Ther. 2011;34:274-285. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2405] [Cited by in RCA: 2293] [Article Influence: 163.8] [Reference Citation Analysis (0)] |
| 5. | Adams LA, Lymp JF, St Sauver J, Sanderson SO, Lindor KD, Feldstein A, Angulo P. The natural history of nonalcoholic fatty liver disease: a population-based cohort study. Gastroenterology. 2005;129:113-121. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2092] [Cited by in RCA: 2129] [Article Influence: 106.5] [Reference Citation Analysis (0)] |
| 6. | Musso G, Gambino R, Cassader M, Pagano G. Meta-analysis: natural history of non-alcoholic fatty liver disease (NAFLD) and diagnostic accuracy of non-invasive tests for liver disease severity. Ann Med. 2011;43:617-649. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 886] [Cited by in RCA: 919] [Article Influence: 65.6] [Reference Citation Analysis (0)] |
| 7. | Anstee QM, Daly AK, Day CP. Genetics of alcoholic and nonalcoholic fatty liver disease. Semin Liver Dis. 2011;31:128-146. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 90] [Cited by in RCA: 81] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 8. | Makkonen J, Pietiläinen KH, Rissanen A, Kaprio J, Yki-Järvinen H. Genetic factors contribute to variation in serum alanine aminotransferase activity independent of obesity and alcohol: a study in monozygotic and dizygotic twins. J Hepatol. 2009;50:1035-1042. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 105] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 9. | Struben VM, Hespenheide EE, Caldwell SH. Nonalcoholic steatohepatitis and cryptogenic cirrhosis within kindreds. Am J Med. 2000;108:9-13. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 205] [Cited by in RCA: 195] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 10. | Willner IR, Waters B, Patil SR, Reuben A, Morelli J, Riely CA. Ninety patients with nonalcoholic steatohepatitis: insulin resistance, familial tendency, and severity of disease. Am J Gastroenterol. 2001;96:2957-2961. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 276] [Cited by in RCA: 281] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 11. | Browning JD, Szczepaniak LS, Dobbins R, Nuremberg P, Horton JD, Cohen JC, Grundy SM, Hobbs HH. Prevalence of hepatic steatosis in an urban population in the United States: impact of ethnicity. Hepatology. 2004;40:1387-1395. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2633] [Cited by in RCA: 2695] [Article Influence: 128.3] [Reference Citation Analysis (3)] |
| 12. | Petersen KF, Dufour S, Feng J, Befroy D, Dziura J, Dalla Man C, Cobelli C, Shulman GI. Increased prevalence of insulin resistance and nonalcoholic fatty liver disease in Asian-Indian men. Proc Natl Acad Sci USA. 2006;103:18273-18277. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 322] [Cited by in RCA: 298] [Article Influence: 15.7] [Reference Citation Analysis (0)] |
| 13. | Schneider AL, Lazo M, Selvin E, Clark JM. Racial differences in nonalcoholic fatty liver disease in the U.S. population. Obesity (Silver Spring). 2014;22:292-299. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 84] [Cited by in RCA: 107] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 14. | Lewis CM, Knight J. Introduction to genetic association studies. Cold Spring Harb Protoc. 2012;2012:297-306. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 97] [Cited by in RCA: 118] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 15. | Romeo S, Kozlitina J, Xing C, Pertsemlidis A, Cox D, Pennacchio LA, Boerwinkle E, Cohen JC, Hobbs HH. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat Genet. 2008;40:1461-1465. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2701] [Cited by in RCA: 2599] [Article Influence: 152.9] [Reference Citation Analysis (0)] |
| 16. | Pingitore P, Pirazzi C, Mancina RM, Motta BM, Indiveri C, Pujia A, Montalcini T, Hedfalk K, Romeo S. Recombinant PNPLA3 protein shows triglyceride hydrolase activity and its I148M mutation results in loss of function. Biochim Biophys Acta. 2014;1841:574-580. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 154] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 17. | He S, McPhaul C, Li JZ, Garuti R, Kinch L, Grishin NV, Cohen JC, Hobbs HH. A sequence variation (I148M) in PNPLA3 associated with nonalcoholic fatty liver disease disrupts triglyceride hydrolysis. J Biol Chem. 2010;285:6706-6715. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 433] [Cited by in RCA: 495] [Article Influence: 30.9] [Reference Citation Analysis (0)] |
| 18. | Rae-Whitcombe SM, Kennedy D, Voyles M, Thompson MP. Regulation of the promoter region of the human adiponutrin/PNPLA3 gene by glucose and insulin. Biochem Biophys Res Commun. 2010;402:767-772. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 31] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 19. | Dubuquoy C, Robichon C, Lasnier F, Langlois C, Dugail I, Foufelle F, Girard J, Burnol AF, Postic C, Moldes M. Distinct regulation of adiponutrin/PNPLA3 gene expression by the transcription factors ChREBP and SREBP1c in mouse and human hepatocytes. J Hepatol. 2011;55:145-153. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 109] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 20. | Kumari M, Schoiswohl G, Chitraju C, Paar M, Cornaciu I, Rangrez AY, Wongsiriroj N, Nagy HM, Ivanova PT, Scott SA. Adiponutrin functions as a nutritionally regulated lysophosphatidic acid acyltransferase. Cell Metab. 2012;15:691-702. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 215] [Cited by in RCA: 238] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 21. | Pirazzi C, Valenti L, Motta BM, Pingitore P, Hedfalk K, Mancina RM, Burza MA, Indiveri C, Ferro Y, Montalcini T. PNPLA3 has retinyl-palmitate lipase activity in human hepatic stellate cells. Hum Mol Genet. 2014;23:4077-4085. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 224] [Cited by in RCA: 294] [Article Influence: 26.7] [Reference Citation Analysis (0)] |
| 22. | Wang CW, Lin HY, Shin SJ, Yu ML, Lin ZY, Dai CY, Huang JF, Chen SC, Li SS, Chuang WL. The PNPLA3 I148M polymorphism is associated with insulin resistance and nonalcoholic fatty liver disease in a normoglycaemic population. Liver Int. 2011;31:1326-1331. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 65] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 23. | Sevastianova K, Kotronen A, Gastaldelli A, Perttilä J, Hakkarainen A, Lundbom J, Suojanen L, Orho-Melander M, Lundbom N, Ferrannini E. Genetic variation in PNPLA3 (adiponutrin) confers sensitivity to weight loss-induced decrease in liver fat in humans. Am J Clin Nutr. 2011;94:104-111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 105] [Cited by in RCA: 120] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 24. | Sookoian S, Pirola CJ. Meta-analysis of the influence of I148M variant of patatin-like phospholipase domain containing 3 gene (PNPLA3) on the susceptibility and histological severity of nonalcoholic fatty liver disease. Hepatology. 2011;53:1883-1894. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 756] [Cited by in RCA: 740] [Article Influence: 52.9] [Reference Citation Analysis (1)] |
| 25. | Speliotes EK, Butler JL, Palmer CD, Voight BF, Hirschhorn JN. PNPLA3 variants specifically confer increased risk for histologic nonalcoholic fatty liver disease but not metabolic disease. Hepatology. 2010;52:904-912. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 302] [Cited by in RCA: 293] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 26. | Wagenknecht LE, Palmer ND, Bowden DW, Rotter JI, Norris JM, Ziegler J, Chen YD, Haffner S, Scherzinger A, Langefeld CD. Association of PNPLA3 with non-alcoholic fatty liver disease in a minority cohort: the Insulin Resistance Atherosclerosis Family Study. Liver Int. 2011;31:412-416. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 71] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 27. | Sookoian S, Castaño GO, Burgueño AL, Gianotti TF, Rosselli MS, Pirola CJ. A nonsynonymous gene variant in the adiponutrin gene is associated with nonalcoholic fatty liver disease severity. J Lipid Res. 2009;50:2111-2116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 275] [Cited by in RCA: 312] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 28. | Kollerits B, Coassin S, Kiechl S, Hunt SC, Paulweber B, Willeit J, Brandstätter A, Lamina C, Adams TD, Kronenberg F. A common variant in the adiponutrin gene influences liver enzyme values. J Med Genet. 2010;47:116-119. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 53] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 29. | Rotman Y, Koh C, Zmuda JM, Kleiner DE, Liang TJ. The association of genetic variability in patatin-like phospholipase domain-containing protein 3 (PNPLA3) with histological severity of nonalcoholic fatty liver disease. Hepatology. 2010;52:894-903. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 387] [Cited by in RCA: 381] [Article Influence: 25.4] [Reference Citation Analysis (0)] |
| 30. | Valenti L, Al-Serri A, Daly AK, Galmozzi E, Rametta R, Dongiovanni P, Nobili V, Mozzi E, Roviaro G, Vanni E. Homozygosity for the patatin-like phospholipase-3/adiponutrin I148M polymorphism influences liver fibrosis in patients with nonalcoholic fatty liver disease. Hepatology. 2010;51:1209-1217. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 477] [Cited by in RCA: 530] [Article Influence: 35.3] [Reference Citation Analysis (0)] |
| 31. | Krawczyk M, Grünhage F, Zimmer V, Lammert F. Variant adiponutrin (PNPLA3) represents a common fibrosis risk gene: non-invasive elastography-based study in chronic liver disease. J Hepatol. 2011;55:299-306. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 62] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 32. | Singal AG, Manjunath H, Yopp AC, Beg MS, Marrero JA, Gopal P, Waljee AK. The effect of PNPLA3 on fibrosis progression and development of hepatocellular carcinoma: a meta-analysis. Am J Gastroenterol. 2014;109:325-334. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 297] [Cited by in RCA: 278] [Article Influence: 25.3] [Reference Citation Analysis (0)] |
| 33. | Goran MI, Walker R, Le KA, Mahurkar S, Vikman S, Davis JN, Spruijt-Metz D, Weigensberg MJ, Allayee H. Effects of PNPLA3 on liver fat and metabolic profile in Hispanic children and adolescents. Diabetes. 2010;59:3127-3130. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 86] [Cited by in RCA: 98] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 34. | Lin YC, Chang PF, Hu FC, Yang WS, Chang MH, Ni YH. A common variant in the PNPLA3 gene is a risk factor for non-alcoholic fatty liver disease in obese Taiwanese children. J Pediatr. 2011;158:740-744. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 75] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 35. | Valenti L, Alisi A, Galmozzi E, Bartuli A, Del Menico B, Alterio A, Dongiovanni P, Fargion S, Nobili V. I148M patatin-like phospholipase domain-containing 3 gene variant and severity of pediatric nonalcoholic fatty liver disease. Hepatology. 2010;52:1274-1280. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 229] [Article Influence: 15.3] [Reference Citation Analysis (0)] |
| 36. | Trépo E, Pradat P, Potthoff A, Momozawa Y, Quertinmont E, Gustot T, Lemmers A, Berthillon P, Amininejad L, Chevallier M. Impact of patatin-like phospholipase-3 (rs738409 C& gt; G) polymorphism on fibrosis progression and steatosis in chronic hepatitis C. Hepatology. 2011;54:60-69. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 149] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 37. | Petta S, Vanni E, Bugianesi E, Rosso C, Cabibi D, Cammà C, Di Marco V, Eslam M, Grimaudo S, Macaluso FS. PNPLA3 rs738409 I748M is associated with steatohepatitis in 434 non-obese subjects with hepatitis C. Aliment Pharmacol Ther. 2015;41:939-948. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 17] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 38. | Viganò M, Valenti L, Lampertico P, Facchetti F, Motta BM, D’Ambrosio R, Romagnoli S, Dongiovanni P, Donati B, Fargion S. Patatin-like phospholipase domain-containing 3 I148M affects liver steatosis in patients with chronic hepatitis B. Hepatology. 2013;58:1245-1252. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 57] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 39. | Stickel F, Buch S, Lau K, Meyer zu Schwabedissen H, Berg T, Ridinger M, Rietschel M, Schafmayer C, Braun F, Hinrichsen H. Genetic variation in the PNPLA3 gene is associated with alcoholic liver injury in caucasians. Hepatology. 2011;53:86-95. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 203] [Cited by in RCA: 217] [Article Influence: 15.5] [Reference Citation Analysis (0)] |
| 40. | Trepo E, Guyot E, Ganne-Carrie N, Degre D, Gustot T, Franchimont D, Sutton A, Nahon P, Moreno C. PNPLA3 (rs738409 C& gt; G) is a common risk variant associated with hepatocellular carcinoma in alcoholic cirrhosis. Hepatology. 2012;55:1307-1308. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 61] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 41. | Liu YL, Patman GL, Leathart JB, Piguet AC, Burt AD, Dufour JF, Day CP, Daly AK, Reeves HL, Anstee QM. Carriage of the PNPLA3 rs738409 C & gt; G polymorphism confers an increased risk of non-alcoholic fatty liver disease associated hepatocellular carcinoma. J Hepatol. 2014;61:75-81. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 329] [Cited by in RCA: 414] [Article Influence: 37.6] [Reference Citation Analysis (0)] |
| 42. | Burza MA, Pirazzi C, Maglio C, Sjöholm K, Mancina RM, Svensson PA, Jacobson P, Adiels M, Baroni MG, Borén J. PNPLA3 I148M (rs738409) genetic variant is associated with hepatocellular carcinoma in obese individuals. Dig Liver Dis. 2012;44:1037-1041. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 87] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 43. | Krawczyk M, Stokes CS, Romeo S, Lammert F. HCC and liver disease risks in homozygous PNPLA3 p.I148M carriers approach monogenic inheritance. J Hepatol. 2015;62:980-981. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 44. | Krawczyk M, Portincasa P, Lammert F. PNPLA3-associated steatohepatitis: toward a gene-based classification of fatty liver disease. Semin Liver Dis. 2013;33:369-379. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 68] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 45. | Petta S, Valenti L, Marchesini G, Di Marco V, Licata A, Cammà C, Barcellona MR, Cabibi D, Donati B, Fracanzani A. PNPLA3 GG genotype and carotid atherosclerosis in patients with non-alcoholic fatty liver disease. PLoS One. 2013;8:e74089. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 62] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 46. | Musso G, Cassader M, Gambino R. PNPLA3 rs738409 and TM6SF2 rs58542926 gene variants affect renal disease and function in nonalcoholic fatty liver disease. Hepatology. 2015;62:658-659. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 50] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 47. | Speliotes EK, Yerges-Armstrong LM, Wu J, Hernaez R, Kim LJ, Palmer CD, Gudnason V, Eiriksdottir G, Garcia ME, Launer LJ. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet. 2011;7:e1001324. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 775] [Cited by in RCA: 762] [Article Influence: 54.4] [Reference Citation Analysis (0)] |
| 48. | Kozlitina J, Smagris E, Stender S, Nordestgaard BG, Zhou HH, Tybjærg-Hansen A, Vogt TF, Hobbs HH, Cohen JC. Exome-wide association study identifies a TM6SF2 variant that confers susceptibility to nonalcoholic fatty liver disease. Nat Genet. 2014;46:352-356. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 724] [Cited by in RCA: 931] [Article Influence: 84.6] [Reference Citation Analysis (0)] |
| 49. | Mahdessian H, Taxiarchis A, Popov S, Silveira A, Franco-Cereceda A, Hamsten A, Eriksson P, van’t Hooft F. TM6SF2 is a regulator of liver fat metabolism influencing triglyceride secretion and hepatic lipid droplet content. Proc Natl Acad Sci USA. 2014;111:8913-8918. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 217] [Cited by in RCA: 286] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 50. | Liu YL, Reeves HL, Burt AD, Tiniakos D, McPherson S, Leathart JB, Allison ME, Alexander GJ, Piguet AC, Anty R. TM6SF2 rs58542926 influences hepatic fibrosis progression in patients with non-alcoholic fatty liver disease. Nat Commun. 2014;5:4309. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 362] [Cited by in RCA: 461] [Article Influence: 41.9] [Reference Citation Analysis (0)] |
| 51. | Wong VW, Wong GL, Tse CH, Chan HL. Prevalence of the TM6SF2 variant and non-alcoholic fatty liver disease in Chinese. J Hepatol. 2014;61:708-709. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 64] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 52. | Sookoian S, Castaño GO, Scian R, Mallardi P, Fernández Gianotti T, Burgueño AL, San Martino J, Pirola CJ. Genetic variation in transmembrane 6 superfamily member 2 and the risk of nonalcoholic fatty liver disease and histological disease severity. Hepatology. 2015;61:515-525. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 144] [Cited by in RCA: 167] [Article Influence: 16.7] [Reference Citation Analysis (0)] |
| 53. | Wang X, Liu Z, Peng Z, Liu W. The TM6SF2 rs58542926 T allele is significantly associated with non-alcoholic fatty liver disease in Chinese. J Hepatol. 2015;62:1438-1439. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 40] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 54. | Dongiovanni P, Petta S, Maglio C, Fracanzani AL, Pipitone R, Mozzi E, Motta BM, Kaminska D, Rametta R, Grimaudo S. Transmembrane 6 superfamily member 2 gene variant disentangles nonalcoholic steatohepatitis from cardiovascular disease. Hepatology. 2015;61:506-514. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 348] [Cited by in RCA: 418] [Article Influence: 41.8] [Reference Citation Analysis (0)] |
| 55. | Grimsby J, Coffey JW, Dvorozniak MT, Magram J, Li G, Matschinsky FM, Shiota C, Kaur S, Magnuson MA, Grippo JF. Characterization of glucokinase regulatory protein-deficient mice. J Biol Chem. 2000;275:7826-7831. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 100] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 56. | Peter A, Stefan N, Cegan A, Walenta M, Wagner S, Königsrainer A, Königsrainer I, Machicao F, Schick F, Häring HU. Hepatic glucokinase expression is associated with lipogenesis and fatty liver in humans. J Clin Endocrinol Metab. 2011;96:E1126-E1130. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 90] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 57. | Gorden A, Yang R, Yerges-Armstrong LM, Ryan KA, Speliotes E, Borecki IB, Harris TB, Chu X, Wood GC, Still CD. Genetic variation at NCAN locus is associated with inflammation and fibrosis in non-alcoholic fatty liver disease in morbid obesity. Hum Hered. 2013;75:34-43. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 76] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 58. | Lin YC, Chang PF, Chang MH, Ni YH. Genetic variants in GCKR and PNPLA3 confer susceptibility to nonalcoholic fatty liver disease in obese individuals. Am J Clin Nutr. 2014;99:869-874. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 87] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 59. | Tan HL, Zain SM, Mohamed R, Rampal S, Chin KF, Basu RC, Cheah PL, Mahadeva S, Mohamed Z. Association of glucokinase regulatory gene polymorphisms with risk and severity of non-alcoholic fatty liver disease: an interaction study with adiponutrin gene. J Gastroenterol. 2014;49:1056-1064. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 62] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 60. | Yang Z, Wen J, Tao X, Lu B, Du Y, Wang M, Wang X, Zhang W, Gong W, Ling C. Genetic variation in the GCKR gene is associated with non-alcoholic fatty liver disease in Chinese people. Mol Biol Rep. 2011;38:1145-1150. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 52] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 61. | Petta S, Miele L, Bugianesi E, Cammà C, Rosso C, Boccia S, Cabibi D, Di Marco V, Grimaudo S, Grieco A. Glucokinase regulatory protein gene polymorphism affects liver fibrosis in non-alcoholic fatty liver disease. PLoS One. 2014;9:e87523. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 116] [Cited by in RCA: 110] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 62. | Zain SM, Mohamed Z, Mohamed R. Common variant in the glucokinase regulatory gene rs780094 and risk of nonalcoholic fatty liver disease: a meta-analysis. J Gastroenterol Hepatol. 2015;30:21-27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 64] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 63. | Chalasani N, Guo X, Loomba R, Goodarzi MO, Haritunians T, Kwon S, Cui J, Taylor KD, Wilson L, Cummings OW, Chen YD, Rotter JI; Nonalcoholic Steatohepatitis Clinical Research Network. Genome-wide association study identifies variants associated with histologic features of nonalcoholic Fatty liver disease. Gastroenterology. 2010;139:1567-1576, 1576.e1-e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 224] [Cited by in RCA: 256] [Article Influence: 17.1] [Reference Citation Analysis (0)] |
| 64. | Michael MD, Kulkarni RN, Postic C, Previs SF, Shulman GI, Magnuson MA, Kahn CR. Loss of insulin signaling in hepatocytes leads to severe insulin resistance and progressive hepatic dysfunction. Mol Cell. 2000;6:87-97. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 246] [Cited by in RCA: 136] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 65. | Grarup N, Urhammer SA, Ek J, Albrechtsen A, Glümer C, Borch-Johnsen K, Jørgensen T, Hansen T, Pedersen O. Studies of the relationship between the ENPP1 K121Q polymorphism and type 2 diabetes, insulin resistance and obesity in 7,333 Danish white subjects. Diabetologia. 2006;49:2097-2104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 87] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 66. | McGettrick AJ, Feener EP, Kahn CR. Human insulin receptor substrate-1 (IRS-1) polymorphism G972R causes IRS-1 to associate with the insulin receptor and inhibit receptor autophosphorylation. J Biol Chem. 2005;280:6441-6446. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 53] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 67. | Jellema A, Zeegers MP, Feskens EJ, Dagnelie PC, Mensink RP. Gly972Arg variant in the insulin receptor substrate-1 gene and association with Type 2 diabetes: a meta-analysis of 27 studies. Diabetologia. 2003;46:990-995. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 105] [Cited by in RCA: 102] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 68. | Dongiovanni P, Valenti L, Rametta R, Daly AK, Nobili V, Mozzi E, Leathart JB, Pietrobattista A, Burt AD, Maggioni M. Genetic variants regulating insulin receptor signalling are associated with the severity of liver damage in patients with non-alcoholic fatty liver disease. Gut. 2010;59:267-273. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 137] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 69. | Hara K, Yamauchi T, Kadowaki T. Adiponectin: an adipokine linking adipocytes and type 2 diabetes in humans. Curr Diab Rep. 2005;5:136-140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 51] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 70. | Hui JM, Hodge A, Farrell GC, Kench JG, Kriketos A, George J. Beyond insulin resistance in NASH: TNF-alpha or adiponectin? Hepatology. 2004;40:46-54. [PubMed] |
| 71. | Ma H, Gomez V, Lu L, Yang X, Wu X, Xiao SY. Expression of adiponectin and its receptors in livers of morbidly obese patients with non-alcoholic fatty liver disease. J Gastroenterol Hepatol. 2009;24:233-237. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 59] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 72. | Kamada Y, Tamura S, Kiso S, Matsumoto H, Saji Y, Yoshida Y, Fukui K, Maeda N, Nishizawa H, Nagaretani H. Enhanced carbon tetrachloride-induced liver fibrosis in mice lacking adiponectin. Gastroenterology. 2003;125:1796-1807. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 358] [Cited by in RCA: 354] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 73. | Matsumoto H, Tamura S, Kamada Y, Kiso S, Fukushima J, Wada A, Maeda N, Kihara S, Funahashi T, Matsuzawa Y. Adiponectin deficiency exacerbates lipopolysaccharide/D-galactosamine-induced liver injury in mice. World J Gastroenterol. 2006;12:3352-3358. [PubMed] |
| 74. | Musso G, Gambino R, De Michieli F, Durazzo M, Pagano G, Cassader M. Adiponectin gene polymorphisms modulate acute adiponectin response to dietary fat: Possible pathogenetic role in NASH. Hepatology. 2008;47:1167-1177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 100] [Cited by in RCA: 103] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 75. | Tokushige K, Hashimoto E, Noto H, Yatsuji S, Taniai M, Torii N, Shiratori K. Influence of adiponectin gene polymorphisms in Japanese patients with non-alcoholic fatty liver disease. J Gastroenterol. 2009;44:976-982. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 76. | Wang ZL, Xia B, Shrestha U, Jiang L, Ma CW, Chen Q, Chen H, Hu ZG. Correlation between adiponectin polymorphisms and non-alcoholic fatty liver disease with or without metabolic syndrome in Chinese population. J Endocrinol Invest. 2008;31:1086-1091. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 35] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 77. | Denver RJ, Bonett RM, Boorse GC. Evolution of leptin structure and function. Neuroendocrinology. 2011;94:21-38. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 148] [Cited by in RCA: 157] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 78. | Chitturi S, Farrell G, Frost L, Kriketos A, Lin R, Fung C, Liddle C, Samarasinghe D, George J. Serum leptin in NASH correlates with hepatic steatosis but not fibrosis: a manifestation of lipotoxicity? Hepatology. 2002;36:403-409. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 247] [Cited by in RCA: 237] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 79. | Uygun A, Kadayifci A, Yesilova Z, Erdil A, Yaman H, Saka M, Deveci MS, Bagci S, Gulsen M, Karaeren N. Serum leptin levels in patients with nonalcoholic steatohepatitis. Am J Gastroenterol. 2000;95:3584-3589. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 171] [Cited by in RCA: 165] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 80. | Nakao K, Nakata K, Ohtsubo N, Maeda M, Moriuchi T, Ichikawa T, Hamasaki K, Kato Y, Eguchi K, Yukawa K. Association between nonalcoholic fatty liver, markers of obesity, and serum leptin level in young adults. Am J Gastroenterol. 2002;97:1796-1801. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 54] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 81. | Lahlou N, Clement K, Carel JC, Vaisse C, Lotton C, Le Bihan Y, Basdevant A, Lebouc Y, Froguel P, Roger M. Soluble leptin receptor in serum of subjects with complete resistance to leptin: relation to fat mass. Diabetes. 2000;49:1347-1352. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 58] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 82. | Wauters M, Mertens I, Rankinen T, Chagnon M, Bouchard C, Van Gaal L. Leptin receptor gene polymorphisms are associated with insulin in obese women with impaired glucose tolerance. J Clin Endocrinol Metab. 2001;86:3227-3232. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 31] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 83. | Lu H, Sun J, Sun L, Shu X, Xu Y, Xie D. Polymorphism of human leptin receptor gene is associated with type 2 diabetic patients complicated with non-alcoholic fatty liver disease in China. J Gastroenterol Hepatol. 2009;24:228-232. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 32] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 84. | Swellam M, Hamdy N. Association of nonalcoholic fatty liver disease with a single nucleotide polymorphism on the gene encoding leptin receptor. IUBMB Life. 2012;64:180-186. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 34] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 85. | Zain SM, Mohamed Z, Mahadeva S, Cheah PL, Rampal S, Chin KF, Mahfudz AS, Basu RC, Tan HL, Mohamed R. Impact of leptin receptor gene variants on risk of non-alcoholic fatty liver disease and its interaction with adiponutrin gene. J Gastroenterol Hepatol. 2013;28:873-879. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 32] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 86. | Aller R, De Luis DA, Izaola O, González Sagrado M, Conde R, Pacheco D, Velasco MC, Ovalle HF. Lys656Asn polymorphism of leptin receptor, leptin levels and insulin resistance in patients with non alcoholic fatty liver disease. Eur Rev Med Pharmacol Sci. 2012;16:335-341. [PubMed] |
| 87. | Muse ED, Obici S, Bhanot S, Monia BP, McKay RA, Rajala MW, Scherer PE, Rossetti L. Role of resistin in diet-induced hepatic insulin resistance. J Clin Invest. 2004;114:232-239. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 262] [Cited by in RCA: 249] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 88. | Song H, Shojima N, Sakoda H, Ogihara T, Fujishiro M, Katagiri H, Anai M, Onishi Y, Ono H, Inukai K. Resistin is regulated by C/EBPs, PPARs, and signal-transducing molecules. Biochem Biophys Res Commun. 2002;299:291-298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 51] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 89. | Silswal N, Singh AK, Aruna B, Mukhopadhyay S, Ghosh S, Ehtesham NZ. Human resistin stimulates the pro-inflammatory cytokines TNF-alpha and IL-12 in macrophages by NF-kappaB-dependent pathway. Biochem Biophys Res Commun. 2005;334:1092-1101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 410] [Cited by in RCA: 456] [Article Influence: 22.8] [Reference Citation Analysis (0)] |
| 90. | Bertolani C, Sancho-Bru P, Failli P, Bataller R, Aleffi S, DeFranco R, Mazzinghi B, Romagnani P, Milani S, Ginés P. Resistin as an intrahepatic cytokine: overexpression during chronic injury and induction of proinflammatory actions in hepatic stellate cells. Am J Pathol. 2006;169:2042-2053. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 120] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 91. | Tan MS, Chang SY, Chang DM, Tsai JC, Lee YJ. Association of resistin gene 3’-untranslated region +62G--& gt; A polymorphism with type 2 diabetes and hypertension in a Chinese population. J Clin Endocrinol Metab. 2003;88:1258-1263. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 47] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 92. | Zhang LY, Jin YJ, Jin QS, Lin LY, Zhang DD, Kong LL. Association between resistin +299A/A genotype and nonalcoholic fatty liver disease in Chinese patients with type 2 diabetes mellitus. Gene. 2013;529:340-344. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 13] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 93. | Vance DE, Walkey CJ, Cui Z. Phosphatidylethanolamine N-methyltransferase from liver. Biochim Biophys Acta. 1997;1348:142-150. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 154] [Cited by in RCA: 160] [Article Influence: 5.7] [Reference Citation Analysis (1)] |
| 94. | Zhu X, Song J, Mar MH, Edwards LJ, Zeisel SH. Phosphatidylethanolamine N-methyltransferase (PEMT) knockout mice have hepatic steatosis and abnormal hepatic choline metabolite concentrations despite ingesting a recommended dietary intake of choline. Biochem J. 2003;370:987-993. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 94] [Cited by in RCA: 103] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 95. | Song J, da Costa KA, Fischer LM, Kohlmeier M, Kwock L, Wang S, Zeisel SH. Polymorphism of the PEMT gene and susceptibility to nonalcoholic fatty liver disease (NAFLD). FASEB J. 2005;19:1266-1271. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 183] [Cited by in RCA: 190] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 96. | Dong H, Wang J, Li C, Hirose A, Nozaki Y, Takahashi M, Ono M, Akisawa N, Iwasaki S, Saibara T. The phosphatidylethanolamine N-methyltransferase gene V175M single nucleotide polymorphism confers the susceptibility to NASH in Japanese population. J Hepatol. 2007;46:915-920. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 95] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 97. | Jun DW, Han JH, Jang EC, Kim SH, Kim SH, Jo YJ, Park YS, Chae JD. Polymorphisms of microsomal triglyceride transfer protein gene and phosphatidylethanolamine N-methyltransferase gene in alcoholic and nonalcoholic fatty liver disease in Koreans. Eur J Gastroenterol Hepatol. 2009;21:667-672. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 22] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 98. | Romeo S, Cohen JC, Hobbs HH. No association between polymorphism in PEMT (V175M) and hepatic triglyceride content in the Dallas Heart Study. FASEB J. 2006;20:2180; author reply 2181-2182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 17] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 99. | Victor RG, Haley RW, Willett DL, Peshock RM, Vaeth PC, Leonard D, Basit M, Cooper RS, Iannacchione VG, Visscher WA. The Dallas Heart Study: a population-based probability sample for the multidisciplinary study of ethnic differences in cardiovascular health. Am J Cardiol. 2004;93:1473-1480. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 404] [Cited by in RCA: 436] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 100. | Wu Q, Ortegon AM, Tsang B, Doege H, Feingold KR, Stahl A. FATP1 is an insulin-sensitive fatty acid transporter involved in diet-induced obesity. Mol Cell Biol. 2006;26:3455-3467. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 221] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 101. | Hirsch D, Stahl A, Lodish HF. A family of fatty acid transporters conserved from mycobacterium to man. Proc Natl Acad Sci USA. 1998;95:8625-8629. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 317] [Cited by in RCA: 326] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 102. | Doege H, Baillie RA, Ortegon AM, Tsang B, Wu Q, Punreddy S, Hirsch D, Watson N, Gimeno RE, Stahl A. Targeted deletion of FATP5 reveals multiple functions in liver metabolism: alterations in hepatic lipid homeostasis. Gastroenterology. 2006;130:1245-1258. [PubMed] |
| 103. | Auinger A, Valenti L, Pfeuffer M, Helwig U, Herrmann J, Fracanzani AL, Dongiovanni P, Fargion S, Schrezenmeir J, Rubin D. A promoter polymorphism in the liver-specific fatty acid transport protein 5 is associated with features of the metabolic syndrome and steatosis. Horm Metab Res. 2010;42:854-859. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 38] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 104. | Rubin D, Helwig U, Nothnagel M, Fölsch UR, Schreiber S, Schrezenmeir J. Association of postprandial and fasting triglycerides with traits of the metabolic syndrome in the Metabolic Intervention Cohort Kiel. Eur J Endocrinol. 2010;162:719-727. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 105. | Bao X, Mills PJ, Rana BK, Dimsdale JE, Schork NJ, Smith DW, Rao F, Milic M, O’Connor DT, Ziegler MG. Interactive effects of common beta2-adrenoceptor haplotypes and age on susceptibility to hypertension and receptor function. Hypertension. 2005;46:301-307. [PubMed] |
| 106. | Iwamoto N, Ogawa Y, Kajihara S, Hisatomi A, Yasutake T, Yoshimura T, Mizuta T, Hara T, Ozaki I, Yamamoto K. Gln27Glu beta2-adrenergic receptor variant is associated with hypertriglyceridemia and the development of fatty liver. Clin Chim Acta. 2001;314:85-91. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 40] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 107. | Vardeny O, Detry MA, Moran JJ, Johnson MR, Sweitzer NK. The beta2 adrenergic receptor Gln27Glu polymorphism affects insulin resistance in patients with heart failure: possible modulation by choice of beta blocker. J Cardiovasc Pharmacol. 2008;52:500-506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 10] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 108. | Isaza CA, Henao J, Sánchez JC, Porras GL, Cardona J, Bedoya G. Beta-2-adrenergic receptor polymorphisms and changes in lipids induced by metoprolol. Pharmacology. 2007;80:279-285. [PubMed] |
| 109. | Nozaki Y, Saibara T, Nemoto Y, Ono M, Akisawa N, Iwasaki S, Hayashi Y, Hiroi M, Enzan H, Onishi S. Polymorphisms of interleukin-1 beta and beta 3-adrenergic receptor in Japanese patients with nonalcoholic steatohepatitis. Alcohol Clin Exp Res. 2004;28:106S-110S. [PubMed] |
| 110. | Loomba R, Rao F, Zhang L, Khandrika S, Ziegler MG, Brenner DA, O’Connor DT. Genetic covariance between gamma-glutamyl transpeptidase and fatty liver risk factors: role of beta2-adrenergic receptor genetic variation in twins. Gastroenterology. 2010;139:836-845, 845.e1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 47] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 111. | Petta S, Macaluso FS, Barcellona MR, Cammà C, Cabibi D, Di Marco V, Craxì A. Serum γ-glutamyl transferase levels, insulin resistance and liver fibrosis in patients with chronic liver diseases. PLoS One. 2012;7:e51165. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 35] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 112. | Jo SK, Lee WY, Rhee EJ, Won JC, Jung CH, Park CY, Oh KW, Park SW, Kim SW. Serum gamma-glutamyl transferase activity predicts future development of metabolic syndrome defined by 2 different criteria. Clin Chim Acta. 2009;403:234-240. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 37] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 113. | Pawlak M, Lefebvre P, Staels B. Molecular mechanism of PPARα action and its impact on lipid metabolism, inflammation and fibrosis in non-alcoholic fatty liver disease. J Hepatol. 2015;62:720-733. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 793] [Cited by in RCA: 1145] [Article Influence: 114.5] [Reference Citation Analysis (0)] |
| 114. | Di Rosa M, Malaguarnera L. Genetic variants in candidate genes influencing NAFLD progression. J Mol Med (Berl). 2012;90:105-118. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 40] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 115. | Chen S, Li Y, Li S, Yu C. A Val227Ala substitution in the peroxisome proliferator activated receptor alpha (PPAR alpha) gene associated with non-alcoholic fatty liver disease and decreased waist circumference and waist-to-hip ratio. J Gastroenterol Hepatol. 2008;23:1415-1418. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 52] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 116. | Dongiovanni P, Rametta R, Fracanzani AL, Benedan L, Borroni V, Maggioni P, Maggioni M, Fargion S, Valenti L. Lack of association between peroxisome proliferator-activated receptors alpha and gamma2 polymorphisms and progressive liver damage in patients with non-alcoholic fatty liver disease: a case control study. BMC Gastroenterol. 2010;10:102. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 47] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 117. | Yen CJ, Beamer BA, Negri C, Silver K, Brown KA, Yarnall DP, Burns DK, Roth J, Shuldiner AR. Molecular scanning of the human peroxisome proliferator activated receptor gamma (hPPAR gamma) gene in diabetic Caucasians: identification of a Pro12Ala PPAR gamma 2 missense mutation. Biochem Biophys Res Commun. 1997;241:270-274. [PubMed] |
| 118. | Wang J, Guo X, Wu P, Song J, Ye C, Yu S, Zhang J, Dong W. Association between the Pro12Ala polymorphism of PPAR-γ gene and the non-alcoholic fatty liver disease: a meta-analysis. Gene. 2013;528:328-334. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 12] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 119. | Bhatt SP, Nigam P, Misra A, Guleria R, Luthra K, Pandey RM, Pasha MA. Association of peroxisome proliferator activated receptor-γ gene with non-alcoholic fatty liver disease in Asian Indians residing in north India. Gene. 2013;512:143-147. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 24] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 120. | Cao CY, Li YY, Zhou YJ, Nie YQ, Wan YJ. The C-681G polymorphism of the PPAR-γ gene is associated with susceptibility to non-alcoholic fatty liver disease. Tohoku J Exp Med. 2012;227:253-262. [PubMed] |
| 121. | Gupta AC, Chaudhory AK, Sukriti C, Sakhuja P, Singh Y, Basir SF, Sarin SK. Peroxisome proliferators-activated receptor γ2 Pro12Ala variant is associated with body mass index in non-alcoholic fatty liver disease patients. Hepatol Int. 2010;5:575-580. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 23] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 122. | Yang Z, Wen J, Li Q, Tao X, Ye Z, He M, Zhang W, Huang Y, Chen L, Ling C. PPARG gene Pro12Ala variant contributes to the development of non-alcoholic fatty liver in middle-aged and older Chinese population. Mol Cell Endocrinol. 2012;348:255-259. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 28] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 123. | Ye Q, Lv ZS. [A study of the association between PPARr2 gene Pro12Ala polymorphism and NAFLD]. Zhonghua Ganzangbing Zazhi. 2007;15:228-229. [PubMed] |
| 124. | Gawrieh S, Marion MC, Komorowski R, Wallace J, Charlton M, Kissebah A, Langefeld CD, Olivier M. Genetic variation in the peroxisome proliferator activated receptor-gamma gene is associated with histologically advanced NAFLD. Dig Dis Sci. 2012;57:952-957. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 40] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 125. | Rey JW, Noetel A, Hardt A, Canbay A, Alakus H, Zur Hausen A, Dienes HP, Drebber U, Odenthal M. Pro12Ala polymorphism of the peroxisome proliferator-activated receptor γ2 in patients with fatty liver diseases. World J Gastroenterol. 2010;16:5830-5837. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 28] [Cited by in RCA: 31] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 126. | Soyal S, Krempler F, Oberkofler H, Patsch W. PGC-1alpha: a potent transcriptional cofactor involved in the pathogenesis of type 2 diabetes. Diabetologia. 2006;49:1477-1488. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 113] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 127. | Barroso I, Luan J, Sandhu MS, Franks PW, Crowley V, Schafer AJ, O’Rahilly S, Wareham NJ. Meta-analysis of the Gly482Ser variant in PPARGC1A in type 2 diabetes and related phenotypes. Diabetologia. 2006;49:501-505. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 77] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 128. | Vimaleswaran KS, Luan J, Andersen G, Muller YL, Wheeler E, Brito EC, O’Rahilly S, Pedersen O, Baier LJ, Knowler WC. The Gly482Ser genotype at the PPARGC1A gene and elevated blood pressure: a meta-analysis involving 13,949 individuals. J Appl Physiol (1985). 2008;105:1352-1358. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 21] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 129. | Lagou V, Scott RA, Manios Y, Chen TL, Wang G, Grammatikaki E, Kortsalioudaki C, Liarigkovinos T, Moschonis G, Roma-Giannikou E. Impact of peroxisome proliferator-activated receptors gamma and delta on adiposity in toddlers and preschoolers in the GENESIS Study. Obesity (Silver Spring). 2008;16:913-918. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 34] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 130. | Chen Y, Mu P, He S, Tang X, Guo X, Li H, Xu H, Woo SL, Qian X, Zeng L. Gly482Ser mutation impairs the effects of peroxisome proliferator-activated receptor γ coactivator-1α on decreasing fat deposition and stimulating phosphoenolpyruvate carboxykinase expression in hepatocytes. Nutr Res. 2013;33:332-339. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 14] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 131. | Lin YC, Chang PF, Chang MH, Ni YH. A common variant in the peroxisome proliferator-activated receptor-γ coactivator-1α gene is associated with nonalcoholic fatty liver disease in obese children. Am J Clin Nutr. 2013;97:326-331. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 39] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 132. | Yoneda M, Hotta K, Nozaki Y, Endo H, Uchiyama T, Mawatari H, Iida H, Kato S, Hosono K, Fujita K. Association between PPARGC1A polymorphisms and the occurrence of nonalcoholic fatty liver disease (NAFLD). BMC Gastroenterol. 2008;8:27. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 59] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 133. | Hui Y, Yu-Yuan L, Yu-Qiang N, Wei-Hong S, Yan-Lei D, Xiao-Bo L, Yong-Jian Z. Effect of peroxisome proliferator-activated receptors-gamma and co-activator-1alpha genetic polymorphisms on plasma adiponectin levels and susceptibility of non-alcoholic fatty liver disease in Chinese people. Liver Int. 2008;28:385-392. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 58] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 134. | Mahley RW, Nathan BP, Pitas RE. Apolipoprotein E. Structure, function, and possible roles in Alzheimer’s disease. Ann N Y Acad Sci. 1996;777:139-145. [PubMed] |
| 135. | Demirag MD, Onen HI, Karaoguz MY, Dogan I, Karakan T, Ekmekci A, Guz G. Apolipoprotein E gene polymorphism in nonalcoholic fatty liver disease. Dig Dis Sci. 2007;52:3399-3403. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 27] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 136. | Sazci A, Akpinar G, Aygun C, Ergul E, Senturk O, Hulagu S. Association of apolipoprotein E polymorphisms in patients with non-alcoholic steatohepatitis. Dig Dis Sci. 2008;53:3218-3224. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 62] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 137. | Lee DM, Lee SO, Mun BS, Ahn HS, Park HY, Lee HS, Kim DG. [Relation of apolipoprotein E polymorphism to clinically diagnosed fatty liver disease]. Taehan Kan Hakhoe Chi. 2002;8:355-362. [PubMed] |
| 138. | Yang MH, Son HJ, Sung JD, Choi YH, Koh KC, Yoo BC, Paik SW. The relationship between apolipoprotein E polymorphism, lipoprotein (a) and fatty liver disease. Hepatogastroenterology. 2005;52:1832-1835. [PubMed] |
| 139. | De Feo E, Cefalo C, Arzani D, Amore R, Landolfi R, Grieco A, Ricciardi W, Miele L, Boccia S. A case-control study on the effects of the apolipoprotein E genotypes in nonalcoholic fatty liver disease. Mol Biol Rep. 2012;39:7381-7388. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 21] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 140. | Ginsberg HN, Le NA, Goldberg IJ, Gibson JC, Rubinstein A, Wang-Iverson P, Norum R, Brown WV. Apolipoprotein B metabolism in subjects with deficiency of apolipoproteins CIII and AI. Evidence that apolipoprotein CIII inhibits catabolism of triglyceride-rich lipoproteins by lipoprotein lipase in vivo. J Clin Invest. 1986;78:1287-1295. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 333] [Cited by in RCA: 343] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 141. | Miller M, Rhyne J, Chen H, Beach V, Ericson R, Luthra K, Dwivedi M, Misra A. APOC3 promoter polymorphisms C-482T and T-455C are associated with the metabolic syndrome. Arch Med Res. 2007;38:444-451. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 58] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 142. | Petersen KF, Dufour S, Hariri A, Nelson-Williams C, Foo JN, Zhang XM, Dziura J, Lifton RP, Shulman GI. Apolipoprotein C3 gene variants in nonalcoholic fatty liver disease. N Engl J Med. 2010;362:1082-1089. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 340] [Cited by in RCA: 320] [Article Influence: 21.3] [Reference Citation Analysis (0)] |
| 143. | Puppala J, Bhrugumalla S, Kumar A, Siddapuram SP, Viswa PD, Kondawar M, Akka J, Munshi A. Apolipoprotein C3 gene polymorphisms in Southern Indian patients with nonalcoholic fatty liver disease. Indian J Gastroenterol. 2014;33:524-529. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 9] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 144. | Li MR, Zhang SH, Chao K, Liao XH, Yao JY, Chen MH, Zhong BH. Apolipoprotein C3 (-455T& gt; C) polymorphism confers susceptibility to nonalcoholic fatty liver disease in the Southern Han Chinese population. World J Gastroenterol. 2014;20:14010-14017. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 26] [Cited by in RCA: 26] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 145. | Valenti L, Nobili V, Al-Serri A, Rametta R, Leathart JB, Zappa MA, Dongiovanni P, Fracanzani AL, Alterio A, Roviaro G. The APOC3 T-455C and C-482T promoter region polymorphisms are not associated with the severity of liver damage independently of PNPLA3 I148M genotype in patients with nonalcoholic fatty liver. J Hepatol. 2011;55:1409-1414. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 65] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 146. | Kozlitina J, Boerwinkle E, Cohen JC, Hobbs HH. Dissociation between APOC3 variants, hepatic triglyceride content and insulin resistance. Hepatology. 2011;53:467-474. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 96] [Cited by in RCA: 97] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 147. | Hyysalo J, Stojkovic I, Kotronen A, Hakkarainen A, Sevastianova K, Makkonen J, Lundbom N, Rissanen A, Krauss RM, Melander O. Genetic variation in PNPLA3 but not APOC3 influences liver fat in non-alcoholic fatty liver disease. J Gastroenterol Hepatol. 2012;27:951-956. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 44] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 148. | Peter A, Kantartzis K, Machicao F, Machann J, Wagner S, Templin S, Königsrainer I, Königsrainer A, Schick F, Fritsche A. Visceral obesity modulates the impact of apolipoprotein C3 gene variants on liver fat content. Int J Obes (Lond). 2012;36:774-782. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 27] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 149. | Verrijken A, Beckers S, Francque S, Hilden H, Caron S, Zegers D, Ruppert M, Hubens G, Van Marck E, Michielsen P. A gene variant of PNPLA3, but not of APOC3, is associated with histological parameters of NAFLD in an obese population. Obesity (Silver Spring). 2013;21:2138-2145. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 54] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 150. | Niu TH, Jiang M, Xin YN, Jiang XJ, Lin ZH, Xuan SY. Lack of association between apolipoprotein C3 gene polymorphisms and risk of nonalcoholic fatty liver disease in a Chinese Han population. World J Gastroenterol. 2014;20:3655-3662. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 27] [Cited by in RCA: 28] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 151. | Zhang H, Chen L, Xin Y, Lou Y, Liu Y, Xuan S. Apolipoprotein c3 gene polymorphisms are not a risk factor for developing non-alcoholic Fatty liver disease: a meta-analysis. Hepat Mon. 2014;14:e23100. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 17] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 152. | Hussain MM, Rava P, Walsh M, Rana M, Iqbal J. Multiple functions of microsomal triglyceride transfer protein. Nutr Metab (Lond). 2012;9:14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 190] [Cited by in RCA: 194] [Article Influence: 14.9] [Reference Citation Analysis (0)] |
| 153. | Karpe F, Lundahl B, Ehrenborg E, Eriksson P, Hamsten A. A common functional polymorphism in the promoter region of the microsomal triglyceride transfer protein gene influences plasma LDL levels. Arterioscler Thromb Vasc Biol. 1998;18:756-761. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 101] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 154. | Namikawa C, Shu-Ping Z, Vyselaar JR, Nozaki Y, Nemoto Y, Ono M, Akisawa N, Saibara T, Hiroi M, Enzan H. Polymorphisms of microsomal triglyceride transfer protein gene and manganese superoxide dismutase gene in non-alcoholic steatohepatitis. J Hepatol. 2004;40:781-786. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 175] [Cited by in RCA: 174] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 155. | Gambino R, Cassader M, Pagano G, Durazzo M, Musso G. Polymorphism in microsomal triglyceride transfer protein: a link between liver disease and atherogenic postprandial lipid profile in NASH? Hepatology. 2007;45:1097-1107. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 89] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 156. | Bernard S, Touzet S, Personne I, Lapras V, Bondon PJ, Berthezène F, Moulin P. Association between microsomal triglyceride transfer protein gene polymorphism and the biological features of liver steatosis in patients with type II diabetes. Diabetologia. 2000;43:995-999. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 111] [Cited by in RCA: 107] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 157. | Oliveira CP, Stefano JT, Cavaleiro AM, Zanella Fortes MA, Vieira SM, Rodrigues Lima VM, Santos TE, Santos VN, de Azevedo Salgado AL, Parise ER. Association of polymorphisms of glutamate-cystein ligase and microsomal triglyceride transfer protein genes in non-alcoholic fatty liver disease. J Gastroenterol Hepatol. 2010;25:357-361. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 59] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 158. | Peng XE, Wu YL, Lu QQ, Hu ZJ, Lin X. MTTP polymorphisms and susceptibility to non-alcoholic fatty liver disease in a Han Chinese population. Liver Int. 2014;34:118-128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 33] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 159. | Carulli L, Canedi I, Rondinella S, Lombardini S, Ganazzi D, Fargion S, De Palma M, Lonardo A, Ricchi M, Bertolotti M. Genetic polymorphisms in non-alcoholic fatty liver disease: interleukin-6-174G/C polymorphism is associated with non-alcoholic steatohepatitis. Dig Liver Dis. 2009;41:823-828. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 66] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 160. | Zheng W, Wang L, Su X, Hu XF. MTP -493G& gt; T polymorphism and susceptibility to nonalcoholic fatty liver disease: a meta-analysis. DNA Cell Biol. 2014;33:361-369. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 16] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 161. | Reue K, Zhang P. The lipin protein family: dual roles in lipid biosynthesis and gene expression. FEBS Lett. 2008;582:90-96. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 167] [Cited by in RCA: 162] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 162. | Wiedmann S, Fischer M, Koehler M, Neureuther K, Riegger G, Doering A, Schunkert H, Hengstenberg C, Baessler A. Genetic variants within the LPIN1 gene, encoding lipin, are influencing phenotypes of the metabolic syndrome in humans. Diabetes. 2008;57:209-217. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 61] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 163. | Fawcett KA, Grimsey N, Loos RJ, Wheeler E, Daly A, Soos M, Semple R, Syddall H, Cooper C, Siniossoglou S. Evaluating the role of LPIN1 variation in insulin resistance, body weight, and human lipodystrophy in U.K. Populations. Diabetes. 2008;57:2527-2533. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 41] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 164. | Valenti L, Motta BM, Alisi A, Sartorelli R, Buonaiuto G, Dongiovanni P, Rametta R, Pelusi S, Fargion S, Nobili V. LPIN1 rs13412852 polymorphism in pediatric nonalcoholic fatty liver disease. J Pediatr Gastroenterol Nutr. 2012;54:588-593. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 51] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 165. | Hotamisligil GS, Spiegelman BM. Tumor necrosis factor alpha: a key component of the obesity-diabetes link. Diabetes. 1994;43:1271-1278. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 297] [Cited by in RCA: 251] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 166. | Wong VW, Hui AY, Tsang SW, Chan JL, Tse AM, Chan KF, So WY, Cheng AY, Ng WF, Wong GL. Metabolic and adipokine profile of Chinese patients with nonalcoholic fatty liver disease. Clin Gastroenterol Hepatol. 2006;4:1154-1161. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 148] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 167. | Valenti L, Fracanzani AL, Dongiovanni P, Santorelli G, Branchi A, Taioli E, Fiorelli G, Fargion S. Tumor necrosis factor alpha promoter polymorphisms and insulin resistance in nonalcoholic fatty liver disease. Gastroenterology. 2002;122:274-280. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 215] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 168. | Huang J, Li YY, Zhou YJ. [Association between tumor necrosis factor-alpha gene polymorphism and insulin resistance in nonalcoholic fatty liver disease]. Zhonghua Ganzangbing Zazhi. 2006;14:613-615. [PubMed] |
| 169. | Kroeger KM, Carville KS, Abraham LJ. The -308 tumor necrosis factor-alpha promoter polymorphism effects transcription. Mol Immunol. 1997;34:391-399. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 641] [Cited by in RCA: 674] [Article Influence: 24.1] [Reference Citation Analysis (0)] |
| 170. | Wilson AG, Symons JA, McDowell TL, McDevitt HO, Duff GW. Effects of a polymorphism in the human tumor necrosis factor alpha promoter on transcriptional activation. Proc Natl Acad Sci USA. 1997;94:3195-3199. [PubMed] |
| 171. | Tokushige K, Takakura M, Tsuchiya-Matsushita N, Taniai M, Hashimoto E, Shiratori K. Influence of TNF gene polymorphisms in Japanese patients with NASH and simple steatosis. J Hepatol. 2007;46:1104-1110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 122] [Cited by in RCA: 112] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 172. | Wong VW, Wong GL, Tsang SW, Hui AY, Chan AW, Choi PC, So WY, Tse AM, Chan FK, Sung JJ. Genetic polymorphisms of adiponectin and tumor necrosis factor-alpha and nonalcoholic fatty liver disease in Chinese people. J Gastroenterol Hepatol. 2008;23:914-921. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 53] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 173. | Wang JK, Feng ZW, Li YC, Li QY, Tao XY. Association of tumor necrosis factor-α gene promoter polymorphism at sites -308 and -238 with non-alcoholic fatty liver disease: a meta-analysis. J Gastroenterol Hepatol. 2012;27:670-676. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 44] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 174. | Hu ZW, Luo HB, Xu YM, Guo JW, Deng XL, Tong YW, Tang X. Tumor necrosis factor--alpha gene promoter polymorphisms in Chinese patients with nonalcoholic fatty liver diseases. Acta Gastroenterol Belg. 2009;72:215-221. [PubMed] |
| 175. | Aller R, de Luis DA, Izaola O, González Sagrado M, Conde R, Alvarez Gago T, Pacheco D, González JM, Velasco MC. G308A polymorphism of TNF-alpha gene is associated with insulin resistance and histological changes in non alcoholic fatty liver disease patients. Ann Hepatol. 2010;9:439-444. [PubMed] |
| 176. | Zhou YJ, Li YY, Nie YQ, Yang H, Zhan Q, Huang J, Shi SL, Lai XB, Huang HL. Influence of polygenetic polymorphisms on the susceptibility to non-alcoholic fatty liver disease of Chinese people. J Gastroenterol Hepatol. 2010;25:772-777. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 63] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 177. | Trujillo-Murillo K, Bosques-Padilla FJ, Caldero?n-Lozano I, Navar-Vizcarra S, Garza-Gonza?lez E, Niderhauser-Garci?a A, Flores-Gutie?rrez JP, Zorrilla-Blanco P, Salinas-Garza R, Rivas-Estilla AM. Association of tumor necrosis factor a and manganese superoxide dismutase polymorphisms in patients with non-alcoholic steatohepatitis from northeast Mexico. Open Hepatol J. 2011;3:1-6. |
| 178. | Yan X, Xu L, Qi J, Liang X, Ma C, Guo C, Zhang L, Sun W, Zhang J, Wei X. sTRAIL levels and TRAIL gene polymorphisms in Chinese patients with fatty liver disease. Immunogenetics. 2009;61:551-556. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 179. | Stojsavljević S, Gomerčić Palčić M, Virović Jukić L, Smirčić Duvnjak L, Duvnjak M. Adipokines and proinflammatory cytokines, the key mediators in the pathogenesis of nonalcoholic fatty liver disease. World J Gastroenterol. 2014;20:18070-18091. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 252] [Cited by in RCA: 252] [Article Influence: 22.9] [Reference Citation Analysis (0)] |
| 180. | Fitzpatrick E, Dew TK, Quaglia A, Sherwood RA, Mitry RR, Dhawan A. Analysis of adipokine concentrations in paediatric non-alcoholic fatty liver disease. Pediatr Obes. 2012;7:471-479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 181. | García-Galiano D, Sánchez-Garrido MA, Espejo I, Montero JL, Costán G, Marchal T, Membrives A, Gallardo-Valverde JM, Muñoz-Castañeda JR, Arévalo E. IL-6 and IGF-1 are independent prognostic factors of liver steatosis and non-alcoholic steatohepatitis in morbidly obese patients. Obes Surg. 2007;17:493-503. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 85] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 182. | Wieckowska A, Papouchado BG, Li Z, Lopez R, Zein NN, Feldstein AE. Increased hepatic and circulating interleukin-6 levels in human nonalcoholic steatohepatitis. Am J Gastroenterol. 2008;103:1372-1379. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 395] [Cited by in RCA: 443] [Article Influence: 26.1] [Reference Citation Analysis (0)] |
| 183. | Kubaszek A, Pihlajamäki J, Punnonen K, Karhapää P, Vauhkonen I, Laakso M. The C-174G promoter polymorphism of the IL-6 gene affects energy expenditure and insulin sensitivity. Diabetes. 2003;52:558-561. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 99] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 184. | Berthier MT, Paradis AM, Tchernof A, Bergeron J, Prud’homme D, Després JP, Vohl MC. The interleukin 6-174G/C polymorphism is associated with indices of obesity in men. J Hum Genet. 2003;48:14-19. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 72] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 185. | Chang YH, Huang CN, Shiau MY. The C-174G promoter polymorphism of the interleukin-6 (IL-6) gene that affects insulin sensitivity in Caucasians is not involved in the pathogenesis of Taiwanese type 2 diabetes mellitus. Eur Cytokine Netw. 2004;15:117-119. [PubMed] |
| 186. | Vozarova B, Fernández-Real JM, Knowler WC, Gallart L, Hanson RL, Gruber JD, Ricart W, Vendrell J, Richart C, Tataranni PA. The interleukin-6 (-174) G/C promoter polymorphism is associated with type-2 diabetes mellitus in Native Americans and Caucasians. Hum Genet. 2003;112:409-413. [PubMed] |
| 187. | Cardellini M, Perego L, D’Adamo M, Marini MA, Procopio C, Hribal ML, Andreozzi F, Frontoni S, Giacomelli M, Paganelli M. C-174G polymorphism in the promoter of the interleukin-6 gene is associated with insulin resistance. Diabetes Care. 2005;28:2007-2012. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 57] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 188. | Tilg H, Moschen AR. IL-1 cytokine family members and NAFLD: neglected in metabolic liver inflammation. J Hepatol. 2011;55:960-962. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 34] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 189. | Kamari Y, Shaish A, Vax E, Shemesh S, Kandel-Kfir M, Arbel Y, Olteanu S, Barshack I, Dotan S, Voronov E. Lack of interleukin-1α or interleukin-1β inhibits transformation of steatosis to steatohepatitis and liver fibrosis in hypercholesterolemic mice. J Hepatol. 2011;55:1086-1094. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 241] [Cited by in RCA: 239] [Article Influence: 17.1] [Reference Citation Analysis (0)] |
| 190. | El-Omar EM, Carrington M, Chow WH, McColl KE, Bream JH, Young HA, Herrera J, Lissowska J, Yuan CC, Rothman N. Interleukin-1 polymorphisms associated with increased risk of gastric cancer. Nature. 2000;404:398-402. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1690] [Cited by in RCA: 1676] [Article Influence: 67.0] [Reference Citation Analysis (0)] |
| 191. | Rivera CA, Adegboyega P, van Rooijen N, Tagalicud A, Allman M, Wallace M. Toll-like receptor-4 signaling and Kupffer cells play pivotal roles in the pathogenesis of non-alcoholic steatohepatitis. J Hepatol. 2007;47:571-579. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 566] [Cited by in RCA: 560] [Article Influence: 31.1] [Reference Citation Analysis (0)] |
| 192. | Decker K. Biologically active products of stimulated liver macrophages (Kupffer cells). Eur J Biochem. 1990;192:245-261. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 660] [Cited by in RCA: 628] [Article Influence: 17.9] [Reference Citation Analysis (0)] |
| 193. | Arbour NC, Lorenz E, Schutte BC, Zabner J, Kline JN, Jones M, Frees K, Watt JL, Schwartz DA. TLR4 mutations are associated with endotoxin hyporesponsiveness in humans. Nat Genet. 2000;25:187-191. [PubMed] |
| 194. | Steinhardt AP, Aranguren F, Tellechea ML, Gómez Rosso LA, Brites FD, Martínez-Larrad MT, Serrano-Ríos M, Frechtel GD, Taverna MJ. A functional nonsynonymous toll-like receptor 4 gene polymorphism is associated with metabolic syndrome, surrogates of insulin resistance, and syndromes of lipid accumulation. Metabolism. 2010;59:711-717. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 26] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 195. | Ferwerda B, McCall MB, Verheijen K, Kullberg BJ, van der Ven AJ, Van der Meer JW, Netea MG. Functional consequences of toll-like receptor 4 polymorphisms. Mol Med. 2008;14:346-352. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 164] [Cited by in RCA: 183] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 196. | Guo J, Loke J, Zheng F, Hong F, Yea S, Fukata M, Tarocchi M, Abar OT, Huang H, Sninsky JJ. Functional linkage of cirrhosis-predictive single nucleotide polymorphisms of Toll-like receptor 4 to hepatic stellate cell responses. Hepatology. 2009;49:960-968. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 175] [Cited by in RCA: 183] [Article Influence: 11.4] [Reference Citation Analysis (0)] |
| 197. | Kiziltas S, Ata P, Colak Y, Mesçi B, Senates E, Enc F, Ulasoglu C, Tuncer I, Oguz A. TLR4 gene polymorphism in patients with nonalcoholic fatty liver disease in comparison to healthy controls. Metab Syndr Relat Disord. 2014;12:165-170. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 33] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 198. | Ge D, Fellay J, Thompson AJ, Simon JS, Shianna KV, Urban TJ, Heinzen EL, Qiu P, Bertelsen AH, Muir AJ. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature. 2009;461:399-401. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2776] [Cited by in RCA: 2723] [Article Influence: 170.2] [Reference Citation Analysis (0)] |
| 199. | Thomas DL, Thio CL, Martin MP, Qi Y, Ge D, O’Huigin C, Kidd J, Kidd K, Khakoo SI, Alexander G. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature. 2009;461:798-801. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1706] [Cited by in RCA: 1687] [Article Influence: 105.4] [Reference Citation Analysis (0)] |
| 200. | Suppiah V, Moldovan M, Ahlenstiel G, Berg T, Weltman M, Abate ML, Bassendine M, Spengler U, Dore GJ, Powell E. IL28B is associated with response to chronic hepatitis C interferon-alpha and ribavirin therapy. Nat Genet. 2009;41:1100-1104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1505] [Cited by in RCA: 1504] [Article Influence: 94.0] [Reference Citation Analysis (0)] |
| 201. | Tanaka Y, Nishida N, Sugiyama M, Kurosaki M, Matsuura K, Sakamoto N, Nakagawa M, Korenaga M, Hino K, Hige S. Genome-wide association of IL28B with response to pegylated interferon-alpha and ribavirin therapy for chronic hepatitis C. Nat Genet. 2009;41:1105-1109. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1779] [Cited by in RCA: 1775] [Article Influence: 110.9] [Reference Citation Analysis (0)] |
| 202. | Rauch A, Kutalik Z, Descombes P, Cai T, Di Iulio J, Mueller T, Bochud M, Battegay M, Bernasconi E, Borovicka J. Genetic variation in IL28B is associated with chronic hepatitis C and treatment failure: a genome-wide association study. Gastroenterology. 2010;138:1338-1345, 1345.e1-e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 850] [Cited by in RCA: 867] [Article Influence: 57.8] [Reference Citation Analysis (0)] |
| 203. | Tillmann HL, Patel K, Muir AJ, Guy CD, Li JH, Lao XQ, Thompson A, Clark PJ, Gardner SD, McHutchison JG. Beneficial IL28B genotype associated with lower frequency of hepatic steatosis in patients with chronic hepatitis C. J Hepatol. 2011;55:1195-1200. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 73] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 204. | Cai T, Dufour JF, Muellhaupt B, Gerlach T, Heim M, Moradpour D, Cerny A, Malinverni R, Kaddai V, Bochud M, Negro F, Bochud PY; Swiss Hepatitis C Cohort Study Group. Viral genotype-specific role of PNPLA3, PPARG, MTTP, and IL28B in hepatitis C virus-associated steatosis. J Hepatol. 2011;55:529-535. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 85] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 205. | Abe H, Ochi H, Maekawa T, Hayes CN, Tsuge M, Miki D, Mitsui F, Hiraga N, Imamura M, Takahashi S. Common variation of IL28 affects gamma-GTP levels and inflammation of the liver in chronically infected hepatitis C virus patients. J Hepatol. 2010;53:439-443. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 117] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 206. | Marabita F, Aghemo A, De Nicola S, Rumi MG, Cheroni C, Scavelli R, Crimi M, Soffredini R, Abrignani S, De Francesco R. Genetic variation in the interleukin-28B gene is not associated with fibrosis progression in patients with chronic hepatitis C and known date of infection. Hepatology. 2011;54:1127-1134. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 104] [Cited by in RCA: 104] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 207. | Falleti E, Bitetto D, Fabris C, Cussigh A, Fornasiere E, Cmet S, Fumolo E, Bignulin S, Fontanini E, Cerutti A. Role of interleukin 28B rs12979860 C/T polymorphism on the histological outcome of chronic hepatitis C: relationship with gender and viral genotype. J Clin Immunol. 2011;31:891-899. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 61] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 208. | Thompson AJ, Clark PJ, Fellay J, Muir AJ, Tillmann HL, Patel K, Naggie S, Afdhal NH, Jacobson IM, Esteban R. IL28B genotype is not associated with advanced hepatic fibrosis in chronic hepatitis C patients enrolled in the ideal study. Hepatology. 2010;52:437A. |
| 209. | Petta S, Grimaudo S, Cammà C, Cabibi D, Di Marco V, Licata G, Pipitone RM, Craxì A. IL28B and PNPLA3 polymorphisms affect histological liver damage in patients with non-alcoholic fatty liver disease. J Hepatol. 2012;56:1356-1362. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 75] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 210. | Eslam M, Hashem AM, Leung R, Romero-Gomez M, Berg T, Dore GJ, Chan HL, Irving WL, Sheridan D, Abate ML. Interferon-λ rs12979860 genotype and liver fibrosis in viral and non-viral chronic liver disease. Nat Commun. 2015;6:6422. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 152] [Cited by in RCA: 149] [Article Influence: 14.9] [Reference Citation Analysis (0)] |
| 211. | Garrett ME, Abdelmalek MF, Ashley-Koch A, Hauser MA, Moylan CA, Pang H, Diehl AM, Tillmann HL. IL28B rs12979860 is not associated with histologic features of NAFLD in a cohort of Caucasian North American patients. J Hepatol. 2013;58:402-403. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 212. | Macmillan-Crow LA, Cruthirds DL. Invited review: manganese superoxide dismutase in disease. Free Radic Res. 2001;34:325-336. [PubMed] |
| 213. | Ratziu V, Bellentani S, Cortez-Pinto H, Day C, Marchesini G. A position statement on NAFLD/NASH based on the EASL 2009 special conference. J Hepatol. 2010;53:372-384. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 723] [Cited by in RCA: 789] [Article Influence: 52.6] [Reference Citation Analysis (1)] |
| 214. | Sutton A, Imbert A, Igoudjil A, Descatoire V, Cazanave S, Pessayre D, Degoul F. The manganese superoxide dismutase Ala16Val dimorphism modulates both mitochondrial import and mRNA stability. Pharmacogenet Genomics. 2005;15:311-319. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 184] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 215. | Sutton A, Khoury H, Prip-Buus C, Cepanec C, Pessayre D, Degoul F. The Ala16Val genetic dimorphism modulates the import of human manganese superoxide dismutase into rat liver mitochondria. Pharmacogenetics. 2003;13:145-157. [PubMed] |
| 216. | Nahon P, Sutton A, Pessayre D, Rufat P, Degoul F, Ganne-Carrie N, Ziol M, Charnaux N, N’kontchou G, Trinchet JC. Genetic dimorphism in superoxide dismutase and susceptibility to alcoholic cirrhosis, hepatocellular carcinoma, and death. Clin Gastroenterol Hepatol. 2005;3:292-298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 46] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 217. | Stewart SF, Leathart JB, Chen Y, Daly AK, Rolla R, Vay D, Mottaran E, Vidali M, Albano E, Day CP. Valine-alanine manganese superoxide dismutase polymorphism is not associated with alcohol-induced oxidative stress or liver fibrosis. Hepatology. 2002;36:1355-1360. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 11] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 218. | El-Koofy NM, El-Karaksy HM, Mandour IM, Anwar GM, El-Raziky MS, El-Hennawy AM. Genetic polymorphisms in non-alcoholic fatty liver disease in obese Egyptian children. Saudi J Gastroenterol. 2011;17:265-270. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 25] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 219. | Al-Serri A, Anstee QM, Valenti L, Nobili V, Leathart JB, Dongiovanni P, Patch J, Fracanzani A, Fargion S, Day CP. The SOD2 C47T polymorphism influences NAFLD fibrosis severity: evidence from case-control and intra-familial allele association studies. J Hepatol. 2012;56:448-454. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 124] [Cited by in RCA: 136] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 220. | Huang YS, Chang CH, Lin TL, Perng CL. Genetic variations of superoxide dismutase 2 and cytochrome P450 2E1 in non-alcoholic steatohepatitis. Liver Int. 2014;34:931-936. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 13] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 221. | Leung TM, Nieto N. CYP2E1 and oxidant stress in alcoholic and non-alcoholic fatty liver disease. J Hepatol. 2013;58:395-398. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 311] [Cited by in RCA: 386] [Article Influence: 32.2] [Reference Citation Analysis (0)] |
| 222. | Varela NM, Quiñones LA, Orellana M, Poniachik J, Csendes A, Smok G, Rodrigo R, Cáceres DD, Videla LA. Study of cytochrome P450 2E1 and its allele variants in liver injury of nondiabetic, nonalcoholic steatohepatitis obese women. Biol Res. 2008;41:81-92. [PubMed] |
| 223. | Piao YF, Li JT, Shi Y. Relationship between genetic polymorphism of cytochrome P450IIE1 and fatty liver. World J Gastroenterol. 2003;9:2612-2615. [PubMed] |
| 224. | Samec S, Seydoux J, Dulloo AG. Role of UCP homologues in skeletal muscles and brown adipose tissue: mediators of thermogenesis or regulators of lipids as fuel substrate? FASEB J. 1998;12:715-724. [PubMed] |
| 225. | de Luis DA, Aller R, Izaola O, Sagrado MG, Conde R. Modulation of adipocytokines response and weight loss secondary to a hypocaloric diet in obese patients by -55CT polymorphism of UCP3 gene. Horm Metab Res. 2008;40:214-218. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 11] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 226. | de Luis DA, Aller R, Izaola O, González Sagrado M, Conde R, Pérez Castrillón JL. Lack of association of -55CT polymorphism of UCP3 gene with fat distribution in obese patients. Ann Nutr Metab. 2007;51:374-378. [PubMed] |
| 227. | Liu YJ, Liu PY, Long J, Lu Y, Elze L, Recker RR, Deng HW. Linkage and association analyses of the UCP3 gene with obesity phenotypes in Caucasian families. Physiol Genomics. 2005;22:197-203. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 40] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 228. | Otabe S, Clement K, Dina C, Pelloux V, Guy-Grand B, Froguel P, Vasseur F. A genetic variation in the 5’ flanking region of the UCP3 gene is associated with body mass index in humans in interaction with physical activity. Diabetologia. 2000;43:245-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 110] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 229. | Meirhaeghe A, Amouyel P, Helbecque N, Cottel D, Otabe S, Froguel P, Vasseur F. An uncoupling protein 3 gene polymorphism associated with a lower risk of developing Type II diabetes and with atherogenic lipid profile in a French cohort. Diabetologia. 2000;43:1424-1428. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 57] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 230. | Aller R, De Luis DA, Izaola O, González Sagrado M, Conde R, Alvarez T, Pacheco D, Velasco MC. Role of -55CT polymorphism of UCP3 gene on non alcoholic fatty liver disease and insulin resistance in patients with obesity. Nutr Hosp. 2010;25:572-576. [PubMed] |
| 231. | Xu YP, Liang L, Wang CL, Fu JF, Liu PN, Lv LQ, Zhu YM. Association between UCP3 gene polymorphisms and nonalcoholic fatty liver disease in Chinese children. World J Gastroenterol. 2013;19:5897-5903. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 25] [Cited by in RCA: 25] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 232. | Serviddio G, Bellanti F, Tamborra R, Rollo T, Capitanio N, Romano AD, Sastre J, Vendemiale G, Altomare E. Uncoupling protein-2 (UCP2) induces mitochondrial proton leak and increases susceptibility of non-alcoholic steatohepatitis (NASH) liver to ischaemia-reperfusion injury. Gut. 2008;57:957-965. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 183] [Cited by in RCA: 177] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 233. | Jia JJ, Zhang X, Ge CR, Jois M. The polymorphisms of UCP2 and UCP3 genes associated with fat metabolism, obesity and diabetes. Obes Rev. 2009;10:519-526. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 105] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 234. | Fares R, Petta S, Lombardi R, Grimaudo S, Dongiovanni P, Pipitone R, Rametta R, Fracanzani AL, Mozzi E, Craxì A. The UCP2 -866 G& gt; A promoter region polymorphism is associated with nonalcoholic steatohepatitis. Liver Int. 2015;35:1574-1580. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 39] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 235. | Ventura P, Rosa MC, Abbati G, Marchini S, Grandone E, Vergura P, Tremosini S, Zeneroli ML. Hyperhomocysteinaemia in chronic liver diseases: role of disease stage, vitamin status and methylenetetrahydrofolate reductase genetics. Liver Int. 2005;25:49-56. [PubMed] |
| 236. | Ji C, Kaplowitz N. Hyperhomocysteinemia, endoplasmic reticulum stress, and alcoholic liver injury. World J Gastroenterol. 2004;10:1699-1708. [PubMed] |
| 237. | Kim YI. 5,10-Methylenetetrahydrofolate reductase polymorphisms and pharmacogenetics: a new role of single nucleotide polymorphisms in the folate metabolic pathway in human health and disease. Nutr Rev. 2005;63:398-407. [PubMed] |
| 238. | van der Put NM, Gabreëls F, Stevens EM, Smeitink JA, Trijbels FJ, Eskes TK, van den Heuvel LP, Blom HJ. A second common mutation in the methylenetetrahydrofolate reductase gene: an additional risk factor for neural-tube defects? Am J Hum Genet. 1998;62:1044-1051. [PubMed] |
| 239. | Sazci A, Ergul E, Aygun C, Akpinar G, Senturk O, Hulagu S. Methylenetetrahydrofolate reductase gene polymorphisms in patients with nonalcoholic steatohepatitis (NASH). Cell Biochem Funct. 2008;26:291-296. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 28] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 240. | Catalano D, Trovato GM, Ragusa A, Martines GF, Tonzuso A, Pirri C, Buccheri MA, Trovato FM. Non-alcoholic fatty liver disease (NAFLD) and MTHFR 1298A & gt; C gene polymorphism. Eur Rev Med Pharmacol Sci. 2014;18:151-159. [PubMed] |
| 241. | Franco Brochado MJ, Domenici FA, Candolo Martinelli Ade L, Zucoloto S, de Carvalho da Cunha SF, Vannucchi H. Methylenetetrahydrofolate reductase gene polymorphism and serum homocysteine levels in nonalcoholic fatty liver disease. Ann Nutr Metab. 2013;63:193-199. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 20] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 242. | Serin E, Güçlü M, Ataç FB, Verdi H, Kayaselçuk F, Ozer B, Bilezikçi B, Yilmaz U. Methylenetetrahydrofolate reductase C677T mutation and nonalcoholic fatty liver disease. Dig Dis Sci. 2007;52:1183-1186. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 14] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 243. | Koide S, Kugiyama K, Sugiyama S, Nakamura S, Fukushima H, Honda O, Yoshimura M, Ogawa H. Association of polymorphism in glutamate-cysteine ligase catalytic subunit gene with coronary vasomotor dysfunction and myocardial infarction. J Am Coll Cardiol. 2003;41:539-545. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 103] [Cited by in RCA: 102] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 244. | Fernández-Checa JC, Kaplowitz N, García-Ruiz C, Colell A, Miranda M, Marí M, Ardite E, Morales A. GSH transport in mitochondria: defense against TNF-induced oxidative stress and alcohol-induced defect. Am J Physiol. 1997;273:G7-17. [PubMed] |
| 245. | Valenti L, Dongiovanni P, Fargion S, Fracanzani AL. Iron genes, dysmetabolism and fibrosis in chronic hepatitis C. J Hepatol. 2008;48:513-514; author reply 514-515. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 246. | Dongiovanni P, Fracanzani AL, Fargion S, Valenti L. Iron in fatty liver and in the metabolic syndrome: a promising therapeutic target. J Hepatol. 2011;55:920-932. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 221] [Cited by in RCA: 267] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 247. | Bonkovsky HL, Jawaid Q, Tortorelli K, LeClair P, Cobb J, Lambrecht RW, Banner BF. Non-alcoholic steatohepatitis and iron: increased prevalence of mutations of the HFE gene in non-alcoholic steatohepatitis. J Hepatol. 1999;31:421-429. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 251] [Cited by in RCA: 236] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 248. | George DK, Goldwurm S, MacDonald GA, Cowley LL, Walker NI, Ward PJ, Jazwinska EC, Powell LW. Increased hepatic iron concentration in nonalcoholic steatohepatitis is associated with increased fibrosis. Gastroenterology. 1998;114:311-318. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 470] [Cited by in RCA: 447] [Article Influence: 16.6] [Reference Citation Analysis (1)] |
| 249. | Lee SH, Jeong SH, Lee D, Lee JH, Hwang SH, Cho YA, Park YS, Hwang JH, Kim JW, Kim N. An epidemiologic study on the incidence and significance of HFE mutations in a Korean cohort with nonalcoholic fatty liver disease. J Clin Gastroenterol. 2010;44:e154-e161. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 8] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 250. | Nelson JE, Bhattacharya R, Lindor KD, Chalasani N, Raaka S, Heathcote EJ, Miskovsky E, Shaffer E, Rulyak SJ, Kowdley KV. HFE C282Y mutations are associated with advanced hepatic fibrosis in Caucasians with nonalcoholic steatohepatitis. Hepatology. 2007;46:723-729. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 86] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 251. | Bugianesi E, Manzini P, D’Antico S, Vanni E, Longo F, Leone N, Massarenti P, Piga A, Marchesini G, Rizzetto M. Relative contribution of iron burden, HFE mutations, and insulin resistance to fibrosis in nonalcoholic fatty liver. Hepatology. 2004;39:179-187. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 302] [Cited by in RCA: 302] [Article Influence: 14.4] [Reference Citation Analysis (0)] |
| 252. | Raszeja-Wyszomirska J, Kurzawski G, Lawniczak M, Miezynska-Kurtycz J, Lubinski J. Nonalcoholic fatty liver disease and HFE gene mutations: a Polish study. World J Gastroenterol. 2010;16:2531-2536. [PubMed] |
| 253. | Hernaez R, Yeung E, Clark JM, Kowdley KV, Brancati FL, Kao WH. Hemochromatosis gene and nonalcoholic fatty liver disease: a systematic review and meta-analysis. J Hepatol. 2011;55:1079-1085. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 29] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 254. | Valenti L, Canavesi E, Galmozzi E, Dongiovanni P, Rametta R, Maggioni P, Maggioni M, Fracanzani AL, Fargion S. Beta-globin mutations are associated with parenchymal siderosis and fibrosis in patients with non-alcoholic fatty liver disease. J Hepatol. 2010;53:927-933. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 47] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 255. | Ramsay AJ, Hooper JD, Folgueras AR, Velasco G, López-Otín C. Matriptase-2 (TMPRSS6): a proteolytic regulator of iron homeostasis. Haematologica. 2009;94:840-849. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 87] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 256. | Valenti L, Rametta R, Dongiovanni P, Motta BM, Canavesi E, Pelusi S, Pulixi EA, Fracanzani AL, Fargion S. The A736V TMPRSS6 polymorphism influences hepatic iron overload in nonalcoholic fatty liver disease. PLoS One. 2012;7:e48804. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 41] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 257. | Suzuki T, Aizawa K, Matsumura T, Nagai R. Vascular implications of the Krüppel-like family of transcription factors. Arterioscler Thromb Vasc Biol. 2005;25:1135-1141. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 99] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 258. | Ratziu V, Lalazar A, Wong L, Dang Q, Collins C, Shaulian E, Jensen S, Friedman SL. Zf9, a Kruppel-like transcription factor up-regulated in vivo during early hepatic fibrosis. Proc Natl Acad Sci USA. 1998;95:9500-9505. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 189] [Cited by in RCA: 193] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 259. | Miele L, Beale G, Patman G, Nobili V, Leathart J, Grieco A, Abate M, Friedman SL, Narla G, Bugianesi E. The Kruppel-like factor 6 genotype is associated with fibrosis in nonalcoholic fatty liver disease. Gastroenterology. 2008;135:282-291.e1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 151] [Cited by in RCA: 149] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 260. | Bechmann LP, Gastaldelli A, Vetter D, Patman GL, Pascoe L, Hannivoort RA, Lee UE, Fiel I, Muñoz U, Ciociaro D. Glucokinase links Krüppel-like factor 6 to the regulation of hepatic insulin sensitivity in nonalcoholic fatty liver disease. Hepatology. 2012;55:1083-1093. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 54] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 261. | Bechmann LP, Vetter D, Ishida J, Hannivoort RA, Lang UE, Kocabayoglu P, Fiel MI, Muñoz U, Patman GL, Ge F. Post-transcriptional activation of PPAR alpha by KLF6 in hepatic steatosis. J Hepatol. 2013;58:1000-1006. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 45] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 262. | Baker SJ, Reddy EP. Transducers of life and death: TNF receptor superfamily and associated proteins. Oncogene. 1996;12:1-9. [PubMed] |
| 263. | Kim S, Ohta K, Hamaguchi A, Omura T, Yukimura T, Miura K, Inada Y, Ishimura Y, Chatani F, Iwao H. Angiotensin II type I receptor antagonist inhibits the gene expression of transforming growth factor-beta 1 and extracellular matrix in cardiac and vascular tissues of hypertensive rats. J Pharmacol Exp Ther. 1995;273:509-515. [PubMed] |
| 264. | Li B, Khanna A, Sharma V, Singh T, Suthanthiran M, August P. TGF-beta1 DNA polymorphisms, protein levels, and blood pressure. Hypertension. 1999;33:271-275. [PubMed] |
| 265. | Powell EE, Edwards-Smith CJ, Hay JL, Clouston AD, Crawford DH, Shorthouse C, Purdie DM, Jonsson JR. Host genetic factors influence disease progression in chronic hepatitis C. Hepatology. 2000;31:828-833. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 292] [Cited by in RCA: 305] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 266. | Dixon JB, Bhathal PS, Jonsson JR, Dixon AF, Powell EE, O’Brien PE. Pro-fibrotic polymorphisms predictive of advanced liver fibrosis in the severely obese. J Hepatol. 2003;39:967-971. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 69] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 267. | Yoshiji H, Kuriyama S, Yoshii J, Ikenaka Y, Noguchi R, Nakatani T, Tsujinoue H, Fukui H. Angiotensin-II type 1 receptor interaction is a major regulator for liver fibrosis development in rats. Hepatology. 2001;34:745-750. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 290] [Cited by in RCA: 309] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 268. | Zain SM, Mohamed Z, Mahadeva S, Rampal S, Basu RC, Cheah PL, Salim A, Mohamed R. Susceptibility and gene interaction study of the angiotensin II type 1 receptor (AGTR1) gene polymorphisms with non-alcoholic fatty liver disease in a multi-ethnic population. PLoS One. 2013;8:e58538. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 30] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 269. | Yoneda M, Hotta K, Nozaki Y, Endo H, Uchiyama T, Mawatari H, Iida H, Kato S, Fujita K, Takahashi H. Association between angiotensin II type 1 receptor polymorphisms and the occurrence of nonalcoholic fatty liver disease. Liver Int. 2009;29:1078-1085. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 69] [Article Influence: 4.3] [Reference Citation Analysis (0)] |