Published online Jan 7, 2011. doi: 10.3748/wjg.v17.i1.79
Revised: July 29, 2010
Accepted: August 5, 2010
Published online: January 7, 2011
AIM: To investigate the expression profile of miRNA in esophageal squamous cell carcinoma (ESCC).
METHODS: The expression profile of miRNA in ESCC tissues was analyzed by miRNA microarray. The expression levels of miR-143 and miR-145 in 86 ESCC patients were determined by real-time polymerase chain reaction (PCR) using TaqMan assay. The mobility effect was estimated by wound-healing using esophageal carcinoma cells transfected with miRNA expression plasmids.
RESULTS: A set of miRNAs was found to be deregulated in the ESCC tissues, and the expression levels of miR-143 and -145 were significantly decreased in most of the ESCC tissues examined. Both miR-143 and miR-145 expression correlated with tumor invasion depth. The transfection of human esophageal carcinoma cells with miR-143 and miR-145 expression plasmids resulted in a greater inhibition of cell mobility, however, the protein level of the previously reported target of miR-145, FSCN1, did not show any significant downregulation.
CONCLUSION: These findings suggest that the deregulation of miRNAs plays an important role in the progression of ESCC. Both miR-143 and miR-145 might act as anti-oncomirs common to ESCC.
- Citation: Wu BL, Xu LY, Du ZP, Liao LD, Zhang HF, Huang Q, Fang GQ, Li EM. MiRNA profile in esophageal squamous cell carcinoma: Downregulation of miR-143 and miR-145. World J Gastroenterol 2011; 17(1): 79-88
- URL: https://www.wjgnet.com/1007-9327/full/v17/i1/79.htm
- DOI: https://dx.doi.org/10.3748/wjg.v17.i1.79
MicroRNAs (miRNAs) are an endogenous conserved class of non-coding 20-22 nt small RNAs that regulate gene expression at post-transcriptional level by mainly binding to 3′-UTR of target mRNAs, leading to mRNA degradation or translation inhibition[1]. Many miRNAs show sequence and function conservation between distantly related organisms, suggesting that this class of small RNAs is an integral part of essential cellular processes[2]. It was predicted that about 30% of human genes are regulated by miRNAs[3]. miRNAs regulate a variety of biological processes, including developmental timing, signal transduction, cell growth, and cell death[4]. The importance of microRNA in cancer is highlighted by the observation that about 50% of miRNAs are located in cancer-associated genomic regions or fragile sites, which are frequently amplified or deleted in tumorigenesis[5]. Moreover, accumulated evidence shows that miRNAs are aberrantly expressed in various cancers, suggesting that they play a vital role as a novel class of oncogenes or tumor suppressor genes, depending on the targets they regulate[6]. Recent reports demonstrate a role for miRNA expression in disease progression and outcome[7].
Esophageal carcinoma is one of the most lethal malignancies in China and other Asian areas, with a significant low 5-year survival rate after curative surgery[8,9]. There are two major histologic types of esophageal cancer: esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma. To date, many studies focusing on miRNA expression profiles in Barrett’s esophagus and esophageal adenocarcinoma have been reported[10]. Yang et al[11] identified 11 miRNAs showing statistically significant differences between the different progression stages of esophageal adenocarcinoma. Nevertheless, there is still little information available on specific miRNA expression patterns and their roles in ESCC. Feber et al[12] identified a set of differentially expressed miRNAs that could distinguish different esophageal tissue types and also discriminate malignant from normal esophageal tissue, including adenocarcinoma, squamous cell carcinoma, Barrett’s esophagus and high-grade dysplasia. These data suggest that miRNA expression profiling is now a promising method to identify key miRNAs which play important roles in esophageal carcinogenesis.
Chaoshan Area in China is the main coastal area and has a high ESCC morbidity rate[13]. The specific geographical environment and dietary habits of the population may characterize some of the specific features of ESCC in this area, which might also be reflected in the miRNA expression profile in ESCC tissues. To develop novel diagnostic and therapeutic targets for esophageal squamous cell cancer, we first investigated the expression profile of miRNA in three pairs of clinical ESCC samples and confirmed the differences in expression of relevant miRNAs using real-time quantitative reverse transcription-polymerase chain reaction (qRT-PCR) in 86 pairs of ESCC samples. The associations between miRNA expression and clinicopathological features were analyzed. Furthermore, we tried to identify the targets of these differentially expressed miRNAs.
Human ESCC cell lines EC8712, KYSE150, EC109, EC18, SHEEC, KYSE180, KYSE70, and KYSE140 were cultured in 199 or DMEM medium (Invitrogen, Carlsbad, CA, USA) plus 10% newly born calf serum.
ESCC tissues and matched normal tissues were obtained from surgical specimens immediately after resection from patients undergoing primary surgical treatment of esophageal carcinoma from Oct 2007 to Dec 2008 in the Department of Tumor Surgery of Shantou Central Hospital, China. No patient had received preoperative irradiation or chemotherapy. The samples were flash frozen in liquid nitrogen and stored at -80°C until RNA extraction. Among these samples, three were used for microRNA microarray analysis and 86 were using for qRT-PCR analysis. Tumor specimens underwent histological examination by a pathologist to confirm the diagnosis, verify the presence of tumor, select those samples with at least 75% tumor tissue, and establish the pathological stage. Clinical and pathological information was extracted from the patients’ medical charts and pathology reports. The clinical data used for qRT-PCR analysis are shown in Table 1. Written consent for tissue donation (for research purposes) was obtained from the patients before tissue collection and the protocol was approved by the Institutional Review Board of Shantou Central Hospital.
| Clinical parameter | n |
| Gender | |
| Male | 62 |
| Female | 24 |
| Age (yr) | |
| < 55 | 35 |
| ≥ 55 | 51 |
| Diameter | |
| < 5 cm | 57 |
| ≥ 5 cm | 29 |
| LNM | |
| N0 | 53 |
| N1 | 33 |
| Invasion | |
| T1 | 4 |
| T2 | 18 |
| T3 | 64 |
| Histological type | |
| Ulcerative | 49 |
| Medullary | 24 |
| Fungating | 10 |
| Others | 3 |
| Differentiation | |
| I | 23 |
| II | 51 |
| III | 12 |
| TNM stage | |
| I | 3 |
| IIa | 49 |
| IIb | 4 |
| III | 30 |
RNA labeling and hybridization were completed by KangChen Bio-tech Inc. (Shanghai, China) according to the manufacturer’s instructions. Briefly, total RNA from three pairs of esophageal carcinoma and matched normal tissues were isolated using Trizol (Invitrogen, USA) and purified using the RNeasy mini kit (QIAGEN, Germany). The concentration and quality of total RNA were measured by NanoDrop ND-1000 at 260 and 280 nm (A260/280) and confirmed by gel electrophoresis. Each RNA sample from three pairs of ESCC was separately labeled using the miRCURY Hy3/Hy5 labeling kit and hybridized on the six miRCURYTM locked nucleic acid (LNA) array version 11.0 (Exiqon, Denmark), which contains probes for 1700 mature miRNAs. Scans were quantified using GenePix software (Molecular Devices). The data were exported to Microsoft Excel worksheets, log2 transformed, normalized using global Lowess (Locally Weighted Scatter plot Smoothing) regression algorithm (MIDAS, TIGR Microarray Data Analysis System), which we previously found to produce the best within-slide normalization to minimize the intensity-dependent differences between the dyes. Replicated spots on the same slide were averaged by obtaining a median ratio of replicated spots. Between slides normalization was performed by scale normalization to reduce between-slide variability. Only those with a greater than 2-fold increase or 2-fold decrease in expression in two samples were considered significantly changed. Samples were clustered according to their miRNA profile using Cluster 3.0 and shown using Treeview.
qRT-PCR analysis of miRNA expression was carried out using TaqMan MicroRNA Assay kits according to the manufacturer’s protocol (Applied Biosystems, USA). Briefly, total RNA was extracted using TRIzol Reagent (Invitrogen, USA) from clinical samples and ESCC cell lines. cDNAs were synthesized from total RNA using gene-specific primers. Reverse transcriptase reactions contained 10 ng RNA samples, 50 nmol/L stem-loop RT primer, 1 × RT buffer, 0.25 mmol/L each of the dNTPs, 3.33 U/μL MultiScribe reverse transcriptase and 0.25 U/μL RNase inhibitor. The 15 μL reactions were incubated for 30 min at 16°C, 30 min at 42°C, 5 min at 85°C, and then held at 4°C. The 20 μL PCR reaction included 1.33 μL RT product, 1 × TaqMan Universal PCR master mix and 1 μL primers and probe mix of the TaqMan MicroRNA Assay kit. Reactions were incubated in a 96-well optical plate at 95°C for 10 min, followed by 40 cycles at 95°C for 15 s and 60°C for 1 min. PCR reactions were run on a 7500 Real Time PCR machine (Applied Biosystems) and analyzed using 7500 System SDS software.
U6 small nuclear RNA was used as an internal control to normalize RNA input. The Ct value is defined as the fractional cycle number at which the fluorescence passes the fixed threshold. The fold change was calculated using the 2-ΔΔCt method, presented as the fold-expression change in tumors relative to their corresponding normal tissues after normalization to the endogenous control. All experiments were carried out in triplicate.
Human genomic fragments of miR-145 and miR-143 precursors with flanking about 200bp were amplified by PCR using human genomic DNA as a template. PCR primers were designed as follows: miR143 forward: 5′-AAGCTTAAGGTCAAGGTTTGGTCCT-3′; miR143 reverse: 5′-CTCGAGTGCTAAGATGGACACACTGG-3′; miR145 forward: 5′-AAGCTTCAGAGGGTTTCCGGTACTT -3′; miR-145 reverse: 5′-CTCGAGAGCCTCACAGGGATGTTATG-3′. The PCR products were cloned into the pDNA3.0 vector and named pcDNA-miR145 and pcDNA-miR143, respectively.
Approximately 2 × 105 of KYSE150 and KYSE180 cells were seeded and cultured in 6-well plates, respectively. For each well, 2.0 μg of plasmids were added to 100 μL Opti-MEM medium and 10 μL of Superfect (QIAGEN, Germany). The mixture was added to the cells and incubated for 2 h before replacing the medium. Stable clones were generated by selection in complete culture medium containing 400 mg/L of G418.
KYSE150 and KYSE180 cells were seeded in a 6-well dish, and incubated overnight yielding a confluent monolayer for wounding. Wound healing was performed using a tip with a flat point. An image was taken at different time points in each visual field at 50 ×.
TargetScan (release 5.1, http://www.targetscan.org/) was used to analyze potential target genes for the deregulated microRNAs.
Total cell lysates were prepared in RIPA buffer [50 mmol/L TrisHCl, pH 8.0, 150 mmol/L NaCl, 1% (vol/vol) Nonidet P-40, 0.5% (wt/vol) sodium desoxycholate, 0.1% (wt/vol) SDS] containing the complete protease inhibitor cocktail. Western blotting analysis was performed as described with the following primary antibodies: monoclonal mouse anti-fascin (DAKO, Denmark) and mouse anti-β-actin (Sigma, MO, USA). Experiments were repeated in triplicate.
Statistical differences between tumor and normal tissue were evaluated using the paired t-test. Statistical differences between clinicopathologic parameters and miRNA fold change were evaluated using ANOVA. The correlation coefficients of miR-143 and miR-145 were calculated using the Spearman correlation. P values less than 0.05 were considered statistically significant. All calculations were performed using Statistical Program for Social Sciences (SPSS) software 13.0 (SPSS Inc., Chicago, IL, USA).
The group specimen used for qRT-PCR was obtained from 62 males and 24 females and their details are shown in Table 1. The average age of these patients was 54 years and ranged from 40 to 75 years. Fifty-seven patients (66.3%) had tumors smaller than 5 cm, and 29 (33.7%) had tumors greater than 5 cm. Lymph node metastases were observed in 38.4% of the patients (n = 33), while the remaining patients had no lymph node metastases. Of these patients, 4 (4.7%) of 86 were diagnosed at invasion T1, 18 (20.9%) at invasion T2 and 64 (74.4%) at invasion T3. Histological type was ulcerative in 49 cases, medullary in 24 cases, fungating in 10 cases and 3 cases were unidentified. Seventy-four cases were well differentiated (I + II) and 12 cases were poorly differentiated. Taken together, the numbers diagnosed in the four clinical stages were 3, 49, 4 and 30, respectively. After two years of follow-up, the overall survival rate of the 86 ESCC patients was as high as 84.9% with only 13 deaths reported. The impact of the expression of miRNAs on patient survival will be analyzed in a future study.
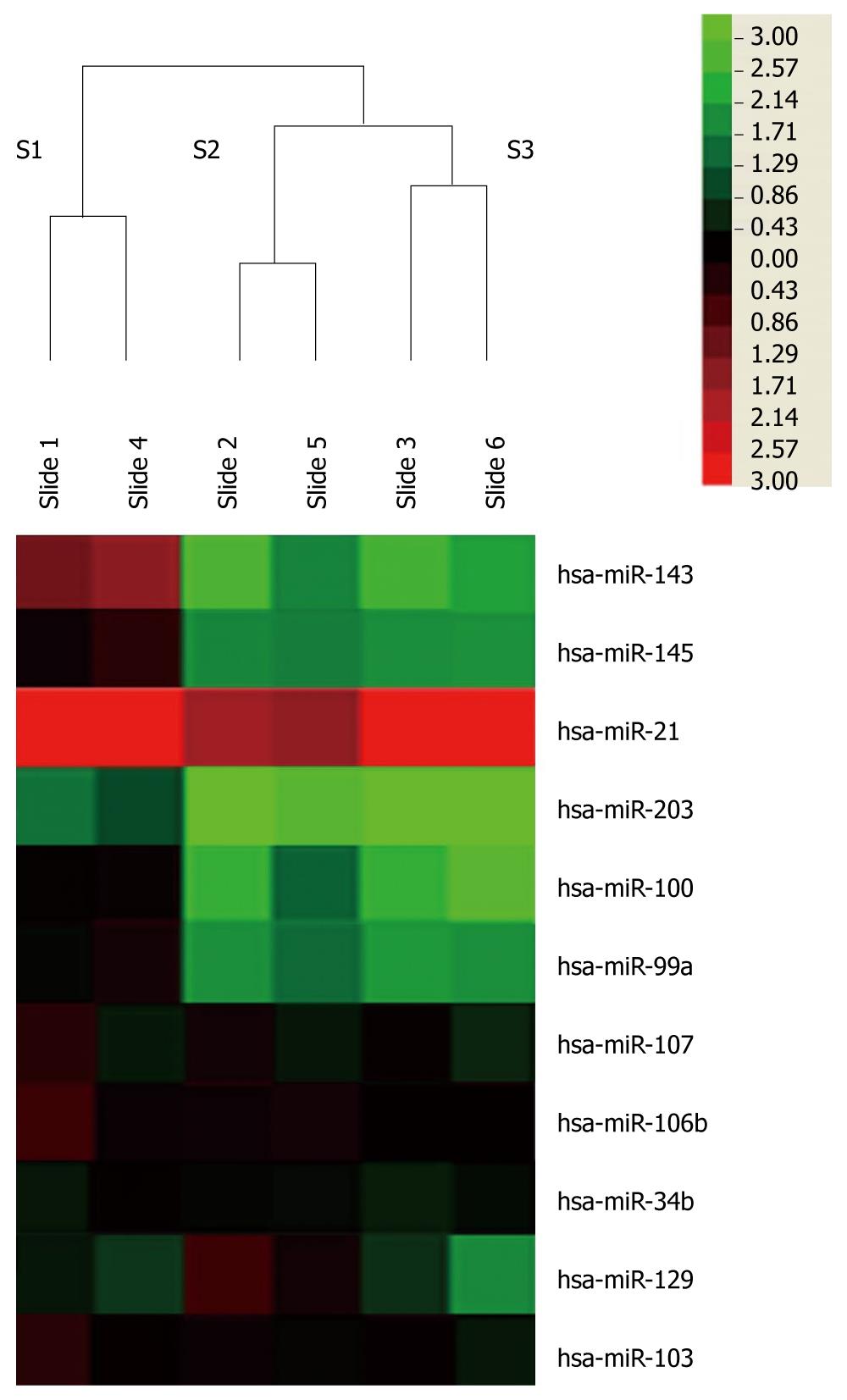
Using miRNA microarray, we then identified the miRNAs that were differentially expressed in tumor and non-tumor samples. Only miRNAs that were altered by at least 2-fold in at least two of the samples were considered significant candidates. Using these strict criteria, we identified 33 upregulated miRNAs and 40 downregulated miRNAs between normal and cancer tissues. Heat maps depict the relative expression level of mature miRNAs indicated by microarray analyses of the samples from three patients (Figure 1). This showed that the changes from two chips in each clinical case were consistent.
In accordance with previous reports[12,14], miR-21 was observed to be the most upregulated miRNA with an average 24.4-fold change. miR-203 was the most downregulated with an average 4.3-fold change. miR-203, miR-99a and miR-100 were also found to be downregulated in this study. With the exception of these, several miRNAs such as miR-143 and miR-145 were found to be changed in esophageal squamous cell carcinoma (Figure 1). On average miR-143 was downregulated 4.3-fold, while miR-145 was downregulated 3.2-fold. miR-25 was only detected in one chip and was upregulated 2.3-fold. However, a previous report of significant differential expression of miRNAs in ESCC[12,14-16], including miR-106b, -103, -107, -34b, -139 and -129, did not show significant changes in our study. All raw and normalized miRNA expression data are available from GEO publicly accessible server (http://www.ncbi.nlm.nih.gov/geo/) with the accession number: GSE23142.
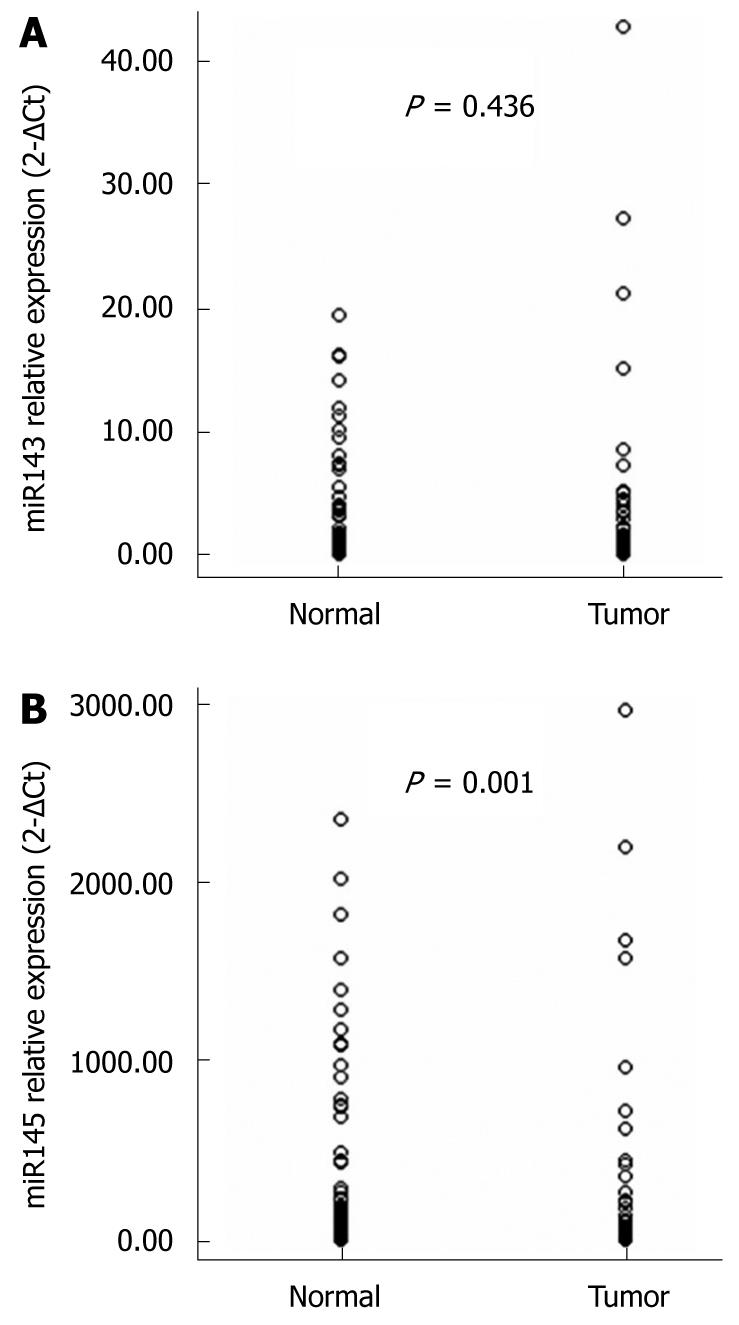
To confirm our microarray data and to determine the clinical significance of deregulated expression of miRNAs in esophageal carcinoma, we evaluated the expression of two cancer-associated miRNAs, miR-143 and miR-145, in 86 clinical samples of esophageal carcinoma and their matched normal tissues using qRT-PCR. These microRNAs were chosen as they had previously been shown to be differentially expressed in various tumors. U6 RNA expression did not differ significantly between tumor and non-tumor tissue in our study population (data not shown). As shown in Figure 2, the expression level of miR-145 was significantly downregulated in ESCC compared to matched normal tissues (P = 0.001), and no statistically significant difference in miR-143 expression between the two groups was observed (P = 0.436).
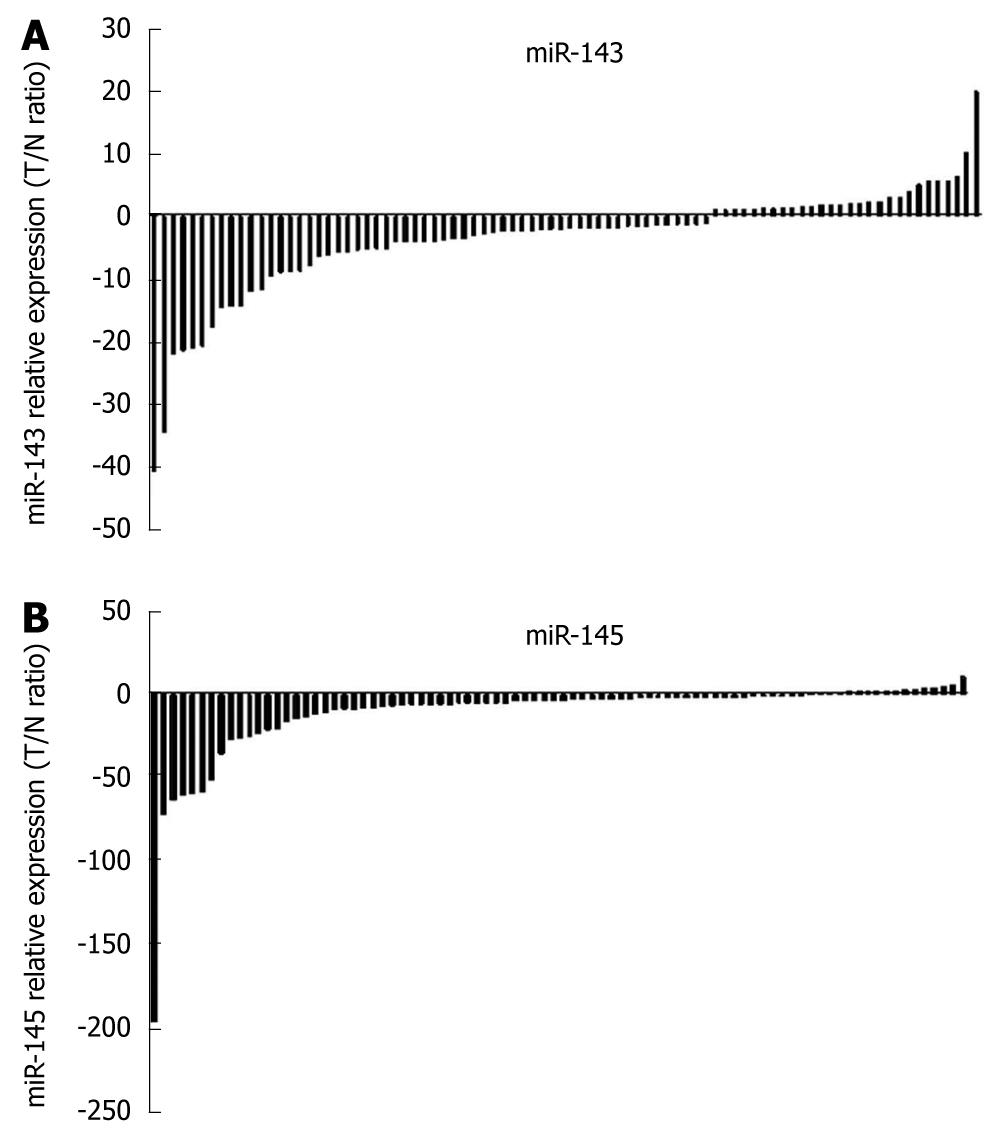
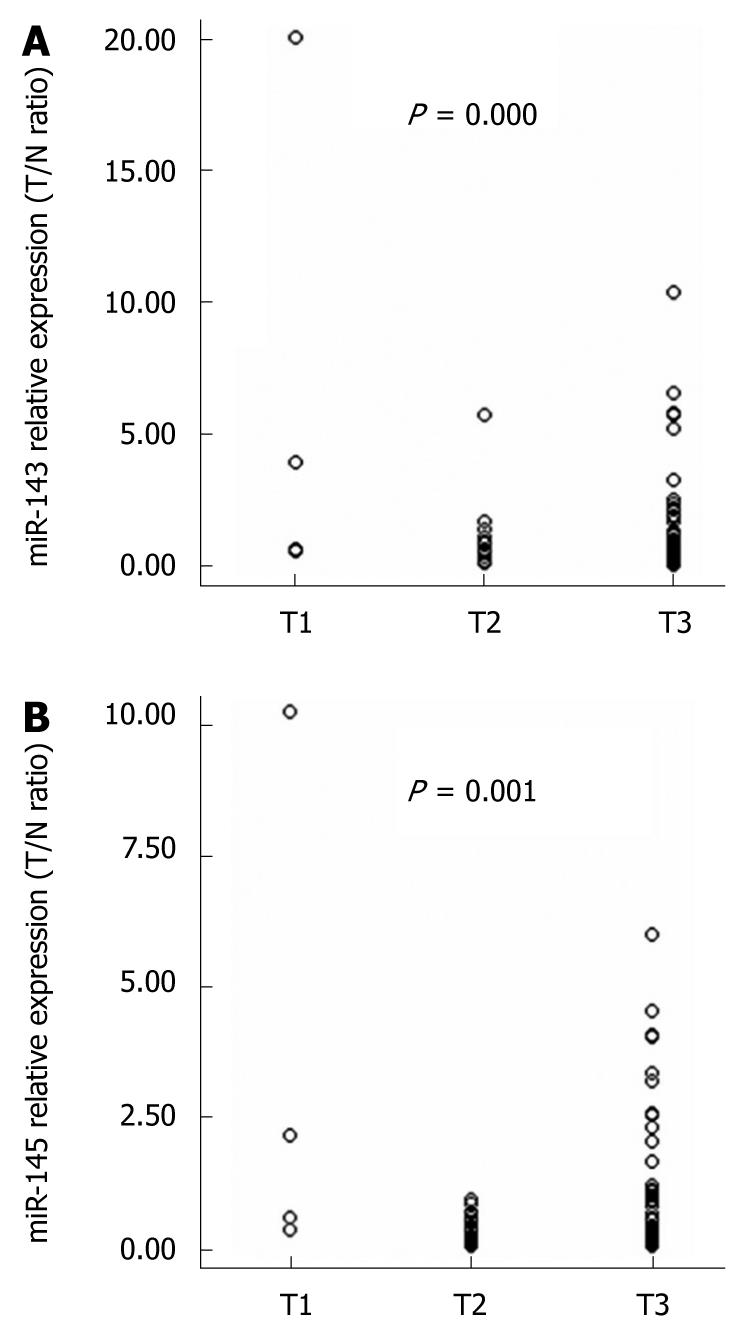
Using the 2-ΔΔCt method and a 2-fold change criterion, the qRT-PCR results showed that the expression of miR-143 was downregulated in 47.7% (41/86) and miR-145 was downregulated in 61.6% (53/86) of the clinical samples of esophageal carcinoma (Figure 3). The qRT-PCR results showed good consistency with the microRNA microarray results. In order to investigate the clinical values of miR-143 and miR-145, we analyzed the relationships between the expression levels of these two miRNAs in cancer tissues and clinicopathological factors of patients with esophageal carcinoma. The mean fold changes of miRNAs in esophageal carcinoma samples and their possible connections with cancer are presented in Table 2. Significant correlations between miR-143 and miR-145 levels in the primary tumors and tumor invasion were observed (P = 0.000 and P = 0.001, respectively, Figure 4). Furthermore, the co-expression of miR-143 and miR-145 was analyzed with the Spearman correction and showed a value of 0.967 (P = 0.000).
| miR-143 | miR-145 | |||
| median ± SD | P value | median ± SD | P value | |
| Gender | ||||
| Male | 1.43 ± 2.99 | 0.805 | 0.82 ± 1.61 | 0.718 |
| Female | 1.27 ± 1.72 | 0.95 ± 1.32 | ||
| Age (yr) | ||||
| < 55 | 1.67 ± 3.54 | 0.418 | 1.15 ± 1.99 | 0.138 |
| ≥ 55 | 1.19 ± 1.92 | 0.65 ± 1.08 | ||
| Diameter (cm) | ||||
| < 5 | 1.47 ± 2.02 | 0.701 | 0.93 ± 1.31 | 0.497 |
| ≥ 5 | 1.23 ± 3.71 | 0.69 ± 1.89 | ||
| LNM | ||||
| N0 | 1.47 ± 3.17 | 0.717 | 0.88 ± 1.71 | 0.815 |
| N1 | 1.25 ± 1.70 | 0.80 ± 1.16 | ||
| Tumor stage | ||||
| T1 | 6.31 ± 9.33 | 0.000b | 3.33 ± 4.70 | 0.001b |
| T2 | 0.80 ± 1.32 | 0.34 ± 0.30 | ||
| T3 | 1.25 ± 1.87 | 0.84 ± 1.26 | ||
| Histological type | ||||
| Ulcerative | 1.45 ± 3.28 | 0.996 | 0.79 ± 1.76 | 0.955 |
| Medullary | 1.29 ± 1.72 | 0.90 ± 1.23 | ||
| Fungating | 1.32 ± 1.70 | 0.90 ± 1.13 | ||
| Others | 1.48 ± 1.18 | 1.27 ± 0.98 | ||
| Differentiation | ||||
| I | 1.60 ± 2.11 | 0.907 | 0.87 ± 1.31 | 0.925 |
| II | 1.30 ± 3.12 | 0.81 ± 1.70 | ||
| III | 1.38 ± 1.60 | 1.00 ± 1.15 | ||
| TNM stage | ||||
| I | 0.76 ± 1.00 | 0.971 | 0.94 ± 1.39 | 0.998 |
| IIa | 1.41 ± 2.18 | 0.83 ± 1.31 | ||
| IIb | 1.06 ± 1.47 | 0.95 ± 1.48 | ||
| III | 1.46 ± 3.60 | 0.87 ± 1.90 | ||
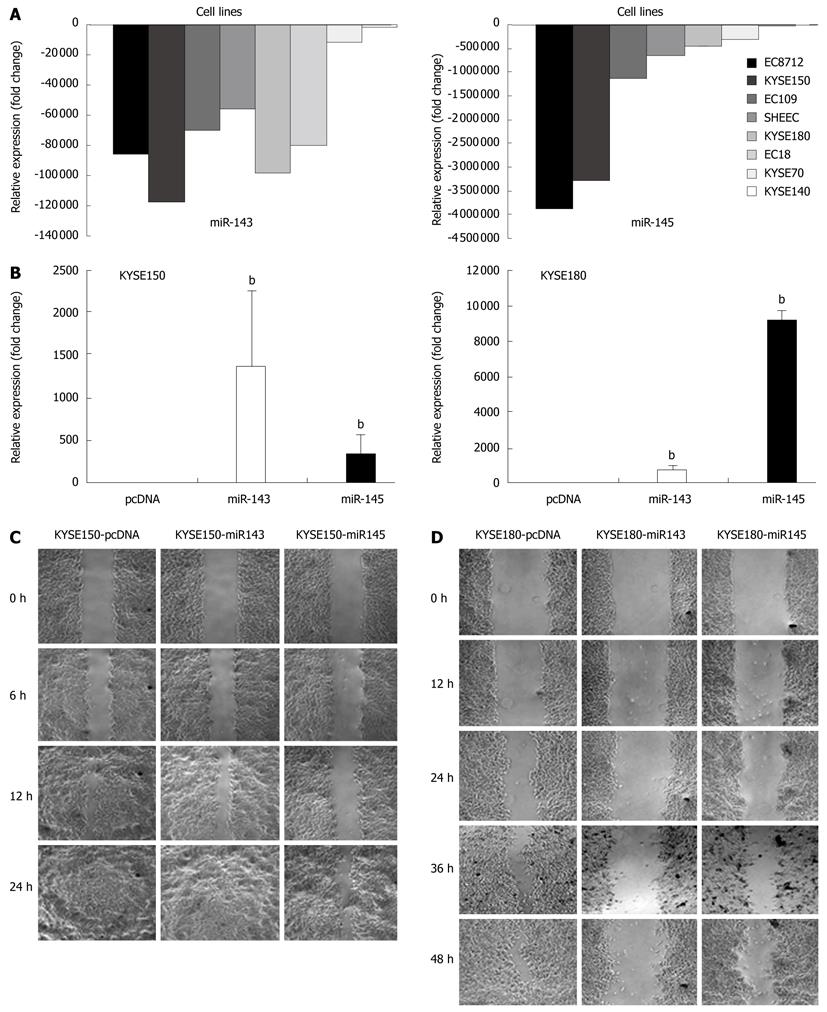
The expression levels of miR-143 and miR-145 were significantly downregulated in ESCC cell lines (Figure 5A). Following transfection with pcDNA-miR143 and pcDNA-miR145, the expression levels of these two microRNAs were significantly increased (Figure 5B). In the wound-healing assays, the distance KYSE150 and KYSE180 stably transfected cells moved in a wounded cell monolayer on plastic was determined, together with pcDNA transfected cells which were used as controls. The results showed that the cells transfected with miR-145 and miR-143 migrated very small distances and were unable to achieve wound closure (Figure 5C and D, respectively). For stably transfected KYSE150 cells, 89% of the wounds containing the control cells (16/18 visual fields) were closed or were healing, while only 58% of the miR-143 transfectant (7/12 visual fields) and 18% of the miR-145 transfectant (3/17 visual fields) were healing. For stably transfected KYSE180 cells, the percentage of healing in the control, miR-143 transfectant and miR-145 transfectant was 50% (6/12), 42% (5/12) and 0% (0/6), respectively.
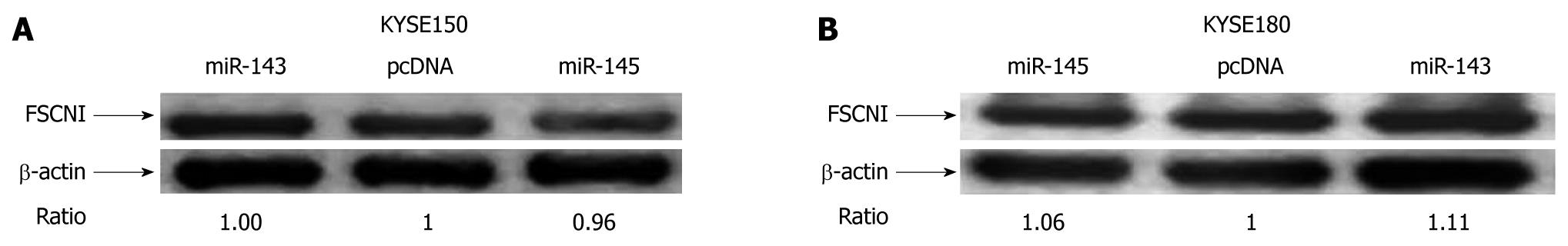
Since miR-143 and miR-145 can inhibit the mobility of esophageal carcinoma cells, we tried to identify their targets that related to cell mobility. Recently, FSCN1 has been proved to be a target of miR-145 both in the esophageal squamous cell lines TE2 and TE13 and in bladder cancer[17,18]. However, our results showed that the protein level of FSCN1 did not significantly change in the miR-143 and miR-145 stable expressing KYSE150 and KYSE180 cells, respectively (Figure 6A and B). Using TargetScan, we found that the 3′-UTR of FSCN1 contained hundreds of miRNA binding sites, especially four conserved binding sites for miR-145 and two non-conserved binding sites for miR-143 (Figure 7). Moreover, we also found other predicted targets of miR-143 and miR-145 that related to cell mobility using TargetScan (Table 3).
| Predicted targets | Functions | |
| miR-143 | LASP1 (LIM and SH3 protein 1) | Functions as an actin-binding protein and possibly in cytoskeletal organization |
| RICTOR (RPTOR independent companion of MTOR, complex 2) | Functions upstream of Rho GTPases to regulate the actin cytoskeleton | |
| ARHGAP26 (Rho GTPase activating protein 26) | Regulates the organization of the actin-cytoskeleton | |
| EPB41 [erythrocyte membrane protein band 4.1 (elliptocytosis 1, RH-linked)] | Together with spectrin and actin, constitute the red cell membrane cytoskeletal network. | |
| MYO3A (myosin IIIA) | Belongs to the myosin superfamily, an actin-dependent motor protein | |
| MARCKS (myristoylated alanine-rich protein kinase C substrate) | An actin filament crosslinking protein involved in cell motility | |
| SVIL (supervillin) | Tightly associated with both actin filaments and plasma membranes | |
| miR-145 | ARF6 (ADP-ribosylation factor 6) | Regulates vesicular trafficking, remodeling of membrane lipids, and signaling pathways that lead to actin remodeling. |
| ABLIM2 (actin binding LIM protein family, member 2) | Bound strongly to F-actin, localized to actin stress fibers | |
| ADD3 [adducin 3 (γ)] | Involved in assembly of the spectrin-actin network in erythrocytes and at sites of cell-cell contact in epithelial tissues | |
| CAPZB (c apping protein (actin filament) muscle Z-line, β) | Regulates growth of the actin filament by capping the barbed end of growing actin filaments. | |
| ELMO1 (engulfment and cell motility 1) | Mediates cytoskeletal rearrangements during phagocytosis of apoptotic cells and cell motility | |
| PHACTR2 (phosphatase and actin regulator 2) | Coprecipitated with both PP1 and actin | |
| TMOD1 (tropomodulin 1) | Inhibits depolymerization and elongation of the pointed end of actin filaments |
Accumulated studies have indicated that microRNAs, posttranscriptional modulators of gene expression, are involved in the initiation and progression of various malignancies. Many studies have demonstrated that some human miRNAs are consistently deregulated in human cancer, suggesting a role for these genes in tumorigenesis[19]. A comparison between human cancers and adjacent normal tissues have revealed distinct miRNA expression profiles. To develop novel diagnostic and therapeutic targets for ESCC, we investigated the expression profile of miRNA in ESCC.
In this study, our miRNA microarray results partially agree with Feber’s and Guo’s findings, including miR-21 and miR-203, miR-99a and miR-100, respectively, which further support the robustness of our results. With the exception of deregulated miRNAs identified in Feber’s and Guo’s reports, we also found common differentially expressed miRNAs as reported recently by Ogawa et al[16], including the upregulation of miR-21 and the downregulation of miR-133b and miR-145. Nevertheless, this is the first time that the downregulation of both miR-143 and miR-145 in esophageal squamous cell carcinoma has been reported. All these results indicate that the miRNA expression profile in ESCC from the Chaoshan Area in China was significantly altered and showed apparent regional features. These results have provided new ways of understanding the mechanism of ESCC in Chaoshan Area.
A number of reports have shown that miR-143 and miR-145 are downregulated in various cancer cells, including colorectal cancer, nasopharyngeal carcinoma, lung cancer and B-cell malignancies[20-23]. To our knowledge, we have performed the largest study to date which assesses the potential diagnostic and prognostic utility of miRNAs in esophageal squamous cell cancer. Consistent with the aforementioned previous reports, our miRNA microarray and qRT-PCR results revealed that miR-143 and miR-145 were under-expressed in most esophageal carcinoma specimens, compared with matched normal tissue samples. These results imply that these two miRNAs might serve as novel diagnostic and therapeutic targets for esophageal squamous cell cancer.
However, the pathobiological significance of aberrant miRNA expression in human esophageal squamous cell carcinoma has not been well documented. In this study, we found that both miR-143 and miR-145 correlated with tumor invasion. This suggested that both of these miRNAs might be associated with esophageal carcinoma progression. To date, little information is available on the correlations between miR-143 and miR-145 and the clinicopathologic features of other tumors. It was reported than there was no relationship between miR-143 and miR-145 expression and other clinicopathological features, except that miR-145 expression was related to cancer site following the analysis of 98 primary colorectal cancer specimens[20]. Slaby et al[24] also found that colorectal cancer tumors larger than 50 mm in maximum diameter were characterized by low expression of miR-143 and miR-145. Clapé et al[25] showed that miR-143 expression levels were inversely correlated with advanced stages of prostate cancer. Therefore, the roles of miR-143 and miR-145 in esophageal squamous cell carcinoma progression remain to be revealed.
Xin et al[26] showed that miR-143 and miR-145 selectively target genes that regulate the actin cytoskeleton in smooth muscle cells, which is intimately coupled to cell migration. Migration of esophageal squamous carcinoma cells was consistently reduced by over-expression of miR-143 and miR-145 in our wound-healing experiments. It has been reported that FSCN1 is a target of miR-145 both in bladder cancer and esophageal carcinoma[17,18]. However, no significant downregulation of FSCN1 protein was found in ESCC KYSE150 and KYSE180 cells, while the expression of miR-143 and miR-145 was rescued. This controversial result may be due to the specificity of these cell lines. Using TargetScan, we also found other predicted targets of miR-143 and miR-145 related to cell mobility, such as LASP1 and ARF6. These results implied that FSCN1 could be regulated by other miRNAs in ESCC cell lines except the TE series[16]. The targets of miR-143 and miR-145 in KYSE150 and KYSE180 cells remain to be identified.
Interestingly, the chromosome loci of both miR-143 and miR-145 are very close to each other within approximately 2 kb on 5q32, which led us to speculate that both precursors originate from the same primary transcript. In our study, we found that these two miRNAs were highly co-expressed in 86 esophageal tumors, with Spearman correlation coefficients of 0.967 (P = 0.000). This indicated that miR-143 and miR-145 might be regulated by the same factor(s) and generated from the same primary transcript (pri-miRNAs). However, various results have suggested that there may be different mechanisms for the transcriptional regulation of this locus. miR-143 and miR-145 could be transcribed individually and co-transcribed. Zhang et al[27] demonstrated that miR-143 is transcribed by nuclear factor κB, and the expression levels of miR-143 were dramatically increased in metastatic HBV-HCC and HCC patients. Xu et al[28] found that the 1.5 kb sequence upstream of miR-145 has an intrinsic promoter activity, and such activity is repressed by OCT4, which is one of the targets of miR-145. Cordes et al[29] also found that miR-143 and miR-145 were direct transcriptional targets of the serum response factors, myocardin and Nkx2-5, in multipotent murine cardiac progenitors. The transcription regulation of miR-143/-145 seems to be tissue-specific.
In summary, we report that miRNAs were deregulated and miR-143 and miR-145 were downregulated in ESCC. Furthermore, rescued expression of miR-143 and miR-145 can inhibit cell mobility. This study provides the first evidence for the anti-oncogenic activity of miR-143 and miR-145 in the development of esophageal squamous cell cancer. Targets of these two miRNAs remain to be defined. These results indicate that miRNAs may eventually constitute useful biomarkers as well as therapeutic targets.
MicroRNAs (miRNAs) regulate gene expression by mainly binding to the 3′-UTR of target mRNAs, leading to mRNA degradation or translation inhibition. miRNAs are aberrantly expressed in various cancers, suggesting that they play a vital role as a novel class of oncogenes or tumor suppressor genes, depending on the targets they regulate.
Esophageal squamous cell carcinoma is one of the most lethal malignancies in China. Many studies have reported the miRNA expression profiles in Barrett’s esophagus and esophageal adenocarcinoma. In this study, the authors report the expression profile of miRNA in esophageal squamous cell carcinoma (ESCC) and investigate the expression and functions of miR-143 and miR-145 in ESCC.
Some human miRNAs are consistently deregulated in human cancer, suggesting a role for these genes in tumorigenesis. A set of miRNAs was found to be deregulated in ESCC and the expression levels of miR-143 and miR-145 were significantly decreased in most of the ESCC tissues examined. The authors performed the largest study to date which assessed the potential diagnostic and prognostic utility of miRNAs in ESCC. It is the first study to show that both miR-143 and miR-145 were correlated with tumor invasion depth. The anti-oncogenic role of miR-143 and miR-145 was demonstrated as significant cell mobility inhibition.
This study provides the first evidence of the anti-oncogenic activity of miR-143 and miR-145 in the development of ESCC. These results indicated that miRNAs may eventually constitute useful biomarkers as well as therapeutic targets.
Many miRNAs are found deregulated in ESCC tissue and miR-143 and -145 are significantly decreased in ESCC. Both miR-143 and miR-145 correlated with tumor invasion depth. This study provides the evidence for an anti-oncogenic activity of miR-143 and miR-145 in the development of ESCC and may develop to be useful biomarkers or therapeutic targets in ESCC. It’s an excellent study.
Peer reviewer: Leonidas G Koniaris, Professor, Alan Livingstone Chair in Surgical Oncology, 3550 Sylvester Comprehensive Cancer Center (310T), 1475 NW 12th Ave., Miami, FL 33136, United States
S- Editor Wang JL L- Editor Webster JR E- Editor Ma WH
| 1. | Roush SF, Slack FJ. Micromanagement: a role for microRNAs in mRNA stability. ACS Chem Biol. 2006;1:132-134. |
| 2. | Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda MI, Maller B, Hayward DC, Ball EE, Degnan B, Müller P. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature. 2000;408:86-89. |
| 4. | Hwang HW, Mendell JT. MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer. 2006;94:776-780. |
| 5. | Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA. 2004;101:2999-3004. |
| 6. | Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10:704-714. |
| 7. | Tricoli JV, Jacobson JW. MicroRNA: Potential for Cancer Detection, Diagnosis, and Prognosis. Cancer Res. 2007;67:4553-4555. |
| 8. | Pisani P, Parkin DM, Bray F, Ferlay J. Estimates of the worldwide mortality from 25 cancers in 1990. Int J Cancer. 1999;83:18-29. |
| 9. | Enzinger PC, Mayer RJ. Esophageal cancer. N Engl J Med. 2003;349:2241-2252. |
| 10. | Kan T, Meltzer SJ. MicroRNAs in Barrett’s esophagus and esophageal adenocarcinoma. Curr Opin Pharmacol. 2009;9:727-732. |
| 11. | Yang H, Gu J, Wang KK, Zhang W, Xing J, Chen Z, Ajani JA, Wu X. MicroRNA expression signatures in Barrett’s esophagus and esophageal adenocarcinoma. Clin Cancer Res. 2009;15:5744-5752. |
| 12. | Feber A, Xi L, Luketich JD, Pennathur A, Landreneau RJ, Wu M, Swanson SJ, Godfrey TE, Litle VR. MicroRNA expression profiles of esophageal cancer. J Thorac Cardiovasc Surg. 2008;135:255-260; discussion 260. |
| 13. | Su M, Liu M, Tian DP, Li XY, Zhang GH, Yang HL, Fan X, Huang HH, Gao YX. Temporal trends of esophageal cancer during 1995-2004 in Nanao Island, an extremely high-risk area in China. Eur J Epidemiol. 2007;22:43-48. |
| 14. | Guo Y, Chen Z, Zhang L, Zhou F, Shi S, Feng X, Li B, Meng X, Ma X, Luo M. Distinctive microRNA profiles relating to patient survival in esophageal squamous cell carcinoma. Cancer Res. 2008;68:26-33. |
| 15. | Lee KH, Goan YG, Hsiao M, Lee CH, Jian SH, Lin JT, Chen YL, Lu PJ. MicroRNA-373 (miR-373) post-transcriptionally regulates large tumor suppressor, homolog 2 (LATS2) and stimulates proliferation in human esophageal cancer. Exp Cell Res. 2009;315:2529-2538. |
| 16. | Ogawa R, Ishiguro H, Kuwabara Y, Kimura M, Mitsui A, Katada T, Harata K, Tanaka T, Fujii Y. Expression profiling of micro-RNAs in human esophageal squamous cell carcinoma using RT-PCR. Med Mol Morphol. 2009;42:102-109. |
| 17. | Kano M, Seki N, Kikkawa N, Fujimura L, Hoshino I, Akutsu Y, Chiyomaru T, Enokida H, Nakagawa M, Matsubara H. miR-145, miR-133a and miR-133b: Tumor suppressive miRNAs target FSCN1 in esophageal squamous cell carcinoma. Int J Cancer. 2010;Epub ahead of print. |
| 18. | Chiyomaru T, Enokida H, Tatarano S, Kawahara K, Uchida Y, Nishiyama K, Fujimura L, Kikkawa N, Seki N, Nakagawa M. miR-145 and miR-133a function as tumour suppressors and directly regulate FSCN1 expression in bladder cancer. Br J Cancer. 2010;102:883-891. |
| 19. | Zhang B, Pan X, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302:1-12. |
| 20. | Wang CJ, Zhou ZG, Wang L, Yang L, Zhou B, Gu J, Chen HY, Sun XF. Clinicopathological significance of microRNA-31, -143 and -145 expression in colorectal cancer. Dis Markers. 2009;26:27-34. |
| 21. | Chen HC, Chen GH, Chen YH, Liao WL, Liu CY, Chang KP, Chang YS, Chen SJ. MicroRNA deregulation and pathway alterations in nasopharyngeal carcinoma. Br J Cancer. 2009;100:1002-1011. |
| 22. | Liu X, Sempere LF, Galimberti F, Freemantle SJ, Black C, Dragnev KH, Ma Y, Fiering S, Memoli V, Li H. Uncovering growth-suppressive MicroRNAs in lung cancer. Clin Cancer Res. 2009;15:1177-1183. |
| 23. | Akao Y, Nakagawa Y, Kitade Y, Kinoshita T, Naoe T. Downregulation of microRNAs-143 and -145 in B-cell malignancies. Cancer Sci. 2007;98:1914-1920. |
| 24. | Slaby O, Svoboda M, Fabian P, Smerdova T, Knoflickova D, Bednarikova M, Nenutil R, Vyzula R. Altered expression of miR-21, miR-31, miR-143 and miR-145 is related to clinicopathologic features of colorectal cancer. Oncology. 2007;72:397-402. |
| 25. | Clapé C, Fritz V, Henriquet C, Apparailly F, Fernandez PL, Iborra F, Avancès C, Villalba M, Culine S, Fajas L. miR-143 interferes with ERK5 signaling, and abrogates prostate cancer progression in mice. PLoS One. 2009;4:e7542. |
| 26. | Xin M, Small EM, Sutherland LB, Qi X, McAnally J, Plato CF, Richardson JA, Bassel-Duby R, Olson EN. MicroRNAs miR-143 and miR-145 modulate cytoskeletal dynamics and responsiveness of smooth muscle cells to injury. Genes Dev. 2009;23:2166-2178. |
| 27. | Zhang X, Liu S, Hu T, Liu S, He Y, Sun S. Up-regulated microRNA-143 transcribed by nuclear factor kappa B enhances hepatocarcinoma metastasis by repressing fibronectin expression. Hepatology. 2009;50:490-499. |