Peer-review started: January 11, 2022
First decision: May 31, 2022
Revised: June 11, 2022
Accepted: July 27, 2022
Article in press: July 27, 2022
Published online: August 30, 2022
Processing time: 231 Days and 2.2 Hours
Early diagnosis and prognosis of ischemic stroke remains a critical challenge in clinical settings. A blood biomarker can be a promising quantitative tool to represent the clinical manifestations in ischemic stroke. Cell-free DNA (cfDNA) has recently turned out to be a popular circulating biomarker due to its potential relevance for diagnostic applications in a variety of disorders. Despite bright outlook of cfDNA in clinical applications, very less is known about its origin, composition, or function. Several recent studies have identified cell-derived mitochondrial components including mitochondrial DNA (mtDNA) in the extracellular spaces including blood and cerebrospinal fluid. However, the time course of alterations in plasma mtDNA concentrations in patients after an ischemic stroke is poorly understood. DNA is thought to be freed into the plasma shortly after the commencement of an ischemic stroke and then gradually decreased. However, the importance of cell-free mtDNA (cf-mtDNA) in ischemic stroke is still unknown. This review summarizes about the utility of biomarkers which has been standardized in clinical settings and role of cfDNA including cf-mtDNA as a non-invasive potential biomarker of ischemic stroke.
Core Tip: Early and accurate diagnosis of ischemic stroke is critical to achieve favorable clinical outcome. Cell-free DNA can be used as a useful biomarker for early diagnosis and prognosis of ischemic stroke for saving time and increasing the likelihood of successful intervention. Discriminative quantification of cell free mitochondrial DNA instead of overall circulating DNA may provide more significant value for identifying real-time host response. The future practical adoption of this strategy may be aided by reliable and standardized quantification of cell-free mitochondrial DNAs in ischemic stroke patients to design more effective diagnosis, prognosis and therapeutic strategies.
- Citation: Fathima N, Manorenj S, Vishwakarma SK, Khan AA. Role of cell-free DNA for predicting incidence and outcome of patients with ischemic stroke. World J Neurol 2022; 8(1): 1-9
- URL: https://www.wjgnet.com/2218-6212/full/v8/i1/1.htm
- DOI: https://dx.doi.org/10.5316/wjn.v8.i1.1
Stroke is associated with significant morbidity and mortality with continuously increasing incidence globally, owing to the rising prevalence of different forms of cardiovascular diseases[1]. Acute brain injuries, particularly traumatic brain injury, and brain stroke are among the main causes of death and disability worldwide[2]. Currently, stroke has become a major public health burden that is expected to rise in the next decades as a result of demographic transitions, particularly in developing countries[3]. It affects 13.7 million people and kills 5.5 million people per year. Ischemic infarctions accounts for approximately 87% of strokes, a prevalence that increased significantly between 1990 and 2016, owing to lower mortality and improved therapeutic management. Over the same period, the incidence of stroke in low- and middle-income countries increased by two-folds, while it reduced by 42% in high-income countries[4]. According to the Global Cost of Disease Study, the socio-economic burden of stroke has increased over time. The risk of stroke rises to more than 3-fold over the age of 55 years. While, younger adults of age 20 years to 54 years the incidence of stroke has been reported 12.9% to 18.6% of all cases globally between 1990 and 2016. Nonetheless, over the same time period, age-standardized attributable death rates declined by 36.2%[5,6].
Among different types of strokes, ischemic stroke is the most common type that includes cryptogenic, lacunae, and thromboembolic forms of strokes. Ischemic stroke results due to blockage in blood flow to a part of the brain and accounts for approximately 87% of all the strokes. Despite the high rate of morbidity and mortality caused by ischemic stroke, the varied etiology and intricate pathophysiology make clinical diagnosis and prognosis prediction very difficult. Efficient prognosis following an ischemic stroke remains a major obstacle in clinical settings.
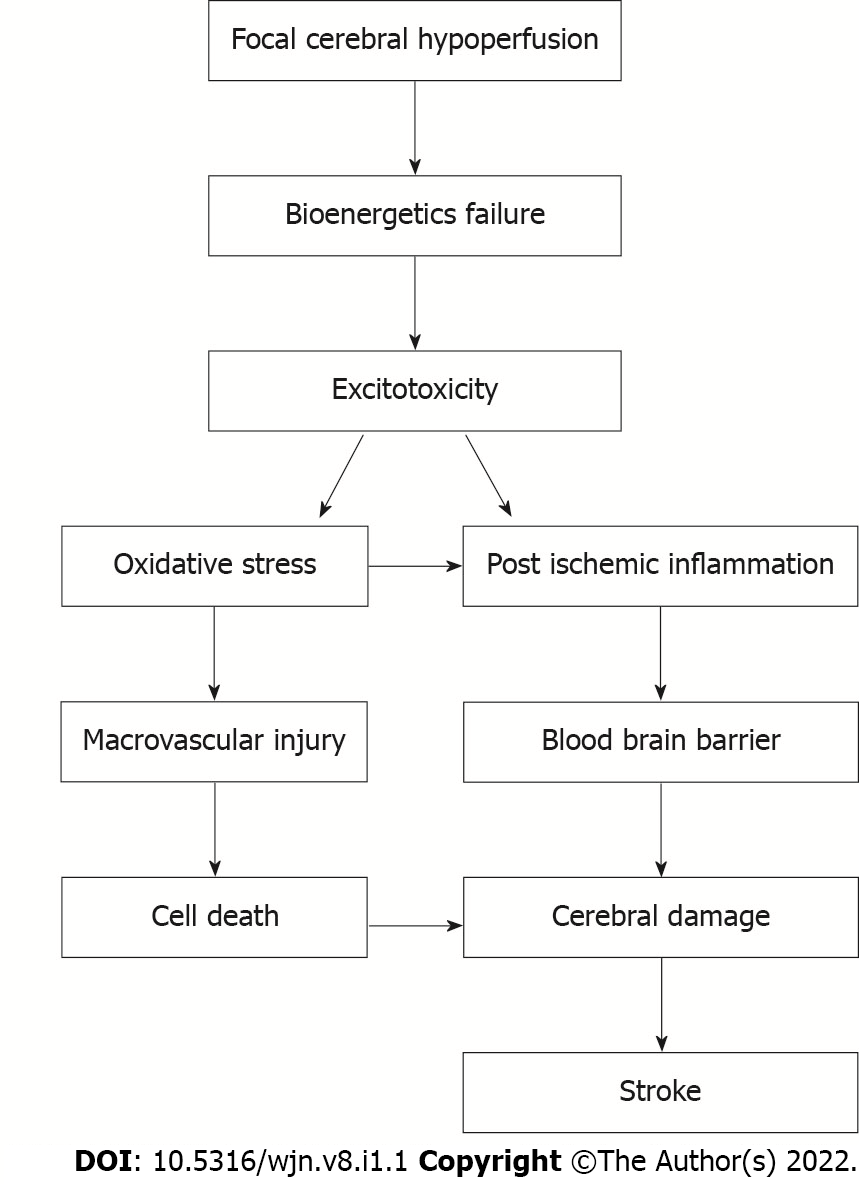
Because acute ischemic stroke is often isodense on computed tomography (CT), its initial utility is mainly limited to excluding a haemorrhage[7]. Currently, magnetic resonance imaging (MRI) with diffusion has been considered a gold standard tool for acute diagnosis. However, MRI is expensive and less widely available to rural and general population[7]. The major hitches in cost-effective diagnosis, prognosis, and improved therapeutic efficacy is accompanied by the complicated pathophysiological processes including energy failure, ion homeostasis imbalance, acidosis, intracellular calcium overload, brain excitoxicity, free radical-mediated lipid oxidation, inflammatory cell infiltration, and glial cell activation (Figure 1). As a result of the combination of aforementioned complicated pathophysiological processes alterations in the blood-brain barrier and the release of different neurological markers into circulation has been reported[8]. In addition, ischemic occlusion causes thrombotic and embolic situations in the brain[9]. The narrowing of veins due to atherosclerosis affects blood flow in thrombosis. Plaque buildup will eventually narrow the vascular chamber and cause clots, resulting in thrombotic stroke. Reduced blood supply to the brain region creates an embolism in an embolic stroke; blood flow to the brain decreases, producing acute stress and premature cell death (necrosis). Following necrosis, the plasma membrane is disrupted, organelles enlarge, and cellular contents leak into the extracellular environment, resulting in loss of neuronal function[8,10].
Owing to the lack of real-time tracking of ischemic stroke pathogenesis and treatment response, the extracellular components released into circulation or secretion may provide an important tool for evolving better approach. Recent studies have demonstrated that circulating cell-free DNA (cfDNA) molecules are significantly increased in stroke condition and act as important tool for non-invasive monitoring of disease progression and prognosis[11-13]. Although various mechanisms of release of cfDNA has been predicted, activated neutrophils have been demonstrated to produce neutrophil extracellular traps (NETs) in response to a variety of stimuli resulting in the release of detectable amount of cfDNA in thrombi. It's also likely that the link between cfDNA levels and neutrophil count is due to activated neutrophils' tendency to leak significant amounts of DNA when edifice NETs[14]. As a result, it is possible that circulating cfDNA in stroke comes from both the damaged Neurovascular Unit and NET formation, which could explain the link between circulating cfDNA levels and neutrophil count[15]. However, more detailed exploration is required to understand the proper mechanism for the occurrence and progression of ischemic stroke.
Determining the cause of a stroke can be difficult. Based upon the current progress towards diagnostic investigations and classification characteristics, the cause of a stroke is usually unclear or even unknown. Natriuretic peptides, glial fibrilliary acidic protein, S100b, neuron specific enolase, myelin basic protein, interleukin-6, matrix metalloproteinase (MMP-9), D-dimer, and fibrinogen have all been examined as biomarkers in stroke. Despite notable research, a troponin-like biomarker to aid in the diagnosis of stroke has been eluded by the researchers. This could be due to the fact that stroke is a heterogeneous disease with a wide range of infarct size, location, and origin[16].
In the case of ischemic stroke, a variety of biomarker panels have been tested for the diagnosis and causation. A panel of five proteins (MMP-9, BNGF, vWF, MCP-1, and S100B)[17], another panel of four proteins (MMP-9, brain natriuretic factor, D-dimer, and S100B)[18], and panel of five proteins (eotaxin, epidermal growth factor receptor, S100A12, metalloproteinase inhibitor-4, and prolactin)[19] are among them. When compared to individual indicators, the panel of numerous markers has consistently exhibited better sensitivity and specificity in diagnosing the ischemic stroke. The results support the concept of merging many markers into a panel, even though none have produced enough evidence to indicate clinical usefulness[20].
Protein assays in plasma or serum have been a frequent method of measuring biomarkers in stroke, and they have provided useful information in the establishment of suitable stroke biomarkers[7,17]. The search for biomarkers has been spurred by the need to better stroke diagnosis and to identify the more appropriate cause. Several markers have shown potential, but there is no enough evidence to warrant their use in clinical practice. The known biomarkers of stroke in patients typically reflect the magnitude of tissue damage and inflammation. A recent genome-wide meta-analysis tried to explore the crucial disease-related pathways and molecular regulatory networks after combining messenger RNA and miRNA expression analyses to identify the candidate target molecule for early diagnosis of stroke[18]. The findings of this study identified six considerably increased genes (PTGS2, IL1B, STAT3, MMP9, SOCS3, and CXCL1), as well as two significantly upregulated miRNAs (miR-320b and miR-320d), as possible clinical diagnostic indicators. These all molecules are linked with the release of extracellular molecules into the circulation and need further investigation.
Although various mechanisms such as oxidative stress, cell death, inflammation and changes in the peripheral blood have been identified, but none have found a place in clinical practice largely because of the stringent criteria that must be met by biomarkers before application. Another systematic review on scalable biomarker combinations for early stroke diagnosis also revealed the need of more comprehensive research on circulatory molecules to evaluate, identify, and systematically implement identified biomarker panels into medical practice for stroke recurrence and diagnosis[19]. Other crucial studies have also suggested that evaluating different blood-based molecules to offer diagnostic accuracy for health interventions is the most promising method. These requirements include the precision, accuracy, sensitivity and specificity of the biomarker for the outcome decided the structured data acquisition protocol and the ease of interpretation (Table 1).
| Ref. | Panel | Source of cases | Source of marker | Panel of marker | Sensitivity/specificity (%) |
| Stamova et al[20], 2010 | Gene panel | Ischemic stroke vs control | RNA, whole blood | BCL6, PYGL, RNASE2, S100A9, S100A12, S100P, SLC16A6,ARG1, CA4, CKAP4, ETS-2, HIST2H2AA | 93/95 |
| Montaner et al[21], 2008 | Protein panel | Cardioembolic vs noncardioembolic stroke | Protein, plasma | D-dimer, BNP | 87/85 |
| Laskowitz et al[22], 2009 | Protein panel | Ischemic + hemorrhagic stroke vs control | Protein, plasma | BNP, D-dimer, MMP9, S100B | 86/37 |
| Jickling and Sharp[23], 2011 | Gene panel | Large vessel vs cardioembolic stroke | RNA, Whole Blood | FCRL1, FLJ40125, GRM5, GSTK1, HLA-DOA, IRF6, LHFP, LHFP, LOC284751, LRRC37A3, P2RX5, PIK3C2B, PTPN20A, TFDP1, TMEM19, TSKS, ZNF185, ZNF254, ADAMTSL4, AP3S2 | 100/96 |
| Barr et al[24], 2010 | Gene panel | Ischemic stroke vs control | RNA, whole blood | SCA4, ARG1, S100A12, ORM1, CCR7, CSPG2, IQGAP1, LY96, MMP9, ORM1 | - |
| Montaner et al[25], 2011 | Protein panel | Ischemic + hemorrhagic stroke vs control | Protein, plasma | D-dimer, MMP-9, Caspase-3, Chimerin, Secretagogin, sRAGE | 17/98 |
| Vanni et al[26], 2011 | Protein panel | Ischemic stroke vs control | Protein, whole blood | BNP, D-dimer, MMP9, S100B | 86/37 |
| Sharma et al[27], 2014 | Protein panel | Ischemic + hemorrhagic stroke vs control | Protein, plasma | S100A12, Metalloproteinase inhibitor-4, Prolactin, Eotaxin, EGFR | 90/84 |
Recently, a number of studies have been conducted that examine cfDNA in biological fluids during pathological processes in the human brain[28-31]. The cfDNA is highly fragmented in which released into circulation during apoptosis, cell death, necrosis, inflammation-like conditions. cfDNA contains nuclear DNA and mitochondrial DNA (mtDNA). And it has been thought that cf-DNA would be released into the bloodstream shortly after the ischemic stroke.
Although several biomarkers distinctive to stroke subtypes have been discovered; however, the association of cfDNA as a novel biomarker demands additional investigation into its alliance with stroke subtypes. Such biomarkers could also be used to non-invasively assess stroke severity, which varies between subtypes. The therapeutic window for stroke is limited, occurring only 3-6 h following the onset of symptoms, early recognition of severe patients, and prompt adoption of appropriate therapeutic interventions have enormous prognostic value. In clinical relevance, it could be critical in predicting patient death or functional outcomes, so it is necessary to elucidate further research with stroke. Even though the blood cfDNA levels rise after numerous clinical processes in the body, cfDNA has not traditionally been considered an excellent marker with specificity to a condition like a stroke.
On admission, cfDNA concentrations in the CSF, but not in plasma, are shown to be considerably higher in patients who had poorer outcomes. The diverse origins of cfDNA in both fluids, as well as differences in the dynamics of the blood-brain barrier and CSF-brain barrier integrity changes during a stroke, can explain the observed disparities in cfDNA levels between plasma and CSF at the same time point. In general, the results of different sample types, such as serum, plasma, or CSF, should not be compared[32].
Several studies have demonstrated that cfDNA concentration correlates well with the outcome of stroke intervention in acute ischemic stroke patients (Table 2). In the setting of recanalization, post-intravenous thrombolysis IVT, and mechanical thrombectomy, using cfDNA as a predictive substitute marker to envisage outcome will aid in the most efficient use of limited resources and reduce the load on the healthcare system[30].
| Ref. | Disease specification | Source | Nucleic acid type | Outcome of the study | |
| 1 | Tsai et al[29], 2011 | Acute ischemic stroke | Plasma | cfDNA | Elevated plasma cf-nuclear and mitochondrial DNA in acute ischemic stroke patients than healthy controls |
| 2 | O’Connell et al[34], 2017 | Ischemic stroke | Plasma | cfDNA | Elevated cfDNA in stroke patients relative to those diagnosed as stroke mimics (P = 0.001) |
| 3 | Naumann et al[28], 2017 | Acute ischemic stroke | Plasma | cfDNA | IVT was associated with improved outcome in patients with cfDNA < 10000 kilogenome-equivalents/L (P < 0.05) |
| 4 | Vajpeyee et al[30], 2018 | Acute ischemic stroke | Plasma | Higher cfDNA levels were associated with severity at the time of admission (P = 0.003) and poor outcome as measured by modified Rankin scale 3-mo scores (P = 0.001). Therapeutic intervention mechanical thrombectomy or IV thrombolysis associated with improved outcome in patients with cfDNA < 10000 kilogenome equivalents/L (P ≤ 0.01) | |
| 5 | Kananen et al[36], 2020 | Mortality rate | Plasma | cfDNA | cfDNA associated with increased risk of mortality (hazard ratio 0.1 μg increase in cfDNA, P = 0.0003) |
Extracellular DNA, on the other hand, has been shown in numerous studies to perform as a hazard signal and drive immune responses. Circulating cfDNA appears to be drawn in endothelium pathophysiological changes in trauma patients, the extent of endothelium injury and an augment in cfDNA release are connected. The cfDNA levels in the blood are also discriminating in relation with greater endothelium damage after cardiac surgery, as well as epithelial and endothelial cell death in the lungs, in a dose-dependent manner[28,33]. Neutrophils use a pathogen-clearance system called NETs, and cfDNA is a key component of these traps. As a result, circulating cfDNA could be both an indicator of the severity of the damage and a contributor in the damage-causing pathways.
Regardless of years of research, the sources and processes of tissue injury that results in high cfDNA levels are only partially understood. Apoptosis and necrosis appear in contribute to circulating cfDNA, however even live cells can release DNA into the circulation under specific conditions. The biological properties of cfDNA are still to some extent understood. To improve the accuracy of stroke diagnosis, cfDNA can be used in concurrence with clinical evaluation and imaging methods[35,36]. It is noted that plasma cf-nucleic acids are increased after acute ischemic stroke and studies also showed correlation with clinical parameters like white blood cell count, diabetes milletus, hemoglobin A1c, blood pressure. Along with this an increased sample size and follow up with duration since window period assessment may give a noninvasive prognostic implement. It can be used to supplement the diagnostic workup and aid triage patients for intervention as a stroke biomarker. With the addition of this unique marker, it is now possible to make clued-up predictions about the outcome of mechanical thrombectomy or intravenous thrombolysis in acute stroke patients[29]. Patients and their families can make informed decisions in patients with negative imaging results and before consenting to invasive or medicinal treatment using markers like cfDNA assessment with prognostic usefulness.
Mortality is one of the most important patient-relevant outcomes that have been explored in several earlier studies. Rainer et al[37] found a 100% sensitivity and > 74% specificity in using cfDNA as a predictive biomarker for both ischemic and hemorrhagic stroke. A cfDNA level of > 1400 kg-equiv/L suggested a substantial 60% increase in the probability of mortality at 6-mo in categorical analysis. Furthermore, in a 2007 update by the same group, a substantial difference in cfDNA, 48 h after a stroke was found to be a robust predictor of 6-mo mortality, with 50% lower cfDNA levels for individuals who did not have an event[38].
There are two major types of cfDNA namely cell-free nuclear DNA (cf-ncDNA), and cell-free mtDNA (cf-mtDNA). Recent studies have discovered that in healthy individual's plasma samples, cf-mtDNA can be present in about 50000-fold more copies than cf-ncDNA[39,40]. This indicates that intact cf-mtDNA molecules are present in circulation in higher amount, allowing for the identification and quantification of circulating cf-mtDNA from circulating cf-ncDNA, which could provide more accurate diagnostic information in a variety of physiological and pathological circumstances including ischemic stroke. Because mtDNA is a small circular genome without protective histones, it is more susceptible to breakdown in the circulation, and it is expected that ncDNA and mtDNA will have considerable configuration differences. Hence, the current focus has been on recognizing the potential of cir-mtDNA as a powerful potential biological source in the field of molecular diagnostics and prognosis which is more precise and non-invasive[41].
Recently a few studies have demonstrated the importance of studying cf-mtDNA quantification in various clinical conditions using various samples such as peripheral blood mononuclear cells, whole blood, plasma, urine, and tissue samples[42]. However, extracting pure cell-free cir-mtDNA from plasma samples without contaminating the sample with cir-ncDNA is a significant problem. In this direction, our recent study has reported a fruitful in-house protocol to extract both cf-mtDNA as well as cf-ncDNA using a single plasma sample without contamination of each type of DNA (unpublished). Our data has shown significance of this procedure to clearly identify the potential of quantifying cf-mtDNA to clearly differentiate healthy and diseased individuals at high levels of sensitivity and specificity compared to cf-ncDNA. Apart from our study, a recent report has indicated that studying mitochondrial dynamics may provide a potential therapeutic target for ischemic stroke[43]. However, such investigations are further desired in ischemic stroke to predict the significance of cf-mtDNA as precise, non-invasive diagnostic and prognostic tool.
Plasma mtDNA measurement's utility in acute medicine may be enhanced by technological advancements. For example, using a column-based DNA extraction technology and real-time quantitative polymerase chain reaction (RT-qPCR) analysis, plasma mtDNA findings can currently be obtained within 3 h of sampling. Recent advances in quick capillary-based instrumentation for RT-qPCR analysis may allow this period to be cut in half, 30 min to 90 min[43]. The cfDNA is thought to be freed into the plasma shortly after the commencement of an ischemic stroke and then gradually decreased. Circulating cfDNA levels in plasma surged quickly after an acute ischemic stroke and then gradually reduced. The clinical severity of ischemic stroke is reflected in the level of plasma mtDNA in the acute stage. Circulating mtDNA in plasma has been researched in a variety of disorders during the last few decades, including sepsis, cancer, myocardial infarction, and serious trauma. Plasma cfDNA's prognostic and diagnostic value has been recognized in a number of important situations.
Ischemic stroke is a major public health problem that is expected to worsen in the next decades as a result of population shifts, particularly in developing nations. Ischemic stroke is linked to major morbidity and mortality, and its prevalence is increasing over the world. Simple cf-mtDNA quantification test in the form of liquid biopsy can assist to estimate likely mortality or functional result when CT and/or MRI are either unavailable or show no obvious acute abnormalities.
It is apparent that cfDNA has clinical utility in predicting functional results and long-term survival. Plasma cfDNA as a prognostic marker has the advantage of being non-invasive and straight forward to use. It has also been shown to be able to distinguish between hemorrhagic and ischemic stroke and to be an independent predictor of stroke outcome in patients with negative neuroimaging. cfDNA can help doctors with patient evaluation and complement imaging technologies to improve stroke diagnostic accuracy. It can also help priorities patients for action by supplementing the diagnostic workup. When imaging is negative or not necessary, the predictive value of cfDNA can help patients and physicians make educated decisions about invasive or medicinal treatment.
The track for biomarkers evolution is prompted by the desire to early and non-invasive diagnosis and prognosis. Although numerous markers have showed promise, there is currently insufficient evidence to warrant their use in clinical practice. Patient biases and blood sample collection have not been taken into account in clinical trials using serum biomarkers. The use of cf-mtDNA as a diagnostic and therapeutic marker for important phases in the ischemic stroke cascade should improve the accuracy of acute stroke diagnosis and provide more reliable stroke prognosis predictions.
In some earlier studies, cfDNA was detected in serum and in some in blood plasma. It has long been recognized that cfDNA is more plentiful in serum than in plasma samples, and that the amount of cfDNA in serum varies greatly from patient to patient. The coagulation cascade, which leads to the lysis of white blood cells, may produce a significant amount of cfDNA in the blood serum. As a result, this effect has the potential to induce inaccuracies into the produced results[37]. Thus, data from different sample types, such as serum, plasma, or CSF, should not be compared.
Furthermore, sampling time discrepancies significantly limit the application and coalescence of evidence. As long as the patient is admitted, sampling frequency can range from once to daily. Given that cfDNA is a transient molecule, standardizing the collection time may be advantageous in ensuring that cfDNA is captured within the time frame of rising, peaking, or declining levels. Because the therapeutic window for stroke is small following the onset of symptoms, the timing of diagnosis is critical.
The use of multiple procedures for collecting and quantifying cfDNA is another aspect that restricts the accuracy of biomarker interpretation. For DNA extraction, the majority of the research in our review used the QIAamp circulating nucleic acid kit. The QIA kit is shown to be quite efficient and yields a lot of cfDNA. While the overall goal was to measure cell death, multiple procedures were utilized across investigations, including quantitative PCR, an enzyme-linked immunosorbent assay for cell death detection, and a nucleic acid immunofluorescent counterstain. The present cfDNA assay procedure in stroke is not standardized, and there has been little research into the consistency of the quantitative PCR methods used to quantify cfDNA.
The use of cfDNA as a biomarker for stroke diagnosis and monitoring is hampered by a lack of uniformity and suitable controls. Because of changes in sample processing methods, storage conditions, and extraction and quantification methodologies, the results from different research are not comparable. This can lead to mistakes in setting cut-off points, as well as sensitivity and specificity of assays. As a result, precise and uniform quantification of cfDNA will benefit in the clinical deployment of this strategy in the future.
Circulating cfDNA levels are increased after insult of acute ischemic stroke and correlate with the clinical severity. Considering after thrombolysis and anti-platelets treatment evaluation of cfDNA may provide a crucial evidence to detect the disease severity in earlier phases of the stroke. The cfDNA is a non-invasive, cost-effective and easy to detect using simple procedures. A limited number of studies have shown that cfDNA has predictive significance in providing functional outcomes and hospital mortality. Comparability between experiments is hampered by inconsistencies in DNA extraction and measurement methods. This necessitates the performance of additional strong cohort studies in the future to determine the best collection periods for stroke prediction as well as the best cfDNA processing for the most accurate outcome. Further studies with follow-up and with window period are required to find exact severity and mortality prediction with ischemic stroke.
Provenance and peer review: Unsolicited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Clinical neurology
Country/Territory of origin: India
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): 0
Grade C (Good): C
Grade D (Fair): D
Grade E (Poor): 0
P-Reviewer: Zhang S, China; Zhu YL, China S-Editor: Gao CC L-Editor: A P-Editor: Gao CC
| 1. | Tieu PT, Lee MH, Dhawan T, Ngyen HH, Afraz S, Chung J, Khan S, Yusuf I, Liu SSH. Cell-free DNA as a potential biomarker in stroke: a comprehensive review of observational studies. J Transl Genet Genom. 2020;4:133-134. [DOI] [Full Text] |
| 2. | Donnan GA, Fisher M, Macleod M, Davis SM. Stroke. Lancet. 2008;371:1612-1623. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1829] [Cited by in RCA: 1807] [Article Influence: 106.3] [Reference Citation Analysis (0)] |
| 3. | Adogu POU, Ubajaka CF, Emelumadu OF, Alutu COC. Epidemiologic Transition of Diseases and Health Related Events in Developing Countries: A Review. Am J Med Medic Sci. 2015;5:150-157. |
| 4. | GBD 2016 Neurology Collaborators. Global, regional, and national burden of neurological disorders, 1990-2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2019;18:459-480. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3259] [Cited by in RCA: 2793] [Article Influence: 465.5] [Reference Citation Analysis (1)] |
| 5. | Kelly-Hayes M. Influence of age and health behaviors on stroke risk: lessons from longitudinal studies. J Am Geriatr Soc. 2010;58 Suppl 2:S325-S328. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 125] [Cited by in RCA: 111] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 6. | Boehme AK, Esenwa C, Elkind MS. Stroke Risk Factors, Genetics, and Prevention. Circ Res. 2017;120:472-495. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 771] [Cited by in RCA: 1028] [Article Influence: 128.5] [Reference Citation Analysis (0)] |
| 7. | Hasan N, McColgan P, Bentley P, Edwards RJ, Sharma P. Towards the identification of blood biomarkers for acute stroke in humans: a comprehensive systematic review. Br J Clin Pharmacol. 2012;74:230-240. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 85] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 8. | Glebova KV, Veiko NN, Nikonov AA, Porokhovnik LN, Kostuyk SV. Cell-free DNA as a biomarker in stroke: Current status, problems and perspectives. Crit Rev Clin Lab Sci. 2018;55:55-70. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 10] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 9. | Musuka TD, Wilton SB, Traboulsi M, Hill MD. Diagnosis and management of acute ischemic stroke: speed is critical. CMAJ. 2015;187:887-893. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 197] [Cited by in RCA: 193] [Article Influence: 19.3] [Reference Citation Analysis (0)] |
| 10. | Broughton BR, Reutens DC, Sobey CG. Apoptotic mechanisms after cerebral ischemia. Stroke. 2009;40:e331-e339. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 835] [Cited by in RCA: 963] [Article Influence: 60.2] [Reference Citation Analysis (0)] |
| 11. | Falcione SR, Jickling GC. Cell-Free DNA in Ischemic Stroke. Stroke. 2022;53:1245-1246. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 4] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 12. | Grosse GM, Blume N, Abu-Fares O, Götz F, Ernst J, Leotescu A, Gabriel MM, van Gemmeren T, Worthmann H, Lichtinghagen R, Imker R, Falk CS, Weissenborn K, Schuppner R, de Buhr N. Endogenous Deoxyribonuclease Activity and Cell-Free Deoxyribonucleic Acid in Acute Ischemic Stroke: A Cohort Study. Stroke. 2022;53:1235-1244. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 22] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 13. | Vajpeyee A, Wijatmiko T, Vajpeyee M, Taywade O, Pandey S, Chauhan PS. Clinical Usefulness of Cell-Free DNA as a Prognostic Marker in Acute Ischemic Stroke. Neurologist. 2020;25:11-13. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 14. | Leavy O. Innate immunity: Multitasking NET makers. Nat Rev Immunol. 2012;12:684-685. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 15. | Martinod K, Wagner DD. Thrombosis: tangled up in NETs. Blood. 2014;123:2768-2776. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 501] [Cited by in RCA: 622] [Article Influence: 51.8] [Reference Citation Analysis (0)] |
| 16. | Jickling GC, Sharp FR. Biomarker panels in ischemic stroke. Stroke. 2015;46:915-920. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 96] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 17. | Whiteley W, Tseng MC, Sandercock P. Blood biomarkers in the diagnosis of ischemic stroke: a systematic review. Stroke. 2008;39:2902-2909. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 141] [Cited by in RCA: 128] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 18. | Xie Q, Zhang X, Peng S, Sun J, Chen X, Deng Y, Yi L. Identification of novel biomarkers in ischemic stroke: a genome-wide integrated analysis. BMC Med Genet. 2020;21:66. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 19. | Baez SC, García Del Barco D, Hardy-Sosa A, Guillen Nieto G, Bringas-Vega ML, Llibre-Guerra JJ, Valdes-Sosa P. Scalable Bio Marker Combinations for Early Stroke Diagnosis: A Systematic Review. Front Neurol. 2021;12:638693. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 15] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 20. | Stamova B, Xu H, Jickling G, Bushnell C, Tian Y, Ander BP, Zhan X, Liu D, Turner R, Adamczyk P, Khoury JC, Pancioli A, Jauch E, Broderick JP, Sharp FR. Gene expression profiling of blood for the prediction of ischemic stroke. Stroke. 2010;41:2171-2177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 107] [Cited by in RCA: 111] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 21. | Montaner J, Perea-Gainza M, Delgado P, Ribó M, Chacón P, Rosell A, Quintana M, Palacios ME, Molina CA, Alvarez-Sabín J. Etiologic diagnosis of ischemic stroke subtypes with plasma biomarkers. Stroke. 2008;39:2280-2287. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 213] [Cited by in RCA: 227] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 22. | Laskowitz DT, Kasner SE, Saver J, Remmel KS, Jauch EC; BRAIN Study Group. Clinical usefulness of a biomarker-based diagnostic test for acute stroke: the Biomarker Rapid Assessment in Ischemic Injury (BRAIN) study. Stroke. 2009;40:77-85. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 156] [Cited by in RCA: 152] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 23. | Jickling GC, Sharp FR. Blood biomarkers of ischemic stroke. Neurotherapeutics. 2011;8:349-360. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 138] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 24. | Barr TL, Conley Y, Ding J, Dillman A, Warach S, Singleton A, Matarin M. Genomic biomarkers and cellular pathways of ischemic stroke by RNA gene expression profiling. Neurology. 2010;75:1009-1014. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 136] [Cited by in RCA: 176] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 25. | Montaner J, Mendioroz M, Ribó M, Delgado P, Quintana M, Penalba A, Chacón P, Molina C, Fernández-Cadenas I, Rosell A, Alvarez-Sabín J. A panel of biomarkers including caspase-3 and D-dimer may differentiate acute stroke from stroke-mimicking conditions in the emergency department. J Intern Med. 2011;270:166-174. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 55] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 26. | Vanni S, Polidori G, Pepe G, Chiarlone M, Albani A, Pagnanelli A, Grifoni S. Use of biomarkers in triage of patients with suspected stroke. J Emerg Med. 2011;40:499-505. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 30] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 27. | Sharma R, Macy S, Richardson K, Lokhnygina Y, Laskowitz DT. A blood-based biomarker panel to detect acute stroke. J Stroke Cerebrovasc Dis. 2014;23:910-918. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 28] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 28. | Naumann DN, Hazeldine J, Dinsdale RJ, Bishop JR, Midwinter MJ, Harrison P, Hutchings SD, Lord JM. Endotheliopathy is associated with higher levels of cell-free DNA following major trauma: A prospective observational study. PLoS One. 2017;12:e0189870. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 33] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 29. | Tsai NW, Lin TK, Chen SD, Chang WN, Wang HC, Yang TM, Lin YJ, Jan CR, Huang CR, Liou CW, Lu CH. The value of serial plasma nuclear and mitochondrial DNA levels in patients with acute ischemic stroke. Clin Chim Acta. 2011;412:476-479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 125] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 30. | Vajpeyee A, Wijatmiko T, Vajpeyee M, Taywade O. Cell free DNA: A Novel Predictor of Neurological Outcome after Intravenous Thrombolysis and/or Mechanical Thrombectomy in Acute Ischemic Stroke Patients. Neurointervention. 2018;13:13-19. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 12] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 31. | Bustamante A, Mancha F, Macher HC, García-Berrocoso T, Giralt D, Ribó M, Guerrero JM, Montaner J. Circulating cell-free DNA is a predictor of short-term neurological outcome in stroke patients treated with intravenous thrombolysis. J Circ Biomark. 2016;5:1849454416668791. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 20] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 32. | Bronkhorst AJ, Aucamp J, Pretorius PJ. Cell-free DNA: Preanalytical variables. Clin Chim Acta. 2015;450:243-253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 130] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 33. | Saffarzadeh M, Juenemann C, Queisser MA, Lochnit G, Barreto G, Galuska SP, Lohmeyer J, Preissner KT. Neutrophil extracellular traps directly induce epithelial and endothelial cell death: a predominant role of histones. PLoS One. 2012;7:e32366. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 760] [Cited by in RCA: 1016] [Article Influence: 78.2] [Reference Citation Analysis (0)] |
| 34. | O'Connell GC, Petrone AB, Tennant CS, Lucke-Wold N, Kabbani Y, Tarabishy AR, Chantler PD, Barr TL. Circulating extracellular DNA levels are acutely elevated in ischaemic stroke and associated with innate immune system activation. Brain Inj. 2017;31:1369-1375. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 50] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 35. | Jylhävä J, Jylhä M, Lehtimäki T, Hervonen A, Hurme M. Circulating cell-free DNA is associated with mortality and inflammatory markers in nonagenarians: the Vitality 90+ Study. Exp Gerontol. 2012;47:372-378. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 52] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 36. | Kananen L, Hurme M, Jylhä M, Härkänen T, Koskinen S, Stenholm S, Kähönen M, Lehtimäki T, Ukkola O, Jylhävä J. Circulating cell-free DNA level predicts all-cause mortality independent of other predictors in the Health 2000 survey. Sci Rep. 2020;10:13809. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 37. | Rainer TH, Wong LK, Lam W, Yuen E, Lam NY, Metreweli C, Lo YM. Prognostic use of circulating plasma nucleic acid concentrations in patients with acute stroke. Clin Chem. 2003;49:562-569. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 213] [Cited by in RCA: 221] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 38. | Rainer TH, Wong KS, Lam W, Lam NY, Graham CA, Lo YM. Comparison of plasma beta-globin DNA and S-100 protein concentrations in acute stroke. Clin Chim Acta. 2007;376:190-196. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 38] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 39. | Otandault A, Abraham JD, Al Amir Dache Z, Khalyfa A, Jariel-Encontre I, Forné T, Prévostel C, Chouaib S, Gozal D, Thierry AR. Hypoxia differently modulates the release of mitochondrial and nuclear DNA. Br J Cancer. 2020;122:715-725. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 20] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 40. | Trumpff C, Michelson J, Lagranha CJ, Taleon V, Karan KR, Sturm G, Lindqvist D, Fernström J, Moser D, Kaufman BA, Picard M. Stress and circulating cell-free mitochondrial DNA: A systematic review of human studies, physiological considerations, and technical recommendations. Mitochondrion. 2021;59:225-245. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 120] [Article Influence: 30.0] [Reference Citation Analysis (0)] |
| 41. | Castellani CA, Longchamps RJ, Sun J, Guallar E, Arking DE. Thinking outside the nucleus: Mitochondrial DNA copy number in health and disease. Mitochondrion. 2020;53:214-223. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 230] [Article Influence: 46.0] [Reference Citation Analysis (0)] |
| 42. | Zhou X, Chen H, Wang L, Lenahan C, Lian L, Ou Y, He Y. Mitochondrial Dynamics: A Potential Therapeutic Target for Ischemic Stroke. Front Aging Neurosci. 2021;13:721428. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 51] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 43. | Wittwer CT, Ririe KM, Andrew RV, David DA, Gundry RA, Balis UJ. The LightCycler: a microvolume multisample fluorimeter with rapid temperature control. Biotechniques. 1997;22:176-181. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 642] [Cited by in RCA: 612] [Article Influence: 21.9] [Reference Citation Analysis (0)] |