Published online Aug 21, 2015. doi: 10.3748/wjg.v21.i31.9413
Peer-review started: July 25, 2014
First decision: September 3, 2014
Revised: May 21, 2015
Accepted: June 16, 2015
Article in press: June 16, 2015
Published online: August 21, 2015
Processing time: 391 Days and 16.7 Hours
AIM: To characterize APC gene mutations and correlate them with patient phenotypes in individuals diagnosed with familial adenomatous polyposis (FAP) in northern Brazil.
METHODS: A total of 15 individuals diagnosed with FAP from 5 different families from the north of Brazil were analyzed in this study. In addition to patients with histopathological diagnosis of FAP, family members who had not developed the disease were also tested in order to identify mutations and for possible genetic counseling. All analyzed patients or their guardians signed a consent form approved by the Research Ethics Committee of the João de Barros Barreto University Hospital (Belem, Brazil). DNA extracted from the peripheral blood of a member of each of the affected families was subjected to direct sequencing. The proband of each family was sequenced to identify germline mutations using the Ion Torrent platform. To validate the detected mutations, Sanger sequencing was also performed. The samples from all patients were also tested for the identification of mutations by real-time quantitative polymerase chain reaction using the amplification refractory mutation system.
RESULTS: Through interviews with relatives and a search of medical records, it was possible to construct genograms for three of the five families included in the study. All 15 patients from the five families with FAP exhibited mutations in the APC gene, and all mutations were detected in exon 15 of the APC gene. In addition to the patients with a histological diagnosis of FAP, family members without disease symptoms showed the mutation in the APC gene. In the present study, we detected two of the three most frequent germline mutations in the literature: the mutation at codon 1309 and the mutation at codon 1061. The presence of c.3956delC mutation was found in all families from this study, and suggests that this mutation was introduced in the population of the State of Pará through ancestor immigration (i.e., a de novo mutation that arose in one member belonging to this state from Brazil).
CONCLUSION: Regardless of its origin, the c.3956delC mutation is a strong candidate biomarker of this hereditary cancer syndrome in families of northern Brazil.
Core tip: In the northern region of Brazil, gastrointestinal tumors are the second most frequent type of cancer among men and the third most frequent among women. These tumors are considered a serious public health problem because they are often diagnosed in advanced stages and have extremely low survival rates. Evaluation of family history to determine the number of relatives affected and genetic screening analysis are important preventive measures to assist in the early diagnosis of patients who have not yet developed the disease, as was the case of some patients analyzed in this study.
- Citation: Moreira-Nunes CA, Alcântara DDF&, Lima-Júnior SF, Cavalléro SRA, Rey JA, Pinto GR, Assumpção PP, Burbano RR. Presence of c.3956delC mutation in familial adenomatous polyposis patients from Brazil. World J Gastroenterol 2015; 21(31): 9413-9419
- URL: https://www.wjgnet.com/1007-9327/full/v21/i31/9413.htm
- DOI: https://dx.doi.org/10.3748/wjg.v21.i31.9413
Familial adenomatous polyposis (FAP) is a hereditary cancer predisposition syndrome with autosomal dominant inheritance caused by germline mutations, mainly in the adenomatous polyposis coli (APC) gene. The main clinical feature of FAP is the development, in the second and third decades of life, of multiple (hundreds to thousands) adenomatous polyps in the colon with the capacity for malignant transformation[1-3].
The tumor suppressor gene APC, located on chromosome 5q21, has 16 exons and encodes a 300-kDa protein that participates in the Wnt signaling pathway, which is important in signal transduction and the control of apoptosis[4-6]. APC gene inactivation occurs through allelic loss, primarily through mutations, which generally produces a truncated protein lacking the carboxyl-terminal region and loss of function[7,8].
To date, 858 different mutations in the APC gene have been recorded in the Human Gene Mutation Database (HGMD). Twenty-three percent of the germline alterations of this gene occur between codons 1055 and 1309. The three most common germline mutations are the deletion of 5 base pairs (bp) from codons 1309 and 1061 and the deletion of 4 bp from codon 1064[9].
The FAP phenotype (number of polyps and disease aggressiveness) can be predicted from the APC gene mutation. The FAP phenotypes can be defined as the following: (1) severe/profuse, for mutations located between codons 1250 and 1464; (2) intermediate, for mutations located between codons 158 and 1595, except for mutations located between codons 312 and 412 and for mutations located between codons 1250 and 1464; and (3) attenuated, for mutations located in exon 9 and near the 5’ and 3’ ends of the APC gene[10-16].
In the north and northeast regions of Brazil, gastrointestinal tumors are the second most frequent type of cancer among men and the third most frequent among women. These tumors are considered a serious public health problem because they are often diagnosed in advanced stages and have extremely low survival rates[17].
The aim of this study was to characterize the mutations present in the APC gene, correlate them with patient phenotypes, and evaluate genomic alterations in individuals diagnosed with FAP in northern Brazil.
The study was approved by the Research Ethics Committee of the João de Barros Barreto University Hospital (Belém, Pará, Brazil; approval number: 274/12). All analyzed patients or their guardians signed a consent form, and it was assured that the use of biological material and study participation would not be harmful or negatively influence the patients’ treatment.
A total of 15 patients belonging to 5 different families were analyzed in this study (Table 1). All patients resided in the State of Pará and were assisted at the Coloproctology Outpatient Clinic of the João de Barros Barreto University Hospital (Belém, Pará, Brazil). Peripheral blood samples were collected from all individuals for analysis.
| Patients | Gender | Age at diagnosis | Histopathology |
| Family 1 (FAP1) | |||
| 01A | Female | 23 | FAP |
| 01B | Female | 25 | FAP |
| 01C | Female | 18 | FAP |
| 01D | Male | 14 | FAP |
| 01E | Male | 17 | FAP |
| Family 2 (FAP2) | |||
| 02F | Female | 40 | FAP |
| 02G | Male | 1 | FAP |
| 02H | Female | 15 | FAP |
| 02I | Male | NA2 | - |
| Family 3 (FAP3) | |||
| 03J | Male | 30 | FAP |
| 03H | Female | NA | - |
| 03L | Female | NA | - |
| 03M | Female | NA | - |
| Family 4 (FAP4) | |||
| 04N | Male | 40 | FAP |
| Family 5 (FAP5) | |||
| 05O | Male | 25 | FAP |
In addition to patients with histopathological diagnosis of FAP, family members who had not developed the disease were also tested to identify mutations and for possible genetic counseling in the same manner as provided to members of the FAP2 and FAP3 families (Table 1).
Genomic DNA was extracted from human peripheral blood samples using the QIAamp DNA Blood Kit (Qiagen®), following the manufacturer’s instructions.
The DNA extracted from the peripheral blood of a member of each of the affected families was subjected to direct sequencing.
Direct sequencing of all exons of the APC gene (NM_000038.5) was performed by the next-generation sequencing platform Ion Torrent™ (Life Technologies™), following the Ion AmpliSeq™ Library Preparation methodology (Life Technologies™).
The primers for detection of changes in the nucleotide sequence of the gene were designed through the Ion Torrent™ platform using the Ion AmpliSeq Designer software (available at: http://www.ampliseq.com).
Analysis of the sequenced data was performed using the analysis software available in the Ion Torrent™ platform.
To validate the detected mutations, Sanger sequencing was performed using the BigDye Terminator v1.1 Cycle Sequencing Kit. After ethanol purification, the samples were run on an ABI 3730 sequencer. The chromatogram of the Sanger sequencing results was analyzed using Sequencing Analysis vs 5.2 Program (Applied Biosystem, United States).
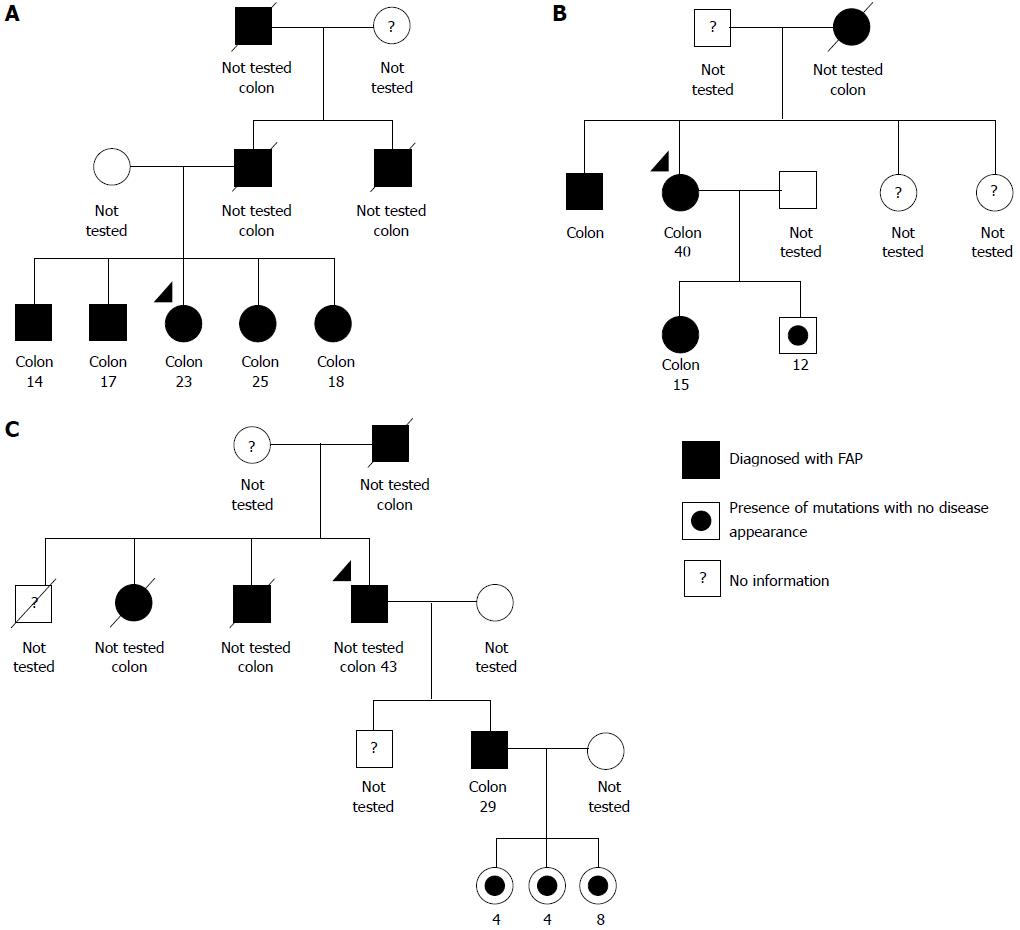
Through interviews with relatives and a search of medical records, it was possible to construct genograms for three of the five families included in the study - FAP1, FAP2, and FAP3 - which, coincidentally, are the families with the largest number of individuals affected by the disease in subsequent generations (Figure 1). The genealogical analysis of the remaining two families was not possible due to a lack of information from patients regarding their relatives.
All 15 patients from the five families with FAP exhibited mutations in the APC gene, and all mutations were detected in exon 15 of the APC gene. The presence of c.3956delC mutation was found in all families from this study. In addition to the patients with a histological diagnosis of FAP, family members without disease symptoms showed the mutation in the APC gene (Table 2).
| Families | Exon | Codon | Mutation | Phenotype1 | Ref. |
| Frameshift mutations | |||||
| FAP1 | 15 | 1061 | c.3183_3187delACAAA | Intermediate | Ficari et al[18], Jarry et al[19] |
| FAP2 | 15 | 1309 | c.3927_3931delAAAGA | Severe | Miyoshi et al[20], Jarry et al[19] |
| FAP5 | 15 | 1309 | c.3921_3925delAAAAG | Severe | Miyaki et al[21], Mulkens et al[22] |
| FAP1-FAP5 | 15 | 1319 | c.3956delC | Severe | Miyaki et al[21] |
FAP is caused by germline mutations primarily in the APC gene and shows autosomal dominant inheritance. The main clinical feature of the disease is the development, in the second and third decades of life, of multiple (hundreds to thousands) adenomatous polyps in the colon and/or rectum capable of malignant transformation from the fourth decade of life[1-3].
Literature reports reveal that most mutations found in FAP (≥ 60%) are located in the central region of the protein (between codons 1281-1556), the so-called “mutation cluster region” (MCR). The MCR corresponds to the region of the APC gene encoding the protein domain responsible for β-catenin regulation[20,23-25]. Mutations that occur in this region generally produce a truncated protein responsible for an increase in the free pool of β-catenin, which is transported to the nucleus, thus activating the transcription of genes involved in cell proliferation[26]. In this study, all mutations discovered in the families with FAP occurred in the MCR region, confirming this region as a hotspot for mutations in the APC gene.
In the present study, we detected two of the three most frequent germline mutations: the mutation at codon 1309 (found in the FAP2 and FAP5 families) and the mutation at codon 1061 (found in the FAP1 family). Half et al[2] found an association between the mutation at codon 1309 and 10% of patients with FAP in the literature, which is in agreement with the study by Torrezan et al[27], who identified this mutation in 9% of the 23 families with FAP in southeastern Brazil. In the present study, the mutation at codon 1309 occurred in all patients with FAP. This frequency is higher than the global average, and, for this reason, we are currently increasing the sample size of families with FAP to assess the validity of this frequency.
The mutation at codon 1061 affects 5% of patients in the literature, according to the survey performed by Half et al[2]. This mutation was found in 20% of the five families with FAP from the State of Pará analyzed in this study, although it was not detected in the studies with families from southeastern Brazil performed by Torrezan et al[27] and De Queiroz Rossanese et al[28]. This result reinforces the difference found between the types of APC gene alterations and the geographic region of Brazil to which the FAP patients belong.
FAP is caused by a highly heterogeneous spectrum of mutations, which are shared among patients from different families, as shown in this study[29,30]. In a comprehensive study performed in a Canadian population by Jarry et al[19], it was demonstrated that several families with FAP exhibited c.3183_3187delACAAA and c.3927_3931delAAAGA mutations, which were also described in this study.
Through direct sequencing of the APC gene, we identified the pathogenic mutation c.3956delC in all families analyzed in this study. This mutation, considered as a frameshift mutation type, and was not found in other Brazilian studies conducted with patients from southeastern Brazil[27,28]. This disparity demonstrates a difference in the spectrum of APC gene alterations in families with FAP according to the geographic region of Brazil. This phenomenon most likely occurs because of miscegenation-related ethnic differences between the populations of the south and north of Brazil[31,32].
Thus, the c.3956delC mutation proved to be an important cause of FAP in northern Brazil and results from a founder effect, most likely originating from Japanese communities, where this mutation was first described by Miyaki et al[21] in colorectal tumors from three unrelated families. To our knowledge, there are no reports describing the c.3956delC mutation in the germline of families with FAP, as shown in this study.
It is likely that either this germline mutation was introduced into the State of Pará after the 1929 landing of Japanese immigrants in Belém[33] or that c.3956delC is a de novo mutation that emerged in a member of the Pará population. Regardless of its origin, the c.3956delC mutation is a strong candidate biomarker for this hereditary cancer syndrome in families of northern Brazil.
According to Crabtree et al[34], the presence of a mutation in the MCR region confers to FAP the severe phenotype, regardless of the contribution of mutations characteristic of other phenotypes. In this study, this assumption held because all families with FAP had in common the presence of mutation at codon 1319, in the MCR region.
The genotype-phenotype relationship in FAP is a determining factor for clinical guidance and genetic counseling, as well as for simplifying the search for mutations in patients with FAP and their relatives[14,15,35].
The diagnosis of severe FAP in the study population occurred around the second and third decade of life, while intermediate FAP was diagnosed in the fourth decade of life. This timeline is in accordance with the guidelines for the clinical management of FAP published by Vasen et al[36], who demonstrated that the onset of the disease in patients with the severe form of FAP occurred, on average, 10 years before the onset in carriers of intermediate and attenuated forms. As an example, in this study the FAP1 family exhibited the early form of the disease, and one of the members had the syndrome diagnosed at 14 years of age, with the presence of thousands of polyps throughout the intestine. This aggressive phenotype is justified by the presence of the c.3956delC mutation, associated with the severe phenotype. For this reason, and in accordance with the abovementioned guidelines, clinical and diagnostic evaluations should be performed in all FAP family members, starting at the second decade of life.
In conclusion, the presence of the c.3956delC mutation in all families studied leads us to believe that this mutation was introduced in the population of the State of Pará through the immigration of ancestors (i.e., a de novo mutation that emerged in a member belonging to this State). Regardless of its origin, the c.3956delC mutation is a strong candidate biomarker of this hereditary cancer syndrome in families of northern Brazil.
It is noteworthy that different germline mutations were found in families of different ethnic groups. Therefore, there is a need to identify the alterations responsible for hereditary cancer syndromes in the Brazilian population, especially in populations of the northern and northeastern regions, where a high incidence of gastrointestinal tumors is observed.
The main clinical feature of familial adenomatous polyposis (FAP) is the development, in the second and third decades of life, of multiple (hundreds to thousands) adenomatous polyps in the colon with the capacity for malignant transformation. It is diagnosed when a person develops more than 100 adenomatous colon polyps. An adenomatous polyp is an area where normal cells that line the inside of the colon become mucous and form a mass on the inside of the intestinal tract. The average age for polyps to develop in people with FAP is in the mid-teens. More than 95% of individuals with FAP will have multiple colon polyps by 35 years of age.
The incidence of FAP is associated with mutations in the q21-q22 region of the long arm of chromosome 5 in 80% of patients. From then onwards, large deletions in the APC gene germline have been reported in families with FAP from different geographic regions. Screening of family members of patients with FAP should begin by 12 years of age. Genetic testing for germline mutations may eliminate the need for surveillance in some family members. Visualization of more than 100 polyps usually establishes the diagnosis because of the diffuse nature of the polyposis.
Cancers of the rectum in patients who have had subtotal colectomy with ileorectal anastomosis have been reported with sulindac and celecoxib therapy. Because of the inability to control polyps medically, eventual rectal resection is usually necessary.
The mutations found in this study have been extensively described in patients with colorectal malignancies of different ethnic groups and geographic regions, mainly Asian and European populations. This fact reinforces the role of miscegenation in our study population in the appearance of several germline mutations in patients with FAP.
Next-generation sequencing: refers to non-Sanger-based high-throughput DNA sequencing technologies. Millions or billions of DNA strands can be sequenced in parallel, yielding substantially more throughput and minimizing the need for the fragment-cloning methods that are often used in Sanger sequencing of genomes.
FAP is caused by a highly heterogeneous spectrum of mutations, which are shared among patients from different families, as shown in this study. The identification of alterations responsible for the onset of hereditary gastrointestinal cancer syndrome in patients allows the assessment, through molecular techniques, of whether their relatives are carriers of these alterations and of their risk of developing the syndrome. The genotype-phenotype relationship in FAP is a determining factor for clinical guidance and genetic counseling, as well as for simplifying the search for mutations in patients with FAP and their relatives.
The authors found that the c.3956delC mutation is a strong candidate biomarker of this hereditary cancer syndrome in families of northern Brazil. It is noteworthy that different germline mutations were found in families of different ethnic groups. Therefore, there is a need to identify the alterations responsible for hereditary cancer syndromes in the Brazilian population, especially in populations of the northern and northeastern regions, where a high incidence of gastrointestinal tumors is observed.
P- Reviewer: Zhao YS S- Editor: Yu J L- Editor: Rutherford A E- Editor: Zhang DN
| 1. | Pineda M, González S, Lázaro C, Blanco I, Capellá G. Detection of genetic alterations in hereditary colorectal cancer screening. Mutat Res. 2010;693:19-31. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 29] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 2. | Half E, Bercovich D, Rozen P. Familial adenomatous polyposis. Orphanet J Rare Dis. 2009;4:22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 308] [Cited by in RCA: 374] [Article Influence: 23.4] [Reference Citation Analysis (0)] |
| 3. | Hosogi H, Nagayama S, Kanamoto N, Yoshizawa A, Suzuki T, Nakao K, Sakai Y. Biallelic APC inactivation was responsible for functional adrenocortical adenoma in familial adenomatous polyposis with novel germline mutation of the APC gene: report of a case. Jpn J Clin Oncol. 2009;39:837-846. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 15] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 4. | Polakis P. The adenomatous polyposis coli (APC) tumor suppressor. Biochim Biophys Acta. 1997;1332:F127-F147. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 159] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 5. | Fearnhead NS, Britton MP, Bodmer WF. The ABC of APC. Hum Mol Genet. 2001;10:721-733. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 635] [Cited by in RCA: 622] [Article Influence: 25.9] [Reference Citation Analysis (0)] |
| 6. | Virmani AK, Rathi A, Sathyanarayana UG, Padar A, Huang CX, Cunnigham HT, Farinas AJ, Milchgrub S, Euhus DM, Gilcrease M. Aberrant methylation of the adenomatous polyposis coli (APC) gene promoter 1A in breast and lung carcinomas. Clin Cancer Res. 2001;7:1998-2004. [PubMed] |
| 7. | Toyota M, Ahuja N, Suzuki H, Itoh F, Ohe-Toyota M, Imai K, Baylin SB, Issa JP. Aberrant methylation in gastric cancer associated with the CpG island methylator phenotype. Cancer Res. 1999;59:5438-5442. [PubMed] |
| 8. | Lesko AC, Goss KH, Prosperi JR. Exploiting APC function as a novel cancer therapy. Curr Drug Targets. 2014;15:90-102. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 44] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 9. | Plawski A, Banasiewicz T, Borun P, Kubaszewski L, Krokowicz P, Skrzypczak-Zielinska M, Lubinski J. Familial adenomatous polyposis of the colon. Hered Cancer Clin Pract. 2013;11:15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 32] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 10. | Newton CR, Graham A, Heptinstall LE, Powell SJ, Summers C, Kalsheker N, Smith JC, Markham AF. Analysis of any point mutation in DNA. The amplification refractory mutation system (ARMS). Nucleic Acids Res. 1989;17:2503-2516. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1716] [Cited by in RCA: 1749] [Article Influence: 48.6] [Reference Citation Analysis (0)] |
| 11. | Bunyan DJ, Shea-Simonds J, Reck AC, Finnis D, Eccles DM. Genotype-phenotype correlations of new causative APC gene mutations in patients with familial adenomatous polyposis. J Med Genet. 1995;32:728-731. [PubMed] |
| 12. | Soravia C, Berk T, Madlensky L, Mitri A, Cheng H, Gallinger S, Cohen Z, Bapat B. Genotype-phenotype correlations in attenuated adenomatous polyposis coli. Am J Hum Genet. 1998;62:1290-1301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 227] [Cited by in RCA: 209] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 13. | Wallis YL, Morton DG, McKeown CM, Macdonald F. Molecular analysis of the APC gene in 205 families: extended genotype-phenotype correlations in FAP and evidence for the role of APC amino acid changes in colorectal cancer predisposition. J Med Genet. 1999;36:14-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 14. | Groves C, Lamlum H, Crabtree M, Williamson J, Taylor C, Bass S, Cuthbert-Heavens D, Hodgson S, Phillips R, Tomlinson I. Mutation cluster region, association between germline and somatic mutations and genotype-phenotype correlation in upper gastrointestinal familial adenomatous polyposis. Am J Pathol. 2002;160:2055-2061. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 87] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 15. | Vandrovcová J, Stekrová J, Kebrdlová V, Kohoutová M. Molecular analysis of the APC and MYH genes in Czech families affected by FAP or multiple adenomas: 13 novel mutations. Hum Mutat. 2004;23:397. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 28] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 16. | Nieuwenhuis MH, Vasen HF. Correlations between mutation site in APC and phenotype of familial adenomatous polyposis (FAP): a review of the literature. Crit Rev Oncol Hematol. 2007;61:153-161. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 260] [Cited by in RCA: 245] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 17. | Silva JAG. Estimate/2012-Cancer Incidence in Brazil. From National Cancer Institute. Rio de Janeiro. 2011; Available from: http://www.inca.gov.br/estimativa/2012. |
| 18. | Ficari F, Cama A, Valanzano R, Curia MC, Palmirotta R, Aceto G, Esposito DL, Crognale S, Lombardi A, Messerini L. APC gene mutations and colorectal adenomatosis in familial adenomatous polyposis. Br J Cancer. 2000;82:348-353. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 64] [Cited by in RCA: 59] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 19. | Jarry J, Brunet JS, Laframboise R, Drouin R, Latreille J, Richard C, Gekas J, Maranda B, Monczak Y, Wong N. A survey of APC mutations in Quebec. Fam Cancer. 2011;10:659-665. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 10] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 20. | Miyoshi Y, Ando H, Nagase H, Nishisho I, Horii A, Miki Y, Mori T, Utsunomiya J, Baba S, Petersen G. Germ-line mutations of the APC gene in 53 familial adenomatous polyposis patients. Proc Natl Acad Sci USA. 1992;89:4452-4456. [PubMed] |
| 21. | Miyaki M, Konishi M, Kikuchi-Yanoshita R, Enomoto M, Igari T, Tanaka K, Muraoka M, Takahashi H, Amada Y, Fukayama M. Characteristics of somatic mutation of the adenomatous polyposis coli gene in colorectal tumors. Cancer Res. 1994;54:3011-3020. [PubMed] |
| 22. | Mulkens J, Poncin J, Arends JW, De Goeij AF. APC mutations in human colorectal adenomas: analysis of the mutation cluster region with temperature gradient gel electrophoresis and clinicopathological features. J Pathol. 1998;185:360-365. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 23. | Cheadle JP, Krawczak M, Thomas MW, Hodges AK, Al-Tassan N, Fleming N, Sampson JR. Different combinations of biallelic APC mutation confer different growth advantages in colorectal tumours. Cancer Res. 2002;62:363-366. [PubMed] |
| 24. | De Rosa M, Scarano MI, Panariello L, Morelli G, Riegler G, Rossi GB, Tempesta A, Romano G, Renda A, Pettinato G. The mutation spectrum of the APC gene in FAP patients from southern Italy: detection of known and four novel mutations. Hum Mutat. 2003;21:655-656. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 29] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 25. | Christie M, Jorissen RN, Mouradov D, Sakthianandeswaren A, Li S, Day F, Tsui C, Lipton L, Desai J, Jones IT. Different APC genotypes in proximal and distal sporadic colorectal cancers suggest distinct WNT/β-catenin signalling thresholds for tumourigenesis. Oncogene. 2013;32:4675-4682. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 104] [Cited by in RCA: 112] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 26. | Minde DP, Anvarian Z, Rüdiger SG, Maurice MM. Messing up disorder: how do missense mutations in the tumor suppressor protein APC lead to cancer? Mol Cancer. 2011;10:101. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 114] [Cited by in RCA: 129] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 27. | Torrezan GT, da Silva FC, Santos EM, Krepischi AC, Achatz MI, Aguiar S, Rossi BM, Carraro DM. Mutational spectrum of the APC and MUTYH genes and genotype-phenotype correlations in Brazilian FAP, AFAP, and MAP patients. Orphanet J Rare Dis. 2013;8:54. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 37] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 28. | De Queiroz Rossanese LB, De Lima Marson FA, Ribeiro JD, Coy CS, Bertuzzo CS. APC germline mutations in families with familial adenomatous polyposis. Oncol Rep. 2013;30:2081-2088. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 14] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 29. | Powell SM, Zilz N, Beazer-Barclay Y, Bryan TM, Hamilton SR, Thibodeau SN, Vogelstein B, Kinzler KW. APC mutations occur early during colorectal tumorigenesis. Nature. 1992;359:235-237. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1308] [Cited by in RCA: 1301] [Article Influence: 39.4] [Reference Citation Analysis (0)] |
| 30. | Goss KH, Groden J. Biology of the adenomatous polyposis coli tumor suppressor. J Clin Oncol. 2000;18:1967-1979. [PubMed] |
| 31. | Batista dos Santos SE, Rodrigues JD, Ribeiro-dos-Santos AK, Zago MA. Differential contribution of indigenous men and women to the formation of an urban population in the Amazon region as revealed by mtDNA and Y-DNA. Am J Phys Anthropol. 1999;109:175-180. [PubMed] |
| 32. | Pena SD, Bastos-Rodrigues L, Pimenta JR, Bydlowski SP. DNA tests probe the genomic ancestry of Brazilians. Braz J Med Biol Res. 2009;42:870-876. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 133] [Cited by in RCA: 130] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 33. | Jordan CF. Working with Nature: Resource Management for Sustainability. Amsterdam: Harwood Academic Publishers 1998; . |
| 34. | Crabtree MD, Fletcher C, Churchman M, Hodgson SV, Neale K, Phillips RK, Tomlinson IP. Analysis of candidate modifier loci for the severity of colonic familial adenomatous polyposis, with evidence for the importance of the N-acetyl transferases. Gut. 2004;53:271-276. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 35. | Munck A, Gargouri L, Alberti C, Viala J, Peuchmaur M, Lenaerts C, Michaud L, Lamireau T, Mougenot JF, Dabadie A. Evaluation of guidelines for management of familial adenomatous polyposis in a multicenter pediatric cohort. J Pediatr Gastroenterol Nutr. 2011;53:296-302. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 18] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 36. | Vasen HF, Möslein G, Alonso A, Aretz S, Bernstein I, Bertario L, Blanco I, Bülow S, Burn J, Capella G. Guidelines for the clinical management of familial adenomatous polyposis (FAP). Gut. 2008;57:704-713. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 564] [Cited by in RCA: 470] [Article Influence: 27.6] [Reference Citation Analysis (0)] |