Published online Oct 15, 2023. doi: 10.4251/wjgo.v15.i10.1823
Peer-review started: June 5, 2023
First decision: July 8, 2023
Revised: July 14, 2023
Accepted: August 28, 2023
Article in press: August 28, 2023
Published online: October 15, 2023
Processing time: 125 Days and 1.5 Hours
Multiple primary colorectal carcinoma (MPCC) is a rare clinical disease, which is challenging to differentiate from metastatic disease using histopathological methods. Next-generation sequencing (NGS) has been employed to identify multiple primary cancers.
This study a rare case of a 63-year-old male patient diagnosed with MPCC by targeted NGS, which was initially missed by radiological evaluation. The patient was found to have two tumors located on the surface of the colorectum which had distinct genomic alterations. Based on wild-type KRAS detected in the unresected tumor, the patient benefited from the epidermal growth factor receptor (EGFR) inhibitor cetuximab treatment, but developed novel mutations including KIF5B-RET fusion, which provides a possible resistance mechanism to anti-EGFR therapy.
Our case highlights the necessity of using genetic testing for primary tumor diagnosis and the application of serial plasma circulating tumor DNA profiling for dynamic disease monitoring.
Core Tip: We report a rare case of a 63-year-old male patient diagnosed with multiple primary colorectal carcinomas through targeted next-generation sequencing, which was initially missed by diagnostic imaging. The patient was found to have two tumors located on the colorectal surface which had different genomic alterations, as evidenced by immunohistochemistry staining. The patient benefited from treatment with the epidermal growth factor receptor inhibitor cetuximab due to the wild-type KRAS detected in the unresected tumor. This case emphasizes the importance of genetic testing for primary tumor diagnosis and the need for longitudinal circulating tumor DNA profiling to develop effective therapeutic strategies.
- Citation: Qu YJ, Zhang QS, Wang B, Zhang F, Pan E, Zhao CY, Liu SY, Fang LP. Comprehensive next-generation sequencing reveals double primary colorectal carcinoma missed by diagnostic imaging: A case report. World J Gastrointest Oncol 2023; 15(10): 1823-1828
- URL: https://www.wjgnet.com/1948-5204/full/v15/i10/1823.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v15.i10.1823
Colorectal cancer (CRC) is one of the most lethal and prevalent malignancies worldwide, with approximately half of CRC patients eventually developing metastatic CRC[1]. However, the occurrence of multiple primary colorectal carcinoma (MPCC) is rare (between 1.1% and 8.1%)[2]. MPCC is defined as the discovery of two or more primary colorectal carcinomas in an individual occurring either synchronously or metachronously[3]. Preoperative detection of multiple primary cancers is important when planning treatment. Nevertheless, current diagnostic criteria may not identify all MPCC patients, leading to inappropriate treatment and follow-up plans[4]. To reduce the rate of misdiagnosis or missed diagnosis, recent studies have proposed the use of molecular testing and genomic profiling[5]. Herein, we report a case of MPCC that was initially missed via imaging but was diagnosed using next-generation sequencing (NGS). The patient was found to have two tumors on the surface of the colorectum which had completely different mutation patterns. Furthermore, a KIF5B-RET fusion was identified following cetuximab resistance, which has not been previously reported in CRC.
A 63-year-old man presented with hematochezia and abdominal pain in June 2020.
He was diagnosed with colorectal carcinoma initially.
He had no major illnesses in the past.
He and his family both had no history of cancer.
The patient's vital signs were normal.
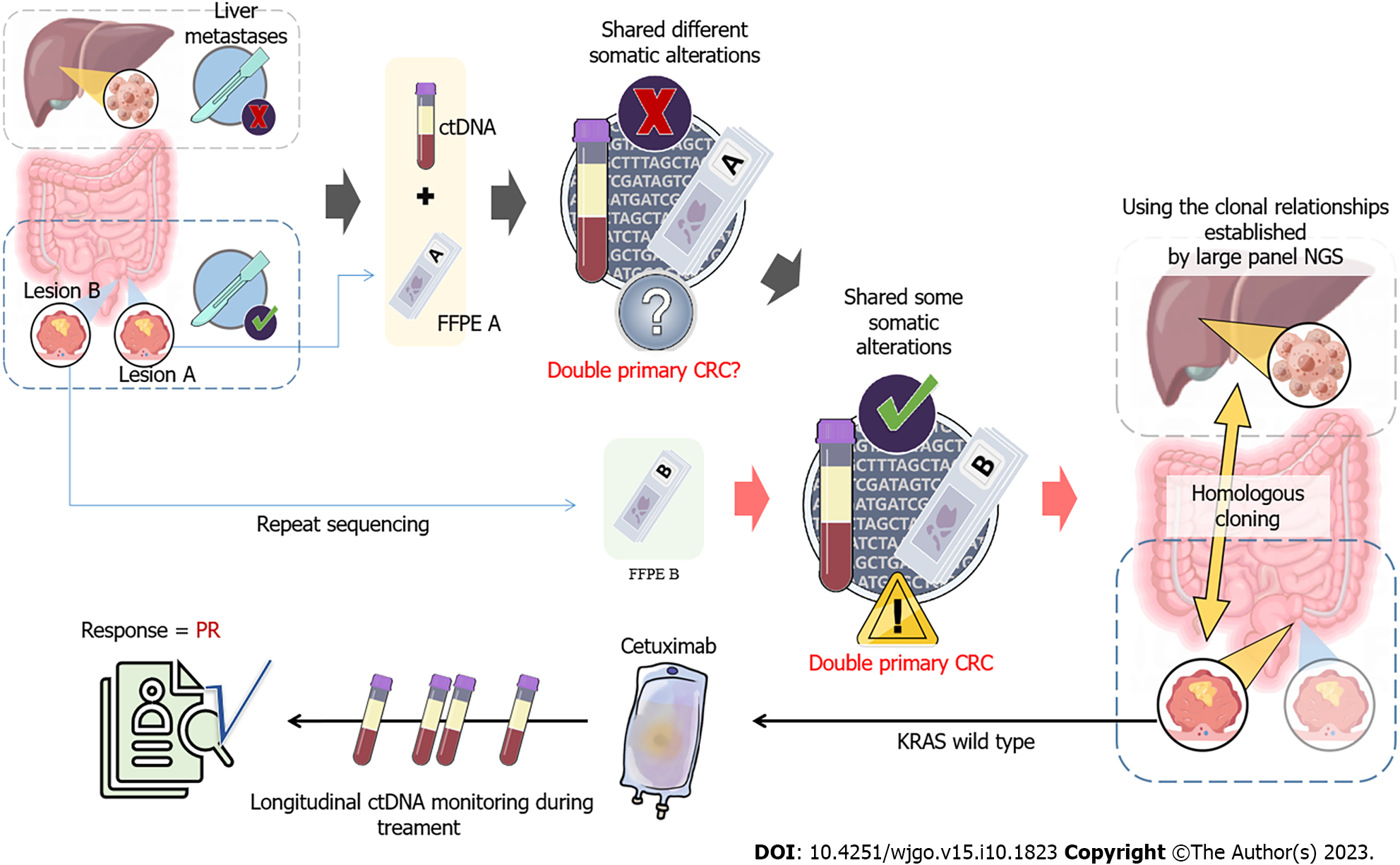
To search for an efficient therapeutic strategy, genomic DNA from the formalin-fixed paraffin-embedded sample of lesion A and circulating tumor DNA (ctDNA) from plasma were subjected to targeted NGS of 425 cancer-related genes (Nanjing Geneseeq Technology Inc.) (Figure 1A). A comparison of the genetic alterations in lesion A and B can be found in Supplementary Table 1.
The computed tomography (CT) scan showed two lesions (defined as A and B) on the liver. Endoscopic resection revealed that the two lesions were located on the surface of the colon, and only lesion A was removed (Figure 2).
He was diagnosed with stage Ⅳ (pT3N1M1) colorectal carcinoma with liver metastases in June 2020.
As the patient presented with intestinal bleeding, endoscopic resection was performed to alleviate his symptoms. Based on the identification of KRAS G12D with a mutation allelic frequency (MAF) of 41.9% identified in lesion A, the patient was administered XELOX plus bevacizumab (oxaliplatin 130 mg/m2 on day 1, capecitabine 1500 mg/m2 twice daily for 14 d, bevacizumab 7.5 mg/kg day 1) every 3 wk as first-line treatment. The patient achieved an initial partial response (PR) with sustained response ongoing for 11 mo. In January 2021, the tumor was evaluated as progressive disease (PD), and second-line chemotherapy was initiated with irinotecan (180 mg/m2 day 1), raltitrexed (3 mg/m2 day 1) and bevacizumab (5 mg/kg day 1) every 2 wk. Unfortunately, the liver lesion size increased by 35% compared to baseline, indicating a PD (Figure 1B). In April 2021, both plasma and lesion B were subjected to NGS and four identical mutation types were identified, with no KRAS mutations (Supplementary Table 1). A comparison of genomic alterations between lesion A and B revealed completely different mutation landscapes. Furthermore, immunohistochemistry illustrated significant differences between the two lesions (Supplementary Figure 1), which confirmed the diagnosis of MPCC. As the patient had wild-type KRAS, he was treated with irinotecan (180 mg/m2 day 1), raltitrexed (3 mg/m2 day 1), plus cetuximab (500 mg/m2 day 1), a monoclonal antibody that blocks the epidermal growth factor receptor (EGFR), every 2 wk in April 2021. Plasma ctDNA sequencing and CT scans were conducted every 2 wk and 2 mo, respectively (Figure 1B and C). Two months later, it was observed that the size of liver metastases had decreased by 75% compared to the previous examination and ctDNA had rapidly decreased to less than 1% (Figure 1B and D). Moreover, the tumor markers for CRC, carcinoembryonic antigen (CEA) and carbohydrate antigen 19-9 (CA19-9) also significantly decreased to normal levels (Figure 1C), indicating a PR.
In August 2021, stable disease was observed with a 4% decrease in liver metastasis size compared to the previous examination, and no significant increase in CEA and CA19-9 Levels was observed (Figure1B and C). However, the allele frequencies (AFs) of ctDNA alterations in plasma samples were considerably elevated (Figure 1D and Supple
MPCC was initially discovered by Warren and Gates in 1941[6]. Despite its rarity, the occurrence of MPCC is showing an upward trend[2]. Due to a lack of understanding regarding MPCC and limited diagnostic techniques, it is always challenging to distinguish between multiple primary cancers and tumor metastasis. The emergence of NGS has already changed the landscape of cancer studies and is now widely used in the diagnosis of multiple primary cancers[7,8]. In our case, lesions A and B, were initially misdiagnosed as a primary lesion with metastasis due to their similar features. Thus, only tumor A and plasma samples were subjected to targeted NGS, revealing a completely different mutation pattern. Further targeted NGS on lesion B showed that these two tumor lesions, A and B, did not share any mutations. Through genetic profiling, it was confirmed that lesions A and B were independent primary lesions. Of note, the molecular variations identified by NGS aided in the diagnosis of both primary tumors. It was also hypothesized that tumor B may have played a role in the development of liver metastases, but the patient declined a liver biopsy. Subsequent NGS supported this hypothesis, as there was a strong correlation between tumor B and plasma ctDNA.
Surgical intervention has long been the ideal choice for cancer patients[9], but not for patients with metastatic lesions. Our patient presented with intestinal bleeding, and we opted for surgery to ease his symptoms. For those patients who cannot undergo surgery, radiotherapy and chemotherapy are the primary methods for disease control[10]. In addition, targeted therapy is an alternative approach that has proven to be effective in prolonging the overall survival rate of CRC patients[1]. The first targeted agent for CRC approved by the Food and Drug Administration was cetuximab, a monoclonal antibody that blocks EGFR, in 2004[11]. The efficacy of anti-EGFR therapy is dependent on the mutational status of downstream signaling molecules of the EGFR pathway, such as KRAS, NRAS, PIK3CA, and BRAF. Patients with a KRAS wild-type tumor are more likely to respond to this therapy[12]. In our patient, lesion B and ctDNA showed wild-type KRAS, while lesion A, which was removed, had a KRAS G12D mutation. Therefore, cetuximab was administered and the patient benefited from this treatment, with a decrease in liver tumor size, a reduction in AF of ctDNA, and lower serum tumor markers (CEA and CA19-9). Initially, the patient was found to have drug resistance by NGS, followed by serum tumor markers and a CT scan. Previous reports indicate that serial ctDNA profiling can detect disease progression earlier than CT scans[13]. Additionally, continuous monitoring of ctDNA can provide a more accurate understanding of the tumor, which can improve personalized treatment decisions[14]. To the best of our knowledge, this is the first time KIF5B-RET fusion has been discovered in a CRC patient with resistance to cetuximab. The emerging RET fusion variant is a significant driver gene for drug resistance in multiple progressive cancers, such as non-small cell lung cancer. Zhu et al[15] reported that the emergence of the KIF5B-RET fusion gene may cause acquired resistance to EGFR-tyrosine kinase inhibitors in EGFR-mutant lung adenocarcinomas[15]. Hence, we propose that this KIF5B-RET fusion gene may be a novel factor contributing to acquired resistance to cetuximab in KRAS wild-type CRCs. Nevertheless, the patient declined treatment with pralsetinib, a targeted RET inhibitor.
The limitations of the single case presentation in this study should be noted. While the KIF5B-RET fusion is a possible resistance mechanism to cetuximab, more pre-clinical research and clinical data are required to confirm its potential. NGS is a powerful tool that can provide valuable insights into an individual's genetic composition[16]. It can help identify genomic variations potentially linked to certain diseases or conditions, allowing for earlier diagnosis and selection of more effective treatment[17]. In this particular case, NGS played a pivotal role in diagnosing MPCC and offered direction for its treatment.
In summary, we report the rare case of a 63-year-old male patient with MPCC diagnosed through genetic profiling. The patient was treated with cetuximab based on wild-type KRAS identified on the lesion and later developed novel mutations including KIF5B-RET fusion, which provides a possible resistance mechanism to anti-EGFR therapy. This case highlights the necessity of using genetic testing for identifying primary tumors and the importance of longitudinal ctDNA profiling, which may trigger the development of effective therapeutic strategies.
Provenance and peer review: Unsolicited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Oncology
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Sridharan G, India; Tanabe H, Japan S-Editor: Fan JR L-Editor: A P-Editor: Yu HG
| 1. | Xie YH, Chen YX, Fang JY. Comprehensive review of targeted therapy for colorectal cancer. Signal Transduct Target Ther. 2020;5:22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 905] [Cited by in RCA: 1014] [Article Influence: 202.8] [Reference Citation Analysis (0)] |
| 2. | Lam AK, Chan SS, Leung M. Synchronous colorectal cancer: clinical, pathological and molecular implications. World J Gastroenterol. 2014;20:6815-6820. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 118] [Cited by in RCA: 114] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 3. | Wang HZ, Huang XF, Wang Y, Ji JF, Gu J. Clinical features, diagnosis, treatment and prognosis of multiple primary colorectal carcinoma. World J Gastroenterol. 2004;10:2136-2139. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 39] [Cited by in RCA: 37] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 4. | Lee SH, Ahn BK, Baek SU. Multiple primary cancers in extracolonic sites with colorectal cancer. Int J Colorectal Dis. 2009;24:301-304. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 5. | Lin MW, Wu CT, Kuo SW, Chang YL, Yang PC. Clinicopathology and genetic profile of synchronous multiple small adenocarcinomas: implication for surgical treatment of an uncommon lung malignancy. Ann Surg Oncol. 2014;21:2555-2562. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 14] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 6. | Warren S, Gates O. Carcinoma of ceruminous gland. Am J Pathol. 1941;17:821-826.3. [PubMed] |
| 7. | Ravella L, Barritault M, Bringuier PP, Chalabreysse L, Thivolet-Bejui F, Maury JM, Duruisseaux M, Brevet M. [Multiple lung carcinoma: Primary or intrapulmonary metastasis?]. Ann Pathol. 2018;38:202-205. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 8. | Chang JC, Alex D, Bott M, Tan KS, Seshan V, Golden A, Sauter JL, Buonocore DJ, Vanderbilt CM, Gupta S, Desmeules P, Bodd FM, Riely GJ, Rusch VW, Jones DR, Arcila ME, Travis WD, Ladanyi M, Rekhtman N. Comprehensive Next-Generation Sequencing Unambiguously Distinguishes Separate Primary Lung Carcinomas From Intrapulmonary Metastases: Comparison with Standard Histopathologic Approach. Clin Cancer Res. 2019;25:7113-7125. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 75] [Article Influence: 12.5] [Reference Citation Analysis (0)] |
| 9. | Schreuders EH, Ruco A, Rabeneck L, Schoen RE, Sung JJ, Young GP, Kuipers EJ. Colorectal cancer screening: a global overview of existing programmes. Gut. 2015;64:1637-1649. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 674] [Cited by in RCA: 908] [Article Influence: 90.8] [Reference Citation Analysis (0)] |
| 10. | Brown KGM, Solomon MJ, Mahon K, O'Shannassy S. Management of colorectal cancer. BMJ. 2019;366:l4561. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 48] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 11. | Chu E. An update on the current and emerging targeted agents in metastatic colorectal cancer. Clin Colorectal Cancer. 2012;11:1-13. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 107] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 12. | Saridaki Z, Georgoulias V, Souglakos J. Mechanisms of resistance to anti-EGFR monoclonal antibody treatment in metastatic colorectal cancer. World J Gastroenterol. 2010;16:1177-1187. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 19] [Cited by in RCA: 19] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 13. | Zhang C, Chen Z, Chong X, Chen Y, Wang Z, Yu R, Sun T, Chen X, Shao Y, Zhang X, Gao J, Shen L. Clinical implications of plasma ctDNA features and dynamics in gastric cancer treated with HER2-targeted therapies. Clin Transl Med. 2020;10:e254. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 24] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 14. | Kim S, Lim Y, Kang JK, Kim HP, Roh H, Kim SY, Lee D, Bang D, Jeong SY, Park KJ, Han SW, Kim TY. Dynamic changes in longitudinal circulating tumour DNA profile during metastatic colorectal cancer treatment. Br J Cancer. 2022;127:898-907. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 23] [Reference Citation Analysis (0)] |
| 15. | Zhu YC, Wang WX, Zhang QX, Xu CW, Zhuang W, Du KQ, Chen G, Lv TF, Song Y. The KIF5B-RET Fusion Gene Mutation as a Novel Mechanism of Acquired EGFR Tyrosine Kinase Inhibitor Resistance in Lung Adenocarcinoma. Clin Lung Cancer. 2019;20:e73-e76. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 12] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 16. | Hussen BM, Abdullah ST, Salihi A, Sabir DK, Sidiq KR, Rasul MF, Hidayat HJ, Ghafouri-Fard S, Taheri M, Jamali E. The emerging roles of NGS in clinical oncology and personalized medicine. Pathol Res Pract. 2022;230:153760. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 35] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 17. | Mosele F, Remon J, Mateo J, Westphalen CB, Barlesi F, Lolkema MP, Normanno N, Scarpa A, Robson M, Meric-Bernstam F, Wagle N, Stenzinger A, Bonastre J, Bayle A, Michiels S, Bièche I, Rouleau E, Jezdic S, Douillard JY, Reis-Filho JS, Dienstmann R, André F. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: a report from the ESMO Precision Medicine Working Group. Ann Oncol. 2020;31:1491-1505. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 569] [Cited by in RCA: 761] [Article Influence: 152.2] [Reference Citation Analysis (0)] |