Published online Sep 26, 2017. doi: 10.5662/wjm.v7.i3.101
Peer-review started: July 17, 2017
First decision: August 4, 2017
Revised: August 22, 2017
Accepted: September 3, 2017
Article in press: September 4, 2017
Published online: September 26, 2017
Processing time: 70 Days and 13.6 Hours
Different causative factors acting on the pancreas can result in diseases such as pancreatitis, diabetes and pancreatic tumors. The high incidence and mortality of pancreatic diseases have placed diagnostic imaging in a crucial position in daily clinical practice. In this mini-review article different pancreatic imaging techniques are discussed, from the standard clinical imaging modalities and state of the art clinical magnetic resonance imaging techniques to current situations in pre-clinical pancreatic imaging studies. In particular, the challenges of pre-clinical rodent pancreatic imaging are addressed, with both the image acquisition techniques and the post-processing methods for rodent pancreatic imaging elaborated.
Core tip: In this minireview, the challenges of pre-clinical rodent pancreatic imaging are addressed, basic clinical magnetic resonance imaging techniques and post-processing methods for rodent pancreatic imaging are also elaborated.
- Citation: Yin T, Liu Y, Peeters R, Feng Y, Ni Y. Pancreatic imaging: Current status of clinical practices and small animal studies. World J Methodol 2017; 7(3): 101-107
- URL: https://www.wjgnet.com/2222-0682/full/v7/i3/101.htm
- DOI: https://dx.doi.org/10.5662/wjm.v7.i3.101
The pancreas is an important visceral organ performing both endocrine and exocrine functions. Abnormalities of the pancreas result in diseases such as pancreatitis, diabetes, and pancreatic tumors[1,2]. The onset of diabetes is usually long after beta cell dysfunction and insulin resistance[3,4]; pancreatic cancer is generally asymptomatic and frequently diagnosed at a late stage[5]; acute pancreatitis is a painful inflammatory condition often with severe complications and high mortality despite treatment[6], while chronic pancreatitis can mimic the symptoms of pancreatic cancer and lead to misdiagnosis[7]. The high incidence of pancreatitis and diabetes, and poor survival rate of pancreatic cancers have increased the demand for new diagnostic and therapeutic strategies[8,9] Herein multimodality multi-parametric imaging plays an indispensable role in disease detection, therapy guidance and patient follow-up. In this mini-review, current situations of common clinical practices and recent development of pre-clinical rodent studies in pancreatic imaging are inspected and discussed with the emphasis on basic magnetic resonance imaging (MRI) techniques and post-processing methods for rodent pancreatic studies.
As an initial step, abdominal ultrasound is most commonly used in screening for biliary stones and tumors, as this equipment is widely available at relatively low costs[10]. However, the quality of ultrasound images and diagnostic accuracy are highly user-dependent, and the retroperitoneal location of the pancreas may impose image artifacts and hamper the ultrasound diagnosis[11]. For further confirmation and staging of pancreatic diseases, imaging modalities with higher quality and sensitivity are needed.
Contrast-enhanced multi-detector computed tomography (MDCT) remains the standard modality in clinic for the assessment of pancreatitis and pancreatic cancer[12,13]. Due to its high spatial resolution and fast image acquisition, MDCT combined with contrast agents injection, has shown its powerful capacity in the staging of pancreatitis and pancreatic cancer with high sensitivity and specificity[7,12].
MRI as a non-ionizing imaging modality has been increasingly utilized in clinic due to its multi-parametric capability[14]. With the constant improvement of the new MRI hardware and imaging reconstruction algorithms, MRI is currently capable of acquiring images of spatial resolution approaching to that of CT. Meanwhile, with the application of accelerated parallel imaging techniques, most MRI protocols have the feasibility to be accomplished in one or a few breath-holds[14,15].
Traditionally, T2-weighted MRI sequences are commonly used to provide structural information on the anatomy of the pancreatic ductal system and lesions[14]. MR cholangiopancreatography (MRCP) using heavily T2-weighted sequences has been widely applied as non-invasive alternative to endoscopic retrograde cholangiopancreatography (ERCP) for biliopancreatic duct system evaluation[16]. The combination of dual-echo (in/opposed-phase) T1-weighted MRI sequences is useful for hemorrhage and fat content assessment[14]. Dynamic contrast-enhanced (DCE) MRI scans are performed for differential diagnosis and grading of solid pancreatic lesions and pancreatitis by analyzing the pharmacokinetic parameters or contrast concentration curves[17,18]. In addition, diffusion-weighted MRI (DWI) protocols have also shown a great potential to depict and characterize pancreatic diseases including acute/chronic pancreatitis and benign or malignant tumors[19] without a need to use contrast agents.
Other more advanced but less popular pancreatic imaging modalities, often with a certain invasiveness or radiation exposure, include endoscopic and contrast enhanced US (EUS and CEUS), positron emission computed tomography and single-photon emission computed tomography incorporated with X-ray based CT (PET/CT and SPECT/CT) for improved spatial resolution and co-localization of imaging findings, etc. For a more comprehensive overview, a recent review article about imaging pancreatic diseases is recommended[5].
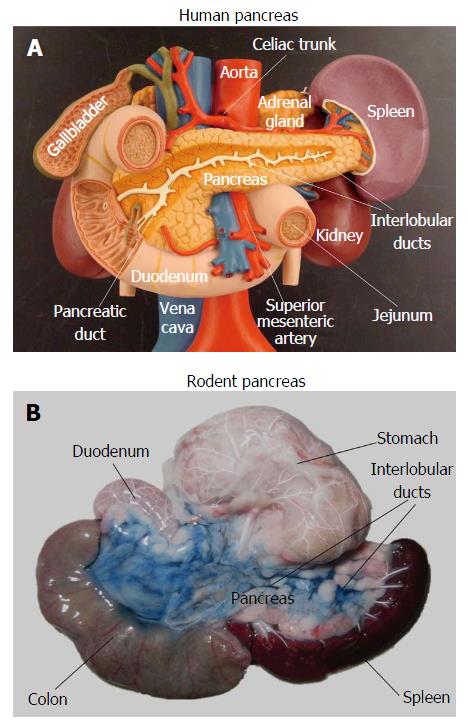
In order to develop new diagnostic and therapeutic strategies against pancreatic diseases, rodent models are commonly used in preclinical studies. However, imaging the pancreas in rodents proves to be extremely challenging due to motion artifacts and uncertain anatomy. The pancreas in humans represents a retroperitoneal solid organ, which can be identified by imaging modalities even without contrast enhancement, as stated above. However, unlike the human pancreas, the rodent pancreas appears as a soft, diffuse and irregularly lobulated organ, which is very difficult to discern from surrounding tissues[19-22], even during open abdominal surgery (Figure 1). Pancreas-specific contrast agents would facilitate pancreas visualization, but currently those pancreatic specific markers are unavailable yet. Without specific labeling, rodent caudate liver lobes and abdominal fat tissue are frequently identified as pancreas by mistake. In some animal studies, pancreas associated injuries in other solid organs, instead of the pancreas itself, were investigated using contrast-enhanced protocols and MR spectroscopy (MRS)[23,24].
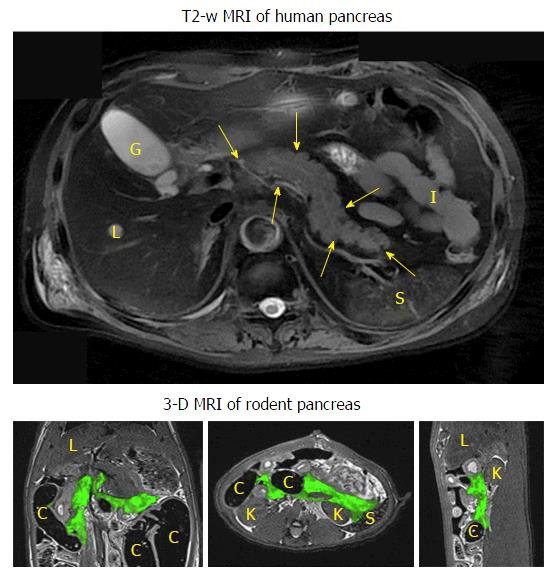
To avoid the misdiagnosis and to have a detailed overview of the pancreas anatomy, two pancreatic imaging studies were performed using contrast-enhanced high-resolution three dimensional (3D) modalities to provide more precise anatomical information of the rodent pancreas[20,22]. In a micro-CT study[20], the in vivo rat pancreatic tail portion could be identified after a two-step contrast injection. Unlike the human pancreas that can be readily depicted by MRI even without using any contrast agent (Figure 2), detailed 3D rodent pancreatic anatomy and surrounding landmarks could only be demonstrated (Figure 2) by biliopancreatic local infusion of mixed contrast media in a post-mortem model[22].
In order to achieve early diagnosis in diabetes, the development of pancreatic specific contrast agents became a hot topic. Among others, some contrast agents were used to target pancreatic beta cells for diabetes related research subjects, for instance, glucagon-like peptide-1 (GLP1) receptor and GLP1 analogues have been frequently studied in rodent diabetes imaging[25,26]. The micro-vascular changes in case of diabetes and pancreatic inflammation were also investigated[27,28].
The first attempt of rodent pancreatitis imaging started during the 1980’s, conducted by Paajanen et al[29], in which Gd-DTPA was applied as a contrast agent for T2/T1 relaxation measurements in an acute hemorrhagic pancreatitis model. In 1989, Kushnir et al[30] performed several MRS experiments to identify imaging bio-markers in an acute pancreatitis model. More recently, specific nanoparticles were developed for pancreatitis imaging, with lipase as the target[31]. Imaging of acute edematous pancreatitis can also be performed with MRCP, T2 relaxation measurement and non-specific contrast enhancement using modified protocols on a state of the art clinical MRI scanner[32].
The direct visualization of rodent pancreas and corresponding pancreatic landmarks could facilitate more precise diagnosis of a pancreatic tumor in the early stages. As a tumor grows to a certain volume, the identification of the solid tumor mass is much easier to perform than imaging other pancreatic disorders. Quantitative T2 and T1 relaxation measurements, DWI parameters and perfusion information can be obtained using multi-parametric MRI[33]. Currently, rodent pancreatic tumor models are increasingly used to investigate new therapeutic strategies in longitudinal follow-up studies by non-invasive MRI.
Due to the small size of the rodent pancreas, it is necessary to use high resolution 3D anatomical images for precise pancreas localization. Misdiagnosis could be avoided by carefully tracking the anatomy of the surrounding organs or tissues in 3D mode. Unfortunately, 3D anatomical MRI in the abdominal region is extremely challenging in commonly used high-field pre-clinical scanners, due to their high sensitivity to motion artifacts at high magnetic field and unavoidable long scanning durations. Alternatively, by the combined use of dedicated multi-channel coils and accelerated parallel imaging, clinical MR scanners have shown the feasibility and flexibility for rodent abdominal imaging[22,32,33].
The biggest problem using clinical scanners for small animal studies is the limited gradient strength[34]. Most clinical MRI scanners have a gradient amplitude of 40 mT/m and slew rates up to 200 T/m per second. Although the maximum gradient amplitude in the recent 3T Siemens Prisma scanner has been increased to 80 mT/m, the gradient strengths are still up to 10 times lower than that of the current state of the art pre-clinical scanners. Consequently, the minimum slice thickness and minimal field of view (FOV) in the clinical systems are more restricted. In our studies, to maintain enough signal-to-noise-ratio (SNR), most 2D scans were performed with a slice thickness of 2 mm, which is identical to those acquired from small animal scanners. However, the minimal FOV is usually limited to around 70 mm. The limited gradient strengths also hamper the use of echo planar imaging (EPI) due to the prolonged readout, which leads to severe image distortions.
On the other hand, current state of the art clinical MRI scanners provide an excellent hardware stability, higher field homogeneity and a dedicated user interface, and are more widely accessible compared to small animal scanners. With the combination of dedicated clinical multi-channel surface coils and the self-calibrated parallel imaging techniques GRAPPA (GeneRalized Autocalibrating Partial Parallel Acquisition), it is possible to acquire high SNR images in rodent heterogeneous abdominal region with a sufficiently short scan duration. Moreover, lower magnetic field and application of GRAPPA provide a higher feasibility to rodent abdominal imaging.
In our serial studies[22,32,33], clinical scanners were used for pancreatic imaging: The Magnetom Tim Trio (Siemens, Erlangen, 45 mT/m, 200 T/m per second) combined with an 8-channel wrist coil; and the Magnetom Prisma (Siemens, Erlangen, 80 mT/m, 200 T/m per second) together with a 16-channel wrist coil. There are several standard clinical protocols which can be directly translated to pre-clinical research, including T2-weighted 3D turbo spin-echo (TSE) based SPACE (3D TSE with variable-flip-angle refocusing RF pulses) imaging and MRCP protocols, standard 2D multi-echo spin-echo sequences for T2 relaxation, as well as diffusion and perfusion sequences. The animal models introduced in these studies[22,32,33] are a rat acute pancreatitis model and a rat orthotopic pancreatic tumor model, in which we intended to characterize pathological changes including edema, hemorrhage and necrosis by those modified MRI methods.
Three-dimensional volumetric images: As mentioned above, 3D imaging would facilitate the localization of rodent pancreas. The other benefit is the possibility of volumetric measurements in 3D. In case of edematous pancreatitis, edema volume is a biomarker for pancreatitis. Meanwhile, 3D views could also provide a more accurate measurement for irregularly shaped abdominal tumors (Figure 2). The volume of the target tissue can be obtained from post-process image segmentation. The most important 3D imaging protocols used here are T2-weighted SPACE and MRCP, which are also widely used in clinical MRI abdominal examinations.
Quantitative MRI measurements: Quantitative MRI relaxation measurements are useful in organ/tissue characterization. T2 mapping is helpful in the assessment of fluid content and hemorrhage; and native T1 mapping is essential for the calculation of the tissue concentration time curve (CTC) in DCE protocols. In these studies, mono-exponential T2 mapping and inversion recovery based T1 mapping were performed.
Measurements of Gaussian and non-Gaussian diffusion for water in biological tissues can be accomplished using DWI with different combinations of diffusion weightings. Mean diffusivity and diffusion kurtosis were obtained from 3-trace diffusion images.
Moreover, tissue perfusion can be characterized using DCE protocols, after the injection of a gadolinium based MRI contrast agent. After the signal conversion to gadolinium concentration using pre-contrast native T1 relaxation information, either semi-quantitative information or quantitative parameters from the pharmacokinetic Tofts model were extracted. Detailed MRI protocol parameters are elaborated in the different serial studies[22,32,33].
In these studies, open-source software and in-house built programs were used for data processing. This includes image segmentation, registration and 3D image visualization in open-source software ITK-SNAP (http://www.itksnap.org) and MeVisLab (MeVis Medical Solutions, Bremen, Germany); DICOM process, MRI mathematical modeling and quantitative analysis in Matlab programs (MathWorks, Natick, Massachusetts); and statistical analysis and data visualization using programs combining both Matlab and R (https://cran.r-project.org). Detailed image processing equations are included in the next section.
Present research project aims at providing practical solutions to rodent pancreatic imaging using clinical facilities, from ex vivo to noninvasive in vivo imaging with the following systematic objectives identified: (1) To overcome the limitations of clinical MRI scanner for small rodent imaging studies; (2) detailed visualization of a complete pancreas and topographic landmarks through contrast enhanced CT and MR imaging in a rat postmortem model; (3) to explore noninvasive MR imaging methods for characterization of the Caerulein induced acute pancreatitis in rats; (4) to estimate the reliability of 3D isotropic MRI and quantitative multi-parametric MRI for characterization of an orthotopic pancreatic head tumor model in rats; and (5) to investigate therapeutic response of a vascular disruption agent in rat pancreatic tumor models with further modified quantitative multi-parametric methods.
MRI quantitative parameters can be obtained from advanced image processing methods using machine learning algorithms[35-37]. For practical considerations, quantitative parameters are re-generated from in-house built Matlab programs using non-linear least square methods with CPU (Central Processing Unit) acceleration.
T2 mapping: Traditionally, the transverse relaxation time T2 is obtained using multi-echo spin-echo pulse sequences, by sampling signals at several different echo-times (TE), and fitted to either multi- or single-exponential decay functions[38]. Fast T2 mapping can be obtained using balanced steady-state free precession (SSFP) readout[39].
T1 mapping: On a clinical scanner, fast T1 mapping can be measured using inversion recovery methods or from variable flip angles experiments[39,40]. Since the MRI acquisition has to be synchronized with animal respiration, the effective repetition time (TR) is usually longer than 1 s. Thus, inversion recovery based protocols would be suggested for T1 mapping in rodent pancreatic imaging. Typically, the equation for measured signal in the inversion recovery T1 mapping experiment is a three-parameter function: SI(t) = a + b × exp(-t/T1*), where SI(t) is the signal intensity after each inversion time t, and T1* is the effective longitudinal relaxation time. The actual T1 relaxation time can be obtained after correction for the flip angles[41], or the Look-Locker readout[42].
Diffusion-weighted model: In DWI experiments, the simple Gaussian diffusion can be assumed using a mono-exponential model. The two-compartment intravoxel incoherent motion model on the other hand is currently widely used in clinical pancreatitis and pancreatic tumor studies[43,44], and separates diffusion into the true-diffusion and the pseudo-diffusion fraction. Alternatively, sampling with high b-values above 1000 s/mm2 can be applied for non-Gaussian diffusion estimation using a diffusion kurtosis model[45].
Post-processing for DCE model: The first step in DCE data post-processing is the conversion of the raw MRI signal to the tissue concentration time curve (CTC). The tissue concentration Ct of contrast agent (CA) during the DCE perfusion experiment is solved as: 1/T1(t) = 1/T10 + r1 × Ct(t), where T10 is the T1 value before contrast injection, obtained from inversion recovery T1 mapping, and r1 is the longitudinal relativity of the applied CA. In a high temporal resolution DCE experiment, the T1 relaxation T1(t) after CA injection can be converted from the signal intensity (SI) time curve as described previously[33]. Alternatively, CTC information can be directly extracted from the dynamic T1 mapping.
The vascular input function (VIF) Cp is determined by the CA concentration in blood Cb: Cp = Cb/(1 - Hct), which is obtained from CTC of the aorta or a major vein, and the hematocrit level Hct which is set to 42% in our studies. VIF is usually fitted into a bi-exponential function for further kinetic modeling. Perfusion indices, the transfer coefficient Ktrans and the rate constant kep, can be obtained from the standard or the modified Tofts model[46]. In practice, the discrete convolution can be constructed as a matrix multiplication. The fraction volume ve of extravascular extracellular space is calculated as: ve = Ktrans/kep.
The diagnosis of pancreatic diseases and their management have been largely facilitated by ever advancing multimodal and multi-parametric imaging technologies in clinical settings. Likewise, thanks to the above-mentioned efforts, preclinical research on rodent models of pancreatic pathologies are also rapidly progressing in terms of visual identification of rodent pancreas on 2D and 3D images, imaging characterization of common pancreatic disorders such as pancreatitis and pancreatic malignancies, and noninvasive imaging follow-up of investigative therapies for new drug development. Eventually clinical practices in patients suffering from those often deadly diseases on this complex visceral organ of pancreas will benefit from all these translational studies.
Manuscript source: Unsolicited manuscript
Specialty type: Medical laboratory technology
Country of origin: Belgium
Peer-review report classification
Grade A (Excellent): A, A
Grade B (Very good): 0
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Barauskas G, Weng CF S- Editor: Ji FF L- Editor: A E- Editor: Li D
| 1. | Nakamura T, Takeuchi T, Tando Y. Pancreatic dysfunction and treatment options. Pancreas. 1998;16:329-336. [PubMed] |
| 2. | Cascinu S, Falconi M, Valentini V, Jelic S; ESMO Guidelines Working Group. Pancreatic cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2010;21 Suppl 5:v55-v58. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 107] [Cited by in RCA: 127] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 3. | Weyer C, Tataranni PA, Bogardus C, Pratley RE. Insulin resistance and insulin secretory dysfunction are independent predictors of worsening of glucose tolerance during each stage of type 2 diabetes development. Diabetes Care. 2001;24:89-94. [PubMed] |
| 4. | Souza F, Freeby M, Hultman K, Simpson N, Herron A, Witkowsky P, Liu E, Maffei A, Harris PE. Current progress in non-invasive imaging of beta cell mass of the endocrine pancreas. Curr Med Chem. 2006;13:2761-2773. [PubMed] |
| 5. | Dimastromatteo J, Brentnall T, Kelly KA. Imaging in pancreatic disease. Nat Rev Gastroenterol Hepatol. 2017;14:97-109. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 58] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 6. | Whitcomb DC. Clinical practice. Acute pancreatitis. N Engl J Med. 2006;354:2142-2150. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 549] [Cited by in RCA: 518] [Article Influence: 27.3] [Reference Citation Analysis (1)] |
| 7. | Tummala P, Junaidi O, Agarwal B. Imaging of pancreatic cancer: An overview. J Gastrointest Oncol. 2011;2:168-174. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 42] [Reference Citation Analysis (0)] |
| 8. | Garrido-Laguna I, Hidalgo M. Pancreatic cancer: from state-of-the-art treatments to promising novel therapies. Nat Rev Clin Oncol. 2015;12:319-334. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 398] [Cited by in RCA: 444] [Article Influence: 44.4] [Reference Citation Analysis (0)] |
| 9. | Coté GA, Smith J, Sherman S, Kelly K. Technologies for imaging the normal and diseased pancreas. Gastroenterology. 2013;144:1262-1271.e1. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 33] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 10. | Erchinger FG, Dimcevski G, Engjom T, Gilja OH. Transabdominal ultrasonography of the pancreas: basic and new aspects. Imaging Med. 2011;3:412-422. [RCA] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 11] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 11. | Dimcevski G, Erchinger FG, Havre R, Gilja OH. Ultrasonography in diagnosing chronic pancreatitis: new aspects. World J Gastroenterol. 2013;19:7247-7257. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 31] [Cited by in RCA: 31] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 12. | Poruk KE, Firpo MA, Adler DG, Mulvihill SJ. Screening for pancreatic cancer: why, how, and who? Ann Surg. 2013;257:17-26. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 143] [Cited by in RCA: 185] [Article Influence: 15.4] [Reference Citation Analysis (0)] |
| 13. | Working Party of the British Society of Gastroenterology. Association of Surgeons of Great Britain and Ireland; Pancreatic Society of Great Britain and Ireland; Association of Upper GI Surgeons of Great Britain and Ireland. UK guidelines for the management of acute pancreatitis. Gut. 2005;54 Suppl 3:iii1-iii9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 184] [Cited by in RCA: 330] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 14. | Sandrasegaran K, Lin C, Akisik FM, Tann M. State-of-the-art pancreatic MRI. AJR Am J Roentgenol. 2010;195:42-53. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 15. | Matos C, Cappeliez O, Winant C, Coppens E, Devière J, Metens T. MR imaging of the pancreas: a pictorial tour. Radiographics. 2002;22:e2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 56] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 16. | Mandelia A, Gupta AK, Verma DK, Sharma S. The Value of Magnetic Resonance Cholangio-Pancreatography (MRCP) in the Detection of Choledocholithiasis. J Clin Diagn Res. 2013;7:1941-1945. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 12] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 17. | Kim JH, Lee JM, Park JH, Kim SC, Joo I, Han JK, Choi BI. Solid pancreatic lesions: characterization by using timing bolus dynamic contrast-enhanced MR imaging assessment--a preliminary study. Radiology. 2013;266:185-196. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 62] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 18. | Coenegrachts K, Van Steenbergen W, De Keyzer F, Vanbeckevoort D, Bielen D, Chen F, Dockx S, Maes F, Bosmans H. Dynamic contrast-enhanced MRI of the pancreas: initial results in healthy volunteers and patients with chronic pancreatitis. J Magn Reson Imaging. 2004;20:990-997. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 47] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 19. | Barral M, Taouli B, Guiu B, Koh DM, Luciani A, Manfredi R, Vilgrain V, Hoeffel C, Kanematsu M, Soyer P. Diffusion-weighted MR imaging of the pancreas: current status and recommendations. Radiology. 2015;274:45-63. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 144] [Cited by in RCA: 147] [Article Influence: 14.7] [Reference Citation Analysis (0)] |
| 20. | Akladios CY, Bour G, Raykov Z, Mutter D, Marescaux J AM. Structural imaging of the pancreas in rat using micro-CT: application to a non-invasive longitudinal evaluation of pancreatic ductal carcinoma monitoring. J Cancer Res Ther. 2013;1:70-76. [RCA] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 21. | Tchokonte-Nana V, Longo-Mbenza B, Page BJ, Du Toit DF. Morphogenetic and clinical perspectives on the neogenesis of pancreatic duct ligation-induced islet cells : a review. Adv Clin Exp Med. 2011;20:5-14. |
| 22. | Yin T, Coudyzer W, Peeters R, Liu Y, Cona MM, Feng Y, Xia Q, Yu J, Jiang Y, Dymarkowski S. Three-dimensional contrasted visualization of pancreas in rats using clinical MRI and CT scanners. Contrast Media Mol Imaging. 2015;10:379-387. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 11] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 23. | Zhang JX, Dang SC, Zhang Y, Sha X, Zhang LR, Wei CS, Chen M, Jiang DL. MRI shows clodronate-liposomes attenuating liver injury in rats with severe acute pancreatitis. Hepatobiliary Pancreat Dis Int. 2010;9:192-200. [PubMed] |
| 24. | Santhakumari R, Reddy IY, Archana R. Effect of type 2 diabetes mellitus on brain metabolites by using proton magnetic resonance spectroscopy-a systematic review. Int J Pharma Bio Sci. 2014;5:1118-1123. [PubMed] |
| 25. | Wang P, Yoo B, Yang J, Zhang X, Ross A, Pantazopoulos P, Dai G, Moore A. GLP-1R-targeting magnetic nanoparticles for pancreatic islet imaging. Diabetes. 2014;63:1465-1474. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 51] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 26. | Smits MM, Tonneijck L, Muskiet MH, Kramer MH, Diamant M, Pieters-van den Bos IC, van Raalte DH, Cahen DL. Glucagon-like peptide-1 receptor agonist exenatide has no acute effect on MRI-measured exocrine pancreatic function in patients with type 2 diabetes: a randomized trial. Diabetes Obes Metab. 2016;18:281-288. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 27. | Medarova Z, Castillo G, Dai G, Bolotin E, Bogdanov A, Moore A. Noninvasive magnetic resonance imaging of microvascular changes in type 1 diabetes. Diabetes. 2007;56:2677-2682. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 39] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 28. | Medarova Z, Greiner DL, Ifediba M, Dai G, Bolotin E, Castillo G, Bogdanov A, Kumar M, Moore A. Imaging the pancreatic vasculature in diabetes models. Diabetes Metab Res Rev. 2011;27:767-772. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 11] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 29. | Paajanen H, Brasch RC, Dean PB. Experimental acute pancreatitis: MR relaxation time studies using gadolinium-DTPA. Magn Reson Med. 1988;6:63-73. [PubMed] |
| 30. | Kushnir T, Kaplan O, Askenasy N, Navon G. Identification of a characteristic 31P NMR signal in acute experimental pancreatitis with the aid of 1H-31P correlated 2D measurements of intact pancreas. Magn Reson Med. 1989;10:119-124. [PubMed] |
| 31. | Zhang HW, Wang LQ, Xiang QF, Zhong Q, Chen LM, Xu CX, Xiang XH, Xu B, Meng F, Wan YQ. Specific lipase-responsive polymer-coated gadolinium nanoparticles for MR imaging of early acute pancreatitis. Biomaterials. 2014;35:356-367. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 31] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 32. | Yin T, Peeters R, Liu Y, Feng Y, Zhang X, Jiang Y, Yu J, Dymarkowski S, Himmelreich U, Oyen R. Visualization, Quantification and Characterization of Caerulein-Induced Acute Pancreatitis in Rats by 3.0T Clinical MRI, Biochemistry and Histomorphology. Theranostics. 2017;7:285-294. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 13] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 33. | Yin T, Peeters R, Feng Y, Liu Y, Yu J, Dymarkowski S, Himmelreich U, Oyen R, Ni Y. Characterization of a rat orthotopic pancreatic head tumor model using three-dimensional and quantitative multi-parametric MRI. NMR Biomed. 2017;30:e3637. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 14] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 34. | Herrmann KH, Schmidt S, Kretz A, Haenold R, Krumbein I, Metzler M, Gaser C, Witte OW, Reichenbach JR. Possibilities and limitations for high resolution small animal MRI on a clinical whole-body 3T scanner. MAGMA. 2012;25:233-244. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 40] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 35. | Jambor I, Pesola M, Merisaari H, Taimen P, Boström PJ, Liimatainen T, Aronen HJ. Relaxation along fictitious field, diffusion-weighted imaging, and T2 mapping of prostate cancer: Prediction of cancer aggressiveness. Magn Reson Med. 2016;75:2130-2140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 15] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 36. | Duan C, Kallehauge JF, Bretthorst GL, Tanderup K, Ackerman JJ, Garbow JR. Are complex DCE-MRI models supported by clinical data? Magn Reson Med. 2017;77:1329-1339. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 33] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 37. | Wurnig MC, Kenkel D, Filli L, Boss A. A Standardized Parameter-Free Algorithm for Combined Intravoxel Incoherent Motion and Diffusion Kurtosis Analysis of Diffusion Imaging Data. Invest Radiol. 2016;51:203-210. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 14] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 38. | Dortch RD, Yankeelov TE, Yue Z, Quarles CC, Gore JC, Does MD. Evidence of multiexponential T2 in rat glioblastoma. NMR Biomed. 2009;22:609-618. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 9] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 39. | Schmitt P, Griswold MA, Jakob PM, Kotas M, Gulani V, Flentje M, Haase A. Inversion recovery TrueFISP: quantification of T(1), T(2), and spin density. Magn Reson Med. 2004;51:661-667. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 191] [Cited by in RCA: 198] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 40. | Cheng HL, Stikov N, Ghugre NR, Wright GA. Practical medical applications of quantitative MR relaxometry. J Magn Reson Imaging. 2012;36:805-824. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 174] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 41. | Deichmann R, Hahn D, Haase A. Fast T1 mapping on a whole-body scanner. Magn Reson Med. 1999;42:206-209. [PubMed] |
| 42. | Cooper MA, Nguyen TD, Spincemaille P, Prince MR, Weinsaft JW, Wang Y. How accurate is MOLLI T1 mapping in vivo? Validation by spin echo methods. PLoS One. 2014;9:e107327. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 14] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 43. | Kuru TH, Roethke MC, Stieltjes B, Maier-Hein K, Schlemmer HP, Hadaschik BA, Fenchel M. Intravoxel incoherent motion (IVIM) diffusion imaging in prostate cancer - what does it add? J Comput Assist Tomogr. 2014;38:558-564. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 38] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 44. | Kang KM, Lee JM, Yoon JH, Kiefer B, Han JK, Choi BI. Intravoxel incoherent motion diffusion-weighted MR imaging for characterization of focal pancreatic lesions. Radiology. 2014;270:444-453. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 134] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 45. | Rosenkrantz AB, Padhani AR, Chenevert TL, Koh DM, De Keyzer F, Taouli B, Le Bihan D. Body diffusion kurtosis imaging: Basic principles, applications, and considerations for clinical practice. J Magn Reson Imaging. 2015;42:1190-1202. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 258] [Cited by in RCA: 290] [Article Influence: 29.0] [Reference Citation Analysis (0)] |
| 46. | Tofts PS. T1-weighted DCE Imaging Concepts: Modelling, Acquisition and Analysis. MAGNETOM Flash, 2010: 1-5. . |