INTRODUCTION
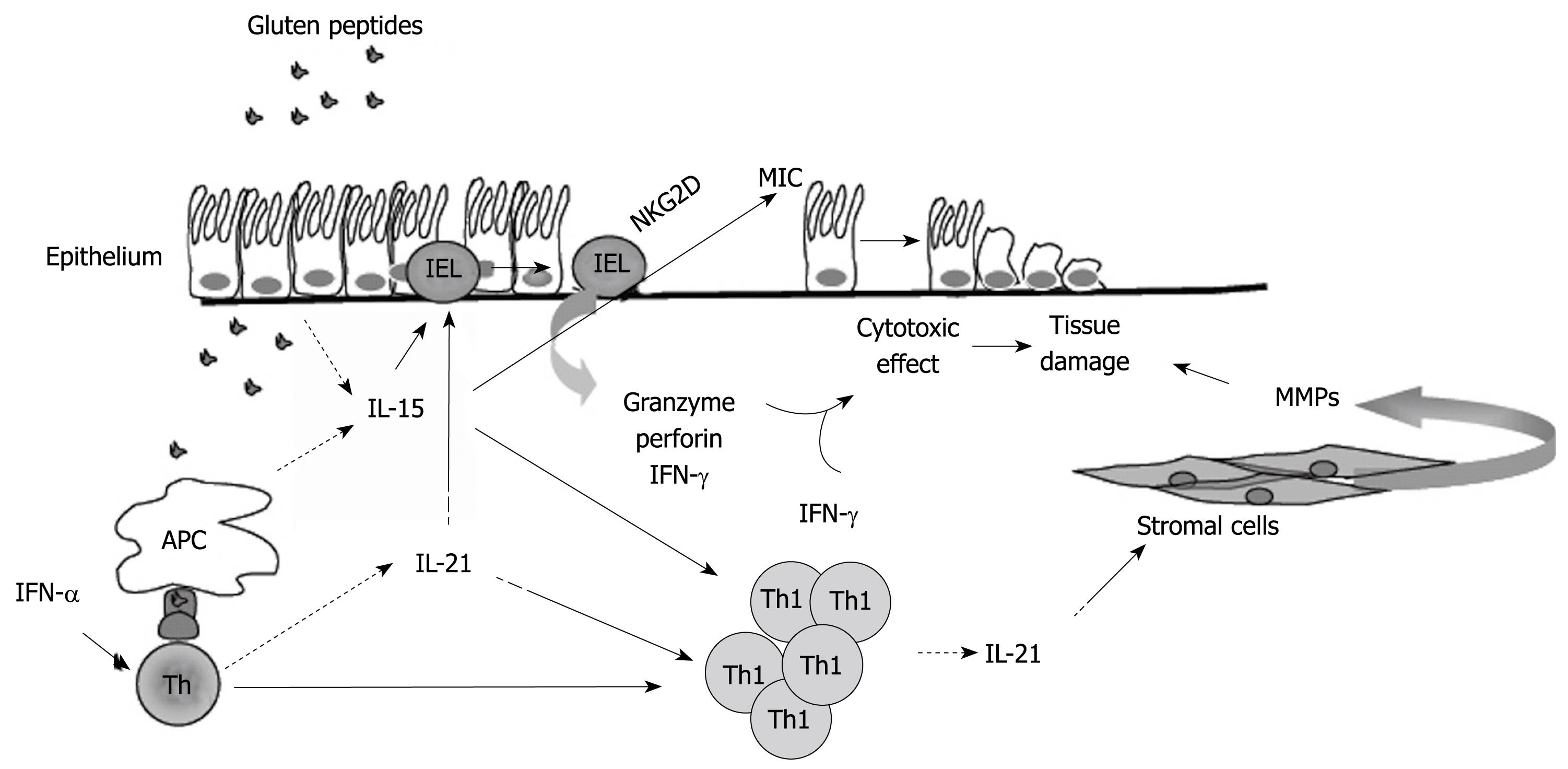
Figure 1 Hypothetical view of the pathogenic effects of IL-15 and IL-21 in celiac disease.
In active celiac disease mucosa, IL-15 is produced by epithelial cells and antigen presenting cells (APC) in response to gluten stimulation. APC also produce interferon (IFN)-α, which stimulates CD4+ T cells to polarize along the Th1 pathway. IL-21 is produced by activated CD4+ T cells, including Th1 cells. IL-15 activates various pathways, which lead to the lysis and death of epithelial cells. IL-15 augments the production of granzyme B by IELs and IFN-γ by IELs and Th1 cells; enhances the expression of the activating NGK2D receptor on IELs; induces the NKG2D ligand, MICA, on epithelial cells. IL-21 seems to contribute to the tissue-damaging immune response in CD, given that this cytokine is able to enhance the production of perforin by IELs and IFN-γ by T cells, and to stimulate stromal cells to make extracellular matrix-degrading metalloproteinases.
Celiac disease (CD) is a chronic gastrointestinal disorder caused, in genetically predisposed individuals, by ingestion of the gluten proteins of wheat, rye, and barley. The prevalence of CD is approximately 1% in the Western world[1-3]. The disease is strongly associated with HLA-DQ genes. Most patients carry a variant of DQ2 (alleles DQA1*05/DQB1*02) and others carry a variant of DQ8 (alleles DQA1*03/DQB1*0302)[4,5]. These haplotypes are relatively common in the healthy population, and only 1:20 individuals who express HLA-DQ2/DQ8 have CD[4,5]. These observations and the fact that the concordance rate between HLA-identical siblings is much lower than between monozygotic twins suggest that most probably other non-HLA genes are involved in the pathogenesis of CD[6-8].
In CD patients, ingestion of gluten triggers a mucosal inflammatory response, which leads to the tissue damage. The histological features of CD are villous atrophy, crypt cell hyperplasia, and increased number of intra-epithelial lymphocytes (IELs). The clinical manifestations can range from asymptomatic to severe malabsorption. Patients can also manifest symptoms of other immune-mediated diseases, which can associate with CD. The only treatment is a lifelong gluten-free diet, which results, in the vast majority of patients, in complete remission of symptoms and recovery of the normal mucosal histology[9].
Although the pathogenesis of CD is not fully understood, it is known that gluten peptides are deamidated by tissue transglutaminase and presented by DQ2+ or DQ8+ antigen-presenting cells to lamina propria CD4+ T cells[9,10]. Upon activation, CD4+ T cells polarize along the T helper (Th)1-type pathway, as substantiated by their ability to produce large amounts of interferon (IFN)-γ, the signature cytokine of Th1 responses[11,12]. In CD patients on a gluten-free diet, IFN-γ production is as low as in healthy controls but it can be stimulated in vitro by gluten to reach levels of untreated CD patients. In these mucosal cultures, neutralization of IFN-γ prevents gliadin-mediated morphological changes thus supporting the role of the adaptive immune response and IFN-γ in CD immunopathology[11,12]. More recently it has become clear that some gluten peptides can induce mucosal damage by directly activating innate immune mechanisms[13]. These observations collectively underline the complexity of the pathogenic mechanism in CD and suggest that the CD-associated mucosal damage relies on the activation of multiple rather than single cell pathways.
GLUTEN PEPTIDES STIMULATE INNATE IMMUNITY AND CAUSE EPITHELIAL DAMAGE VIA AN INTERLEUKIN (IL)-15-DEPENDENT MECHANISM
It has long been known that gluten peptides cause epithelial damage when added to ex vivo organ cultures of biopsies taken from CD patients on a gluten-free diet, but not from controls[14]. As previously mentioned, this pathogenic response was initially thought to be secondary to the activation of lamina propria CD4+ T cells and production of inflammatory cytokines, such as IFN-γ[11]. However, more recently Maiuri et al[13] showed that the gluten p31-43 peptide is able to induce mucosal damage by directly activating innate immune cells. In particular, it was shown that the p31-43 peptide elicits the production of IL-15 by lamina propria macrophages and dendritic cells in ex vivo organ cultures of CD biopsies, thus triggering a sequence of events that culminates in epithelial damage (Figure 1). These findings correlate well with the demonstration that IL-15 is over-expressed in both the lamina propria and intestinal epithelium of patients with active, untreated CD as compared with normal controls or inactive, gluten-free diet treated CD patients[15,16].
The majority of intestinal IELs are T cell receptor (TcR)αβ+ CD8+CD4- and a significant proportion are TcRγδ+ CD8-CD4-[14]. IELs express a variety of natural killer (NK) lineage receptors, supporting their involvement in epithelial cell damage[14]. Corroborating Maiuri’s results, Hüe et al[17] showed that treatment of biopsies taken from inactive CD patients with the gluten p31-49 peptide enhances the enterocyte expression of the non-conventional HLA molecules MICA/B. MIC molecules are known to be induced on enterocytes by stress and are up-regulated in active CD mucosa[18]. MICA/B are ligands of the activating NKG2D receptor, which signals through the adaptor protein DAP10 and is expressed on most NK cells, CD8+ TcRαβ+ and TcRγδ+, but normally not on CD4+ T cells[19,20]. Expression of NKG2D in IELs is increased in active CD mucosa, and IELs lyse epithelial cells via NKG2D[17]. Up-regulation of MICA/B molecules also occurs in CD biopsies treated with exogenous IL-15, and a neutralizing IL-15 antibody blocks the gluten-mediated MIC-inducing effect in organ cultures of CD biopsies[17,21]. Meresse et al[22] showed that IL-15 also enhances the expression of both NKG2D and DAP10 in IELs, and that the cytolytic attack of the epithelium by IELs may be perpetuated via NKG2D independently of TcR specificity (Figure 1). Taken together these findings emphasize the critical role of IL-15 in intestinal epithelial cell death induced by NKG2D-expressing IELs.
IL-15 AND COUNTER-REGULATORY MECHANISMS IN CD
In the gut, mucosal homeostasis arises from a highly dynamic balance between host protective immunity and regulatory mechanisms[23]. One such counter-regulatory mechanism involves transforming growth factor-β1 (TGF-β1), a cytokine that is able to exert a number of negative effects on immune cells, including inhibition of T cell proliferation and differentiation, as well as down-regulation of macrophage activation and dendritic cell maturation[24]. Consistently, mice with global TGF-β1 defects, such as TGF-β1-deficient mice or transgenic mice expressing a dominant-negative TGF-βRII chain that are unresponsive to TGF-β1 signaling, develop intestinal mucosal inflammation[25,26]. On the other hand, studies in mouse models of gut inflammation have shown that production of TGF-β1 is consistently associated with greatly diminished severity of inflammation[27]. Because TGF-β1 regulates both lymphoid and myeloid cells[24], several researchers have examined the expression and activity of TGF-β1 in chronic gastrointestinal inflammatory diseases, including CD. Initial studies conducted by Lahat et al[28] showed that in active CD mucosa there was enhanced expression of transcripts for TGF-β1 as compared to controls. In contrast, Lionetti et al[29] reported that the total RNA expression of TGF-β1 did not differ between active CD and controls. However, in active CD, TGF-β1 production was mostly confined to lamina propria T cells and macrophages, while in controls it was mostly produced by epithelial cells[29].
TGF-β1 initiates signaling through the ligand-dependent activation of a complex of heterodimeric transmembrane serine/threonine kinases, consisting of type I (TGF-βRI) and type II (TGF-βRII) receptors[30]. Upon TGF-β1 binding there is phosphorylation and activation of TGF-βRI by the constitutively active and auto-phosphorylating TGF-βRII. TGF-βRI in turn phosphorylates two proteins, termed Smad 2 and Smad3. Once phosphorylated, Smad2 and Smad3 associate with Smad4 and translocate to the nucleus where Smad protein complexes participate in transcriptional control of target genes[30]. This pathway has been reported to be antagonized by inflammatory cytokines, which can impair the regulatory functions of TGF-β1 and alter immune homeostasis[31,32]. Therefore, it is conceivable that, during chronic inflammatory processes, there may be deficient TGF-β1 activity despite this cytokine being highly produced. In line with this, Benahmed et al[33] have recently shown that TGF-β1 inhibited the proliferative response of normal intestinal IELs and lamina propria lymphocytes induced by IL-2 but not that triggered by IL-15. The inhibitory effect of IL-15 on TGF-β1 activity was substantiated further by the demonstration that IL-15 impaired the formation of Smad3-DNA complexes in response to TGF-β1 stimulation[33]. In contrast, IL-15 enhanced the activation of the Janus kinase (JNK) pathway thus promoting the inhibitory effect of phospho-c-jun on the formation of Smad-DNA complexes. Further analysis revealed that the transcription levels of tristetraprolin, a TGF-β target gene under the control of the Smad3 pathway, were decreased in duodenal biopsies from patients with active CD compared with controls. This defect was associated with no significant change in the intracellular levels of phosphorylated Smad3 and Smad7, clearly indicating that initial steps of TGF-β signaling are preserved in patients with active CD[33]. This is in contrast to the Smad7-dependent inhibition of TGF-β1/Smad3 signaling described in patients with inflammatory bowel diseases[34,35].
IELs, lamina propria lymphocytes, and epithelial cells of active CD contain high levels of phospho-c-jun, and inhibition of this protein in organ cultures of CD biopsies enhances tristetraprolin[33]. Moreover, treatment of ex vivo organ cultures of CD biopsies with anti-IL-15 reduces phospho-c-jun and increases tristetraprolin, thus confirming the prominent role of IL-15 in the phospho-c-jun-mediated inhibition of TGF-β1 signaling[33].
While NKG2C and NKG2D are activating NK receptors, NKG2A is considered an inhibiting NK receptor. NKG2A can associate with CD94 and bind the non-classical MHC Ib molecule, HLA-E, which is expressed on epithelial cells of CD patients but not on epithelial cells of healthy individuals[36,37]. Although NKG2A competes with NKG2C for its interaction with HLA-E, the former has a higher binding affinity for HLA-E compared with NKG2C. Upon binding to this ligand, CD94/NKG2A delivers negative signals to cytotoxic cells[38]. A higher percentage of small intestinal TcRγδ IELs express CD94/NKG2A compared with TcRαβ IELs, and TcRγδ IELs exert negative effects on the IL-15-mediated induction of IFN-γ, granzyme B, and NKG2D in CD8+ TcRαβ IELs[39]. These observations suggest that TcRγδ IELs have the ability to suppress the cytotoxic programming of TcRαβ IELs. This inhibitory effect requires interaction of NKG2A with its ligand HLA-E, a phenomenon which is followed by enhanced secretion of TGF-β1. Notably, blockade of TGF-β1 activity with a neutralizing human TGF-β1 antibody partially abrogates the suppressive capability of TcRγδ IELs, and exogenous TGF-β1 dose-dependently inhibits the IL-15-mediated induction of IFN-γ, granzyme B, and NKG2D in CD8+ TcRαβ IELs[39]. Therefore, in this cell context, IL-15 is not sufficient to abrogate the immunosuppressive action of TGF-β1. These findings conflict with the aforementioned data published by Benahmed et al[33]. It is likely that this discrepancy may simply be due to differences in cell culture conditions used in these two studies. Another possibility is that the IL-15-mediated negative regulation of TGF-β1 signaling occurs only in specific cell types. Since TGF-β1 can trigger both Smad-dependent and -independent intracellular pathways[39,40], it is also plausible that IL-15 can interfere with some and not all TGF-β1-activated signals.
Expression of NKG2A is decreased on TcRαβ IELs and TcRγδ IELs from CD patients as compared to normal subjects[39-41]. The mechanism that removes this negative regulator from CD IELs remains unknown. NKG2A possesses in its promoter a binding site for Smad3, and there is evidence that TGF-β1 cooperates with the TcR in enhancing the expression of NKG2A on CD8+ T cells[18,42]. Therefore, molecules that disrupt TGF-β1 signaling could, at least in theory, contribute to the down-regulation of NKG2A in CD. IL-15 is not able to directly regulate NKG2A expression[43], even though it remains possible that it could inhibit the TGF-β1-induced NKG2A expression.
IL-21 as a positive regulator of Th1 cell response in CD
The demonstration that CD lesions are associated with a marked infiltration of IFN-γ-secreting cells has boosted intensive research aimed at identifying the factors that promote the ongoing mucosal Th1 cell response. Paradoxically, IL-12, the major Th1-inducing factor in man, is not over-produced in CD mucosa[12]. However, analysis of transcription factors that drive Th cell differentiation revealed that in active CD mucosa there is enhanced expression of T-bet, a member of the T-box family of transcription factors that directs Th1 lineage commitment and is essential for IFN-γ production in CD4+ T cells[43]. In contrast, no increase in GATA3 and active STAT6, two transcription factors that specifically regulate Th2 differentiation, is seen in active CD samples as compared to controls[43]. Interestingly, active CD biopsies do not exhibit enhanced activation of STAT4[43], an IL-12-dependent Th1-inducing transcription factor, thus confirming the lack of IL-12 activity. These data strongly support the existence of a regulatory pathway for Th1 differentiation that starts from activation of T-bet and is independent of IL-12-driven STAT-4 signaling. So the critical question is: what induces and sustains T-bet in CD? We have recently shown that in the duodenal mucosa of patients with active but not inactive CD there is enhanced production of IL-21[44]. Interestingly, neutralization of IL-21 activity in ex vivo organ cultures of CD biopsies reduces both T-bet expression and IFN-γ production[44]. Since IFN-γ enhances T-bet via a STAT1-dependent mechanism[43], it is tempting to speculate that IL-21 is part of a positive feedback loop that helps amplify and stabilize the committed Th1 cell phenotype in CD (Figure 1).
Recent genome-wide association studies have provided convincing evidence that the chromosomal 4q27 region harboring the IL-2 and IL-21 genes is associated with CD[7]. A similar genetic association has been described in other immune-mediated diseases, such as psoriasis, type 1 diabetes, and inflammatory bowel diseases[45,46]. However, it is not yet known if such polymorphisms can influence the tissue levels of IL-21.
Biopsies taken from CD patients on a gluten-free diet over-express IL-21 when challenged in vitro with gluten peptides[44]; however, the basic mechanisms that control the expression of IL-21 in CD mucosa remain to be elucidated. Previously, we showed that IFN-α is up-regulated in the mucosa of active CD patients, where it most probably contributes to intensifying IFN-γ production[47]. Indeed, neutralization of IFN-α activity drastically reduces the gliadin peptide-driven IFN-γ expression in biopsies of inactive CD patients[48]. Interestingly, IFN-α enhances the mRNA expression of IL-21 in activated human T cells[49], thus suggesting a role for this cytokine in the positive control of IL-21 in CD (Figure 1).
Potential involvement of IL-21 in the activity of other mucosal cells types
The biological functions of IL-21 are mediated by a cell-surface class I cytokine receptor, formed by the common γ-chain subunit (shared with IL-2, IL-4, IL-7, IL-9, IL-13 and IL-15 receptors) and its own unique receptor (designated IL-21 receptor)[50]. Since this receptor is expressed by both immune and non-immune cells, it is plausible that IL-21 maintains chronic inflammation and/or favors tissue damage in CD by targeting additional cell types other than Th1 lymphocytes. In fact, IL-21 has been shown to stimulate epithelial cells to secrete chemokines and facilitate recruitment of immune cells within the inflamed tissue, induce fibroblasts to make tissue-damaging proteases, and make effector CD4+ T cells resistant to regulatory T cell-mediated immunosuppression[51-53]. In light of the role of IL-21 in the control of B cell and plasma cell function[54], IL-21 may also contribute to the production of CD-associated autoantibodies. Since its discovery, it has become clear that IL-21 is also able to modulate the proliferation and/or effector function of CD8+ T cells and NK cells[55]. Using IELs isolated from human jejunal mucosa, Ebert has recently shown that IL-21 increases perforin-mediated cytotoxicity and serine esterase release without affecting the growth and survival of these cells[56]. The relevance of this finding for CD pathogenesis remains to be ascertained, although IL-21-mediated cytotoxic T cell activation could contribute to the intestinal epithelial cell death and villous atrophy in CD.
CONCLUSION
A growing body of evidence suggests that IL-15 and IL-21 may play important roles in the immune response associated with inflammation and tissue damage in CD, even though some intriguing questions remain to be resolved. For instance, it remains to be determined whether these two cytokines contribute qualitatively and quantitatively differently to the initiation and progress of the inflammatory cascade in CD. The fact that IL-15 and IL-21 share a receptor subunit and target the same cell types raises the possibility that these cytokines may cooperate in regulating specific cell immune responses. Indeed, both IL-15 and IL-21 are known to stimulate the growth of T cells, enhance the production of IFN-γ by Th1 cells, and promote the activation of cytotoxic cells[21,44,50,57] (Figure 1). As outlined in this article, these two cytokines might also act in concert to disrupt local mechanisms of immune tolerance. This hypothesis is supported by the demonstration that IL-15 inhibits TGF-β1 activity, and IL-21 renders effector CD4+ T cells resistant to the suppressive effects of CD4+CD25+ regulatory T cells[33,53]. Nonetheless it is noteworthy that IL-15 and IL-21 seem to differentially modulate some aspects of both innate and adaptive immunity, such as dendritic cell maturation and T cell apoptosis[50,57]. Therefore, we cannot exclude the possibility that, at least in some stages of the inflammatory process, IL-15 and IL-21 can exert opposing effects on the gluten-driven immune response.