Published online Jun 21, 2013. doi: 10.3748/wjg.v19.i23.3615
Revised: April 2, 2013
Accepted: April 9, 2013
Published online: June 21, 2013
Processing time: 121 Days and 18.5 Hours
AIM: To identify genes associated with gastric precancerous lesions in Helicobacter pylori (H. pylori)-susceptible ethnic Malays.
METHODS: Twenty-three Malay subjects with H. pylori infection and gastric precancerous lesions identified during endoscopy were included as “cases”. Thirty-seven Malay subjects who were H. pylori negative and had no precancerous lesions were included as “controls”. Venous blood was collected for genotyping with Affymetrix 50K Xba1 kit. Genotypes with call rates < 90% for autosomal single nucleotide polymorphisms (SNPs) were excluded. For each precancerous lesion, associated SNPs were identified from Manhattan plots, and only SNPs with a χ2P value < 0.05 and Hardy Weinberg Equilibrium P value > 0.5 was considered as significant markers.
RESULTS: Of the 23 H. pylori-positive subjects recruited, one sample was excluded from further analysis due to a low genotyping call rate. Of the 22 H. pylori-positive samples, atrophic gastritis only was present in 50.0%, complete intestinal metaplasia was present in 18.25%, both incomplete intestinal metaplasia and dysplasia was present in 22.7%, and dysplasia only was present in 9.1%. SNPs rs9315542 (UFM1 gene), rs6878265 (THBS4 gene), rs1042194 (CYP2C19 gene) and rs10505799 (MGST1 gene) were significantly associated with atrophic gastritis, complete intestinal metaplasia, incomplete metaplasia with foci of dysplasia and dysplasia, respectively. Allele frequencies in “cases”vs“controls” for rs9315542, rs6878265, rs1042194 and rs10505799 were 0.4 vs 0.06, 0.6 vs 0.01, 0.6 vs 0.01 and 0.5 vs 0.02, respectively.
CONCLUSION: Genetic variants possibly related to gastric precancerous lesions in ethnic Malays susceptible to H. pylori infection were identified for testing in subsequent trials.
Core tip: Gastric cancer and its precancerous lesions are exceptionally rare among ethnic Malays. Gene variants may be associated with precancerous lesions in Helicobacter pylori-susceptible Malays. Genome-wide association was performed to identify gene variants in Malays with a spectrum of gastric precancerous lesions. Results indicated that at different phases of the Correa cascade, different gene variants were manifest, but they followed a pattern of progression similar to their histological and clinical stages. It is possible that, in addition to histological staging, gene variant markers may serve to identify different phases of gastric cancer progression in the near future.
-
Citation: Maran S, Lee YY, Xu S, Rajab NS, Hasan N, Syed Abdul Aziz SH, Majid NA, Zilfalil BA. Gastric precancerous lesions are associated with gene variants in
Helicobacter pylori -susceptible ethnic Malays. World J Gastroenterol 2013; 19(23): 3615-3622 - URL: https://www.wjgnet.com/1007-9327/full/v19/i23/3615.htm
- DOI: https://dx.doi.org/10.3748/wjg.v19.i23.3615
Gastric cancers are thought to arise from a cascade of histological changes or precancerous lesions (atrophic gastritis, intestinal metaplasia and dysplasia) before developing into full-blown malignancy[1]. In Japan, studies have shown that surveillance of these precancerous lesions is associated with increased detection of early gastric cancers and improved survival rates[2,3].
These precancerous lesions are associated with Helicobacter pylori (H. pylori) infection acquired since childhood[4]. In populations with a high prevalence of H. pylori infection, including those in China and Japan, precancerous lesions can be detected in up to 80% of adults[5]. Eradication of H. pylori infection at this stage has not been shown to be effective in these high risk populations[6].
Ethnic Malays residing in the north-eastern region of Peninsular Malaysia (state of Kelantan) have an exceptionally low prevalence of H. pylori infection[7,8]. Exact reasons for this low prevalence are unknown, but it could be a combination of unique environmental, host and strain virulence factors shaped by the population’s evolutionary history[9-12]. Due to the extremely low acquisition of H. pylori infection, gastric cancer and its precancerous lesions are extremely rare in this population[13-15].
In a survey of 234 subjects undergoing upper endoscopy in a tertiary hospital from the state of Kelantan, the reported rate of atrophic gastritis was 42.3% and intestinal metaplasia was present in 7.7% (14/234) of all biopsies, but was only present in 1.4% (2/146) of the ethnic Malays[15]. This low rate of gastric precancerous lesions observed was a result of a low prevalence of H. pylori infection in the studied population of only 6.8%. As shown in a multivariable analysis, the risk of intestinal metaplasia and dysplasia was only significant in the presence of H. pylori infection[15].
A minority of this Malay population is genetically susceptible to H. pylori infection, and DCC gene polymorphism has recently been found to be responsible[16]. An aberrant methylation of this tumor suppressor gene has been observed to occur in the course of gastric carcinogenesis[17]. As such, this population may also be genetically susceptible to the development of gastric precancerous lesions.
The current study aimed to determine the gene polymorphisms associated with gastric precancerous lesions in the Malay population from north-eastern region of Peninsular Malaysia using the genome-wide association approach.
Only those ethnic Malay subjects (age range 20-80 years) whose gastrointestinal symptoms required upper endoscopy were screened for study eligibility. To avoid ascertainment bias, subjects had upper gastrointestinal symptoms (including dyspepsia and/or abdominal discomfort) and required upper endoscopy to exclude gastro-duodenal diseases before being included into the study.
All Malay subjects included in the study were born in the state of Kelantan, had resided within the region for at least 3 generations and were from different families but had similar socio-economic and socio-cultural backgrounds. Subjects positive for H. pylori infection according to a urease test and histology and with gastric precancerous lesions identified during endoscopy were categorized as “cases”, while those negative for H. pylori infection and precancerous lesions were categorized as “controls”. “Cases” and “controls” were matched for age and gender. Subjects satisfying the above inclusion criteria were recruited into the study. Exclusion criteria included an intake of antibiotics 3 mo prior to the upper endoscopy test, upper gastrointestinal bleeding, a positive family history of H. pylori infection and gastric cancer, a previous history of H. pylori infection and chronic psychiatric and medical conditions, including cancer. Informed consent was obtained from all subjects prior to their enrolment into the study.
Cases with H. pylori infection and positive for precancerous lesions were extremely limited in number due to an exceptionally low rate of H. pylori infection among ethnic Malays. Only 23 Malay subjects were eventually included as “cases”. A larger sample size for the “controls” was sought to compensate for the low sample size in “cases”. Furthermore, stringent criteria were set to ensure that only subjects of similar age, socio-economic and socio-cultural backgrounds were included in the study. From a total of 45 screened subjects, 37 Malay subjects were recruited as “controls” with eight subjects being excluded as they did not meet the inclusion criteria, they did not give consent or blood samples were poor.
The study was approved by the Human Research and Ethics Committee of Universiti Sains Malaysia (USM).
All upper endoscopies (model GIF-140 and GIF-160; Olympus Medical Systems, Tokyo, Japan) during this period were performed by one endoscopist with at least 5 years’ experience. If needed, patients were sedated accordingly. Subjects who did not stop proton pump inhibitors 2 wk before endoscopy, those who had received antibiotics prior to study, and patients who had upper gastrointestinal bleeding shortly before the study were excluded.
Endoscopic findings of gastritis and atrophy were recorded and classified based on established Sydney criteria[18] and Atrophy Club criteria[19]. Biopsies were taken using standard biopsy forceps at the antrum, incisura and body. A minimum of 2 to 4 biopsies (size between 2 to 4 mm) were taken in each sites and these gastric biopsies, preserved in formalin containers, were transported to the pathology laboratory on the same day.
Only one histopathologist was involved in reviewing all the slides. All biopsies were stained with routine hematoxylin and eosin (HE) stain followed by Alcian blue-periodic acid Schiff stain for the detection of intestinal metaplasia. The Warthin Starry stain would be used in sections where the H. pylori bacterium was not detected in the routine HE stain.
Chronic atrophic gastritis was identified based on the updated 1994 Sydney system[20] and Atrophy Club definitions[19]. Intestinal metaplasia was identified by replacing glandular epithelium with goblet cells[21]. Intestinal metaplasia was classified into complete or incomplete types. Complete type resembled the small intestinal phenotype with well-formed goblet cells while incomplete type resembled the colonic phenotype with irregular mucin droplets and absence of a brush border. Dysplasia was identified by epithelium disarray and increased nucleo-cytoplasmic ratio[22].
For the purpose of the genotyping study, subjects were grouped as follows: atrophic gastritis only, complete intestinal metaplasia, incomplete metaplasia with foci of dysplasia, or dysplasia only.
All recruited subjects were called up by one of the investigators (SM) to have 1 mL of venous blood taken during the study day. Unlike conventional methods of DNA extraction, 1 mL of blood was sufficient for the commercially available kits. The blood was collected in an EDTA tube and was transported immediately to a facility (Human Genome Centre, USM, Kubang Kerian, Malaysia) to be stored at 4 °C. Subsequently, DNA for all recruited cases and controls was isolated using QIAamp DNA Blood Mini Kit (QIAGEN, Hilden, Germany).
The isolated DNA from all recruited cases (n = 23) and controls (n = 37) were processed and genotyped using Affymetrix 50k Xba1 array (Affymetrix, United States) following the instructions provided in the Affymetrix GeneChip Human Mapping 100K Assay Manual[23]. Genotypes with call rates < 90% for autosomal single nucleotide polymorphisms (SNPs) were excluded. SNPs that had a minor allele frequency < 5%, that failed to genotype in > 5% of samples, or had a Hardy-Weinberg Equilibrium (HWE) P-value < 0.5 were also excluded from the analysis.
Genotype calling to assess the normalization of the SNPs was performed with the Bayesian Robust Linear Model with Mahalanobis distance classifier (BRLMM) algorithm from the Affymetrix® Genotyping Console™ software version 4.0 (Affymetrix, United States). Quality control for genetic markers was assessed using the Genotype filtering tool in the SVS Golden Helix Bioinformatics Tools version 7.4 (Golden Helix Inc., Bozeman, MT, United States).
Association was evaluated for every single SNP in each gene with SVS Golden Helix Bioinformatics Tools. False Discovery Rate, and Bonferroni adjustments were used for multiple-testing corrections. A Manhattan plot for each phenotype was generated to determine SNPs with the highest significant value associated with that phenotype using SVS Golden Helix Bioinformatics Tools (version 7.4). A significant genomic threshold of 3 × 10-7 in Manhattan plots was set in this study and a χ2P value for each SNP was calculated based on Fisher’s exact χ2 test. For each type of precancerous lesion studied, of which a group of associated SNPs were identified from Manhattan plots, only a SNP with χ2P value < 0.05 and HWE P value > 0.5 was considered as a significant marker.
Of the 23 H. pylori-positive subjects recruited, one sample was excluded from further analysis due to a low genotyping call rate (< 90%). The mean age of the remaining 22 “cases” was 56.5 ± 16.5 years, and was 53.2 ± 15.2 years for the 37 “controls”. Cases were 54.5% (12/22) male compared with 50% (17/37) in “controls”. Of subjects positive for H. pylori infection (cases), atrophic gastritis only was present in 50.0% (11/22), complete intestinal metaplasia was present in 18.2% (4/22), both incomplete intestinal metaplasia and dysplasia was present in 22.7% (5/22) and dysplasia only was present in 9.1% (2/22). None of the gastric precancerous lesions were present in subjects negative for H. pylori infection (controls).
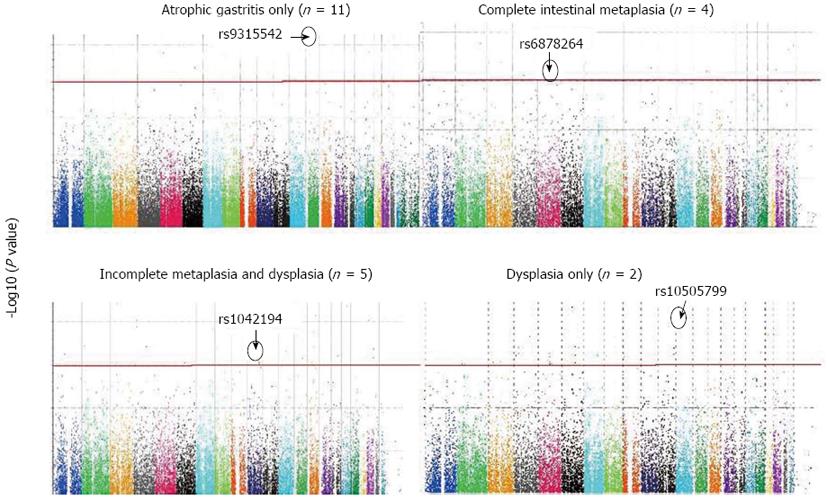
In 10 “cases” with atrophic gastritis only, compared with controls, 26 SNPs were above the significant genomic threshold. Five of the identified 26 SNPs were in HWE, of which rs9315542, located in chromosome 13 q13.3 (UFM1 gene), was the most significant associated SNP (χ2P value = 0.007) (Table 1, Figure 1). The allele frequency for rs9315542 in cases vs controls was 0.4 vs 0.06.
| rs ID | Chromosome | Gene | Minor allele | Allele frequency | χ2 | HWE | |
| Cases | Control | P-value1 | P-value | ||||
| rs2614074 | 8 p21.1 | PNOC | B | 0.409 | 0.062 | 0.008 | 0.983 |
| rs10504944 | 8 q22.1 | GDF6 | A | 0.5 | 0 | 0.034 | 0.516 |
| rs9315542 | 13 q13.3 | UFM1 | B | 0.409 | 0.064 | 0.007 | 0.994 |
| rs4943552 | 13 q13.3 | UFM1 | A | 0.318 | 0.075 | 0.040 | 0.815 |
| rs489977 | 18 q12.3 | KC6 | A | 0.181 | 0 | 0.064 | 0.752 |
In 4 “cases” with intestinal metaplasia only, compared with controls, 13 SNPs were above the genomic threshold and were in HWE, of which rs6878264, located in intron 4 of the thrombospondin 4 (THBS4) gene, was the most significant associated SNP (χ2P value = 0.01) (Table 2, Figure 1). The allele frequency for rs6878264 in cases vs controls was 0.6 vs 0.01.
| rs ID | Chromosome | Gene | Minor allele | Allele frequency | χ2 | HWE | |
| Cases | Control | P-value | P-value | ||||
| rs1166704 | 1p31.1 | NEXN | B | 0.375 | 0.031 | 0.0694 | 0.654 |
| rs2191508 | 2q32.3 | SLC39A10 | A | 0.081 | 0.375 | 0.297 | 0.591 |
| rs1992736 | 3p24.3 | TBC1D5 | A | 0.750 | 0.193 | 0.192 | 0.78 |
| rs10511297 | 3q13.13 | CD96 | A | 0.375 | 0.030 | 0.171 | 0.659 |
| rs2615485 | 4q22.1 | DSPP | A | 0.750 | 0.136 | 0.190 | 0.625 |
| rs2434316 | 5q14.1 | THBS4 | A | 0.750 | 0.257 | 0.118 | 0.743 |
| rs6878264 | 5q14.1 | THBS4 | B | 0.625 | 0.010 | 0.010 | 1.000 |
| rs7800141 | 7p15.3 | DNAH11 | B | 0.666 | 0.096 | 0.154 | 0.717 |
| rs4746259 | 10q22.2 | PPIAL4G | B | 0.750 | 0.015 | 0.062 | 0.794 |
| rs9300471 | 13q32.2 | FARP1 | A | 0.375 | 0.030 | 0.145 | 0.659 |
| rs1881344 | 16p13.2 | C16orf68 | A | 0.375 | 0.030 | 0.078 | 0.659 |
| rs2253429 | 20p13 | SIRPB1 | B | 0.375 | 0.030 | 0.0119 | 0.659 |
| rs2834681 | 21q22.12 | RUNX1 | B | 0.250 | 0 | 0.0623 | 0.859 |
In 6 “cases” with both intestinal metaplasia and dysplasia, compared with controls, 17 SNPs were above the genomic threshold and in HWE, of which rs1042194, located in exon 8 of the CYP2C19 gene, was the most significant associated SNP (χ2P value = 0.00536) (Table 3, Figure 1). The allele frequency for rs1042194 in cases vs controls was 0.6 vs 0.01.
| rs ID | Chromosome | Gene | Minor allele | Allele frequency | χ2 | HWE | |
| Cases | Control | P-value | P-value | ||||
| rs10493872 | 1p21.3 | ABCD3 | A | 0.375 | 0.062 | 0.00601 | 0.518 |
| rs10510792 | 3p14.3 | DNAH12 | A | 0.25 | 0.030 | 0.341 | 0.728 |
| rs2889259 | 4p14 | KIAA1239 | A | 0.375 | 0.015 | 0.0623 | 0.724 |
| rs6837437 | 4p14 | KIAA1239 | A | 0.375 | 0.015 | 0.0623 | 0.728 |
| rs10498879 | 6q13 | RIMS1 | A | 0.75 | 0.196 | 0.160 | 0.629 |
| rs4073894 | 7q22.1 | LHFPL3 | A | 0.375 | 0.015 | 0.0623 | 0.728 |
| rs10487929 | 7q35 | CNTNAP2 | B | 0.25 | 0.031 | 0.260 | 0.724 |
| rs10503727 | 8p21.2 | SLC25A37 | B | 0.75 | 0.183 | 0.151 | 0.908 |
| rs2251417 | 10q21.3 | ANXA2P1 | A | 0.375 | 0.045 | 0.579 | 0.591 |
| rs1042194 | 10q23.33 | CYP2C19 | B | 0.625 | 0.011 | 0.00536 | 0.972 |
| rs10506855 | 12q21.31 | CCDC59 | B | 0.375 | 0.046 | 0.0269 | 0.585 |
| rs10492652 | 13q22.1 | KLF12 | B | 0.25 | 0.015 | 0.0623 | 0.797 |
| rs1565946 | 14q23.1 | SLC35F4 | A | 0.625 | 0.156 | 0.0151 | 0.658 |
| rs10483683 | 14q23.1 | SLC35F4 | B | 0.625 | 0.181 | 0.314 | 0.964 |
| rs10483837 | 14q24.2 | RGS6 | B | 0.5 | 0.031 | 0.171 | 0.585 |
| rs245615 | 16q21 | CDH8 | A | 0.5 | 0.140 | 0.183 | 0.844 |
| rs2058879 | 19q13.32 | IGFL2 | A | 0.375 | 0.015 | 0.0623 | 0.728 |
Finally, in 2 “cases” with dysplasia only, compared with controls, 2 SNPs were above the genomic threshold and in HWE, of which rs10505799, located in chromosome 12p12.3 (MGST1 gene), was the most significant associated SNP (χ2P value = 0.006) (Table 4, Figure 1). The allele frequency for rs10505799 in cases vs controls was 0.5 vs 0.02.
| rs ID | Chromosome | Gene | Minor allele | Allele frequency | χ2 | HWE | |
| Cases | Control | P-value | P-value | ||||
| rs10505799 | 12p12.3 | MGST1 | B | 0.5 | 0.016 | 0.006 | 0.855 |
| rs10498391 | 14q21.2 | FSCB | B | 0.5 | 0 | 0.069 | 0.928 |
In this Malay population with an extremely low risk of H. pylori infection, gastric cancer and its precancerous lesions are very rare. However, in the current study of subjects susceptible to H. pylori infection and those who developed precancerous lesions, certain gene polymorphisms were found to be more commonly associated with precancerous lesions. Notwithstanding the low sample size, resulting from the extremely rare occurrence of gastric precancerous lesions in this population, the current study, with the use of the genome-wide association approach, allowed identification of genetic markers that can be tested in a larger cohort in the near future[24].
In the 50% of H. pylori-infected subjects with atrophic gastritis, the earliest lesion in the Correa cascade, rs9315542, located in chromosome 13 q13.3 (UFM1 gene), was the identified marker. A recently identified expressed protein, ubiquitin-fold modifier 1 or UFM1, is a member of a large family of ubiquitin-like proteins or Ubls[25]. Ubiquitin, a small protein, is associated with the process of “ubiquitination”, a target of proteins for degradation by the proteasome. At the moment, the exact cellular functions of proteins modified by UFM1 remain elusive. A recent report indicated that components of the UFM1 conjugation pathway are highly expressed in the beta cells of the pancreas and some other protein secretory tissues[26]. In the same report, UFM1 conjugate prevented endoplasmic reticulum (ER) stress-induced apoptosis. While UFM1 in gastric tissue has not been investigated, it is known that gastric mucosa secretes a number of peptides and hormones, including pepsinogen and ghrelin, whose levels are reduced in atrophic gastritis[27]. Speculatively, UFM1 may be a marker of the secretory status of the gastric mucosa, similar to pepsinogen, and remains to be tested and validated.
Complete-type or type I intestinal metaplasia, considered as the benign version compared with the incomplete-type, was present in 18.2% of H. pylori-infected subjects. In these subjects, rs6878264 located in intron 4 of the THBS4 gene, was the identified marker. THBS4 is a member of the THBS protein family, a glycoprotein in the extracellular matrix, which mediates cell-to-cell and cell-to-matrix interactions. Although the exact physiological functions of THBS4 are unknown, the published literature indicate that it promotes neurite outgrowth, stimulates proliferation of erythroid cells, skin fibroblasts and kidney epithelial cells, as well as myoblast adhesion and interaction with other extracellular matrix proteins[28-30]. Recently, THBS4 has been found to be associated with gastric adenocarcinomas especially of the diffuse type[31]. While H. pylori infection is associated with atrophic gastritis and intestinal metaplasia, it is not commonly associated with diffuse-type gastric adenocarcinoma[32]. Complete-type intestinal metaplasia represents a reparative process of the epithelium following H. pylori-induced gastritis, and in this context, THBS4 may act as an early proliferative marker but may have a more aggressive pro-oncogenic role in advanced stages. Again, this is speculative in the absence of any published studies, but it is a worthwhile marker for further studies.
Incomplete type intestinal metaplasia is more advanced compared with complete type, and therefore is more closely associated with dysplasia. In 22.7% of cases with both incomplete type intestinal metaplasia and dysplasia, rs1042194 located in exon 8 of CYP2C19 gene was the identified marker. Cytochrome (CYP) P450 2C19, one of the isoforms of the CYP enzyme (phase I detoxification enzyme), plays an important role in metabolism of drugs and also detoxification of potential carcinogens[33]. Several studies indicated that CYP2C19 gene polymorphism is associated with increased cancer susceptibility including hepatocellular carcinoma, and lung, esophageal and gastric cancer, especially in patients having a poor metabolizer (PM) genotype[34-36]. A study from Malaysia found that the PM genotype was uncommon among ethnic Malays (5.9%), compared with Chinese (19.1%) and Indians (10%)[37]. This may be one of the reasons for reduced susceptibility to gastric cancer and its precancerous lesions among ethnic Malays. The finding of CYP2C19 in incomplete type intestinal metaplasia and dysplasia in a group of Malay subjects susceptible to H. pylori infection is therefore important and merits further study.
Dysplasia, a histological stage with high risk of malignant transformation, was present in only 9.1% or 2/23 subjects infected with H. pylori. Compared with controls, rs10505799 located in chromosome 12p12.3 (MGST1 gene) was found to be the SNP marker associated with dysplasia. Microsomal glutathione S-transferase 1 (MGST1) is one of the glutathione S-transferase (GST) family of enzymes, and GSTs are phase II detoxification enzymes, which, similar to CYP enzymes, are involved in the detoxification of potential carcinogens[38,39]. Recently, MGST1 gene polymorphism was found to be involved in colorectal carcinogenesis in the Chinese population but there is no data as yet on gastric cancer[40]. However, since MGST1 and CYP2C19 are both carcinogen detoxification enzymes, with evidence supporting their involvement in gastrointestinal tract carcinogenesis, the role of MGST1 in gastric precancerous lesions is likely to be valid. The limited number of cases with dysplasia in the current study means that the results need to be interpreted cautiously, but the potential of MGST1 as a marker for dysplasia should not be disregarded.
There are a number of studies on gene polymorphisms associated with gastric precancerous lesions in high prevalence populations, but our study covered the entire spectrum of the Correa cascade in a population with an extremely low burden of gastric cancer and H. pylori infection. Development of gastric cancer is thought to involve multi-step carcinogenesis and follows a progressive pattern of pathological stages described by Correa. Our results indicated that, at different phases of the Correa cascade, different gene variants are manifest, but they follow a pattern of progression similar to their histological and clinical stages. During the stage of atrophic gastritis, UFM1 expression reflects the secretory status of epithelium. With early development of intestinal metaplasia, THBS4 acts as a proliferative marker but at more advanced stages, incomplete intestinal metaplasia and dysplasia involve polymorphisms of detoxification enzymes, CYP2C19 and MGST1. Based on the current study, it is possible that, in addition to histological staging, gene variant markers may also serve to identify different phases of progression of gastric cancer in the near future. Recently, epigenetic silencing of FOXD3 has been shown to be an early event in gastric carcinogenesis[41] and, together with genomic changes, it would allow a greater understanding of the pathogenesis of gastric cancer.
We acknowledge from the outset that the current study, based upon a genome-wide approach, was extremely limited in sample size, as gastric precancerous lesions are extremely rare among ethnic Malays from the north-eastern region of Peninsular Malaysia. In this respect, bioinformatics and statistical approaches were taken into consideration for a more reliable analysis of the data. To reduce false-positive results, a more stringent significance threshold of 3 × 10-7 was set for Manhattan plots in the current study. Only SNPs in HWE P-value > 0.5 were selected to reduce occasionality. In addition to being long-term residents within the studied region, cases and controls were similar in age, socio-cultural and economic backgrounds. The current study only identified SNPs associated with gastric precancerous lesions, and further validation studies are in progress to confirm their regulatory role in carcinogenesis.
In conclusion, we have shown that, compared with controls, susceptible ethnic Malays with H. pylori infection expressed different SNP markers at different spectrums of gastric precancerous lesions. These markers may allow efficient screening of precancerous lesions in larger cohorts of H. pylori-infected individuals.
Shuhua Xu is Max-Planck Independent Research Group Leader and member of CAS Youth Innovation Promotion Association. Shuhua Xu also gratefully acknowledges the support of the National Program for Top-notch Young Innovative Talents and the support of K.C. Wong Education Foundation, Hong Kong.
Gastric cancer and its precancerous lesions are exceptionally rare among ethnic Malays. Gene variants may be associated with precancerous lesions in Helicobacter pylori (H. pylori)-susceptible Malays.
In a case-control study, genome-wide association was performed to identify gene variants in the Malay population with a spectrum of H. pylori-associated gastric precancerous lesions.
Results indicated that at different phases of the Correa cascade, different gene variants were manifest, but they followed a pattern of progression similar to the histological and clinical stages.
It is possible that, in addition to histological staging, gene variant markers may also serve to identify different phases of progression of gastric cancer in the near future.
The genome-wide approach utilises microarray technology to identify thousands of single nucleotide polymorphisms (SNPs). Using novel bioinformatics and statistical approaches, the association between SNPs and the studied disease can be determined reliably.
Current study indicates that different gene variants exist that reflect different stages of progression during different spectrums of gastric carcinogenesis. These gene variants, appropriately confirmed in later studies, may be useful markers, in addition to histological staging, of gastric precancerous lesions.
P- Reviewer Kozarek RA S- Editor Wen LL L- Editor Cant MR E- Editor Zhang DN
| 1. | Correa P, Piazuelo MB. The gastric precancerous cascade. J Dig Dis. 2012;13:2-9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 355] [Cited by in RCA: 525] [Article Influence: 40.4] [Reference Citation Analysis (0)] |
| 2. | Kampschöer GH, Fujii A, Masuda Y. Gastric cancer detected by mass survey. Comparison between mass survey and outpatient detection. Scand J Gastroenterol. 1989;24:813-817. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 27] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 3. | Yamashita K, Sakuramoto S, Nemoto M, Shibata T, Mieno H, Katada N, Kikuchi S, Watanabe M. Trend in gastric cancer: 35 years of surgical experience in Japan. World J Gastroenterol. 2011;17:3390-3397. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 51] [Cited by in RCA: 55] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 4. | Bourke B. Will treatment of Helicobacter pylori infection in childhood alter the risk of developing gastric cancer? Can J Gastroenterol. 2005;19:409-411. [PubMed] |
| 5. | Weck MN, Brenner H. Prevalence of chronic atrophic gastritis in different parts of the world. Cancer Epidemiol Biomarkers Prev. 2006;15:1083-1094. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 105] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 6. | Derakhshan MH, Lee YY. Gastric cancer prevention through eradication of helicobacter pylori infection: feasibility and pitfalls. Arch Iran Med. 2012;15:662-663. [PubMed] |
| 7. | Lee YY, Mahendra Raj S, Graham DY. Helicobacter pylori Infection - A Boon or a Bane: Lessons from Studies in a Low-Prevalence Population. Helicobacter. 2013;Epub ahead of print. [PubMed] |
| 8. | Raj SM, Lee YY, Choo KE, Noorizan AM, Zulkifli A, Radzi M, Ang SC. Further observations in an area with an exceptionally low prevalence of Helicobacter pylori infection. Trans R Soc Trop Med Hyg. 2008;102:1163-1164. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 9. | Graham DY, Yamaoka Y, Malaty HM. Thoughts about populations with unexpected low prevalences of Helicobacter pylori infection. Trans R Soc Trop Med Hyg. 2007;101:849-851. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 20] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 10. | Lee YY, Ismail AW, Mustaffa N, Musa KI, Majid NA, Choo KE, Mahendra Raj S, Derakhshan MH, Malaty HM, Graham DY. Sociocultural and dietary practices among Malay subjects in the north-eastern region of Peninsular Malaysia: a region of low prevalence of Helicobacter pylori infection. Helicobacter. 2012;17:54-61. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 40] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 11. | Maran S, Lee YY, Xu S, Raj SM, Noorizan AM, Choo KE, Zilfalil BA, Graham DY. Toward understanding the low prevalence of Helicobacter pylori infection in Malays: Genetic variants differ among Helicobacter pylori negative ethnic Malays in the north-eastern region of Peninsular Malaysia and Han Chinese and South Indians. J Dig Dis. 2013;14:196-202. [RCA] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 9] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 12. | Rahim AA, Lee YY, Majid NA, Choo KE, Raj SM, Derakhshan MH, Graham DY. Helicobacter pylori infection among Aborigines (the Orang Asli) in the northeastern region of Peninsular Malaysia. Am J Trop Med Hyg. 2010;83:1119-1122. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 18] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 13. | Radzi M, Raj SM. The incidence of gastric cancer in Kelantan Malaysia is the lowest reported in the world (abstract). Med J Malaysia. 2000;55:13. |
| 14. | Moore MA, Manan AA, Chow KY, Cornain SF, Devi CR, Triningsih FX, Laudico A, Mapua CA, Mirasol-Lumague MR, Noorwati S. Cancer epidemiology and control in peninsular and island South-East Asia - past, present and future. Asian Pac J Cancer Prev. 2010;11 Suppl 2:81-98. [PubMed] |
| 15. | Yeh LY, Raj M, Hassan S, Aziz SA, Othman NH, Mutum SS, Naik VR. Chronic atrophic antral gastritis and risk of metaplasia and dysplasia in an area with low prevalence of Helicobacter pylori. Indian J Gastroenterol. 2009;28:49-52. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 18] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 16. | Maran S, Lee YY, Xu S, Rajab NS, Hasan N, Mustaffa N, Majid NA, Alwi Z. Deleted in Colorectal Cancer (DCC) Gene Polymorphism is Associated with H. pylori Infection among Susceptible Malays from the North-Eastern Region of Peninsular Malaysia. Hepatogastroenterology. 2012;60. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 17. | Hibi K, Sakata M, Sakuraba K, Kitamura YH, Shirahata A, Goto T, Mizukami H, Saito M, Ishibashi K, Kigawa G. Methylation of the DCC gene is lost in advanced gastric cancer. Anticancer Res. 2010;30:107-109. [PubMed] |
| 18. | Tytgat GN. The Sydney System: endoscopic division. Endoscopic appearances in gastritis/duodenitis. J Gastroenterol Hepatol. 1991;6:223-234. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 143] [Cited by in RCA: 148] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 19. | Rugge M, Correa P, Dixon MF, Fiocca R, Hattori T, Lechago J, Leandro G, Price AB, Sipponen P, Solcia E. Gastric mucosal atrophy: interobserver consistency using new criteria for classification and grading. Aliment Pharmacol Ther. 2002;16:1249-1259. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 246] [Cited by in RCA: 265] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 20. | Dixon MF, Genta RM, Yardley JH, Correa P. Classification and grading of gastritis. The updated Sydney System. International Workshop on the Histopathology of Gastritis, Houston 1994. Am J Surg Pathol. 1996;20:1161-1181. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3221] [Cited by in RCA: 3557] [Article Influence: 122.7] [Reference Citation Analysis (3)] |
| 21. | Antoniolli DA. Gastric carcinoma and its precursors. Gastrointestinal Pathology. United States and Canadian Academy of Pathology Monograph in Pathology No. 31. Baltimore: Williams and Wilkins 1990; 144. |
| 22. | Ming SC, Bajtai A, Correa P, Elster K, Jarvi OH, Munoz N, Nagayo T, Stemmerman GN. Gastric dysplasia. Significance and pathologic criteria. Cancer. 1984;54:1794-1801. [PubMed] |
| 23. | Available from: http://www.affymetrix.com/. |
| 24. | Maran S, Lee YY, Zilfalil BA, Noorizan AM. A new paradigm in medicine: Genome wide association studies. Bulletin of the Genetics Society of Malaysia. 2011;18:3-6. |
| 25. | Hochstrasser M. Origin and function of ubiquitin-like proteins. Nature. 2009;458:422-429. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 642] [Cited by in RCA: 625] [Article Influence: 39.1] [Reference Citation Analysis (0)] |
| 26. | Lemaire K, Moura RF, Granvik M, Igoillo-Esteve M, Hohmeier HE, Hendrickx N, Newgard CB, Waelkens E, Cnop M, Schuit F. Ubiquitin fold modifier 1 (UFM1) and its target UFBP1 protect pancreatic beta cells from ER stress-induced apoptosis. PLoS One. 2011;6:e18517. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 126] [Cited by in RCA: 156] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 27. | Agréus L, Kuipers EJ, Kupcinskas L, Malfertheiner P, Di Mario F, Leja M, Mahachai V, Yaron N, van Oijen M, Perez Perez G, Rugge M, Ronkainen J, Salaspuro M, Sipponen P, Sugano K, Sung J. Rationale in diagnosis and screening of atrophic gastritis with stomach-specific plasma biomarkers. Scand J Gastroenterol. 2012;47:136-147. [PubMed] |
| 28. | Arber S, Caroni P. Thrombospondin-4, an extracellular matrix protein expressed in the developing and adult nervous system promotes neurite outgrowth. J Cell Biol. 1995;131:1083-1094. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 115] [Cited by in RCA: 123] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 29. | Congote LF, Difalco MR, Gibbs BF. The C-terminal peptide of thrombospondin-4 stimulates erythroid cell proliferation. Biochem Biophys Res Commun. 2004;324:673-678. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 16] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 30. | Narouz-Ott L, Maurer P, Nitsche DP, Smyth N, Paulsson M. Thrombospondin-4 binds specifically to both collagenous and non-collagenous extracellular matrix proteins via its C-terminal domains. J Biol Chem. 2000;275:37110-37117. [PubMed] |
| 31. | Förster S, Gretschel S, Jöns T, Yashiro M, Kemmner W. THBS4, a novel stromal molecule of diffuse-type gastric adenocarcinomas, identified by transcriptome-wide expression profiling. Mod Pathol. 2011;24:1390-1403. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 85] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 32. | Adachi Y, Yasuda K, Inomata M, Sato K, Shiraishi N, Kitano S. Pathology and prognosis of gastric carcinoma: well versus poorly differentiated type. Cancer. 2000;89:1418-1424. [PubMed] |
| 33. | Agundez JA. Cytochrome P450 gene polymorphism and cancer. Curr Drug Metab. 2004;5:211-224. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 208] [Cited by in RCA: 217] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 34. | Sugimoto M, Furuta T, Shirai N, Nakamura A, Kajimura M, Sugimura H, Hishida A, Ishizaki T. Poor metabolizer genotype status of CYP2C19 is a risk factor for developing gastric cancer in Japanese patients with Helicobacter pylori infection. Aliment Pharmacol Ther. 2005;22:1033-1040. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 30] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 35. | Shi WX, Chen SQ. Frequencies of poor metabolizers of cytochrome P450 2C19 in esophagus cancer, stomach cancer, lung cancer and bladder cancer in Chinese population. World J Gastroenterol. 2004;10:1961-1963. [PubMed] |
| 36. | Chau TK, Marakami S, Kawai B, Nasu K, Kubota T, Ohnishi A. Genotype analysis of the CYP2C19 gene in HCV-seropositive patients with cirrhosis and hepatocellular carcinoma. Life Sci. 2000;67:1719-1724. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 17] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 37. | Pang YS, Wong LP, Lee TC, Mustafa AM, Mohamed Z, Lang CC. Genetic polymorphism of cytochrome P450 2C19 in healthy Malaysian subjects. Br J Clin Pharmacol. 2004;58:332-335. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 41] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 38. | Andersson C, Mosialou E, Weinander R, Morgenstern R. Enzymology of microsomal glutathione S-transferase. Adv Pharmacol. 1994;27:19-35. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 65] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 39. | Kelner MJ, Bagnell RD, Montoya MA, Estes LA, Forsberg L, Morgenstern R. Structural organization of the microsomal glutathione S-transferase gene (MGST1) on chromosome 12p13.1-13.2. Identification of the correct promoter region and demonstration of transcriptional regulation in response to oxidative stress. J Biol Chem. 2000;275:13000-13006. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 64] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 40. | Zhang H, Liao LH, Liu SM, Lau KW, Lai AK, Zhang JH, Wang Q, Chen XQ, Wei W, Liu H. Microsomal glutathione S-transferase gene polymorphisms and colorectal cancer risk in a Han Chinese population. Int J Colorectal Dis. 2007;22:1185-1194. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 18] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 41. | Cheng AS, Li MS, Kang W, Cheng VY, Chou JL, Lau SS, Go MY, Lee CC, Ling TK, Ng EK. Helicobacter pylori causes epigenetic dysregulation of FOXD3 to promote gastric carcinogenesis. Gastroenterology. 2013;144:122-133.e9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 112] [Article Influence: 9.3] [Reference Citation Analysis (0)] |