Published online Nov 5, 2021. doi: 10.5495/wjcid.v11.i3.49
Peer-review started: February 27, 2021
First decision: March 31, 2021
Revised: April 3, 2021
Accepted: September 10, 2021
Article in press: September 10, 2021
Published online: November 5, 2021
Processing time: 247 Days and 20.6 Hours
In December 2019, coronavirus disease 2019 (COVID-19) was reported firstly in Wuhan, China. COVID-19 is currently a global pandemic.
To assess the suitability of lymphocyte count as a biomarker of COVID-19 severity.
Five literature databases (PubMed/MEDLINE, Web of Science, Google Scholar, Embase, and Scopus) were searched to identify eligible articles. A meta-analysis was performed to calculate the standard mean difference (SMD) and 95% confidence interval (CI) of lymphocyte counts in coronaviral pneumonia cases.
Eight studies, including 1057 patients, were integrated in the meta-analysis. Lymphocyte counts were associated with severe coronavirus (CoV) infection (SMD = 1.35, 95%CI: 1.97 to 0.37, P < 0.001, I2 = 92.6%). In the subgroup analysis stratified by prognosis, lymphocytes were associated with CoV infection mortality (n = 2, SMD = 0.42, 95%CI: 0.66 to 0.19, P < 0.001, I2 = 0.0%), severity (n = 2, SMD = 0.93, 95%CI: 1.20 to 0.67, P < 0.001, I2 = 0.0%), and diagnostic rate (n = 4, SMD = 2.32, 95%CI: 3.60 to 1.04, P < 0.001, I2 = 91.2%).
Lymphocyte count may represent a simple, rapid, and commonly available laboratory index with which to diagnosis infection and predict the severity of CoV infections, including COVID-19.
Core tip: Lymphocyte count reflects immune function and inflammatory state in infectious disease. Severe acute respiratory syndrome coronavirus 2 spreads and invades through respiratory mucosa, triggers a series of immune responses and induces a cytokine storm, resulting in changes in immune components such as lymphocytes. Previous studies have shown that the decrease in lymphocyte count can be used as an indicator of severity for both severe acute respiratory syndrome and Middle East respiratory syndrome; both of which are coronavirus (CoV) infections. Therefore, this systematic review and meta-analysis evaluate the diagnostic and prognostic utility of the lymphocyte count in patients with viral pneumonia by CoV infections.
- Citation: Zhao YS, Yu YX. Lymphocyte count predicts the severity of COVID-19: Evidence from a meta-analysis. World J Clin Infect Dis 2021; 11(3): 49-59
- URL: https://www.wjgnet.com/2220-3176/full/v11/i3/49.htm
- DOI: https://dx.doi.org/10.5495/wjcid.v11.i3.49
In December 2019, in Wuhan, China, a novel CoV was identified as the causative agent of a novel pneumonia. The disease and the virus were subsequently termed coron
Lymphocytes play a decisive role in the maintenance of immune homeostasis and the inflammatory response[8]. During the progression of an infectious disease, lymphocyte count reflects immune function and inflammatory state. SARS-CoV-2 invades via the respiratory mucosa, triggering a series of immune responses, including a cytokine storm, and affecting immune components, such as peripheral blood leukocytes and lymphocytes[9]. Consistent with this, studies have shown that lymphopenia, particularly the depletion of CD4 and CD8 lymphocytes, is a clinical characteristic of COVID-19 patients, especially those with severe infections[10-12]. Indeed, previous studies have shown that the decrease in lymphocyte count can be used to indicate the severity of SARS and MERS, both of which are CoV infections[13,14]. However, due to the lack of analytical data, as might be provided by case–control or cohort studies, it is unclear whether lymphocyte count can also be used to reflect COVID-19 severity. Although Tan et al[15] had reported that lymphocyte count predicted the severity of COVID-19, this study had too few samples. However, relying on the association of CoV infection, we found some evidence in a small number of observational studies of COVID-19. Here, we performed a systematic review and meta-analysis to evaluate the diagnostic and prognostic utility of lymphocyte count in patients with viral pneumonia caused by CoV infections. Our aim was to explore the possibility that lymphocyte counts can predict COVID-19 severity and provide associated evidence.
This protocol was registered with the PROSPERO international prospective register of systematic reviews (CRD42020177132, available from: https://www.crd.york.ac.uk/prospero/display_record.php?RecordID=177132) and followed the recommendations established by the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement[16]. We conducted a systematic review across five literature databases: PubMed/MEDLINE, Web of Science, Google Scholar, Embase, and Scopus. The search terms used were as follows: “lymphocyte count,” and “pneumonia, viral” or “SARS” or “MERS” or “COVID-19”. All of the searches were concluded by March 21, 2020, and two researchers independently evaluated the search results.
We included published peer-reviewed articles describing case–control, cohort, or cross-sectional studies, where lymphocyte counts were measured in peripheral blood samples from humans of any age with CoV infections. Case reports, conference abstracts, and review articles were excluded.
COVID-19 infections were diagnosed using next-generation sequencing or real-time reverse transcription polymerase chain reactions (RT-PCRs)[17]. MERS-CoV was diagnosed according to the WHO criteria: a confirmed case was defined as a suspected case that was positive for MERS-CoV based on RT-PCR results[18]. SARS infections were confirmed based on a definite exposure history, as well as either a positive RT-PCR test during acute infection or detectable CoV-specific antibodies during convalescence[19]. We excluded studies conducted exclusively in patients with active cancer, chronic liver disease, HIV, or immunosuppression. When an article reported duplicate information from the same patient, the reports were combined to obtain complete data, but the case was only counted once.
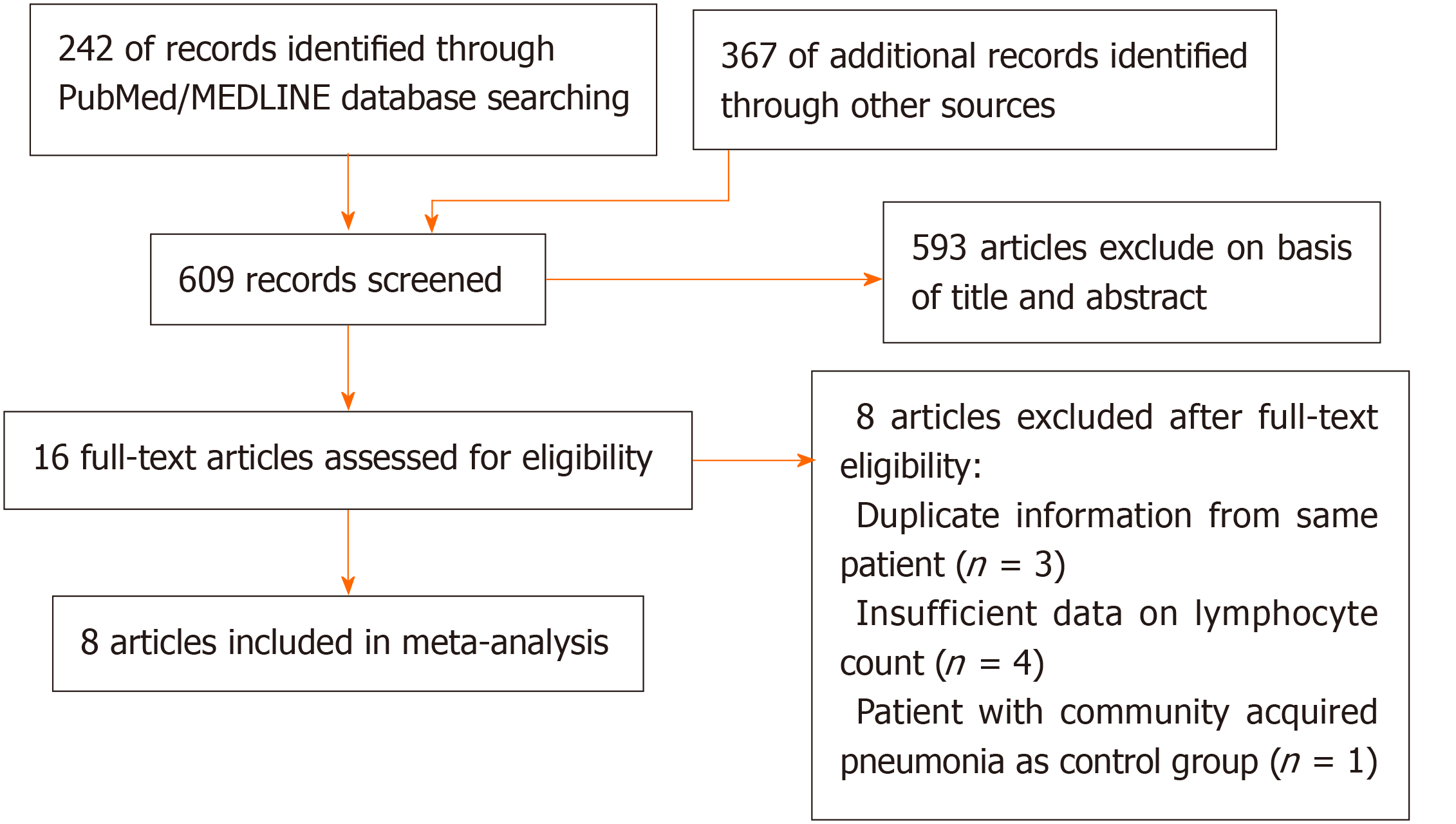
All of the titles and abstracts returned by the database search (Figure 1) were reviewed by first author (Zhao YS) independently to assess the need for a full-text review. Any disagreements were resolved through discussion between the same author and another author (Yu YX). Reasons for exclusion were recorded.
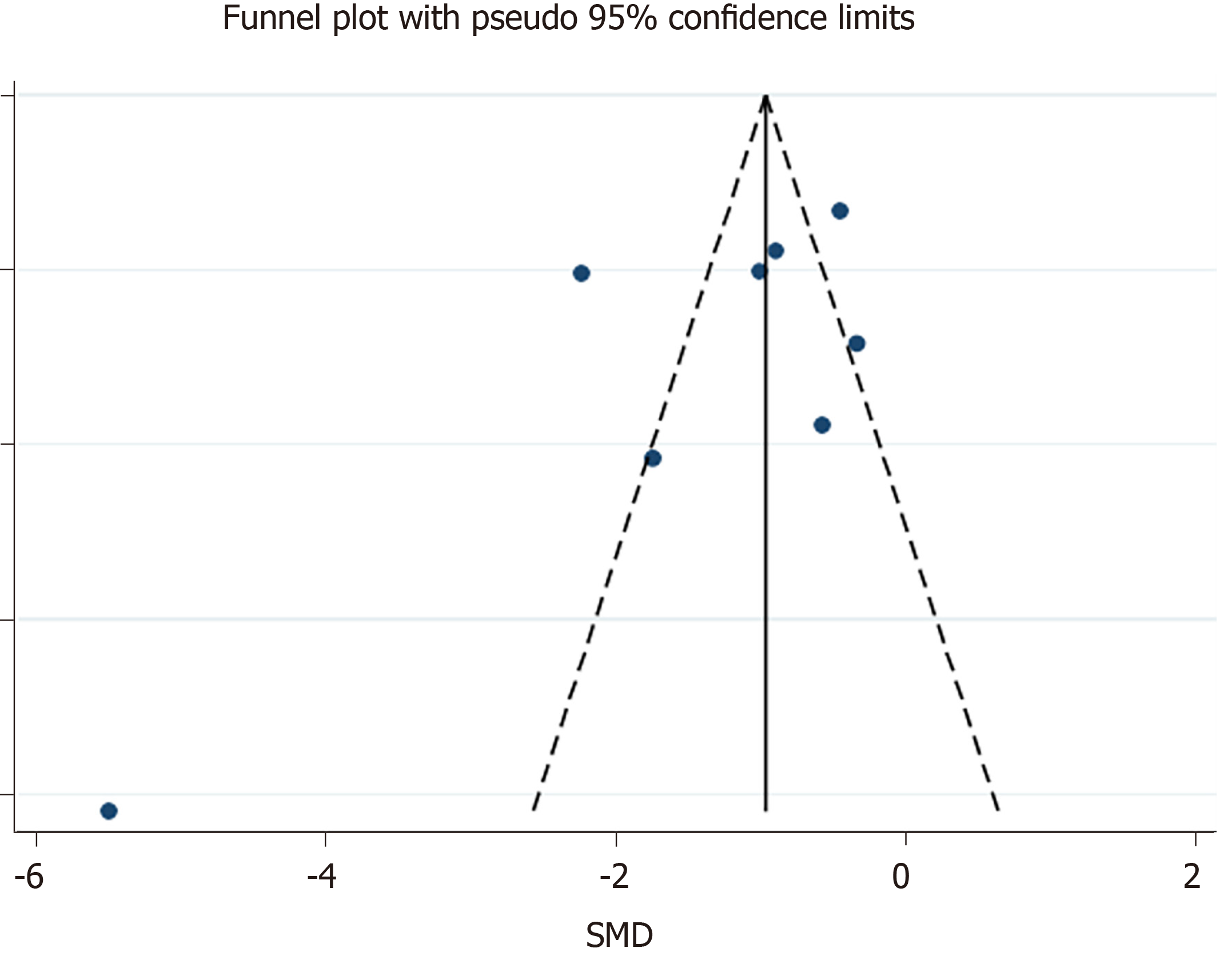
Publication bias was assessed using a funnel plot, Begg’s test and Egger’s test.
According to a study by Hozo et al[20], we estimating the mean and standard deviation from the median and range, and the size of a sample as data extrapolation. When sample size was small (n < 25), we used a simple formula: x = (a + 2m + b)/4 to estimate the mean (x) using the values of the median (m), low and high end of the range (a and b, respectively). As soon as sample size exceeded 25, the median itself was the best estimator. When sample size was small (n < 15), we used the formula: S2 = 1/12 × (((a - 2m + b)2)/4 + (b - a) 2) to estimate the standard deviation (S2). When the sample size increased (15 < n < 70), Range/4 was the best estimator for S2. For large samples (n > 70) Range/6 was the best estimator for S2[20]. To indicate the severity of CoV infection, we designed a cohort that combined three different measures of prognosis with respect to the control group: diagnosed vs nondiagnosed, severe vs nonsevere, and death vs survival. The control group included nondiagnosed, nonsevere and survival groups. The case group was severe, including diagnosed, severe and death groups.
We used the random-effects model to calculate the standard mean difference (SMD) and 95% confidence interval (CI) for lymphocyte count in the CoV-infection patients and to draw a forest plot. Subgroup analysis was performed based on the study definition of severity. Heterogeneity between pairs of studies was quantified using the I2 statistic. We investigated potential sources of heterogeneity, including prognosis and data source (original data vs extrapolated data), by performing subgroup analyses. One was prognosis subgroup that was divided into mortality, severity and diagnostic rate subgroups according to different prognosis investigated in included studies. One was data source subgroup that was divided into original data and extrapolated data subgroups. The criterion was whether the data extrapolated according to the study by Hozo et al[20]. All of the analyses were performed using Stata version 14 (Stata Corp., College Station, TX, USA).
We identified 609 articles using database searches. After screening, 16 articles remained. After examination of the full-text articles, we excluded eight of these articles: three contained duplicate information from same patient; four articles had insufficient data on lymphocyte count; and one article chose a patient with community-acquired pneumonia as a control group. Therefore, eight studies were included in our meta-analysis[10,13,14,19,21-24].
The characteristics of the eight included studies are summarized in Table 1. All of the studies included were prospective or retrospective case–control studies, cohort studies, or cross-sectional studies. The patient population in each study ranged from 30 to 346 (1057 patients in total). The earliest publications were from 2003[22,24], and the most recent article was published in 2020[10,21,23]. Most studies were based in China (n = 7), including Taiwan (n = 3)[13,19,24], Wuhan (n = 2)[10], Beijing (n = 1)[22], and Shanghai (n = 1)[23]. One study was based in Saudi Arabia[14]. The CoV diseases studied were COVID-19 (n = 3; 37.5%)[10,21,23], MERS (n = 1; 12.5%)[14], and SARS (n = 4; 50%)[13,19,22,24]. Four studies (50%) investigated prognosis with respect to diagnostic rate (diagnosed vs nondiagnosed)[19,22-24], two studies (25%) investigated prognosis with respect to severity (severe vs nonsevere)[10,21], and two studies (25%) investigated prognosis with respect to mortality (death vs survival)[13,14].
| Ref. | City, country | Coronavirus disease | Outcome | n (case) | Lymphocyte count of case, mean ± SD (× 109/L) | n (control) | Lymphocyte count of control, mean ± SD (× 109/L) |
| Das et al[14], 2015 | Saudi Arabia | MERS | Mortality | 19 | 18.25 ± 11.75 | 36 | 24.00 ± 19.75 |
| Chang et al[13], 2006 | Taiwan, China | SARS | Mortality | 73 | 0.81 ± 0.38 | 273 | 1.02 ± 0.48 |
| Wang et al[10], 2020 | Wuhan, China | COVID-19 | Severity | 36 | 0.80 ± 0.10 | 102 | 0.90 ± 0.10 |
| Zhang et al[21], 2020 | Wuhan, China | COVID-19 | Severity | 58 | 0.70 ± 0.13 | 82 | 0.80 ± 0.10 |
| Cui et al[22], 2003 | Beijing, China | SARS | Diagnostic rate | 38 | 0.91 ± 0.44 | 200 | 3.00 ± 1.00 |
| Chen et al[19], 2006 | Taiwan, China | SARS | Diagnostic rate | 15 | 0.60 ± 0.30 | 15 | 2.00 ± 0.20 |
| Li et al[23], 2020 | Shanghai, China | COVID-19 | Diagnostic rate | 10 | 1.22 ± 0.20 | 30 | 1.75 ± 0.33 |
| Chen et al[24], 2004 | Taiwan, China | SARS | Diagnostic rate | 8 | 0.90 ± 0.30 | 62 | 1.50 ± 1.10 |
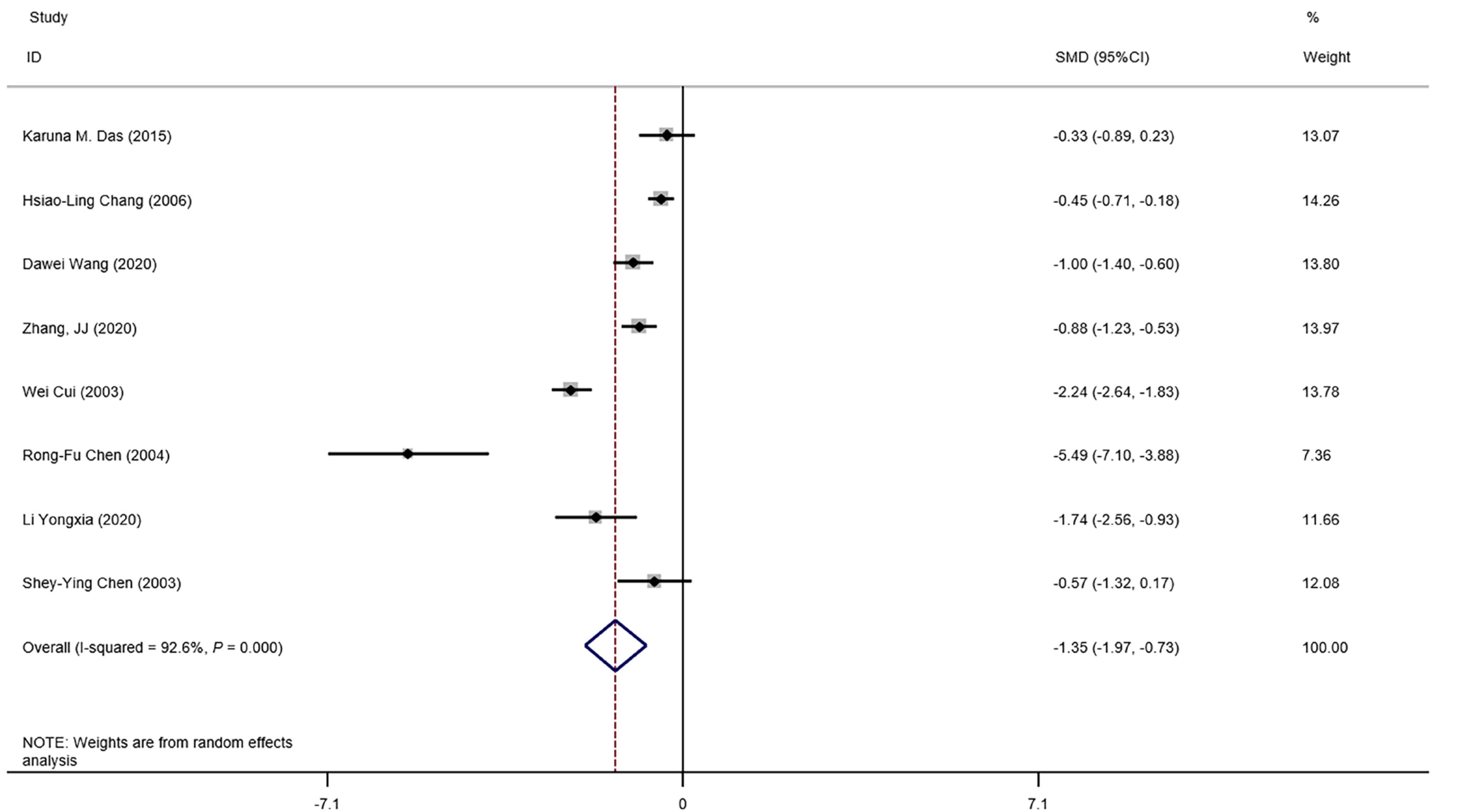
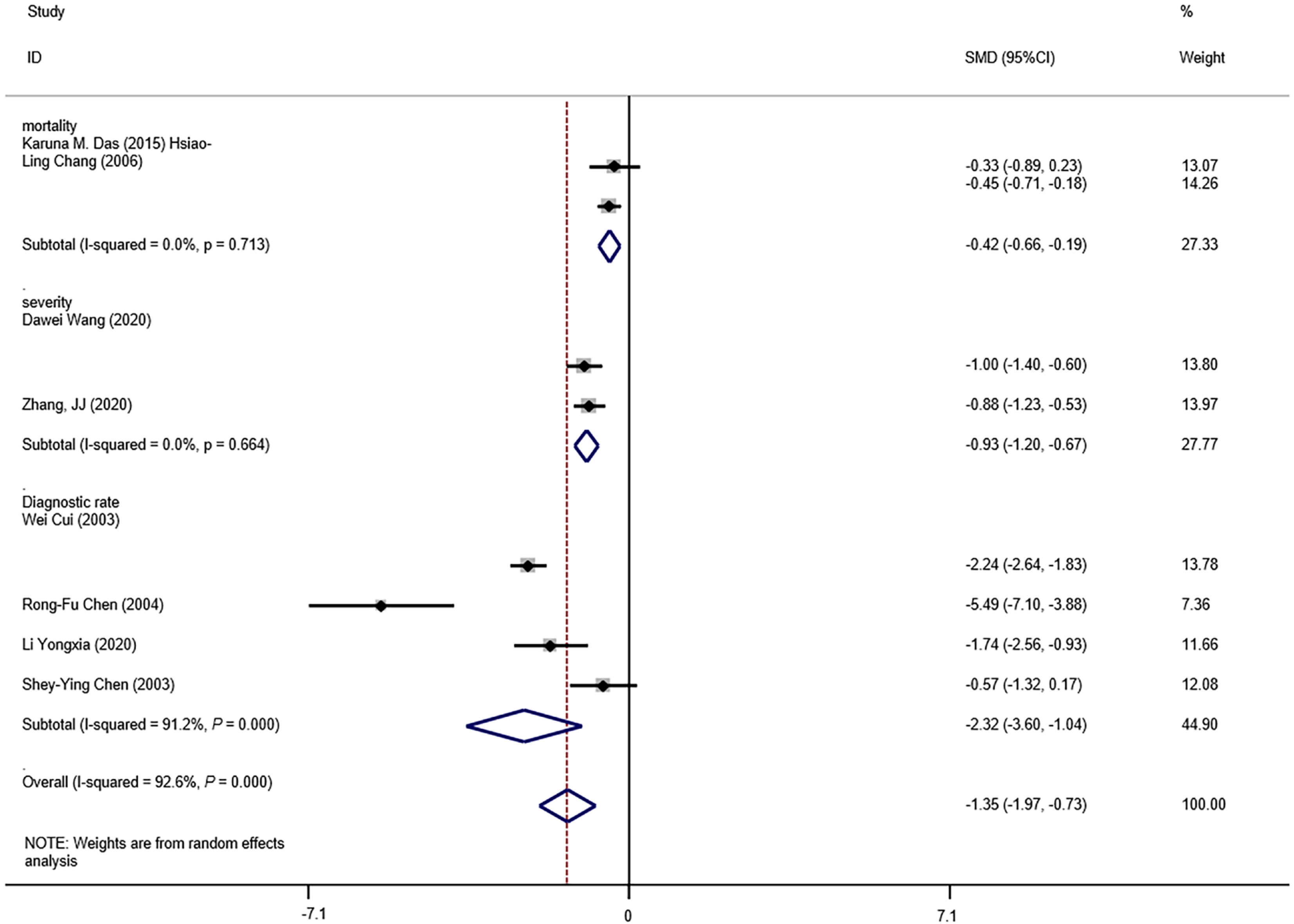
The means and standard deviations of lymphocyte counts from four studies[10,14,21,23] were extrapolated from sample size, median, and IQR (Table 1). In the forest plot, if the SMD (95%CI) was < 0, it showed that the mean and standard deviation of lymphocyte counts in the case group were less than those in the control group. That meant lymphocyte counts were less in severe CoV infection. Analysis showed that lymphocyte counts were associated with severe CoV infection (SMD = 1.35, 95%CI: 1.97 to 0.37, P < 0.001, I2= 92.6%) (Figure 2). There was heterogeneity among studies. Therefore, subgroup analysis, stratified by prognosis and data source, was performed. In the subgroup analysis stratified by prognosis, lymphocytes were associated with mortality due to CoV infection (n = 2, SMD = 0.42, 95%CI: 0.66 to 0.19, P < 0.001, I2= 0.0%), with severity of CoV infection (n = 2, SMD = 0.93, 95%CI: 1.20 to 0.67, P < 0.001, I2= 0.0%), and with the diagnostic rate of CoV infection (n = 4, SMD = 2.32, 95%CI: 3.60 to 1.04, P < 0.001, I2= 91.2%) (Figure 3). In the subgroup analysis stratified by data source, lymphocytes were associated with both the extrapolated data (n = 4, SMD = 0.92, 95%CI: 1.33 to 0.51, P < 0.001, I2= 64.0%) and the original data (n = 4, SMD = 1.97, 95%CI: 3.35 to 0.60, P < 0.001, I2= 96.5%) (see Supplementary Figure 1, which illustrates the forest plot of subgroup analysis stratified by data source), explaining the observed overall heterogeneity among studies. In addition, because few studies were included in our analysis, it was unclear whether the funnel plot was symmetrical (Figure 4), but Begg’s test and Egger’s test showed that publication bias had no significant effects on the results of the meta-analysis (P = 0.174).
COVID-19 is rapidly infectious and highly severe, with a high mortality rate[3]. Patients with COVID-19 exhibit a wide range of variability in disease severity. In clinical practice, we believe that low levels of lymphocytes are disadvantageous for COVID-19 patients. Because lymphocyte count reflects disease characteristics, it might potentially help in the evaluation of COVID-19 severity. Usefully, this measure is easily available in laboratory tests.
Lymphocytes are vital cells that maintain immune function and execute the immune response in the body[8]. In the best-case scenario, the cellular immune response rapidly clears CoV with little or no clinical signs of infection. Alternatively, the virus causes a state of immunosuppression, which debilitates and sometimes overwhelms the host’s defenses[25]. Currently, no detailed study of the immunological response to SARS-CoV-2 is available. Thus, we must rely on previous studies of other CoVs, especially SARS-CoV and MERS-CoV[26]. Based on other CoVs, SARS-CoV-2 might induce a T-lymphocyte-mediated protective immune response[25]. However, lymph
In this study, we designed a meta-analysis to explore the feasibility of using lymphocyte count to predict COVID-19 severity. All of the eight studies included in our meta-analysis involved SARS, MERS or COVID-19, and were prospective or retrospective case–control, cohort or cross-sectional studies (Table 1). Existing research results do not always rank the severity of COVID-19 infections using the same criteria. Even during the earliest part of the Chinese outbreak, the guidelines for COVID-19 diagnosis and treatment issued by the Chinese National Health Commission were revised seven times based on clinical experiences[30]. Therefore, we roughly evaluated disease severity using paired concepts: diagnosed vs nondiagnosed [19,22-24]; severe vs nonsevere[10], and death vs survival[13,14].
We found that lymphocyte count was associated with severe CoV infection (Figure 2). Lymphocytes were also associated with CoV mortality, severity and diagnostic rate (Figure 3). Thus, lymphocyte count may represent a simple, rapid, and commonly available diagnostic and severity-prediction tool for CoV infections, including COVID-19. Because we included studies of SARS-CoV-2 infections, our results indicate the value of lymphocyte count for the assessment of COVID-19 severity. A recent related study suggested that lymphocyte percentage might be a reliable indicator of moderate, severe or critical infections, independent of auxiliary indicators[15]. However, this study included few severe (n = 39) and critical (n = 28) cases[15]. In combination, our analysis and the most recent research results suggested that lymphocyte counts potentially reflect COVID-19 severity. Meanwhile, some research has shown that the viral load of SARS-CoV-2 is associated with the lymphocyte count: A study of interleukin (IL)-3 (a cytokine produced by T lymp
For the acute outbreak of COVID-19, the meta-analysis has limitations. We faced a shortage of data sources during the research. Even now, the data on COVID-19 are insufficient. The research results in the previous response to CoVs should be used for reference. The most mainstream of these were SARS and MERS. As other important CoV diseases, they have similarities to COVID-19. From the perspective of the important characteristics of respiratory infections, SARS and MERS could be included in the reference category. The prevention and treatment methods of COVID-19 all often refer to methods of SARS and MERS. A meta-analysis about prevention of person-to-person transmission of COVID-19 also included SARS and MERS coincidentally[37]. Studies have shown that lymphopenia is related to apoptosis in SARS[28]. Lymphopenia is the result of direct T cell infection and infection-induced apoptosis in MERS[29]. Although we think that this conclusion is insufficient, because there are some treatments (for example: hormone use, antibody titer, etc.) that affect the lymphocyte count in the clinic. However, from the current situation of prevalence, spread and treatment, it is of reference value. In studies that were included in the meta-analysis, five referred to lymphocyte counts that were obtained on the day of hospital admission, and two studies did not report clearly the time that lymphocyte counts were obtained. In clinical research, because it is difficult to make a complete review of the course of disease admission, it is generally accepted to choose the day/time of hospital admission as the sampling point[10]. It has been shown that the degree of lymphopenia is steady during the first week[38], and lymphocyte counts progressively decrease and reach their lowest point on day 14 in most cases[38]. Therefore, our analysis did not divide these studies by time of lymphocyte count acquisition. One week of hospital admission could be recognized as a stable interval for lymphocyte counts. However, a recent study showed that lymphocyte count varied at different times of hospitalization[39]. Thus, further study of lymphocyte counts at varied times could increase the predicted efficiency in COVID-19.
Our meta-analysis provides information on three simple and common interventions to combat the immediate threat of COVID-19, while new evidence on pharmacological treatments, vaccines, hematology monitoring and other personal protective strategies is being generated[37]. It showed that lymphocyte count may represent a simple, rapid and commonly available laboratory index with which to diagnosis infection and predict the severity of CoV infections, including COVID-19. However, our study was limited by the lack of COVID-19 studies, especially those with homogeneous or uniform standards. Therefore, it was necessary to include SARS and MERS in our study; both of which can lead to coronaviral pneumonia. Thus, these diseases might represent a reference for COVID-19. During the current severe COVID-19 pandemic, we hope to provide a framework for the future predictions of COVID-19 severity. Indeed, if critical infections could be predicted and treated earlier, mortality would be lower. Lymphocyte count is a potential indicator of COVID-19 severity and deserves further examination. However, the high heterogeneity among studies suggests that additional research, including case–control and cohort studies, is required for detailed analyses of the relationship between lymphocyte count and COVID-19 severity.
In December 2019, coronavirus disease 2019 (COVID-19) was reported first in Wuhan, China. COVID-19 is currently a global pandemic.
COVID-19 with high morbidity is a life-threatening disease globally. It is important to develop a rapid, simple clinical method to identify severe COVID-19 cases.
The aim of this study was to assess the suitability of lymphocyte count as a biomarker of COVID-19 severity.
We searched five literature databases (PubMed/MEDLINE, Web of Science, Google Scholar, Embase, and Scopus) to identify eligible articles. A meta-analysis was performed to calculate the standard mean difference (SMD) and 95% confidence interval (CI) of lymphocyte counts in coronaviral pneumonia cases.
Our research integrated eight studies, including 1057 patients. Lymphocyte counts were associated with severe coronavirus (CoV) infection (SMD = 1.35, 95%CI: 1.97 to 0.37, P < 0.001, I2 = 92.6%). In the subgroup analysis stratified by prognosis, lymphocytes were associated with coronavirus infection mortality (n = 2, SMD = 0.42, 95%CI: 0.66 to 0.19, P < 0.001, I2 = 0.0%), severity (n = 2, SMD = 0.93, 95%CI: 1.20 to 0.67, P < 0.001, I2 = 0.0%), and diagnostic rate (n = 4, SMD = 2.32, 95%CI: 3.60 to 1.04, P < 0.001, I2 = 91.2%).
Lymphocyte count may represent a simple, rapid and commonly available laboratory index with which to diagnosis infection and predict the severity of CoV infections, including COVID-19.
As a CoV hypotype, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is similar to the CoVs causing severe acute respi
This article also commemorates the Chinese medical staff who gave their lives in the fight against COVID-19.
Manuscript source: Unsolicited manuscript
Specialty type: Infectious diseases
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B, B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Shimizu Y S-Editor: Liu M L-Editor: Kerr C P-Editor: Yu HG
| 1. | Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W; China Novel Coronavirus Investigating and Research Team. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020;382:727-733. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18987] [Cited by in RCA: 17639] [Article Influence: 3527.8] [Reference Citation Analysis (0)] |
| 2. | Sohrabi C, Alsafi Z, O'Neill N, Khan M, Kerwan A, Al-Jabir A, Iosifidis C, Agha R. World Health Organization declares global emergency: A review of the 2019 novel coronavirus (COVID-19). Int J Surg. 2020;76:71-76. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3213] [Cited by in RCA: 2654] [Article Influence: 530.8] [Reference Citation Analysis (0)] |
| 3. | World Health Organization. Coronavirus disease 2019 (COVID-19) Situation Report –66. 2020 [cited 20 December 2020]. Available from: https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200326-sitrep-66-covid-19.pdf?sfvrsn=81b94e61_2. |
| 4. | Peiris JS, Guan Y, Yuen KY. Severe acute respiratory syndrome. Nat Med. 2004;10:S88-S97. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 805] [Cited by in RCA: 740] [Article Influence: 35.2] [Reference Citation Analysis (0)] |
| 5. | Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med. 2012;367:1814-1820. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4030] [Cited by in RCA: 4029] [Article Influence: 309.9] [Reference Citation Analysis (0)] |
| 6. | Badawi A, Ryoo SG. Prevalence of Diabetes in the 2009 Influenza A (H1N1) and the Middle East Respiratory Syndrome Coronavirus: A Systematic Review and Meta-Analysis. J Public Health Res. 2016;5:733. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 58] [Cited by in RCA: 60] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 7. | Epidemiology Working Group for NCIP Epidemic Response; Chinese Center for Disease Control and Prevention. The epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19) in China. Zhonghua Liu Xing Bing Xue Za Zhi. 2020;41:145-151. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1341] [Reference Citation Analysis (0)] |
| 8. | Jalkanen S, Salmi M. Lymphatic endothelial cells of the lymph node. Nat Rev Immunol. 2020;20:566-578. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 146] [Article Influence: 29.2] [Reference Citation Analysis (0)] |
| 9. | Wu J, Liu J, Zhao X, Liu C, Wang W, Wang D, Xu W, Zhang C, Yu J, Jiang B, Cao H, Li L. Clinical Characteristics of Imported Cases of Coronavirus Disease 2019 (COVID-19) in Jiangsu Province: A Multicenter Descriptive Study. Clin Infect Dis. 2020;71:706-712. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 393] [Cited by in RCA: 413] [Article Influence: 82.6] [Reference Citation Analysis (1)] |
| 10. | Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA. 2020;323:1061-1069. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14113] [Cited by in RCA: 14764] [Article Influence: 2952.8] [Reference Citation Analysis (0)] |
| 11. | Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395:507-513. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14869] [Cited by in RCA: 12971] [Article Influence: 2594.2] [Reference Citation Analysis (1)] |
| 12. | Rodriguez-Morales AJ, Cardona-Ospina JA, Gutiérrez-Ocampo E, Villamizar-Peña R, Holguin-Rivera Y, Escalera-Antezana JP, Alvarado-Arnez LE, Bonilla-Aldana DK, Franco-Paredes C, Henao-Martinez AF, Paniz-Mondolfi A, Lagos-Grisales GJ, Ramírez-Vallejo E, Suárez JA, Zambrano LI, Villamil-Gómez WE, Balbin-Ramon GJ, Rabaan AA, Harapan H, Dhama K, Nishiura H, Kataoka H, Ahmad T, Sah R; Latin American Network of Coronavirus Disease 2019-COVID-19 Research (LANCOVID-19). Clinical, laboratory and imaging features of COVID-19: A systematic review and meta-analysis. Travel Med Infect Dis. 2020;34:101623. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1647] [Cited by in RCA: 1432] [Article Influence: 286.4] [Reference Citation Analysis (0)] |
| 13. | Chang HL, Chen KT, Lai SK, Kuo HW, Su IJ, Lin RS, Sung FC. Hematological and biochemical factors predicting SARS fatality in Taiwan. J Formos Med Assoc. 2006;105:439-450. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 40] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 14. | Das KM, Lee EY, Al Jawder SE, Enani MA, Singh R, Skakni L, Al-Nakshabandi N, AlDossari K, Larsson SG. Acute Middle East Respiratory Syndrome Coronavirus: Temporal Lung Changes Observed on the Chest Radiographs of 55 Patients. AJR Am J Roentgenol. 2015;205:W267-W274. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 148] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 15. | Tan L, Wang Q, Zhang D, Ding J, Huang Q, Tang YQ, Miao H. Lymphopenia predicts disease severity of COVID-19: a descriptive and predictive study. Signal Transduct Target Ther. 2020;5:33. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 759] [Cited by in RCA: 1048] [Article Influence: 209.6] [Reference Citation Analysis (0)] |
| 16. | Moher D, Liberati A, Tetzlaff J, Altman DG; PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. Int J Surg. 2010;8:336-341. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9207] [Cited by in RCA: 8047] [Article Influence: 536.5] [Reference Citation Analysis (2)] |
| 17. | Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497-506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35178] [Cited by in RCA: 30106] [Article Influence: 6021.2] [Reference Citation Analysis (3)] |
| 18. | Who Mers-Cov Research Group. State of Knowledge and Data Gaps of Middle East Respiratory Syndrome Coronavirus (MERS-CoV) in Humans. PLoS Curr. 2013;5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 235] [Article Influence: 19.6] [Reference Citation Analysis (0)] |
| 19. | Chen RF, Chang JC, Yeh WT, Lee CH, Liu JW, Eng HL, Yang KD. Role of vascular cell adhesion molecules and leukocyte apoptosis in the lymphopenia and thrombocytopenia of patients with severe acute respiratory syndrome (SARS). Microbes Infect. 2006;8:122-127. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 49] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 20. | Hozo SP, Djulbegovic B, Hozo I. Estimating the mean and variance from the median, range, and the size of a sample. BMC Med Res Methodol. 2005;5:13. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4895] [Cited by in RCA: 6890] [Article Influence: 344.5] [Reference Citation Analysis (0)] |
| 21. | Zhang JJ, Dong X, Cao YY, Yuan YD, Yang YB, Yan YQ, Akdis CA, Gao YD. Clinical characteristics of 140 patients infected with SARS-CoV-2 in Wuhan, China. Allergy. 2020;75:1730-1741. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2139] [Cited by in RCA: 2341] [Article Influence: 468.2] [Reference Citation Analysis (0)] |
| 22. | Cui W, Fan Y, Wu W, Zhang F, Wang JY, Ni AP. Expression of lymphocytes and lymphocyte subsets in patients with severe acute respiratory syndrome. Clin Infect Dis. 2003;37:857-859. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 130] [Cited by in RCA: 138] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 23. | Li YX, Wu W, Yang T, Zhou W, Fu YM, Feng QM, Ye JM. [Characteristics of peripheral blood leukocyte differential counts in patients with COVID-19]. Zhonghua Nei Ke Za Zhi. 2020;59:372-374. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 24. | Chen SY, Su CP, Ma MH, Chiang WC, Hsu CY, Ko PC, Tsai KC, Yen ZS, Shih FY, Chen SC, Chen WJ. Predictive model of diagnosing probable cases of severe acute respiratory syndrome in febrile patients with exposure risk. Ann Emerg Med. 2004;43:1-5. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 21] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 25. | Jin W, Zhu X, Yao F, Xu X, Chen X, Luo Z, Zhao D, Li X, Leng X, Sun L. Corrigendum to "Cytoprotective effect of Fufang Lurong Jiangu capsule against hydrogen peroxide-induced oxidative stress in bone marrow stromal cell-derived osteoblasts through the Nrf2/HO-1 signaling pathway" [Biomed. Pharmacother. 121 (2020) 109676]. Biomed Pharmacother. 2020;122:109808. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 26. | Hu H, Tao L, Wang Y, Chen L, Yang J, Wang H. Enhancing immune responses against SARS-CoV nucleocapsid DNA vaccine by co-inoculating interleukin-2 expressing vector in mice. Biotechnol Lett. 2009;31:1685-1693. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 17] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 27. | Chan MH, Wong VW, Wong CK, Chan PK, Chu CM, Hui DS, Suen MW, Sung JJ, Chung SS, Lam CW. Serum LD1 isoenzyme and blood lymphocyte subsets as prognostic indicators for severe acute respiratory syndrome. J Intern Med. 2004;255:512-518. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 30] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 28. | O'Donnell R, Tasker RC, Roe MF. SARS: understanding the coronavirus: apoptosis may explain lymphopenia of SARS. BMJ. 2003;327:620. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 32] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 29. | Chu H, Zhou J, Wong BH, Li C, Chan JF, Cheng ZS, Yang D, Wang D, Lee AC, Yeung ML, Cai JP, Chan IH, Ho WK, To KK, Zheng BJ, Yao Y, Qin C, Yuen KY. Middle East Respiratory Syndrome Coronavirus Efficiently Infects Human Primary T Lymphocytes and Activates the Extrinsic and Intrinsic Apoptosis Pathways. J Infect Dis. 2016;213:904-914. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 288] [Cited by in RCA: 386] [Article Influence: 38.6] [Reference Citation Analysis (0)] |
| 30. | National Health Commission of China. The guidelines for diagnosis and treatment of novel coronavirus (2019-nCoV) infected pneumonia (the seventh edition draft). [cited 20 December 2020]. Available from: http://www.gov.cn/zhengce/zhengceku/2020-03/04/5486705/files/ae61004f930d47598711a0d4cbf874a9.pdf. |
| 31. | Bénard A, Jacobsen A, Brunner M, Krautz C, Klösch B, Swierzy I, Naschberger E, Podolska MJ, Kouhestani D, David P, Birkholz T, Castellanos I, Trufa D, Sirbu H, Vetter M, Kremer AE, Hildner K, Hecker A, Edinger F, Tenbusch M, Mühl-Zürbes P, Steinkasserer A, Richter E, Streeck H, Berger MM, Brenner T, Weigand MA, Swirski FK, Schett G, Grützmann R, Weber GF. Interleukin-3 is a predictive marker for severity and outcome during SARS-CoV-2 infections. Nat Commun. 2021;12:1112. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 47] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 32. | Graham JB, Swarts JL, Leist SR, Schäfer A, Menachery VD, Gralinski LE, Jeng S, Miller DR, Mooney MA, McWeeney SK, Ferris MT, Pardo-Manuel de Villena F, Heise MT, Baric RS, Lund JM. Baseline T cell immune phenotypes predict virologic and disease control upon SARS-CoV infection in Collaborative Cross mice. PLoS Pathog. 2021;17:e1009287. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 24] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 33. | Corrao S, Gervasi F, Di Bernardo F, Natoli G, Raspanti M, Catalano N, Argano C. Immunological Characteristics of Non-Intensive Care Hospitalized COVID-19 Patients: A Preliminary Report. J Clin Med. 2021;10:849. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 8] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 34. | Gao Y, Chen L, Chi J, Zeng S, Feng X, Li H, Liu D, Wang S, Wang Y, Yu R, Yuan Y, Xu S, Li C, Zhang W, Li S, Gao Q. Development and validation of an online model to predict critical COVID-19 with immune-inflammatory parameters. J Intensive Care. 2021;9:19. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 35. | Xu H, Zhong L, Deng J, Peng J, Dan H, Zeng X, Li T, Chen Q. High expression of ACE2 receptor of 2019-nCoV on the epithelial cells of oral mucosa. Int J Oral Sci. 2020;12:8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1709] [Cited by in RCA: 1732] [Article Influence: 346.4] [Reference Citation Analysis (0)] |
| 36. | Fischer K, Hoffmann P, Voelkl S, Meidenbauer N, Ammer J, Edinger M, Gottfried E, Schwarz S, Rothe G, Hoves S, Renner K, Timischl B, Mackensen A, Kunz-Schughart L, Andreesen R, Krause SW, Kreutz M. Inhibitory effect of tumor cell-derived lactic acid on human T cells. Blood. 2007;109:3812-3819. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1001] [Cited by in RCA: 1406] [Article Influence: 78.1] [Reference Citation Analysis (0)] |
| 37. | Chu DK, Akl EA, Duda S, Solo K, Yaacoub S, Schünemann HJ; COVID-19 Systematic Urgent Review Group Effort (SURGE) study authors. Physical distancing, face masks, and eye protection to prevent person-to-person transmission of SARS-CoV-2 and COVID-19: a systematic review and meta-analysis. Lancet. 2020;395:1973-1987. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2542] [Cited by in RCA: 2385] [Article Influence: 477.0] [Reference Citation Analysis (0)] |
| 38. | Wong RS, Wu A, To KF, Lee N, Lam CW, Wong CK, Chan PK, Ng MH, Yu LM, Hui DS, Tam JS, Cheng G, Sung JJ. Haematological manifestations in patients with severe acute respiratory syndrome: retrospective analysis. BMJ. 2003;326:1358-1362. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 424] [Cited by in RCA: 433] [Article Influence: 19.7] [Reference Citation Analysis (0)] |
| 39. | Petrilli CM, Jones SA, Yang J, Rajagopalan H, O'Donnell L, Chernyak Y, Tobin KA, Cerfolio RJ, Francois F, Horwitz LI. Factors associated with hospital admission and critical illness among 5279 people with coronavirus disease 2019 in New York City: prospective cohort study. BMJ. 2020;369:m1966. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1590] [Cited by in RCA: 1818] [Article Influence: 363.6] [Reference Citation Analysis (1)] |