Published online Nov 21, 2012. doi: 10.3748/wjg.v18.i43.6226
Revised: August 27, 2012
Accepted: September 12, 2012
Published online: November 21, 2012
AIM: To gain insights into the molecular action of erlotinib in pancreatic cancer (PC) cells.
METHODS: Two PC cell lines, BxPC-3 and Capan-1, were treated with various concentrations of erlotinib, the specific mitogen-activated protein kinase kinase (MEK) inhibitor U0126, and protein kinase B (AKT) inhibitor XIV. DNA synthesis was measured by 5-bromo-2'-deoxyuridine (BrdU) assays. Expression and phosphorylation of the epidermal growth factor receptor (EGFR) and downstream signaling molecules were quantified by Western blot analysis. The data were processed to calibrate a mathematical model, based on ordinary differential equations, describing the EGFR-mediated signal transduction.
RESULTS: Erlotinib significantly inhibited BrdU incorporation in BxPC-3 cells at a concentration of 1 μmol/L, whereas Capan-1 cells were much more resistant. In both cell lines, MEK inhibitor U0126 and erlotinib attenuated DNA synthesis in a cumulative manner, whereas the AKT pathway-specific inhibitor did not enhance the effects of erlotinib. While basal phosphorylation of EGFR and extracellular signal-regulated kinase (ERK) did not differ much between the two cell lines, BxPC-3 cells displayed a more than five-times higher basal phospho-AKT level than Capan-1 cells. Epidermal growth factor (EGF) at 10 ng/mL induced the phosphorylation of EGFR, AKT and ERK in both cell lines with similar kinetics. In BxPC-3 cells, higher levels of phospho-AKT and phospho-ERK (normalized to the total protein levels) were observed. Independent of the cell line, erlotinib efficiently inhibited phosphorylation of EGFR, AKT and ERK. The mathematical model successfully simulated the experimental findings and provided predictions regarding phosphoprotein levels that could be verified experimentally.
CONCLUSION: Our data suggest basal AKT phosphorylation and the degree of EGF-induced activation of AKT and ERK as molecular determinants of erlotinib efficiency in PC cells.
- Citation: Lange F, Rateitschak K, Kossow C, Wolkenhauer O, Jaster R. Insights into erlotinib action in pancreatic cancer cells using a combined experimental and mathematical approach. World J Gastroenterol 2012; 18(43): 6226-6234
- URL: https://www.wjgnet.com/1007-9327/full/v18/i43/6226.htm
- DOI: https://dx.doi.org/10.3748/wjg.v18.i43.6226
Pancreatic cancer (PC) is the fourth leading cause of cancer death in the western hemisphere, with an overall five-year survival rate less than 6%[1]. The most common kind of PC is pancreatic ductal adenocarcinoma, which accounts for more than 90% of all cases. Major reasons for this poor outcome are a late diagnosis and the lack of appropriate therapy approaches. Aside from genetic alterations in oncogenes like KRAS, or tumor suppressor genes such as TP53, p16/CDNK2A, and SMAD4/DPC4[2], an increased expression of protein kinase B 2 (AKT2) and epidermal growth factor receptor (EGFR) can be found in a broad range of patient samples[3-8]. Overexpression of EGFR was accompanied by a worse overall survival[9]. Most carcinomas are diagnosed in an advanced non-resectable state, with palliative care remaining as the only treatment option.
Erlotinib, a small molecule inhibitor of the EGFR, is approved for the treatment of advanced PC. Combination treatment with gemcitabine has a moderate, but significant, survival benefit over standard treatment with gemcitabine alone[10]. Rash is a prominent side effect of EGFR-targeted therapies with monoclonal antibodies and small molecule inhibitors. In various studies, a correlation of the efficacy of a targeted therapy with erlotinib and rash was observed[10,11].
In non-small-cell lung cancer (NSCLC), activating EGFR mutations were identified as an indicator of a good response to small molecule inhibitors targeting this receptor[12]. However, EGFR activating mutations are uncommon in PC[13-15]. Unlike in NSCLC, no predictive marker (besides rash) for a response to erlotinib has been established to date in PC. It is believed that the identification of such markers holds promise for the classification of patient subgroups that would benefit most from targeted therapy in PC.
Erlotinib binds to the adenosine-5’-triphosphate (ATP) binding site of the EGFR and prevents ligand-induced receptor activation. Hence, no transphosphorylation of receptor complexes takes place, and executive downstream signaling pathways, like Ras-Raf-mitogen-activated protein kinase kinase (MEK)-extracellular signal-regulated kinase (ERK) and phosphatidylinositol 3-kinase (PI3K)-AKT, are not activated. To this day, erlotinib-attenuated signal transduction in PC is poorly understood.
Computational approaches for analyzing biochemical reactions are increasingly recognized as useful tools for the study of signaling networks and gaining deeper insights into dynamic processes[16,17]. We, along with others, have previously shown that a combination of experimental and mathematical approaches can also be successfully applied to the analysis of pathophysiological mechanisms in PC and pancreatic fibrosis[18-20].
In this study, we have addressed the question of how erlotinib modulates signal transduction via the EGFR, in order to determine molecular predictors of erlotinib sensitivity and resistance. To this end, two commonly-used and well-characterized human PC cell lines that differ in their biological sensitivity to erlotinib were chosen for a comparison of the molecular effects of the small molecule inhibitor. Experimental findings were used to establish a mathematical model that simulated major signaling pathways downstream of the EGFR. Together, our data suggest basal AKT phosphorylation and the degree of EGF-induced activation of downstream signaling pathways as molecular determinants of erlotinib efficiency.
Iscove’s modified Dulbecco’s medium (IMDM) was from Biochrom (Berlin, Germany), and RPMI 1640 and fetal calf serum (FCS) was from PAA Laboratories (Pasching, Austria). Erlotinib was supplied by Biaffin (Kassel, Germany), and AKT inhibitor XIV and U0126 by Merck (Darmstadt, Germany). Recombinant human EGF and bovine serum albumin (BSA) were delivered by Sigma-Aldrich (St Louis, MO, United States).
Phospho-EGFR (pEGFR) (Tyr1068) rabbit mAb, phospho-AKT (pAKT) (Ser473) rabbit mAb, phospho-p44/42 mitogen-activated protein kinase (phospho-extracellular signal-regulated kinase 1/2, pERK1/2) (Thr202/Tyr204) rabbit pAb, AKT rabbit mAB, and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) rabbit mAb, were purchased from New England Biolabs (Frankfurt, Germany). EGFR rabbit mAb was from Epitomics (Burlingame, CA, United States) and MAP Kinase 1/2 (ERK1/2) rabbit pAb from Millipore (Billerica, MA, United States). Fluorescently-labeled secondary antibodies for immunoblot analysis were delivered by LI-COR (Lincoln, NE, United States). Polyvinylidene fluoride (PVDF) membrane was obtained from Millipore. Standard laboratory chemicals were from Sigma-Aldrich.
The human PC cell lines BxPC-3 and Capan-1 were obtained from the American Type Culture Collection. BxPC-3 was cultured in RPMI 1640 medium supplemented with 10% FCS, 105 U/L penicillin, and 100 mg/L streptomycin. Capan-1 was cultured in IMDM medium supplemented with 17% FCS, 10 mL/L non-essential amino acids (dilution of a 100 × stock solution), 105 U/L penicillin, and 100 mg/L streptomycin. The cells were grown at 37 °C in a 5% CO2 humidified atmosphere.
To analyze the inhibitory effects of erlotinib, AKT inhibitor XIV, U0126, and combinations thereof on cell proliferation, DNA synthesis was measured using a 5-bromo-2’-deoxy-uridine (BrdU) incorporation assay (Roche Applied Science, Mannheim, Germany). Therefore, the cells were seeded in 96 half-area plates. The following day, the cells were serum-starved and the inhibitors alone or in combination were applied as indicated. After 24 h, BrdU labeling solution was added for an additional 8 h and DNA synthesis was measured following the instructions of the manufacturer.
Serum-starved BxPC-3 and Capan-1 cells were preincubated with different doses of erlotinib, AKT inhibitor XIV, or U0126, for 4 h before they were stimulated with 10 ng/mL human recombinant EGF. The cells were harvested by medium aspiration and boiled in lysis buffer [2% sodium dodecyl sulphate (SDS), 10% glycerol, 5 mmol/L ethylenediaminetetraacetic acid (pH 8.0), 62.5 mmol/L Tris-HCl (pH 6.8), 0.01% 3,3’,5,5’-tetrabromophenolsulfonphthalein, 5% β-mercaptoethanol]. The cellular proteins were separated by SDS-polyacrylamide gel electrophoresis (PAGE) and blotted onto PVDF membrane. Afterwards, the blots were blocked with 1% BSA dissolved in tris-buffered saline [TBS; 20 mmol/L Tris-HCl (pH 7.6), 140 mmol/L NaCl] for 1 h, followed by incubation with primary antibody in TBST (TBS + 0.1% Tween 20) overnight at 4 °C. After washing with TBST, the blots were incubated in secondary antibody solution for 30 min. Immunofluorescence was detected using an Odyssey Infrared Imaging system. Signals for pERK, pAKT, pEGFR, the corresponding total proteins, and GAPDH were quantified using Odyssey® Application Software 3.16.
Phosphoprotein and total protein fluorescence intensities were adjusted to GAPDH and the potential effects of gel inhomogeneities were minimized by normalizing the individually-adjusted signal intensities to the mean of all samples of the gel. At least six independent experiments were performed to calculate mean and SE.
Potential inhibitory effects of erlotinib on the basal phosphorylation of the proteins (prior to the application of EGF) were considered as follows: the time curves of EGF stimulation were adjusted with experimentally-measured basal pERK/ERK and pAKT/AKT ratios for different erlotinib concentrations in both cell lines. Results for Capan-1 cells were related to the corresponding data for BxPC-3 cells (Figure 1). However, no meaningful scaling of pEGFR for different erlotinib concentrations without EGF stimulation could be performed, as basal phosphorylation of the EGFR was too weak.
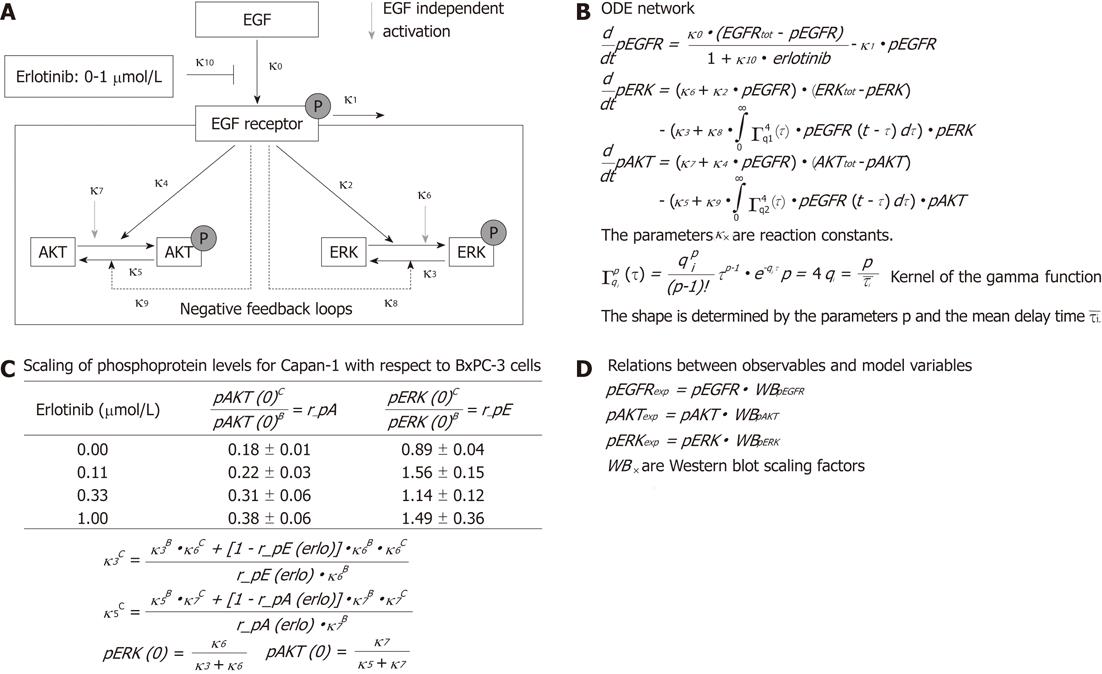
To examine the signal transduction dynamics downstream of the EGFR, we combined experimental data and mathematical modeling in a systems biological approach. A simplified network of EGFR signaling (Figure 1A) was chosen to describe the different steps in Figure 1A with the help of ordinary differential equations (ODE), whose terms are interpreted using mass action kinetics (Figure 1B). In the ODE model, EGF binds to the EGFR and triggers the phosphorylation of the receptor. Experimentally, only a low level of phosphorylated receptor was found in the absence of EGF. Therefore, no EGF-independent receptor phosphorylation was assumed. The receptor activation can be attenuated directly by erlotinib in a dose-dependent manner and by dephosphorylation. In turn, the phosphorylated receptor triggers the activation of downstream signaling pathways, where AKT and ERK were chosen as representative components. For simplicity, only two individual EGFR-induced feedback loops enhancing the dephosphorylation of AKT and ERK were assumed, although both kinases are targets of multiple inhibitory pathways[21,22].
It was shown that PC cells may secrete EGF in an autocrine loop[23]. Taking this into account, and to simulate an oncogenic KRAS-driven activation of the Ras-Raf-MEK-ERK pathway in Capan-1 cells, a phosphorylation of AKT and ERK independent of exocrine EGF was considered in the model.
We also included the experimentally measured ratios of basal phosphorylated AKT and ERK in Capan-1 versus BxPC-3 cells for all erlotinib concentrations. This information led to algebraic relations between the model parameters, and between the initial conditions and the model parameters (Figure 1C).
The relationship between the observables which are fitted to the experimental time series and the variables of the mathematical model include scaling parameters, since the levels of the phosphorylated proteins could not be quantified in an absolute manner (Figure 1D).
To optimize the parameter values, the mathematical model was trained against quantitative immunoblot data. The optimization was done with a hybrid algorithm combining a global and a local search implemented in pwFitBoost of the MATLAB Toolbox PottersWheel[24].
All experimental results represent mean ± SE for the indicated number of experiments. The Wilcoxon rank-sum test was used to test differences for statistical significance. P < 0.05 was considered statistically significant.
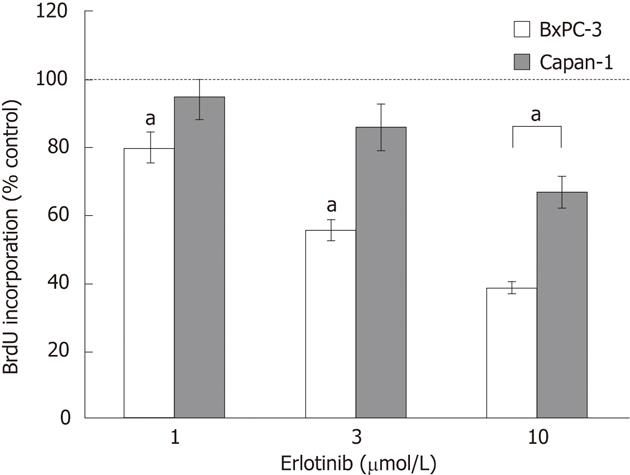
In initial experiments, two PC cell lines with different KRAS status, BxPC-3 (wild-type) and Capan-1 (harboring mutant KRAS), were tested for their sensitivity to erlotinib. Therefore, the effects of clinically achievable concentrations of erlotinib[25,26] on DNA synthesis were measured using a BrdU assay (Figure 2). Erlotinib significantly inhibited the incorporation of BrdU into newly synthesized DNA in BxPC-3 cells in a dose-dependent manner. Capan-1 cells were much more resistant to erlotinib treatment, and only the highest concentration of 10 μmol/L significantly reduced the DNA synthesis of the cells. Neither of the two cell lines carried genetic alterations in exons 19 and 21 of EGFR, the sites of hotspot mutations sensitizing the receptor to erlotinib in NSCLC[27] (data not shown).
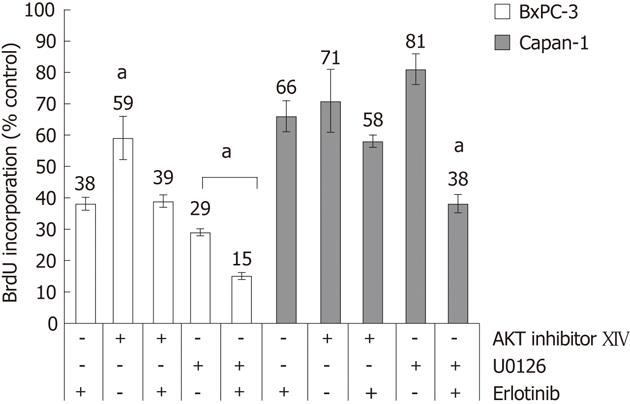
Next, the question was addressed if one of the two major pathways downstream of the EGFR, Ras-Raf-MEK-ERK and PI3K-AKT, is more sensitive against a perturbation at the EGFR level than the other. Therefore, two pathway-specific inhibitors were used. As shown in Figure 3, AKT inhibitor XIV at a concentration of 10 μmol/L diminished DNA synthesis in both cell lines. When AKT inhibitor XIV and erlotinib treatment were combined, no additional growth reduction over erlotinib alone was observed.
At a concentration of 10 μmol/L, the MEK inhibitor U0126 inhibited the DNA synthesis of both cell types. Unlike in the case of AKT inhibitor XIV, an additional treatment with erlotinib further increased the inhibition of DNA synthesis of the cells. Comparing both cell lines, BxPC-3 cells were much more sensitive to U0126 than Capan-1.
The differences in the biological response of BxPC-3 and Capan-1 cells to erlotinib and the two pathway-specific inhibitors raised the question of the underlying molecular mechanism. In our approach, we focused on the EGFR and the major downstream signaling cascades, where AKT and ERK were chosen as representative components.
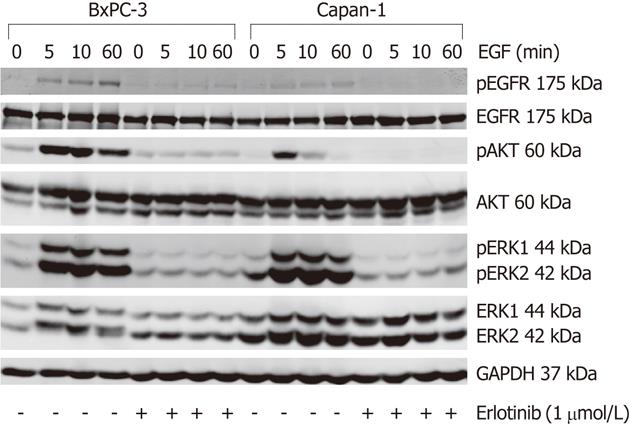
The basal (EGF-independent) phosphorylation level of the EGFR and ERK did not differ much between both cell lines; while pEGFR was barely detectable, pERK1/2 was present at a low level. In contrast, BxPC-3 cells displayed a more than five-times higher basal pAKT/AKT ratio than Capan-1 cells (Figures 4 and 5).
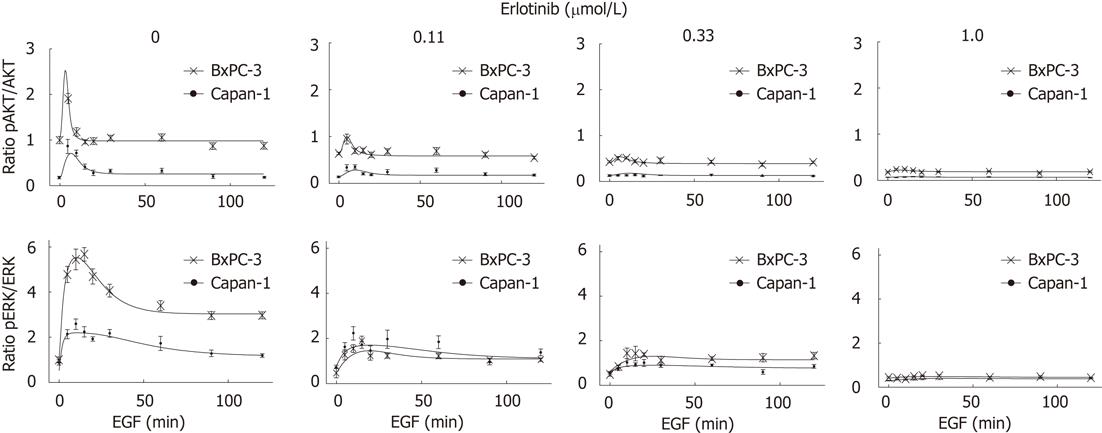
To activate the EGFR and its related pathways, cells were stimulated with 10 ng/mL human EGF. As shown in Figure 4, in response to EGF, a phosphorylation of EGFR, AKT and ERK was observed in both cell lines. The pEGFR level increased over the first 60 min of stimulation (Figure 4) and only slightly attenuated afterwards (data not shown). AKT had its maximum phosphorylation at 5 min and ERK at 10-15 min, respectively. Subsequently, pAKT decreased to the initial phosphoprotein level, while ERK phosphorylation remained above the basal level until the end of treatment (Figures 4 and 5). As shown in Figure 5, BxPC-3 cells displayed, at all time points, higher levels of pAKT and pERK than Capan-1 cells.
The phosphorylation of all three signaling components was efficiently inhibited by preincubation of the cells with 1 μmol/L erlotinib (Figures 4 and 5).
To further characterize the signal transduction dynamics downstream of the EGFR, a mathematical model was established. Towards this goal, the experimental data set presented in Figure 4 was extended with phosphoprotein data obtained by using additional erlotinib concentrations and time points of EGF stimulation. Figure 5 shows for pAKT and pERK the comparison of the experimental data and model simulations, using optimized parameter values for all erlotinib concentrations and both cell lines. Parameter values and initial conditions of fixed parameters are summarized in Table 1.
| Model parameter | Value | |
| BxPC-3 | Capan-1 | |
| Global parameter | ||
| κ0 = EGFR phosphorylation (min-1) | 0.293 | 0.438 |
| κ1 = pEGFR attenuation (min-1) | 0.067 | 0.155 |
| κ2 = EGF-dependent ERK phosphorylation (a.u.-.1min-1) | 0.388 | 0.482 |
| κ3 = basal pERK dephosphorylation (min-1) | 3.688 | a |
| κ4 = EGF-dependent AKT phosphorylation (a.u.-1min-1) | 0.225 | 0.044 |
| κ5 = basal pAKT dephosphorylation (min-1) | 0.149 | a |
| κ6 = basal ERK phosphorylation (min-1) | 0.067 | 0.104 |
| κ7 = basal AKT phosphorylation (min-1) | 0.013 | 0.007 |
| κ8 = EGF-dependent pERK dephosphorylation (a.u.-1min-1) | 4.512 | 8.520 |
| κ9 = EGF-dependent pAKT dephosphorylation (a.u.-1min-1) | 2.914 | 1.771 |
| κ10 = pEGFR inhibition by erlotinib (μmol/L-1) | 39.066 | 62.143 |
| tau1 = delay for pERK (min) | 29.766 | 66.809 |
| tau2 = delay for pAKT (min) | 3.109 | 13.803 |
| Scaling parameter | ||
| erlotinib = 0 μmol/L | ||
| scale_pEGFR | 0.858 | 0.540 |
| scale_pERK | 61.339 | 33.035 |
| scale_pAKT | 13.483 | 12.039 |
| erlotinib = 0.11 μmol/L | ||
| scale_pEGFR | 1.594 | 1.191 |
| scale_pERK | 27.224 | 29.561 |
| scale_pAKT | 8.231 | 8.254 |
| erlotinib = 0.33 μmol/L | ||
| scale_pEGFR | 2.522 | 1.955 |
| scale_pERK | 35.326 | 29.479 |
| scale_pAKT | 5.528 | 5.237 |
| erlotinib = 1.0 μmol/L | ||
| scale_pEGFR | 6.242 | 9.296 |
| scale_pERK | 15.631 | 15.464 |
| scale_pAKT | 2.569 | 2.340 |
| Fixed parameter and initial condition | ||
| pEGFR (a.u.) | 0 | 0 |
| EGFRtot = total EGFR (a.u.) | 1 | 1 |
| AKTtot = total AKT protein (a.u.) | 1 | 1 |
| ERKtot = total ERK protein (a.u.) | 1 | 1 |
As shown by experimental data and model simulation (Figure 5), erlotinib attenuated the activation of AKT and ERK in a dose-dependent manner in both cell types. Phosphoprotein levels in the two cell lines were diminished to a similar degree, except for ERK phosphorylation being more sensitive to erlotinib treatment in BxPC-3 than in Capan-1 cells.
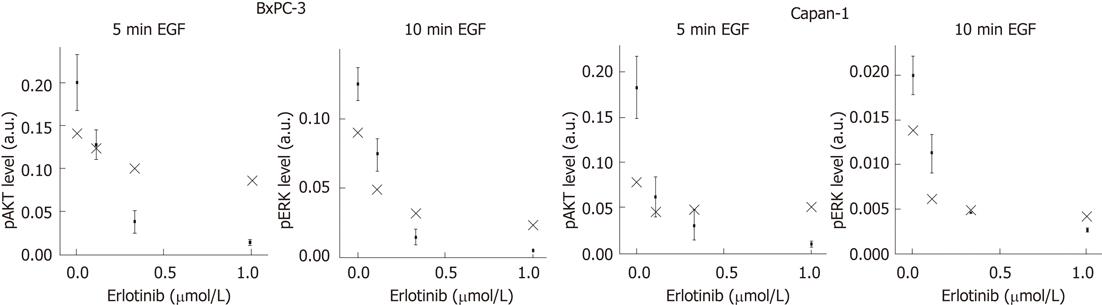
To perform a validation of our mathematical model, we experimentally verified the peaks of the phosphoprotein levels of AKT and ERK for different doses of erlotinib that were previously predicted by computational simulation. Therefore, pAKT and pERK levels were quantified for the indicated times and compared with the model calculations (Figure 6). As shown, the model was able to provide suitable predictions of phosphoprotein peaks of both signaling components in BxPC-3 and Capan-1 cells.
In this study, a combined experimental and mathematical approach was chosen to gain deeper insights into the mechanisms of EGFR signaling and erlotinib action in PC cells. In agreement with previous studies, we observed a high biological erlotinib sensitivity of BxPC-3 cells and a lesser sensitivity for Capan-1 cells[28-31].
In the two cell lines, EGF induced the phosphorylation of EGFR, AKT, and ERK with similar kinetics, but different amplitudes (higher ratios of pAKT/AKT and pERK/ERK in BxPC-3 than in Capan-1 cells). Furthermore, BxPC-3 cells displayed a more than five-times higher basal pAKT level than Capan-1 cells. Factors that may contribute to the increased AKT phosphorylation are the amplification of AKT2, and the existence of an autocrine EGF loop in BxPC-3 but not Capan-1 cells[32,33]. Despite the presence of an oncogenic KRAS allele in Capan-1 cells, basal pERK levels were similarly low in both cell lines. This seemingly surprising finding is in agreement with previous studies[34], where low levels of pERK in KRAS mutant PC cells were linked to the activity of MKP-2, a member of the dual-specificity phosphatase family that acts in a negative feedback loop[35].
In both cell lines, erlotinib efficiently inhibited phosphorylation of EGFR, ERK and AKT.
Next, we analyzed if both the PI3K-AKT and the Ras-Raf-MEK-ERK pathway were involved in mediating the anti-proliferative effects of erlotinib. We therefore challenged the cells with erlotinib or additional MEK and AKT-specific inhibitors to compare the effects on cell proliferation. In both cell types, erlotinib enhanced inhibition of cell proliferation by the pathway-specific inhibitors. U0126, but not AKT inhibitor XIV, was able to increase the growth-inhibitory effect of erlotinib. Together, these data are compatible with the hypothesis that both the AKT and ERK pathway are involved in the mediation of the antiproliferative effects of erlotinib in BxPC-3 and Capan-1 cells. The additional growth inhibitory effect of U0126 plus erlotinib versus erlotinib alone might possibly be explained by off-target effects of the MEK inhibitor, which have previously been described[36]. Our observations of BxPC-3 cells are in agreement with a previous study by Diep et al[37], who showed that in KRAS wild-type PC cells, erlotinib-attenuated cell proliferation could be further diminished with MEK inhibitors. The results of the two studies, however, differ in that we observed a similar effect of the drug combination in KRAS mutant cells, while Diep et al[37] did not. These contradictory findings are possibly due to the fact that different KRAS mutant cell lines and non-identical MEK inhibitors were used.
The antiproliferative effect of erlotinib in PC cells has previously already been linked to the expression of HER3 (ErbB3)[28,38,39]. Interestingly, HER3 has also been shown to act upstream of AKT in a pathway that is activated by EGF-induced formation of EGFR/HER3 heterodimers[39]. Thus, by blocking EGFR-mediated transphosphorylation of HER3, erlotinib may effectively interfere with AKT activation in EGF-treated PC cells. Furthermore, using a systems biology approach, Schoeberl et al[40] found that, in ovarian cancer cells, HER3 and AKT are particularly sensitive components of the HER receptor signaling network.
In conclusion, our data are in agreement with recent publications suggesting inhibition of EGF-induced ERK and AKT signaling as key components of erlotinib action in PC cells. A new finding of this study is that cells with high and low erlotinib sensitivity differed in their basal pAKT level. Furthermore, erlotinib displayed a stronger growth-inhibitory effect in the cells with a more pronounced activation of AKT and ERK in response to EGF (BxPC-3). Although we found a correlation between KRAS status and the growth-inhibitory effect of erlotinib, our data did not reveal a causal relationship, since the drug blocked ERK phosphorylation in KRAS wild-type and mutant cells with equal efficiency.
The ODE model of EGFR signaling in PC cells, established in the course of this study, accurately reflected the experimental findings. This observation suggested that the model, despite its simplifications, still contained all the components crucial to reproducing EGFR-triggered activation of AKT and ERK in silico. In support of this conclusion, the model also provided predictions regarding erlotinib-dependent changes of the phosphoprotein levels that could be verified experimentally. We consider the introduction of a mathematical model of EGFR signaling in PC cells as a first step on a path towards the identification of promising drug targets by means of computational modeling. Potential applications also include the in silico-testing of novel therapeutics targeting the EGFR pathway, and the further analysis of mechanisms of drug sensitivity and resistance.
Taken together, the results of this study may facilitate the search for molecular markers of erlotinib efficiency in PC patients. Mathematical models, like the one established here, are helping to gain further insights into signaling processes in cancer cells. In the long run, they may also become useful for predicting drug efficiencies in a clinical setting. In this regard, the particular advantages of mathematical models are that they can be based on a relatively small number of measurable parameters, and provide information about dynamic processes.
We thank Katja Bergmann for expert technical support and Andreas Frost for helping us to analyze the experimental data.
Of all common malignancies, pancreatic cancer (PC) has the lowest survival rate. The epidermal growth factor receptor (EGFR) tyrosine kinase inhibitor erlotinib is the only new drug for the treatment of locally advanced, unresectable, or metastatic PC that has been successfully introduced into the clinics in recent years, but its positive effect on survival time is small.
To date, only rash, a common side effect of EGFR targeted therapies, is an indicator of a possible response to an erlotinib treatment. Identification of molecular markers holds promise for classifying subgroups of patients who would benefit most from erlotinib therapy.
This is the first study combining experimental data and mathematical modeling to elucidate epidermal growth factor and erlotinib action in PC cells. The authors observed that PC cells with a high biological sensitivity to erlotinib displayed a higher basal phospho-protein kinase B level and increased activation of EGFR-induced downstream signaling pathways than less sensitive cells. The mathematical model not only reflected the experimental findings, but also provided predictions regarding phosphoprotein levels that could be verified experimentally.
The results of this study may facilitate the search for molecular markers of erlotinib efficiency in PC patients. Mathematical models, like the one established here, are currently applicable in order to gain molecular insights into signaling processes in cancer cells. In the long run, they may also become useful for predicting drug efficiencies in a clinical setting.
The authors present a manuscript with important data which may be helpful for patient stratification for the treatment of pancreatic carcinoma using erlotinib. Of course, the model as established in this study should be confirmed through further in vivo tests, including clinical trials, and a group of biomarkers may be developed in the future and applied for a selection of potentially sensitive patients.
Peer reviewers: Qin Su, Professor, Department of Pathology, Cancer Hospital and Cancer Institute, Chinese Academy of Medical Sciences and Peking Medical College, PO Box 2258, Beijing 100021, China; Hikaru Nagahara, MD, PhD, Professor, Aoyama Hospital, Tokyo Women’s Medical University, 2-7-13 Kita-Aoyama, Minatoku, Tokyo 107-0061, Japan; Pradyumna Mishra, Professor, Department of Translational Research, Tata Memorial Centre, Navi Mumbai 410210, India
S- Editor Lv S L- Editor Rutherford A E- Editor Lu YJ
| 1. | Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62:10-29. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8406] [Cited by in RCA: 8966] [Article Influence: 689.7] [Reference Citation Analysis (0)] |
| 2. | Hong SM, Park JY, Hruban RH, Goggins M. Molecular signatures of pancreatic cancer. Arch Pathol Lab Med. 2011;135:716-727. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 15] [Reference Citation Analysis (0)] |
| 3. | Korc M, Chandrasekar B, Yamanaka Y, Friess H, Buchier M, Beger HG. Overexpression of the epidermal growth factor receptor in human pancreatic cancer is associated with concomitant increases in the levels of epidermal growth factor and transforming growth factor alpha. J Clin Invest. 1992;90:1352-1360. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 345] [Cited by in RCA: 352] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 4. | Friess H, Wang L, Zhu Z, Gerber R, Schröder M, Fukuda A, Zimmermann A, Korc M, Büchler MW. Growth factor receptors are differentially expressed in cancers of the papilla of vater and pancreas. Ann Surg. 1999;230:767-774; discussion 774-775. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 58] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 5. | Yamamoto S, Tomita Y, Hoshida Y, Morooka T, Nagano H, Dono K, Umeshita K, Sakon M, Ishikawa O, Ohigashi H. Prognostic significance of activated Akt expression in pancreatic ductal adenocarcinoma. Clin Cancer Res. 2004;10:2846-2850. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 196] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 6. | Cheng JQ, Ruggeri B, Klein WM, Sonoda G, Altomare DA, Watson DK, Testa JR. Amplification of AKT2 in human pancreatic cells and inhibition of AKT2 expression and tumorigenicity by antisense RNA. Proc Natl Acad Sci USA. 1996;93:3636-3641. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 550] [Cited by in RCA: 579] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 7. | Ruggeri BA, Huang L, Wood M, Cheng JQ, Testa JR. Amplification and overexpression of the AKT2 oncogene in a subset of human pancreatic ductal adenocarcinomas. Mol Carcinog. 1998;21:81-86. [PubMed] |
| 8. | Altomare DA, Tanno S, De Rienzo A, Klein-Szanto AJ, Tanno S, Skele KL, Hoffman JP, Testa JR. Frequent activation of AKT2 kinase in human pancreatic carcinomas. J Cell Biochem. 2002;87:470-476. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 120] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 9. | Ueda S, Ogata S, Tsuda H, Kawarabayashi N, Kimura M, Sugiura Y, Tamai S, Matsubara O, Hatsuse K, Mochizuki H. The correlation between cytoplasmic overexpression of epidermal growth factor receptor and tumor aggressiveness: poor prognosis in patients with pancreatic ductal adenocarcinoma. Pancreas. 2004;29:e1-e8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 175] [Cited by in RCA: 191] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 10. | Moore MJ, Goldstein D, Hamm J, Figer A, Hecht JR, Gallinger S, Au HJ, Murawa P, Walde D, Wolff RA. Erlotinib plus gemcitabine compared with gemcitabine alone in patients with advanced pancreatic cancer: a phase III trial of the National Cancer Institute of Canada Clinical Trials Group. J Clin Oncol. 2007;25:1960-1966. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2835] [Cited by in RCA: 2768] [Article Influence: 153.8] [Reference Citation Analysis (0)] |
| 11. | Aranda E, Manzano JL, Rivera F, Galán M, Valladares-Ayerbes M, Pericay C, Safont MJ, Mendez MJ, Irigoyen A, Arrivi A. Phase II open-label study of erlotinib in combination with gemcitabine in unresectable and/or metastatic adenocarcinoma of the pancreas: relationship between skin rash and survival (Pantar study). Ann Oncol. 2012;23:1919-1925. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 41] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 12. | Wheeler DL, Dunn EF, Harari PM. Understanding resistance to EGFR inhibitors-impact on future treatment strategies. Nat Rev Clin Oncol. 2010;7:493-507. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 547] [Cited by in RCA: 531] [Article Influence: 35.4] [Reference Citation Analysis (0)] |
| 13. | Kwak EL, Jankowski J, Thayer SP, Lauwers GY, Brannigan BW, Harris PL, Okimoto RA, Haserlat SM, Driscoll DR, Ferry D. Epidermal growth factor receptor kinase domain mutations in esophageal and pancreatic adenocarcinomas. Clin Cancer Res. 2006;12:4283-4287. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 119] [Cited by in RCA: 127] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 14. | Tzeng CW, Frolov A, Frolova N, Jhala NC, Howard JH, Buchsbaum DJ, Vickers SM, Heslin MJ, Arnoletti JP. Epidermal growth factor receptor (EGFR) is highly conserved in pancreatic cancer. Surgery. 2007;141:464-469. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 53] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 15. | Lee J, Jang KT, Ki CS, Lim T, Park YS, Lim HY, Choi DW, Kang WK, Park K, Park JO. Impact of epidermal growth factor receptor (EGFR) kinase mutations, EGFR gene amplifications, and KRAS mutations on survival of pancreatic adenocarcinoma. Cancer. 2007;109:1561-1569. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 110] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 16. | Kholodenko B, Yaffe MB, Kolch W. Computational approaches for analyzing information flow in biological networks. Sci Signal. 2012;5:re1. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 141] [Cited by in RCA: 131] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 17. | Pe'er D, Hacohen N. Principles and strategies for developing network models in cancer. Cell. 2011;144:864-873. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 155] [Cited by in RCA: 125] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 18. | Haeno H, Gonen M, Davis MB, Herman JM, Iacobuzio-Donahue CA, Michor F. Computational modeling of pancreatic cancer reveals kinetics of metastasis suggesting optimum treatment strategies. Cell. 2012;148:362-375. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 299] [Cited by in RCA: 305] [Article Influence: 23.5] [Reference Citation Analysis (2)] |
| 19. | Lange F, Rateitschak K, Fitzner B, Pöhland R, Wolkenhauer O, Jaster R. Studies on mechanisms of interferon-gamma action in pancreatic cancer using a data-driven and model-based approach. Mol Cancer. 2011;10:13. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 23] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 20. | Rateitschak K, Karger A, Fitzner B, Lange F, Wolkenhauer O, Jaster R. Mathematical modelling of interferon-gamma signalling in pancreatic stellate cells reflects and predicts the dynamics of STAT1 pathway activity. Cell Signal. 2010;22:97-105. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 29] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 21. | Avraham R, Yarden Y. Feedback regulation of EGFR signalling: decision making by early and delayed loops. Nat Rev Mol Cell Biol. 2011;12:104-117. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 491] [Cited by in RCA: 513] [Article Influence: 36.6] [Reference Citation Analysis (0)] |
| 22. | Carracedo A, Pandolfi PP. The PTEN-PI3K pathway: of feedbacks and cross-talks. Oncogene. 2008;27:5527-5541. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 624] [Cited by in RCA: 689] [Article Influence: 40.5] [Reference Citation Analysis (0)] |
| 23. | Murphy LO, Cluck MW, Lovas S, Otvös F, Murphy RF, Schally AV, Permert J, Larsson J, Knezetic JA, Adrian TE. Pancreatic cancer cells require an EGF receptor-mediated autocrine pathway for proliferation in serum-free conditions. Br J Cancer. 2001;84:926-935. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 37] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 24. | Maiwald T, Timmer J. Dynamical modeling and multi-experiment fitting with PottersWheel. Bioinformatics. 2008;24:2037-2043. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 177] [Cited by in RCA: 147] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 25. | Hidalgo M, Siu LL, Nemunaitis J, Rizzo J, Hammond LA, Takimoto C, Eckhardt SG, Tolcher A, Britten CD, Denis L. Phase I and pharmacologic study of OSI-774, an epidermal growth factor receptor tyrosine kinase inhibitor, in patients with advanced solid malignancies. J Clin Oncol. 2001;19:3267-3279. [PubMed] |
| 26. | Tan AR, Yang X, Hewitt SM, Berman A, Lepper ER, Sparreboom A, Parr AL, Figg WD, Chow C, Steinberg SM. Evaluation of biologic end points and pharmacokinetics in patients with metastatic breast cancer after treatment with erlotinib, an epidermal growth factor receptor tyrosine kinase inhibitor. J Clin Oncol. 2004;22:3080-3090. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 132] [Cited by in RCA: 120] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 27. | Sharma SV, Bell DW, Settleman J, Haber DA. Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer. 2007;7:169-181. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2198] [Cited by in RCA: 2356] [Article Influence: 130.9] [Reference Citation Analysis (0)] |
| 28. | Buck E, Eyzaguirre A, Haley JD, Gibson NW, Cagnoni P, Iwata KK. Inactivation of Akt by the epidermal growth factor receptor inhibitor erlotinib is mediated by HER-3 in pancreatic and colorectal tumor cell lines and contributes to erlotinib sensitivity. Mol Cancer Ther. 2006;5:2051-2059. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 100] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 29. | Buck E, Eyzaguirre A, Barr S, Thompson S, Sennello R, Young D, Iwata KK, Gibson NW, Cagnoni P, Haley JD. Loss of homotypic cell adhesion by epithelial-mesenchymal transition or mutation limits sensitivity to epidermal growth factor receptor inhibition. Mol Cancer Ther. 2007;6:532-541. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 30. | Lu YY, Jing DD, Xu M, Wu K, Wang XP. Anti-tumor activity of erlotinib in the BxPC-3 pancreatic cancer cell line. World J Gastroenterol. 2008;14:5403-5411. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 10] [Cited by in RCA: 13] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 31. | El-Rayes BF, Ali S, Ali IF, Philip PA, Abbruzzese J, Sarkar FH. Potentiation of the effect of erlotinib by genistein in pancreatic cancer: the role of Akt and nuclear factor-kappaB. Cancer Res. 2006;66:10553-10559. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 93] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 32. | Shirk AJ, Kuver R. Epidermal growth factor mediates detachment from and invasion through collagen I and Matrigel in Capan-1 pancreatic cancer cells. BMC Gastroenterol. 2005;5:12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 19] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 33. | Han L, Ma Q, Li J, Liu H, Li W, Ma G, Xu Q, Zhou S, Wu E. High glucose promotes pancreatic cancer cell proliferation via the induction of EGF expression and transactivation of EGFR. PLoS One. 2011;6:e27074. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 79] [Cited by in RCA: 112] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 34. | Yip-Schneider MT, Lin A, Barnard D, Sweeney CJ, Marshall MS. Lack of elevated MAP kinase (Erk) activity in pancreatic carcinomas despite oncogenic K-ras expression. Int J Oncol. 1999;15:271-279. [PubMed] |
| 35. | Yip-Schneider MT, Lin A, Marshall MS. Pancreatic tumor cells with mutant K-ras suppress ERK activity by MEK-dependent induction of MAP kinase phosphatase-2. Biochem Biophys Res Commun. 2001;280:992-997. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 68] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 36. | Davies SP, Reddy H, Caivano M, Cohen P. Specificity and mechanism of action of some commonly used protein kinase inhibitors. Biochem J. 2000;351:95-105. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 37. | Diep CH, Munoz RM, Choudhary A, Von Hoff DD, Han H. Synergistic effect between erlotinib and MEK inhibitors in KRAS wild-type human pancreatic cancer cells. Clin Cancer Res. 2011;17:2744-2756. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 60] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 38. | Liles JS, Arnoletti JP, Tzeng CW, Howard JH, Kossenkov AV, Kulesza P, Heslin MJ, Frolov A. ErbB3 expression promotes tumorigenesis in pancreatic adenocarcinoma. Cancer Biol Ther. 2010;10:555-563. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 53] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 39. | Frolov A, Schuller K, Tzeng CW, Cannon EE, Ku BC, Howard JH, Vickers SM, Heslin MJ, Buchsbaum DJ, Arnoletti JP. ErbB3 expression and dimerization with EGFR influence pancreatic cancer cell sensitivity to erlotinib. Cancer Biol Ther. 2007;6:548-554. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 75] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 40. | Schoeberl B, Pace EA, Fitzgerald JB, Harms BD, Xu L, Nie L, Linggi B, Kalra A, Paragas V, Bukhalid R. Therapeutically targeting ErbB3: a key node in ligand-induced activation of the ErbB receptor-PI3K axis. Sci Signal. 2009;2:ra31. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 281] [Cited by in RCA: 266] [Article Influence: 16.6] [Reference Citation Analysis (0)] |