Copyright
©The Author(s) 2020.
World J Gastroenterol. Sep 21, 2020; 26(35): 5223-5247
Published online Sep 21, 2020. doi: 10.3748/wjg.v26.i35.5223
Published online Sep 21, 2020. doi: 10.3748/wjg.v26.i35.5223
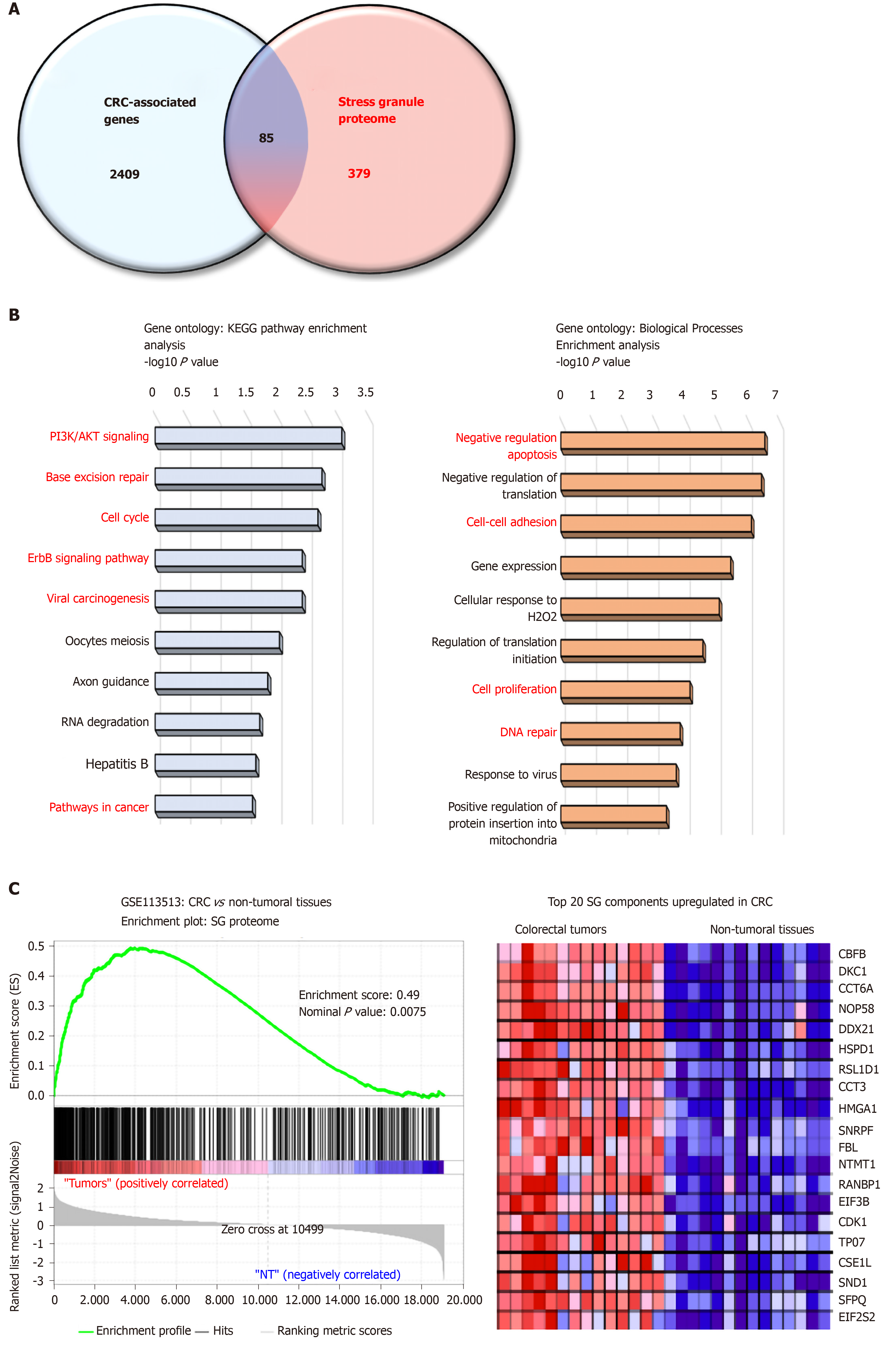
Figure 1 The stress granule proteome contains several colorectal cancer-associated proteins.
A: Venn diagram merging a list of colorectal cancer (CRC) -associated genes (retrieved from Metacore software) and the mammalian stress granule (SG) proteome from https://msgp.pt; B: Gene ontology analysis of SG proteome using KEGG pathway and biological processes analysis. Enrichment is represented with a-log10 P value. Processes and pathways in red are those involved in cancer development; C: Gene-Set Enrichment Analysis (version 3.0, Broad Institute, Cambridge, MA, United States) of the SG proteome on CRC patients (GSE113513). The top 20 genes upregulated in CRC patients as compared to non-tumoral tissues are represented in a heatmap. The enrichment score was calculated using the number of genes ranking at the top or the bottom of the gene list (permutation type: Phenotype; with 1000 permutations). The Signal2Noise was used for ranking genes. A nominal P value < 0.05 and an FDR < 0.2 were considered significant.
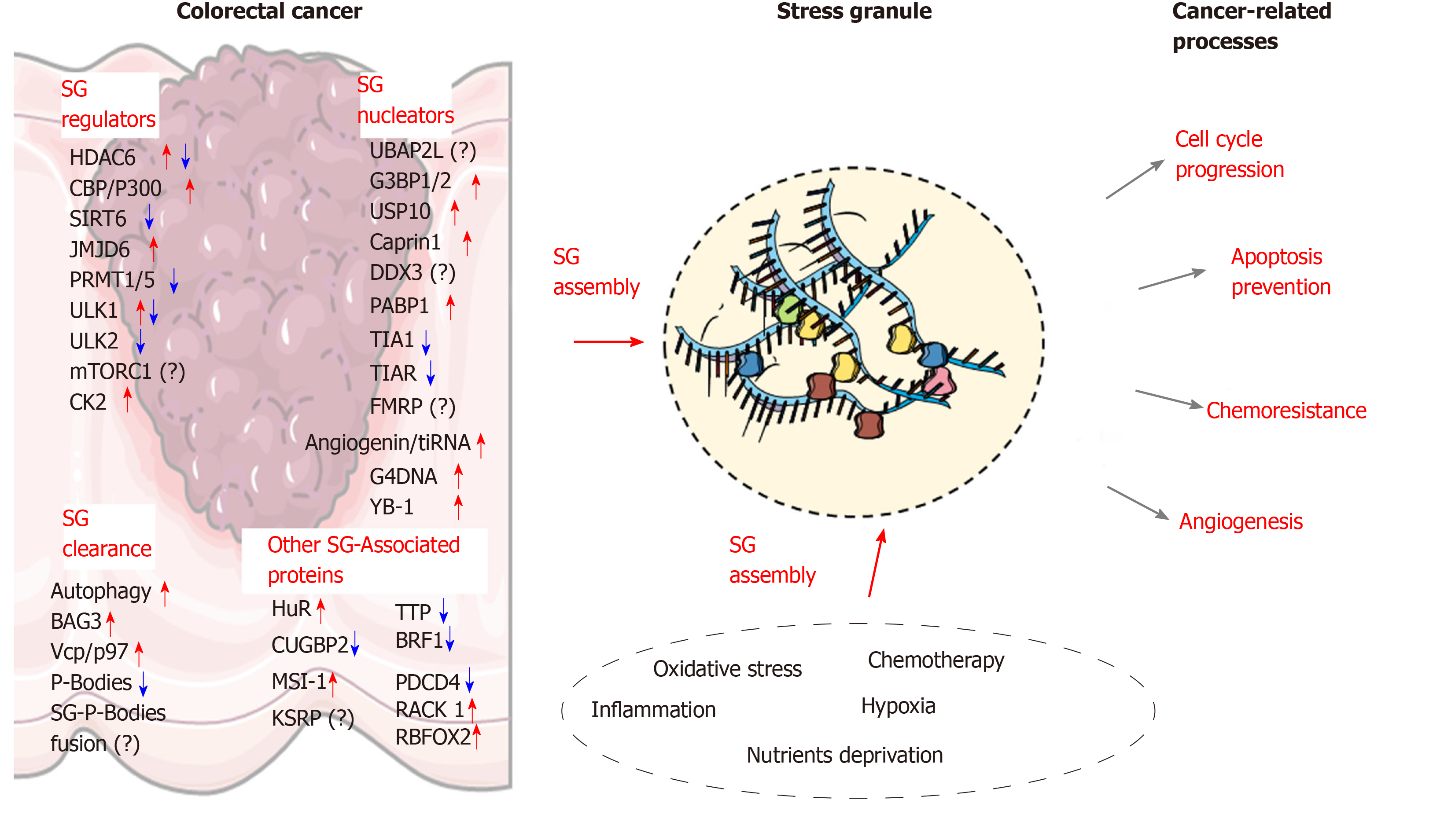
Figure 2 The molecular landscape underlying stress granules formation in colorectal cancer.
Stress granules (SGs) assembly in colorectal cancer cells is associated with several alterations in the expression of proteins involved in SG nucleation or clearance. The stress-related conditions within the tumor microenvironment and various antitumor agents can further promote SGs assembly. Several SG-associated proteins (RNA-binding proteins or others) contribute to various cancer-related processes such as cell cycle progression, apoptosis inhibition, angiogenesis, and chemoresistance. Illustrations were retrieved from Servier Medical art (https://smart.servier.com/). SG: Stress granules.
- Citation: Legrand N, Dixon DA, Sobolewski C. Stress granules in colorectal cancer: Current knowledge and potential therapeutic applications. World J Gastroenterol 2020; 26(35): 5223-5247
- URL: https://www.wjgnet.com/1007-9327/full/v26/i35/5223.htm
- DOI: https://dx.doi.org/10.3748/wjg.v26.i35.5223