Published online May 20, 2022. doi: 10.5662/wjm.v12.i3.107
Peer-review started: October 26, 2021
First decision: December 17, 2021
Revised: December 18, 2021
Accepted: March 16, 2021
Article in press: March 16, 2021
Published online: May 20, 2022
Processing time: 204 Days and 6.2 Hours
Lutetium has been shown to be an important potential innovation in pre-treated metastatic castration-resistant prostate cancer. Two clinical trials have evaluated lutetium thus far (therap and vision with 99 and 385 patients, respectively), but their results are discordant.
To synthetize the available evidence on the effectiveness of lutetium in pre-treated metastatic castration-resistant prostate cancer; and to test the application of a new artificial intelligence technique that synthetizes effectiveness based on reconstructed patient-level data.
We employed a new artificial intelligence method (shiny method) to pool the survival data of these two trials and evaluate to what extent the lutetium cohorts differed from one another. The shiny technique employs an original reconstruction of individual patient data from the Kaplan-Meier curves. The progression-free survival graphs of the two lutetium cohorts were analyzed and compared.
The hazard ratio estimated was in favor of the vision trial; the difference was statistically significant (P < 0.001). These results indicate that further studies on lutetium are needed because the survival data of the two trials published thus far are conflicting.
Our study confirms the feasibility of reconstructing patient-level data from survival graphs in order to generate a survival statistics.
Core Tip: This paper describes the application of a new technique of individual-patient data reconstruction to the progression-free survival curves published in two trials evaluating lutetium in metastatic prostate cancer. Our analysis interpreted these survival data and showed discordant results between the two trials, that need to be addressed by further clinical research.
- Citation: Messori A. Lutetium in prostate cancer: Reconstruction of patient-level data from published trials and generation of a multi-trial Kaplan-Meier curve. World J Methodol 2022; 12(3): 107-112
- URL: https://www.wjgnet.com/2222-0682/full/v12/i3/107.htm
- DOI: https://dx.doi.org/10.5662/wjm.v12.i3.107
Lutetium has been shown to be an important potential innovation in pre-treated metastatic prostate cancer, but the extent to which outcomes are improved by this treatment still needs to a fully investigated. Three studies have evaluated lutetium in this disease condition. One was phase II (therap trial[1]), the second was phase III (vision trial[2]); the third, which was an observational real-world study[3], differed from the first two because lutetium was given after radium-223.
In recent times, techniques that reconstruct individual patient data from the graphs of Kaplan-Meier curves have considerably improved in terms of performance and easy applicability[4]. One advantage is that the availability of these techniques permits to combine multiple survival curves published in different trials without using any meta-analytical statistics. An example of this approach is presented herein. Our objective was two-fold: 1) to quantify the gain in progression-free survival determined by lutetium: 2) to demonstrate the applicability of techniques of patient-level data reconstruction in addressing specific questions based on time-to-event endpoints without the need to employ any meta-analytic statistics.
We applied the shiny technique of individual patient data reconstruction[4] to the Kaplan-Meier graphs of progression-free survival reported in the therap phase-II trial[1] and in the vision phase III trial[2]. Both trials were conducted in patients with metastatic castration-resistant prostate cancer previously treated for their metastatic disease. In the therap trial, the treatment group received Lu-PSMA-617 (6·0–8·5 GBq intravenously every 6 wk for up to six cycles) while the controls were given cabazitaxel (20 mg/m² intravenously every 3 wk for up to ten cycles). In the vision trial, the treatment group received 177Lu-PSMA-617 (7.4 GBq every 6 wk for four to six cycles) plus protocol-permitted standard care while the controls received standard care alone. In the therap trial, progression-free survival was defined as the interval from randomisation to first evidence of pupil-size artefact progression defined by an increase of at least 25% and at least 2 ng/mL after 12 wk (as per PCWG316), radiographic progression using locally reported computed tomography and bone scanning [Response Evaluation Criteria In Solid Tumors (RECIST) 1.1 and PCWG3 criteria for bone lesions], commencement of non-protocol anticancer treatment, or death from any cause. In the vision trial, the end-point was image based progression free survival.
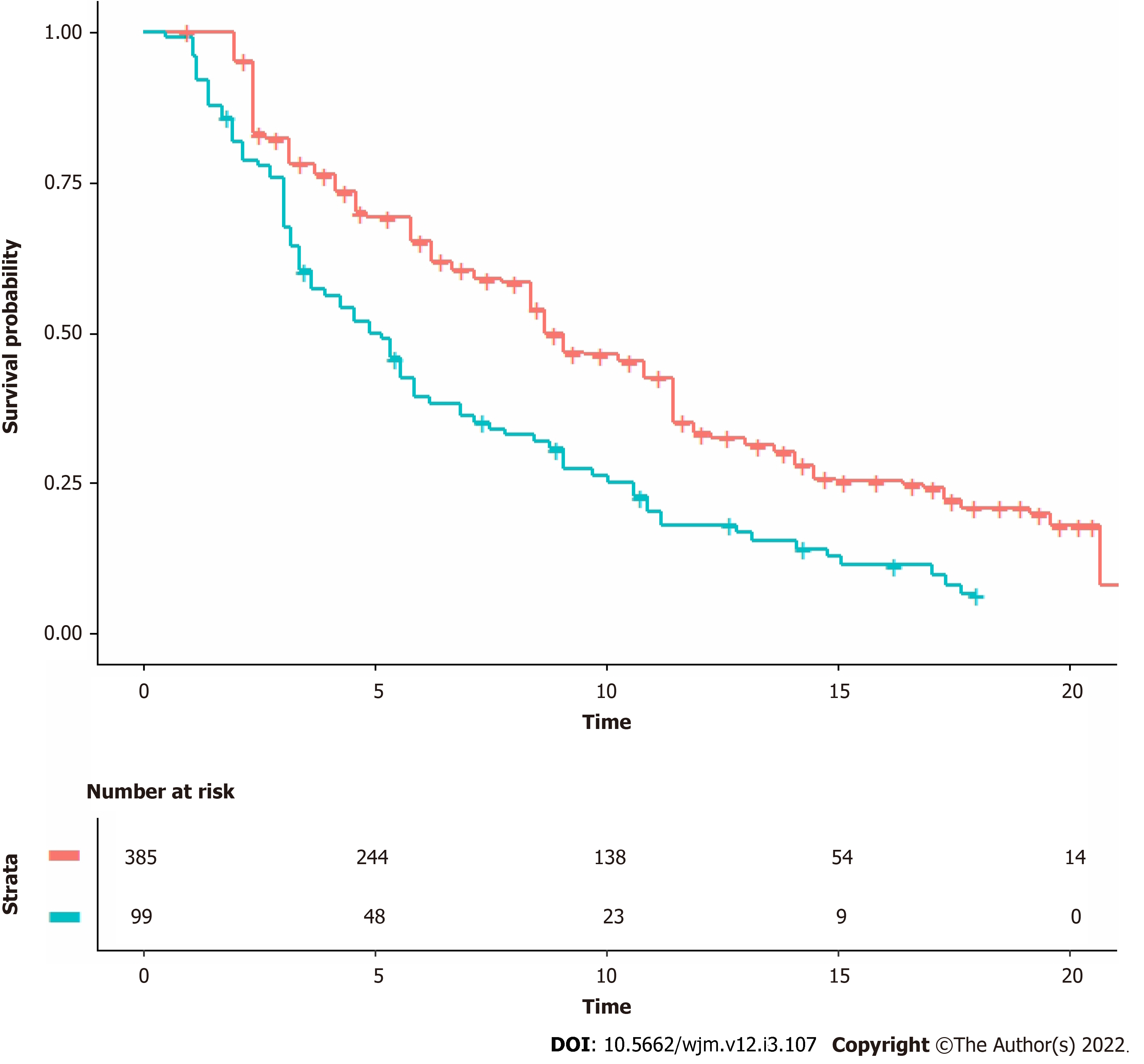
The progression-free survival graphs of the two lutenium cohorts by Hofman et al[1] for the therap trial (99 patients; follow-up of 18 mo; 90 events) and Sartor et al[2] for the vision trial (385 patients; follow-up of 30 mo; 254 events). For each of these two Kaplan-Meier curves, the graph was digitalized and converted into x-y data pairs using Webplotdigitizer (version 4.5, https://apps.automeris.io/wpd/). Then, the shiny package (version: 1.2.2.0; subprogram “Reconstruct Individual Patient Data”; https://www.trialdesign.org/one-page-shell.html#IPDfromKM, see reference[4]) was used to reconstruct patient-level data on the basis of x-y data pairs, total number of enrolled patients, and total number of events. Finally, the pooled survival curves were generated from the reconstructed patient-level data and analyzed through standard Cox statistics. For this purpose we used three packages (“coxph”, “survfit”, and “ggsurvplot”) under the R-platform. The hazard ratio (HR) was estimated.
The shiny procedure combined with standard Kaplan-Meier statistics allowed us to compare the 99 patients given lutetium in the therap trial with the 385 patients given lutetium in the vision trial.
Figure 1 shows the two Kaplan-Meier curves generated from reconstructed patient-level data. The HR estimated from these curves favored the patients of the vision trial and was 0.59 (95%CI, 0.46 to 0.75). The difference was statistically significant (P < 0.001).
When two or more randomised trials are available on a therapeutic issue and the clinical end-point is the form of time-to-event, synthetising the clinical evidence is a complex issue, and there is presently no consensus on which methodological approach should be preferred[5,6]. Pooling the values of HR is certainly the method most commonly used, but its important limitations have been widely recognised for many years (e.g. the inability to account for the length of follow-up, the inability to model variations of risk over time, the dimensionless nature of HR as opposed to the greater informative value of absolute parameters such as medians, etc.)[8]. The development of the restricted mean survival time has represented an advancement in this field[8,9], but the use of this parameter unfortunately remains low.
In this context, the marked improvement in performance of techniques that reconstruct individual-patient data[4] represents an important innovation, the role of which still needs to be fully evaluated. On the one hand, reconstructing individual-patient data is a mandatory pre-requisite to determine the RMST, and this explains the increased use of these reconstruction techniques when a single trial needs to be analysed[7]. On the other hand, another potential use of these techniques is being recognised when multiple trials are available: in such cases, these techniques offer a new methodological alternative to standard meta-analytic methods[5,6] and also to the more recent approaches where meta-analysis is based on the use of RMSTs[8,9].
The various parameters mentioned above (especially HR, RMST, and median) have been investigated for many years to identify their respective advantages and disadvantages, and the literature on this issue is wide[7]. In contrast, the literature on the use of reconstructed survival curves is still in its early stages[4,6], and this holds true particularly when multiple trials are analysed and pooled together.
The experience described herein offers a limited but useful contribution to the development of meta-analysis-like methods based on reconstructed survival curves.
The two control groups of the two trials differed in the treatment they received, and so were not included in our analysis, which was focused only on the two lutetium groups of the two trials. In comparing these two group with one another, our results raise the need to explain the statistically different outcomes shown by the HR and presented in Figure 1.
The inclusion criteria of the therap and vision trials were very similar, and so they likely had no substantial role in determining this difference. In fact, in the therap trial, patients had metastatic castration-resistant cancer and PET eligibility criteria for the trial were PSMA-positive disease, and no sites of metastatic disease with discordant FDG-positive and PSMA-negative findings; previous treatment with androgen receptor-directed therapy was allowed. In the vision trial, patients had metastatic castration-resistant prostate cancer previously treated with at least one androgen-receptor–pathway inhibitor and one or two taxane regimens and had PSMA-positive gallium-68 (68Ga)–labeled PSMA-11 and PET scans. While these differences in the inclusion criteria do not seem to suggest a better prognosis for patients included in either trial, a number of factors (e.g. environmental and lifestyle factors, tissue biomarkers, molecular pathological epidemiology, the microbiota, etc.) might have influenced tumor development and response to therapy. Hence, the discrepancies observed across the two trials included in our analysis might be explained by these factors. As regards innovative treatments such as lutetium, it should be stressed that molecular pathological epidemiology research has a growing role and is increasingly recognized to be a promising strategy to improve prediction of response to therapy.
In summary, the main strength of our analysis lies in the originality of the methodological approach that reflects the recent availability of very efficient patient data reconstruction techniques. The main limitation is represented by the indirect nature of the comparison between the two lutetium cohorts.
Our study indicates that further studies on Lu-PSMA-617 are needed because the survival data of the two trials published thus far demonstrate quite conflicting results. The example described in this paper confirms the feasibility of reconstructing patient-level data from survival graphs in order to generate a survival statistics from these reconstructed data. To evaluate the advantages and disadvantages of this new methodological approach, further analyses will be needed.
Two trials have been published to assess the effectiveness of lutetium in metastatic prostate cancer. The need to convert these effectiveness data into a pooled estimate represents a useful opportunity to test an innovative technique of individual patient reconstruction based on the analysis of Kaplan-Meier curves (shiny method).
The main motivation was to test the performance of the shiny method based on a real data-set.
Clarifying the effectiveness of lutetium in metastatic prostate cancer and confirm the reliability of the shiny method as a tool for reconstructing individual patient data.
The clinical trials that have thus far evaluated lutetium in metastatic prostate cancer have been identified by standard literature search. A pooled survival curve has been generated from these trials by using the shiny technique of individual patient data reconstruction.
Two clinical trials were identified. A pooled Kaplan-Meier survival curve was generated that synthesizes the current evidence on the effectiveness of this treatment in this disease condition.
A two-fold conclusion: First, lutetium is effective in metastatic prostate cancer; second, the Shiny technique can successfully be used to pool survival data from two trials without employing any meta-analytical method.
The shiny technique has been confirmed to be a useful new tool for analyzing survival data from multiple trials and therefore deserves to be further applied in the analysis of clinical evidence.
Provenance and peer review: Unsolicited article; Externally peer reviewed.
Peer-review model: Single blind
Corresponding Author's Membership in Professional Societies: Sifact
Specialty type: Microscopy
Country/Territory of origin: Italy
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Liang L, China; Ogino S, United States S-Editor: Xing YX L-Editor: A P-Editor: Xing YX
| 1. | Hofman MS, Emmett L, Sandhu S, Iravani A, Joshua AM, Goh JC, Pattison DA, Tan TH, Kirkwood ID, Ng S, Francis RJ, Gedye C, Rutherford NK, Weickhardt A, Scott AM, Lee ST, Kwan EM, Azad AA, Ramdave S, Redfern AD, Macdonald W, Guminski A, Hsiao E, Chua W, Lin P, Zhang AY, McJannett MM, Stockler MR, Violet JA, Williams SG, Martin AJ, Davis ID; TheraP Trial Investigators and the Australian and New Zealand Urogenital and Prostate Cancer Trials Group. [177Lu]Lu-PSMA-617 vs cabazitaxel in patients with metastatic castration-resistant prostate cancer (TheraP): a randomised, open-label, phase 2 trial. Lancet. 2021;397:797-804. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 341] [Cited by in RCA: 667] [Article Influence: 166.8] [Reference Citation Analysis (0)] |
| 2. | Sartor O, de Bono J, Chi KN, Fizazi K, Herrmann K, Rahbar K, Tagawa ST, Nordquist LT, Vaishampayan N, El-Haddad G, Park CH, Beer TM, Armour A, Pérez-Contreras WJ, DeSilvio M, Kpamegan E, Gericke G, Messmann RA, Morris MJ, Krause BJ; VISION Investigators. Lutetium-177-PSMA-617 for Metastatic Castration-Resistant Prostate Cancer. N Engl J Med. 2021;385:1091-1103. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 548] [Cited by in RCA: 1518] [Article Influence: 379.5] [Reference Citation Analysis (0)] |
| 3. | Sartor AO, la Fougère C, Essler M, Ezziddin S, Kramer G, Elllinger J, Nordquist L, Sylvester J, Paganelli G, Peer A, Bögemann M, Meltzer J, Sandström P, Verholen F, Song DY. Lutetium-177-prostate-specific membrane antigen ligand following radium-223 treatment in men with bone-metastatic castration-resistant prostate cancer: real-world clinical experience. J Nucl Med. 2021;. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 20] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 4. | Liu N, Zhou Y, Lee JJ. IPDfromKM: reconstruct individual patient data from published Kaplan-Meier survival curves. BMC Med Res Methodol. 2021;21:111. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 392] [Cited by in RCA: 390] [Article Influence: 97.5] [Reference Citation Analysis (0)] |
| 5. | Batson S, Greenall G, Hudson P. Review of the Reporting of Survival Analyses within Randomised Controlled Trials and the Implications for Meta-Analysis. PLoS One. 2016;11:e0154870. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 18] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 6. | Messori A, Trippoli S, Vaiani M. Survival Meta-Analysis of Individual Patient Data and Survival Meta-Analysis of Published (Aggregate) Data. Clin Drug Investig. 2000;20: 309-316. |
| 7. | Messori A. The advantages of restricted mean survival time in analysing Kaplan-Meier survival curves: analysis of 55 articles published in the last 12 mo (preprint). Open Science Framework, 2021. |
| 8. | Messori A, Bartoli L, Trippoli S. The restricted mean survival time as a replacement for the hazard ratio and the number needed to treat in long-term studies. ESC Heart Fail. 2021;8:2345-2348. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 3] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 9. | Messori A, Bartoli L, Ferracane E, Trippoli S. Medical therapy, radiofrequency ablation or cryoballoon ablation as first-line treatment for paroxysmal atrial fibrillation: interpreting efficacy through restricted mean survival time and network meta-analysis. Rev Cardiovasc Med. 2021;22:557-561. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 1.3] [Reference Citation Analysis (0)] |