Published online Mar 21, 2013. doi: 10.3748/wjg.v19.i11.1718
Revised: January 11, 2013
Accepted: January 23, 2013
Published online: March 21, 2013
Processing time: 189 Days and 3.1 Hours
AIM: To evaluate the utility of measuring epigenetic alterations in pancreatic and biliary fluids in determining molecular markers for pancreatobiliary cancers.
METHODS: DNA was extracted from undiluted pancreatic and biliary fluids. As a surrogate for a genome-wide hypomethylation assay, levels of long interspersed nuclear element-1 (LINE-1) methylation were analyzed using bisulfite pyrosequencing. CpG island hypermethylation of 10 tumor-associated genes, aryl-hydrocarbon receptor repressor, adenomatous polyposis coli, calcium channel, voltage dependent, T type α1G subunit, insulin-like growth factor 2, O-6-methyl-guanine-DNA methyltransferase, neurogenin 1, CDKN2A, runt-related transcription factor 3 (RUNX3), secreted frizzled-related protein 1, and ubiquitin carboxyl-terminal esterase L1 (UCHL1), was analyzed using MethyLight. To examine the role of CpG methylation and histone deacetylation in the silencing of UCHL1, human gallbladder carcinoma cell lines and pancreatic carcinoma cell lines were treated with 2 or 5 μmol/L 5-AZA-dC for 72 h or 100 nmol/L Trichostatin A for 24 h. After the treatment, UCHL1 expression was analyzed by real-time reverse transcription-polymerase chain reaction.
RESULTS: Pancreatobiliary cancers exhibited significantly lower LINE-1 methylation levels in pancreatic and biliary fluids than did noncancerous pancreatobiliary disease (58.7% ± 4.3% vs 61.7% ± 2.2%, P = 0.027; 53.8% ± 6.6% vs 57.5% ± 1.7%, P = 0.007); however, LINE-1 hypomethylation was more evident in pancreatic cancer tissues than in pancreatic fluids (45.4% ± 5.5% vs 58.7% ± 4.3%, P < 0.001). CpG island hypermethylation of tumor-associated genes was detected at various frequencies, but it was not correlated with LINE-1 hypomethylation. Hypermethylation of the UCHL1 gene was cancer-specific and most frequently detected in pancreatic (67%) or biliary (70%) fluids from patients with pancreatobiliary cancer. As a single marker, hypermethylation of the UCHL1 gene in pancreatic and biliary fluids was most useful for the detection of pancreatic and pancreatobiliary cancers, respectively (100% specificity). Hypermethylation of the UCHL1 and RUNX3 genes in pancreatic and biliary fluids was the most useful combined marker for pancreatic (87% sensitivity and 100% specificity) and pancreatobiliary (97% sensitivity and 100% specificity) cancers. Treatment with a demethylating agent, 5-AZA-2’-deoxycytidine, restored UCHL1 expression in pancreatobiliary cancer cell lines.
CONCLUSION: Our results suggest that hypermethylation of UCHL1 and RUNX3 in pancreatobiliary fluid might be useful for the diagnosis of pancreatobiliary cancers.
- Citation: Kato N, Yamamoto H, Adachi Y, Ohashi H, Taniguchi H, Suzuki H, Nakazawa M, Kaneto H, Sasaki S, Imai K, Shinomura Y. Cancer detection by ubiquitin carboxyl-terminal esterase L1 methylation in pancreatobiliary fluids. World J Gastroenterol 2013; 19(11): 1718-1727
- URL: https://www.wjgnet.com/1007-9327/full/v19/i11/1718.htm
- DOI: https://dx.doi.org/10.3748/wjg.v19.i11.1718
Despite recent advances in diagnosis and treatment, the prognosis of patients with pancreatobiliary cancer is still poor. Surgical resection is possible in only a small proportion of patients[1,2]. Consequently, elucidation of the biological characteristics of pancreatobiliary carcinomas is necessary to improve the prognosis of patients and to devise better treatment strategies. Various genetic and epigenetic alterations play a role in pancreatobiliary cancer[3-6].
Two contradicting epigenetic alterations often coexist in cancer: global or genome-wide hypomethylation, which is mainly observed in repetitive sequences within the genome, and regional hypermethylation, which is frequently associated with CpG islands within gene promoters[7]. Hypermethylation of CpG islands is a common feature of cancer that is associated with gene silencing[7,8]. A number of genes are aberrantly methylated and silenced in pancreatobiliary cancer that are rarely methylated in non-neoplastic counterparts[4-6,8], and this methylation is detectable in pancreatic and/or biliary fluids[9-12]. The detection and/or quantification of these alterations in pancreatic and/or biliary fluids has promise for facilitating the differentiation of benign and malignant pancreatic and/or biliary strictures.
In contrast to CpG islands, repetitive DNA elements are normally heavily methylated in somatic tissues. Approximately 45% of the human genome is composed of repetitive sequences, including long interspersed nuclear elements (LINEs) and short interspersed nuclear elements[13]. Liquid chromatography-mass spectrometry analysis has shown that levels of LINE-1 methylation strongly correlate with methyl cytosine content. This strong correlation enables LINE-1 methylation to be used as a proxy for genome-wide methylation[14]. Moreover, LINE-1 hypomethylation is known to occur during the development and progression of various human malignancies[15,16]. Additionally, we recently reported that LINE-1 hypomethylation correlates significantly with the aggressiveness of gastrointestinal stromal tumors and that LINE-1 methylation could be a useful marker for risk assessment[17]. Array comparative genomic hybridization analysis revealed a significant correlation between LINE-1 hypomethylation and chromosomal aberrations[17]. Chromosomal gains and losses are also common in pancreatic and biliary cancers[18,19]; their detection by fluorescence in situ hybridization modestly improves the prediction of cancer using biliary brushings[20,21]. Gene hypomethylation has been reported to be a frequent epigenetic event in pancreatic cancer and is commonly associated with the overexpression of affected genes[22]. A previous study showed that hypomethylation is more common in carcinoid tumors than in pancreatic endocrine tumors and is associated with clinicopathologic features, including lymph node metastasis, as well as genetic and epigenetic alterations in these tumors[23].
To date, however, only a few groups have reported the methylation of LINE-1 and/or other repetitive sequences in pancreatobiliary cancer[23,24], and there are no published studies analyzing LINE-1 methylation in pancreatic and/or biliary fluids. We found correlations between the level of LINE-1 methylation and the methylation of other repetitive sequences[17]. In this study, we analyzed LINE-1 methylation and its relationship with hypermethylation of CpG islands in pancreatic and biliary fluids, and we investigated whether the detection and/or quantification of these epigenetic alterations can be used as markers for pancreatobiliary cancer.
Pancreatic and biliary fluids were obtained at the time of endoscopic retrograde cholangiopancreatography (ERCP) and ERCP/percutaneous transhepatic cholangiography and drainage, respectively[9-12]. Pancreatic and biliary fluids were collected from 30 and 48 patients, respectively. Informed consent was obtained from each subject. Tumors were classified according to the tumor-node-metastasis classification system of the International Union Against Cancer. The absence of cancer was based on clinical evaluation and follow-up of one or more years. Human gallbladder carcinoma cell lines TGBC1TKB and TGBC2TKB and pancreatic carcinoma cell lines PANC-1, PK-1, PK-45P and PK59 were purchased from Riken Cell Bank (Tsukuba, Japan). Cells were cultured in RPMI1640 or DMEM supplemented with 10% fetal bovine serum.
DNA was extracted from undiluted pancreatic and biliary fluids using a DNeasy Tissue Kit. Extracted DNA was quantified, and 500 ng of DNA was modified with sodium bisulfite using a MethylampTM DNA modification kit.
Bisulfite-pyrosequencing analysis was performed as described previously[17,24]. Briefly, polymerase chain reaction (PCR) was run in a 25 μL volume containing 50 ng bisulfite-treated DNA, 1× MSP buffer, 1.25 mmol/L dNTP, 0.4 μmol/L of each primer and 0.5 U of JumpStart REDTaq DNA Polymerase. The PCR protocol for bisulfite sequencing entailed 5 min at 95 °C; 40 cycles of 1 min at 95 °C, 1 min at 60 °C and 1 min at 72 °C; and a 7 min final extension at 72 °C. The biotinylated PCR product was purified, made single-stranded and used as a template in a pyrosequencing reaction run according to the manufacturer’s instructions. The PCR products were bound to Streptavidin Sepharose beads HP; then, the beads containing the immobilized PCR product were purified, washed and denatured using a 0.2 mol/L NaOH solution. After adding 0.3 μmol/L sequencing primer to the purified PCR product, pyrosequencing was performed using a PSQ96MA system and Pyro Q-CpG software. Primer sequences for LINE-1 methylation were as previously described[25].
The MethyLight assay was performed as previously described[25,26]. Based on previous studies and our preliminary results, we analyzed 10 promoter CpG island loci: aryl-hydrocarbon receptor repressor, adenomatous polyposis coli, calcium channel, voltage dependent, T type α1G subunit, insulin-like growth factor 2, O-6-methylguanine-DNA methyltransferase, neurogenin 1, CDKN2A, runt-related transcription factor 3 (RUNX3), secreted frizzled-related protein 1, and ubiquitin carboxyl-terminal esterase L1 (UCHL1). β-actin was used as the internal reference gene to quantify modified DNA levels in the samples[26]. Primers, probes and the percentage of methylated reference (PMR, i.e., the degree of methylation) were as previously described[27-29]. We used a PMR cutoff of 4 to distinguish methylation-positive (PMR > 4) from methylation-negative (PMR ≤ 4) samples based on previously validated data[29].
To examine the role of CpG methylation and histone deacetylation in the silencing of UCHL1, cancer cells were treated with 2 or 5 mol/L 5-AZA-2’-deoxycytidine (5-AZA-dC) (Sigma) for 72 h or 100 nmol/L Trichostatin A (TSA) for 24 h. The cells were also treated with 2 μmol/L 5-AZA-dC for 72 h, followed by 100 nmol/L TSA for an additional 24 h. The timing and sequencing of 5-AZA-dC and/or TSA were based on similar preliminary studies, as well as published studies[30]. After the treatment, UCHL1 expression was analyzed by real-time RT-PCR.
Total RNA from cell lines was extracted using an extraction kit. cDNA was synthesized from 1 μg of total RNA using SuperScript III reverse transcriptase with random hexamers. qRT-PCR was performed using the TaqMan real-time PCR system as previously described[31]. A comparative threshold cycle (CT) was used to determine the gene expression relative to the control (calibrator). Control reactions were performed without reverse transcriptase.
Mean methylation levels of LINE-1 were compared using t tests, the Welch test, or one-way ANOVA with a post hoc Games-Howell test. LINE-1 methylation levels and hypermethylation of tumor-associated genes were assessed for associations with clinicopathological parameters using t tests, the Welch test, the χ2 two-tailed test, Fisher’s exact test, the Mann-Whitney test, or one-way ANOVA. A P value < 0.05 was considered statistically significant. A P value between 0.05 and 0.10 was considered to indicate a trend toward an association.
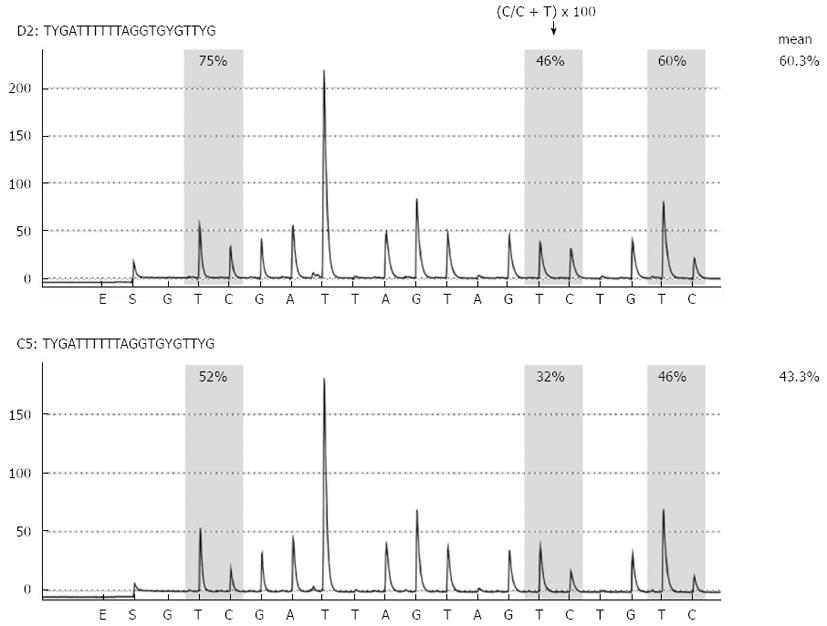
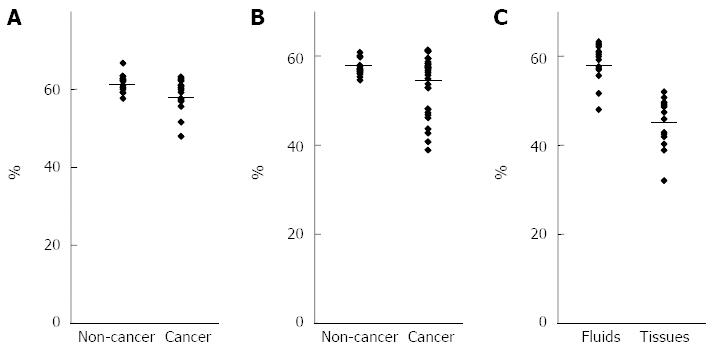
We performed bisulfite pyrosequencing to quantitatively analyze LINE-1 promoter methylation as a surrogate for genome-wide methylation (Figure 1). The mean level of LINE-1 methylation in pancreatic fluids was slightly but significantly lower in patients with pancreatic cancer than in those with noncancerous pancreatic disease (Figure 2). The mean level of LINE-1 methylation in biliary fluids was significantly lower in patients with pancreatobiliary cancer than in those with noncancerous pancreatobiliary disease. There was no correlation between LINE-1 methylation levels and clinicopathological characteristics in patients with pancreatic or pancreatobiliary cancers. The mean level of LINE-1 methylation in pancreatic cancer tissues was significantly lower than that in pancreatic fluids from the corresponding patients.
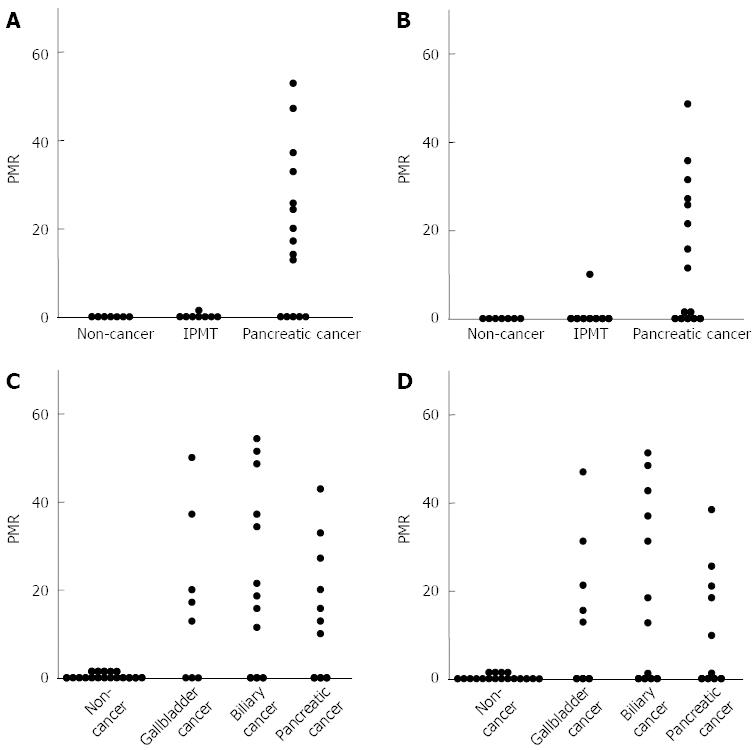
We next assessed the methylation levels of CpG islands of well-characterized tumor-suppressor and tumor-associated genes. Using MethyLight assays, we analyzed 10 genes. CpG island hypermethylation of tumor-associated genes was detected at various frequencies. The results are summarized on the basis of each individual marker (Tables 1 and 2). Methylation of several genes, such as UCHL1 and RUNX3, was cancer-specific (Figure 3). Some cancer samples showed methylation in many genes, suggesting that these cancers have a CpG island hypermethylator phenotype. We failed to find any significant correlation between the methylation of these genes and clinicopathological features or between CpG island methylation and LINE-1 methylation levels.
| Age (yr) | Sex | Stage | UCHL1 | RUNX3 | CDKN2A | IGF2 | CACNA1G | AHRR | SFRP1 | MGMT | APC | NEUROG1 | |
| NC1 | 68 | M | MN | MN | MN | MN | MN | MN | MP | MP | MN | MN | |
| NC2 | 65 | M | MN | MN | MN | MN | MP | MN | MN | MN | MN | MN | |
| NC3 | 59 | M | MN | MN | MP | MN | MN | MN | MN | MN | MN | MP | |
| NC4 | 70 | F | MN | MN | MN | MP | MN | MN | MN | MN | MN | MN | |
| NC5 | 43 | F | MN | MN | MN | MN | MN | MN | MN | MP | MN | MN | |
| NC6 | 55 | M | MN | MN | MN | MN | MP | MN | MN | MN | MN | MN | |
| NC7 | 49 | M | MN | MN | MN | MN | MN | MP | MN | MN | MP | MN | |
| MN | MN | MN | MN | MN | MN | MN | MN | MN | MN | ||||
| IP1 | 71 | F | MN | MN | MP | MN | MN | MN | MN | MN | MN | MN | |
| IP2 | 79 | F | MN | MN | MN | MN | MN | MN | MN | MN | MN | MP | |
| IP3 | 76 | M | MN | MP | MN | MN | MN | MN | MN | MP | MN | MN | |
| IP4 | 75 | F | MN | MN | MN | MP | MN | MP | MP | MN | MP | MN | |
| IP5 | 84 | F | MN | MN | MN | MN | MN | MN | MP | MN | MN | MN | |
| IP6 | 52 | M | MN | MN | MP | MN | MN | MN | MP | MP | MN | MN | |
| IP7 | 62 | M | MN | MN | MN | MP | MN | MN | MN | MP | MP | MN | |
| IP8 | 63 | M | MN | MN | MP | MN | MN | MN | MN | MP | MN | MN | |
| MN | MN | MN | MN | MN | MN | MN | MN | MN | MN | ||||
| PC1 | 66 | F | IIA | MP | MN | MN | MN | MN | MN | MN | MP | MN | MN |
| PC2 | 54 | F | III | MN | MP | MN | MN | MN | MN | MN | MN | MN | MN |
| PC3 | 66 | M | IIB | MP | MP | MN | MN | MN | MN | MP | MN | MP | MP |
| PC4 | 67 | M | IIB | MP | MN | MN | MN | MP | MN | MN | MN | MP | MN |
| PC5 | 71 | M | IIB | MN | MP | MP | MN | MN | MN | MN | MN | MN | MN |
| PC6 | 49 | F | IIA | MP | MP | MN | MN | MN | MN | MP | MN | MP | MP |
| PC7 | 73 | F | IIB | MP | MN | MN | MN | MN | MP | MP | MN | MN | MN |
| PC8 | 75 | M | III | MP | MP | MN | MN | MN | MN | MN | MN | MP | MN |
| PC9 | 59 | M | IIB | MN | MN | MN | MN | MP | MN | MP | MP | MP | MP |
| PC10 | 78 | F | IIA | MP | MP | MP | MN | MN | MN | MP | MP | MN | MP |
| PC11 | 75 | F | IIB | MN | MN | MN | MP | MN | MN | MP | MN | MN | MN |
| PC12 | 66 | M | IV | MP | MP | MP | MN | MN | MN | MP | MN | MP | MP |
| PC13 | 70 | F | IV | MP | MN | MN | MN | MN | MP | MP | MP | MP | MP |
| PC14 | 68 | M | IB | MN | MP | MN | MN | MN | MN | MN | MP | MN | MN |
| PC15 | 62 | M | IIB | MP | MN | MP | MN | MN | MN | MN | MN | MN | MN |
| Age (yr) | Sex | Stage | UCHL1 | RUNX3 | CDKN2A | IGF2 | CACNA1G | AHRR | SFRP1 | MGMT | APC | NEUROG1 | |
| NC8 | 77 | M | MN | MN | MN | MN | MN | MN | MN | MP | MN | MN | |
| NC9 | 69 | F | MN | MN | MN | MN | MN | MN | MP | MN | MN | MN | |
| NC10 | 76 | M | MN | MN | MN | MN | MN | MN | MN | MP | MN | MN | |
| NC11 | 58 | M | MN | MN | MN | MN | MN | MN | MN | MN | MN | MN | |
| NC12 | 41 | M | MN | MN | MN | MN | MN | MP | MN | MP | MN | MN | |
| NC13 | 66 | M | MN | MN | MN | MN | MN | MP | MP | MP | MP | MP | |
| NC14 | 69 | F | MN | MN | MN | MN | MN | MP | MP | MP | MN | MP | |
| NC15 | 71 | F | MN | MN | MN | MN | MN | MN | MN | MP | MP | MN | |
| NC16 | 59 | M | MN | MN | MN | MN | MN | MP | MP | MN | MN | MN | |
| NC17 | 54 | F | MN | MN | MN | MN | MN | MN | MN | MN | MP | MP | |
| NC18 | 67 | M | MN | MN | MN | MN | MN | MP | MP | MN | MN | MN | |
| NC19 | 85 | M | MN | MN | MN | MN | MN | MN | MN | MN | MN | MN | |
| NC20 | 73 | F | MN | MN | MN | MN | MN | MN | MP | MN | MN | MN | |
| NC21 | 80 | F | MN | MN | MN | MN | MN | MN | MN | MP | MN | MN | |
| NC22 | 74 | F | MN | MN | MN | MN | MN | MN | MN | MP | MN | MN | |
| NC23 | 52 | M | MN | MN | MN | MN | MN | MN | MN | MN | MN | MN | |
| NC24 | 64 | M | MN | MN | MN | MN | MN | MN | MP | MP | MP | MN | |
| NC25 | 64 | F | MN | MN | MN | MN | MN | MP | MP | MN | MN | MN | |
| MN | MN | MN | MN | MN | MN | MN | MN | MN | MN | ||||
| GB1 | 75 | F | IIIB | MP | MP | MP | MP | MP | MP | MN | MN | MN | MN |
| GB | 76 | F | IIIB | MN | MP | MP | MP | MP | MP | MN | MN | MP | MN |
| GB3 | 62 | M | IIIA | MP | MP | MP | MP | MN | MN | MP | MP | MN | MN |
| GB4 | 67 | M | IVA | MN | MP | MN | MN | MN | MN | MN | MN | MN | MN |
| GB5 | 59 | F | IIIB | MP | MN | MP | MN | MN | MP | MP | MN | MP | MP |
| GB6 | 63 | F | II | MP | MN | MP | MN | MN | MP | MP | MN | MP | MP |
| GB7 | 77 | M | I | MN | MP | MP | MN | MN | MN | MP | MP | MN | MN |
| GB8 | 78 | M | IIIA | MP | MN | MP | MN | MN | MN | MP | MP | MN | MN |
| BC1 | 73 | M | I | MN | MP | MN | MP | MN | MP | MP | MN | MN | MN |
| BC2 | 80 | F | IIA | MP | MP | MN | MN | MN | MP | MP | MP | MN | MN |
| BC3 | 71 | M | IIB | MP | MN | MN | MN | MP | MP | MP | MN | MN | MN |
| BC4 | 75 | M | III | MP | MN | MN | MN | MN | MN | MP | MP | MN | MN |
| BC5 | 77 | M | IIB | MN | MP | MN | MN | MN | MN | MN | MP | MN | MN |
| BC6 | 65 | M | IIA | MP | MN | MP | MN | MN | MN | MN | MN | MN | MN |
| BC7 | 72 | M | IIA | MP | MN | MN | MN | MN | MN | MP | MN | MN | MN |
| BC8 | 73 | F | IV | MP | MP | MN | MN | MN | MP | MN | MN | MN | MN |
| BC9 | 76 | F | IIA | MP | MP | MN | MP | MP | MP | MP | MN | MN | MN |
| BC10 | 74 | M | IIB | MP | MN | MN | MN | MN | MP | MN | MP | MN | MP |
| BC11 | 66 | M | III | MP | MN | MN | MN | MN | MN | MN | MN | MP | MN |
| BC12 | 58 | F | IIB | MN | MP | MN | MN | MN | MN | MN | MN | MN | MN |
| PC1 | 66 | F | IIA | MP | MN | MN | MN | MN | MN | MP | MP | MN | MN |
| PC5 | 71 | M | IIB | MN | MP | MP | MN | MN | MN | MN | MP | MN | MN |
| PC7 | 73 | F | IIB | MP | MN | MN | MN | MN | MP | MN | MN | MN | MN |
| PC8 | 75 | M | III | MP | MP | MN | MN | MN | MN | MN | MP | MN | MN |
| PC13 | 70 | F | IV | MP | MN | MN | MN | MN | MP | MP | MP | MP | MN |
| PC16 | 59 | M | IIA | MP | MP | MN | MN | MP | MN | MN | MN | MN | MN |
| PC17 | 80 | F | IIB | MP | MP | MP | MP | MN | MP | MP | MN | MN | MN |
| PC18 | 67 | M | IB | MN | MP | MN | MN | MN | MP | MN | MP | MN | MN |
| PC19 | 63 | M | IIA | MP | MN | MN | MN | MN | MN | MN | MP | MP | MN |
| PC20 | 78 | F | IIB | MN | MN | MP | MN | MN | MP | MN | MN | MN | MN |
Among the cancer-specific hypermethylated genes in pancreatic fluids, the UCHL1 gene was most frequently (67%) detected in pancreatic cancer and served as the most useful single marker for the detection of pancreatic cancer (Figure 3A). Hypermethylation of the UCHL1 and RUNX3 genes in pancreatic fluids was the most useful combined marker for the detection of pancreatic cancer.
Among the cancer-specific hypermethylated genes in biliary fluids, the UCHL1 gene was most frequently (70%) detected in pancreatobiliary cancer and served as the most useful single marker for the detection of pancreatobiliary cancer (Figure 3C). Hypermethylation of the UCHL1 and RUNX3 genes in biliary fluids was the most useful combined marker for detection of pancreatobiliary cancer.
The pancreatic and biliary fluids obtained from pancreatic cancer patients were compared with regard to the methylation patterns of 10 tumor-associated genes (n = 5). The methylation patterns were similar in both the pancreatic and biliary fluids from the same patients (Tables 1 and 2). The methylation patterns of the UCHL1 and RUNX3 genes were identical in the pancreatic and biliary fluids from the same patients.
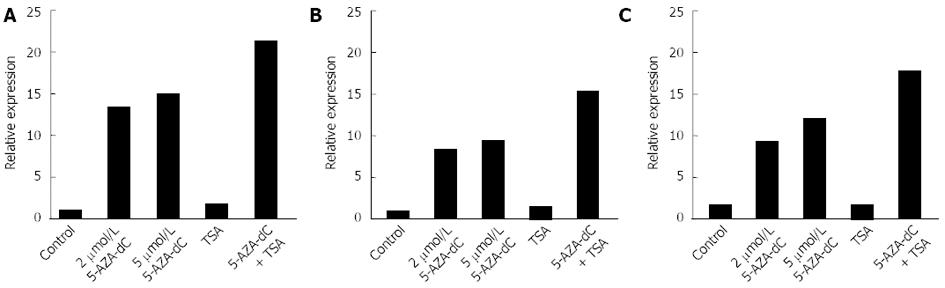
To further examine the role of CpG methylation and histone deacetylation in silencing UCHL1, cancer cells were treated with 5-AZA-dC and/or TSA. 5-AZA-dC restored UCHL1 expression, and combined treatment with 5-AZA-dC and TSA restored UCHL1 expression synergistically at the mRNA level in pancreatobiliary cancer cell lines (Figure 4 and data not shown). TSA alone did not restore UCHL1 expression in cell lines.
In the present study, we found that the levels of methylation of LINE-1 were reduced in pancreatobiliary cancers compared to those in noncancerous pancreatobiliary disease. Genome-wide hypomethylation is known to be a common feature of human cancer, and genome-wide hypomethylation has recently been studied in various human malignancies using LINE-1 and other repetitive sequences as surrogates. Our results suggest that pancreatobiliary cancers exhibit a pattern of genome-wide hypomethylation that can be detected using pancreatic and biliary fluids.
Genome-wide hypomethylation is thought to be associated with tumor malignancy through a variety of mechanisms. For example, global hypomethylation is associated with genomic instability[32], which may confer a poor prognosis. Hypomethylation can also lead to the activation of proto-oncogenes, endogenous retroviruses or transposable elements; such transcriptional dysregulation could affect tumor aggressiveness. Although we found correlations between the level of LINE-1 methylation and the methylation of other repetitive sequences[17], it is possible that there are functional and/or biological differences in the regulation of repetitive DNA sequences. Further analysis is necessary to clarify the role of genome-wide hypomethylation in pancreatobiliary cancers.
The LINE-1 hypomethylation in pancreatic cancers determined using pancreatic fluids was less significant than that determined using tissue samples, although the number of samples analyzed was limited. It is possible that some of the pancreatic fluid samples did not contain sufficient concentrations of cancer DNA[12]. Given the relatively poor diagnostic yield of cytology in this setting, a problem that is likely to be related to the highly scirrhous nature of pancreatic ductal adenocarcinomas, sample adequacy is likely to be one of the limiting factors in the molecular analysis of these samples[12]. Serum LINE-1 hypomethylation has been reported to be a potential prognostic marker for hepatocellular carcinoma[33]. It would be interesting to analyze serum LINE-1 methylation levels in patients with pancreatobiliary cancers.
CpG island hypermethylation of tumor-associated genes was detected at various frequencies in pancreatobiliary cancers using pancreatobiliary fluids. Although genome-wide hypomethylation and regional hypermethylation of 5’ CpG islands are common features of neoplasias, the link between the two remains controversial[17]. In the current study, we did not find a significant correlation between 5’ CpG island hypermethylation of tumor-associated genes and global hypomethylation.
Hypermethylation of the UCHL1 gene was cancer-specific and most frequently detected in pancreatobiliary cancers. Hypermethylation of the UCHL1 gene in pancreatic and biliary fluids was the most useful single marker of pancreatic and pancreatobiliary cancers, respectively. Hypermethylation of the UCHL1 and RUNX3 genes in pancreatic and biliary fluids was the most useful combined marker for pancreatic and pancreatobiliary cancers, respectively. Epigenetic inactivation of UCHL1 has been reported in a variety of human cancers[34]. Epigenetic inactivation of RUNX3 is known to play an important role in the pathogenesis of pancreatobiliary cancer[35,36].
LINE-1 and SAT2 methylation levels have been reported to be significantly lower in extrahepatic cholangiocarcinoma than in normal duct and biliary intraepithelial neoplasias (BilINs). BilINs showed a decrease of SAT2 methylation levels, but no decrease of LINE-1 methylation levels was found compared to those in normal samples[24]. Most of the cancer-specific CpG island hypermethylation is thought to occur in the BilIN stage, before LINE-1 hypomethylation. Our results also suggest that CpG island hypermethylation analyzed in pancreatobiliary fluids is more useful than LINE-1 methylation for the detection of pancreatobiliary cancer.
Importantly, the methylation patterns of 10 tumor-associated genes were similar in both the pancreatic and biliary fluids from the same patients with pancreatic cancer. Moreover, the methylation patterns of the UCHL1 and RUNX3 genes were identical in both the pancreatic and biliary fluids from the same patients. These results further support the notion that hypermethylation of UCHL1 and RUNX3 in pancreatobiliary fluids is a useful marker for the detection of pancreatobiliary cancer.
To confirm the role of epigenetic alterations in transcriptional repression of the UCHL1 gene, we treated pancreatobiliary cancer cell lines, in which UCHL1 was methylated, with 5-AZA-dC alone or in combination with TSA. Treatment with 5-AZA-dC restored the UCHL1 expression in cancer cell lines. Moreover, combined treatment with 5-AZA-dC and TSA restored UCHL1 expression synergistically, indicating that CpG methylation and histone deacetylation play important roles in silencing the UCHL1 gene.
Not only the clinical utility but also the pathobiological effects of nucleic acids in circulation (nucleosomes, DNA, RNA, microRNA etc.) are receiving increasing attention[37,38]. Further analysis is necessary to clarify the possible detrimental effects of nucleic acids in the tumor microenvironment, including the contribution of methylated DNA in pancreatobiliary fluids to disease progression.
In conclusion, our results suggest that hypermethylation of the UCHL1 gene plays a key role in the pathogenesis of pancreatobiliary cancers and that detection of hypermethylation of UCHL1 and RUNX3 in pancreatobiliary fluids is useful for the diagnosis of these malignancies. Our MethyLight panel (UCHL1 and RUNX3) is simple and accurate for differentiating between neoplastic and non-neoplastic samples and compares favorably with other quantitative MSP panels and with the identification of mutant KRAS or telomerase, which have been used previously to differentiate between malignant and benign pancreatic samples[39,40]. Moreover, newer assays that can detect low concentrations of mutations in pancreatic juice[41], as well as novel assays and technologies, are likely to improve the detection of low concentrations of mutant DNA for cancer diagnosis in the future. Although we focused on epigenetic alterations in the current study, a combination of highly specific epigenetic and genetic markers might provide the best diagnostic utility.
Despite recent advances in diagnosis and treatment, the prognosis of patients with pancreatobiliary cancer is still poor. Elucidation of the biological characteristics of these carcinomas has become necessary to improve the prognosis of patients and to devise better treatment strategies.
Roles of epigenetic alterations in pancreatobiliary cancer are receiving increasing attention. Two contradicting epigenetic alterations often coexist in cancer: global or genome-wide hypomethylation, which is mainly observed in repetitive sequences within the genome, and regional hypermethylation, which is frequently associated with CpG islands within gene promoters. Long interspersed nuclear element-1 (LINE-1) methylation status and its relationship with the hypermethylation of CpG islands in pancreatic and biliary fluids taken from patients with pancreatobiliary cancer is not known.
This is the first study to report that pancreatobiliary cancers exhibit a pattern of genome-wide hypomethylation that can be detected using pancreatic and biliary fluids. CpG island hypermethylation of tumor-associated genes was detected at various frequencies. Hypermethylation of the ubiquitin carboxyl-terminal esterase L1 (UCHL1) gene may play a key role in the pathogenesis of pancreatobiliary cancers.
Hypermethylation of UCHL1 and runt-related transcription factor 3 in pancreatobiliary fluids might be useful for the diagnosis of pancreatobiliary cancers. A combination of highly specific epigenetic and genetic markers might provide the best diagnostic utility.
LINEs are 6-8 kb long, GC-poor sequences encoding an RNA-binding protein and a reverse transcriptase/endonuclease; these sequences constitute approximately 20% of the human genome. LINE-1 elements are most abundant, and over half a million copies of these elements are present in the human genome; UCHL1, which is also known as PARK5/PGP9.5, is a member of the ubiquitin carboxy terminal hydrolase family targeting the ubiquitin-dependent protein degradation pathway. With both ubiquitin hydrolase and dimerization-dependent ubiquitin ligase activities, UCHL1 plays important roles in multiple cellular processes. UCHL1 is a tumor-suppressor gene that is inactivated by promoter methylation or gene deletion in several types of human cancers.
The presence of such high amounts of methylated DNA in pancreatobiliary fluid is intriguing. The study has translational significance.
P- Reviewer Mishra PK S- Editor Gou SX L- Editor A E- Editor Zhang DN
| 1. | Cleary SP, Dawson LA, Knox JJ, Gallinger S. Cancer of the gallbladder and extrahepatic bile ducts. Curr Probl Surg. 2007;44:396-482. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 19] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 2. | Thomas MB. Biological characteristics of cancers in the gallbladder and biliary tract and targeted therapy. Crit Rev Oncol Hematol. 2007;61:44-51. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 33] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 3. | Hezel AF, Deshpande V, Zhu AX. Genetics of biliary tract cancers and emerging targeted therapies. J Clin Oncol. 2010;28:3531-3540. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 162] [Cited by in RCA: 172] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 4. | Kim SG, Chan AO, Wu TT, Issa JP, Hamilton SR, Rashid A. Epigenetic and genetic alterations in duodenal carcinomas are distinct from biliary and ampullary carcinomas. Gastroenterology. 2003;124:1300-1310. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 36] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 5. | Morris JP, Wang SC, Hebrok M. KRAS, Hedgehog, Wnt and the twisted developmental biology of pancreatic ductal adenocarcinoma. Nat Rev Cancer. 2010;10:683-695. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 455] [Cited by in RCA: 454] [Article Influence: 30.3] [Reference Citation Analysis (0)] |
| 6. | Ueki T, Toyota M, Sohn T, Yeo CJ, Issa JP, Hruban RH, Goggins M. Hypermethylation of multiple genes in pancreatic adenocarcinoma. Cancer Res. 2000;60:1835-1839. [PubMed] |
| 7. | Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet. 2002;3:415-428. [PubMed] |
| 8. | Sato N, Goggins M. The role of epigenetic alterations in pancreatic cancer. J Hepatobiliary Pancreat Surg. 2006;13:286-295. [PubMed] |
| 9. | Sato N, Fukushima N, Maitra A, Matsubayashi H, Yeo CJ, Cameron JL, Hruban RH, Goggins M. Discovery of novel targets for aberrant methylation in pancreatic carcinoma using high-throughput microarrays. Cancer Res. 2003;63:3735-3742. [PubMed] |
| 10. | Fukushima N, Walter KM, Uek T, Sato N, Matsubayashi H, Cameron JL, Hruban RH, Canto M, Yeo CJ, Goggins M. Diagnosing pancreatic cancer using methylation specific PCR analysis of pancreatic juice. Cancer Biol Ther. 2003;2:78-83. [PubMed] |
| 11. | Matsubayashi H, Canto M, Sato N, Klein A, Abe T, Yamashita K, Yeo CJ, Kalloo A, Hruban R, Goggins M. DNA methylation alterations in the pancreatic juice of patients with suspected pancreatic disease. Cancer Res. 2006;66:1208-1217. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 159] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 12. | Parsi MA, Li A, Li CP, Goggins M. DNA methylation alterations in endoscopic retrograde cholangiopancreatography brush samples of patients with suspected pancreaticobiliary disease. Clin Gastroenterol Hepatol. 2008;6:1270-1278. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 54] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 13. | Cordaux R, Batzer MA. The impact of retrotransposons on human genome evolution. Nat Rev Genet. 2009;10:691-703. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1100] [Cited by in RCA: 1186] [Article Influence: 74.1] [Reference Citation Analysis (0)] |
| 14. | Yang AS, Estécio MR, Doshi K, Kondo Y, Tajara EH, Issa JP. A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucleic Acids Res. 2004;32:e38. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 813] [Cited by in RCA: 794] [Article Influence: 37.8] [Reference Citation Analysis (0)] |
| 15. | Chalitchagorn K, Shuangshoti S, Hourpai N, Kongruttanachok N, Tangkijvanich P, Thong-ngam D, Voravud N, Sriuranpong V, Mutirangura A. Distinctive pattern of LINE-1 methylation level in normal tissues and the association with carcinogenesis. Oncogene. 2004;23:8841-8846. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 297] [Cited by in RCA: 314] [Article Influence: 15.7] [Reference Citation Analysis (0)] |
| 16. | Ogino S, Kawasaki T, Nosho K, Ohnishi M, Suemoto Y, Kirkner GJ, Fuchs CS. LINE-1 hypomethylation is inversely associated with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Int J Cancer. 2008;122:2767-2773. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 208] [Cited by in RCA: 213] [Article Influence: 12.5] [Reference Citation Analysis (0)] |
| 17. | Igarashi S, Suzuki H, Niinuma T, Shimizu H, Nojima M, Iwaki H, Nobuoka T, Nishida T, Miyazaki Y, Takamaru H. A novel correlation between LINE-1 hypomethylation and the malignancy of gastrointestinal stromal tumors. Clin Cancer Res. 2010;16:5114-5123. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 71] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 18. | Iacobuzio-Donahue CA, van der Heijden MS, Baumgartner MR, Troup WJ, Romm JM, Doheny K, Pugh E, Yeo CJ, Goggins MG, Hruban RH. Large-scale allelotype of pancreaticobiliary carcinoma provides quantitative estimates of genome-wide allelic loss. Cancer Res. 2004;64:871-875. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 51] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 19. | Calhoun ES, Hucl T, Gallmeier E, West KM, Arking DE, Maitra A, Iacobuzio-Donahue CA, Chakravarti A, Hruban RH, Kern SE. Identifying allelic loss and homozygous deletions in pancreatic cancer without matched normals using high-density single-nucleotide polymorphism arrays. Cancer Res. 2006;66:7920-7928. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 64] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 20. | Kipp BR, Stadheim LM, Halling SA, Pochron NL, Harmsen S, Nagorney DM, Sebo TJ, Therneau TM, Gores GJ, de Groen PC. A comparison of routine cytology and fluorescence in situ hybridization for the detection of malignant bile duct strictures. Am J Gastroenterol. 2004;99:1675-1681. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 280] [Cited by in RCA: 255] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 21. | Barr Fritcher EG, Kipp BR, Slezak JM, Moreno-Luna LE, Gores GJ, Levy MJ, Roberts LR, Halling KC, Sebo TJ. Correlating routine cytology, quantitative nuclear morphometry by digital image analysis, and genetic alterations by fluorescence in situ hybridization to assess the sensitivity of cytology for detecting pancreatobiliary tract malignancy. Am J Clin Pathol. 2007;128:272-279. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 55] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 22. | Sato N, Maitra A, Fukushima N, van Heek NT, Matsubayashi H, Iacobuzio-Donahue CA, Rosty C, Goggins M. Frequent hypomethylation of multiple genes overexpressed in pancreatic ductal adenocarcinoma. Cancer Res. 2003;63:4158-4166. [PubMed] |
| 23. | Choi IS, Estecio MR, Nagano Y, Kim do H, White JA, Yao JC, Issa JP, Rashid A. Hypomethylation of LINE-1 and Alu in well-differentiated neuroendocrine tumors (pancreatic endocrine tumors and carcinoid tumors). Mod Pathol. 2007;20:802-810. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 118] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 24. | Kim BH, Cho NY, Shin SH, Kwon HJ, Jang JJ, Kang GH. CpG island hypermethylation and repetitive DNA hypomethylation in premalignant lesion of extrahepatic cholangiocarcinoma. Virchows Arch. 2009;455:343-351. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 42] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 25. | Yamamoto E, Toyota M, Suzuki H, Kondo Y, Sanomura T, Murayama Y, Ohe-Toyota M, Maruyama R, Nojima M, Ashida M. LINE-1 hypomethylation is associated with increased CpG island methylation in Helicobacter pylori-related enlarged-fold gastritis. Cancer Epidemiol Biomarkers Prev. 2008;17:2555-2564. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 58] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 26. | Weisenberger DJ, Campan M, Long TI, Kim M, Woods C, Fiala E, Ehrlich M, Laird PW. Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res. 2005;33:6823-6836. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 543] [Cited by in RCA: 562] [Article Influence: 28.1] [Reference Citation Analysis (0)] |
| 27. | Nosho K, Yamamoto H, Takahashi T, Mikami M, Taniguchi H, Miyamoto N, Adachi Y, Arimura Y, Itoh F, Imai K. Genetic and epigenetic profiling in early colorectal tumors and prediction of invasive potential in pT1 (early invasive) colorectal cancers. Carcinogenesis. 2007;28:1364-1370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 39] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 28. | Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, Kang GH, Widschwendter M, Weener D, Buchanan D. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet. 2006;38:787-793. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1423] [Cited by in RCA: 1499] [Article Influence: 78.9] [Reference Citation Analysis (0)] |
| 29. | Ogino S, Kawasaki T, Brahmandam M, Cantor M, Kirkner GJ, Spiegelman D, Makrigiorgos GM, Weisenberger DJ, Laird PW, Loda M. Precision and performance characteristics of bisulfite conversion and real-time PCR (MethyLight) for quantitative DNA methylation analysis. J Mol Diagn. 2006;8:209-217. [PubMed] |
| 30. | Taniguchi H, Yamamoto H, Hirata T, Miyamoto N, Oki M, Nosho K, Adachi Y, Endo T, Imai K, Shinomura Y. Frequent epigenetic inactivation of Wnt inhibitory factor-1 in human gastrointestinal cancers. Oncogene. 2005;24:7946-7952. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 316] [Cited by in RCA: 361] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 31. | Oki M, Yamamoto H, Taniguchi H, Adachi Y, Imai K, Shinomura Y. Overexpression of the receptor tyrosine kinase EphA4 in human gastric cancers. World J Gastroenterol. 2008;14:5650-5656. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 51] [Cited by in RCA: 52] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 32. | Eden A, Gaudet F, Waghmare A, Jaenisch R. Chromosomal instability and tumors promoted by DNA hypomethylation. Science. 2003;300:455. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1012] [Cited by in RCA: 944] [Article Influence: 42.9] [Reference Citation Analysis (0)] |
| 33. | Tangkijvanich P, Hourpai N, Rattanatanyong P, Wisedopas N, Mahachai V, Mutirangura A. Serum LINE-1 hypomethylation as a potential prognostic marker for hepatocellular carcinoma. Clin Chim Acta. 2007;379:127-133. [PubMed] |
| 34. | Tokumaru Y, Yamashita K, Kim MS, Park HL, Osada M, Mori M, Sidransky D. The role of PGP9.5 as a tumor suppressor gene in human cancer. Int J Cancer. 2008;123:753-759. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 52] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 35. | Wada M, Yazumi S, Takaishi S, Hasegawa K, Sawada M, Tanaka H, Ida H, Sakakura C, Ito K, Ito Y. Frequent loss of RUNX3 gene expression in human bile duct and pancreatic cancer cell lines. Oncogene. 2004;23:2401-2407. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 74] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 36. | Nomoto S, Kinoshita T, Mori T, Kato K, Sugimoto H, Kanazumi N, Takeda S, Nakao A. Adverse prognosis of epigenetic inactivation in RUNX3 gene at 1p36 in human pancreatic cancer. Br J Cancer. 2008;98:1690-1695. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 28] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 37. | Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011;11:426-437. [PubMed] |
| 38. | Mittra I, Nair NK, Mishra PK. Nucleic acids in circulation: are they harmful to the host? J Biosci. 2012;37:301-312. [PubMed] |
| 39. | Logsdon CD, Simeone DM, Binkley C, Arumugam T, Greenson JK, Giordano TJ, Misek DE, Kuick R, Hanash S. Molecular profiling of pancreatic adenocarcinoma and chronic pancreatitis identifies multiple genes differentially regulated in pancreatic cancer. Cancer Res. 2003;63:2649-2657. [PubMed] |
| 40. | Ohuchida K, Mizumoto K, Ogura Y, Ishikawa N, Nagai E, Yamaguchi K, Tanaka M. Quantitative assessment of telomerase activity and human telomerase reverse transcriptase messenger RNA levels in pancreatic juice samples for the diagnosis of pancreatic cancer. Clin Cancer Res. 2005;11:2285-2292. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 45] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 41. | Bian Y, Matsubayashi H, Li CP, Abe T, Canto M, Murphy KM, Goggins M. Detecting low-abundance p16 and p53 mutations in pancreatic juice using a novel assay: heteroduplex analysis of limiting dilution PCRs. Cancer Biol Ther. 2006;5:1392-1399. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 11] [Article Influence: 0.6] [Reference Citation Analysis (0)] |