Published online Jan 14, 2008. doi: 10.3748/wjg.14.292
Revised: October 13, 2007
Published online: January 14, 2008
AIM: To investigate the effect of pituitary homeobox 1 (PITX1) expression in cases of human gastric cancer on cancer differentiation and progression, and carcinogenesis.
METHODS: Using polyclonal PITX1 antibodies, we studied the expression of PITX1 in normal gastric mucosa, atypical hyperplasia, intestinal metaplasia, and cancer tissue samples from 83 gastric cancer patients by immunohistochemistry. Moreover, semi-reverse transcription polymerase chain reaction (semi-RT-PCR) was performed to detect the mRNA level of PITX1 in three gastric cancer cell lines and a normal gastric epithelial cell line. Subsequently, somatic mutations of the PITX1 gene in 71 gastric cancer patients were analyzed by a combination of denaturing high performance liquid chromatography (DHPLC) and DNA sequencing.
RESULTS: Immunohistochemistry showed that PITX1 was strongly or moderately expressed in the parietal cells of normal gastric mucosa (100%), while 55 (66.3%) out of 83 samples of gastric cancers showed decreased PITX1 expression. Moreover, PITX1 expression was reduced in 20 out of 28 cases (71.5%) of intestinal metaplasia, but in only 1 out of 9 cases (11%) of atypical hyperplasia. More importantly, PITX1 expression was significantly associated with the differentiation, position and invasion depth of gastric cancers (r = -0.316, P < 0.01; r = 0.213, P < 0.05; r = -0.259, P < 0.05, respectively). Similarly, levels of PITX1 mRNA were significantly decreased in 2 gastric cancer cell lines, BGC-823 and SGC-7901, compared with the normal gastric epithelial cell line GES-1 (0.306 ± 0.060 vs 0.722 ± 0.102, P < 0.05; 0.356 ± 0.081 vs 0.722 ± 0.102, P < 0.05, respectively). Nevertheless, no somatic mutation of PITX1 gene was found in 71 samples of gastric cancer by DHPLC analysis followed by sequencing.
CONCLUSION: Down-regulation of PITX1 may be a frequent molecular event in gastric carcinogenesis. Aberrant levels of PITX1 expression may be closely correlated with the progression and differentiation of gastric cancer.
- Citation: Chen YN, Chen H, Xu Y, Zhang X, Luo Y. Expression of pituitary homeobox 1 gene in human gastric carcinogenesis and its clinicopathological significance. World J Gastroenterol 2008; 14(2): 292-297
- URL: https://www.wjgnet.com/1007-9327/full/v14/i2/292.htm
- DOI: https://dx.doi.org/10.3748/wjg.14.292
Cancer develops as a result of multiple genetic and epigenetic alterations[12]. Functional imbalances between oncogenes and tumor suppressor genes have been identified in most cancers, including gastric cancer. Better knowledge of the changes in gene expression that occur during gastric carcinogenesis may lead to improvements in diagnosis, treatment and prevention[3]. Recently, PITX1, first identified as a bicoid-related transcription factor involved in proopiomelanocortin (POMC) gene expression[4], was reported to be a tumor suppressor in many parenchyma tumors.
The evidence that PITX1 gene is a significant tumor suppressor gene in human cancer remains largely circumstantial, but is clearly worth further study. As little is known about the tumor-inhibitory roles of PITX1 in gastric cancer, we focused on the expression of PITX1 in gastric cancer tissues and gastric cancer cell lines; the potential roles of PITX1 in cancer differentiation and carcinogenesis of the stomach will be discussed.
For immunohistochemical analysis, we used archival formalin-fixed, paraffin-embedded tissues from 83 patients with gastric cancer. The average age of patients was 61 ± 1.5 years and the male/female ratio was 63/20. Histological classification of gastric cancers was performed according to the general rules established by the Japanese Gastric Cancer Association and staging was performed according to the TNM classification of Malignant Tumors (5th edition) in 1997[5].
Tissues from 71 cases of gastric cancer were used for DNA mutation analysis. Tissue fragments were frozen and stored at -80°C for DNA extraction. DNA was isolated using a standard proteinase K digestion and phenol-choloroform extraction procedure.
All of the above tissues were obtained from the patients who underwent surgery for gastric cancer at the First Affiliated Hospital of China Medical University between December 2001 and November 2004.
Three human gastric cancer cell lines, including the poorly-differentiated MGC-803 and BGC-823 lines and the well-differentiated SGC-7901 line, as well as the normal gastric epithelial cell line GES-1, were cultured by the Research Center for Medical Genomics of China Medical University. Among the three gastric cell lines, SGC-7901 was originally obtained from metastatic lymph nodes of gastric cancer while the other two were obtained from gastric primary loci. All cell lines were grown in RPMI 1640 medium containing 10% heat-inactivated fetal calf serum at 37°C, in 99% humidity under 5% CO2, subcultured every 3 d, and checked routinely to avoid microbial contamination. Those cells growing logarithmically were harvested for total RNA isolation and RT-PCR.
An S-P Kit (Zhongshan Golden Bridge Biotech Co., Ltd) was used for immunohistochemical analysis. In brief, paraffin sections were deparaffinized in xylene and gradually rehydrated in alcohol. Antigen retrieval was performed by microwave treatment in 0.01 mol/L sodium citrate buffer (pH = 6.0) for 15 min, cooled to room temperature for 1 h, and then washed with phosphate-buffered saline (PBS). After endogenous peroxidase activity was blocked by incubation with 3% methanol for 10 min and washed with PBS, sections were incubated with normal rabbit serum for 1 h to block non-specific staining. Subsequently, sections were treated with polyclonal PITX1 antibodies (Santa Cruz, CA, USA) diluted to 1:300 with PBS, overnight at 4°C in a humidified chamber. Sections then were incubated with biotinylated anti-rabbit IgG for 10 min and then with peroxidase-labeled streptavidin for 10 min. Staining was completed with a 10-min incubation with 3,3’-diaminobenzidine. Sections were counterstained with 0.1% haematoxylin. Intracytoplasmic and/or intranuclear buffy granular staining in gastric mucous cells was regarded as positive. Ten visual fields were randomly selected from each section, and at least 100 cells were counted in each field.
Omission of primary antibodies acted as a negative control, while normal gastric tissue was regarded as a positive control. Immunohistochemical results were assessed as ‘negative’ if fewer than 50% of cancer cells were stained. When at least 50% of cancer cells were stained, immunostaining was considered to be ‘positive’.
Total RNA was isolated from the cell lines using Trizol (Invitrogen, CA, USA), and 1 &mgr;g of total RNA was converted to cDNA using a First Strand cDNA Synthesis Kit (Takara, Japan). Then, PCR reactions were performed in volumes of 20 &mgr;L. A 1 &mgr;L aliquot of the cDNA was used for PCR amplification with the primers 5’-ATGGACGCCTTCAAGGGGGGC-3’ (forward) and 5’-TGCAACTGCTGGCTTGTGAAG-3’ (reverse), resulting in a DNA product of 305 bp. PCR conditions were 95°C for 1 min, 55°C for 30 s and 72°C for 45 s, for 30 cycles. Under the same PCR conditions, glyceraldehyde-3-phosphate dehydrogenase (GAPDH) transcripts (227 bp) were amplified as an external control for RT-PCR analysis from the same cDNA samples, using the following primer pair: 5’-GACCACAGTCCATGCCATCAC-3’ (forward) and 5’-GTCCACCACCCTGTTGCTGTA-3’ (reverse).
PCR products were separated by 1.5% agarose gel electrophoresis, stained with ethidium bromide, and visualized under UV light. Electrophoresis strips were analyzed by Fluor-S software. By calculating the ratio of the IDVs (integrated density value) for PITX1 transcripts to those for GAPDH transcripts, we compared the PITX1 mRNA levels between gastric cancer cell lines and the normal gastric epithelial cell line.
DNA was amplified in a final reaction volume of 20 &mgr;L using 100 ng of genomic DNA, 10 × buffer, 1.5 mmol/L MgCl2, 200 &mgr;mol/L dNTPs, 0.27 &mgr;mol/L (100 ng) primers and 1.25 units of rTaq polymerase (Takara, Japan).
In order to form the presumed heteroduplex, unpurified PCR products were pretreated by heating at 94°C followed by cooling down to 25°C at a rate of 0.03°C/s. DHPLC was carried out using the Transgenomic WAVEMAKE DNA fragment analysis system (Transgenomic, Ohama, USA). PCR products showing aberrant chromatograms on DHPLC were subjected to direct DNA sequencing.
The software SSPS for Windows version 11.5 was used for statistical analysis. Correlations between clinicopathological parameters and PITX1 expression were analyzed by χ2-test, and comparisons of the mRNA levels in different cell lines were performed using t-tests. P values of less than 0.05 were considered to be statistically significant.
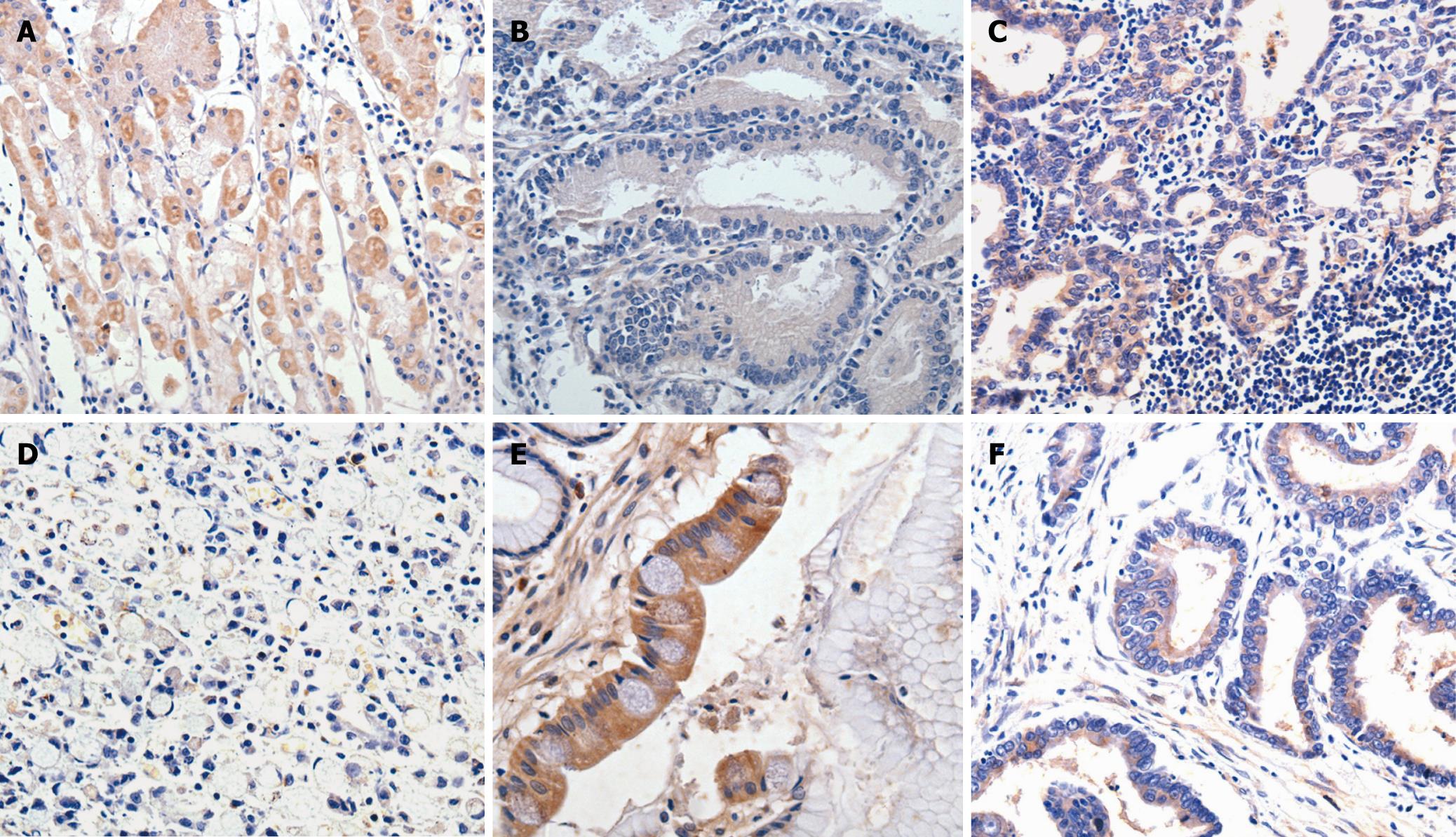
As shown in Table 1, all parietal cells (100%) of normal gastric mucosa showed strong or moderate PITX1 staining, while 55 (66.3%) out of 83 gastric cancer samples showed decreased staining for PITX1 (P < 0.001). In addition, the expression of PITX1 in intestinal metaplasia tissue samples was reduced in 20 out of 28 cases (71.5%), but in only 1 out of 9 (11%) cases of atypical hyperplasia.
The correlations between clinicopathologic parameters and PITX1 expression in 83 patients are shown in Table 2. PITX1 expression did not differ significantly by age, gender, size, clinical staging and lymph node metastasis. However, significant differences in PITX1 expression were found in tumors with different degrees of differentiation, position or depth of invasion. That is, reduced expression of PITX1 was significantly linked with tumors with poor differentiation, located in the upper and middle parts of the stomach, or with T3/T4 invasion depth (P < 0.01, P < 0.05, and P < 0.05 respectively). Representative results of PITX1 expression in gastric cancer samples are depicted in Figure 1.
| PITX1 | |||
| n | Negative (%) | Positive(%) | |
| Gender | |||
| Male | 53 | 42 (66.7) | 21 (33.3) |
| Female | 20 | 13 (65.0) | 7 (35.0) |
| Age (yr) | |||
| < 60 | 32 | 22 (68.7) | 10 (31.3) |
| ≥ 60 | 51 | 33 (64.7) | 7 (35.3) |
| Position | |||
| Upper (cardia & fundus) | 16 | 12 (75) | 4 (25) |
| Middle (body) | 15 | 14 (93.3) | 1 (6.7) |
| Lower (antrum) | 53 | 30 (56.6)a | 23 (43.4)a |
| Depth of invasion | |||
| T1/T2 | 29 | 15 (48.4) | 16 (51.6) |
| T3/T4 | 54 | 40 (74.1)c | 14 (25.9)c |
| Clinical stage | |||
| Early stage | 17 | 8 (47.1) | 9 (52.9) |
| Advanced stage | 66 | 47 (71.2) | 19 (28.8) |
| Differentiation type | |||
| Well1 | 38 | 19 (50) | 19 (50) |
| Poorly2 | 45 | 36 (80)b | 9 (20)b |
| Lymph-node metastasis | |||
| Yes | 47 | 33 (70.2) | 14 (29.8) |
| No | 36 | 22 (61.1) | 14 (38.9) |
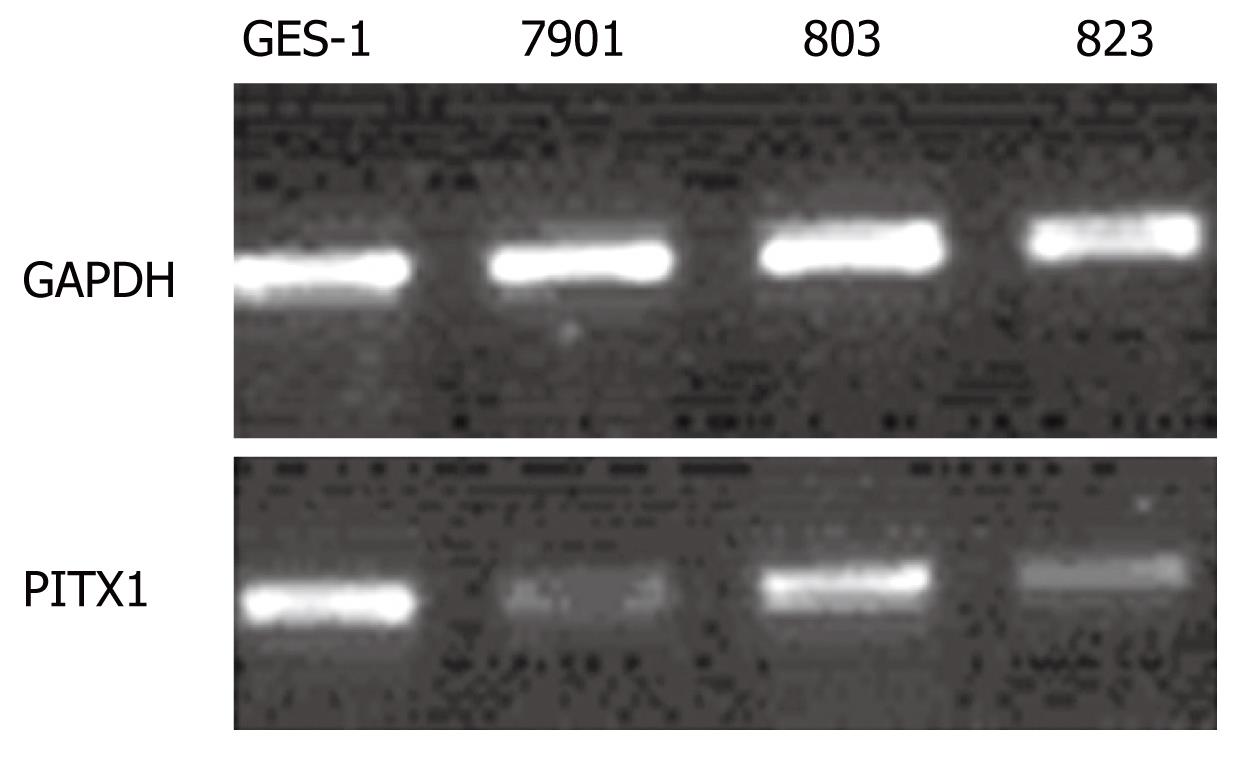
RT-PCR was repeated 12 times to analyze the differences in PITX1 mRNA expression between the three gastric cancer cell lines and the normal GES-1 cell line (Figure 2). The average IDV ratios of the gastric cancer cell lines BGC-823 and SGC-7901 were significantly lower than that in the normal gastric cell line GES-1 (P < 0.05); there was no statistically significant decrease in IDV ratio in the MGC-803 cell line (Table 3).
All of the DHPLC chromatographies in coding area from 71 tumor tissues showed a normal single peak shape except that the deletion of a repeated sequence “gggcgc” in 3’UTR in one sample was found through DHPLC screening and sequencing.
Carcinogenesis refers to changes of multiple genes, and is closely related to the corresponding transcription factors (TFs) and their mutual regulation. As a family of TFs, homeobox genes not only play important roles in embryonic development and differentiation, but also control the differentiation and proliferation of mature tissues[67]. ClassI human homeobox-containing genes (HOX genes) have been reported to act as a network of transcriptional regulators involved in the process of cell to cell communication during normal morphogenesis, the alteration of which may contribute to the evolution of cancer[8]. Moreover, a recent study concluded that HOX genes may play a critical role in the genesis and progression of gastric cancer[9].
PITX1 and PITX2 are members of the PITX (pituitary homeobox) family, a family of homeobox genes; they have been widely researched and contain a highly conserved homeodomain (HD) with a lysine residue at amino acid position 50, which is responsible for recognition of the TAA (T/G) CC motif in the promoter regions of several target genes[10]. Besides sharing similar DNA binding specificities, PITX1 and PITX2 are co-expressed in the stomodeum and have overlapping expression patterns in stomodeal derivatives[11–13]. Specifically, PITX1 is responsible for the morphology of muscles, tendons and the bones of hind limbs, and is involved in the development of anterior structures as well as pituitary and craniofacial morphogenesis[1415].
Although PITX1 has essential roles in human development, the possible involvement of PITX1 in human cancer has been suspected since an elegant study by Gidekel et al[16] showed that a homeobox gene normally required for the differentiation of a particular tissue can play a causal role in the carcinogenesis of the same tissue if expressed at the wrong time, at the wrong levels, or in the wrong contexts. For instance, PITX1 was found in adrenocorticotropic hormone (ACTH or corticotrophin) -positive tumors as well as in ACTH-negative carcinoids[17]. Expression of PITX1 mRNA might be a potential biomarker of malignancy in patients with Barrett’s esophagus[18]. Moreover, PITX1 expression is also decreased in lung, prostate and bladder tumor tissues compared with normal control tissues[19–22].
Considering that the expression of the homeobox genes, such as BARX1 and PITX1, is confined to the stomach mesenchyme during gastrointestinal development[23], we studied PITX1 expression in human gastric cancer tissues. In our study, PITX1 expression declined in 63.3% cases of gastric cancer, compared with adjacent normal tissues (P < 0.001). Together with former reports on the role of PITX1 in various cancers, our results hinted the negative regulation of PITX1 during carcinogenesis and the development of gastric cancer.
In addition, we found that PITX1 mRNA expression in gastric cancer cell lines was in accordance with its protein expression in immunohistochemistrical results, which agrees with the research of Chen et al[19] in lung cancer. Given the insufficient evidence of gastric cancer cell lines, a further study on PITX1 mRNA expression in gastric cancer tissues is necessary to confirm the results of immunohistochemistry and RT-PCR in cell lines.
The low expression of PITX1 suggested that PITX1 may have a tumor suppressive function, inspiring us to examine its potential role in carcinogenesis. Kolfschoten et al[22], reported that PITX1 regulates the RAS (rat sarcoma) signaling pathway and, therefore, tumorigenesis. As a transcriptional activator, PITX1 might activate RASAL1 to inhibit RAS activity, because PITX1 consensus binding sites, TAA(T/G)CC, were found in the 2 kb upstream promoter region of RASAL1, a member of the RAS-GAP family of genes (GTP-activating factors-negative regulators of RAS). Another novel explanation for PITX1 tumorigenesis came from a study on the role of PITX1 in the transcription of the p53 tumor suppressor gene in human mammary carcinoma (MCF-7) cells, implying that p53 is a direct transcriptional target of PITX1[24].
It has been long recognized that the inactivation of tumor suppressor genes results from mutation, loss of heterozygosity (LOH), aberrant methylation and haploinsufficiency[2526], among which aberrant promoter hypermethylation is well-known to participate in homeobox gene silencing[2728]; however, PITX1 promoter hypermethylation has not been found in studies of lung cancer[19]. Therefore, we examined the PITX1 gene for the presence of mutations in order to explore the mechanism of inactivation of this candidate tumor suppressor gene. We failed to find any mutation in the coding area of PITX1, although we did identify the deletion of a repeated sequence in 3’UTR of only one gastric cancer sample. Hence, the absence of detectable mutations or epigenetic silencing suggest that haploinsufficiency is behind the loss of PITX1 expression[29]. Indeed, it was recently shown that the striking phenotype observed in patients with Rieger’s syndrome resulted from decreased PITX2 expression due to deletion mutations in one allele of the gene[30]. As a close relative of PITX2, is PITX1 also subject to haploinsufficiency in gastric carcinogenesis? Further studies are required to investigate the mechanism of PITX1 down-regulation in gastric cancers.
Our study showed that the levels of PITX1 protein are decreased in gastric cancer tissues and that PITX1 mRNA levels are decreased in gastric cancer cell lines, suggesting that PITX1 is a candidate tumor suppressor gene in gastric cancers, and that its expression is related to the differentiation, position and depth of invasion of gastric cancers.
Knowledge of transcription factors is crucial for understanding the molecular basis of neoplasia. PITX1, a pituitary transcription factor, mediates a plethora of embryonic functions, and its down-regulation in parenchyma tumors may be associated with carcinogenesis. Our study is the first time to explore the link between PITX1 and gastric cancer. Down-regulation of PITX1 both in gastric cancer and gastric cancer cell lines may be involved in the development of gastric cancer and may be a frequent molecular event in gastric carcinogenesis; thus, PITX1 represents a candidate tumor suppressor gene. Therefore, further study should investigate potential mechanisms for modulating PITX1 expression during carcinogenesis and the progression of gastric cancer.
Gastric cancer is one of the most common types of cancer worldwide. A more complete molecular genetic understanding of gastric cancer will allow the development of specific therapies. Recently, studies on tumor suppressor genes have been highly valued in gastric cancer research owing to their widespread involvement in diverse cellular activities.
A family of homeobox genes, the PITX family, contains a highly-conserved homeodomain responsible for recognition of the TAA(T/G)CC motif in the promoter regions of several target genes. By virtue of its DNA-binding specificity, PITX1, an important member of this family, has been reported to be involved in many human cancers, such as Barrett’s esophagus and lung cancer. The carcinogenic mechanism of PITX1 has been explored in terms of regulation of the RAS signaling pathway and direct activation of p53 transcription.
The study observed a change in the expression levels of PITX1 that parallels the progression of gastric carcinogenesis; we discussed the possible carcinogenic mechanisms used by PITX1 and its clinicopathological significance in gastric cancer. This is the first study to explore the link between PITX1 and gastric cancer.
The present study suggests that down-regulation of PITX1 may be a frequent molecular event in gastric carcinogenesis. However, the exact function and mechanism of PITX1 in gastric cancer remain to be elucidated.
Homeobox genes contain a 180-base-pair segment (the homeobox) that encodes a protein domain (homeodomain) that is involved in binding to and, thus, regulating the expression of DNA; members of the Ras family of small G proteins have been implicated as key intermediates mediating signals from upstream tyrosine kinases to downstream cascades of serine/threonine kinases, which then activate the nuclear factors that control gene expression and protein synthesis.
An excellent paper - well written, nicely researched. The results support the hypothesis but further research across multiple centers is needed to establish a positive correlation.
| 1. | Yasui W, Yokozaki H, Fujimoto J, Naka K, Kuniyasu H, Tahara E. Genetic and epigenetic alterations in multistep carcinogenesis of the stomach. J Gastroenterol. 2000;35 Suppl 12:111-115. |
| 2. | Ushijima T, Sasako M. Focus on gastric cancer. Cancer Cell. 2004;5:121-125. |
| 3. | Sutter AP, Fechner H. Gene therapy for gastric cancer: is it promising? World J Gastroenterol. 2006;12:380-387. |
| 4. | Lamonerie T, Tremblay JJ, Lanctot C, Therrien M, Gauthier Y, Drouin J. Ptx1, a bicoid-related homeo box transcription factor involved in transcription of the pro-opiomelanocortin gene. Genes Dev. 1996;10:1284-1295. |
| 5. | Sobin LH, Fleming ID. TNM Classification of Malignant Tumors, fifth edition (1997). Union Internationale Contre le Cancer and the American Joint Committee on Cancer. Cancer. 1997;80:1803-1804. |
| 6. | Imoto I, Pimkhaokham A, Watanabe T, Saito-Ohara F, Soeda E, Inazawa J. Amplification and overexpression of TGIF2, a novel homeobox gene of the TALE superclass, in ovarian cancer cell lines. Biochem Biophys Res Commun. 2000;276:264-270. |
| 7. | Duboule D. Vertebrate Hox genes and proliferation: an alternative pathway to homeosis? Curr Opin Genet Dev. 1995;5:525-528. |
| 8. | Cillo C, Faiella A, Cantile M, Boncinelli E. Homeobox genes and cancer. Exp Cell Res. 1999;248:1-9. |
| 9. | Rossi Degl’innocenti D, Castiglione F, Buccoliero AM, Bechi P, Taddei GL, Freschi G, Taddei A. Quantitative expression of the homeobox and integrin genes in human gastric carcinoma. Int J Mol Med. 2007;20:621-629. |
| 10. | Tremblay JJ, Goodyer CG, Drouin J. Transcriptional properties of Ptx1 and Ptx2 isoforms. Neuroendocrinology. 2000;71:277-286. |
| 11. | Tremblay JJ, Lanctot C, Drouin J. The pan-pituitary activator of transcription, Ptx1 (pituitary homeobox 1), acts in synergy with SF-1 and Pit1 and is an upstream regulator of the Lim-homeodomain gene Lim3/Lhx3. Mol Endocrinol. 1998;12:428-441. |
| 12. | Gage PJ, Camper SA. Pituitary homeobox 2, a novel member of the bicoid-related family of homeobox genes, is a potential regulator of anterior structure formation. Hum Mol Genet. 1997;6:457-464. |
| 13. | Lanctot C, Lamolet B, Drouin J. The bicoid-related homeoprotein Ptx1 defines the most anterior domain of the embryo and differentiates posterior from anterior lateral mesoderm. Development. 1997;124:2807-2817. |
| 14. | Szeto DP, Rodriguez-Esteban C, Ryan AK, O’Connell SM, Liu F, Kioussi C, Gleiberman AS, Izpisua-Belmonte JC, Rosenfeld MG. Role of the Bicoid-related homeodomain factor Pitx1 in specifying hindlimb morphogenesis and pituitary development. Genes Dev. 1999;13:484-494. |
| 15. | DeLaurier A, Schweitzer R, Logan M. Pitx1 determines the morphology of muscle, tendon, and bones of the hindlimb. Dev Biol. 2006;299:22-34. |
| 16. | Gidekel S, Pizov G, Bergman Y, Pikarsky E. Oct-3/4 is a dose-dependent oncogenic fate determinant. Cancer Cell. 2003;4:361-370. |
| 17. | Messager M, Carriere C, Bertagna X, de Keyzer Y. RT-PCR analysis of corticotroph-associated genes expression in carcinoid tumours in the ectopic-ACTH syndrome. Eur J Endocrinol. 2006;154:159-166. |
| 18. | Lord RV, Brabender J, Wickramasinghe K, DeMeester SR, Holscher A, Schneider PM, Danenberg PV, DeMeester TR. Increased CDX2 and decreased PITX1 homeobox gene expression in Barrett's esophagus and Barrett's-associated adenocarcinoma. Surgery. 2005;138:924-931. |
| 19. | Chen Y, Knosel T, Ye F, Pacyna-Gengelbach M, Deutschmann N, Petersen I. Decreased PITX1 homeobox gene expression in human lung cancer. Lung Cancer. 2007;55:287-294. |
| 20. | Singh D, Febbo PG, Ross K, Jackson DG, Manola J, Ladd C, Tamayo P, Renshaw AA, D’Amico AV, Richie JP. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell. 2002;1:203-209. |
| 21. | Ramaswamy S, Tamayo P, Rifkin R, Mukherjee S, Yeang CH, Angelo M, Ladd C, Reich M, Latulippe E, Mesirov JP. Multiclass cancer diagnosis using tumor gene expression signatures. Proc Natl Acad Sci USA. 2001;98:15149-15154. |
| 22. | Kolfschoten IG, van Leeuwen B, Berns K, Mullenders J, Beijersbergen RL, Bernards R, Voorhoeve PM, Agami R. A genetic screen identifies PITX1 as a suppressor of RAS activity and tumorigenicity. Cell. 2005;121:849-858. |
| 23. | Kim BM, Buchner G, Miletich I, Sharpe PT, Shivdasani RA. The stomach mesenchymal transcription factor Barx1 specifies gastric epithelial identity through inhibition of transient Wnt signaling. Dev Cell. 2005;8:611-622. |
| 24. | Liu DX, Lobie PE. Transcriptional activation of p53 by Pitx1. Cell Death Differ. 2007;14:1893-1907. |
| 26. | Tamura G. Alterations of tumor suppressor and tumor-related genes in the development and progression of gastric cancer. World J Gastroenterol. 2006;12:192-198. |
| 27. | Pilozzi E, Onelli MR, Ziparo V, Mercantini P, Ruco L. CDX1 expression is reduced in colorectal carcinoma and is associated with promoter hypermethylation. J Pathol. 2004;204:289-295. |
| 28. | Nagai H, Li Y, Hatano S, Toshihito O, Yuge M, Ito E, Utsumi M, Saito H, Kinoshita T. Mutations and aberrant DNA methylation of the PROX1 gene in hematologic malignancies. Genes Chromosomes Cancer. 2003;38:13-21. |