Published online Jun 28, 2021. doi: 10.35712/aig.v2.i3.69
Peer-review started: January 27, 2021
First decision: March 29, 2021
Revised: April 9, 2021
Accepted: June 4, 2021
Article in press: June 4, 2021
Published online: June 28, 2021
Processing time: 158 Days and 13.9 Hours
Artificial intelligence (AI) applications are growing in medicine. It is important to understand the current state of the AI applications prior to utilizing in disease research and treatment. In this review, AI application in the diagnosis and treatment of gastrointestinal diseases are studied and summarized. In most cases, AI studies had large amounts of data, including images, to learn to distinguish disease characteristics according to a human’s perspectives. The detailed pros and cons of utilizing AI approaches should be investigated in advance to ensure the safe application of AI in medicine. Evidence suggests that the collaborative usage of AI in both diagnosis and treatment of diseases will increase the precision and effectiveness of medicine. Recent progress in genome technology such as genome editing provides a specific example where AI has revealed the diagnostic and therapeutic possibilities of RNA detection and targeting.
Core Tip: The application of artificial intelligence (AI) in the diagnosis and treatment of disease is a promising approach in medicine. The application of AI approaches in gastrointestinal diseases is summarized and reviewed. AI holds great promise in medicine, but to safely and efficiently apply AI in medicine, the advantages and limitations should first be carefully considered.
- Citation: Tanabe S, Perkins EJ, Ono R, Sasaki H. Artificial intelligence in gastrointestinal diseases. Artif Intell Gastroenterol 2021; 2(3): 69-76
- URL: https://www.wjgnet.com/2644-3236/full/v2/i3/69.htm
- DOI: https://dx.doi.org/10.35712/aig.v2.i3.69
Recent studies have developed RNA editing using the Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) system, which has made genome editing more accessible and has resulted in the development of many applications[1-3]. These new technologies have many advan
| Disease of interest | Purpose of AI | User | Limitation of use | Ref. |
| Acute appendicitis | Diagnosis | Specialist | The study is designed in retrospective nature | Reismann et al[5] |
| Colon cancer | Diagnosis | Specialist | The design of the analysis is post hoc and the number of patients is limited | Reichling et al[6] |
| Ulcerative colitis | Diagnosis | Specialist | Long-term clinical prognosis is not clear | Maeda et al[7] |
| Spinal stenosis in degenerative lumbar kyphoscoliosis | Surgery navigation | Specialist | The number of patients is limited. Long-term follow-up data is needed | Ho et al[9] |
| Coronavirus infectious disease (COVID-19) | Screening, diagnosis | Specialist | Privacy of the patient data should be considered | Bhattacharya et al[10] |
| Diseases in general | Diagnosis | Specialist | The burden on specialists may increase | Karako et al[11] |
| Diseases in general | Screening | Specialist | Careful and thorough investigation is necessary | Shiyam Sundar et al[12] |
| Gastrointestinal disease | Diagnosis | Specialist | There is a difference in the definition of anomaly detection between the area of computer science and medical domain | de Lange et al[13] |
| Gastrointestinal disease, hepatic diseases | Diagnosis | Specialist | High-quality datasets are needed | Le Berre et al[14] |
| Colorectal cancer | Diagnosis | Specialist | The quality of previous study designs is limited, and practical usefulness of computer-associated diagnosis systems is unknown | Kudo et al[15] |
| Colorectal cancer, bladder cancer | Prediction of anti-cancer drug efficacy | Specialist | Further molecular layer profiling in organoids may be needed | Kong et al[17] |
| Hypopharyngeal squamous cell carcinoma, esophageal squamous cell carcinoma | Identification of diagnostic and therapeutic targets | Specialist | Further studies are needed to validate the findings of the study | Zhou et al[18] |
| Arterial stenosis, coronary arterial diseases, stricture of the gastrointestinal tract | Guiding of balloon catheter | Specialist | The systemic performance needs to be improved | Kim et al[19] |
| Gastrointestinal disease | Diagnosis | Specialist | Further studies are needed to improve the performance | Marlicz et al[20] |
| Colorectal cancer | Prediction of liver metastasis | Specialist | The investigation of another dataset is needed | Lee et al[21] |
| Colon cancer | Diagnosis | Specialist | The change of protein expression level needs to be investigated | Xue et al[22] |
| Gastrointestinal disease | Diagnosis | Specialist | Investigation and development of newly improved methods are encouraged | Borgli et al[23] |
| Gastrointestinal disease | Diagnosis | Specialist | Further development is needed | Adler and Bjarnason[24] |
| Upper gastrointestinal cancer | Diagnosis | Specialist | Only high-quality endoscopic images for the training and validation analyses were used | Luo et al[25] |
| Gastric cancer | Diagnosis | Specialist | The associations of the quality or the number of training images and the CNN accuracy needs to be examined | Hirasawa et al[26] |
| Gastrointestinal disease | Diagnosis | Specialist | The possibilities to improve the medical performance, to reduce the medical cost, and to improve the satisfaction of the patient and medical staff are unknown | Min et al[27] |
| Functional gastrointestinal disorder | Diagnosis | Specialist | Evaluation of the feasibility of AI on studies on the gut-brain-microbiome axis is needed | Mukhtar et al[28] |
| Colorectal cancer | Diagnosis | Specialist | The uncertainty about the true efficacy of CAD in “real-world” practice remains | Ahmad et al[29] |
| Colorectal cancer | Diagnosis | Specialist | Further accumulation of lesion images for training is needed | Yamada et al[30] |
| Small-bowel disease | Diagnosis | Specialist | Further multicenter, prospective studies and external validation are needed | Yang[31] |
| Colorectal cancer | Diagnosis | Specialist | Complaints of system malfunctions and reports of patient injuries could lead to lawsuits against stakeholders | Ciuti et al[32] |
| Cholangiocarcinoma, pancreatic adenocarcinoma | Diagnosis | Specialist | Case-control and single-center design, and the lack of an independent validation cohort should be considered | Urman et al[33] |
| Colorectal cancer | Screening | Specialist | The applicability to other types of cancer needs optimization | Misawa et al[34] |
| Gastrointestinal disease | Diagnosis | Specialist | Most studies were designed in retrospective manner. Ethical issues on misdiagnosis or misclassification need to be handled | Yang and Bang[35] |
| Gastrointestinal cancer | Prediction of microsatellite instability for immunotherapy | Specialist | Larger training cohorts are needed | Kather et al[36] |
| Colorectal cancer | Diagnosis | Specialist | The CNN architecture needs to be improved for colonoscopy | Azer[37] |
| Barrett esophagus cancer | Diagnosis | Specialist | The number of patients is limited. Further optimization is needed | Ebigbo et al[38] |
| Celiac disease | Diagnosis | Specialist | The preliminary results need to be followed-up with a real clinical setting | Tenório et al[39] |
| Esophageal squamous cell carcinoma | Prediction of prognosis | Specialist | Further experimental studies to verify the results are needed | Zhang et al[40] |
| Advanced rectal adenocarcinoma | Prediction of response to neoadjuvant chemoradiotherapy | Specialist | The size of the cohort is limited. The confirmation of the findings with another data set is needed | Ferrando et al[41] |
| Inflammatory bowel disease | Prediction of prognosis | Specialist | Interventional study to confirm the efficacy of the stratifying therapy is needed | Biasci et al[42] |
| Inflammatory bowel disease | Mapping | Specialist | The application of advanced natural language processing algorithms to the text-mining step may improve the current process | Sarntivijai et al[43] |
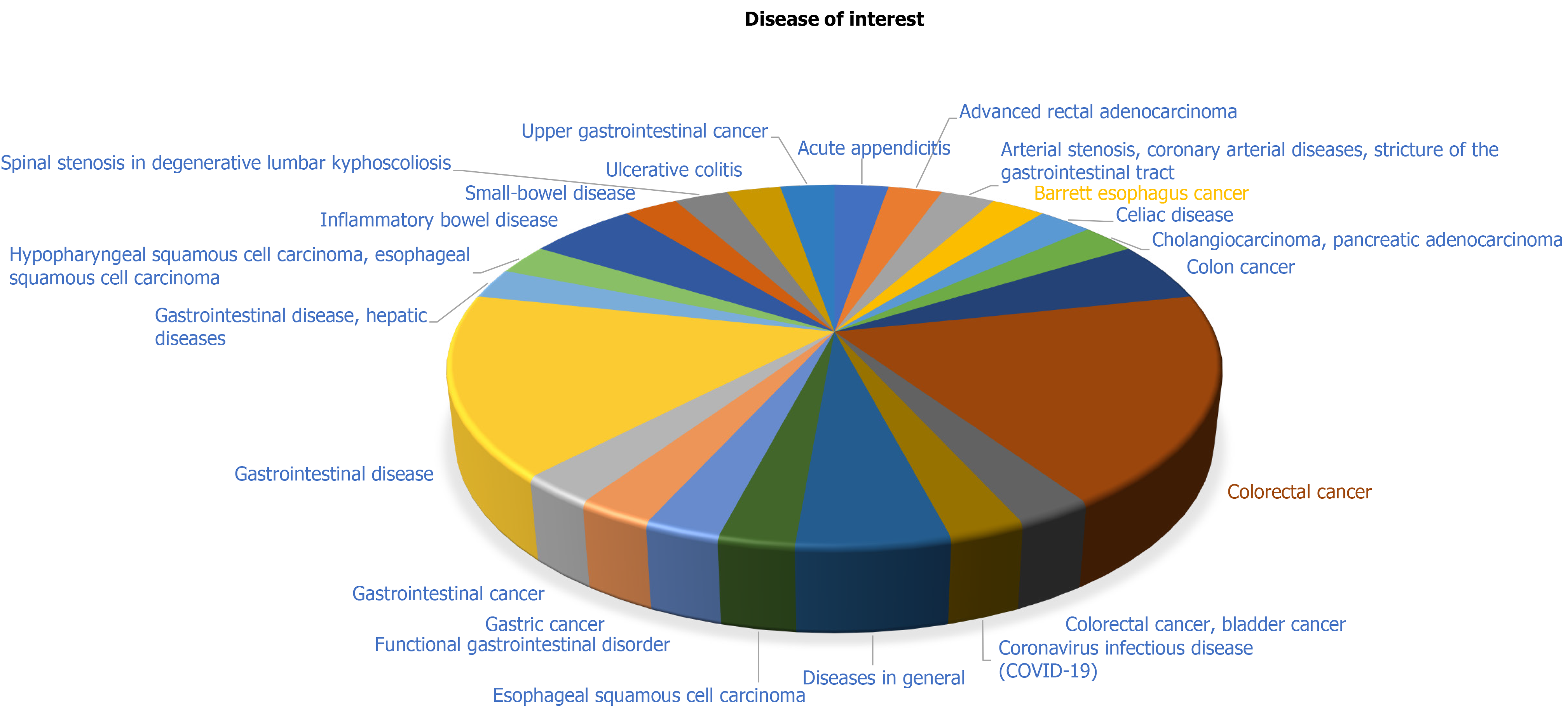
There are several areas in which AI can advance the diagnosis of GI diseases. Diseases of interest for AI-oriented disease management are summarized in Figure 1.
The diagnosis of GI diseases such as inflammatory bowel disease (IBD) including Crohn’s disease, a chronic inflammatory condition in the GI tract, and ulcerative colitis, which occurs in the colon, includes several fundamental laboratory tests including measurement of hemoglobin, hematocrit, blood urea nitrogen, creatinine, liver enzymes and C-reactive protein[16].
Recent progress in AI has resulted in predictive tools for the diagnosis of GI cancer classification, where network-based machine learning in colorectal and bladder organoid models predicts drug responders and non-responders using network analysis of pharmacogenomics data and the patient’s transcriptome[17]. Bioinformatic analyses of gene expression data have revealed common gene signatures in hypopharyngeal and esophageal squamous cell carcinoma, which may serve as diagnostic and therapeutic targets[18]. Balloon catheter tracking and visualization in GI tracking could be made more precise with AI guidance using image recognition[19]. Deep learning algorithms for image recognition can lead to more precise endoscopic diagnosis with improved sensitivity and specificity in upper GI tract diseases such as gastric cancer and Barrett’s esophagus[20]. Convolutional neural networks (CNNs) have generated liver imaging features and shown promise in predicting the metachronous liver metastasis in stage I-III colorectal cancer patients[21]. Deep learning of immunohistochemistry images of human colon tissues are used to improve the performance in detection of protein subcellular localization[22]. AI is poised to have a greater impact on GI endoscopy with publication of large datasets including multi-class images and video datasets that are useful for AI deep learning[23]. It seems that the performance of capsule endoscopy for diagnosing small bowel disease is improved using AI approaches[24]. An AI deep learning algorithm that can diagnose upper GI cancers with clinical endoscopic imaging data has been developed and validated[25]. CNNs in AI deep learning using numerous endoscopic image data have been developed that can detect and diagnose gastric cancer[26].
Min et al[27] pointed out that one drawback of AI approaches is the need for large datasets to train the system; therefore, the quality of CNN-based AI endoscopy is limited by the need for a large number of high-quality endoscopic images. Machine learning and AI are important to diagnose functional GI disorders and aid healthcare professionals and researchers[28]. Ahmad et al[29] suggested that the level of AI and CAD in colonoscopy has reached that of human expert performance. A real-time AI system with deep learning technology has been developed to diagnose colorectal cancer[30]. An AI-oriented automated CAD system can identify histologic inflammation associated with ulcerative colitis[7]. Reismann et al[5] used AI to identify biomarker signatures to diagnose and classify the pediatric acute appendicitis.
The application of AI in therapeutics of GI diseases has been expanding. The roles of AI in capsule endoscopy and other recent advanced diagnostic technologies have increased in therapeutics of GI diseases[31,32]. AI analysis was implemented to build neural network models enabling the classification of patients with biliary strictures and identify potential biomarkers in human bile[33]. Machine learning on medical examination records has stimulated the development of preventative measures for colorectal cancer[34]. Retrospective and prospective clinical studies have been conducted to diagnose and predict the prognosis of GI diseases including gastroesophageal reflux disease, atrophic corpus gastritis, acute pancreatitis, acute lower GI bleeding, esophageal cancer, nonvariceal upper GI bleeding, ulcerative colitis after cytoapheresis therapy, IBD, lymph node metastasis in T1 colorectal cancer and postoperative distant metastasis in esophageal squamous cell carcinoma[35]. Kather et al[36] found that deep learning can be used to predict microsatellite instability from histology in GI cancer. Azer[37] developed CNN models that can detect and classify colorectal polyps, which may increase colonoscopy application in appropriate colorectal cancer therapeutics. AI-guided tissue analysis has been developed that predicts stage III colon cancer outcomes, which may improve patient care with pathologists’ assistance[6]. Ebigbo et al[38] found that AI utilization can be used to classify the Barret esophagus cancer. An AI-based clinical decision-support system has been developed to diagnose celiac disease[39]. Bioinformatics analyses have identified important genes associated with the pathogenesis and prognosis of esophageal squamous cell carcinoma, which may contribute to the molecular-targeted therapy[40]. Long non-coding RNA signature has been identified in locally advanced rectal adenocarcinoma, which may predict the response to neoadjuvant chemoradiotherapy in the patients[41]. Machine learning has been utilized for identifying prognostic biomarkers in the whole blood of IBD patients to support the personalized therapy[42]. Ontology tools such as Experimental Factor Ontology or the Ontology of Biomedical Association may be useful for mining the disease-phenotype associations for IBD[43]. Since the responsiveness toward drug alters in cancer cell phenotypes such as epithelial-mesenchymal transition in diffuse-type gastric cancer, the AI application in the identification of cancer subtype would lead to establish therapeutic strategy[44,45]. The machine learning algorithms may be applied to the therapy of the GI diseases in terms of gut-brain axis[28,46].
Despite the rapid advances of the application of AI in GI diseases, there still remains some concern in terms of the precision of AI-based diagnosis and the criteria for the therapeutics. Further evidence is needed to solely rely on CAD in colonoscopy to determine an appropriate endpoint[15]. Some regulatory coordination may be needed to use the combination of an AI-assisted device and CAD software[15]. The differences in levels of AI performance would be considered and adjusted for application in clinical situations[14]. More high-quality datasets are needed to establish deep learning algorithms[14].
The area for AI application is rapidly expanding in the diagnosis and therapeutics of GI diseases. AI utilization in image recognition is currently being used to diagnose diseases and assist with personalized therapy. Future studies on disease-phenotype association are needed to maximize the capacity and performance of AI to aide in practical situations.
The authors would like to acknowledge the colleagues for their support.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Japan
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): E, E
P-Reviewer: Eccher A, Hanada E, Liu Y S-Editor: Gao CC L-Editor: Filipodia P-Editor: Li JH
| 1. | Sternberg SH, Redding S, Jinek M, Greene EC, Doudna JA. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature. 2014;507:62-67. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1254] [Cited by in RCA: 1349] [Article Influence: 122.6] [Reference Citation Analysis (0)] |
| 2. | Pickar-Oliver A, Gersbach CA. The next generation of CRISPR-Cas technologies and applications. Nat Rev Mol Cell Biol. 2019;20:490-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 932] [Cited by in RCA: 937] [Article Influence: 156.2] [Reference Citation Analysis (0)] |
| 3. | Jansen R, Embden JD, Gaastra W, Schouls LM. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol. 2002;43:1565-1575. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1236] [Cited by in RCA: 1215] [Article Influence: 52.8] [Reference Citation Analysis (0)] |
| 4. | Chen SC, Lo CM, Wang SH, Su EC. RNA editing-based classification of diffuse gliomas: predicting isocitrate dehydrogenase mutation and chromosome 1p/19q codeletion. BMC Bioinformatics. 2019;20:659. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 5. | Reismann J, Romualdi A, Kiss N, Minderjahn MI, Kallarackal J, Schad M, Reismann M. Diagnosis and classification of pediatric acute appendicitis by artificial intelligence methods: An investigator-independent approach. PLoS One. 2019;14:e0222030. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 48] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 6. | Reichling C, Taieb J, Derangere V, Klopfenstein Q, Le Malicot K, Gornet JM, Becheur H, Fein F, Cojocarasu O, Kaminsky MC, Lagasse JP, Luet D, Nguyen S, Etienne PL, Gasmi M, Vanoli A, Perrier H, Puig PL, Emile JF, Lepage C, Ghiringhelli F. Artificial intelligence-guided tissue analysis combined with immune infiltrate assessment predicts stage III colon cancer outcomes in PETACC08 study. Gut. 2020;69:681-690. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 84] [Article Influence: 16.8] [Reference Citation Analysis (0)] |
| 7. | Maeda Y, Kudo SE, Mori Y, Misawa M, Ogata N, Sasanuma S, Wakamura K, Oda M, Mori K, Ohtsuka K. Fully automated diagnostic system with artificial intelligence using endocytoscopy to identify the presence of histologic inflammation associated with ulcerative colitis (with video). Gastrointest Endosc. 2019;89:408-415. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 124] [Cited by in RCA: 165] [Article Influence: 27.5] [Reference Citation Analysis (0)] |
| 8. | LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436-444. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36149] [Cited by in RCA: 20049] [Article Influence: 2004.9] [Reference Citation Analysis (0)] |
| 9. | Ho TY, Lin CW, Chang CC, Chen HT, Chen YJ, Lo YS, Hsiao PH, Chen PC, Lin CS, Tsou HK. Percutaneous endoscopic unilateral laminotomy and bilateral decompression under 3D real-time image-guided navigation for spinal stenosis in degenerative lumbar kyphoscoliosis patients: an innovative preliminary study. BMC Musculoskelet Disord. 2020;21:734. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 6] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 10. | Bhattacharya S, Reddy Maddikunta PK, Pham QV, Gadekallu TR, Krishnan S SR, Chowdhary CL, Alazab M, Jalil Piran M. Deep learning and medical image processing for coronavirus (COVID-19) pandemic: A survey. Sustain Cities Soc. 2021;65:102589. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 310] [Cited by in RCA: 169] [Article Influence: 42.3] [Reference Citation Analysis (0)] |
| 11. | Karako K, Song P, Chen Y, Tang W. Realizing 5G- and AI-based doctor-to-doctor remote diagnosis: opportunities, challenges, and prospects. Biosci Trends. 2020;14:314-317. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 12. | Shiyam Sundar LK, Muzik O, Buvat I, Bidaut L, Beyer T. Potentials and caveats of AI in hybrid imaging. Methods. 2021;188:4-19. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 13. | de Lange T, Halvorsen P, Riegler M. Methodology to develop machine learning algorithms to improve performance in gastrointestinal endoscopy. World J Gastroenterol. 2018;24:5057-5062. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 29] [Cited by in RCA: 23] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 14. | Le Berre C, Sandborn WJ, Aridhi S, Devignes MD, Fournier L, Smaïl-Tabbone M, Danese S, Peyrin-Biroulet L. Application of Artificial Intelligence to Gastroenterology and Hepatology. Gastroenterology 2020; 158: 76-94. e2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 230] [Cited by in RCA: 324] [Article Influence: 64.8] [Reference Citation Analysis (1)] |
| 15. | Kudo SE, Mori Y, Misawa M, Takeda K, Kudo T, Itoh H, Oda M, Mori K. Artificial intelligence and colonoscopy: Current status and future perspectives. Dig Endosc. 2019;31:363-371. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 83] [Article Influence: 13.8] [Reference Citation Analysis (0)] |
| 16. | Wilkins T, Jarvis K, Patel J. Diagnosis and management of Crohn's disease. Am Fam Physician. 2011;84:1365-1375. [PubMed] |
| 17. | Kong J, Lee H, Kim D, Han SK, Ha D, Shin K, Kim S. Network-based machine learning in colorectal and bladder organoid models predicts anti-cancer drug efficacy in patients. Nat Commun. 2020;11:5485. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 93] [Cited by in RCA: 126] [Article Influence: 25.2] [Reference Citation Analysis (0)] |
| 18. | Zhou R, Liu D, Zhu J, Zhang T. Common gene signatures and key pathways in hypopharyngeal and esophageal squamous cell carcinoma: Evidence from bioinformatic analysis. Medicine (Baltimore). 2020;99:e22434. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 19. | Kim J, Mai TT, Kim JY, Min JJ, Kim C, Lee C. Feasibility Study of Precise Balloon Catheter Tracking and Visualization with Fast Photoacoustic Microscopy. Sensors (Basel). 2020;20. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 20. | Marlicz W, Ren X, Robertson A, Skonieczna-Żydecka K, Łoniewski I, Dario P, Wang S, Plevris JN, Koulaouzidis A, Ciuti G. Frontiers of Robotic Gastroscopy: A Comprehensive Review of Robotic Gastroscopes and Technologies. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 25] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 21. | Lee S, Choe EK, Kim SY, Kim HS, Park KJ, Kim D. Liver imaging features by convolutional neural network to predict the metachronous liver metastasis in stage I-III colorectal cancer patients based on preoperative abdominal CT scan. BMC Bioinformatics. 2020;21:382. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 26] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 22. | Xue ZZ, Wu Y, Gao QZ, Zhao L, Xu YY. Automated classification of protein subcellular localization in immunohistochemistry images to reveal biomarkers in colon cancer. BMC Bioinformatics. 2020;21:398. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 23. | Borgli H, Thambawita V, Smedsrud PH, Hicks S, Jha D, Eskeland SL, Randel KR, Pogorelov K, Lux M, Nguyen DTD, Johansen D, Griwodz C, Stensland HK, Garcia-Ceja E, Schmidt PT, Hammer HL, Riegler MA, Halvorsen P, de Lange T. HyperKvasir, a comprehensive multi-class image and video dataset for gastrointestinal endoscopy. Sci Data. 2020;7:283. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 297] [Cited by in RCA: 136] [Article Influence: 27.2] [Reference Citation Analysis (0)] |
| 24. | Adler SN, Bjarnason I. What we have learned and what to expect from capsule endoscopy. World J Gastrointest Endosc. 2012;4:448-452. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 12] [Cited by in RCA: 12] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 25. | Luo H, Xu G, Li C, He L, Luo L, Wang Z, Jing B, Deng Y, Jin Y, Li Y, Li B, Tan W, He C, Seeruttun SR, Wu Q, Huang J, Huang DW, Chen B, Lin SB, Chen QM, Yuan CM, Chen HX, Pu HY, Zhou F, He Y, Xu RH. Real-time artificial intelligence for detection of upper gastrointestinal cancer by endoscopy: a multicentre, case-control, diagnostic study. Lancet Oncol. 2019;20:1645-1654. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 155] [Cited by in RCA: 253] [Article Influence: 42.2] [Reference Citation Analysis (0)] |
| 26. | Hirasawa T, Aoyama K, Tanimoto T, Ishihara S, Shichijo S, Ozawa T, Ohnishi T, Fujishiro M, Matsuo K, Fujisaki J, Tada T. Application of artificial intelligence using a convolutional neural network for detecting gastric cancer in endoscopic images. Gastric Cancer. 2018;21:653-660. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 568] [Cited by in RCA: 427] [Article Influence: 61.0] [Reference Citation Analysis (0)] |
| 27. | Min JK, Kwak MS, Cha JM. Overview of Deep Learning in Gastrointestinal Endoscopy. Gut Liver. 2019;13:388-393. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 74] [Cited by in RCA: 116] [Article Influence: 23.2] [Reference Citation Analysis (0)] |
| 28. | Mukhtar K, Nawaz H, Abid S. Functional gastrointestinal disorders and gut-brain axis: What does the future hold? World J Gastroenterol. 2019;25:552-566. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 105] [Cited by in RCA: 98] [Article Influence: 16.3] [Reference Citation Analysis (3)] |
| 29. | Ahmad OF, Soares AS, Mazomenos E, Brandao P, Vega R, Seward E, Stoyanov D, Chand M, Lovat LB. Artificial intelligence and computer-aided diagnosis in colonoscopy: current evidence and future directions. Lancet Gastroenterol Hepatol. 2019;4:71-80. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 126] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 30. | Yamada M, Saito Y, Imaoka H, Saiko M, Yamada S, Kondo H, Takamaru H, Sakamoto T, Sese J, Kuchiba A, Shibata T, Hamamoto R. Development of a real-time endoscopic image diagnosis support system using deep learning technology in colonoscopy. Sci Rep. 2019;9:14465. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 114] [Cited by in RCA: 161] [Article Influence: 26.8] [Reference Citation Analysis (0)] |
| 31. | Yang YJ. The Future of Capsule Endoscopy: The Role of Artificial Intelligence and Other Technical Advancements. Clin Endosc. 2020;53:387-394. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 25] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 32. | Ciuti G, Skonieczna-Żydecka K, Marlicz W, Iacovacci V, Liu H, Stoyanov D, Arezzo A, Chiurazzi M, Toth E, Thorlacius H, Dario P, Koulaouzidis A. Frontiers of Robotic Colonoscopy: A Comprehensive Review of Robotic Colonoscopes and Technologies. J Clin Med. 2020;9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 83] [Cited by in RCA: 50] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 33. | Urman JM, Herranz JM, Uriarte I, Rullán M, Oyón D, González B, Fernandez-Urién I, Carrascosa J, Bolado F, Zabalza L, Arechederra M, Alvarez-Sola G, Colyn L, Latasa MU, Puchades-Carrasco L, Pineda-Lucena A, Iraburu MJ, Iruarrizaga-Lejarreta M, Alonso C, Sangro B, Purroy A, Gil I, Carmona L, Cubero FJ, Martínez-Chantar ML, Banales JM, Romero MR, Macias RIR, Monte MJ, Marín JJG, Vila JJ, Corrales FJ, Berasain C, Fernández-Barrena MG, Avila MA. Pilot Multi-Omic Analysis of Human Bile from Benign and Malignant Biliary Strictures: A Machine-Learning Approach. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 48] [Cited by in RCA: 45] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 34. | Misawa D, Fukuyoshi J, Sengoku S. Cancer Prevention Using Machine Learning, Nudge Theory and Social Impact Bond. Int J Environ Res Public Health. 2020;17. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 15] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 35. | Yang YJ, Bang CS. Application of artificial intelligence in gastroenterology. World J Gastroenterol. 2019;25:1666-1683. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 211] [Cited by in RCA: 160] [Article Influence: 26.7] [Reference Citation Analysis (5)] |
| 36. | Kather JN, Pearson AT, Halama N, Jäger D, Krause J, Loosen SH, Marx A, Boor P, Tacke F, Neumann UP, Grabsch HI, Yoshikawa T, Brenner H, Chang-Claude J, Hoffmeister M, Trautwein C, Luedde T. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat Med. 2019;25:1054-1056. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 992] [Cited by in RCA: 760] [Article Influence: 126.7] [Reference Citation Analysis (0)] |
| 37. | Azer SA. Challenges Facing the Detection of Colonic Polyps: What Can Deep Learning Do? Medicina (Kaunas). 2019;55. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 24] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 38. | Ebigbo A, Mendel R, Probst A, Manzeneder J, Prinz F, de Souza LA Jr, Papa J, Palm C, Messmann H. Real-time use of artificial intelligence in the evaluation of cancer in Barrett's oesophagus. Gut. 2020;69:615-616. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 84] [Cited by in RCA: 124] [Article Influence: 24.8] [Reference Citation Analysis (0)] |
| 39. | Tenório JM, Hummel AD, Cohrs FM, Sdepanian VL, Pisa IT, de Fátima Marin H. Artificial intelligence techniques applied to the development of a decision-support system for diagnosing celiac disease. Int J Med Inform. 2011;80:793-802. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 45] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 40. | Zhang H, Zhong J, Tu Y, Liu B, Chen Z, Luo Y, Tang Y, Xiao F. Integrated Bioinformatics Analysis Identifies Hub Genes Associated with the Pathogenesis and Prognosis of Esophageal Squamous Cell Carcinoma. Biomed Res Int. 2019;2019:2615921. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 22] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 41. | Ferrando L, Cirmena G, Garuti A, Scabini S, Grillo F, Mastracci L, Isnaldi E, Marrone C, Gonella R, Murialdo R, Fiocca R, Romairone E, Ballestrero A, Zoppoli G. Development of a long non-coding RNA signature for prediction of response to neoadjuvant chemoradiotherapy in locally advanced rectal adenocarcinoma. PLoS One. 2020;15:e0226595. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 42. | Biasci D, Lee JC, Noor NM, Pombal DR, Hou M, Lewis N, Ahmad T, Hart A, Parkes M, McKinney EF, Lyons PA, Smith KGC. A blood-based prognostic biomarker in IBD. Gut. 2019;68:1386-1395. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 125] [Cited by in RCA: 147] [Article Influence: 24.5] [Reference Citation Analysis (0)] |
| 43. | Sarntivijai S, Vasant D, Jupp S, Saunders G, Bento AP, Gonzalez D, Betts J, Hasan S, Koscielny G, Dunham I, Parkinson H, Malone J. Linking rare and common disease: mapping clinical disease-phenotypes to ontologies in therapeutic target validation. J Biomed Semantics. 2016;7:8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 24] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 44. | Tanabe S, Quader S, Cabral H, Ono R. Interplay of EMT and CSC in Cancer and the Potential Therapeutic Strategies. Front Pharmacol. 2020;11:904. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 121] [Cited by in RCA: 112] [Article Influence: 22.4] [Reference Citation Analysis (0)] |
| 45. | Tanabe S, Quader S, Ono R, Cabral H, Aoyagi K, Hirose A, Yokozaki H, Sasaki H. Molecular Network Profiling in Intestinal- and Diffuse-Type Gastric Cancer. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 23] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 46. | Kano M, Dupont P, Aziz Q, Fukudo S. Understanding Neurogastroenterology From Neuroimaging Perspective: A Comprehensive Review of Functional and Structural Brain Imaging in Functional Gastrointestinal Disorders. J Neurogastroenterol Motil. 2018;24:512-527. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 62] [Article Influence: 8.9] [Reference Citation Analysis (0)] |