Published online Jul 7, 2009. doi: 10.3748/wjg.15.3161
Revised: May 27, 2009
Accepted: June 3, 2009
Published online: July 7, 2009
AIM: To characterize the types of mutations present in the 23S rRNA genes of Malaysian isolates of clarithromycin-resistant Helicobacter pylori (H pylori).
METHODS: Clarithromycin susceptibility of H pylori isolates was determined by E test. Analyses for point mutations in the domain V of 23S rRNA genes in clarithromycin-resistant and -sensitive strains were performed by sequence analysis of amplified polymerase chain reaction products. Restriction fragment length polymorphism was performed using BsaI and MboII enzymes to detect restriction sites that correspond to the mutations in the clarithromycin-resistant strains.
RESULTS: Of 187 isolates from 120 patients, four were resistant to clarithromycin, while 183 were sensitive. The MIC of the resistant strains ranged from 1.5 to 24 &mgr;g/mL. Two isolates had an A2142G mutation and another two had A2143G mutations. A T2182C mutation was detected in two out of four clarithromycin-resistant isolates and in 13 of 14 clarithromycin-sensitive isolates. Restriction enzyme analyses with BsaI and MboII were able to detect the mutations.
CONCLUSION: Clarithromycin resistance is an uncommon occurrence among Malaysian isolates of H pylori strains and the mutations A2142G and A2143G detected were associated with low-level resistance.
-
Citation: Ahmad N, Zakaria WR, Abdullah SA, Mohamed R. Characterization of clarithromycin resistance in Malaysian isolates of
Helicobacter pylori . World J Gastroenterol 2009; 15(25): 3161-3165 - URL: https://www.wjgnet.com/1007-9327/full/v15/i25/3161.htm
- DOI: https://dx.doi.org/10.3748/wjg.15.3161
Helicobacter pylori (H pylori) is a microaerophilic, Gram-negative bacterium, and is implicated often as the causative agent of chronic gastritis, duodenal and gastric ulcers, and gastric carcinoma[12]. H pylori-associated disorders usually regress or heal completely after treatment with antibiotics[3]. Clarithromycin is a macrolide that has been used frequently in combination with other antimicrobial agents for the treatment of H pylori infection[4–6]. However, resistance to clarithromycin has become one of the major reasons for treatment failure[7]. The prevalence of H pylori resistance to clarithromycin varies among different countries, such as 12% in Japan, 1.7%-23.4% in Europe and 10.6%-25% in North America[89].
Clarithromycin acts by binding to the peptidyl transferase region of 23S rRNA and inhibits bacterial protein synthesis. Resistance to clarithromycin results from structural changes in the 23S rRNA molecule caused by mutation of the 23S rRNA gene[1011]. Adenine to guanine transitions at positions 2142 and 2143 are the main 23S rRNA mutations in clarithromycin-resistant isolates[12–14]. All these mutations have been shown to confer resistance to this macrolide by mutagenesis study[15]. Other mutations that have been observed in clarithromycin-resistant H pylori isolates are A2515G and T2717C, A2116G, G2141A, A2144T, T2182C, G2224A, C2245T, and T2289C[16]. The A2142C/G and A2143G mutation also generate specific restriction sites, namely MboII and BsaI, and these may be used for the rapid screening of clarithromycin resistance.
In the present study, we determined the prevalence of clarithromycin resistance in our local H pylori strains and characterized the types of mutations that occurred in the resistant strains. In addition, we determined whether the Restriction fragment length polymorphism (RFLP) technique was suitable for rapid detection of the mutations among our local isolates.
The patients enrolled in this study were the patients who underwent endoscopy for gastrointestinal symptoms at the Gastroenterology Unit, National University of Malaysia Hospital between 2005 and 2007. Written informed consent was obtained from all patients before biopsies were taken from the antrum and corpus of each patient.
Biopsy samples were cultured on Columbia agar supplemented with 10% ox blood. Plates were incubated at 37°C under microaerophilic condition for 5-7 d. Bacterial isolates were identified according to colony morphology, Gram-staining, urease, catalase and oxidase. The cultures were stored at -80°C in Brucella broth supplemented with 15% glycerol and fetal calf serum (Invitrogen, USA).
The MIC of clarithromycin was determined by the E-test method. E tests (AB Biodisk, Sweden) were performed on Columbia agar supplemented with 10% ox blood. The plates were incubated under microaerophilic condition for 3-5 d. Isolates were classified as clarithromycin-resistant if MIC was > 1 &mgr;g/mL. H pylori strain ATCC43504 was included as a control clarithromycin-sensitive strain.
DNA extraction was carried out using the Nucleospin® Tissue Kit (Macherey-Nagel, BD Biosciences, USA) and stored at -20°C until use. The primers used for PCR amplification were Hp5 forward, 5'-GTCGTGCCAAGAAAAGCGTCT-3' (positions 1672-1693; GenBank accession number U27270), and Hp2 reverse, 5'-TGTGTGCTACCCAGCGATGCTC-3' (positions 2811-2790; GenBank accession number U27270). PCR was carried out on 10 mmol/L dNTP (Promega, USA), 10 pmol of each primer, 1.25 U Taq DNA polymerase (Promega), and genomic DNA of H pylori. The PCR was performed in a GeneAmp PCR System 2400 thermal cycler (Perkin Elmer, USA) for 30 cycles with the following cycling conditions: 94°C for 1 min, 58°C for 30 s and 72°C for 30 s. The PCR products were then analyzed by electrophoresis using 1% agarose and stained with ethidium bromide.
PCR products were purified using a QIAquick PCR purification kit (Qiagen). Sequencing was performed using BigDye® Terminator v3.1 sequencing kit and analyzed on ABI PRISM® 377 Genetic Analyzer. The sequences were analyzed using free ClustalW software (http://www.ebi.ac.uk/clustalw).
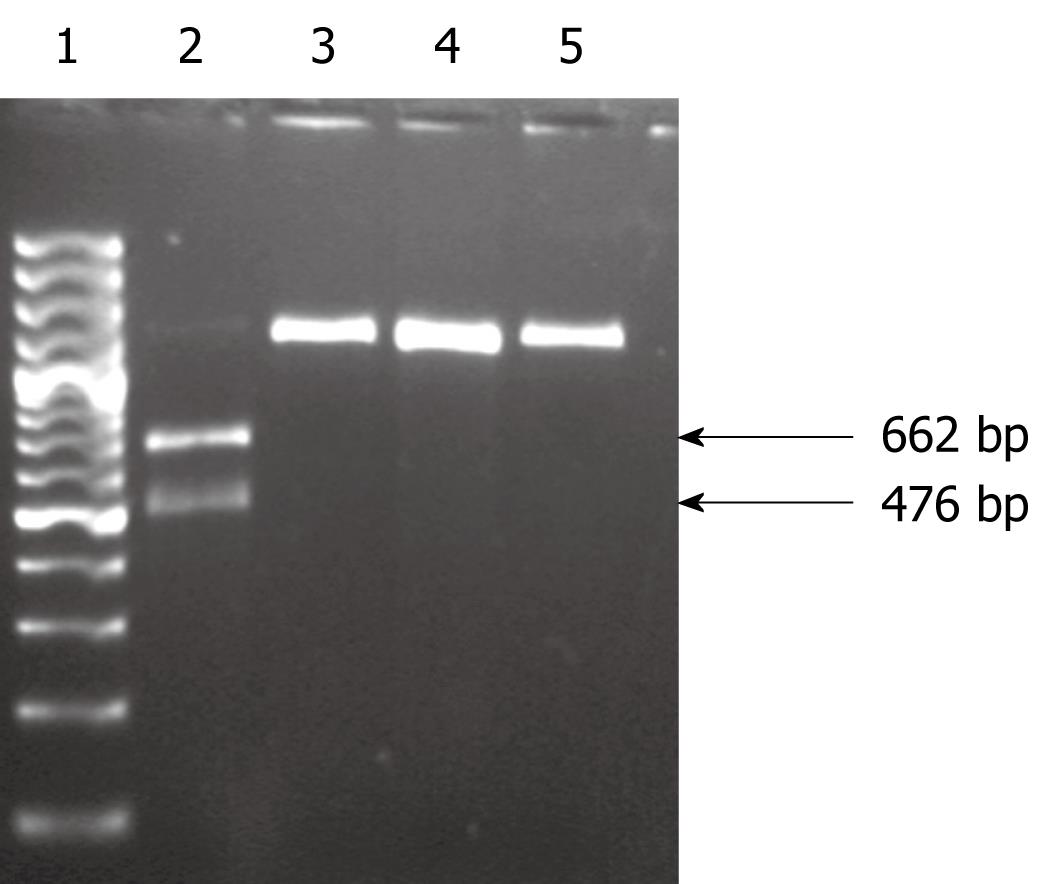
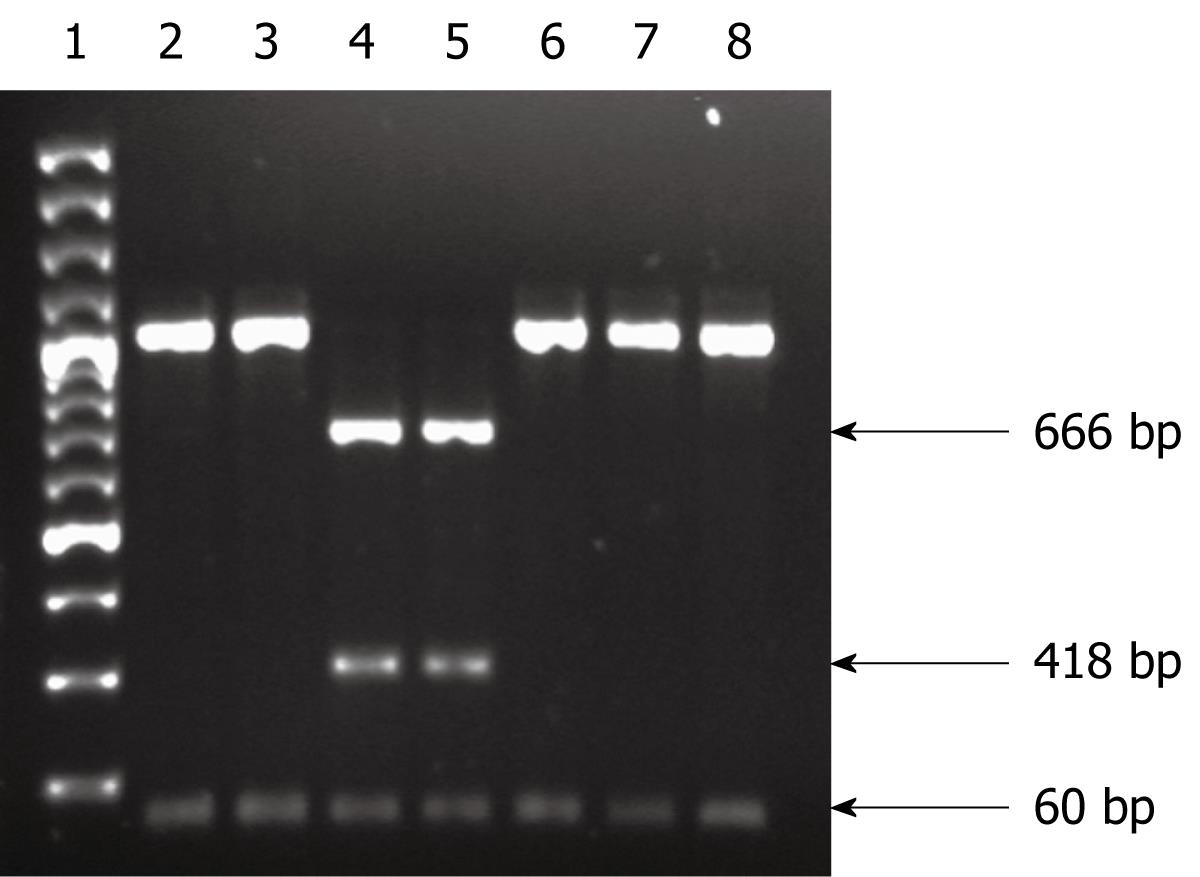
To detect the A-G point mutations at positions 2142 and 2143, the amplified PCR products were digested with the MboII and BsaI (MBI Fermentas, Lithuania) restriction enzymes, respectively, and analyzed on 1.6% agarose gels. MboII-digested PCR products that contained the A2142G mutation were expected to yield two fragments (476 and 662 bp) when digested with MboII, whereas the PCR products that contained the mutation A2143G were expected to yield three fragments (60, 418 and 666 bp) when digested with BsaI.
A total of 187 H pylori were isolated from gastric biopsies of 120 patients. Four of these isolates (2.1%) from four patients were resistant to clarithromycin, while 183 isolates from 116 patients were sensitive to clarithromycin. The MIC values of all the clarithromycin-resistant isolates ranged from 1.5 to 24 &mgr;g/mL (Table 1). The patients with clarithromycin-resistant isolates had never been treated for H pylori infection.
| Clarithromycin MIC (&mgr;g/mL) and phenotype | Mutation site | Number of isolate(s) |
| 24 (R) | A2142G | 1 |
| 24 (R) | A2143G, T2182C | 1 |
| 3 (R) | A2143G | 1 |
| 1.5 (R) | A2142G, T2182C | 1 |
| < 0.016 (S) | T2182C | 13 |
| < 0.016 (S) | No mutation | 1 |
All the clarithromycin-resistant strains and 14 randomly selected clarithromycin-sensitive strains from 14 different patients were subjected to DNA extraction and PCR amplification of domain V of the 23S rRNA gene. The mutations in domain V of 23S rRNA were determined by comparing the sequences of clarithromycin-resistant and -sensitive isolates with the sequences of the reference strain ATCC43504. All of the clarithromycin-resistant isolates were shown to have point mutations of either A2142G or A2143G, while none of the 14 clarithromycin-sensitive isolates had these types of mutations. T2182C mutation was detected in both clarithromycin-resistant and -sensitive isolates.
Sequence analysis of restriction sites of the amplified V domain of the 23S rRNA gene of the resistant strains showed restriction sites that could be digested with BsaI and MboII. Digestion of the amplified PCR product with MboII produced two fragments of 662 and 476 bp sizes in strains with the A2142G mutation. No fragment was produced with the A2143G or T2182C mutation (Figure 1). Resistant isolates with the A2143G mutation produced three fragments of 662, 476 and 60 bp when digested with BsaI. No restriction fragment was produced in strains with A2142G and T2182C mutations (Figure 2). None of the clarithromycin-sensitive isolates and the negative control (ATCC43504) produced restriction fragments when digested with MboII and BsaI, which indicated the absence of A2142G and A2143G mutations.
The prevalence of clarithromycin resistance was found to be low among the isolates in the present study. To the best of our knowledge, this was the first study that reported and characterized clarithromycin-resistant strains in Malaysia.
Clarithromycin is used worldwide for H pylori eradication therapy. H pylori strains that are resistant to clarithromycin have been reported increasingly in several studies[917]. As resistance will often lead to failure of eradication therapy, knowledge of the current susceptibility patterns of local H pylori isolates could help in determining the choice of appropriate treatment for the patient. In the present study, the prevalence of clarithromycin resistance was still low; however, physicians should bear in mind the possibility of clarithromycin resistance if their patient does not respond to this antibiotic.
Versalovic et al[18] have shown that A2142G and 2143G mutations of 23S rRNA genes in H pylori are associated with resistance to clarithromycin. A mutagenesis study performed by Taylor et al[15] has confirmed that the A2142G and A2143G mutations are associated with clarithromycin resistance of H pylori. A review by Mégraud[8] of several studies worldwide has shown that 81.5% of the mutations in chlarithromycin-resistant isolates were the A2142G or A2143C mutations. In our study, either A2142G or A2143G mutations were found in the clarithromycin-resistant isolates, or they were absent in the clarithromycin-sensitive isolates. The mutations found in our local strains were in accordance with the findings in other countries. The mutations were also associated with low-level resistance.
Another type of mutation detected in our isolates was T2182C. This mutation was detected in most of the strains studied, which suggests a common occurrence among our local strains. This mutation is not new and has been described in previous reports[1920]. This mutation is not associated with clarithromycin resistance[21]. However, it has not been discussed whether this type of mutation occurs commonly in H pylori. Mutations other than A2142G and A2143G have been reported in other studies[1622], but among our strains, sequence analysis showed that the mutations were limited to these types only. Although the low number of resistant isolates in our study limit the scope of detection, the results suggest that clarithromycin resistance of H pylori in Malaysia can be predicted by detection of mutations at positions 2142 and 2143 of the 23S rRNA gene. The resistance to clarithromycin among the four patients is considered primary resistance because they were never treated previously for H pylori eradication.
Knowledge about the molecular mechanism of resistance is important as it can be used to facilitate the development of other molecular methods to detect resistance. It has been shown that PCR-RFLP can be used to detect mutations in clarithromycin-resistant isolates[2324]. Using this technique in our study, it was demonstrated that only strains with the A2142G and A2143G mutations produced restriction fragments when digested with MboII or BsaI. This alleviated the need to use the more expensive and time-consuming processes of sequencing and analysis to look for mutation sites.
In conclusion, we showed that there was a low prevalence of clarithromycin resistance in our local strains of H pylori. The A2142G or A2143G mutations detected were in accordance with the findings in other countries. These mutations were not found among the analyzed clarithromycin-sensitive strains. T2182C mutation was a common occurrence in our local isolates. Other types of mutations associated with clarithromycin resistance were not observed in our resistant strains. The A2142G or A2143G mutations were detected also by RFLP, thus making it a rapid method for the detection of clarithromycin-resistant strains in our local isolates.
The incidence of clarithromycin-resistant Helicobacter is increasing worldwide. This limits the therapeutic options available for eradicating the bacterium. Clarithromycin resistance has been attributed to mutations in the 23S rRNA gene.
Clarithromycin acts by binding to the peptidyl transferase region of 23S rRNA and inhibits bacterial protein synthesis. There are many types of mutations observed in the 23S rRNA genes of clarithromycin-resistant Helicobacter pylori (H pylori). There is little information on the prevalence and characteristics of clarithromycin resistance in H pylori strains isolated from Malaysian patients. In the present study, the authors determined the prevalence of resistance and characterized the types of mutations present in their resistant strains.
The study gave an insight into the low prevalence of clarithromycin resistance among the H pylori strains studied. The A2142G and A2143G mutation detected were in accordance with the findings in our other countries. However, other types of mutations associated with clarithromycin resistance were not observed among our resistant strains.
The A2142G and 2143G mutations in the 23S rRNA genes in H pylori can be detected easily by restriction fragment length polymorphism (RFLP) analysis of the polymerase chain reaction (PCR) product of the genes, using MboII or BsaI. This eliminates the need to detect these mutations by sequencing.
This is a straightforward study that determined the prevalence of clarithromycin resistance in H pylori strains isolated from the authors¡¯ institute in Malaysia. The types of mutations that occurred in the resistant strains were further characterized by PCR and sequencing. Finally, the authors evaluated whether the RFLP technique was suitable for rapid detection of the mutations among these isolates. Although the sample size of clarithromycin-resistant isolates was relatively small, the techniques and methodology used were appropriate and standard. The information is potentially useful for the clinical management of H pylori infection in Malaysia, and also for our overall appreciation of the global distribution and prevalence of antibiotic resistance of this important pathogen.
| 1. | Huang JQ, Hunt RH. Review article: Helicobacter pylori and gastric cancer--the clinicians'point of view. Aliment Pharmacol Ther. 2000;14 Suppl 3:48-54. |
| 2. | Sipponen P, Kosunen TU, Valle J, Riihelä M, Seppälä K. Helicobacter pylori infection and chronic gastritis in gastric cancer. J Clin Pathol. 1992;45:319-323. |
| 3. | Sugiyama T, Sakaki N, Kozawa H, Sato R, Fujioka T, Satoh K, Sugano K, Sekine H, Takagi A, Ajioka Y. Sensitivity of biopsy site in evaluating regression of gastric atrophy after Helicobacter pylori eradication treatment. Aliment Pharmacol Ther. 2002;16 Suppl 2:187-190. |
| 4. | Goddard AF, Logan RP. Antimicrobial resistance and Helicobacter pylori. J Antimicrob Chemother. 1996;37:639-643. |
| 5. | Mégraud F, Lamouliatte H. Review article: the treatment of refractory Helicobacter pylori infection. Aliment Pharmacol Ther. 2003;17:1333-1343. |
| 6. | Cavallaro LG, Egan B, O'Morain C, Di Mario F. Treatment of Helicobacter pylori infection. Helicobacter. 2006;11 Suppl 1:36-39. |
| 7. | Miyaji H, Azuma T, Ito S, Suto H, Ito Y, Yamazaki Y, Sato F, Hirai M, Kuriyama M, Kato T. Susceptibility of Helicobacter pylori isolates to metronidazole, clarithromycin and amoxycillin in vitro and in clinical treatment in Japan. Aliment Pharmacol Ther. 1997;11:1131-1136. |
| 8. | Mégraud F. H pylori antibiotic resistance: prevalence, importance, and advances in testing. Gut. 2004;53:1374-1384. |
| 9. | Toracchio S, Marzio L. Primary and secondary antibiotic resistance of Helicobacter pylori strains isolated in central Italy during the years 1998-2002. Dig Liver Dis. 2003;35:541-545. |
| 10. | Vester B, Douthwaite S. Macrolide resistance conferred by base substitutions in 23S rRNA. Antimicrob Agents Chemother. 2001;45:1-12. |
| 11. | Ribeiro ML, Vitiello L, Miranda MC, Benvengo YH, Godoy AP, Mendonca S, Pedrazzoli J Jr. Mutations in the 23S rRNA gene are associated with clarithromycin resistance in Helicobacter pylori isolates in Brazil. Ann Clin Microbiol Antimicrob. 2003;2:11. |
| 12. | Toracchio S, Aceto GM, Mariani-Costantini R, Battista P, Marzio L. Identification of a novel mutation affecting domain V of the 23S rRNA gene in Helicobacter pylori. Helicobacter. 2004;9:396-399. |
| 13. | Alarcón T, Vega AE, Domingo D, Martínez MJ, López-Brea M. Clarithromycin resistance among Helicobacter pylori strains isolated from children: prevalence and study of mechanism of resistance by PCR-restriction fragment length polymorphism analysis. J Clin Microbiol. 2003;41:486-499. |
| 14. | Nakamura A, Furuta T, Shirai N, Sugimoto M, Kajimura M, Soya Y, Hishida A. Determination of mutations of the 23S rRNA gene of Helicobacter pylori by allele specific primer-polymerase chain reaction method. J Gastroenterol Hepatol. 2007;22:1057-1063. |
| 15. | Taylor DE, Ge Z, Purych D, Lo T, Hiratsuka K. Cloning and sequence analysis of two copies of a 23S rRNA gene from Helicobacter pylori and association of clarithromycin resistance with 23S rRNA mutations. Antimicrob Agents Chemother. 1997;41:2621-2628. |
| 16. | Fontana C, Favaro M, Minelli S, Criscuolo AA, Pietroiusti A, Galante A, Favalli C. New site of modification of 23S rRNA associated with clarithromycin resistance of Helicobacter pylori clinical isolates. Antimicrob Agents Chemother. 2002;46:3765-3769. |
| 17. | Torres J, Camorlinga-Ponce M, Pérez-Pérez G, Madrazo-De la Garza A, Dehesa M, González-Valencia G, Muñoz O. Increasing multidrug resistance in Helicobacter pylori strains isolated from children and adults in Mexico. J Clin Microbiol. 2001;39:2677-2680. |
| 18. | Versalovic J, Shortridge D, Kibler K, Griffy MV, Beyer J, Flamm RK, Tanaka SK, Graham DY, Go MF. Mutations in 23S rRNA are associated with clarithromycin resistance in Helicobacter pylori. Antimicrob Agents Chemother. 1996;40:477-480. |
| 19. | Kim KS, Kang JO, Eun CS, Han DS, Choi TY. Mutations in the 23S rRNA gene of Helicobacter pylori associated with clarithromycin resistance. J Korean Med Sci. 2002;17:599-603. |
| 20. | Matsuoka M, Yoshida Y, Hayakawa K, Fukuchi S, Sugano K. Simultaneous colonisation of Helicobacter pylori with and without mutations in the 23S rRNA gene in patients with no history of clarithromycin exposure. Gut. 1999;45:503-507. |
| 21. | Burucoa C, Landron C, Garnier M, Fauchère JL. T2182C mutation is not associated with clarithromycin resistance in Helicobacter pylori. Antimicrob Agents Chemother. 2005;49:868; author reply 868-870. |
| 22. | Garrido L, Toledo H. Novel genotypes in Helicobacter pylori involving domain V of the 23S rRNA gene. Helicobacter. 2007;12:505-509. |
| 23. | Ménard A, Santos A, Mégraud F, Oleastro M. PCR-restriction fragment length polymorphism can also detect point mutation A2142C in the 23S rRNA gene, associated with Helicobacter pylori resistance to clarithromycin. Antimicrob Agents Chemother. 2002;46:1156-1157. |
| 24. | Booka M, Okuda M, Shin K, Miyashiro E, Hayashi H, Yamauchi K, Tamura Y, Yoshikawa N. Polymerase chain reaction--restriction fragment length polymorphism analysis of clarithromycin-resistant Helicobacter pylori infection in children using stool sample. Helicobacter. 2005;10:205-213. |