Published online Dec 21, 2014. doi: 10.3748/wjg.v20.i47.17727
Revised: September 30, 2014
Accepted: November 18, 2014
Published online: December 21, 2014
Processing time: 296 Days and 4.2 Hours
Traditionally bacteria have been considered as either pathogens, commensals or symbionts. The mammal gut harbors 1014 organisms dispersed on approximately 1000 different species. Today, diagnostics, in contrast to previous cultivation techniques, allow the identification of close to 100% of bacterial species. This has revealed that a range of animal models within different research areas, such as diabetes, obesity, cancer, allergy, behavior and colitis, are affected by their gut microbiota. Correlation studies may for some diseases show correlation between gut microbiota composition and disease parameters higher than 70%. Some disease phenotypes may be transferred when recolonizing germ free mice. The mechanistic aspects are not clear, but some examples on how gut bacteria stimulate receptors, metabolism, and immune responses are discussed. A more deeper understanding of the impact of microbiota has its origin in the overall composition of the microbiota and in some newly recognized species, such as Akkermansia muciniphila, Segmented filamentous bacteria and Faecalibacterium prausnitzii, which seem to have an impact on more or less severe disease in specific models. Thus, the impact of the microbiota on animal models is of a magnitude that cannot be ignored in future research. Therefore, either models with specific microbiota must be developed, or the microbiota must be characterized in individual studies and incorporated into data evaluation.
Core tip: Full characterization of the gut microbiota of animal models has revealed that animal models within different research areas, such as diabetes, obesity, cancer, allergy, behavior and colitis, are highly affected by their gut microbiota. The mechanistic aspects are not clear; however, the impact of the microbiota on animal models is of a magnitude that cannot be ignored in future research. Therefore, either models with specific microbiota must be developed, or the microbiota must be characterized in individual studies and incorporated into data evaluation.
- Citation: Hansen AK, Hansen CHF, Krych L, Nielsen DS. Impact of the gut microbiota on rodent models of human disease. World J Gastroenterol 2014; 20(47): 17727-17736
- URL: https://www.wjgnet.com/1007-9327/full/v20/i47/17727.htm
- DOI: https://dx.doi.org/10.3748/wjg.v20.i47.17727
The gut is an ideal incubation chamber for bacteria adapted to the mammal body temperature and the anaerobic environment. Thousands of years of co-existence has led to such adaptation, and the mammal gut harbors 1014 organisms dispersed over approximately 1000 different species, dependent on how the cut-offs are set for similarity. Within the traditional approach to laboratory animal bacteriology, bacteria have been considered as either pathogens, commensals or symbionts; however, there seems to be a need for a broader understanding of this. When first inside the gut, the bacteria will be fed and will be allowed to propagate, while the host organism will benefit from otherwise unavailable products of microbial digestion. Generally, pathogenicity is not in the interest of the microorganism, because it induces a strong and eradicating immune response from the host, and even in the case of microbial victory in this battle, the end result may be the death of the host and the need for the microbe to relocate to a new habitat. The host immune system, on the other hand, needs to protect the host from invasion without being so aggressive that it loses the microbe and thereby all its benefits.
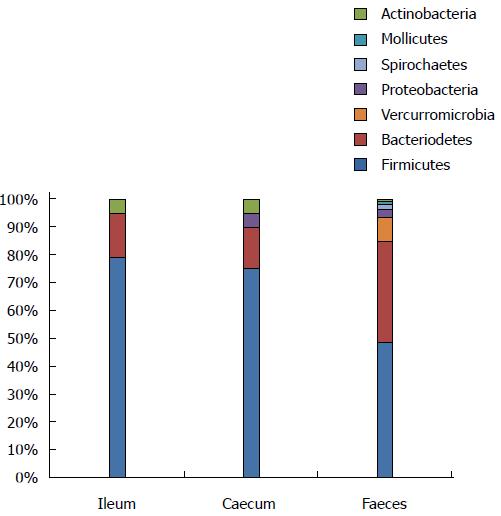
A more advanced understanding of the impact of the microbiota takes into consideration both the overall composition and the balance between the members of the microbiota, as well as some newly recognized species, which, by themselves, seem to have an effect on the specific models. Some of these have a symbiotic effect, while others push disease development in a more detrimental direction. However, same species may act in favor of the development of one disease, while being more protective against another disease, and the mechanistic potential of the species may differ between different parts of the gut. For most of these bacteria, it is their abundance, rather than their qualitative presence or absence, which are responsible for their effect on the host[1-4]. The microbiota is normally not very diverse in the upper part of the gut, e.g. in the ileum, where there is a huge accumulation of lymphatic tissue available for stimulation[3,5-10]. It gradually becomes more diverse as the gut contents pass through the large intestine and become feces (Figure 1)[3,5-11]. In both man and mouse, a microbiota with a low diversity is indicative of an increased risk of developing inflammatory disease[12,13]. Furthermore, in animals, a microbiota that is roughly similar in the upper part of the gut, may differ substantially in the lower part of the gut and vice versa[3,14]. Finally, there might be essential differences between the effects of the various species at different ages of the animals, which may explain why some species favor the development of one disease, while protecting against another.
Over recent decades, new methods based upon molecular biology diagnostics have been developed. Such methods, which include quantitative real-time polymerase chain reaction (qPCR) assays[15], pyrosequencing[16] and metagenomic sequencing, have permitted identification of close to 100% of the gut’s operational taxonomic units (OTU), which include both cultivable and non-cultivable bacterial species, and in principle, viral, eukaryotes and Archea[17], although they are seldom specifically tested for at present. In contrast, previous cultivation techniques only allowed cultivation and identification of 10%-20% of the bacterial species present in the gut[18]. These molecular biology-based tools have enabled detailed correlation studies. Such studies have revealed that a range of animal models within a range of different research areas are affected by their gut microbiota[19].
As described below, the impact of the microbiota on animal models is well documented, while the mechanisms underlying this are less clear. Some hypotheses, though, make more sense than others. As techniques for the full characterization of the microbiota have been developed over the last decade, we are only now beginning to achieve an understanding of how the microbiota actually exerts its effect on the host; however, some examples can be given.
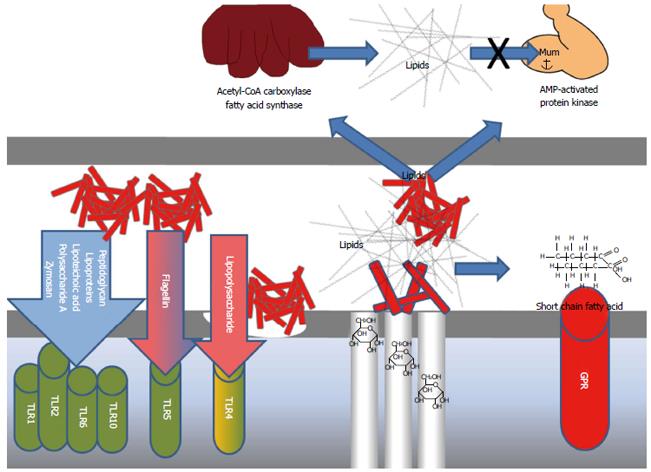
In early life, there is a window for the induction of oral tolerance in the gut[20]. This seems essential to avoid inflammatory disease later in life[21]. Molecular structures in bacteria known as microbial-associated molecular patterns (MAMP) stimulate pattern-recognition receptors (PRR) in the host, thereby inducing innate responses[22]. Among the most important PRRs are the toll-like receptors (TLR), which are present in different types on a range of different cell types[22-29] (Figure 2). An important example of a MAMP is lipopolysaccharides (LPS), which are important parts of the cell wall of Gram negative bacteria[30], such as Proteobacteria[31], from which it stimulates TLR4. Another important example is peptidoglycan, found in the cell walls of Gram positive bacteria, which stimulates TLR2[32] and flagellin deriving from flagellated bacteria, leading to stimulation of TLR5[33]. Therefore, as different types of MAMPs stimulate different TLR’s dispersed on a variety of different cell types[23], and as MAMPs are also dispersed and shared between members of the microbiota[22], there is a vast range of innate host responses to bacteria.
The age of the animal also makes a difference. For example, stimulation of TLR1, TLR2 and TLR4 in early life leads to higher production of interleukin (IL)-6 than stimulation later in life[34]. Germ free animals have more T helper cells type 2 (TH2) and less TH1 cells[35], as the stimulation of the gut lamina propria dendritic cells, e.g. by polysaccharide A (PSA) from Bacteroides fragilis, induces IL-12 secretion, which favors TH1 at the cost of TH2[36]. Host-bacterial interactions, probably mediated through glucagon-like peptide 2 (GLP-2), seem to control the gut barrier function[37]. Metabolic endotoxaemia is responsible for the phenomenon whereby excess intake of dietary fat increases plasma LPS levels[38,39], which in mice, is a sufficient molecular mechanism to trigger metabolic diseases, such as obesity and diabetes[40].
The clearest documentation of a general microbial impact on rodent models is observed when comparing a conventional model with a microbiota with a germ free version. In several studies, this has revealed essential differences in disease expression (Table 1)[22,41-57]. Although germ free mice eat more, they are leaner, and they have less body fat compared with conventional mice because they are less efficient in extracting energy from their diet[50]. Germ free mice have increased expression of obesity-related peptides, such as glucagon-like peptide 1 (GLP-1) in the brain[58], which is relevant, because central GLP-1 reduces food intake in rats[59]. Germ free mice also behave differently from microbiota-harboring mice and this behavior may be normalized by colonization[43]. However, for this phenotype there also seems to be an important time window in early life[60]. Germ free mice with a mutation causing a defect in the skin barrier suffer from a more severe B-lymphoproliferative disorder, because they express significantly higher levels of the proinflammatory cytokine thymic stromal lymphopoietin[61]. Inflammatory bowel disease (IBD) occurs either because of a TH1/TH17 response (Crohn’s disease) or a TH2 response (ulcerative colitis) to gut commensals[62]. Therefore, IBD under germ free conditions does not develop at all in, e.g. Human Leucocyte Antigen subtypes B27 (HLA-B27) transgenic rats[53] and IL-10 knockout mice[56]. For the IL-10 knockout mice[63] it does not occur even under barrier protected conditions (Table 1). IL-2 knockout mice may, under germ free conditions, show mild focal intestinal inflammation[64] (Table 1).
| Model | Disease |
| Models with increased disease incidence or severity | |
| β-lactoglobulin induced mouse[51] | Allergy |
| NOD mouse[42] | Type 1 diabetes |
| MyD88 KO NOD mouse[42] | Type 1 diabetes |
| Restrained mouse[43] | Stress |
| Models with decreased disease incidence or severity | |
| Ovalbumin-specific TCR TG mouse[44] | Allergy |
| Swiss-Webster mouse[45] | Anxiety |
| Collagen induced rat[52] | Arthritis |
| HLA-B27 TG rat[53] | IBD |
| IL-2 KO mouse[54,55] | IBD |
| IL-10 KO mouse[56] | IBD |
| TCRα KO mouse[57] | IBD |
| Dextran sulfate sodium induced mouse[46] | IBD |
| SAMP1/Yit mouse[47] | IBD |
| Adoptive T-cell transfer in the mouse[48] | IBD |
| Carrageenan, LPS, or formalin induced mouse[49] | Inflammatory pain |
| C57BL/6 mouse[65] | Obesity |
| C57BL/6 mouse[65] | Type 2 diabetes |
Within animal models of the metabolic syndrome, there seems to be an association between the gut microbiota and at least some of the metabolic parameters. For example, in leptin-deficient obese mice, there is a strong correlation between glycated hemoglobin levels and the composition of the gut microbiota[1]. Further, these mice have significantly more Firmicutes and fewer Bacteriodetes members compared with their wild-type and heterozygous litter mates[10]. Their obese phenotype may be transferred with the microbiota by recolonizing germ free lean wild-type mice[65]. In C57 Black substrain 6 (C57BL/6) mice on both high and low calorie diet, continuous oral ampicillin improves glucose tolerance[66,67]. However, this effect is mainly caused by an early life impact on glucose tolerance, and the effect ceases immediately after termination of treatment; thereafter, the glucose tolerance may even decrease[68,69]. Several studies describe cross-talk between the brain and the gut through both the vagal system and the hypothalamus-pituitary-adrenal (HPA) axis[70]. Stressing animal models changes their microbiota[71], and the composition of the gut microbiota has an impact on responses in rodent stress tests[72,73]. Innate immune system cytokines, such as IL-1, IL-6 and tumor necrosis factor α (TNFα), which may originate from a gut microbiota provocation, induce “sickness behavior”, changing the priorities of the organism to enhance recovery and survival[74]. However, metabolites formed by microbial decomposition in the gut may also have a direct impact on the brain[75]. In mouse models of atopic dermatitis, more than 70% of the variation observed in the local tissue cytokine response may be shared with the variation in gut microbiota[76]. Changes in the structure of the microbial community seem to reduce the number, as well as the size, of tumors in azoxymethane/dextran sodium sulfate (AOM/DSS) colon cancer-induced mice, and tumor induction may be achieved by colonizing germ free mice with microbiota from induced mice[77].
Akkermansia muciniphila (A. muciniphila) is a Gram negative bacterium, which in mice is the only species belonging to the phylum Verrucomicrobia[78]. It interacts via its mucin degrading capabilities with enteroendocrine cells to modulate gut barrier function, and it is capable of producing certain short chain fatty acids (SCFA’s) with a direct action on the receptor G-protein receptor 43 (GPR43)[79]. Abundance of A. muciniphila is reduced in mice with obesity and type 2 diabetes[80], and it gradually disappears as aging leptin deficient obese mice develop insulin resistance[1]. In non-obese diabetic (NOD) mice it becomes more abundant when mice are fed a gluten-free diet, which decreases the incidence of type 1 diabetes[81]. Early life treatment with vancomycin in NOD mice allows A. muciniphila to become a dominant gut microbiota member, which reduces the incidence of type 1 diabetes[3], but enhances susceptibility to allergic asthma[82], which is in accordance with other studies showing allergy and diabetes to counteract one another in NOD mice[83,84]. Induction of IBD in mice with dextran sodium sulfate (DSS) reduces the number of extracellular vesicles derived from A. muciniphila, and feeding DSS induced mice such vesicles reduces the severity of IBD[85], which fits well with observations in humans[4]. However, it not only reduces the severity of diseases: its presence is correlated with higher severity when infecting mice with Salmonella typhimurium[86], and AOM/DSS colon cancer-induced mice have an increased abundance of A. muciniphila[77], which may be explained by its ability to downregulate the natural killer cell receptor, NKG2D, which is part of the anti-carcinogenic defense[87].
Segmented filamentous bacteria (SFB’s) are clostridia-related Gram-positive bacteria[88]. The term has been applied for decades to describe intestinal bacteria of a uniform morphology[89]. However, today the term refers to one single species, also known as Candidatus Savagella[90]. SFBs induce secretion of the pro-inflammatory cytokine IL-17 from TH17 cells[91], which in the adult mouse is correlated with a low number of regulatory T cells[92]. The presence of SFB’s differs between mice from different vendors[92], and SFB positive NOD mice have a significantly lower incidence of type 1 diabetes compared with SFB negative ones[93]. In the adoptive transfer severe combined immune deficiency (SCID) mouse model of IBD, SFBs are essential for the induction of severe inflammation[48]. Furthermore, SFBs and the induced TH17 are important in the defense against intestinal pathogens. For example, mice infected with Citrobacter rodentium, a potent murine colon pathogen, exhibit severe symptoms if they lack SFBs[91].
IBD in IL-10 knockout mice is enhanced by Enterococcus fecalis[94,95], which is probably linked to its production of gelatinase[96].
Faecalibacterium prausnitzii (F. prausnitzii) is a clostridia-related bacterium[97] linked to a protective effect against human Crohn’s disease[98]. Oral feeding of F. prausnitzii reduced the severity of 2,4,6-trinitrobenzenesulfonic acid (TNBS)-induced colitis in mice, and some studies indicated that this may also be the case in both multidrug resistance gene deficient (mdr1a knockout)[99] and in the DSS-induced mouse models of colitis[100].
High abundances of Lactobacillus spp. and bifidobacteria are correlated strongly with low levels of inflammation in mice[101] and leptin in rats[102], which also fits well with these bacteria acting protectively against IBD in IL-10 knockout mice[103], allergic sensitization in mice[104], and myocardial infarction in rats[102]. Lachnospiraceae seems quantitatively correlated to improved glucose tolerance in leptin-deficient obese mice[1].
In stressed mice, there is correlation between their Firmicutes levels and their responses in the stress tests[73]. Ingestion of Lactobacillus rhamnosus in mice regulates their emotional behavior and central γ-aminobutyric acid (GABA) receptor expression via the vagus nerve[72].
A high abundance of the Gram negative family Prevotellaceae, perhaps restricted to one unclassified genus, in the gut of leptin-deficient obese mice correlated with impaired glucose tolerance[1].By contrast, in AOM/DSS induced colon cancer mice, a high abundance of Prevotellaceae correlated with a low tumor burden[77]. P. copri, which has been correlated with the development of arthritis in humans, seems to increase the severity of DSS induced colitis in mice[5]. Caspase-3 knockout mice exhibit a lower inflammatory response to DSS induction of colitis compared with wild-type mice; however, this protective effect of the mutation is decreased by cohousing knockout mice with wild-type mice, which significantly increases the abundance of Prevotella spp. in the knockout mice[105].
Bacteroides vulgatus seems to enhance IBD in HLA-B27 transgenic rats[106] and IL-10 knockout mice[95], and in the Bio Breeding (BB) rat, a spontaneous type 1 diabetes model. The fecal microbiota differ and contain an increased number of Bacteroides spp. before onset of diabetes[107]. As in all other mammals, Bacteroides spp. form an important part of the Bacteroidetes fraction of the rodent gut[16]. These Gram negative bacteria are important for the processing of complex molecules to simpler ones in the gut[108]: complex glycans are their key source of energy[109]. B. fragilis toxins cause symptoms of diarrhea and IBD in germ-free mice[110], and they induce colonic tumors strongly in multiple intestinal neoplasia (MIN) mice[111]. On the other hand, B. fragilis PSA, which is important for the inflammatory gut response to pathogens[36], also protects against Helicobacter hepaticus-induced colitis in mice; probably via the prevention of IL-17 secretion[112]. Feeding the maternal immune activation (MIA) mouse model with B. fragilis reduces symptoms of autism, which is probably linked to the normalization of the levels of a specific gut metabolite[113].
The abundance of Alistipes spp., a bacterium of the Rikenellaceae family, seems to increase when mice are stressed by grid floor housing[73].
Escherichia coli (E. coli) enhances IBD in HLA-B27 overexpressing rats[106], although E. coli Nissle stabilizes the enteric barrier in mice[114]. When reducing type 1 diabetes by pre-weaning treatment of NOD mice with vancomycin, a vast increase in the abundance of Proteobacteria in the pups was observed[3].
Bifidobacterium spp. in rodents have a positive impact on the regulatory and innate immunity[101,115]. Perinatal supplementation of B. longum reduced TH1 and TH2 responses in allergen sensitized mice[104]. On the other hand, their numbers are also increased in gluten-fed NOD mice with a high incidence of type 1 diabetes compared with NOD mice on a gluten-free diet[81].
The information gained over the last decade on how the entire microbiota, as well as some of its individual members, affect animal models of very different types, has prompted the scientific community to incorporate this in future production and quality assurance of animal models. It is not possible to regard these matters from a “Specific pathogen-free” concept, as some of the species act in favor of the development of one disease, while against the development of another disease, e.g. SFB’s both protect against type 1 diabetes and induces a TH17 response in favor of the development of Crohn’s disease. Furthermore, the balance between the different fractions of the microbiota is also likely to make a difference. Ultimately, it is often a quantitative rather than a qualitative presence that makes the difference. Therefore, it is likely that we will see more tailor-made rodent models, i.e. commercial breeders and research groups have sought to produce animals with a specific microbiota for the conditions under test. One obvious idea may be to breed such animals by selective breeding; however, this does not seem to increase the microbiota similarity, although the microbiota of offspring show a clear clustering with the mother’s microbiota[116,117]. It is probably rational to inoculate germ free mice with a tailor-made microbiota around weaning, as they are conventionalized in SPF conditions[118]. The window for induction of oral tolerance in animal models may also be turned around, such that a low bacterial stimulation in the open phase of this window may be essential to develop target diseases in the model. When stimulated later on, the nature of this stimulation is also essential, because commonly used disease models in rodents are driven by specific subsets of T cells[19]. Another alternative will be to characterize the microbiota composition for animals in sensitive studies and incorporate this in the data evaluation by chemometric or multifactorial statistical means. The impact of the gut microbiota on animal models is of a magnitude that cannot be neglected in the future.
P- Reviewer: Bailey MT, Franceschi F S- Editor: Nan J L- Editor: Stewart G E- Editor: Liu XM
| 1. | Ellekilde M, Krych L, Hansen CH, Hufeldt MR, Dahl K, Hansen LH, Sørensen SJ, Vogensen FK, Nielsen DS, Hansen AK. Characterization of the gut microbiota in leptin deficient obese mice - Correlation to inflammatory and diabetic parameters. Res Vet Sci. 2014;96:241-250. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 70] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 2. | Brown CT, Davis-Richardson AG, Giongo A, Gano KA, Crabb DB, Mukherjee N, Casella G, Drew JC, Ilonen J, Knip M. Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PLoS One. 2011;6:e25792. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 496] [Cited by in RCA: 590] [Article Influence: 42.1] [Reference Citation Analysis (0)] |
| 3. | Hansen CH, Krych L, Nielsen DS, Vogensen FK, Hansen LH, Sørensen SJ, Buschard K, Hansen AK. Early life treatment with vancomycin propagates Akkermansia muciniphila and reduces diabetes incidence in the NOD mouse. Diabetologia. 2012;55:2285-2294. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 355] [Cited by in RCA: 383] [Article Influence: 29.5] [Reference Citation Analysis (0)] |
| 4. | Png CW, Lindén SK, Gilshenan KS, Zoetendal EG, McSweeney CS, Sly LI, McGuckin MA, Florin TH. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am J Gastroenterol. 2010;105:2420-2428. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 863] [Cited by in RCA: 1035] [Article Influence: 69.0] [Reference Citation Analysis (0)] |
| 5. | Scher JU, Sczesnak A, Longman RS, Segata N, Ubeda C, Bielski C, Rostron T, Cerundolo V, Pamer EG, Abramson SB. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. Elife. 2013;2:e01202. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1133] [Cited by in RCA: 1434] [Article Influence: 119.5] [Reference Citation Analysis (0)] |
| 6. | Ubeda C, Taur Y, Jenq RR, Equinda MJ, Son T, Samstein M, Viale A, Socci ND, van den Brink MR, Kamboj M. Vancomycin-resistant Enterococcus domination of intestinal microbiota is enabled by antibiotic treatment in mice and precedes bloodstream invasion in humans. J Clin Invest. 2010;120:4332-4341. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 626] [Cited by in RCA: 687] [Article Influence: 45.8] [Reference Citation Analysis (0)] |
| 7. | Hildebrandt MA, Hoffmann C, Sherrill-Mix SA, Keilbaugh SA, Hamady M, Chen YY, Knight R, Ahima RS, Bushman F, Wu GD. High-fat diet determines the composition of the murine gut microbiome independently of obesity. Gastroenterology. 2009;137:1716-1724.e1-e2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1234] [Cited by in RCA: 1160] [Article Influence: 72.5] [Reference Citation Analysis (0)] |
| 8. | Antonopoulos DA, Huse SM, Morrison HG, Schmidt TM, Sogin ML, Young VB. Reproducible community dynamics of the gastrointestinal microbiota following antibiotic perturbation. Infect Immun. 2009;77:2367-2375. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 447] [Cited by in RCA: 414] [Article Influence: 25.9] [Reference Citation Analysis (0)] |
| 9. | Wilson KH, Brown RS, Andersen GL, Tsang J, Sartor B. Comparison of fecal biota from specific pathogen free and feral mice. Anaerobe. 2006;12:249-253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 24] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 10. | Ley RE, Bäckhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI. Obesity alters gut microbial ecology. Proc Natl Acad Sci USA. 2005;102:11070-11075. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4639] [Cited by in RCA: 4462] [Article Influence: 223.1] [Reference Citation Analysis (1)] |
| 11. | Walk ST, Blum AM, Ewing SA, Weinstock JV, Young VB. Alteration of the murine gut microbiota during infection with the parasitic helminth Heligmosomoides polygyrus. Inflamm Bowel Dis. 2010;16:1841-1849. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 242] [Cited by in RCA: 246] [Article Influence: 16.4] [Reference Citation Analysis (0)] |
| 12. | Hildebrand F, Nguyen TL, Brinkman B, Yunta RG, Cauwe B, Vandenabeele P, Liston A, Raes J. Inflammation-associated enterotypes, host genotype, cage and inter-individual effects drive gut microbiota variation in common laboratory mice. Genome Biol. 2013;14:R4. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 340] [Cited by in RCA: 357] [Article Influence: 29.8] [Reference Citation Analysis (0)] |
| 13. | Qin J, Li Y, Cai Z, Li S, Zhu J, Zhang F, Liang S, Zhang W, Guan Y, Shen D. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature. 2012;490:55-60. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3971] [Cited by in RCA: 4826] [Article Influence: 371.2] [Reference Citation Analysis (1)] |
| 14. | Pang W, Vogensen FK, Nielsen DS, Hansen AK. Faecal and caecal microbiota profiles of mice do not cluster in the same way. Lab Anim. 2012;46:231-236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 31] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 15. | Bergström A, Licht TR, Wilcks A, Andersen JB, Schmidt LR, Grønlund HA, Vigsnaes LK, Michaelsen KF, Bahl MI. Introducing GUt low-density array (GULDA): a validated approach for qPCR-based intestinal microbial community analysis. FEMS Microbiol Lett. 2012;337:38-47. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 70] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 16. | Krych L, Hansen CH, Hansen AK, van den Berg FW, Nielsen DS. Quantitatively different, yet qualitatively alike: a meta-analysis of the mouse core gut microbiome with a view towards the human gut microbiome. PLoS One. 2013;8:e62578. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 146] [Cited by in RCA: 178] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 17. | Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464:59-65. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9101] [Cited by in RCA: 7831] [Article Influence: 522.1] [Reference Citation Analysis (4)] |
| 18. | Zoetendal EG, Collier CT, Koike S, Mackie RI, Gaskins HR. Molecular ecological analysis of the gastrointestinal microbiota: a review. J Nutr. 2004;134:465-472. [PubMed] |
| 19. | Bleich A, Hansen AK. Time to include the gut microbiota in the hygienic standardisation of laboratory rodents. Comp Immunol Microbiol Infect Dis. 2012;35:81-92. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 59] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 20. | Hansen CH, Nielsen DS, Kverka M, Zakostelska Z, Klimesova K, Hudcovic T, Tlaskalova-Hogenova H, Hansen AK. Patterns of early gut colonization shape future immune responses of the host. PLoS One. 2012;7:e34043. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 225] [Cited by in RCA: 215] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 21. | Weng M, Walker WA. The role of gut microbiota in programming the immune phenotype. J Dev Orig Health Dis. 2013;4:203-214. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 111] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 22. | Tlaskalová-Hogenová H, Stepánková R, Hudcovic T, Tucková L, Cukrowska B, Lodinová-Zádníková R, Kozáková H, Rossmann P, Bártová J, Sokol D. Commensal bacteria (normal microflora), mucosal immunity and chronic inflammatory and autoimmune diseases. Immunol Lett. 2004;93:97-108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 470] [Cited by in RCA: 478] [Article Influence: 23.9] [Reference Citation Analysis (0)] |
| 23. | Kamdar K, Nguyen V, DePaolo RW. Toll-like receptor signaling and regulation of intestinal immunity. Virulence. 2013;4:207-212. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 50] [Cited by in RCA: 63] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 24. | Souza-Fonseca-Guimaraes F, Parlato M, Philippart F, Misset B, Cavaillon JM, Adib-Conquy M. Toll-like receptors expression and interferon-γ production by NK cells in human sepsis. Crit Care. 2012;16:R206. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 93] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 25. | Sasai M, Yamamoto M. Pathogen recognition receptors: ligands and signaling pathways by Toll-like receptors. Int Rev Immunol. 2013;32:116-133. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 122] [Cited by in RCA: 126] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 26. | Samuel BS, Shaito A, Motoike T, Rey FE, Backhed F, Manchester JK, Hammer RE, Williams SC, Crowley J, Yanagisawa M. Effects of the gut microbiota on host adiposity are modulated by the short-chain fatty-acid binding G protein-coupled receptor, Gpr41. Proc Natl Acad Sci USA. 2008;105:16767-16772. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1270] [Cited by in RCA: 1149] [Article Influence: 67.6] [Reference Citation Analysis (0)] |
| 27. | Denechaud PD, Dentin R, Girard J, Postic C. Role of ChREBP in hepatic steatosis and insulin resistance. FEBS Lett. 2008;582:68-73. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 105] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 28. | Stappenbeck TS, Hooper LV, Gordon JI. Developmental regulation of intestinal angiogenesis by indigenous microbes via Paneth cells. Proc Natl Acad Sci USA. 2002;99:15451-15455. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 771] [Cited by in RCA: 758] [Article Influence: 33.0] [Reference Citation Analysis (0)] |
| 29. | Bjursell M, Admyre T, Göransson M, Marley AE, Smith DM, Oscarsson J, Bohlooly-Y M. Improved glucose control and reduced body fat mass in free fatty acid receptor 2-deficient mice fed a high-fat diet. Am J Physiol Endocrinol Metab. 2011;300:E211-E220. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 210] [Cited by in RCA: 220] [Article Influence: 15.7] [Reference Citation Analysis (0)] |
| 30. | Raetz CR, Whitfield C. Lipopolysaccharide endotoxins. Annu Rev Biochem. 2002;71:635-700. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3576] [Cited by in RCA: 3466] [Article Influence: 144.4] [Reference Citation Analysis (0)] |
| 31. | Huber M, Kalis C, Keck S, Jiang Z, Georgel P, Du X, Shamel L, Sovath S, Mudd S, Beutler B. R-form LPS, the master key to the activation ofTLR4/MD-2-positive cells. Eur J Immunol. 2006;36:701-711. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 118] [Cited by in RCA: 129] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 32. | Ozinsky A, Underhill DM, Fontenot JD, Hajjar AM, Smith KD, Wilson CB, Schroeder L, Aderem A. The repertoire for pattern recognition of pathogens by the innate immune system is defined by cooperation between toll-like receptors. Proc Natl Acad Sci USA. 2000;97:13766-13771. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1562] [Cited by in RCA: 1478] [Article Influence: 59.1] [Reference Citation Analysis (0)] |
| 33. | Feuillet V, Medjane S, Mondor I, Demaria O, Pagni PP, Galán JE, Flavell RA, Alexopoulou L. Involvement of Toll-like receptor 5 in the recognition of flagellated bacteria. Proc Natl Acad Sci USA. 2006;103:12487-12492. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 232] [Cited by in RCA: 250] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 34. | Liao SL, Yeh KW, Lai SH, Lee WI, Huang JL. Maturation of Toll-like receptor 1-4 responsiveness during early life. Early Hum Dev. 2013;89:473-478. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 24] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 35. | Mazmanian SK, Kasper DL. The love-hate relationship between bacterial polysaccharides and the host immune system. Nat Rev Immunol. 2006;6:849-858. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 240] [Cited by in RCA: 252] [Article Influence: 13.3] [Reference Citation Analysis (0)] |
| 36. | Mazmanian SK, Liu CH, Tzianabos AO, Kasper DL. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell. 2005;122:107-118. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1932] [Cited by in RCA: 2093] [Article Influence: 104.7] [Reference Citation Analysis (0)] |
| 37. | Cani PD, Possemiers S, Van de Wiele T, Guiot Y, Everard A, Rottier O, Geurts L, Naslain D, Neyrinck A, Lambert DM. Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut. 2009;58:1091-1103. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1735] [Cited by in RCA: 1886] [Article Influence: 117.9] [Reference Citation Analysis (1)] |
| 38. | Cani PD, Delzenne NM. Gut microflora as a target for energy and metabolic homeostasis. Curr Opin Clin Nutr Metab Care. 2007;10:729-734. [PubMed] |
| 39. | Cani PD, Bibiloni R, Knauf C, Waget A, Neyrinck AM, Delzenne NM, Burcelin R. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes. 2008;57:1470-1481. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3224] [Cited by in RCA: 3541] [Article Influence: 208.3] [Reference Citation Analysis (0)] |
| 40. | Cani PD, Amar J, Iglesias MA, Poggi M, Knauf C, Bastelica D, Neyrinck AM, Fava F, Tuohy KM, Chabo C. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes. 2007;56:1761-1772. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4095] [Cited by in RCA: 4559] [Article Influence: 253.3] [Reference Citation Analysis (1)] |
| 41. | Hazebrouck S, Przybylski-Nicaise L, Ah-Leung S, Adel-Patient K, Corthier G, Wal JM, Rabot S. Allergic sensitization to bovine beta-lactoglobulin: comparison between germ-free and conventional BALB/c mice. Int Arch Allergy Immunol. 2009;148:65-72. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 35] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 42. | Wen L, Ley RE, Volchkov PY, Stranges PB, Avanesyan L, Stonebraker AC, Hu C, Wong FS, Szot GL, Bluestone JA. Innate immunity and intestinal microbiota in the development of Type 1 diabetes. Nature. 2008;455:1109-1113. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1623] [Cited by in RCA: 1495] [Article Influence: 87.9] [Reference Citation Analysis (1)] |
| 43. | Sudo N, Chida Y, Aiba Y, Sonoda J, Oyama N, Yu XN, Kubo C, Koga Y. Postnatal microbial colonization programs the hypothalamic-pituitary-adrenal system for stress response in mice. J Physiol. 2004;558:263-275. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2097] [Cited by in RCA: 1861] [Article Influence: 88.6] [Reference Citation Analysis (1)] |
| 44. | Tsuda M, Hosono A, Yanagibashi T, Kihara-Fujioka M, Hachimura S, Itoh K, Hirayama K, Takahashi K, Kaminogawa S. Intestinal commensal bacteria promote T cell hyporesponsiveness and down-regulate the serum antibody responses induced by dietary antigen. Immunol Lett. 2010;132:45-52. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 23] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 45. | Neufeld KM, Kang N, Bienenstock J, Foster JA. Reduced anxiety-like behavior and central neurochemical change in germ-free mice. Neurogastroenterol Motil. 2011;23:255-264, e119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 887] [Cited by in RCA: 934] [Article Influence: 66.7] [Reference Citation Analysis (0)] |
| 46. | Pils MC, Bleich A, Prinz I, Fasnacht N, Bollati-Fogolin M, Schippers A, Rozell B, Müller W. Commensal gut flora reduces susceptibility to experimentally induced colitis via T-cell-derived interleukin-10. Inflamm Bowel Dis. 2011;17:2038-2046. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 40] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 47. | Matsumoto S, Okabe Y, Setoyama H, Takayama K, Ohtsuka J, Funahashi H, Imaoka A, Okada Y, Umesaki Y. Inflammatory bowel disease-like enteritis and caecitis in a senescence accelerated mouse P1/Yit strain. Gut. 1998;43:71-78. [PubMed] |
| 48. | Stepankova R, Powrie F, Kofronova O, Kozakova H, Hudcovic T, Hrncir T, Uhlig H, Read S, Rehakova Z, Benada O. Segmented filamentous bacteria in a defined bacterial cocktail induce intestinal inflammation in SCID mice reconstituted with CD45RBhigh CD4+ T cells. Inflamm Bowel Dis. 2007;13:1202-1211. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 151] [Cited by in RCA: 159] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 49. | Amaral FA, Sachs D, Costa VV, Fagundes CT, Cisalpino D, Cunha TM, Ferreira SH, Cunha FQ, Silva TA, Nicoli JR. Commensal microbiota is fundamental for the development of inflammatory pain. Proc Natl Acad Sci USA. 2008;105:2193-2197. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 168] [Cited by in RCA: 218] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 50. | Bäckhed F, Ding H, Wang T, Hooper LV, Koh GY, Nagy A, Semenkovich CF, Gordon JI. The gut microbiota as an environmental factor that regulates fat storage. Proc Natl Acad Sci USA. 2004;101:15718-15723. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4530] [Cited by in RCA: 4408] [Article Influence: 209.9] [Reference Citation Analysis (4)] |
| 51. | Rodriguez B, Prioult G, Bibiloni R, Nicolis I, Mercenier A, Butel MJ, Waligora-Dupriet AJ. Germ-free status and altered caecal subdominant microbiota are associated with a high susceptibility to cow’s milk allergy in mice. FEMS Microbiol Ecol. 2011;76:133-144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 69] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 52. | Breban MA, Moreau MC, Fournier C, Ducluzeau R, Kahn MF. Influence of the bacterial flora on collagen-induced arthritis in susceptible and resistant strains of rats. Clin Exp Rheumatol. 1993;11:61-64. [PubMed] |
| 53. | Taurog JD, Richardson JA, Croft JT, Simmons WA, Zhou M, Fernández-Sueiro JL, Balish E, Hammer RE. The germfree state prevents development of gut and joint inflammatory disease in HLA-B27 transgenic rats. J Exp Med. 1994;180:2359-2364. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 841] [Cited by in RCA: 817] [Article Influence: 26.4] [Reference Citation Analysis (0)] |
| 54. | Contractor NV, Bassiri H, Reya T, Park AY, Baumgart DC, Wasik MA, Emerson SG, Carding SR. Lymphoid hyperplasia, autoimmunity, and compromised intestinal intraepithelial lymphocyte development in colitis-free gnotobiotic IL-2-deficient mice. J Immunol. 1998;160:385-394. [PubMed] |
| 55. | Sadlack B, Merz H, Schorle H, Schimpl A, Feller AC, Horak I. Ulcerative colitis-like disease in mice with a disrupted interleukin-2 gene. Cell. 1993;75:253-261. [PubMed] |
| 56. | Sellon RK, Tonkonogy S, Schultz M, Dieleman LA, Grenther W, Balish E, Rennick DM, Sartor RB. Resident enteric bacteria are necessary for development of spontaneous colitis and immune system activation in interleukin-10-deficient mice. Infect Immun. 1998;66:5224-5231. [PubMed] |
| 57. | Dianda L, Hanby AM, Wright NA, Sebesteny A, Hayday AC, Owen MJ. T cell receptor-alpha beta-deficient mice fail to develop colitis in the absence of a microbial environment. Am J Pathol. 1997;150:91-97. [PubMed] |
| 58. | Schéle E, Grahnemo L, Anesten F, Hallén A, Bäckhed F, Jansson JO. The gut microbiota reduces leptin sensitivity and the expression of the obesity-suppressing neuropeptides proglucagon (Gcg) and brain-derived neurotrophic factor (Bdnf) in the central nervous system. Endocrinology. 2013;154:3643-3651. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 131] [Cited by in RCA: 162] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 59. | Turton MD, O’Shea D, Gunn I, Beak SA, Edwards CM, Meeran K, Choi SJ, Taylor GM, Heath MM, Lambert PD. A role for glucagon-like peptide-1 in the central regulation of feeding. Nature. 1996;379:69-72. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1354] [Cited by in RCA: 1374] [Article Influence: 47.4] [Reference Citation Analysis (0)] |
| 60. | Neufeld KA, Kang N, Bienenstock J, Foster JA. Effects of intestinal microbiota on anxiety-like behavior. Commun Integr Biol. 2011;4:492-494. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 111] [Reference Citation Analysis (0)] |
| 61. | Yockey LJ, Demehri S, Turkoz M, Turkoz A, Ahern PP, Jassim O, Manivasagam S, Kearney JF, Gordon JI, Kopan R. The absence of a microbiota enhances TSLP expression in mice with defective skin barrier but does not affect the severity of their allergic inflammation. J Invest Dermatol. 2013;133:2714-2721. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 27] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 62. | Kaser A, Zeissig S, Blumberg RS. Inflammatory bowel disease. Annu Rev Immunol. 2010;28:573-621. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1587] [Cited by in RCA: 1535] [Article Influence: 102.3] [Reference Citation Analysis (0)] |
| 63. | Kühn R, Löhler J, Rennick D, Rajewsky K, Müller W. Interleukin-10-deficient mice develop chronic enterocolitis. Cell. 1993;75:263-274. [PubMed] |
| 64. | Schultz M, Tonkonogy SL, Sellon RK, Veltkamp C, Godfrey VL, Kwon J, Grenther WB, Balish E, Horak I, Sartor RB. IL-2-deficient mice raised under germfree conditions develop delayed mild focal intestinal inflammation. Am J Physiol. 1999;276:G1461-G1472. [PubMed] |
| 65. | Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444:1027-1031. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9796] [Cited by in RCA: 8747] [Article Influence: 460.4] [Reference Citation Analysis (1)] |
| 66. | Membrez M, Blancher F, Jaquet M, Bibiloni R, Cani PD, Burcelin RG, Corthesy I, Macé K, Chou CJ. Gut microbiota modulation with norfloxacin and ampicillin enhances glucose tolerance in mice. FASEB J. 2008;22:2416-2426. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 348] [Cited by in RCA: 368] [Article Influence: 21.6] [Reference Citation Analysis (0)] |
| 67. | Bech-Nielsen GV, Hansen CH, Hufeldt MR, Nielsen DS, Aasted B, Vogensen FK, Midtvedt T, Hansen AK. Manipulation of the gut microbiota in C57BL/6 mice changes glucose tolerance without affecting weight development and gut mucosal immunity. Res Vet Sci. 2012;92:501-508. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 38] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 68. | Rune I, Hansen CH, Ellekilde M, Nielsen DS, Skovgaard K, Rolin BC, Lykkesfeldt J, Josefsen K, Tranberg B, Kihl P. Ampicillin-improved glucose tolerance in diet-induced obese C57BL/6NTac mice is age dependent. J Diabetes Res. 2013;2013:319321. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 24] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 69. | Cho I, Yamanishi S, Cox L, Methé BA, Zavadil J, Li K, Gao Z, Mahana D, Raju K, Teitler I. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature. 2012;488:621-626. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1092] [Cited by in RCA: 1209] [Article Influence: 93.0] [Reference Citation Analysis (0)] |
| 70. | Bonaz BL, Bernstein CN. Brain-gut interactions in inflammatory bowel disease. Gastroenterology. 2013;144:36-49. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 423] [Cited by in RCA: 486] [Article Influence: 40.5] [Reference Citation Analysis (0)] |
| 71. | O’Mahony SM, Marchesi JR, Scully P, Codling C, Ceolho AM, Quigley EM, Cryan JF, Dinan TG. Early life stress alters behavior, immunity, and microbiota in rats: implications for irritable bowel syndrome and psychiatric illnesses. Biol Psychiatry. 2009;65:263-267. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 753] [Cited by in RCA: 796] [Article Influence: 49.8] [Reference Citation Analysis (0)] |
| 72. | Bravo JA, Forsythe P, Chew MV, Escaravage E, Savignac HM, Dinan TG, Bienenstock J, Cryan JF. Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proc Natl Acad Sci USA. 2011;108:16050-16055. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2500] [Cited by in RCA: 2629] [Article Influence: 187.8] [Reference Citation Analysis (0)] |
| 73. | Bangsgaard Bendtsen KM, Krych L, Sørensen DB, Pang W, Nielsen DS, Josefsen K, Hansen LH, Sørensen SJ, Hansen AK. Gut microbiota composition is correlated to grid floor induced stress and behavior in the BALB/c mouse. PLoS One. 2012;7:e46231. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 227] [Cited by in RCA: 234] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 74. | Dantzer R, Kelley KW. Twenty years of research on cytokine-induced sickness behavior. Brain Behav Immun. 2007;21:153-160. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1057] [Cited by in RCA: 1008] [Article Influence: 56.0] [Reference Citation Analysis (0)] |
| 75. | Hanstock TL, Mallet PE, Clayton EH. Increased plasma d-lactic acid associated with impaired memory in rats. Physiol Behav. 2010;101:653-659. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 76. | Lundberg R, Clausen SK, Pang W, Nielsen DS, Möller K, Josefsen KE, Hansen AK. Gastrointestinal microbiota and local inflammation during oxazolone-induced dermatitis in BALB/cA mice. Comp Med. 2012;62:371-380. [PubMed] |
| 77. | Zackular JP, Baxter NT, Iverson KD, Sadler WD, Petrosino JF, Chen GY, Schloss PD. The gut microbiome modulates colon tumorigenesis. MBio. 2013;4:e00692-e00613. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 433] [Cited by in RCA: 534] [Article Influence: 44.5] [Reference Citation Analysis (0)] |
| 78. | Hedlund BP, Gosink JJ, Staley JT. Verrucomicrobia div. nov., a new division of the bacteria containing three new species of Prosthecobacter. Antonie Van Leeuwenhoek. 1997;72:29-38. [PubMed] |
| 79. | Derrien M, Collado MC, Ben-Amor K, Salminen S, de Vos WM. The Mucin degrader Akkermansia muciniphila is an abundant resident of the human intestinal tract. Appl Environ Microbiol. 2008;74:1646-1648. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 389] [Cited by in RCA: 503] [Article Influence: 27.9] [Reference Citation Analysis (0)] |
| 80. | Everard A, Belzer C, Geurts L, Ouwerkerk JP, Druart C, Bindels LB, Guiot Y, Derrien M, Muccioli GG, Delzenne NM. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc Natl Acad Sci USA. 2013;110:9066-9071. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2639] [Cited by in RCA: 3306] [Article Influence: 275.5] [Reference Citation Analysis (0)] |
| 81. | Marietta EV, Gomez AM, Yeoman C, Tilahun AY, Clark CR, Luckey DH, Murray JA, White BA, Kudva YC, Rajagopalan G. Low incidence of spontaneous type 1 diabetes in non-obese diabetic mice raised on gluten-free diets is associated with changes in the intestinal microbiome. PLoS One. 2013;8:e78687. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 96] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 82. | Russell SL, Gold MJ, Hartmann M, Willing BP, Thorson L, Wlodarska M, Gill N, Blanchet MR, Mohn WW, McNagny KM. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 2012;13:440-447. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 582] [Cited by in RCA: 666] [Article Influence: 51.2] [Reference Citation Analysis (0)] |
| 83. | Engkilde K, Buschard K, Hansen AK, Menné T, Johansen JD. Prevention of diabetes in NOD mice by repeated exposures to a contact allergen inducing a sub-clinical dermatitis. PLoS One. 2010;5:e10591. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 9] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 84. | Engkilde K, Johansen JD, Hansen AK, Menné T, Buschard K. Prevention of type 1 diabetes by inducing subclinical dermatitis on a small area. Diabetes Metab Res Rev. 2011;27:954-958. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 85. | Kang CS, Ban M, Choi EJ, Moon HG, Jeon JS, Kim DK, Park SK, Jeon SG, Roh TY, Myung SJ. Extracellular vesicles derived from gut microbiota, especially Akkermansia muciniphila, protect the progression of dextran sulfate sodium-induced colitis. PLoS One. 2013;8:e76520. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 313] [Cited by in RCA: 403] [Article Influence: 33.6] [Reference Citation Analysis (0)] |
| 86. | Ganesh BP, Klopfleisch R, Loh G, Blaut M. Commensal Akkermansia muciniphila exacerbates gut inflammation in Salmonella Typhimurium-infected gnotobiotic mice. PLoS One. 2013;8:e74963. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 269] [Cited by in RCA: 374] [Article Influence: 31.2] [Reference Citation Analysis (0)] |
| 87. | Hansen CH, Holm TL, Krych Ł, Andresen L, Nielsen DS, Rune I, Hansen AK, Skov S. Gut microbiota regulates NKG2D ligand expression on intestinal epithelial cells. Eur J Immunol. 2013;43:447-457. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 47] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 88. | Kuwahara T, Ogura Y, Oshima K, Kurokawa K, Ooka T, Hirakawa H, Itoh T, Nakayama-Imaohji H, Ichimura M, Itoh K. The lifestyle of the segmented filamentous bacterium: a non-culturable gut-associated immunostimulating microbe inferred by whole-genome sequencing. DNA Res. 2011;18:291-303. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 70] [Cited by in RCA: 69] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 89. | Margulis L, Jorgensen JZ, Dolan S, Kolchinsky R, Rainey FA, Lo SC. The Arthromitus stage of Bacillus cereus: intestinal symbionts of animals. Proc Natl Acad Sci USA. 1998;95:1236-1241. [PubMed] |
| 90. | Ivanov II, Littman DR. Segmented filamentous bacteria take the stage. Mucosal Immunol. 2010;3:209-212. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 141] [Cited by in RCA: 136] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 91. | Ivanov II, Atarashi K, Manel N, Brodie EL, Shima T, Karaoz U, Wei D, Goldfarb KC, Santee CA, Lynch SV. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell. 2009;139:485-498. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3529] [Cited by in RCA: 3519] [Article Influence: 219.9] [Reference Citation Analysis (0)] |
| 92. | Ivanov II, Frutos Rde L, Manel N, Yoshinaga K, Rifkin DB, Sartor RB, Finlay BB, Littman DR. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe. 2008;4:337-349. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1480] [Cited by in RCA: 1399] [Article Influence: 82.3] [Reference Citation Analysis (0)] |
| 93. | Kriegel MA, Sefik E, Hill JA, Wu HJ, Benoist C, Mathis D. Naturally transmitted segmented filamentous bacteria segregate with diabetes protection in nonobese diabetic mice. Proc Natl Acad Sci USA. 2011;108:11548-11553. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 355] [Cited by in RCA: 342] [Article Influence: 24.4] [Reference Citation Analysis (0)] |
| 94. | Balish E, Warner T. Enterococcus faecalis induces inflammatory bowel disease in interleukin-10 knockout mice. Am J Pathol. 2002;160:2253-2257. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 241] [Cited by in RCA: 246] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 95. | Kim SC, Tonkonogy SL, Albright CA, Tsang J, Balish EJ, Braun J, Huycke MM, Sartor RB. Variable phenotypes of enterocolitis in interleukin 10-deficient mice monoassociated with two different commensal bacteria. Gastroenterology. 2005;128:891-906. [PubMed] |
| 96. | Steck N, Hoffmann M, Sava IG, Kim SC, Hahne H, Tonkonogy SL, Mair K, Krueger D, Pruteanu M, Shanahan F. Enterococcus faecalis metalloprotease compromises epithelial barrier and contributes to intestinal inflammation. Gastroenterology. 2011;141:959-971. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 198] [Cited by in RCA: 234] [Article Influence: 16.7] [Reference Citation Analysis (0)] |
| 97. | Duncan SH, Hold GL, Harmsen HJ, Stewart CS, Flint HJ. Growth requirements and fermentation products of Fusobacterium prausnitzii, and a proposal to reclassify it as Faecalibacterium prausnitzii gen. nov., comb. nov. Int J Syst Evol Microbiol. 2002;52:2141-2146. [PubMed] |
| 98. | Sokol H, Pigneur B, Watterlot L, Lakhdari O, Bermúdez-Humarán LG, Gratadoux JJ, Blugeon S, Bridonneau C, Furet JP, Corthier G. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA. 2008;105:16731-16736. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2747] [Cited by in RCA: 3195] [Article Influence: 187.9] [Reference Citation Analysis (0)] |
| 99. | Paturi G, Mandimika T, Butts CA, Zhu S, Roy NC, McNabb WC, Ansell J. Influence of dietary blueberry and broccoli on cecal microbiota activity and colon morphology in mdr1a(-/-) mice, a model of inflammatory bowel diseases. Nutrition. 2012;28:324-330. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 77] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 100. | Carlsson AH, Yakymenko O, Olivier I, Håkansson F, Postma E, Keita AV, Söderholm JD. Faecalibacterium prausnitzii supernatant improves intestinal barrier function in mice DSS colitis. Scand J Gastroenterol. 2013;48:1136-1144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 119] [Cited by in RCA: 148] [Article Influence: 12.3] [Reference Citation Analysis (0)] |
| 101. | Hansen CH, Frøkiær H, Christensen AG, Bergström A, Licht TR, Hansen AK, Metzdorff SB. Dietary xylooligosaccharide downregulates IFN-γ and the low-grade inflammatory cytokine IL-1β systemically in mice. J Nutr. 2013;143:533-540. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 71] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 102. | Lam V, Su J, Koprowski S, Hsu A, Tweddell JS, Rafiee P, Gross GJ, Salzman NH, Baker JE. Intestinal microbiota determine severity of myocardial infarction in rats. FASEB J. 2012;26:1727-1735. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 185] [Cited by in RCA: 234] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 103. | McCarthy J, O’Mahony L, O’Callaghan L, Sheil B, Vaughan EE, Fitzsimons N, Fitzgibbon J, O’Sullivan GC, Kiely B, Collins JK. Double blind, placebo controlled trial of two probiotic strains in interleukin 10 knockout mice and mechanistic link with cytokine balance. Gut. 2003;52:975-980. [PubMed] |
| 104. | Schwarzer M, Srutkova D, Schabussova I, Hudcovic T, Akgün J, Wiedermann U, Kozakova H. Neonatal colonization of germ-free mice with Bifidobacterium longum prevents allergic sensitization to major birch pollen allergen Bet v 1. Vaccine. 2013;31:5405-5412. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 105. | Brinkman BM, Becker A, Ayiseh RB, Hildebrand F, Raes J, Huys G, Vandenabeele P. Gut microbiota affects sensitivity to acute DSS-induced colitis independently of host genotype. Inflamm Bowel Dis. 2013;19:2560-2567. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 58] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 106. | Rath HC, Wilson KH, Sartor RB. Differential induction of colitis and gastritis in HLA-B27 transgenic rats selectively colonized with Bacteroides vulgatus or Escherichia coli. Infect Immun. 1999;67:2969-2974. [PubMed] |
| 107. | Brugman S, Klatter FA, Visser JT, Wildeboer-Veloo AC, Harmsen HJ, Rozing J, Bos NA. Antibiotic treatment partially protects against type 1 diabetes in the Bio-Breeding diabetes-prone rat. Is the gut flora involved in the development of type 1 diabetes? Diabetologia. 2006;49:2105-2108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 256] [Cited by in RCA: 249] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 108. | Xu J, Mahowald MA, Ley RE, Lozupone CA, Hamady M, Martens EC, Henrissat B, Coutinho PM, Minx P, Latreille P. Evolution of symbiotic bacteria in the distal human intestine. PLoS Biol. 2007;5:e156. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 396] [Cited by in RCA: 409] [Article Influence: 22.7] [Reference Citation Analysis (0)] |
| 109. | Martens EC, Chiang HC, Gordon JI. Mucosal glycan foraging enhances fitness and transmission of a saccharolytic human gut bacterial symbiont. Cell Host Microbe. 2008;4:447-457. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 799] [Cited by in RCA: 706] [Article Influence: 41.5] [Reference Citation Analysis (0)] |
| 110. | Nakano V, Gomes DA, Arantes RM, Nicoli JR, Avila-Campos MJ. Evaluation of the pathogenicity of the Bacteroides fragilis toxin gene subtypes in gnotobiotic mice. Curr Microbiol. 2006;53:113-117. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 20] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 111. | Wu S, Rhee KJ, Albesiano E, Rabizadeh S, Wu X, Yen HR, Huso DL, Brancati FL, Wick E, McAllister F. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med. 2009;15:1016-1022. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1113] [Cited by in RCA: 1303] [Article Influence: 81.4] [Reference Citation Analysis (0)] |
| 112. | Mazmanian SK, Round JL, Kasper DL. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature. 2008;453:620-625. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1625] [Cited by in RCA: 1773] [Article Influence: 104.3] [Reference Citation Analysis (0)] |
| 113. | Hsiao EY, McBride SW, Hsien S, Sharon G, Hyde ER, McCue T, Codelli JA, Chow J, Reisman SE, Petrosino JF. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell. 2013;155:1451-1463. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2084] [Cited by in RCA: 2339] [Article Influence: 194.9] [Reference Citation Analysis (0)] |
| 114. | Ukena SN, Singh A, Dringenberg U, Engelhardt R, Seidler U, Hansen W, Bleich A, Bruder D, Franzke A, Rogler G. Probiotic Escherichia coli Nissle 1917 inhibits leaky gut by enhancing mucosal integrity. PLoS One. 2007;2:e1308. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 310] [Cited by in RCA: 348] [Article Influence: 19.3] [Reference Citation Analysis (0)] |
| 115. | Turroni F, Taverniti V, Ruas-Madiedo P, Duranti S, Guglielmetti S, Lugli GA, Gioiosa L, Palanza P, Margolles A, van Sinderen D. Bifidobacterium bifidum PRL2010 modulates the host innate immune response. Appl Environ Microbiol. 2014;80:730-740. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 65] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 116. | Hufeldt MR, Nielsen DS, Vogensen FK, Midtvedt T, Hansen AK. Family relationship of female breeders reduce the systematic inter-individual variation in the gut microbiota of inbred laboratory mice. Lab Anim. 2010;44:283-289. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 38] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 117. | Pang W, Stradiotto D, Krych L, Karlskov-Mortensen P, Vogensen FK, Nielsen DS, Fredholm M, Hansen AK. Selective inbreeding does not increase gut microbiota similarity in BALB/c mice. Lab Anim. 2012;46:335-337. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 118. | Hansen CH, Metzdorff SB, Hansen AK. Customizing laboratory mice by modifying gut microbiota and host immunity in an early “window of opportunity”. Gut Microbes. 2013;4:241-245. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 26] [Article Influence: 2.2] [Reference Citation Analysis (0)] |