Published online Apr 28, 2015. doi: 10.3748/wjg.v21.i16.4864
Peer-review started: October 11, 2014
First decision: October 29, 2014
Revised: November 24, 2014
Accepted: December 14, 2014
Article in press: December 16, 2014
Published online: April 28, 2015
Processing time: 200 Days and 16.3 Hours
AIM: To study the inflammatory microenvironment and expression of chemokines in hepatocellular carcinoma (HCC) in nude mice.
METHODS: CBRH-7919 HCC cells were injected into the subcutaneous region of nude mice. Beginning two weeks after the challenge, tumor growth was measured every week for six weeks. The stromal microenvironment and inflammatory cell infiltration was assessed by immunohistochemistry in paired tumor and adjacent peritumoral samples, and macrophage phenotype was assessed using double-stain immunohistochemistry incorporating expression of an intracellular enzyme. A chemokine PCR array, comprised of 98 genes, was used to screen differential gene expressions, which were validated by Western blotting. Additionally, expression of identified chemokines was knocked-down by RNA interference, and the effect on tumor growth was assessed.
RESULTS: Inflammatory cell infiltrates are a key feature of adjacent peritumoral tissues with increased macrophage, neutrophil, and T cell (specifically helper and activated subsets) infiltration. Macrophages within adjacent peritumoral tissues express inducible nitric oxide synthase, suggestive of a proinflammatory phenotype. Fifty-one genes were identified in tumor tissues during the progression period, including 50 that were overexpressed (including CXCL1, CXCL2 and CXCL3) and three that were underexpressed (CXCR1, Ifg and Actb). RNA interference of CXCL1 in the CBRH-7919 cells decreased the growth of tumors in nude mice and inhibited expression of CXCL2, CXCL3 and interleukin-1β protein.
CONCLUSION: These findings suggest that CXCL1 plays a critical role in tumor growth and may serve as a potential molecular target for use in HCC therapy.
Core tip: An orthotopic transplantation tumor model of hepatocellular carcinoma (HCC) with CBRH-7919 cells was established. Inflammatory cell infiltration and macrophage phenotype were assessed by immunohistochemistry. A chemokine PCR array was used to identify differentially expressed genes, and tumor growth was assessed after knockdown with RNA interference. This study describes the inflammatory microenvironment and differential expression of chemokines in hepatocellular carcinoma. The data suggest that CXCL1 plays a critical role in tumor growth and may serve as a potential molecular target for use in HCC therapy.
- Citation: Han KQ, He XQ, Ma MY, Guo XD, Zhang XM, Chen J, Han H, Zhang WW, Zhu QG, Nian H, Ma LJ. Inflammatory microenvironment and expression of chemokines in hepatocellular carcinoma. World J Gastroenterol 2015; 21(16): 4864-4874
- URL: https://www.wjgnet.com/1007-9327/full/v21/i16/4864.htm
- DOI: https://dx.doi.org/10.3748/wjg.v21.i16.4864
Hepatocellular carcinoma (HCC) is one of the most common tumors worldwide, and the fifth leading cause of cancer-related deaths[1,2] due to its rapid growth, early metastasis, and relationships with chronic hepatitis. At the time of diagnosis, surgical resection remains the most effective treatment for early disease. However, more than 75% of patients relapse within five years, and the overall survival for HCC patients has not yet been improved[3].
Recent studies have indicated that the tumor inflammatory microenvironment plays an essential role in the progression of HCC. The tumor microenvironment plays a critical role in modulating the process of liver fibrosis, hepatocarcinogenesis, epithelial-mesenchymal transition, tumor invasion and metastasis[4-8]. The tumor microenvironment consists of: (1)hepatic stellate cells, fibroblasts, immune cells (including regulatory and cytotoxic T cells and tumor-associated macrophages), and endothelial cells; (2) growth actors (including transforming growth factor [TGF]-1 and platelet-derived growth factor); (3) proteolytic enzymes (such as matrix metalloproteinases and tissue inhibitor of metalloproteinases); and (4) extracellular matrix proteins and inflammatory cytokines. These play a critical role in HCC development, tumor control and response to treatment[9,10].
The aim of this study was to elucidate the mechanisms that underlie chronic inflammation in HCC disease progression in order to identify potential therapeutic targets. We also discuss the current understanding of each component of the tumor microenvironment and their roles in the pathogenesis of HCC. Thus, understanding the inflammatory microenvironment is critical to promote understanding of the molecular, cellular and pathophysiological mechanisms of HCC, and is essential for the development of new therapeutic strategies. Nevertheless, to date, the inflammatory microenvironment and differential expression pattern of chemokines in hepatocellular carcinoma is still not clear.
In this study, infiltration of inflammatory cells in the stromal microenvironment was assessed by immunohistochemistry in paired tumor and adjacent peritumoral samples. Macrophage phenotype was assessed using double-staining immunohistochemistry, incorporating expression of an intracellular enzyme. PCR array analysis was used to evaluate the expression profiles of chemokines and their receptors in subcutaneous CBRH-7919 cell xenograft tumors[11] and peritumoral tissues. The expression of chemokines identified by the PCR array were verified by Western blotting and immunohistochemistry. Additionally, knockdown of chemokines by RNA interference (RNAi) was used to assess the effect on tumor growth.
The human hepatocellular carcinoma cell line CBRH-7919 (Chinese Academy of Science, Shanghai, China) was used in this study. Cells were cultured at 37 °C in a humidified atmosphere with 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM; Gibco of Thermo Fisher Scientific, Waltham, MA, United States) supplemented with 10% fetal bovine serum, 100 mg/mL penicillin G, and 50 μg/mL streptomycin (Life Technologies of Thermo Fisher Scientific).
Male Balb/c nude mice were obtained from Laboratory Animal Service Center of the Medical College of Shanghai. All mice were maintained under specific pathogen free conditions and had free access to sterilized food and autoclaved water. These experimental procedures were approved by the Animal Ethics Committee of Shanghai University of Traditional Chinese Medicine.
Male Balb/c nude mice (4 wk of age, 15-18 g) were subcutaneously injected with 0.1 mL of a CBRH-7919 cell suspension (1 × 107 cells) using a 21-gauge needle. Mice were observed after two weeks, and their tumors were excised, weighed, and measured. A portion of the tumor tissue was fixed in 10% formalin for subsequent histologic examination, and the remaining tissue was snap frozen in liquid nitrogen and stored at -70 °C for molecular studies.
Inflammatory cell phenotype was assessed by immunohistochemistry on three tumor tissues and adjacent normal biopsies. Tissue was stained using the Envision+ biotin-free system[12] incorporating either the Envision+ peroxidase-linked biotin-free system (K5007) or CSA II biotin-free tyramide signal amplification (K1497; Dako of Agilent Technologies, Santa Clara, CA, United States) depending on antibody requirements. One microscopic high-powered field (HPF) from each sample was digitally imaged (× 640 magnification) with normal tissue as representative of tissue type, distinct from the area of most positive staining. The number of positive cells was counted to give a score of inflammatory cellular infiltrate.
Macrophage phenotype was also assessed in three pairs of tumor and adjacent tissue biopsies as described above. A double-stain immunohistochemical technique was used, incorporating detection of an intracellular enzymatic marker of macrophage function, namely [inducible nitric oxide synthase (iNOS), a proinflammatory classically activated macrophage) or arginase I (alternatively activated macrophage][13,14], identified by peroxidase-linked immunoreactivity. The area of most positive macrophage infiltration within one HPF (× 680 magnification), distinct from lymphoid aggregation, was identified under fluorescent light at 580 nm using a Texas red filter set and digitally imaged, and also captured under standard optical light. The two images were imported into Corel-Paint X3 (version 13; Corel Corp., Ottawa, CA) to assess macrophage infiltrate.
Formalin-fixed tumors were embedded in paraffin, and 4 μm sections were cut and stained with hematoxylin and eosin (HE).
HCC tissue samples were dissolved with 1 mL of Trizol reagent (Invitrogen of Thermo Fisher Scientific) and homogenized; the sample volume did not exceed 10% of the volume of Trizol reagent. The homogenized samples were incubated at 15-30 °C for 5 min in clear polypropylene tubes to permit the complete dissociation of nucleoprotein complexes. Chloroform (0.2 mL) was added to the tube. After vigorous shaking, the mixture was incubated again at 15-30 °C for 2-3 min. After centrifugation at 12000 ×g for 15 min at 4 °C, the RNA in the aqueous phase was moved to a fresh RNA-free tube and mixed with 0.5 mL isopropyl alcohol. The samples were incubated at 15-30 °C for 10 min and centrifuged at 12000 ×g for 10 min at 4 °C. The supernatant was removed and the RNA pellet was washed once with 75% ethanol, redissolved in RNase-free water, and stored at -70 °C. All samples were treated with MinElute (Qiagen, Vinlo, Netherlands) to remove residual DNA. The quality of the RNA was analyzed on an RNA chip by means of a bioanalyzer (model 2100; Agilent Technologies); the 260/280 ratio of array-tested RNA was 1.8-2.0.
Files were extracted from Agilent Feature Extraction Software (version 9.5.3) and imported into the Agilent GeneSpring GX software (version 7.3 or later) for further analysis. The microarray datasets were normalized in GeneSpring GX using the Agilent FE one-color scenario (mainly median normalization). The positive effect of this median normalization is illustrated in a box-plot, and genes marked present (“All Targets Value”) were chosen for data analysis. Finally, a fold-change analysis was carried out by calculating the ratio between the treatment and the control to identify differentially expressed genes. A cutoff value of twofold change was used; genes with expression levels that differed by at least twofold from the mean in at least one sample were selected for further evaluation. The gene expression profiling data complied with the Minimum Information About Microarray Experiments standard. The microarray experiment was completed by Shanghai KangChen Bio-tech Company (Shanghai, China).
Paraffin blocks were cut (6 μm sections), and sections were deparaffinized followed by antigen retrieval using citric acid buffer (pH 6.0, 95 °C for 15 min). Slides were treated with 3% hydrogen peroxide in methanol to block endogenous peroxidase activity. After 20 min of blocking in 1% bovine serum albumin (BSA), the slides were incubated overnight at 4 °C with anti-human CXCL1 (Ab86436), CXCL2 (Ab25130), CXCL3 (Ab10064), and CXCR1 (Ab60254) antibodies (all goat polyclonal antibodies, 1:250 in 1% BSA; Abcam, Cambridge, United Kingdom). Next, the slides were incubated with 2 μg/mL of biotinylated anti-goat IgG secondary antibody (Vector Laboratories, Burlingame, CA) for 40 min at room temperature. Subsequently, the sections were stained using Standard Ultra-Sensitive ABC Peroxidase Staining kit (Pierce of Thermo Fisher Scientific) and 3,3′-diaminobenzidine (Vector Laboratories), and counterstained by hematoxylin. Mouse xenograft tumors from the hepatocellular cancer cell line CBRH-7919, known to stain strongly for CXCL1, CXCL2, CXCL3 and CXCR1[15] were used as a positive control.
HCC cell extracts from tumor and peritumoral tissues were analyzed using antibodies against CXCL1, CXCL2, CXCL3 and their receptor CXCR1. Equal amounts of protein were subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis and the intracellular amount of GAPDH was analyzed as a loading control. Finally, the immunoreactive bands were developed on X-ray film using an ECL Western Blotting Analysis System (Amersham, Buckinghamshire, United Kingdom) according to the manufacturer’s instructions.
Total RNA was isolated as described above, and the concentration was measured by using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific). RNA was converted to cDNA using the RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific). The level of CXCL1 mRNA expression was evaluated by qRT-PCR performed using DyNAmo ColorFlash SYBR Green qPCR Kit on an ABI 7300 system (Applied Biosystems of Thermo Fisher Scientific). The following primers were used: CXCL1, 5′-TAGAAGGTGTTGAGCGGGAAG-3′ (sense) and 5′-TGAGACGAGAAGGAGCATTGG-3′ (antisense); GAPDH, 5′-GTCGGTGTGAACGGATTTG-3′ (sense) and 5′-TCCCATTCTCAGCCTTGAC-3′ (antisense).
The siRNAs against CXCL1 were designed and ordered from Shanghai GenePharma Co., Ltd. CBRH-7919 cells were transfected with the siRNAs using Lipofectamine RNAi Max (Invitrogen) according to manufacturer’s protocol. Cells were incubated for 48 h, and knockdown efficiency was determined by both qRT-PCR and Western blot analysis. The siRNA with the sequence 5’-GTCTCAGGACAGAGAAGTT-3’ showed the highest efficiency in the knockdown of CXCL1 and was used in this study. Western blot analysis was used to assess the expression of CXCL2, CXCL3 and interleukin (IL)-1β.
Data are summarized as mean ± SE. All statistical calculations were performed using either Student’s t or Wilcoxon’s rank sum tests performed with SPSS version 18 (SPSS Inc., Chicago, IL, United States). A P < 0.05 was considered statistically significant. The statistical methods of this study were reviewed by Wenjun Li.
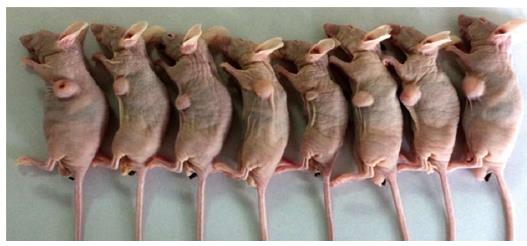
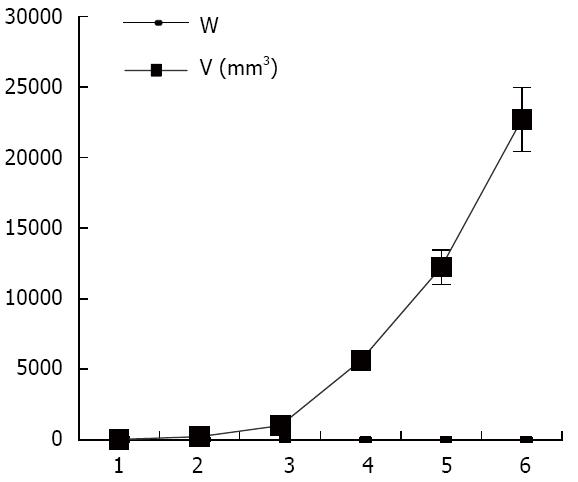
As shown in Figure 1, the tumor growth developed rapidly between weeks 1-6 after xenograft tumor transplantation. The mice were evaluated for the formation of a tumor every week. Fourteen days after challenge with the CBRH-7919 cells, a local tumor was initially observed in the subcutaneous abdominal region of the mice (Figure 2).
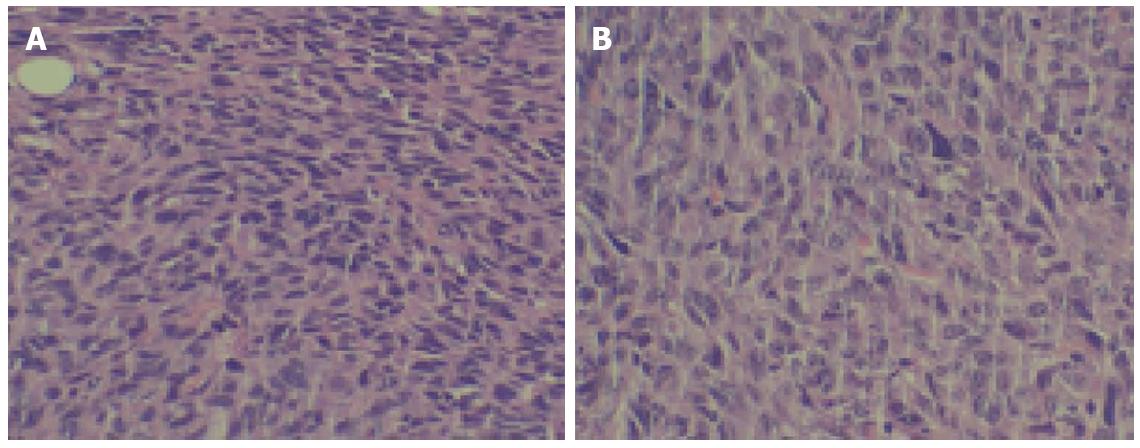
HE staining confirmed tumorigenesis. Tumor tissue characteristics included atypia, a large core, darker colors, various cell sizes, irregular shapes, and nuclear division. Many of these characteristics were similar to those of the original CBRH-7919 cells (Figure 3).
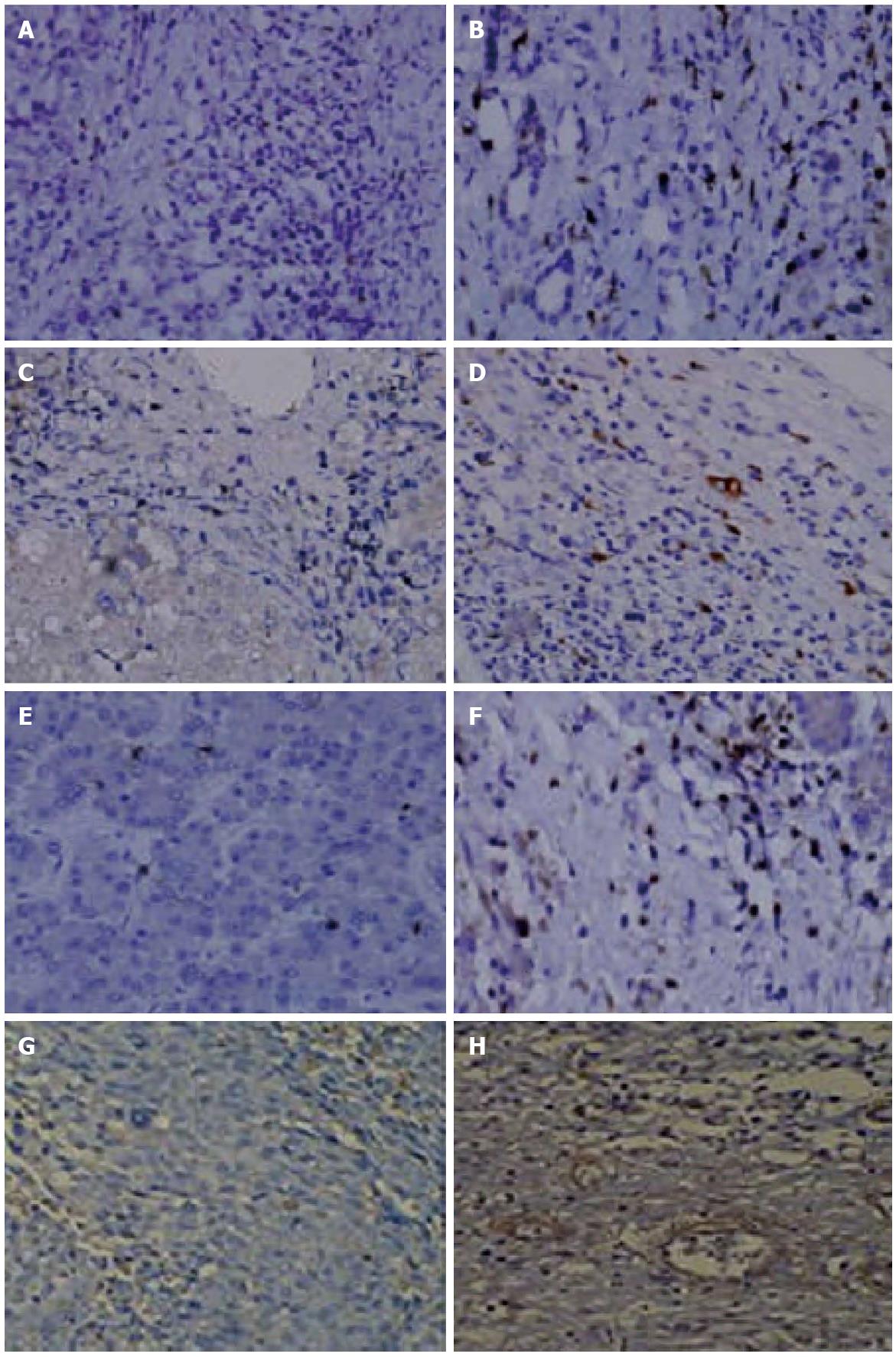
Macrophage, helper T cells, activated T cells, and neutrophils were significantly increased in peritumoral tissue compared to tumor tissue (Ps < 0.005) (Table 1). There were no iNOS+ proinflammatory macrophages in the tumor tissue, whereas an average of 10 cells (range: 6-16) with positive expression were found within peritumoral tissue. The median number of arginase I+ cells within the tumor and peritumoral tissues were 1 (range: 0-2) and 0 (range: 0-1), respectively (P < 0.005). The relative proportion of regulatory to proinflammatory macrophages was higher in peritumoral tissue (Figure 4).
| Inflammatory cell | Tumor sample | Peritumoral samples | Mean difference | (95%CI) | P |
| T helper cell | 12.09 ± 10.74 | 20.08 ± 13.31 | 7.53 | (2.75-13.31) | 0.005 |
| Cytotoxic T cell | 7.25 ± 8.07 | 6.05 ± 8.41 | 11.87 | (21.60-2.14) | 0.800 |
| B cell | 2.88 ± 6.36 | 4.31 ± 6.87 | 10.87 | (19.60-4.07) | 0.600 |
| Activated T cell | 2.12 ± 2.37 | 6.96 ± 5.17 | 5.10 | (3.17-7.02) | 0.005 |
| NK cell | 0.29 ± 1.30 | 2.30 ± 5.18 | 1.83 | (0.21-3.45) | 0.060 |
| Macrophage | 9.03 ± 9.97 | 19.76 ± 9.41 | 8.07 | (4.00-12.13) | 0.005 |
| Mast cell | 9.00 ± 5.35 | 10.44 ± 9.00 | 0.78 | (21.07-3.09) | 0.600 |
| Neutrophil | 1.56 ± 3.22 | 14.83 ± 14.76 | 13.28 | (8.37-18.19) | 0.005 |
| Plasma cell | 5.59 ± 7.11 | 6.50 ± 8.00 | 0.67 | (20.16-4.14) | 0.800 |
| iNOS+ cells median (range) | 0 (0-3) | 10 (6-16) | - | - | 0.005 |
| Arginase I+ cells median (range) | 1 (0-2) | 0 (0-1) | - | - | 0.005 |
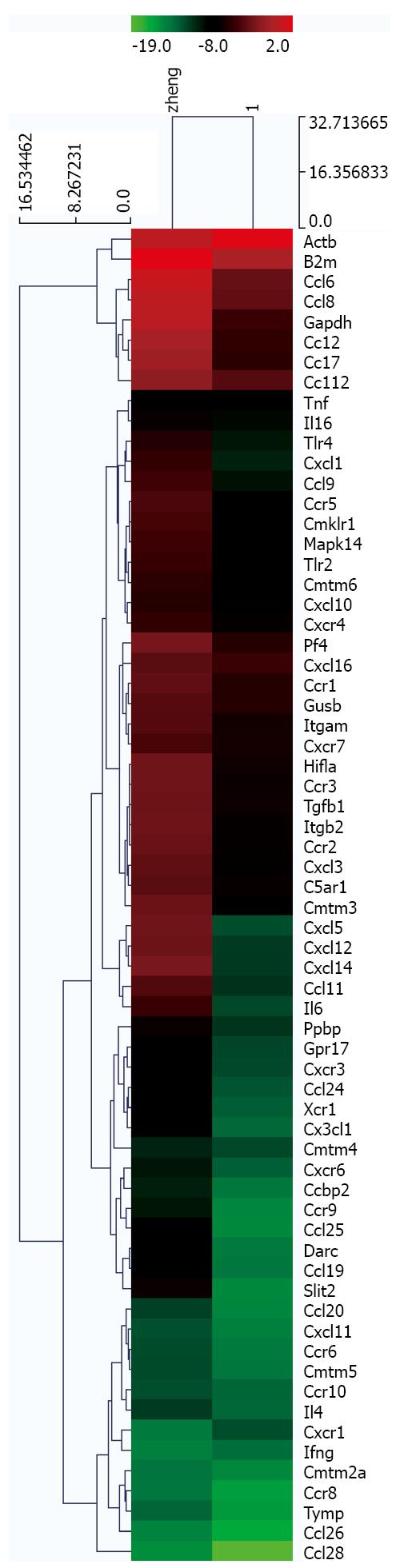
The expressions of chemokine-related genes during HCC progression were evaluated in tumor and peritumoral tissues using a chemokine PCR array. A total of 50 genes were identified as upregulated, including CXCL1, CXCL2, CXCL3 and IL-1β, whereas CXCR1, Ifg and Actb were downregulated in tumor tissues (Tables 2 and 3; Figure 5).
| Accession No. | Gene name | Gene symbol | Fold change |
| A01 | C5aR | C5ar1 | 8.86 |
| A02 | CCR10 | Ccbp2 | 5.39 |
| A04 | Scya11 | Ccl11 | 97.95 |
| A05 | Scya12 | Ccl12 | 4.75 |
| A07 | CKb11 | Ccl19 | 71.66 |
| A08 | HC11 | Ccl2 | 25.81 |
| A11 | CKb-6 | Ccl24 | 17.46 |
| B06 | MRP-1 | Ccl6 | 20.28 |
| B07 | MCP-3 | Ccl7 | 22.47 |
| B08 | MCP-2 | Ccl8 | 16.51 |
| B09 | MRP-2 | Ccl9 | 20.72 |
| B10 | Cmkbr1 | Ccr1 | 5.2 |
| B11 | Cmkbr9 | Ccr10 | 2.01 |
| C01 | Ccr2a | Ccr2 | 14.64 |
| C02 | CKR3 | Ccr3 | 14.14 |
| C04 | CD195 | Ccr5 | 9.77 |
| C05 | CCR-6 | Ccr6 | 4.4 |
| C11 | mcmklrl | Cmklr1 | 8.96 |
| C12 | Cklf | Cmtm2a | 2.25 |
| D01 | Cklfsf3 | Cmtm3 | 28.58 |
| D02 | Cklfsf4 | Cmtm4 | 2.94 |
| D03 | Cklfsf5 | Cmtm5 | 4.14 |
| D04 | Cklfsf6 | Cmtm6 | 5.27 |
| D05 | Cxc3 | Cx3cl1 | 32.58 |
| D07 | KC | Cxcl1 | 23.49 |
| D08 | C7 | Cxcl10 | 3.15 |
| D09 | Cxc11 | Cxcl11 | 4.84 |
| D10 | Pbsf | Cxcl12 | 246.94 |
| D12 | KS1 | Cxcl14 | 323.14 |
| E02 | Zmynd15 | Cxcl16 | 2.3 |
| E04 | Gm1960 | Cxcl3 | 12.76 |
| E05 | LIX | Cxcl5 | 458.57 |
| E09 | Cd183 | Cxxr3 | 12.42 |
| E10 | CD184 | Cxcr4 | 3.16 |
| E12 | B0NZ0 | Cxcr6 | 9.26 |
| F01 | Rdc1 | Cxcr7 | 3.6 |
| F02 | CCBP1 | Darc | 68.4 |
| F04 | AI853548 | Gpr17 | 7.55 |
| F05 | MOP1 | Hifla | 12.83 |
| F07 | KIAA4048 | Il16 | 3.96 |
| F09 | Il-4 | Il4 | 3.73 |
| F10 | Il-6 | Il6 | 87.67 |
| F11 | CR3 | Itgam | 5.51 |
| F12 | 2E6 | Itgbz | 15.7 |
| G02 | Crk1 | Mapk14 | 6.74 |
| G03 | Cxcl4 | Pf4 | 9.21 |
| G04 | Cxcl7 | Ppbp | 14.62 |
| G06 | Tgfb | Tgfb1 | 12.46 |
| G07 | Ly105 | Tlr2 | 8.67 |
| G08 | Lps | Tlr4 | 10.41 |
| G09 | Tnfa | Tnf | 2.37 |
| G12 | Ccxcr1 | Xcr1 | 17.78 |
| H02 | beta2m | B2m | 6.14 |
| H03 | Gapd | Gapdh | 41.27 |
| H04 | Gur | Gusb | 3.63 |
| Accession No. | Gene name | Gene symbol | Fold change |
| E07 | I18ra | Cxcr1 | -4.18 |
| F06 | Ifng | Ifg | -2.02 |
| H01 | β-actin | Actb | -2.80 |
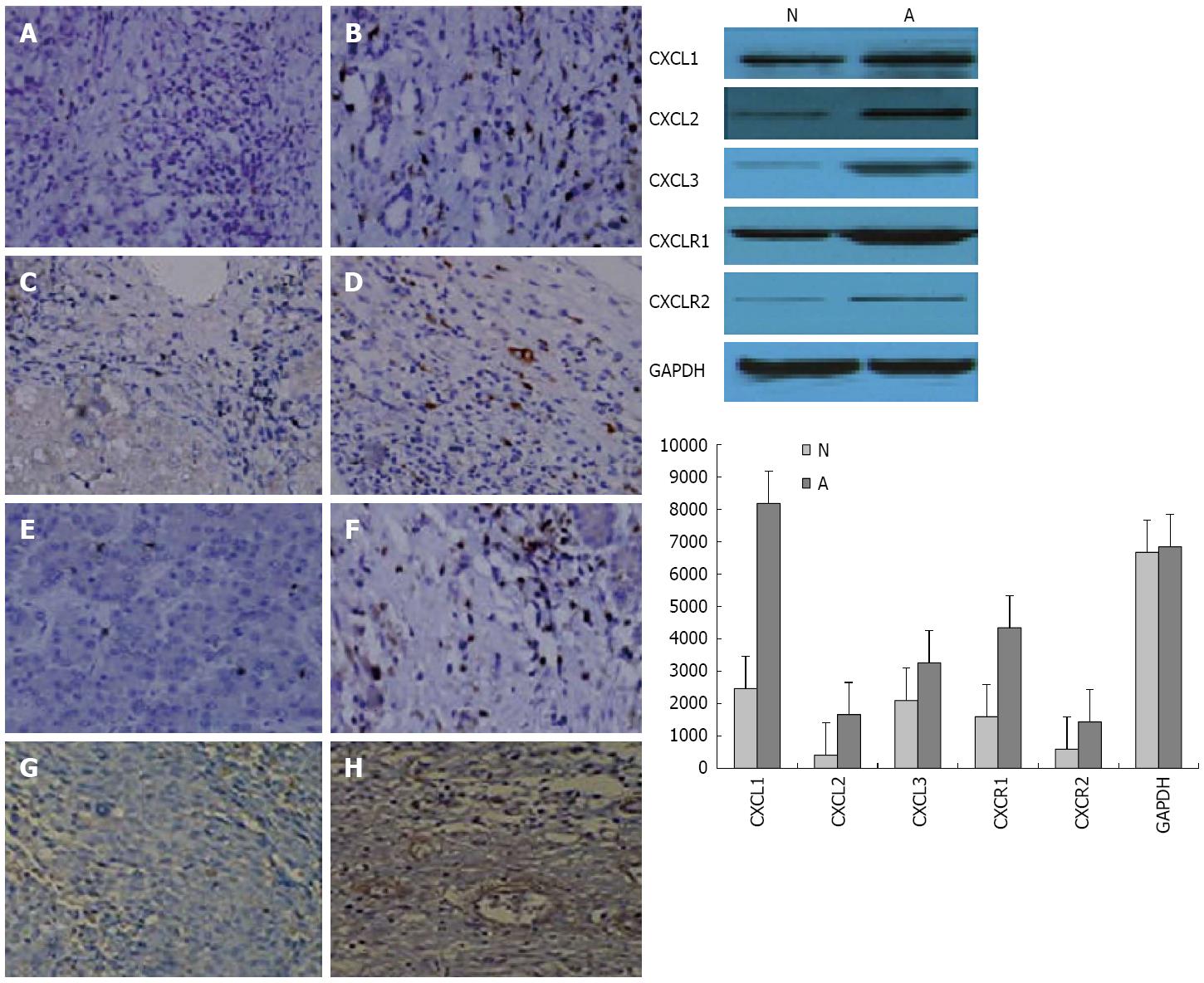
The expression of selected genes was analyzed using immunohistochemistry and Western blotting (Figure 6) to confirm the changes observed by the PCR microarray. These results were consistent with those of the microarray analysis. CXCL1, CXCL2, and CXCL3 expression was higher and CXCR1 expression was lower in tumor tissues during HCC progression compared to the controls (Figure 6).
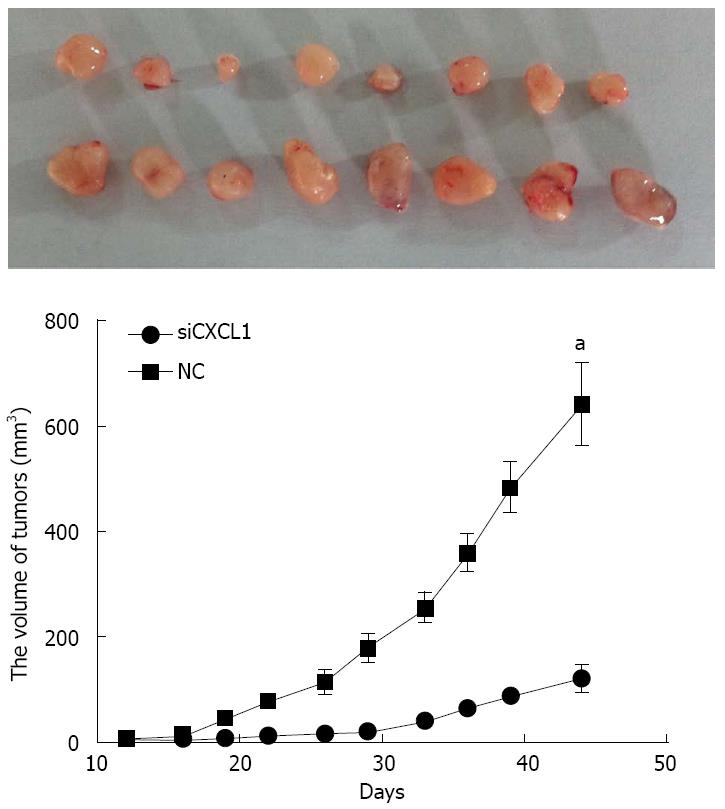
Gene silencing of CXCL1 inhibits the growth of CBRH-7919 tumors in vivo. Dissection of the subcutaneous tumors and evaluation in mice were carried out on the 45th day. Results showed the complete formation of tumors in all mice (Figure 7). The volume of tumors in the group with shCXCL1 were significantly smaller than those in the control group (P < 0.05).
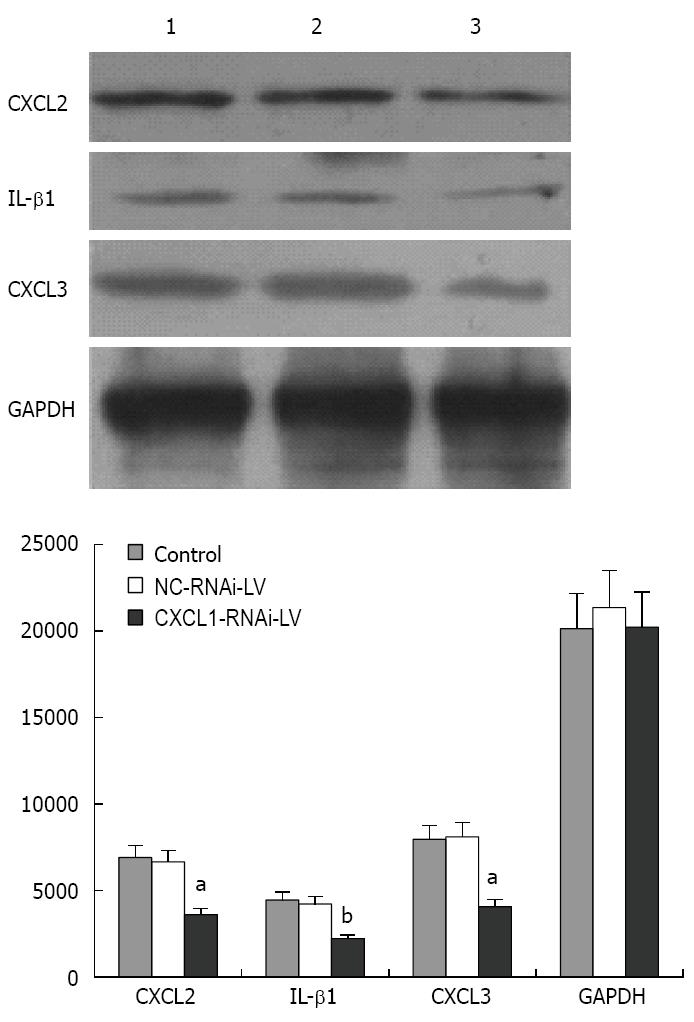
Western blotting analysis was performed to detect the expression of CXCL2, CXCL3 and IL-1β after knockdown of CXCL1. The results indicate that the protein expression levels of CXCL2, CXCL3 and IL-1β were significantly decreased compared with those of the control group and NC-RNAi-LV group (P < 0.01) (Figure 8).
Although efforts have been made to investigate the cellular and molecular pathways involved in cancer-related inflammation, as well as their potential as cancer biomarkers and therapeutic targets, the mechanisms regarding how inflammation contributes to cancer initiation, progression, metastasis, and angiogenesis remain largely unknown. In this study, we investigate the inflammatory microenvironment and expression of chemokines in HCC in nude mice. Our results show that transplanted tumors contained cells with inflammatory and macrophage phenotypes. A chemokine PCR array was used to identify differential gene expressions, which were validated by western blotting. Additionally, knockdown CXCL1 by RNAi suppressed tumor growth and expression of CXCL2, CXCL3 and IL-1β.
Chemokines and their receptors play an intricate role in HCC progression[16,17]. Chemokines are divided by their different structure and function into four families: including CC, CXC, C, and CX3C. CXC chemokines are the second largest family of chemokines, and increasing evidence suggests that chemokine expression is associated with tumor angiogenesis, tumor, progression, and metastasis[18]. CXCL1 is a growth-regulated oncogene with melanoma growth-stimulating activity. Studies have shown that CXCL1 can regulate tumor epithelial-stromal interactions that facilitate tumor growth and invasion, and CXCL1 has also been associated with angiogenesis[19-22]. CXCL1 is primarily regulated by growth factors/mediators, such as vascular endothelial growth factor (VEGF), TGF-β, c-Jun N-terminal kinase, and phosphoinositol 3-kinase. For instance, VEGF can stimulate CXCL1 release in both a time- and concentration-dependent manner, and this can be inhibited by VEGF receptor antagonists[23-26].
Some studies suggest that chemokines also mediate tumor metastasis. Both CXCL1 and CXCL2 are closely related to metastasis[23]. CXCL1 and CXCL2 expression can be suppressed by inhibiting phosphorylated IκBa and nuclear factor (NF)-κB activation. NF-κB promotes the survival of premalignant epithelial cells while also stimulating the release of proinflammatory mediators. NF-κB affects the expression of at least 400 genes with a variety functions, including inflammation, invasion and metastasis. Thus, downregulation of chemokines represents a potential treatment for cancer[4,27-29].
CXCL2 and CXCL3 are upregulated by proinflammatory cytokines. Our study shows that CXCL1, CXCL2, and CXCL3 are upregulated and CXCR1 is downregulated in the tumors of mice with HCC. These CXC chemokines and chemokine receptor are also expressed in several solid tumors. In human colon carcinoma cell lines, CXCL1 and its receptor CXCR2 have been associated with metastatic potential and are thought to modulate cell proliferation and invasion in both an autocrine and paracrine manner. CXCL1 is also overexpressed in human bladder carcinomas, and this increase in expression is associated with higher bladder carcinoma grade and stage. Studies have also reported that CXCL1 is overexpressed in colon, renal, gastric, skin, and breast cancers. However, other studies on CXCL1 expression in non-small cell lung cancer have reported the opposite results[30-32].
CXCR1 binds to only CXCL-8. Several studies have suggested that CXCR1 is an important player in tumor progression. Neutralization CXCR1 using small molecule antagonists affects cell proliferation and migration. Recent reports have also demonstrated CXCR1 expression in all melanoma cases, irrespective of stage and grade, and modulating CXCR1 expression and/or activity has been shown to regulate malignant melanoma growth, angiogenesis, and metastasis[33-35]. In addition, activating both CXCR1 and CXCR2 increased the rate of cell proliferation in prostate cancer[36,37].
RNAi is one of the post-transcriptional gene silencing mechanisms, which has emerged as a powerful tool to induce loss-of-function phenotypes and is now widely used in the research of gene analysis and therapy. A lentivirus is a retrovirus, and lentiviral vectors can efficiently deliver si/shRNA-expression cassettes into various cells with sustained expression and potent function of the encoded siRNAs[38].
Our study shows that 50 chemokine-related genes were upregulated in HCC tissues and three additional genes were downregulated. Western blotting confirmed the changes in CXCL1, CXCL2, CXCL3, and CXCR1 expression in the CBRH-7919 HCC mouse model. Furthermore, we used a chemically modified siRNA, which is thought to be more stable and effective than a non-modified siRNA. The introduction of siRNA targeting CXCL1 into CBRH-7919 resulted in efficient and specific inhibition of CXCL1 expression, as demonstrated by Western blotting. The results show that gene silencing of CXCL1 inhibits the growth of CBRH-7919 tumors in vivo and significantly decreases protein expression levels of CXCL2, CXCL3 and IL-1β. Our study provides the first evidence that the targeted silencing of CXCL1 expression results in inhibited growth in HCC. These findings support the hypothesis that CXCL1 plays an important role in protecting CBRH-7919 cells from cell death, thus indicating CXCL1 is a target for future therapeutic interventions.
Recent reports show that inflammation is a crucial factor in the tumor microenvironment, and has become a hot topic in the past few years. The tumor microenvironment is an indispensable participant in the neoplastic process, promoting proliferation, survival, and migration of such tumors, and consists of cancer cells, tumor-associated fibroblasts, and chemokines. Chemokines play a paramount role in tumor progression, angiogenesis, invasion and metastasis. Hepatocellular carcinoma (HCC) tumors express a number of chemokines/receptors, including CXCL12-CXCR4, CX3CL1-CX3CR1, and CCL20-CCR6. However, the inflammatory microenvironment and differential expression pattern of chemokines in HCC is still not clear.
In this study, inflammatory cell infiltration in the stromal microenvironment was assessed by immunohistochemistry in paired tumor and adjacent peritumoral samples, and macrophage phenotype was assessed using double-stain immunohistochemistry. PCR array analysis was used to evaluate the expression profiles of chemokines and their receptors in tumors grown in nude mice that were challenged with liver cancer (CBRH-7919 cells injected subcutaneously). The expression of some chemokines was regulated in a variety of patterns during the progression of the tumors. The expression of chemokines was confirmed by Western blotting and immunohistochemistry, demonstrating the change in chemokines and receptors at the protein level in HCC. Additionally, the effect of chemokine expression knockdown by RNA interference on tumor growth was assessed.
Recent studies have highlighted that the tumor inflammatory microenvironment plays an essential role in the progression of HCC. The tumor microenvironment plays a critical role in modulating the process of liver fibrosis, hepatocarcinogenesis, epithelial-mesenchymal transition, tumor invasion, and metastasis. Thus, understanding the inflammatory microenvironment is critical to promote understanding of the molecular, cellular, and pathophysiologic mechanisms of HCC, and is essential for the development of new therapeutic strategies.
This study provides insight into the roles of CXCL1, CXCL2, and CXCL3 in a proinflammatory peritumoral microenvironment that affect HCC development and progression. Understanding CXCL1 is involved in tumor progression may facilitate the design of new therapeutic approaches that inhibit tumor cell growth.
CXCL1 is a member of CXC chemokines family, which is identified as growth-regulated oncogene with melanoma growth-stimulating activity. CXCL1 may regulate tumor epithelial stromal interactions that facilitate tumor growth and invasion. CXCL1 is primarily regulated by growth factors/mediators, such as vascular endothelial growth factor, transforming growth factor-β, c-Jun N-terminal kinase, and phosphoinositol 3-kinase.
This paper brings us very important information about the inflammatory microenvironment and expression of chemokines in HCC. This study found that the inflammation microenvironment and differential expression of chemokines, particularly CXCL1, play critical roles in tumor growth. The result is very important for advanced research of HCC.
Animal care and use statement: The animal protocol was designed to minimize pain or discomfort to the animals. The animals were acclimatized to laboratory conditions (23 °C, 12 h/12 h light/dark, 50% humidity, ad libitum access to food and water) for two weeks prior to experimentation. All animals were euthanized by barbiturate overdose (150 mg/kg pentobarbital sodium, iv) for tissue collection.
P- Reviewer: Dirchwolf M, Yan LN S- Editor: Qi Y L- Editor: AmEditor E- Editor: Zhang DN
| 1. | Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11-30. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9215] [Cited by in RCA: 9856] [Article Influence: 821.3] [Reference Citation Analysis (4)] |
| 2. | Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893-2917. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11128] [Cited by in RCA: 11836] [Article Influence: 845.4] [Reference Citation Analysis (4)] |
| 3. | Stefaniuk P, Cianciara J, Wiercinska-Drapalo A. Present and future possibilities for early diagnosis of hepatocellular carcinoma. World J Gastroenterol. 2010;16:418-424. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 124] [Cited by in RCA: 145] [Article Influence: 9.7] [Reference Citation Analysis (1)] |
| 4. | Chew V, Tow C, Teo M, Wong HL, Chan J, Gehring A, Loh M, Bolze A, Quek R, Lee VK. Inflammatory tumour microenvironment is associated with superior survival in hepatocellular carcinoma patients. J Hepatol. 2010;52:370-379. [PubMed] |
| 5. | Schrader J, Iredale JP. The inflammatory microenvironment of HCC - the plot becomes complex. J Hepatol. 2011;54:853-855. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 19] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 6. | Yang JD, Nakamura I, Roberts LR. The tumor microenvironment in hepatocellular carcinoma: current status and therapeutic targets. Semin Cancer Biol. 2011;21:35-43. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 536] [Reference Citation Analysis (0)] |
| 7. | Li N, Grivennikov SI, Karin M. The unholy trinity: inflammation, cytokines, and STAT3 shape the cancer microenvironment. Cancer Cell. 2011;19:429-431. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 197] [Cited by in RCA: 200] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 8. | Porta C, Larghi P, Rimoldi M, Totaro MG, Allavena P, Mantovani A, Sica A. Cellular and molecular pathways linking inflammation and cancer. Immunobiology. 2009;214:761-777. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 183] [Cited by in RCA: 182] [Article Influence: 11.4] [Reference Citation Analysis (0)] |
| 9. | Dieu-Nosjean MC, Antoine M, Danel C, Heudes D, Wislez M, Poulot V, Rabbe N, Laurans L, Tartour E, de Chaisemartin L. Long-term survival for patients with non-small-cell lung cancer with intratumoral lymphoid structures. J Clin Oncol. 2008;26:4410-4417. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 608] [Cited by in RCA: 748] [Article Influence: 44.0] [Reference Citation Analysis (0)] |
| 10. | de Visser KE, Eichten A, Coussens LM. Paradoxical roles of the immune system during cancer development. Nat Rev Cancer. 2006;6:24-37. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1633] [Cited by in RCA: 1641] [Article Influence: 86.4] [Reference Citation Analysis (0)] |
| 11. | Gu W, Han KQ, Su YH, Huang XQ, Ling CQ. [Inhibition action of bufalin on human transplanted hepatocellular tumor and its effects on expressions of Bcl-2 and Bax proteins in nude mice]. Zhong Xi Yi Jie He Xue Bao. 2007;5:155-159. [PubMed] |
| 12. | Kumarakulasingham M, Rooney PH, Dundas SR, Telfer C, Melvin WT, Curran S, Murray GI. Cytochrome p450 profile of colorectal cancer: identification of markers of prognosis. Clin Cancer Res. 2005;11:3758-3765. [PubMed] |
| 13. | Munder M, Eichmann K, Morán JM, Centeno F, Soler G, Modolell M. Th1/Th2-regulated expression of arginase isoforms in murine macrophages and dendritic cells. J Immunol. 1999;163:3771-3777. [PubMed] |
| 14. | Mosser DM. The many faces of macrophage activation. J Leukoc Biol. 2003;73:209-212. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1340] [Cited by in RCA: 1359] [Article Influence: 61.8] [Reference Citation Analysis (0)] |
| 15. | Raman D, Baugher PJ, Thu YM, Richmond A. Role of chemokines in tumor growth. Cancer Lett. 2007;256:137-165. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 452] [Cited by in RCA: 453] [Article Influence: 25.2] [Reference Citation Analysis (0)] |
| 16. | Moore BB, Arenberg DA, Stoy K, Morgan T, Addison CL, Morris SB, Glass M, Wilke C, Xue YY, Sitterding S. Distinct CXC chemokines mediate tumorigenicity of prostate cancer cells. Am J Pathol. 1999;154:1503-1512. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 141] [Cited by in RCA: 144] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 17. | Waugh DJ, Wilson C, Seaton A, Maxwell PJ. Multi-faceted roles for CXC-chemokines in prostate cancer progression. Front Biosci. 2008;13:4595-4604. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 44] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 18. | Miyake M, Goodison S, Urquidi V, Gomes Giacoia E, Rosser CJ. Expression of CXCL1 in human endothelial cells induces angiogenesis through the CXCR2 receptor and the ERK1/2 and EGF pathways. Lab Invest. 2013;93:768-778. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 115] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 19. | Verbeke H, Struyf S, Laureys G, Van Damme J. The expression and role of CXC chemokines in colorectal cancer. Cytokine Growth Factor Rev. 2011;22:345-358. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 99] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 20. | Miyake M, Lawton A, Goodison S, Urquidi V, Gomes-Giacoia E, Zhang G, Ross S, Kim J, Rosser CJ. Chemokine (C-X-C) ligand 1 (CXCL1) protein expression is increased in aggressive bladder cancers. BMC Cancer. 2013;13:322. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 52] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 21. | Vinader V, Afarinkia K. The emerging role of CXC chemokines and their receptors in cancer. Future Med Chem. 2012;4:853-867. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 29] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 22. | Baggiolini M. Chemokines in pathology and medicine. J Intern Med. 2001;250:91-104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 521] [Cited by in RCA: 517] [Article Influence: 21.5] [Reference Citation Analysis (0)] |
| 23. | Lo HM, Shieh JM, Chen CL, Tsou CJ, Wu WB. Vascular Endothelial Growth Factor Induces CXCL1 Chemokine Release via JNK and PI-3K-Dependent Pathways in Human Lung Carcinoma Epithelial Cells. Int J Mol Sci. 2013;14:10090-10106. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 30] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 24. | Tong Q, Zheng L, Lin L, Li B, Wang D, Huang C, Li D. VEGF is upregulated by hypoxia-induced mitogenic factor via the PI-3K/Akt-NF-kappaB signaling pathway. Respir Res. 2006;7:37. [PubMed] |
| 25. | Hutchings H, Ortega N, Plouët J. Extracellular matrix-bound vascular endothelial growth factor promotes endothelial cell adhesion, migration, and survival through integrin ligation. FASEB J. 2003;17:1520-1522. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 26. | Shin JH, Shim JW, Kim DS, Shim JY. TGF-beta effects on airway smooth muscle cell proliferation, VEGF release and signal transduction pathways. Respirology. 2009;14:347-353. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 38] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 27. | Kavandi L, Collier MA, Nguyen H, Syed V. Progesterone and calcitriol attenuate inflammatory cytokines CXCL1 and CXCL2 in ovarian and endometrial cancer cells. J Cell Biochem. 2012;113:3143-3152. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 40] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 28. | Ditsworth D, Zong WX. NF-kappaB: key mediator of inflammation-associated cancer. Cancer Biol Ther. 2004;3:1214-1216. [PubMed] |
| 29. | Vanoirbeek E, Krishnan A, Eelen G, Verlinden L, Bouillon R, Feldman D, Verstuyf A. The anti-cancer and anti-inflammatory actions of 1,25(OH)₂D₃. Best Pract Res Clin Endocrinol Metab. 2011;25:593-604. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 73] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 30. | Li A, Varney ML, Singh RK. Constitutive expression of growth regulated oncogene (gro) in human colon carcinoma cells with different metastatic potential and its role in regulating their metastatic phenotype. Clin Exp Metastasis. 2004;21:571-579. [PubMed] |
| 31. | Tebo J, Der S, Frevel M, Khabar KS, Williams BR, Hamilton TA. Heterogeneity in control of mRNA stability by AU-rich elements. J Biol Chem. 2003;278:12085-12093. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 94] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 32. | Doll D, Keller L, Maak M, Boulesteix AL, Siewert JR, Holzmann B, Janssen KP. Differential expression of the chemokines GRO-2, GRO-3, and interleukin-8 in colon cancer and their impact on metastatic disease and survival. Int J Colorectal Dis. 2010;25:573-581. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 93] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 33. | Singh S, Sadanandam A, Varney ML, Nannuru KC, Singh RK. Small interfering RNA-mediated CXCR1 or CXCR2 knock-down inhibits melanoma tumor growth and invasion. Int J Cancer. 2010;126:328-336. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 34. | Sharma B, Singh S, Varney ML, Singh RK. Targeting CXCR1/CXCR2 receptor antagonism in malignant melanoma. Expert Opin Ther Targets. 2010;14:435-442. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 50] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 35. | Varney ML, Li A, Dave BJ, Bucana CD, Johansson SL, Singh RK. Expression of CXCR1 and CXCR2 receptors in malignant melanoma with different metastatic potential and their role in interleukin-8 (CXCL-8)-mediated modulation of metastatic phenotype. Clin Exp Metastasis. 2003;20:723-731. [PubMed] |
| 36. | Murphy C, McGurk M, Pettigrew J, Santinelli A, Mazzucchelli R, Johnston PG, Montironi R, Waugh DJ. Nonapical and cytoplasmic expression of interleukin-8, CXCR1, and CXCR2 correlates with cell proliferation and microvessel density in prostate cancer. Clin Cancer Res. 2005;11:4117-4127. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 158] [Cited by in RCA: 168] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 37. | Gabellini C, Trisciuoglio D, Desideri M, Candiloro A, Ragazzoni Y, Orlandi A, Zupi G, Del Bufalo D. Functional activity of CXCL8 receptors, CXCR1 and CXCR2, on human malignant melanoma progression. Eur J Cancer. 2009;45:2618-2627. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 105] [Cited by in RCA: 112] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 38. | Li G, Chang H, Zhai YP, Xu W. Targeted silencing of inhibitors of apoptosis proteins with siRNAs: a potential anti-cancer strategy for hepatocellular carcinoma. Asian Pac J Cancer Prev. 2013;14:4943-4952. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 27] [Article Influence: 2.5] [Reference Citation Analysis (0)] |