Published online Jul 24, 2021. doi: 10.5306/wjco.v12.i7.507
Peer-review started: February 5, 2021
First decision: March 31, 2021
Revised: April 13, 2021
Accepted: June 2, 2021
Article in press: June 2, 2021
Published online: July 24, 2021
Processing time: 166 Days and 5.6 Hours
Esophageal squamous cell carcinoma (ESCC) is a highly malignant disease that has a poor prognosis. Its high lethality is mainly due to the lack of symptoms at early stages, which culminates in diagnosis at a late stage when the tumor has already metastasized. Unfortunately, the common cancer biomarkers have low sensitivity and specificity in esophageal cancer. Therefore, a better understanding of the molecular mechanisms underlying ESCC progression is needed to identify novel diagnostic markers and therapeutic targets for intervention. The invasion of cancer cells into the surrounding tissue is a crucial step for metastasis. During metastasis, tumor cells can interact with extracellular components and secrete proteolytic enzymes to remodel the surrounding tumor microenvironment. Proteoglycans are one of the major components of extracellular matrix. They are involved in multiple processes of cancer cell invasion and metastasis by inter
Core Tip: Esophageal squamous cell carcinoma (ESCC) is a highly malignant human cancer because of its early metastasis and late diagnosis. Cancer metastasis involves multiple steps that involve cell-cell and cell-matrix interactions in the tumor microenvironment. Proteoglycans are one of the components of the extracellular matrix that play an important role in cell-matrix interactions. We herein summarize the proteo
- Citation: Zhu Y, Cheung ALM. Proteoglycans and their functions in esophageal squamous cell carcinoma. World J Clin Oncol 2021; 12(7): 507-521
- URL: https://www.wjgnet.com/2218-4333/full/v12/i7/507.htm
- DOI: https://dx.doi.org/10.5306/wjco.v12.i7.507
Esophageal cancer is the 7th most common cancer in the world and the sixth highest ranking cancer in terms of mortality rate[1]. The two major subtypes of esophageal cancer are esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC). People who develop esophageal cancer usually have no specific symptoms at the early stage. The onset of symptoms is often accompanied by difficulty and pain in swallowing (dysphagia), loss of body weight, and heartburn, by which time the tumor is likely already in the advanced stage. Late diagnosis of esophageal cancer is the primary reason for its high lethality. Tobacco use and alcohol con
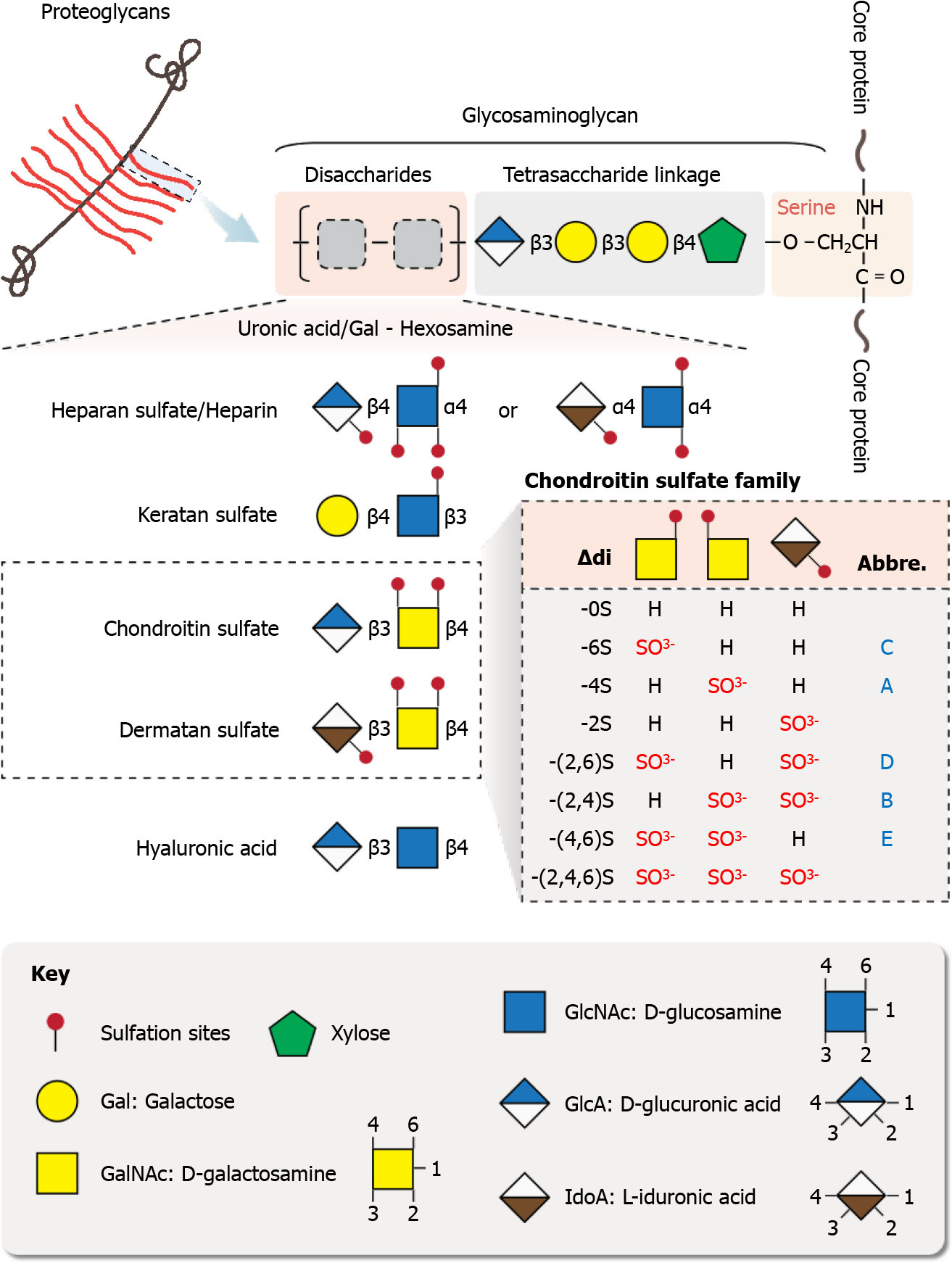
Proteoglycans typically consist of polysaccharide chains termed glycosaminoglycan (GAG) and a core protein. The GAGs are covalently attached to the serine residues on the core protein. Proteoglycans differ from glycoproteins in several aspects. For example, the carbohydrate content in proteoglycans is 50%-60%, which is much higher than that in glycoproteins. The GAGs in proteoglycans are linear, negatively-charged long chains, whereas the oligosaccharides in glycoproteins are branched short chains that may or may not be negatively-charged. The GAGs of proteoglycans are composed of repeating disaccharide units of hexuronic acid and hexosamine. They may be modified with sulfate groups at various positions to achieve multiple biological functions[13]. The major categories of GAGs are chondroitin sulfate (CS), dermatan sulfate (DS), keratan sulfate (KS), heparan sulfate (HS), heparin, and hyaluronan (HA).
CSs consist of repeating disaccharide units of N-acetylgalactosamine (GalNAc) and D-glucuronic acid (GlcA), and can be sulfated at C4 and/or C6 position of the GalNAc unit. The CS chains that contain sulfate group modification at both C4 and C6 positions of the GalNAc unit are named CS-E, whereas those modified exclusively at C4 position of GalNAc are known as CS-A. CS-C contains sulfate group only at C6 position of the GalNAc. CS-D chains have sulfate groups at C6 position of the GalNAc and C2 position of the GlcA. DS chains, also named CS-B, are derived from CS, of which the GlcA residues are epimerized into L-iduronic acid (IdoA). DS chains are sulfated at C2 position of IdoA and C4/C6 position of the GalNAc. KS chains contain repeating disaccharide units of galactose (Gal) and N-acetylglucosamine (GlcNAc). HS chains consist of GlcA and GlcNAc repeats, while heparins consist of repeating disaccharide units of IdoA and GlcNAc. The sulfate modifications in HS and heparins are in clusters. HA, a linear, protein-free, non-sulfated GAG consists of GlcNAc and GlcA repeating units. The structures of GAG chains are shown in Figure 1.
Proteoglycans can be classified according to their predominant GAG types into CS proteoglycans and HS proteoglycans or according to their cellular and subcellular location, overall gene/protein homology, and the presence of specific protein modules within their respective protein cores into intracellular, cell-surface, pericellular, and extracellular proteoglycans[14]. In this review, the functions of proteoglycans in ESCC will be discussed according to this classification. These proteoglycans are classified and summarized in Table 1.
| Location | Classification | Gene symbol | Eponym | Predominant GAG | Studied in ESCC | Ref. |
| Intracellular | Secretory granules | SRGN | Serglycin | Heparin/CS | Serglycin | Zhu et al[57] |
| Cell-surface | Transmembrane | SDC | Syndecan 1-4 | HS/CS | Syndecan-1, -2 | Mikami et al[58], Szumilo et al[60] and Huang et al[61] |
| CSPG4 | NG2 | CS/DS | ||||
| TGFBR3 | Betaglycan | HS/CS | ||||
| PTPRZ1 | Phosphacan | CS/DS | ||||
| GPI-anchored | GPC | Glypican 1-6 | HS | Glypican-1, -3 | Hara et al[62], Li et al[63], Harada et al[64] and Song[65] | |
| Pericellular | Basement membrane zone | HSPG2 | Perlecan | HS/CS | ||
| AGRN | Agrin | HS | ||||
| COL18A1 | Collagen XVIII | HS | Endostatin | Zheng et al[71], Zhong et al[72] and Chen et al[73] | ||
| COL15A1 | Collagen XV | CS/HS | ||||
| Extracellular | Hyalectans | ACAN | Aggrecan | CS/KS | ||
| VCAN | Versican | CS/DS | ||||
| NCAN | Neurocan | CS/DS | ||||
| BCAN | Brevican | CS/DS | ||||
| SLRPs, Canonical Class I | DCN | Decorin | CS/DS | Decorin | Wu et al[83], Ji et al[84] and Augoff et al[85] | |
| BGN | Biglycan | CS/DS | Biglycan | Zhu et al[97] | ||
| ASPN | Asporin | |||||
| ECM2 | ECM2 | |||||
| ECMX | ECMX | |||||
| SLRPs, Canonical Class II | FMOD | Fibromodulin | KS | |||
| LUM | Lumican | KS | Lumican | Kashyap et al[102,103] | ||
| PRELP | PRELP | |||||
| KERA | Keratocan | KS | ||||
| OMD | Osteoadherin | KS | ||||
| SLRPs, Canonical Class III | EPYC | Epiphycan | CS/DS | |||
| OPTC | Opticin | |||||
| OGN | Osteoglycin | |||||
| SLRPs, Non-Canonical Class IV | CHAD | Chondroadherin | ||||
| NYX | Nyctalopin | |||||
| TSKU | Tsukushi | |||||
| SLRPs, Non-Canonical Class V | PODN | Podocan | ||||
| PODNL1 | Podocan-Like 1 | |||||
| SPOCK | SPOCK | Testican 1-3 | HS | Testican-1 | Song et al[108] |
Serglycin is the only known intracellular proteoglycan. It was first isolated from rat yolk carcinoma cell line L2 (named as pPG1) in 1985[15]. Human SRGN (serglycin) was isolated from promyelocytic leukemia HL-60 cells in 1990[16] and from hemato
The human SRGN gene is located on chromosome 10 and spans about 16.7 kb with 7% of the gene comprising of exons[16,17,21]. Nicodemus et al[16] showed that human SRGN gene contains three exons, with exon 1 encoding signal peptide (amino acids 1-27), exon 2 encoding amino acids 28-77, and exon 3 encoding amino acids 77-158, which includes the serine/glycine repeat region (amino acids 94-111). The serine/gly
Serglycin was initially characterized as a hematopoietic proteoglycan[23] and was subsequently found to be present in many other non-hematopoietic cells such as endothelial cells[24], immune cells[25], chondrocytes[26-28], and cancer cells[29,30]. The type and size of the GAG chains of serglycin vary among different cell types and can affect the functions of serglycin. Serglycin synthesized by rat serosal mast cells is mainly modified with heparin or HS, while CS chains are predominant in rat mucosal-like mast cells[31]. HS/CS hybrid GAG chains are found in mouse mastocytoma cells and human erythroleukemia cells[32]. In human umbilical vein endothelial cells, serglycin is modified with CS-GAG chains and has smaller GAG chains than that expressed in platelets[24,33]. In human platelets, the predominant type of GAG chain is CS-4[18].
The enzymes involved in the synthesis of GAG chains have different functions in the particular physiological process of cells. Two enzymes that synthesize CS, namely chondroitin 4-sulfotransferase-1 and GalNAc(4S)-6-O-sulfotansferase, and N-deacety
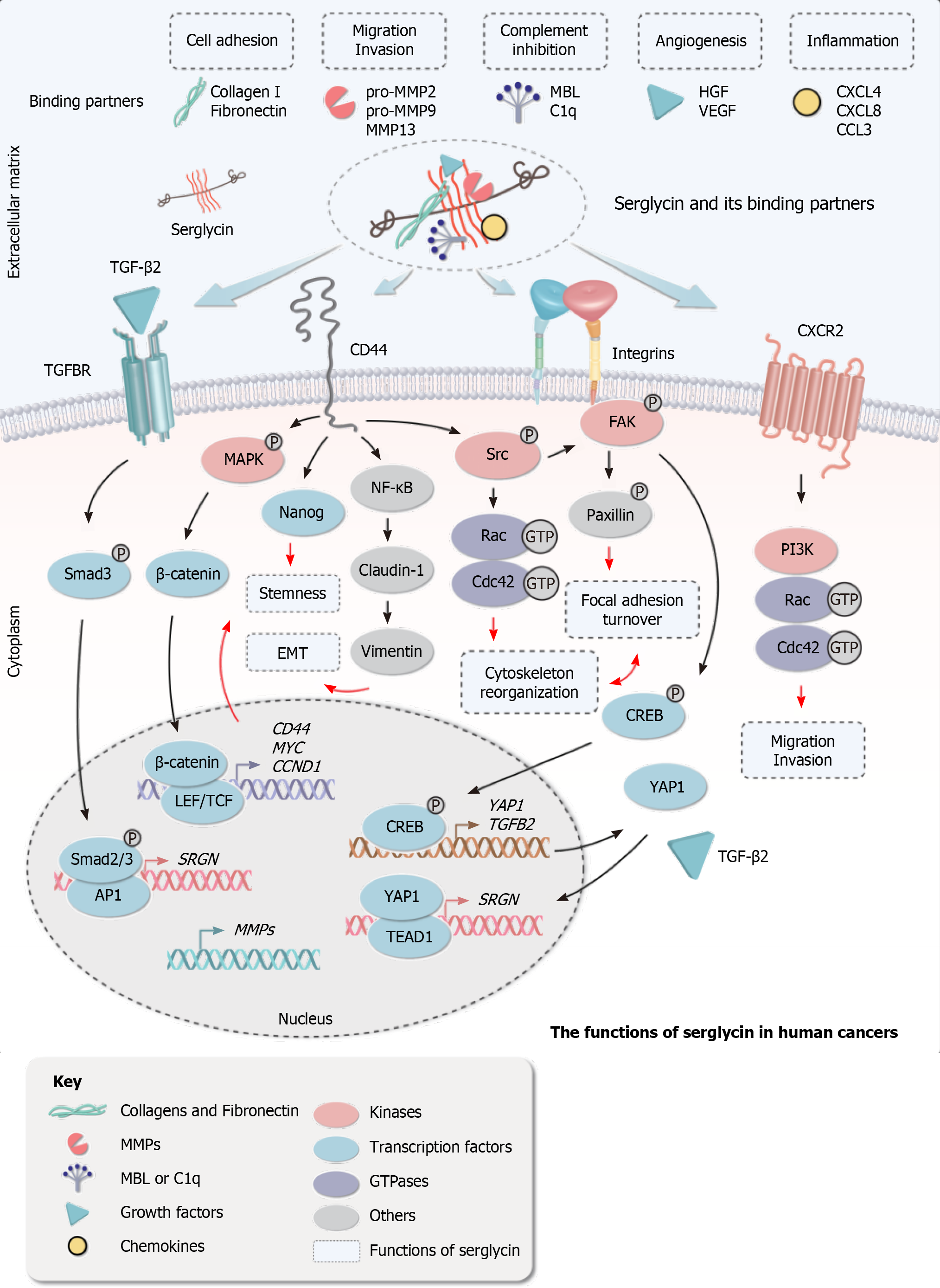
In the past two decades, an ever-increasing number of studies have shown that serglycin plays a significant role in human cancers. It has been reported that serglycin is increased in many human cancers including breast cancer[36-40], multiple myeloma[41-44], acute myeloid leukemia[45], nasopharyngeal carcinoma[46-48], hepatocellular carcinoma (HCC)[49-51], colon cancer[52,53], non-small cell lung cancer[54,55], and glioblastoma[56]. The reported functions of serglycin in promoting cancer progression include regulation of cell adhesion, promoting migration/invasion, inducing angioge
Our recent study showed for the first time that serum serglycin could be a potential non-invasive biomarker with prognostic significance in ESCC[57]. We found that serglycin and its binding partners (including matrix metalloproteinases) could promote ESCC cell invasion, migration, and metastasis. We also identified midkine as a novel GAG-dependent binding partner of serglycin, which contributes to ESCC progression by upregulating mitogen-activated protein kinase/extracellular signal-regulated protein kinase signaling and c-Myc expression in an autocrine manner[57]. This study also highlighted the significance of CS-GAGs of serglycin in ESCC cell invasion. Strategies that target serglycin, its binding partners, or its GAG chains where most of the protein interactions take place may be a new direction in ESCC therapy.
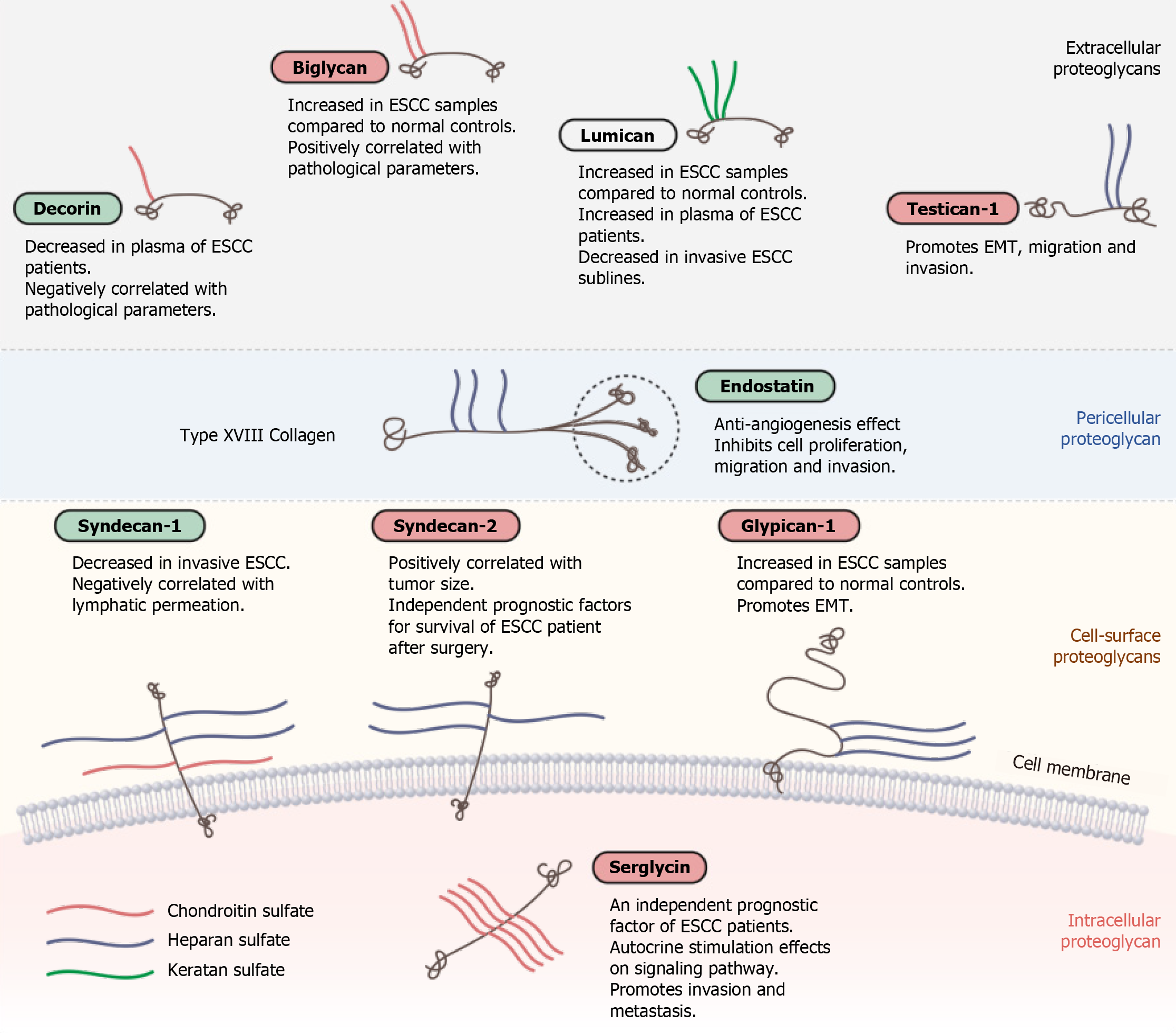
The syndecan family is a group of transmembrane proteoglycans consisting of syndecan-1, -2, -3, and -4. They generally carry HS-GAG chains. In normal esophageal epithelium, syndecan-1 is strongly expressed in cell membrane, and the expression of syndecan-1 core protein and HS-GAG chains are significantly decreased in T2 and T3 stage specimens compared with lower grade specimens[58]. Although the study by Conejo et al[59] found that syndecan-1 messenger RNA level in esophageal maligna
Glypican family includes six members, which are glypican-1 to -6. They also carry HS-GAG chains. Glypican-1 was found to be upregulated in ESCC cell lines compared with normal epithelial cells, and its expression in ESCC tissue is negatively correlated with survival rate of patients[62]. Another study confirmed that ESCC tumor samples have higher expression of glypican-1 than that in para-tumor tissues[63]. Functionally, glypican-1 promotes cell motility and induces epithelial-mesenchymal transition (EMT) in ESCC, possibly through activation of AKT/β-catenin pathway[63]. Knock
Unlike glypican-1, glypican-3 did not show significant correlation with histological type, tumor stage, tumor grade, or patient survival in ESCC[66,67], although it was reported to be a diagnostic molecule and a therapeutic target in HCC[68,69].
Pericellular proteoglycans include perlecan, agrin, collagen XVIII, and collagen XV (Table 1). The functions of perlecan, agrin, and collagen XV in ESCC have not been elucidated. Endostatin, which is a 20 kDa C-terminal fragment of collagen XVIII, was found to have anti-angiogenic activity[70]. It was later shown to have inhibitory effect on formation of ESCC-related lymphatic vessels[71]. The application of recombinant endostatin protein combined with chemoradiotherapy in ESCC treatment increased the overall survival rate of patients[72]. Recombinant endostatin combined with radiotherapy could significantly inhibit proliferation and migration/invasion of ESCC cells as well as reduce angiogenesis, but there was no effect on cell apoptosis[73].
Decorin, also called PG40, belongs to the small leucine-rich proteoglycan (SLRP) family[74]. The core protein of decorin is about 42 kDa. There is a single GAG chain attached to the N-terminus of the core protein[75]. Proteoglycans belonging to SLRP family contain a region with leucine-rich tandem repeats (LRR). The LRR region is modified by N-glycosylation. N-glycosylation and the O-linked GAG side chains are crucial for the interactions of decorin with other molecules. Studies have shown that the DS-GAGs are essential in fibrillar network formation through bridging collagen fibers[76,77]. The GAG chains and LRR region of SLRPs are both involved in ECM assembly.
Decorin was found to be necessary for appropriate fibrillogenesis due to its ability to bind to collagen[74]. The significance of decorin, as well as other SLRPs, in ECM assembly has been intensively investigated and reviewed[78]. Of note, same classes of SLRPs have the same function of binding to collagen and therefore compete with each other. For example, two other class I SLRPs, namely biglycan and asporin, are able to compensate for the loss of decorin[79]. Asporin can compensate for both decorin and biglycan loss[80]. Lumican, a class II SLRP, can compete with fibromodulin, which belongs to the same class[81,82].
The concentration of plasma decorin in 275 ESCC patients was found to be signi
Biglycan, a SLRP protein coded by the BGN gene, is structurally related to decorin but holds two GAG chains rather than one at the N-terminus of the core protein. The molecular weight of biglycan core protein is about 42 kDa. Although biglycan shares similar structure with decorin, the functions of biglycan in human cancers differ from that of decorin. In ESCC, the gene expression of BGN is upregulated in tumor samples compared with non-tumor tissues[96,97], although there is no significant association with patient survival[98]. High expression of biglycan in tumor tissue is positively correlated with tumor invasion, lymph node metastasis, and advanced clinical stage[96,97] and is an independent prognostic marker of ESCC[97]. In addition, higher serum biglycan was found in patients with EAC[99], suggesting that biglycan has diagnostic significance in esophageal cancer. Functionally, biglycan has anti-apoptotic effects on mesangial cells and pro-angiogenesis effects on tumor endothelial cells[100], which contribute to cancer cell survival and metastasis. These studies suggest that targeting biglycan may be a novel approach in anti-angiogenic and anti-tumor therapy for ESCC patients[97,100].
Lumican is a class II SLRP that has up to three KS-GAG chains. There are contradictory reports on the roles of lumican in human cancers. In pancreatic cancer, patients with high expression of stromal lumican have favorable survival after surgery[101]. How
Testican-1, also known as proteoglycan 1, belongs to the SPOCK family and is encoded by the SPOCK1 gene. It has been reported to be able to induce EMT in gastric cancer[106]. In colorectal cancer, knockdown of SPOCK1 could significantly reduce cell proliferation and invasion through the inhibition of phosphatidylinositol-3-kinase/ AKT pathway[107]. In ESCC, upregulation of SPOCK1 also induces EMT and pro
This article reviews the classification and structure of proteoglycans (Table 1 and Figure 1) and the functions of proteoglycans related to ESCC (Figure 3). As one of the components of the ECM, proteoglycans have been studied in several kinds of human cancers due to their roles in matrix organization and regulation of tumor cell-matrix interactions. Proteoglycans have diagnostic and prognostic significance in ESCC and can modulate ESCC cell migration/invasion and EMT. The secreted serglycin can also activate cell signaling in an autocrine manner (Figure 2). In addition, the significance of GAGs attached to the core protein of proteoglycans (e.g., serglycin and syndecan-1) and the expression level of GAGs regulated by multiple enzymes are increasingly gaining attention. Proteoglycans can enhance or inhibit the activity of soluble factors through interacting with them, and such interactions depend largely on the GAG chains. To date, most studies on proteoglycans in ESCC have focused primarily on their diagnostic and/or prognostic significance. In order to utilize or target these dynamic molecules in designing new strategies for treatment of this cancer, more in-depth research is needed to decipher the complex roles of proteoglycans in ESCC, especially their interactions with other ECM components, receptors, and soluble factors.
Manuscript source: Invited manuscript
Specialty type: Oncology
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Li JC S-Editor: Gao CC L-Editor: Filipodia P-Editor: Yuan YY
| 1. | Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394-424. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53206] [Cited by in RCA: 55838] [Article Influence: 7976.9] [Reference Citation Analysis (132)] |
| 2. | Song Y, Li L, Ou Y, Gao Z, Li E, Li X, Zhang W, Wang J, Xu L, Zhou Y, Ma X, Liu L, Zhao Z, Huang X, Fan J, Dong L, Chen G, Ma L, Yang J, Chen L, He M, Li M, Zhuang X, Huang K, Qiu K, Yin G, Guo G, Feng Q, Chen P, Wu Z, Wu J, Zhao J, Luo L, Fu M, Xu B, Chen B, Li Y, Tong T, Wang M, Liu Z, Lin D, Zhang X, Yang H, Zhan Q. Identification of genomic alterations in oesophageal squamous cell cancer. Nature. 2014;509:91-95. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 721] [Cited by in RCA: 855] [Article Influence: 77.7] [Reference Citation Analysis (0)] |
| 3. | Gao YB, Chen ZL, Li JG, Hu XD, Shi XJ, Sun ZM, Zhang F, Zhao ZR, Li ZT, Liu ZY, Zhao YD, Sun J, Zhou CC, Yao R, Wang SY, Wang P, Sun N, Zhang BH, Dong JS, Yu Y, Luo M, Feng XL, Shi SS, Zhou F, Tan FW, Qiu B, Li N, Shao K, Zhang LJ, Xue Q, Gao SG, He J. Genetic landscape of esophageal squamous cell carcinoma. Nat Genet. 2014;46:1097-1102. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 557] [Cited by in RCA: 553] [Article Influence: 50.3] [Reference Citation Analysis (0)] |
| 4. | Zhang L, Zhou Y, Cheng C, Cui H, Cheng L, Kong P, Wang J, Li Y, Chen W, Song B, Wang F, Jia Z, Li L, Yang B, Liu J, Shi R, Bi Y, Zhang Y, Zhao Z, Hu X, Yang J, Li H, Gao Z, Chen G, Huang X, Yang X, Wan S, Chen C, Li B, Tan Y, Chen L, He M, Xie S, Li X, Zhuang X, Wang M, Xia Z, Luo L, Ma J, Dong B, Zhao J, Song Y, Ou Y, Li E, Xu L, Xi Y, Li G, Xu E, Liang J, Guo J, Chen X, Li Q, Liu L, Zhang X, Yang H, Lin D, Cheng X, Guo Y, Zhan Q, Cui Y. Genomic analyses reveal mutational signatures and frequently altered genes in esophageal squamous cell carcinoma. Am J Hum Genet. 2015;96:597-611. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 225] [Cited by in RCA: 276] [Article Influence: 27.6] [Reference Citation Analysis (0)] |
| 5. | Lin DC, Hao JJ, Nagata Y, Xu L, Shang L, Meng X, Sato Y, Okuno Y, Varela AM, Ding LW, Garg M, Liu LZ, Yang H, Yin D, Shi ZZ, Jiang YY, Gu WY, Gong T, Zhang Y, Xu X, Kalid O, Shacham S, Ogawa S, Wang MR, Koeffler HP. Genomic and molecular characterization of esophageal squamous cell carcinoma. Nat Genet. 2014;46:467-473. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 497] [Cited by in RCA: 504] [Article Influence: 45.8] [Reference Citation Analysis (0)] |
| 6. | Cui Y, Chen H, Xi R, Cui H, Zhao Y, Xu E, Yan T, Lu X, Huang F, Kong P, Li Y, Zhu X, Wang J, Zhu W, Ma Y, Zhou Y, Guo S, Zhang L, Liu Y, Wang B, Xi Y, Sun R, Yu X, Zhai Y, Wang F, Yang J, Yang B, Cheng C, Liu J, Song B, Li H, Wang Y, Zhang Y, Cheng X, Zhan Q, Liu Z. Whole-genome sequencing of 508 patients identifies key molecular features associated with poor prognosis in esophageal squamous cell carcinoma. Cell Res. 2020;30:902-913. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 171] [Article Influence: 34.2] [Reference Citation Analysis (0)] |
| 7. | Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646-674. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51728] [Cited by in RCA: 47143] [Article Influence: 3367.4] [Reference Citation Analysis (5)] |
| 8. | Multhaupt HA, Leitinger B, Gullberg D, Couchman JR. Extracellular matrix component signaling in cancer. Adv Drug Deliv Rev. 2016;97:28-40. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 121] [Cited by in RCA: 133] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 9. | Palumbo A Jr, Meireles Da Costa N, Pontes B, Leite de Oliveira F, Lohan Codeço M, Ribeiro Pinto LF, Nasciutti LE. Esophageal Cancer Development: Crucial Clues Arising from the Extracellular Matrix. Cells. 2020;9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 40] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 10. | Theocharis AD, Skandalis SS, Gialeli C, Karamanos NK. Extracellular matrix structure. Adv Drug Deliv Rev. 2016;97:4-27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1153] [Cited by in RCA: 1591] [Article Influence: 176.8] [Reference Citation Analysis (0)] |
| 11. | Theocharis AD, Manou D, Karamanos NK. The extracellular matrix as a multitasking player in disease. FEBS J. 2019;286:2830-2869. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 165] [Cited by in RCA: 291] [Article Influence: 48.5] [Reference Citation Analysis (0)] |
| 12. | Theocharis AD, Karamanos NK. Proteoglycans remodeling in cancer: Underlying molecular mechanisms. Matrix Biol. 2019;75-76:220-259. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 152] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 13. | Soares da Costa D, Reis RL, Pashkuleva I. Sulfation of Glycosaminoglycans and Its Implications in Human Health and Disorders. Annu Rev Biomed Eng. 2017;19:1-26. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 170] [Cited by in RCA: 240] [Article Influence: 30.0] [Reference Citation Analysis (0)] |
| 14. | Iozzo RV, Schaefer L. Proteoglycan form and function: A comprehensive nomenclature of proteoglycans. Matrix Biol. 2015;42:11-55. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 880] [Cited by in RCA: 856] [Article Influence: 85.6] [Reference Citation Analysis (0)] |
| 15. | Bourdon MA, Oldberg A, Pierschbacher M, Ruoslahti E. Molecular cloning and sequence analysis of a chondroitin sulfate proteoglycan cDNA. Proc Natl Acad Sci USA. 1985;82:1321-1325. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 151] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 16. | Nicodemus CF, Avraham S, Austen KF, Purdy S, Jablonski J, Stevens RL. Characterization of the human gene that encodes the peptide core of secretory granule proteoglycans in promyelocytic leukemia HL-60 cells and analysis of the translated product. J Biol Chem. 1990;265:5889-5896. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 35] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 17. | Humphries DE, Nicodemus CF, Schiller V, Stevens RL. The human serglycin gene. Nucleotide sequence and methylation pattern in human promyelocytic leukemia HL-60 cells and T-lymphoblast Molt-4 cells. J Biol Chem. 1992;267:13558-13563. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 26] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 18. | Okayama M, Oguri K, Fujiwara Y, Nakanishi H, Yonekura H, Kondo T, Ui N. Purification and characterization of human platelet proteoglycan. Biochem J. 1986;233:73-81. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 41] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 19. | Périn JP, Bonnet F, Maillet P, Jollès P. Characterization and N-terminal sequence of human platelet proteoglycan. Biochem J. 1988;255:1007-1013. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 35] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 20. | Alliel PM, Périn JP, Maillet P, Bonnet F, Rosa JP, Jollès P. Complete amino acid sequence of a human platelet proteoglycan. FEBS Lett. 1988;236:123-126. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 28] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 21. | Stevens RL, Avraham S, Gartner MC, Bruns GA, Austen KF, Weis JH. Isolation and characterization of a cDNA that encodes the peptide core of the secretory granule proteoglycan of human promyelocytic leukemia HL-60 cells. J Biol Chem. 1988;263:7287-7291. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 60] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 22. | Kolset SO, Tveit H. Serglycin--structure and biology. Cell Mol Life Sci. 2008;65:1073-1085. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 148] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 23. | Matsumoto R, Sali A, Ghildyal N, Karplus M, Stevens RL. Packaging of proteases and proteoglycans in the granules of mast cells and other hematopoietic cells. A cluster of histidines on mouse mast cell protease 7 regulates its binding to heparin serglycin proteoglycans. J Biol Chem. 1995;270:19524-19531. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 100] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 24. | Schick BP, Gradowski JF, San Antonio JD. Synthesis, secretion, and subcellular localization of serglycin proteoglycan in human endothelial cells. Blood. 2001;97:449-458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 76] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 25. | Kolset SO, Pejler G. Serglycin: a structural and functional chameleon with wide impact on immune cells. J Immunol. 2011;187:4927-4933. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 128] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 26. | Zhang L, Yang M, Yang D, Cavey G, Davidson P, Gibson G. Molecular interactions of MMP-13 C-terminal domain with chondrocyte proteins. Connect Tissue Res. 2010;51:230-239. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 39] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 27. | Scuruchi M, D'Ascola A, Avenoso A, Mandraffino G G, Campo S S, Campo GM. Serglycin as part of IL-1β induced inflammation in human chondrocytes. Arch Biochem Biophys. 2019;669:80-86. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 21] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 28. | D'Ascola A, Scuruchi M, Avenoso A, Bruschetta G, Campo S, Mandraffino G, Campo GM. Serglycin is involved in inflammatory response in articular mouse chondrocytes. Biochem Biophys Res Commun. 2018;499:506-512. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 20] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 29. | Korpetinou A, Skandalis SS, Labropoulou VT, Smirlaki G, Noulas A, Karamanos NK, Theocharis AD. Serglycin: at the crossroad of inflammation and malignancy. Front Oncol. 2014;3:327. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 102] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 30. | Manou D, Karamanos NK, Theocharis AD. Tumorigenic functions of serglycin: Regulatory roles in epithelial to mesenchymal transition and oncogenic signaling. Semin Cancer Biol. 2020;62:108-115. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 32] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 31. | Tantravahi RV, Stevens RL, Austen KF, Weis JH. A single gene in mast cells encodes the core peptides of heparin and chondroitin sulfate proteoglycans. Proc Natl Acad Sci USA. 1986;83:9207-9210. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 67] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 32. | Lidholt K, Eriksson I, Kjellén L. Heparin proteoglycans synthesized by mouse mastocytoma contain chondroitin sulphate. Biochem J. 1995;311 ( Pt 1):233-238. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 15] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 33. | Schick BP, Pestina TI, San Antonio JD, Stenberg PE, Jackson CW. Decreased serglycin proteoglycan size is associated with the platelet alpha granule storage defect in Wistar Furth hereditary macrothrombocytopenic rats. Serglycin binding affinity to type I collagen is unaltered. J Cell Physiol. 1997;172:87-93. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 34. | Duelli A, Rönnberg E, Waern I, Ringvall M, Kolset SO, Pejler G. Mast cell differentiation and activation is closely linked to expression of genes coding for the serglycin proteoglycan core protein and a distinct set of chondroitin sulfate and heparin sulfotransferases. J Immunol. 2009;183:7073-7083. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 31] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 35. | Farrugia BL, Mizumoto S, Lord MS, O'Grady RL, Kuchel RP, Yamada S, Whitelock JM. Hyaluronidase-4 is produced by mast cells and can cleave serglycin chondroitin sulfate chains into lower molecular weight forms. J Biol Chem. 2019;294:11458-11472. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 16] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 36. | Kang Y, Siegel PM, Shu W, Drobnjak M, Kakonen SM, Cordón-Cardo C, Guise TA, Massagué J. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell. 2003;3:537-549. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1946] [Cited by in RCA: 1933] [Article Influence: 87.9] [Reference Citation Analysis (0)] |
| 37. | Iida J, Dorchak J, Clancy R, Slavik J, Ellsworth R, Katagiri Y, Pugacheva EN, van Kuppevelt TH, Mural RJ, Cutler ML, Shriver CD. Role for chondroitin sulfate glycosaminoglycan in NEDD9-mediated breast cancer cell growth. Exp Cell Res. 2015;330:358-370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 50] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 38. | Korpetinou A, Skandalis SS, Moustakas A, Happonen KE, Tveit H, Prydz K, Labropoulou VT, Giannopoulou E, Kalofonos HP, Blom AM, Karamanos NK, Theocharis AD. Serglycin is implicated in the promotion of aggressive phenotype of breast cancer cells. PLoS One. 2013;8:e78157. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 50] [Cited by in RCA: 60] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 39. | Bouris P, Manou D, Sopaki-Valalaki A, Kolokotroni A, Moustakas A, Kapoor A, Iozzo RV, Karamanos NK, Theocharis AD. Serglycin promotes breast cancer cell aggressiveness: Induction of epithelial to mesenchymal transition, proteolytic activity and IL-8 signaling. Matrix Biol. 2018;74:35-51. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 51] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 40. | Zhang Z, Deng Y, Zheng G, Jia X, Xiong Y, Luo K, Qiu Q, Qiu N, Yin J, Lu M, Liu H, Gu Y, He Z. SRGN-TGFβ2 regulatory loop confers invasion and metastasis in triple-negative breast cancer. Oncogenesis. 2017;6:e360. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 45] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 41. | Skliris A, Labropoulou VT, Papachristou DJ, Aletras A, Karamanos NK, Theocharis AD. Cell-surface serglycin promotes adhesion of myeloma cells to collagen type I and affects the expression of matrix metalloproteinases. FEBS J. 2013;280:2342-2352. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 48] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 42. | Theocharis AD, Seidel C, Borset M, Dobra K, Baykov V, Labropoulou V, Kanakis I, Dalas E, Karamanos NK, Sundan A, Hjerpe A. Serglycin constitutively secreted by myeloma plasma cells is a potent inhibitor of bone mineralization in vitro. J Biol Chem. 2006;281:35116-35128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 82] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 43. | Skliris A, Happonen KE, Terpos E, Labropoulou V, Børset M, Heinegård D, Blom AM, Theocharis AD. Serglycin inhibits the classical and lectin pathways of complement via its glycosaminoglycan chains: implications for multiple myeloma. Eur J Immunol. 2011;41:437-449. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 46] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 44. | Purushothaman A, Toole BP. Serglycin proteoglycan is required for multiple myeloma cell adhesion, in vivo growth, and vascularization. J Biol Chem. 2014;289:5499-5509. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 47] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 45. | Niemann CU, Kjeldsen L, Ralfkiaer E, Jensen MK, Borregaard N. Serglycin proteoglycan in hematologic malignancies: a marker of acute myeloid leukemia. Leukemia. 2007;21:2406-2410. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 41] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 46. | Chu Q, Huang H, Huang T, Cao L, Peng L, Shi S, Zheng L, Xu L, Zhang S, Huang J, Li X, Qian C, Huang B. Extracellular serglycin upregulates the CD44 receptor in an autocrine manner to maintain self-renewal in nasopharyngeal carcinoma cells by reciprocally activating the MAPK/β-catenin axis. Cell Death Dis. 2016;7:e2456. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 40] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 47. | Chia CS, Ong WS, Li XJ, Soong YL, Chong FT, Tan HK, Soo KC, Qian CN, Teh BT, Iyer NG. Serglycin expression: An independent marker of distant metastases in nasopharyngeal carcinoma. Head Neck. 2016;38:21-28. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 13] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 48. | Li XJ, Ong CK, Cao Y, Xiang YQ, Shao JY, Ooi A, Peng LX, Lu WH, Zhang Z, Petillo D, Qin L, Bao YN, Zheng FJ, Chia CS, Iyer NG, Kang TB, Zeng YX, Soo KC, Trent JM, Teh BT, Qian CN. Serglycin is a theranostic target in nasopharyngeal carcinoma that promotes metastasis. Cancer Res. 2011;71:3162-3172. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 108] [Cited by in RCA: 122] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 49. | He J, Zeng ZC, Xiang ZL, Yang P. Mass spectrometry-based serum peptide profiling in hepatocellular carcinoma with bone metastasis. World J Gastroenterol. 2014;20:3025-3032. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 17] [Cited by in RCA: 18] [Article Influence: 1.6] [Reference Citation Analysis (1)] |
| 50. | He L, Zhou X, Qu C, Tang Y, Zhang Q, Hong J. Serglycin (SRGN) overexpression predicts poor prognosis in hepatocellular carcinoma patients. Med Oncol. 2013;30:707. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 40] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 51. | Chen H, Li Y, Chen Y, McGowan E, Lin Y. Serglycin level in peripheral circulating blood cells has prognostic significance in patients with hepatocellular carcinoma. Ann Oncol. 2019;30 Suppl 4:iv8-iv9. [RCA] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 52. | Korpetinou A, Papachristou DJ, Lampropoulou A, Bouris P, Labropoulou VT, Noulas A, Karamanos NK, Theocharis AD. Increased Expression of Serglycin in Specific Carcinomas and Aggressive Cancer Cell Lines. Biomed Res Int. 2015;2015:690721. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 28] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 53. | Suhovskih AV, Aidagulova SV, Kashuba VI, Grigorieva EV. Proteoglycans as potential microenvironmental biomarkers for colon cancer. Cell Tissue Res. 2015;361:833-844. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 54] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 54. | Guo JY, Chiu CH, Wang MJ, Li FA, Chen JY. Proteoglycan serglycin promotes non-small cell lung cancer cell migration through the interaction of its glycosaminoglycans with CD44. J Biomed Sci. 2020;27:2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 33] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 55. | Guo JY, Hsu HS, Tyan SW, Li FY, Shew JY, Lee WH, Chen JY. Serglycin in tumor microenvironment promotes non-small cell lung cancer aggressiveness in a CD44-dependent manner. Oncogene. 2017;36:2457-2471. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 95] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 56. | Roy A, Attarha S, Weishaupt H, Edqvist PH, Swartling FJ, Bergqvist M, Siebzehnrubl FA, Smits A, Pontén F, Tchougounova E. Serglycin as a potential biomarker for glioma: association of serglycin expression, extent of mast cell recruitment and glioblastoma progression. Oncotarget. 2017;8:24815-24827. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 31] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 57. | Zhu Y, Lam AKY, Shum DKY, Cui D, Zhang J, Yan DD, Li B, Xu WW, Lee NPY, Chan KT, Law S, Tsao SW, Cheung ALM. Significance of serglycin and its binding partners in autocrine promotion of metastasis in esophageal cancer. Theranostics. 2021;11:2722-2741. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 11] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 58. | Mikami S, Ohashi K, Usui Y, Nemoto T, Katsube K, Yanagishita M, Nakajima M, Nakamura K, Koike M. Loss of syndecan-1 and increased expression of heparanase in invasive esophageal carcinomas. Jpn J Cancer Res. 2001;92:1062-1073. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 82] [Cited by in RCA: 86] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 59. | Conejo JR, Kleeff J, Koliopanos A, Matsuda K, Zhu ZW, Goecke H, Bicheng N, Zimmermann A, Korc M, Friess H, Büchler MW. Syndecan-1 expression is up-regulated in pancreatic but not in other gastrointestinal cancers. Int J Cancer. 2000;88:12-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 60. | Szumilo J, Burdan F, Zinkiewicz K, Dudka J, Klepacz R, Dabrowski A, Korobowicz E. Expression of syndecan-1 and cathepsins D and K in advanced esophageal squamous cell carcinoma. Folia Histochem Cytobiol. 2009;47:571-578. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 61. | Huang X, Xiao DW, Xu LY, Zhong HJ, Liao LD, Xie ZF, Li EM. Prognostic significance of altered expression of SDC2 and CYR61 in esophageal squamous cell carcinoma. Oncol Rep. 2009;21:1123-1129. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 16] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 62. | Hara H, Takahashi T, Serada S, Fujimoto M, Ohkawara T, Nakatsuka R, Harada E, Nishigaki T, Takahashi Y, Nojima S, Miyazaki Y, Makino T, Kurokawa Y, Yamasaki M, Miyata H, Nakajima K, Takiguchi S, Morii E, Mori M, Doki Y, Naka T. Overexpression of glypican-1 implicates poor prognosis and their chemoresistance in oesophageal squamous cell carcinoma. Br J Cancer. 2016;115:66-75. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 76] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 63. | Li J, Chen Y, Zhan C, Zhu J, Weng S, Dong L, Liu T, Shen X. Glypican-1 Promotes Tumorigenesis by Regulating the PTEN/Akt/β-Catenin Signaling Pathway in Esophageal Squamous Cell Carcinoma. Dig Dis Sci. 2019;64:1493-1502. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 26] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 64. | Harada E, Serada S, Fujimoto M, Takahashi Y, Takahashi T, Hara H, Nakatsuka R, Sugase T, Nishigaki T, Saito Y, Hiramatsu K, Nojima S, Mitsuo R, Ohkawara T, Morii E, Mori M, Doki Y, Kaneda Y, Naka T. Glypican-1 targeted antibody-based therapy induces preclinical antitumor activity against esophageal squamous cell carcinoma. Oncotarget. 2017;8:24741-24752. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 47] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 65. | Song Q. Glypican-1 as a Therapy Target in Esophageal Squamous Cell Carcinoma. Dig Dis Sci. 2019;64:3355-3356. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 66. | Mounajjed T, Zhang L, Wu TT. Glypican-3 expression in gastrointestinal and pancreatic epithelial neoplasms. Hum Pathol. 2013;44:542-550. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 38] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 67. | Zhu Z, Friess H, Kleeff J, Wang L, Wirtz M, Zimmermann A, Korc M, Büchler MW. Glypican-3 expression is markedly decreased in human gastric cancer but not in esophageal cancer. Am J Surg. 2002;184:78-83. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 19] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 68. | Chen JJ, Xie CM, Wang CR, Wan Y, Dong ZN, Li M, Xu WW. Development of a Time-Resolved Fluorescence Immunoassay for the Diagnosis of Hepatocellular Carcinoma Based on the Detection of Glypican-3. J Fluoresc. 2017;27:1479-1485. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 16] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 69. | Zhou F, Shang W, Yu X, Tian J. Glypican-3: A promising biomarker for hepatocellular carcinoma diagnosis and treatment. Med Res Rev. 2018;38:741-767. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 144] [Cited by in RCA: 257] [Article Influence: 32.1] [Reference Citation Analysis (0)] |
| 70. | O'Reilly MS, Boehm T, Shing Y, Fukai N, Vasios G, Lane WS, Flynn E, Birkhead JR, Olsen BR, Folkman J. Endostatin: an endogenous inhibitor of angiogenesis and tumor growth. Cell. 1997;88:277-285. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3249] [Cited by in RCA: 3111] [Article Influence: 111.1] [Reference Citation Analysis (0)] |
| 71. | Zheng Y, Sun M, Chen J, He L, Zhao N, Chen K. Effect of VEGF-C siRNA and endostatin on ring formation and proliferation of esophageal squamous cell carcinoma lymphatic endothelial cells. Onco Targets Ther. 2016;9:6727-6732. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 72. | Zhong Z, Gu X, Zhang Z, Wang D, Qing Y, Li M, Dai N. Recombinant human endostatin combined with definitive chemoradiotherapy as primary treatment for patients with unresectable but without systemic metastatic squamous cell carcinoma of the oesophagus. Br J Radiol. 2012;85:e1104-e1109. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 10] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 73. | Chen X, Zhang H, Zhu H, Yang X, Yang Y, Min H, Chen G, Liu J, Lu J, Cheng H, Sun X. Endostatin combined with radiotherapy suppresses vasculogenic mimicry formation through inhibition of epithelial-mesenchymal transition in esophageal cancer. Tumour Biol. 2016;37:4679-4688. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 15] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 74. | Danielson KG, Baribault H, Holmes DF, Graham H, Kadler KE, Iozzo RV. Targeted disruption of decorin leads to abnormal collagen fibril morphology and skin fragility. J Cell Biol. 1997;136:729-743. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1143] [Cited by in RCA: 1078] [Article Influence: 38.5] [Reference Citation Analysis (0)] |
| 75. | Krusius T, Ruoslahti E. Primary structure of an extracellular matrix proteoglycan core protein deduced from cloned cDNA. Proc Natl Acad Sci USA. 1986;83:7683-7687. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 352] [Cited by in RCA: 400] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 76. | Henninger HB, Maas SA, Underwood CJ, Whitaker RT, Weiss JA. Spatial distribution and orientation of dermatan sulfate in human medial collateral ligament. J Struct Biol. 2007;158:33-45. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 17] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 77. | Lewis PN, Pinali C, Young RD, Meek KM, Quantock AJ, Knupp C. Structural interactions between collagen and proteoglycans are elucidated by three-dimensional electron tomography of bovine cornea. Structure. 2010;18:239-245. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 115] [Cited by in RCA: 98] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 78. | Chen S, Birk DE. The regulatory roles of small leucine-rich proteoglycans in extracellular matrix assembly. FEBS J. 2013;280:2120-2137. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 247] [Cited by in RCA: 290] [Article Influence: 24.2] [Reference Citation Analysis (0)] |
| 79. | Kalamajski S, Aspberg A, Lindblom K, Heinegård D, Oldberg A. Asporin competes with decorin for collagen binding, binds calcium and promotes osteoblast collagen mineralization. Biochem J. 2009;423:53-59. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 99] [Cited by in RCA: 108] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 80. | Corsi A, Xu T, Chen XD, Boyde A, Liang J, Mankani M, Sommer B, Iozzo RV, Eichstetter I, Robey PG, Bianco P, Young MF. Phenotypic effects of biglycan deficiency are linked to collagen fibril abnormalities, are synergized by decorin deficiency, and mimic Ehlers-Danlos-like changes in bone and other connective tissues. J Bone Miner Res. 2002;17:1180-1189. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 338] [Cited by in RCA: 322] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 81. | Svensson L, Närlid I, Oldberg A. Fibromodulin and lumican bind to the same region on collagen type I fibrils. FEBS Lett. 2000;470:178-182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 141] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 82. | Kalamajski S, Oldberg Å. Homologous sequence in lumican and fibromodulin leucine-rich repeat 5-7 competes for collagen binding. J Biol Chem. 2009;284:534-539. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 84] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 83. | Wu IC, Wu DC, Huang CC, Lin HS, Chen YK, Tsai HJ, Lu CY, Chou SH, Chou YP, Li LH, Tai SY, Wu MT. Plasma decorin predicts the presence of esophageal squamous cell carcinoma. Int J Cancer. 2010;127:2138-2146. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 29] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 84. | Ji C, Liu H, Xiang M, Liu J, Yue F, Wang W, Chu X. Deregulation of decorin and FHL1 are associated with esophageal squamous cell carcinoma progression and poor prognosis. Int J Clin Exp Med. 2015;8:20965-20970. [PubMed] |
| 85. | Augoff K, Grabowski K, Rabczynski J, Kolondra A, Tabola R, Sikorski AF. Expression of decorin in esophageal cancer in relation to the expression of three isoforms of transforming growth factor-beta (TGF-beta1, -beta2, and -beta3) and matrix metalloproteinase-2 activity. Cancer Invest. 2009;27:443-452. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 15] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 86. | Seidler DG, Goldoni S, Agnew C, Cardi C, Thakur ML, Owens RT, McQuillan DJ, Iozzo RV. Decorin protein core inhibits in vivo cancer growth and metabolism by hindering epidermal growth factor receptor function and triggering apoptosis via caspase-3 activation. J Biol Chem. 2006;281:26408-26418. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 148] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 87. | Santra M, Eichstetter I, Iozzo RV. An anti-oncogenic role for decorin. Down-regulation of ErbB2 leads to growth suppression and cytodifferentiation of mammary carcinoma cells. J Biol Chem. 2000;275:35153-35161. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 148] [Cited by in RCA: 157] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 88. | Reed CC, Waterhouse A, Kirby S, Kay P, Owens RT, McQuillan DJ, Iozzo RV. Decorin prevents metastatic spreading of breast cancer. Oncogene. 2005;24:1104-1110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 151] [Cited by in RCA: 166] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 89. | Markmann A, Hausser H, Schönherr E, Kresse H. Influence of decorin expression on transforming growth factor-beta-mediated collagen gel retraction and biglycan induction. Matrix Biol. 2000;19:631-636. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 68] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 90. | Neill T, Schaefer L, Iozzo RV. Decorin as a multivalent therapeutic agent against cancer. Adv Drug Deliv Rev. 2016;97:174-185. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 113] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 91. | Neill T, Schaefer L, Iozzo RV. Oncosuppressive functions of decorin. Mol Cell Oncol. 2015;2:e975645. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 52] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 92. | Neill T, Schaefer L, Iozzo RV. Decoding the Matrix: Instructive Roles of Proteoglycan Receptors. Biochemistry. 2015;54:4583-4598. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 82] [Cited by in RCA: 104] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 93. | Morrione A, Neill T, Iozzo RV. Dichotomy of decorin activity on the insulin-like growth factor-I system. FEBS J. 2013;280:2138-2149. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 53] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 94. | Sainio AO, Järveläinen HT. Decorin-mediated oncosuppression - a potential future adjuvant therapy for human epithelial cancers. Br J Pharmacol. 2019;176:5-15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 36] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 95. | Järvinen TAH, Ruoslahti E. Generation of a multi-functional, target organ-specific, anti-fibrotic molecule by molecular engineering of the extracellular matrix protein, decorin. Br J Pharmacol. 2019;176:16-25. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 33] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 96. | Wong FH, Huang CY, Su LJ, Wu YC, Lin YS, Hsia JY, Tsai HT, Lee SA, Lin CH, Tzeng CH, Chen PM, Chen YJ, Liang SC, Lai JM, Yen CC. Combination of microarray profiling and protein-protein interaction databases delineates the minimal discriminators as a metastasis network for esophageal squamous cell carcinoma. Int J Oncol. 2009;34:117-128. [PubMed] |
| 97. | Zhu YH, Yang F, Zhang SS, Zeng TT, Xie X, Guan XY. High expression of biglycan is associated with poor prognosis in patients with esophageal squamous cell carcinoma. Int J Clin Exp Pathol. 2013;6:2497-2505. [PubMed] |
| 98. | Zhao SF, Yin XJ, Zhao WJ, Liu LC, Wang ZP. Biglycan as a potential diagnostic and prognostic biomarker in multiple human cancers. Oncol Lett. 2020;19:1673-1682. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 25] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 99. | Zaidi AH, Gopalakrishnan V, Kasi PM, Zeng X, Malhotra U, Balasubramanian J, Visweswaran S, Sun M, Flint MS, Davison JM, Hood BL, Conrads TP, Bergman JJ, Bigbee WL, Jobe BA. Evaluation of a 4-protein serum biomarker panel-biglycan, annexin-A6, myeloperoxidase, and protein S100-A9 (B-AMP)-for the detection of esophageal adenocarcinoma. Cancer. 2014;120:3902-3913. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 37] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 100. | Yamamoto K, Ohga N, Hida Y, Maishi N, Kawamoto T, Kitayama K, Akiyama K, Osawa T, Kondoh M, Matsuda K, Onodera Y, Fujie M, Kaga K, Hirano S, Shinohara N, Shindoh M, Hida K. Biglycan is a specific marker and an autocrine angiogenic factor of tumour endothelial cells. Br J Cancer. 2012;106:1214-1223. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 75] [Cited by in RCA: 85] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 101. | Li X, Truty MA, Kang Y, Chopin-Laly X, Zhang R, Roife D, Chatterjee D, Lin E, Thomas RM, Wang H, Katz MH, Fleming JB. Extracellular lumican inhibits pancreatic cancer cell growth and is associated with prolonged survival after surgery. Clin Cancer Res. 2014;20:6529-6540. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 71] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 102. | Kashyap MK, Marimuthu A, Kishore CJ, Peri S, Keerthikumar S, Prasad TS, Mahmood R, Rao S, Ranganathan P, Sanjeeviah RC, Vijayakumar M, Kumar KV, Montgomery EA, Kumar RV, Pandey A. Genomewide mRNA profiling of esophageal squamous cell carcinoma for identification of cancer biomarkers. Cancer Biol Ther. 2009;8:36-46. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 102] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 103. | Kashyap MK, Marimuthu A, Peri S, Kumar GS, Jacob HK, Prasad TS, Mahmood R, Kumar KV, Kumar MV, Meltzer SJ, Montgomery EA, Kumar RV, Pandey A. Overexpression of periostin and lumican in esophageal squamous cell carcinoma. Cancers (Basel). 2010;2:133-142. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 16] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 104. | Wang X, Peng Y, Xie M, Gao Z, Yin L, Pu Y, Liu R. Identification of extracellular matrix protein 1 as a potential plasma biomarker of ESCC by proteomic analysis using iTRAQ and 2D-LC-MS/MS. Proteomics Clin Appl. 2017;11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 17] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 105. | Li B, Xu WW, Lam AKY, Wang Y, Hu HF, Guan XY, Qin YR, Saremi N, Tsao SW, He QY, Cheung ALM. Significance of PI3K/AKT signaling pathway in metastasis of esophageal squamous cell carcinoma and its potential as a target for anti-metastasis therapy. Oncotarget. 2017;8:38755-38766. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 64] [Cited by in RCA: 81] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 106. | Kim HP, Han SW, Song SH, Jeong EG, Lee MY, Hwang D, Im SA, Bang YJ, Kim TY. Testican-1-mediated epithelial-mesenchymal transition signaling confers acquired resistance to lapatinib in HER2-positive gastric cancer. Oncogene. 2014;33:3334-3341. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 93] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 107. | Zhao P, Guan HT, Dai ZJ, Ma YG, Liu XX, Wang XJ. Knockdown of SPOCK1 Inhibits the Proliferation and Invasion in Colorectal Cancer Cells by Suppressing the PI3K/Akt Pathway. Oncol Res. 2016;24:437-445. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 38] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 108. | Song X, Han P, Liu J, Wang Y, Li D, He J, Gong J, Li M, Tu W, Yan W, Liu M, Huang H, Tian D, Liao J. Up-regulation of SPOCK1 induces epithelial-mesenchymal transition and promotes migration and invasion in esophageal squamous cell carcinoma. J Mol Histol. 2015;46:347-356. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 29] [Article Influence: 2.9] [Reference Citation Analysis (0)] |