Published online Mar 14, 2009. doi: 10.3748/wjg.15.1201
Revised: January 14, 2009
Accepted: January 21, 2009
Published online: March 14, 2009
Alcoholic patients have a high incidence of hepatitis C virus (HCV) infection. Alcohol consumption enhances the severity of the HCV disease course and worsens the outcome of chronic hepatitis C. The accumulation of virally infected cells in the liver is related to the HCV-induced inability of the immune system to recognize infected cells and to develop the immune responses. This review covers the effects of HCV proteins and ethanol on major histocompatibility complex (MHC) class I- and class II-restricted antigen presentation. Here, we discuss the liver which functions as an immune privilege organ; factors, which affect cleavage and loading of antigenic peptides onto MHC class I and class II in hepatocytes and dendritic cells, and the modulating effects of ethanol and HCV on antigen presentation by liver cells. Altered antigen presentation in the liver limits the ability of the immune system to clear HCV and infected cells and contributes to disease progression. HCV by itself affects dendritic cell function, switching their cytokine profile to the suppressive phenotype of interleukin-10 (IL-10) and transforming growth factor beta (TGFβ) predominance, preventing cell maturation and allostimulation capacity. The synergistic action of ethanol with HCV results in the suppression of MHC class II-restricted antigen presentation. In addition, ethanol metabolism and HCV proteins reduce proteasome function and interferon signaling, thereby suppressing the generation of peptides for MHC class I-restricted antigen presentation. Collectively, ethanol exposure further impairs antigen presentation in HCV-infected liver cells, which may provide a partial explanation for exacerbations and the poor outcome of HCV infection in alcoholics.
- Citation: Osna NA. Hepatitis C virus and ethanol alter antigen presentation in liver cells. World J Gastroenterol 2009; 15(10): 1201-1208
- URL: https://www.wjgnet.com/1007-9327/full/v15/i10/1201.htm
- DOI: https://dx.doi.org/10.3748/wjg.15.1201
In non-cytolytic viral infections, the immune system (and mainly T-lymphocytes) is necessary to clear the infected cells. The most specialized and effective homeostatic control is provided by cytotoxic T lymphocytes (CTLs) (CD8+ T-lymphocytes). For clonal expansion, CTLs need activating “help” from CD4+ T-lymphocytes, which, in turn, require recognition of viral antigens on professional antigen-presenting cells (APC), including dendritic cells (DC), macrophages and B-lymphocytes.
Recognition of antigens by T-lymphocytes depends upon the expression of the major histocompatibility complex (MHC) on the surface of APC. MHC molecules involved in antigen presentation include MHC class I, MHC class II and MHC-like CD1 molecules[1]. MHC class I molecules are expressed on the nuclear cells and load protease-generated peptides to CD8+ T-lymphocytes. MHC class II molecules are expressed on the surface of specialized APC and load peptides generated in the endocytic compartment to CD4+ T cells. CD1 molecules bind acyl chains, therefore allowing T cells to recognize fatty acids, glycolipids and lipopeptide antigens (self or foreign)[2].
Hepatitis C virus (HCV) is an example of an intracellularly persistent virus, which targets liver cells and is eliminated by immune cells[34]. In acute HCV infection, antigen presentation plays a pivotal role in the activation of immune response, while HCV protein-induced defects in antigen presentation lead to insufficient activation of the immune system, which controls the killing of infected hepatocytes[5]. Accumulation and persistence of virally infected cells provide a basis for the chronic course of HCV infection. The reduced clearance of virus and increased viral load usually observed in immunodeficient and in alcohol-consuming patients are associated with a high risk of hepatocarcinoma development[67].
Liver is considered an immunological organ[8]. Liver cells are involved in the clearance of foreign antigens, which come from the gastrointestinal tract. The physiological role of the liver is to face antigenic flow from the gastrointestinal tract and to respond to this intervention by activating the immune response. To escape immune activation, liver cells use a mechanism of immune tolerance, which prevents unnecessary immune-mediated damage of hepatocytes.
Some liver cells, such as liver dendritic, Kupffer, endothelial and stellate cells serve as antigen presenting cells[9]. Kupffer cells (KC) are the resident macrophage population, which act as incompetent antigen-presenters[10]. They produce IL-10, which down-regulates CD4+ T-cell activation[1112]. Reactive oxygen species (ROS) production is an essential trigger of antigen presentation in KC, accompanied by induction of MHC class II and co-stimulatory molecule expression[13]. In contrast to KC, liver endothelial cells (LSEC) are known as potent APC that present antigens in a MHC class II-restricted manner to CD4+ T-cells[9]. The efficacy of their antigen presentation is comparable to DC[8]; however, IL-10 and TGFβ secreted by KC and LSEC as well as lipopolysaccharide (LPS), which comes with blood from the gastrointestinal tract, can down-regulate their antigen-presenting properties[1415]. Both KC and LSEC express MHC antigens, costimulatory and adhesion molecules and produce IL-1 and IFNγ, suggesting that these are relatively mature cells[16]. In contrast, liver DC are immature cells, which express MHC antigens on their surface, but express a relatively small amount of co-stimulatory molecules, which does not allow them to efficiently stimulate naïve T-cells[17]. Hepatic DC play an important role in the induction and regulation of immune responses, by interacting with CD4+, CD8+ lymphocytes and natural killers (NK) cells. These cells typically reside only around portal triads and like other DC, they capture, process and transport antigens to regional lymphoid tissues[16]. Two subpopulations of DC, myeloid and plasmocitoid cells are found in liver at a low density. Exposure of progenitor DC to IL-10 and TGFβ generates suppressive and tolerogenic effects[18]. In the liver, they are resistant to DC maturation stimuli, such as IFNγ and TNFα, while matrix proteins, such as collagen type 1, induces maturation[19]. However, even being tolerogenic in the liver, hepatic DC are the main professional liver APC, which can further migrate to lymphoid tissue to undergo maturation[20]. In addition to other liver cells, stellate cells have also recently been characterized as having antigen presentation properties[21].
Almost 60% of liver cells are hepatocytes. Due to specific liver architecture, hepatocytes are exposed to a mixture of portal venous and hepatic arterial blood. Liver parenchyma is actively involved in immune response by expressing a variety of receptors. Because liver is a site of apoptotic CD8+ T-lymphocyte accumulation[22], it raises the question whether liver cells, including hepatocytes, induce apoptosis in activated CD8+ T-lymphocytes, thereby promoting a tolerogenic response or they specifically attract apoptotic T-lymphocytes without causing T-lymphocyte death. Resting hepatocytes act as APC for MHC class I-restricted T-cells, while MHC class II and CD1 are not constitutively expressed on hepatocytes[23]. However, the role of hepatocytes as potent APC activating intrahepatic lymphocytes is contradictory. While some studies indicate that activation of naïve CD8+ T-lymphocytes does not happen in case the primary activation occurs within the liver, other studies suggest that the presentation of antigens in the liver showed no immunosuppressive effect on the activation of CD8+ T-cells and that hepatocytes are an excellent priming site for naïve CD8+ T-cells[24–26].
Proteolysis is the first step in the antigen processing and presentation pathway. Peptide products are transported by transporters associated with antigen processing (TAP) to endoplasmic reticulum (ER), where they assemble in a trimolecular complex with β2-microglobulin and the heavy chain of MHC. Assembly is facilitated by TAP and a number of chaperones and allows class I molecule to bind the peptide to achieve optimal MHC class I loading. These complexes are transported to the plasma membrane, where they interact with the T-cell receptor (TCR) of a T-lymphocyte. TAP has sequence preference and peptide-size limitation, which is matched to MHC class I. To be presented in a context of MHC class I, antigenic proteins undergo the processing to peptides and then the peptides are further cleaved, to fit into the MHC class I groove. Certain proteases participate in the cleavage of peptides for antigen presentation, and the major protease is proteasome.
For activation, CD8+ cells require presentation of MHC class I-peptide complexes on professional APC cells. Peptides can be generated from viral proteins sensitized by infected APC or from proteins originally synthesized by other infected “donor” cells (cross-priming, or cross presentation). Cross-priming can be carried by molecular chaperones and come from apoptotic cells[27]. In addition, it is suggested that these peptides are captured by the lysosomal compartment, with further trafficking to cytosol followed by proteasomal processing for MHC class I loading[28]. However, other authors question the role of proteasomal processing in the cross-presentation mechanism[29].
Degradation of intracellular proteins into oligopeptides for antigen presentation is catalyzed by the proteasome[30]. This multicatalytic enzyme exists as a 26S particle, which is ATP-dependent and recognizes ubiquitylated polypeptides. Another form, the 20S proteasome, can degrade substrate proteins in an ubiquitin-independent manner. Both the 26S and 20S proteasome particles degrade antigenic proteins that may later be presented as peptides, depending on the properties of the protein. The chymotrypsin-like and the trypsin-like activities of the proteasome are related to antigen presentation due to their ability to cleave peptide bonds after hydrophobic and basic amino acids, respectively[30]. In cells treated with IFNγ, proteasome particles acquire two novel MHC-encoded low molecular weight subunits, known as LMP-2 and LMP-7, and a third subunit, MECL-1, encoded by genes outside the MHC locus[31]. The acquisition of these three subunits converts the constitutive proteasome to the immunoproteasome and alters its peptidase activities for the generation of uniform sized 8-10-mer peptides[32]. The importance of immunoproteasome induction for antigen presentation has been confirmed using LMP-2 and LMP-7-knockout mice, which demonstrated impaired antigen presentation, reduced expression of MHC class I and poor CTL response to viral epitopes[3334]. The immunoproteasome is responsible for cleavage of peptides at their C-termini[35]. However, some antigenic peptides have N-terminal extensions that are unable to bind to the MHC class I groove. Aminopeptidases, namely, cytosolic leucine aminopeptidase (LAP), finally trims the epitopes to 8-9 amino acid residues necessary to form the complexes with MHC class I[36]. This enzyme is also IFNγ-inducible[37]. Trimmed peptides are transported to the endoplasmic reticulum (ER) by TAP. In the ER lumen, the peptide can be further trimmed by downstream endoplasmic reticulum aminopeptidase I (ERAP1)[38] to generate a stable complex with MHC class I heavy and light chains (β2-microglobulin). These complexes, with the help of molecular chaperones, traffic to the cell surface to be recognized by CD8+ T-cells[3940]. The immune system relies heavily on intracellular protein degradation to recognize a complex of surface MHC class I molecules with short peptide fragments of eight to nine amino acids[41]. Induction of CTLs is programmed by the spectrum of presented MHC class I-peptide complexes on the cell surface[42].
The effects of HCV proteins on MHC class I-restricted antigen presentation are not widely studied and most of the findings are related to MHC class II-restricted antigen presentation. In HCV infected patients, associations between the human MHC and sustained virological response have been found, indicating that the various immunogenetic backgrounds of chronic hepatitis C patients are related to the differences in the course of the disease[43].
Processing of CTL epitopes in HCV infected cells may be altered due to mutations generated by HCV, which interfere with the ability of the peptide to be cleaved by the proteasome[44]. Nevertheless, interferon alpha-mediated induction of immunoproteasome or aminopeptidase expression has been shown as a necessary step for CD8+ T-cell activation in acute HCV infected chimpanzees[4546]. In addition to HCV-restricted mutations of viral proteins, proteasome activity is also modulated by HCV. Thus, proteasome activity in the nuclear compartment is up-regulated by PA28γ activator which forms a complex with HCV core protein[47], while non-structural HCV protein, NS3, interacts with LMP-7, an immunoproteasome subunit, and reduces its activity, potentially providing a negative effect on the generation of peptides for antigen presentation[48]. Our previous studies on HCV core protein expressing Huh7 cells, which also express CYP2E1, revealed that core protein slightly enhances 20S proteasome activity, by 20S proteasome-core protein direct interactions and by induction of low CYP2E1-dependent oxidative stress[49]. However, this proteasome activation is reversed after ethanol exposure, where ethanol treatment considerably reduces proteasome function due to induction of high oxidative stress[49]. Indeed, the dependence of proteasome on the level of oxidative stress has been previously demonstrated[5051], and ethanol is known to suppress proteasome function in liver cells[52–55]. Ethanol-elicited suppression of proteasome activity in the liver ultimately results in reduced generation of antigenic peptides[56] and reduced MHC class I-restricted antigen presentation on hepatocytes (Osna et al, Hepatology, in press). Presentation of peptide-MHC class I complexes on virally infected hepatocytes (HCV, HBV infections, etc) as target cells is crucial for CTLs because when clonal expansion of CTLs is established, the next important restriction for elimination of infected cell is the availability of peptide-MHC class I complexes on the surface of target cells (hepatocytes), which are recognized by CTLs. Thus, even if CD8+ T-cells are activated by the presentation of HCV peptides on professional APC, the recognition of the peptide-MHC class I complexes on hepatocytes (which are target cells for CTLs) may be limited under ethanol-induced oxidative stress.
Loading of MHC class II molecules with antigenic peptides requires the involvement of specialized APC, such as DC, B-lymphocytes and macrophages. Phagocytosis is the main way to capture antigenic proteins by DC or macrophages, while B-lymphocytes use the antigen specific B-cell receptors[57]. Antigens are digested into peptides (which are longer than those required for MHC class I-restricted antigen presentation) by proteases at endosome compartments. Cathepsins, intracellular acidic proteases, play a pivotal role in the processing of internalized antigens into class II-presentable T cell epitopes[58]. These peptidases play a dual role by generating the peptides or by destroying them. The coupling of peptides with MHC class II is controlled by chaperones and takes place in late endocytic vesicules[44].
When pathogen-associated molecular patterns (PAMPs) on macrophages are recognized by innate immunity receptors (such as Toll-like (TLR)-, mannose-, Fc- and complement- receptors), they induce production of pro-inflammatory cytokines, namely, IFNγ and colony-stimulating factor (GM-CSF). These cytokines increase the expression of MHC class II and co-stimulatory molecules in the cells. Chronic TLR signaling induced by LPS may impair MHC class II-restricted antigen presentation[59]. A large number of class II-bound proteins are derived from cytosolic proteins. Because the autophagy inhibitor has been shown to block the presentation of cytosolic peptides by MHC class II molecules, some authors discuss the role of autophagy in delivery of these cytosolic peptides to the endocytic route[60].
After immature DC engulf pathogens, they undergo biochemical changes, such as secretion of TNFα, IL-6, IL-12, IL-10 and IFNα and expression of co-stimulatory molecules, including CD80, CD86 and CD40. These cells traffic to the nearest lymph node for presentation to T-lymphocytes. However, only mature DC can efficiently prime naïve T-lymphocytes, because only these cells generate MHC class II molecules, by redistribution of class II molecules and cathepsins to peptide-loading compartments and by enhancement of lysosomal acidification[61]. It is still unclear how the peptide-MHC class II complex is recruited to the cell surface from the endocytic compartment in DC. Human DC are subdivided into two big categories: myeloid DC and plasmocytoid DC. In addition to phenotypic differences, they express different TLRs and secrete a different spectrum of cytokines. Thus, myeloid DC express TLR3 and upon stimulation, produce IL-12p70, thereby promoting Th1 response, while plasmocytoid DC express TLR7 and TLR9 and secrete IFNα[62]. Therefore, myeloid DC cells are considered classic antigen presenters, while plasmocytoid DC have a limited ability to capture, process and load antigen onto MHC molecules[63]. Activation of DC is positively regulated by IFNγ and additional CD40 ligation; anti-inflammatory cytokines, such as IL-10, inhibit both IL-12 and IFNα expression, which results in pathogen survival[64].
HCV infection affects DC function in many ways. Firstly, there is a decrease in the frequency of peripheral myeloid or plasmocytoid DC in chronically infected patients[65]. This may be, in part, related to an accumulation of intrahepatic DC in HCV infection, due to altered DC trafficking[66]. However, another study suggested that HCV directly targets mature cells because some HCV proteins (core, NS3 and NS5) induce apoptosis in DC[67]. Secondly, myeloid DC from chronic HCV patients have a decreased capacity to stimulate allogenic T-lymphocytes[68]. Furthermore, HCV core and E1 genes introduced in DC obtained from uninfected donors, lower their capacity to stimulate allogenic T-cell response[69]. In addition, core, NS3 and NS4 viral proteins were reported to influence the differentiation of DC from monocytes, due to IL-10 secretion[70]. The function of myeloid DC obtained from HCV patients is also decreased due to a reduced level of co-stimulatory molecules and down-regulated HLA-DR expression[568]. In addition, in chronic HCV patients, the production of IL-12 by DC is suppressed by HCV proteins, but can be restored by successful antiviral therapy[71]. Thirdly, the impairment of plasmocytoid DC function has been reported in HCV patients and reduced IFNα production upon DC stimulation with TLR ligands has also been observed[72]. This decrease in IFNα production can be attributed to monocyte-derived TNFα and IL-10[73]. Interestingly, the other studies did not demonstrate defective abilities in plasmocytoid DC to initiate immune response[74], thereby questioning the altered presentation of HCV proteins by these cells to CD4+ cells.
The proposed mechanisms of ethanol-HCV interactions were summarized elsewhere[75]. Ethanol drastically enhances the negative effects of HCV on DC function. A possible reason for this is that alcohol stimulates HCV expression at HCV RNA and proteins levels, which is, in part, related to up-regulation of the Cox2 pathway[76]. Chronic ethanol feeding suppresses CD4+ T cell proliferation in response to antigens as well as CTL activity after DNA-based vaccine immunization with NS5 and HCV-expressing plasmids[77–79]. This CD4+T-cell response was restored by co-administration of IL-2–expressing plasmid, while the restoration of CTL activity required administration of GM-CSF[7778]. Alcohol doubles the effects of HCV by decreasing expression of co-stimulatory molecule, B7, and IL-12 production, increasing IL-10 production and lowering allostimulatory capacity[80–82]. Furthermore, human studies demonstrated that ethanol affects the allostimulatory activity of resting and activated DC generated from peripheral blood mononuclear cells in the presence of cytokines[81], indicating that HCV infection impairs DC function, which is further exacerbated by ethanol. Recent studies performed on alcohol-fed mice, immunized with HCV DNA-based vaccine that induces immune response to NS5 HCV protein, demonstrated that ethanol exposure reduced the number of splenic DCs without altering endocytosis capacity. It also reduced the lymphoid DC population, as well as expression of co-stimulatory molecules, CD40 and CD86, altered cytokine profile (enhanced production of IL-1 and IL-10 and decreased secretion of IL-12, IFNγ and IL-6)[83]. Finally, CTL response to NS5 was shown to be impaired, but was corrected by syngeneic transfer of control DC[83].
Antigen presentation is so tightly regulated by IFNs that it should be analyzed in the context of IFN signaling. The reported effects of HCV proteins on IFN signaling are quite controversial. Most studies revealed down-regulation of STAT1 phosphorylation in response to IFNα[84–86] attributed to SOCS3 activation by HCV core or NS5A proteins. In addition to that, NS3 protein plays a role in dissociation of the adaptor molecule, MAVS that disrupts the early activation of IFNβ signaling by the virus[8788]. Subsequently, the disruption of STAT1 phosphorylation by the NS3/NS4 complex was documented[89]. In some studies, suppressed STAT1 phosphorylation was linked to either the ability of the core protein to specifically interact with the SH2 domain, preventing STAT1 hetero- or homodimerization[90] or to direct targeting of STAT1 with core protein and subjecting activated STAT1 to degradation by the proteasome[91]. The latter hypothesis is consistent with our recent findings that core protein activates proteasome function[49]. Core protein also inhibits the IFN-induced nuclear import of STAT1 and STAT2 and transcription of antiviral genes[9293]. HCV proteins interfere with IFNα signaling related to the attachment of activated STAT1 to DNA. Methylation on arginine residues, which prevents the formation in STAT1-PIAS1 complexes, is altered in chronic hepatitis C patients[9495]. Protective effects of the methylation regulators, S-adenosyl methionine and betaine, were shown to correct HCV-induced inhibition of IFNα signaling in vitro[96]. Interestingly, in other studies, core protein has been shown to activate interferon stimulated response element (ISRE) and gamma activated site (GAS) sequence promoters in the presence or absence of IFNα and γ and to stimulate INOS promoter, INOS protein induction as well as activation of IFN-inducible 2’5’-oligoadenylate synthetase (2’5’-OAS)[9798]. Less is known about the interference of HCV with IFNγ-induced signaling, which is up-regulated by core protein, due to enhanced IFNγR2 expression[99]. In a cell culture system, prolonged treatment with IFNγ attenuated IFNα-signaling[100]. Acute ethanol treatment of hepatoma cells inhibited anti-viral actions of IFN against HCV replicon, induced serine STAT1 phosphorylation, but blocked tyrosine phosphorylation[101]. Recently, it has been shown that alcohol metabolism induces increased replication of HCV and attenuates the antiviral effects of IFN[102]. However, the number of studies regarding the combined effects of ethanol and HCV proteins on IFN signaling is very limited.
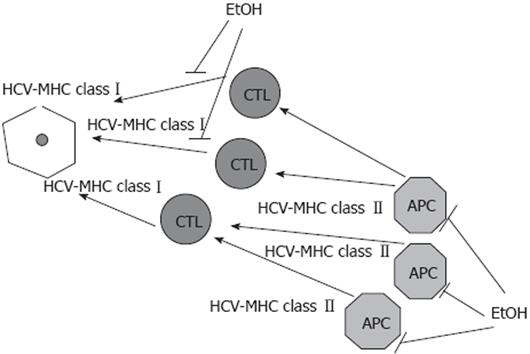
HCV alters antigen-presentation capacities of professional APC, thereby contributing to the persistence of virally infected cells. The most prominent effects were observed on MHC class II-restricted antigen presentation and DC functions. Ethanol potentiates the effects of HCV, providing further suppression of both MHC class I- and class II-restricted antigen presentation on DC, hepatocytes and others hepatic APC, which diminish CTL response. IFNs which support antigen presentation by activating APC and peptidases lose this property in the presence of HCV and alcohol, because HCV and alcohol, separately or in combination, reduce IFN signaling. This, in part, explains why alcohol consumption exacerbates the course, worsens the outcome and reduces the responsiveness to interferon alpha treatment in chronic HCV infection. The effects of alcohol (ethanol) on HCV-restricted antigen presentation are summarized in Figure 1.
| 1. | van den Elsen PJ, Holling TM, Kuipers HF, van der Stoep N. Transcriptional regulation of antigen presentation. Curr Opin Immunol. 2004;16:67-75. [Cited in This Article: ] |
| 2. | Sugita M, Cernadas M, Brenner MB. New insights into pathways for CD1-mediated antigen presentation. Curr Opin Immunol. 2004;16:90-95. [Cited in This Article: ] |
| 3. | Ando K, Hiroishi K, Kaneko T, Moriyama T, Muto Y, Kayagaki N, Yagita H, Okumura K, Imawari M. Perforin, Fas/Fas ligand, and TNF-alpha pathways as specific and bystander killing mechanisms of hepatitis C virus-specific human CTL. J Immunol. 1997;158:5283-5291. [Cited in This Article: ] |
| 4. | Willberg C, Barnes E, Klenerman P. HCV immunology--death and the maiden T cell. Cell Death Differ. 2003;10 Suppl 1:S39-S47. [Cited in This Article: ] |
| 5. | Kanto T, Hayashi N. Immunopathogenesis of hepatitis C virus infection: multifaceted strategies subverting innate and adaptive immunity. Intern Med. 2006;45:183-191. [Cited in This Article: ] |
| 6. | Yu MC, Yuan JM, Lu SC. Alcohol, cofactors and the genetics of hepatocellular carcinoma. J Gastroenterol Hepatol. 2008;23 Suppl 1:S92-S97. [Cited in This Article: ] |
| 7. | Koike K, Tsutsumi T, Miyoshi H, Shinzawa S, Shintani Y, Fujie H, Yotsuyanagi H, Moriya K. Molecular basis for the synergy between alcohol and hepatitis C virus in hepatocarcinogenesis. J Gastroenterol Hepatol. 2008;23 Suppl 1:S87-S91. [Cited in This Article: ] |
| 8. | Racanelli V, Rehermann B. The liver as an immunological organ. Hepatology. 2006;43:S54-S62. [Cited in This Article: ] |
| 9. | Knolle PA, Gerken G. Local control of the immune response in the liver. Immunol Rev. 2000;174:21-34. [Cited in This Article: ] |
| 10. | You Q, Cheng L, Kedl RM, Ju C. Mechanism of T cell tolerance induction by murine hepatic Kupffer cells. Hepatology. 2008;48:978-990. [Cited in This Article: ] |
| 11. | Knolle PA, Uhrig A, Hegenbarth S, Loser E, Schmitt E, Gerken G, Lohse AW. IL-10 down-regulates T cell activation by antigen-presenting liver sinusoidal endothelial cells through decreased antigen uptake via the mannose receptor and lowered surface expression of accessory molecules. Clin Exp Immunol. 1998;114:427-433. [Cited in This Article: ] |
| 12. | Selmi C, Mackay IR, Gershwin ME. The immunological milieu of the liver. Semin Liver Dis. 2007;27:129-139. [Cited in This Article: ] |
| 13. | Maemura K, Zheng Q, Wada T, Ozaki M, Takao S, Aikou T, Bulkley GB, Klein AS, Sun Z. Reactive oxygen species are essential mediators in antigen presentation by Kupffer cells. Immunol Cell Biol. 2005;83:336-343. [Cited in This Article: ] |
| 14. | Steinbrink K, Wolfl M, Jonuleit H, Knop J, Enk AH. Induction of tolerance by IL-10-treated dendritic cells. J Immunol. 1997;159:4772-4780. [Cited in This Article: ] |
| 15. | Knolle P, Lohr H, Treichel U, Dienes HP, Lohse A, Schlaack J, Gerken G. Parenchymal and nonparenchymal liver cells and their interaction in the local immune response. Z Gastroenterol. 1995;33:613-620. [Cited in This Article: ] |
| 16. | Lau AH, Thomson AW. Dendritic cells and immune regulation in the liver. Gut. 2003;52:307-314. [Cited in This Article: ] |
| 17. | Abe M, Akbar SM, Horiike N, Onji M. Induction of cytokine production and proliferation of memory lymphocytes by murine liver dendritic cell progenitors: role of these progenitors as immunogenic resident antigen-presenting cells in the liver. J Hepatol. 2001;34:61-67. [Cited in This Article: ] |
| 18. | Thomson AW, Lu L. Are dendritic cells the key to liver transplant tolerance? Immunol Today. 1999;20:27-32. [Cited in This Article: ] |
| 19. | Drakes ML, Lu L, McKenna HJ, Thomson AW. The influence of collagen, fibronectin, and laminin on the maturation of dendritic cell progenitors propagated from normal or Flt3-ligand-treated mouse liver. Adv Exp Med Biol. 1997;417:115-120. [Cited in This Article: ] |
| 20. | Sumpter TL, Abe M, Tokita D, Thomson AW. Dendritic cells, the liver, and transplantation. Hepatology. 2007;46:2021-2031. [Cited in This Article: ] |
| 21. | Unanue ER. Ito cells, stellate cells, and myofibroblasts: new actors in antigen presentation. Immunity. 2007;26:9-10. [Cited in This Article: ] |
| 22. | Crispe IN, Dao T, Klugewitz K, Mehal WZ, Metz DP. The liver as a site of T-cell apoptosis: graveyard, or killing field? Immunol Rev. 2000;174:47-62. [Cited in This Article: ] |
| 23. | Markiewski MM, DeAngelis RA, Lambris JD. Liver inflammation and regeneration: two distinct biological phenomena or parallel pathophysiologic processes? Mol Immunol. 2006;43:45-56. [Cited in This Article: ] |
| 24. | Bowen DG, McCaughan GW, Bertolino P. Intrahepatic immunity: a tale of two sites? Trends Immunol. 2005;26:512-517. [Cited in This Article: ] |
| 25. | Wuensch SA, Pierce RH, Crispe IN. Local intrahepatic CD8+ T cell activation by a non-self-antigen results in full functional differentiation. J Immunol. 2006;177:1689-1697. [Cited in This Article: ] |
| 26. | Klein I, Gassel HJ, Crispe IN. Cytotoxic T-cell response following mouse liver transplantation is independent of the initial site of T-cell priming. Transplant Proc. 2006;38:3241-3243. [Cited in This Article: ] |
| 27. | Brusa D, Garetto S, Chiorino G, Scatolini M, Migliore E, Camussi G, Matera L. Post-apoptotic tumors are more palatable to dendritic cells and enhance their antigen cross-presentation activity. Vaccine. 2008;26:6422-6432. [Cited in This Article: ] |
| 28. | Basha G, Lizee G, Reinicke AT, Seipp RP, Omilusik KD, Jefferies WA. MHC class I endosomal and lysosomal trafficking coincides with exogenous antigen loading in dendritic cells. PLoS ONE. 2008;3:e3247. [Cited in This Article: ] |
| 29. | Norbury CC, Basta S, Donohue KB, Tscharke DC, Princiotta MF, Berglund P, Gibbs J, Bennink JR, Yewdell JW. CD8+ T cell cross-priming via transfer of proteasome substrates. Science. 2004;304:1318-1321. [Cited in This Article: ] |
| 30. | Goldberg AL, Cascio P, Saric T, Rock KL. The importance of the proteasome and subsequent proteolytic steps in the generation of antigenic peptides. Mol Immunol. 2002;39:147-164. [Cited in This Article: ] |
| 31. | Baumeister W, Walz J, Zuhl F, Seemuller E. The proteasome: paradigm of a self-compartmentalizing protease. Cell. 1998;92:367-380. [Cited in This Article: ] |
| 32. | Hisamatsu H, Shimbara N, Saito Y, Kristensen P, Hendil KB, Fujiwara T, Takahashi E, Tanahashi N, Tamura T, Ichihara A. Newly identified pair of proteasomal subunits regulated reciprocally by interferon gamma. J Exp Med. 1996;183:1807-1816. [Cited in This Article: ] |
| 33. | Fehling HJ, Swat W, Laplace C, Kuhn R, Rajewsky K, Muller U, von Boehmer H. MHC class I expression in mice lacking the proteasome subunit LMP-7. Science. 1994;265:1234-1237. [Cited in This Article: ] |
| 34. | Van Kaer L, Ashton-Rickardt PG, Eichelberger M, Gaczynska M, Nagashima K, Rock KL, Goldberg AL, Doherty PC, Tonegawa S. Altered peptidase and viral-specific T cell response in LMP2 mutant mice. Immunity. 1994;1:533-541. [Cited in This Article: ] |
| 35. | Craiu A, Akopian T, Goldberg A, Rock KL. Two distinct proteolytic processes in the generation of a major histocompatibility complex class I-presented peptide. Proc Natl Acad Sci USA. 1997;94:10850-10855. [Cited in This Article: ] |
| 36. | Kloetzel PM, Ossendorp F. Proteasome and peptidase function in MHC-class-I-mediated antigen presentation. Curr Opin Immunol. 2004;16:76-81. [Cited in This Article: ] |
| 37. | Cascio P, Hilton C, Kisselev AF, Rock KL, Goldberg AL. 26S proteasomes and immunoproteasomes produce mainly N-extended versions of an antigenic peptide. EMBO J. 2001;20:2357-2366. [Cited in This Article: ] |
| 38. | York IA, Chang SC, Saric T, Keys JA, Favreau JM, Goldberg AL, Rock KL. The ER aminopeptidase ERAP1 enhances or limits antigen presentation by trimming epitopes to 8-9 residues. Nat Immunol. 2002;3:1177-1184. [Cited in This Article: ] |
| 39. | Brossart P, Bevan MJ. Presentation of exogenous protein antigens on major histocompatibility complex class I molecules by dendritic cells: pathway of presentation and regulation by cytokines. Blood. 1997;90:1594-1599. [Cited in This Article: ] |
| 40. | Brossart P, Goldrath AW, Butz EA, Martin S, Bevan MJ. Virus-mediated delivery of antigenic epitopes into dendritic cells as a means to induce CTL. J Immunol. 1997;158:3270-3276. [Cited in This Article: ] |
| 41. | Kessler BM, Glas R, Ploegh HL. MHC class I antigen processing regulated by cytosolic proteolysis-short cuts that alter peptide generation. Mol Immunol. 2002;39:171-179. [Cited in This Article: ] |
| 42. | Restifo NP, BacÃk I, Irvine KR, Yewdell JW, McCabe BJ, Anderson RW, Eisenlohr LC, Rosenberg SA, Bennink JR. Antigen processing in vivo and the elicitation of primary CTL responses. J Immunol. 1995;154:4414-4422. [Cited in This Article: ] |
| 43. | Rhodes SL, Erlich H, Im KA, Wang J, Li J, Bugawan T, Jeffers L, Tong X, Su X, Rosen HR. Associations between the human MHC and sustained virologic response in the treatment of chronic hepatitis C virus infection. Genes Immun. 2008;9:328-333. [Cited in This Article: ] |
| 44. | Seifert U, Liermann H, Racanelli V, Halenius A, Wiese M, Wedemeyer H, Ruppert T, Rispeter K, Henklein P, Sijts A. Hepatitis C virus mutation affects proteasomal epitope processing. J Clin Invest. 2004;114:250-259. [Cited in This Article: ] |
| 45. | Shin EC, Seifert U, Kato T, Rice CM, Feinstone SM, Kloetzel PM, Rehermann B. Virus-induced type I IFN stimulates generation of immunoproteasomes at the site of infection. J Clin Invest. 2006;116:3006-3014. [Cited in This Article: ] |
| 46. | Shin EC, Seifert U, Urban S, Truong KT, Feinstone SM, Rice CM, Kloetzel PM, Rehermann B. Proteasome activator and antigen-processing aminopeptidases are regulated by virus-induced type I interferon in the hepatitis C virus-infected liver. J Interferon Cytokine Res. 2007;27:985-990. [Cited in This Article: ] |
| 47. | Miyamoto H, Moriishi K, Moriya K, Murata S, Tanaka K, Suzuki T, Miyamura T, Koike K, Matsuura Y. Involvement of the PA28gamma-dependent pathway in insulin resistance induced by hepatitis C virus core protein. J Virol. 2007;81:1727-1735. [Cited in This Article: ] |
| 48. | Khu YL, Tan YJ, Lim SG, Hong W, Goh PY. Hepatitis C virus non-structural protein NS3 interacts with LMP7, a component of the immunoproteasome, and affects its proteasome activity. Biochem J. 2004;384:401-409. [Cited in This Article: ] |
| 49. | Osna NA, White RL, Krutik VM, Wang T, Weinman SA, Donohue TM Jr. Proteasome activation by hepatitis C core protein is reversed by ethanol-induced oxidative stress. Gastroenterology. 2008;134:2144-2152. [Cited in This Article: ] |
| 50. | Davies KJ. Degradation of oxidized proteins by the 20S proteasome. Biochimie. 2001;83:301-310. [Cited in This Article: ] |
| 51. | Osna NA, Haorah J, Krutik VM, Donohue TM Jr. Peroxy-nitrite alters the catalytic activity of rodent liver proteasome in vitro and in vivo. Hepatology. 2004;40:574-582. [Cited in This Article: ] |
| 52. | Osna NA, Clemens DL, Donohue TM Jr. Interferon gamma enhances proteasome activity in recombinant Hep G2 cells that express cytochrome P4502E1: modulation by ethanol. Biochem Pharmacol. 2003;66:697-710. [Cited in This Article: ] |
| 53. | Kessova IG, Cederbaum AI. The effect of CYP2E1-dependent oxidant stress on activity of proteasomes in HepG2 cells. J Pharmacol Exp Ther. 2005;315:304-312. [Cited in This Article: ] |
| 54. | Bardag-Gorce F, French BA, Nan L, Song H, Nguyen SK, Yong H, Dede J, French SW. CYP2E1 induced by ethanol causes oxidative stress, proteasome inhibition and cytokeratin aggresome (Mallory body-like) formation. Exp Mol Pathol. 2006;81:191-201. [Cited in This Article: ] |
| 55. | Bardag-Gorce F, Li J, French BA, French SW. The effect of ethanol-induced CYP2E1 on proteasome activity: the role of 4-hydroxynonenal. Exp Mol Pathol. 2005;78:109-115. [Cited in This Article: ] |
| 56. | Osna NA, White RL, Todero S, McVicker BL, Thiele GM, Clemens DL, Tuma DJ, Donohue TM Jr. Ethanol-induced oxidative stress suppresses generation of peptides for antigen presentation by hepatoma cells. Hepatology. 2007;45:53-61. [Cited in This Article: ] |
| 57. | Bryant P, Ploegh H. Class II MHC peptide loading by the professionals. Curr Opin Immunol. 2004;16:96-102. [Cited in This Article: ] |
| 58. | Hsieh CS, deRoos P, Honey K, Beers C, Rudensky AY. A role for cathepsin L and cathepsin S in peptide generation for MHC class II presentation. J Immunol. 2002;168:2618-2625. [Cited in This Article: ] |
| 59. | Ramachandra L, Chu RS, Askew D, Noss EH, Canaday DH, Potter NS, Johnsen A, Krieg AM, Nedrud JG, Boom WH. Phagocytic antigen processing and effects of microbial products on antigen processing and T-cell responses. Immunol Rev. 1999;168:217-239. [Cited in This Article: ] |
| 60. | Nimmerjahn F, Milosevic S, Behrends U, Jaffee EM, Pardoll DM, Bornkamm GW, Mautner J. Major histocompatibility complex class II-restricted presentation of a cytosolic antigen by autophagy. Eur J Immunol. 2003;33:1250-1259. [Cited in This Article: ] |
| 61. | Boes M, Cerny J, Massol R, Op den Brouw M, Kirchhausen T, Chen J, Ploegh HL. T-cell engagement of dendritic cells rapidly rearranges MHC class II transport. Nature. 2002;418:983-988. [Cited in This Article: ] |
| 62. | Kadowaki N, Ho S, Antonenko S, Malefyt RW, Kastelein RA, Bazan F, Liu YJ. Subsets of human dendritic cell precursors express different toll-like receptors and respond to different microbial antigens. J Exp Med. 2001;194:863-869. [Cited in This Article: ] |
| 63. | Liu YJ. IPC: professional type 1 interferon-producing cells and plasmacytoid dendritic cell precursors. Annu Rev Immunol. 2005;23:275-306. [Cited in This Article: ] |
| 64. | Liu B, Woltman AM, Janssen HL, Boonstra A. Modulation of dendritic cell function by persistent viruses. J Leukoc Biol. 2009;85:205-214. [Cited in This Article: ] |
| 65. | Bain C, Fatmi A, Zoulim F, Zarski JP, Trepo C, Inchauspe G. Impaired allostimulatory function of dendritic cells in chronic hepatitis C infection. Gastroenterology. 2001;120:512-524. [Cited in This Article: ] |
| 66. | Nattermann J, Zimmermann H, Iwan A, von Lilienfeld-Toal M, Leifeld L, Nischalke HD, Langhans B, Sauerbruch T, Spengler U. Hepatitis C virus E2 and CD81 interaction may be associated with altered trafficking of dendritic cells in chronic hepatitis C. Hepatology. 2006;44:945-954. [Cited in This Article: ] |
| 67. | Siavoshian S, Abraham JD, Thumann C, Kieny MP, Schuster C. Hepatitis C virus core, NS3, NS5A, NS5B proteins induce apoptosis in mature dendritic cells. J Med Virol. 2005;75:402-411. [Cited in This Article: ] |
| 68. | Averill L, Lee WM, Karandikar NJ. Differential dysfunction in dendritic cell subsets during chronic HCV infection. Clin Immunol. 2007;123:40-49. [Cited in This Article: ] |
| 69. | Crotta S, Stilla A, Wack A, D’Andrea A, Nuti S, D’Oro U, Mosca M, Filliponi F, Brunetto RM, Bonino F. Inhibition of natural killer cells through engagement of CD81 by the major hepatitis C virus envelope protein. J Exp Med. 2002;195:35-41. [Cited in This Article: ] |
| 70. | Dolganiuc A, Kodys K, Kopasz A, Marshall C, Do T, Romics L Jr, Mandrekar P, Zapp M, Szabo G. Hepatitis C virus core and nonstructural protein 3 proteins induce pro- and anti-inflammatory cytokines and inhibit dendritic cell differentiation. J Immunol. 2003;170:5615-5624. [Cited in This Article: ] |
| 71. | Tsubouchi E, Akbar SM, Murakami H, Horiike N, Onji M. Isolation and functional analysis of circulating dendritic cells from hepatitis C virus (HCV) RNA-positive and HCV RNA-negative patients with chronic hepatitis C: role of antiviral therapy. Clin Exp Immunol. 2004;137:417-423. [Cited in This Article: ] |
| 72. | Kanto T, Inoue M, Miyatake H, Sato A, Sakakibara M, Yakushijin T, Oki C, Itose I, Hiramatsu N, Takehara T. Reduced numbers and impaired ability of myeloid and plasmacytoid dendritic cells to polarize T helper cells in chronic hepatitis C virus infection. J Infect Dis. 2004;190:1919-1926. [Cited in This Article: ] |
| 73. | Dolganiuc A, Chang S, Kodys K, Mandrekar P, Bakis G, Cormier M, Szabo G. Hepatitis C virus (HCV) core protein-induced, monocyte-mediated mechanisms of reduced IFN-alpha and plasmacytoid dendritic cell loss in chronic HCV infection. J Immunol. 2006;177:6758-6768. [Cited in This Article: ] |
| 74. | Albert ML, Decalf J, Pol S. Plasmacytoid dendritic cells move down on the list of suspects: in search of the immune pathogenesis of chronic hepatitis C. J Hepatol. 2008;49:1069-1078. [Cited in This Article: ] |
| 75. | Szabo G, Aloman C, Polyak SJ, Weinman SA, Wands J, Zakhari S. Hepatitis C infection and alcohol use: A dangerous mix for the liver and antiviral immunity. Alcohol Clin Exp Res. 2006;30:709-719. [Cited in This Article: ] |
| 76. | Trujillo-Murillo K, Alvarez-Martinez O, Garza-Rodriguez L, Martinez-Rodriguez H, Bosques-Padilla F, Ramos-Jimenez J, Barrera-Saldana H, Rincon-Sanchez AR, Rivas-Estilla AM. Additive effect of ethanol and HCV subgenomic replicon expression on COX-2 protein levels and activity. J Viral Hepat. 2007;14:608-617. [Cited in This Article: ] |
| 77. | Geissler M, Gesien A, Tokushige K, Wands JR. Enhancement of cellular and humoral immune responses to hepatitis C virus core protein using DNA-based vaccines augmented with cytokine-expressing plasmids. J Immunol. 1997;158:1231-1237. [Cited in This Article: ] |
| 78. | Geissler M, Gesien A, Wands JR. Inhibitory effects of chronic ethanol consumption on cellular immune responses to hepatitis C virus core protein are reversed by genetic immunizations augmented with cytokine-expressing plasmids. J Immunol. 1997;159:5107-5113. [Cited in This Article: ] |
| 79. | Encke J, Wands JR. Ethanol inhibition: the humoral and cellular immune response to hepatitis C virus NS5 protein after genetic immunization. Alcohol Clin Exp Res. 2000;24:1063-1069. [Cited in This Article: ] |
| 80. | Szabo G, Dolganiuc A, Mandrekar P, White B. Inhibition of antigen-presenting cell functions by alcohol: implications for hepatitis C virus infection. Alcohol. 2004;33:241-249. [Cited in This Article: ] |
| 81. | Dolganiuc A, Kodys K, Kopasz A, Marshall C, Mandrekar P, Szabo G. Additive inhibition of dendritic cell allostimulatory capacity by alcohol and hepatitis C is not restored by DC maturation and involves abnormal IL-10 and IL-2 induction. Alcohol Clin Exp Res. 2003;27:1023-1031. [Cited in This Article: ] |
| 82. | Mandrekar P, Catalano D, Dolganiuc A, Kodys K, Szabo G. Inhibition of myeloid dendritic cell accessory cell function and induction of T cell anergy by alcohol correlates with decreased IL-12 production. J Immunol. 2004;173:3398-3407. [Cited in This Article: ] |
| 83. | Aloman C, Gehring S, Wintermeyer P, Kuzushita N, Wands JR. Chronic ethanol consumption impairs cellular immune responses against HCV NS5 protein due to dendritic cell dysfunction. Gastroenterology. 2007;132:698-708. [Cited in This Article: ] |
| 84. | Bode JG, Ludwig S, Ehrhardt C, Albrecht U, Erhardt A, Schaper F, Heinrich PC, Haussinger D. IFN-alpha antagonistic activity of HCV core protein involves induction of suppressor of cytokine signaling-3. FASEB J. 2003;17:488-490. [Cited in This Article: ] |
| 85. | Miyoshi H, Fujie H, Shintani Y, Tsutsumi T, Shinzawa S, Makuuchi M, Kokudo N, Matsuura Y, Suzuki T, Miyamura T. Hepatitis C virus core protein exerts an inhibitory effect on suppressor of cytokine signaling (SOCS)-1 gene expression. J Hepatol. 2005;43:757-763. [Cited in This Article: ] |
| 86. | Huang Y, Feld JJ, Sapp RK, Nanda S, Lin JH, Blatt LM, Fried MW, Murthy K, Liang TJ. Defective hepatic response to interferon and activation of suppressor of cytokine signaling 3 in chronic hepatitis C. Gastroenterology. 2007;132:733-744. [Cited in This Article: ] |
| 87. | Loo YM, Owen DM, Li K, Erickson AK, Johnson CL, Fish PM, Carney DS, Wang T, Ishida H, Yoneyama M. Viral and therapeutic control of IFN-beta promoter stimulator 1 during hepatitis C virus infection. Proc Natl Acad Sci USA. 2006;103:6001-6006. [Cited in This Article: ] |
| 88. | Liu B, Liao J, Rao X, Kushner SA, Chung CD, Chang DD, Shuai K. Inhibition of Stat1-mediated gene activation by PIAS1. Proc Natl Acad Sci USA. 1998;95:10626-10631. [Cited in This Article: ] |
| 89. | Helbig KJ, Yip E, McCartney EM, Eyre NS, Beard MR. A screening method for identifying disruptions in interferon signaling reveals HCV NS3/4a disrupts Stat-1 phosphorylation. Antiviral Res. 2008;77:169-176. [Cited in This Article: ] |
| 90. | Lin W, Kim SS, Yeung E, Kamegaya Y, Blackard JT, Kim KA, Holtzman MJ, Chung RT. Hepatitis C virus core protein blocks interferon signaling by interaction with the STAT1 SH2 domain. J Virol. 2006;80:9226-9235. [Cited in This Article: ] |
| 91. | Lin W, Choe WH, Hiasa Y, Kamegaya Y, Blackard JT, Schmidt EV, Chung RT. Hepatitis C virus expression suppresses interferon signaling by degrading STAT1. Gastroenterology. 2005;128:1034-1041. [Cited in This Article: ] |
| 92. | de Lucas S, Bartolome J, Carreno V. Hepatitis C virus core protein down-regulates transcription of interferon-induced antiviral genes. J Infect Dis. 2005;191:93-99. [Cited in This Article: ] |
| 93. | Melen K, Fagerlund R, Nyqvist M, Keskinen P, Julkunen I. Expression of hepatitis C virus core protein inhibits interferon-induced nuclear import of STATs. J Med Virol. 2004;73:536-547. [Cited in This Article: ] |
| 94. | Duong FH, Christen V, Berke JM, Penna SH, Moradpour D, Heim MH. Upregulation of protein phosphatase 2Ac by hepatitis C virus modulates NS3 helicase activity through inhibition of protein arginine methyltransferase 1. J Virol. 2005;79:15342-15350. [Cited in This Article: ] |
| 95. | Mowen KA, Tang J, Zhu W, Schurter BT, Shuai K, Herschman HR, David M. Arginine methylation of STAT1 modulates IFNalpha/beta-induced transcription. Cell. 2001;104:731-741. [Cited in This Article: ] |
| 96. | Duong FH, Christen V, Filipowicz M, Heim MH. S-Adenosylmethionine and betaine correct hepatitis C virus induced inhibition of interferon signaling in vitro. Hepatology. 2006;43:796-806. [Cited in This Article: ] |
| 97. | Dansako H, Naganuma A, Nakamura T, Ikeda F, Nozaki A, Kato N. Differential activation of interferon-inducible genes by hepatitis C virus core protein mediated by the interferon stimulated response element. Virus Res. 2003;97:17-30. [Cited in This Article: ] |
| 98. | Miller K, McArdle S, Gale MJ Jr, Geller DA, Tenoever B, Hiscott J, Gretch DR, Polyak SJ. Effects of the hepatitis C virus core protein on innate cellular defense pathways. J Interferon Cytokine Res. 2004;24:391-402. [Cited in This Article: ] |
| 99. | Hosui A, Ohkawa K, Ishida H, Sato A, Nakanishi F, Ueda K, Takehara T, Kasahara A, Sasaki Y, Hori M. Hepatitis C virus core protein differently regulates the JAK-STAT signaling pathway under interleukin-6 and interferon-gamma stimuli. J Biol Chem. 2003;278:28562-28571. [Cited in This Article: ] |
| 100. | Radaeva S, Jaruga B, Kim WH, Heller T, Liang TJ, Gao B. Interferon-gamma inhibits interferon-alpha signalling in hepatic cells: evidence for the involvement of STAT1 induction and hyperexpression of STAT1 in chronic hepatitis C. Biochem J. 2004;379:199-208. [Cited in This Article: ] |
| 101. | Plumlee CR, Lazaro CA, Fausto N, Polyak SJ. Effect of ethanol on innate antiviral pathways and HCV replication in human liver cells. Virol J. 2005;2:89. [Cited in This Article: ] |
| 102. | McCartney EM, Semendric L, Helbig KJ, Hinze S, Jones B, Weinman SA, Beard MR. Alcohol metabolism increases the replication of hepatitis C virus and attenuates the antiviral action of interferon. J Infect Dis. 2008;198:1766-1775. [Cited in This Article: ] |