Peer-review started: September 1, 2016
First decision: October 26, 2016
Revised: November 18, 2016
Accepted: December 16, 2016
Article in press: December 19, 2016
Published online: February 6, 2017
Processing time: 143 Days and 5.7 Hours
To investigat the influence of hemochromatosis gene (Hfe) mutation on 59Fe labelled duodenal heme absorption in mice.
Heme absorption was measured in Hfe wild type and Hfe(-/-) mice by the duodenal tied loop and by oral gavage methods. The mRNA expression of heme oxygenase (HO-1), Abcg2 and Flvcr1 genes and levels were determined by quantitative polymerase chain reaction.
Heme absorption was significantly increased in homozygous Hfe(-/-) mice despite significant hepatic and splenic iron overload. While duodenal HO-1 mRNA was highly expressed in the wild type and Hfe(-/-) heme-treated group following 24 h heme administration, Flvcr1a mRNA decreased. However, Abcg2 mRNA expression levels in duodenum remained unchanged.
Heme absorption was enhanced in Hfe(-/-) mice from both duodenal tied-loop segments and by oral gavage methods. HO-1 mRNA levels were enhanced in mice duodenum after 24 h of heme feeding and may account for enhanced heme absorption in Hfe(-/-) mice. Implications for dietary recommendations on heme intake by Hfe subjects to modulate iron loading are important clinical considerations.
Core tip: These results indicate that loss of hemochromatosis gene (Hfe) protein results in increased dietary heme iron absorption that further contributes to the iron loading of the liver and other tissues of mice. Enhanced heme iron intake by homozygous Hfe subjects may contribute to body iron overload and early manifestation of phenotypic traits. This may have implications for dietary recommendations on heme intake by hemochromatosis subjects to avert tissue iron loading.
- Citation: Laftah AH, Simpson RJ, Latunde-Dada GO. Intestinal heme absorption in hemochromatosis gene knock-out mice. World J Hematol 2017; 6(1): 17-23
- URL: https://www.wjgnet.com/2218-6204/full/v6/i1/17.htm
- DOI: https://dx.doi.org/10.5315/wjh.v6.i1.17
Dietary iron intake from both heme and non-heme sources is a key homeostatic step in iron metabolism, of which deficiency or enhanced absorption is associated with iron disorders in populations all over the world[1]. Heme from animal sources contributes about 10%-25% of total food iron and has a higher bioavailability (about 15%-38%) than non-heme iron[2] in humans. The absorption mode and molecular mechanism of both forms of iron are disparate. While non-heme iron is transported by a divalent metal transporter, a proton coupled symporter, heme is presumed to be transited into the enterocytes by endocytosis (passive pinocytosis or active receptor mediation), or proteins[3,4], that are yet to be fully characterised since HCP1 is a high affinity folate transporter[5]. Internalised heme is trafficked from the cytoplasm into endosomes[6] where it is catabolised by heme oxygenase (HO-1) to yield ferrous iron that converges with the labile non-heme iron pool for transit into circulation by ferroportin, the efflux regulatory protein[7]. On the other hand, basolateral efflux of intact heme has been shown in guinea pigs, and this may be via Flvcr1 or Abcg2[8].
Hereditary hemochromatosis (HH) constitutes heterogeneous mutations of genes in the hepcidin regulatory pathway. Homozygous C282Y mutation in Hemochromatosis gene (Hfe) is predominant in about 1:300 of Caucasian populations[9]. Coincidentally these populations are, in general, avid consumers of meat and animal products. HH patients are characterized by increased heme and non-heme iron absorption from the diet[10] coupled with excessive iron accumulation in parenchyma cells of the liver and the heart. This occurs because of low hepcidin expression due to loss of function of Hfe[11,12]. Hfe protein is vital for iron-sensing in the signal transduction cascade regulating hepcidin expression. Low serum hepcidin in Hfe patients permits sustained functional expression of ferroportin. Consequently, there is enhanced efflux of non-heme and heme iron by ferroportin into circulation[13,14]. There is, however, disparity in the phenotypic expression of HH which may be due to influences of other modifier genes, dietary factors or physiological iron requirements of the subjects[15]. Consequently, iron loading in HH subjects varies in severity[16,17]. Mouse strains have been shown to modulate phenotypic variability of Hfe severity[13].
Epidemiological studies generally agree that red meat consumption leads to higher iron stores in humans[18,19]. Moreover, dietary heme iron intake was found to be associated with high serum ferritin levels in HH subjects[18]. Of particular interest, however, is the question as to whether Hfe patients could benefit from dietary modifications of iron intake during treatment by phlebotomy.
Further work is needed to elucidate the effects of the loss of Hfe on the regulation of intestinal heme absorption[20]. Mouse knock-out models, however have contributed immensely to significant advances in understanding iron metabolism and disorders. The study, therefore, set out to investigate the effects of Hfe knock-out genotype on heme absorption in mice.
Chemicals and biochemicals were of Analar grade and were from either BDH-Merck Ltd (Poole, Dorset) or Sigma Chemical Company Ltd (Poole). 59Fe (supplied as ferric chloride) was from PerkinElmer Life and Analytical Sciences (Wellesley, MA, United States, specific activity 185 GBq/g). 59Fe-heme was prepared as described in[21]. To make 59Fe-heme, a male Wistar rat was injected ip with 3.7 MBq 59Fe citrate and housed in a metabolic cage for 1 wk. The animal was bled and the red cells washed three times in 10 volumes of saline and then lysed in 10 volumes of distilled water. Heme was then isolated from the haemoglobin by crystallization using the method of Labbe and Nishada[22].
Animal care and the regulation of scientific procedures met the criteria laid down by the United Kingdom “Animals (Scientific Procedures) Act 1986”. Mice were housed in a light- and a temperature-controlled room with ad libitum access to standard pelleted diet and water unless stated. Hfe(-/-) breeders (C57/BL6 background strain; donated by Srai K, Department of Biochemistry and Molecular Biology, Royal Free and University College Medical School, London, United Kingdom) were mated and subsequently genotyped by polymerase chain reaction (PCR). Wild-type and Hfe(-/-) homozygote breeders were established to produce age-matched male mice for experimental study. Mice at 3-5 wk of age were maintained on either iron-deficient (3 mg iron per kilogram) diet ad libitum during the treatment with either arginate (control) or heme:Arginate (200 mg/L heme and 3.3 mmol/L arginate) in drinking water for 24 h.
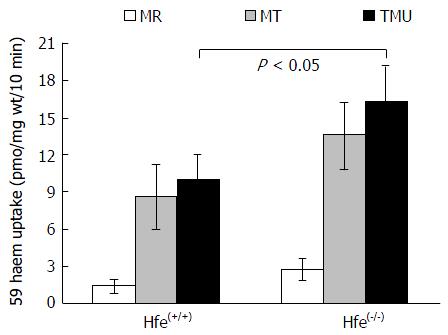
In vivo Fe absorption was measured in tied-off duodenal segments as described previously[23]. In brief, the experiments were conducted in anaesthetised mice. A duodenal segment was tied at both ends followed by the injection of 59Fe-heme arginate (100 μmol/L) into the tied-off segment. The segment was placed back into the abdominal cavity. After 10 min incubation, the duodenal segment was flushed with an ice-cold saline solution and weighed. Blood, liver and spleen were collected. Radioactivity in tissue samples and blood was measured using a gamma counter (1282 Compugamma; LKB Wallac, Turku, Finland), while carcasses were counted for radioactivity by a high-resolution bulk sample counter (J and P Engineering, Reading, United Kingdom). Radioactivity in the duodenum is referred to as mucosal retention while radioactivity in the carcass and other tissues is regarded as mucosal transfer (MT). TMU is the amount of total radioactive Fe absorbed from the gut lumen, and the percentage of MT (% MT) is the relative amount of Fe transfer into the body in comparison with total Fe uptake.
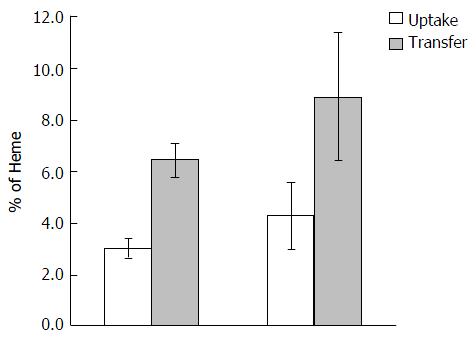
Food was withheld from the mice for 12 h prior to the oral dose, but they had free access to distilled drinking water during that period. Mice were then given 100 μL of physiological solution freshly prepared to contain heme:arginate labelled with 18 kBq 59Fe (FeCl3, in 0.1 M-hydrochloric acid, 1835 MBq/mg Fe; PerkinElmer) to provide target dosages of 4 mmol/kg body weight. This was gavaged as a single dose through the oesophagus and directly into the stomach of the animal through a 40 mm 13 gauge olive-tipped needle. No food was given to the animals after dosing and until tissue collection. The mice were then killed at approximately the same time (of 30 min) after the oral dose was administered. The abdomen was opened and after blood collection via a 1 mL syringe through a puncture into the heart, the whole gut was removed, externally rinsed, and divided into the stomach, duodenum, jejunum, ileum, caecum and colon. The lumen of each section was flushed gently with 3 mL of cold saline (9 g sodium chloride/L). Each section and the collected wash were counted for 1 min in a twin channel γ-counter (LKB, Wallac 1280, Helsinki, Finland). The carcass, minus gut, liver, spleen, kidney and blood, was counted in a high-resolution bulk sample counter for 2 min.
In the present study, it was found that 30 min were sufficient time to allow for passage of approximately 50% of the radiolabelled dose through the duodenum. Mucosal uptake of 59Fe-heme measured only in the duodenum and jejunum were defined as the proportion of the initial dose of the label retained by the carcass plus duodenal and jejunal wall after dosing. Mucosal transfer at a given time was defined as the amount of 59Fe in the carcass expressed as a percentage of the mean mucosal uptake[24].
Total RNA was extracted from tissue samples using Trizol reagent (Invitrogen, United Kingdom) according to manufacturer’s instructions. Quantitative RT-PCR was carried out using an ABI Prism 7000 detection system in a two-step protocol with SYBR Green (ABI, Life Technologies, United Kingdom). The efficacy of the amplification was confirmed by a melting curve analysis and gel electrophoresis to confirm the presence of a single product. Quantitative measurement of each gene was derived from a standard curve constructed from known amounts of PCR product. The results were calculated by the ΔCt method that expresses the difference in threshold for the target gene relative to that of 18S RNA.
Data are presented as means with their standard deviations. The comparison of multiple groups for significant effects of two variables was determined by two-way ANOVA with a Bonferroni post hoc test. P < 0.05 was considered as significant. All statistical analyses were performed using GraphPad Prism 4 software (GraphPad Software, Inc., La Jolla, CA, United States).
Heme absorption, determined by the tied loop method was greater in Hfe(-/-) knockout mice than in control wild type mice (WT) (Figure 1). This was due to a significant increase in both the uptake and transfer phases of absorption. Moreover, a similar trend of absorption was observed when heme absorption was determined by oral gavage (Figure 2). Following the oral administration of 59Fe-heme, intestinal uptake and transfer were elevated in Hfe(-/-) compared to WT after 30 min of heme administration.
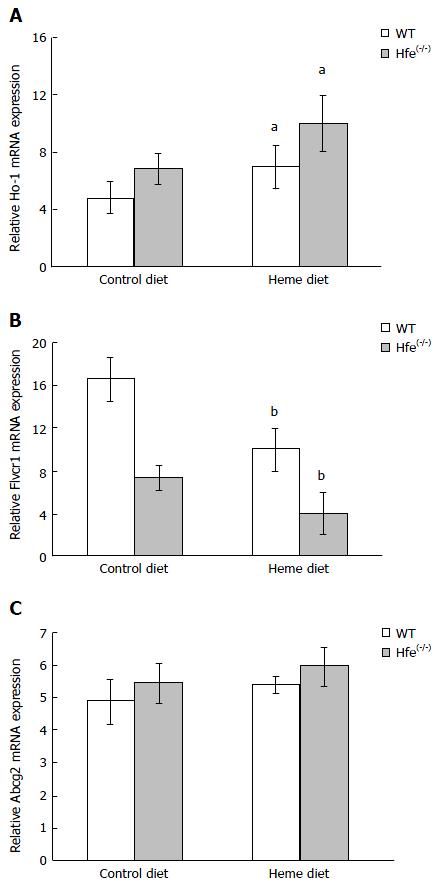
To analyze the expression of genes involved in heme metabolism and transport, WT and Hfe(-/-) mice were administered arginate (control) or heme:arginate (200 mg/L heme and 3.3 mmol/L arginate) in drinking water for 24 h. Hfe(-/-) mice treated with heme and maintained on iron-deficient diets for 24 h showed an induction of HO-1 expression (Figure 3A). The increase in HO-1 expression in the duodenum of mice on the control iron-deficient diet was not significant (Figure 3A). Flvcr1 mRNA level was lower in the duodenum of WT than Hfe(-/-) in mice fed the control iron-deficient diet. Flvcr1 mRNA levels were significantly down regulated after 24 h heme feeding in drinking water (Figure 3B). Abcg2 mRNA expression levels, however, were not significantly altered by heme feeding in drinking water (Figure 3C).
Serum and tissue iron status was determined in the mice after 24 h of heme feeding. Consistent with the literature, serum iron and transferrin saturation were significantly higher in Hfe(-/-) than WT (Table 1). Contrary to expectation, however, feeding heme to WT or Hfe(-/-) mice for 24 h showed no effect on serum iron and transferrin saturation (Table 2).
| Forward | Reverse | |
| Mouse Flvcr1 | ||
| 5’-CAGTTGATAGTCGGGTAGATCCAA-3' 5'-ACACCGGCTTCTTCAGAGTGA-3’ | ||
| Mouse Abcg2 | ||
| 5’-TCGCAGAAGGAGATGTGTTGAG-3 5CCAGAATAGCATTAAGGCCAGG-3’ | ||
| Mouse HO-1 | ||
| 5’-CAAGGAGGTACACATCCAAGCC-3' 5'-TACAAGGAAGCCATCACCAGCT-3’ | ||
| Mouse 18S | ||
| 5’-GAATTCCCAGTAAGTGGCGGG-3' 5'-GGGCAGGGACTTAATCAACG-3’ | ||
| Serum iron (μmol/L) | % Transferrin saturation | ||
| Control | 25.3 ± 4.4 | 32.1 ± 6.5 | |
| WT | Heme-24 h | 26.8 ± 3.1 | 35.9 ± 7.9 |
| Control | 49.8 ± 5.0 | 59.8 ± 6.1 | |
| Hfe(-/-) | Heme-24 h | 50.5 ± 10.6 | 61.0 ± 10.4 |
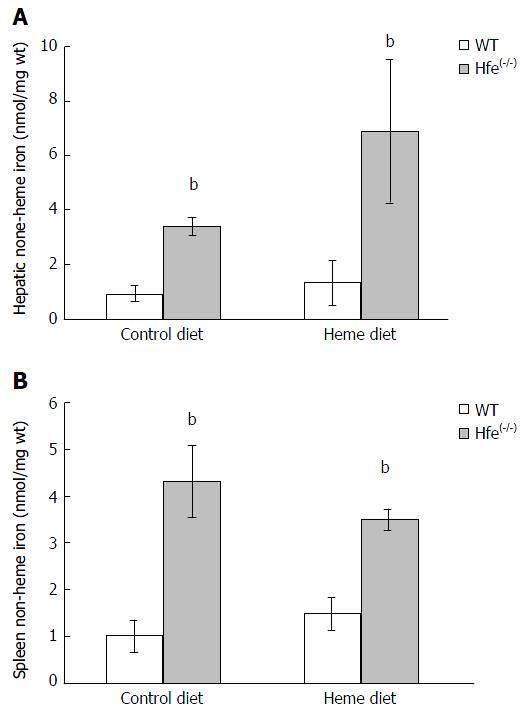
Endogenous non-heme iron levels in liver and spleen homogenates from Hfe(-/-) mice were significantly higher than WT (Figure 4; P < 0.001). Non-heme iron levels in liver homogenates were not significantly influenced by heme feeding. Although liver showed a trend towards being increased in Hfe(-/-) mice.
Heme as an exogenous source of iron is significant in nutrition because it is highly bioavailable for absorption by the gastrointestinal tract. In systemic metabolism, however, heme is derived endogenously from de novo biosynthesis for vital metabolic functions. Consequently, modulation of cytosolic, vesicular, membrane or plasma heme transport is regulated by a variety of extracellular and intracellular proteins[25,26]. While the luminal high affinity heme transport protein is not yet defined, heme absorption is enhanced in HH subjects and it is regulated by iron stores albeit by an order of magnitude less than non-heme iron absorption[10,27].
The current study demonstrates that heme feeding stimulates iron absorption in Hfe KO mice and provides evidence of increased iron storage in the spleen and hepatocytes of the mice. 59Fe-Heme arginate absorption from the duodenal loop of the mice was significantly enhanced in Hfe(-/-) mice after 10 min of exposure (Figure 1). This trend was also confirmed in mice that were given 59Fe-heme arginate by gavage and measuring absorption after 30 min (Figure 2). Hfe has been shown to have an impact on cellular iron trafficking and, indirectly, on intestinal iron absorption. The direct effects of Hfe on heme iron absorption are not clear. Alternatively, low levels of iron in the enterocyte of Hfe mice might induce heme absorption via increased HO-1 expression. It has been speculated that heme degradation by HO-1 might be the rate-liming step of heme absorption in the gut because the HO-1 activity was found to increase during Fe deficiency[28]. Increased heme iron absorption after 24 h, shown in Figure 1, might be induced by enhanced expression of HO-1 (Figure 3A). Augmented catabolism of heme by HO-1 consequently may increase the inorganic iron pool that can be chaperoned into systemic circulation. It has also been speculated that a fraction of the heme in the enterocyte might be transferred intact into the circulation by the heme efflux proteins Flvcr or Abcg2. The expression of Flvcr1 was down-regulated in Hfe mice (Figure 3B). Other modifiers such as TfR1 or TfR2 could interact with Hfe directly or, more likely, modify iron loading through independent mechanisms to increase or depress the effects of Hfe. This possibly might be due to hepatocyte regulation of hepcidin expression. A two-fold decrease was observed in hepcidin mRNA levels in the Hfe(-/-) mice used in the current study[29]. Reduced expression of hepcidin in Hfe(-/-) phenotype would lead to the maximal functional capability of FPN, hence the enhanced absorption of iron[30] in Hfe(-/-) mice. Hepcidin levels in the liver correlate negatively with serum ferritin which in humans is a biomarker of iron intake and iron status.
The increase in plasma iron and percentage transferrin saturation after feeding heme for 24 h might have contributed to increased liver and spleen non-heme iron levels in Hfe(-/-). The phenotype of HH patients of European descent attests to the higher iron absorption due to enhanced duodenal expression of transport proteins despite high iron stores[31]. Previous studies have attempted to use low iron intake and inhibitors of iron absorption as dietary strategies to ameliorate the rate of tissue Fe deposition in Hfe patients[32].
While the feeding of high heme diet did not increase serum and hepatic iron levels of both Hmox1fl/fl and Hmox1Vil-Cre mice[33], Hfe knock-out mice demonstrated increased HO-1 expression and enhanced heme absorption in the current study. Mouse strain differences have been shown to determine the severity of tissue iron deposition in Hfe knockout model of HH[13]. There might be species or strain differences in the absorption of heme iron, an earlier study however, showed that mice have the least heme absorption capacity, while canines are the highest (dog > guinea pig > rat > mouse)[34]. Moreover, to sustain heme in solution, heme arginate was used in the current study to measure absorption[35]. This study has identified HO-1 as a key candidate in the regulation of heme iron transport in the gastrointestinal tract of mice. Increased HO-1 expression in Hfe KO mice contributes to enhanced heme iron absorption.
Enhanced heme iron intake by homozygous Hfe subjects may contribute to body iron overload and early manifestation of phenotypic traits. This may have implications for dietary recommendations on heme intake by HH subjects to avert tissue iron loading. Moreover, since high intake of red meat has been associated with an elevated amount of iron in the body and increased risk of metabolic diseases, an emerging consensus, in general, suggests reduced red meat consumption by the populace.
Hemochromatosis patients are characterized with high level of heme- and inorganic iron absorption from the diet coupled with excessive iron accumulation in parenchyma cells of the liver and the heart due to low hepcidin expression.
Enhanced heme iron intake by homozygous Hfe subjects may contribute to body iron overload and early manifestation of phenotypic traits. Moreover, since high intake of red meat has been associated with elevated amount of iron in the body and increased risk of metabolic diseases, an emerging consensus, in general, suggests reduced red meat consumption by the populace.
These results indicate that loss of Hfe protein results in increased dietary heme iron absorption that further contributes to the iron loading of the liver and other tissues of mice. This may have implications for dietary recommendations on heme intake by HH subjects to avert tissue iron loading.
Implications for dietary recommendations on heme intake by Hfe subjects to modulate iron loading are important clinical considerations.
HH: Hemochromatosis; Hfe: Hemochromatosis gene; Flvcr1: Feline Leukemia Virus Subgroup C Cellular Receptor 1; HO-1: Hemoxygenase-1; Abcg2: ATP-Binding Cassette, Subfamily G, Member 2; MR: Mucosal retention; MT:Mucosal transfer,TMU:Total mucosal uptake.
It is a very well written manuscript investigating the influence of Hfe mutation on Fe labeled duodenal heme absorption in mice and showing that heme absorption was enhanced from both duodenal tied-loop segments and by oral gavage methods.
Manuscript source: Invited manuscript
Specialty type: Hematology
Country of origin: United Kingdom
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B, B, B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Borrione P, Garfield DH, Moschovi MA S- Editor: Qiu S L- Editor: A E- Editor: Li D
| 1. | Ezzati M, Lopez AD, Rodgers A, Vander Hoorn S, Murray CJ. Selected major risk factors and global and regional burden of disease. Lancet. 2002;360:1347-1360. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2283] [Cited by in RCA: 2092] [Article Influence: 91.0] [Reference Citation Analysis (0)] |
| 2. | West AR, Oates PS. Mechanisms of heme iron absorption: current questions and controversies. World J Gastroenterol. 2008;14:4101-4110. [PubMed] |
| 3. | Parmley RT, Barton JC, Conrad ME, Austin RL, Holland RM. Ultrastructural cytochemistry and radioautography of hemoglobin--iron absorption. Exp Mol Pathol. 1981;34:131-144. [PubMed] |
| 4. | Wyllie JC, Kaufman N. An electron microscopic study of heme uptake by rat duodenum. Lab Invest. 1982;47:471-476. [PubMed] |
| 5. | Qiu A, Jansen M, Sakaris A, Min SH, Chattopadhyay S, Tsai E, Sandoval C, Zhao R, Akabas MH, Goldman ID. Identification of an intestinal folate transporter and the molecular basis for hereditary folate malabsorption. Cell. 2006;127:917-928. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 616] [Cited by in RCA: 604] [Article Influence: 33.6] [Reference Citation Analysis (0)] |
| 6. | Yuan X, Protchenko O, Philpott CC, Hamza I. Topologically conserved residues direct heme transport in HRG-1-related proteins. J Biol Chem. 2012;287:4914-4924. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 55] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 7. | Nemeth E, Tuttle MS, Powelson J, Vaughn MB, Donovan A, Ward DM, Ganz T, Kaplan J. Hepcidin regulates cellular iron efflux by binding to ferroportin and inducing its internalization. Science. 2004;306:2090-2093. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3282] [Cited by in RCA: 3527] [Article Influence: 168.0] [Reference Citation Analysis (0)] |
| 8. | Conrad ME, Weintraub LR, Sears DA, Crosby WH. Absorption of hemoglobin iron. Am J Physiol. 1966;211:1123-1130. [PubMed] |
| 9. | Merryweather-Clarke AT, Pointon JJ, Shearman JD, Robson KJ. Global prevalence of putative haemochromatosis mutations. J Med Genet. 1997;34:275-278. [PubMed] |
| 10. | Lynch SR, Skikne BS, Cook JD. Food iron absorption in idiopathic hemochromatosis. Blood. 1989;74:2187-2193. [PubMed] |
| 11. | Ahmad KA, Ahmann JR, Migas MC, Waheed A, Britton RS, Bacon BR, Sly WS, Fleming RE. Decreased liver hepcidin expression in the Hfe knockout mouse. Blood Cells Mol Dis. 2002;29:361-366. [PubMed] |
| 12. | Wallace DF, Summerville L, Crampton EM, Frazer DM, Anderson GJ, Subramaniam VN. Combined deletion of Hfe and transferrin receptor 2 in mice leads to marked dysregulation of hepcidin and iron overload. Hepatology. 2009;50:1992-2000. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 164] [Cited by in RCA: 157] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 13. | Fleming RE, Holden CC, Tomatsu S, Waheed A, Brunt EM, Britton RS, Bacon BR, Roopenian DC, Sly WS. Mouse strain differences determine severity of iron accumulation in Hfe knockout model of hereditary hemochromatosis. Proc Natl Acad Sci USA. 2001;98:2707-2711. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 147] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 14. | Lynch SR, Skikne BS, Cook JD. Food iron absorption in idiopathic hemochromatosis. Blood. 1989;74:2187-2193. [PubMed] |
| 15. | Ajioka RS, Levy JE, Andrews NC, Kushner JP. Regulation of iron absorption in Hfe mutant mice. Blood. 2002;100:1465-1469. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 64] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 16. | Levy JE, Montross LK, Andrews NC. Genes that modify the hemochromatosis phenotype in mice. J Clin Invest. 2000;105:1209-1216. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 173] [Cited by in RCA: 177] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 17. | Fleming RE, Sly WS. Mechanisms of iron accumulation in hereditary hemochromatosis. Annu Rev Physiol. 2002;64:663-680. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 122] [Cited by in RCA: 105] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 18. | van der A DL, Peeters PH, Grobbee DE, Roest M, Voorbij HA, van der Schouw YT. HFE genotypes and dietary heme iron: no evidence of strong gene-nutrient interaction on serum ferritin concentrations in middle-aged women. Nutr Metab Cardiovasc Dis. 2006;16:60-68. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 25] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 19. | Ye K, Cao C, Lin X, O’Brien KO, Gu Z. Natural selection on HFE in Asian populations contributes to enhanced non-heme iron absorption. BMC Genet. 2015;16:61. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 19] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 20. | Moretti D, van Doorn GM, Swinkels DW, Melse-Boonstra A. Relevance of dietary iron intake and bioavailability in the management of HFE hemochromatosis: a systematic review. Am J Clin Nutr. 2013;98:468-479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 23] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 21. | Follett JR, Suzuki YA, Lönnerdal B. High specific activity heme-Fe and its application for studying heme-Fe metabolism in Caco-2 cell monolayers. Am J Physiol Gastrointest Liver Physiol. 2002;283:G1125-G1131. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 12] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 22. | Labbe RF, Nishida G. A new method of hemin isolation. Biochim Biophys Acta. 1957;26:437. [PubMed] |
| 23. | Masaratana P, Laftah AH, Latunde-Dada GO, Vaulont S, Simpson RJ, McKie AT. Iron absorption in hepcidin1 knockout mice. Br J Nutr. 2011;105:1583-1591. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 13] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 24. | Raja KB, Simpson RJ, Peters TJ. Comparison of 59Fe3+ uptake in vitro and in vivo by mouse duodenum. Biochim Biophys Acta. 1987;901:52-60. [PubMed] |
| 25. | Morello N, Tonoli E, Logrand F, Fiorito V, Fagoonee S, Turco E, Silengo L, Vercelli A, Altruda F, Tolosano E. Haemopexin affects iron distribution and ferritin expression in mouse brain. J Cell Mol Med. 2009;13:4192-4204. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 35] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 26. | Chiabrando D, Vinchi F, Fiorito V, Mercurio S, Tolosano E. Heme in pathophysiology: a matter of scavenging, metabolism and trafficking across cell membranes. Front Pharmacol. 2014;5:61. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 216] [Cited by in RCA: 312] [Article Influence: 28.4] [Reference Citation Analysis (0)] |
| 27. | Wheby MS, Spyker DA. Hemoglobin iron absorption kinetics in the iron-deficient dog. Am J Clin Nutr. 1981;34:1686-1693. [PubMed] |
| 28. | Raffin SB, Woo CH, Roost KT, Price DC, Schmid R. Intestinal absorption of hemoglobin iron-heme cleavage by mucosal heme oxygenase. J Clin Invest. 1974;54:1344-1352. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 153] [Cited by in RCA: 137] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 29. | Latunde-Dada GO, Simpson RJ, Mckie AT. Intestinal absorption of iron (2): Fe(II) and heme iron. in ‘Iron Metabolism and Disease’, Fuchs, H. (ed), Research Signpost, Kerala, India, pp 31-51, 2008. . |
| 30. | Stuart KA, Anderson GJ, Frazer DM, Powell LW, McCullen M, Fletcher LM, Crawford DH. Duodenal expression of iron transport molecules in untreated haemochromatosis subjects. Gut. 2003;52:953-959. [PubMed] |
| 31. | Hutchinson C, Conway RE, Bomford A, Hider RC, Powell JJ, Geissler CA. Post-prandial iron absorption in humans: comparison between HFE genotypes and iron deficiency anaemia. Clin Nutr. 2008;27:258-263. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 32. | Bezwoda WR, Disler PB, Lynch SR, Charlton RW, Torrance JD, Derman D, Bothwell TH, Walker RB, Mayet F. Patterns of food iron absorption in iron-deficient white and indian subjects and in venesected haemochromatotic patients. Br J Haematol. 1976;33:425-436. [PubMed] |
| 33. | Fillebeen C, Gkouvatsos K, Fragoso G, Calvé A, Garcia-Santos D, Buffler M, Becker C, Schümann K, Ponka P, Santos MM. Mice are poor heme absorbers and do not require intestinal Hmox1 for dietary heme iron assimilation. Haematologica. 2015;100:e334-e337. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 37] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 34. | Bannerman RM. Quantitative aspects of hemoglobin-iron absorption. J Lab Clin Med. 1965;65:944-950. [PubMed] |
| 35. | Shayeghi M, Latunde-Dada GO, Oakhill JS, Laftah AH, Takeuchi K, Halliday N, Khan Y, Warley A, McCann FE, Hider RC. Identification of an intestinal heme transporter. Cell. 2005;122:789-801. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 558] [Cited by in RCA: 488] [Article Influence: 24.4] [Reference Citation Analysis (0)] |