Published online Jan 27, 2023. doi: 10.4254/wjh.v15.i1.19
Peer-review started: September 28, 2022
First decision: October 30, 2022
Revised: November 17, 2022
Accepted: December 21, 2022
Article in press: December 21, 2022
Published online: January 27, 2023
Processing time: 109 Days and 15.2 Hours
Liver disorders are one of the most common pathological problems worldwide. It affects more than 1.5 billion worldwide. Many types of hepatic cells have been reported to be involved in the initiation and propagation of both acute and ch
Core Tip: Acute and chronic liver diseases are worldwide problems with multifactorial pathogenesis. The exact pathological mechanism of several liver disorders is still unclear. However, many suggested mechanisms are involved, including but not limited to oxidative stress, inflammation, autophagy, and microRNA. The underlying perspective mechanisms are helpful in the discovery of new and effective therapeutic interventions for this annoying problem.
- Citation: Ali FE, Abd El-Aziz MK, Sharab EI, Bakr AG. Therapeutic interventions of acute and chronic liver disorders: A comprehensive review. World J Hepatol 2023; 15(1): 19-40
- URL: https://www.wjgnet.com/1948-5182/full/v15/i1/19.htm
- DOI: https://dx.doi.org/10.4254/wjh.v15.i1.19
Chronic liver diseases are a significantly prevalent health problem contributing to the rising burden on countries daily. Specifically, liver cirrhosis - a result of chronic liver damage - is considered one of the well-known causes of morbidity and mortality all over the globe. According to Cheemerla and Balakrishnan[1], liver cirrhosis was responsible for the worldwide death of approximately 1.32 million patients in 2017. Not only that liver cirrhosis ranked 11th among the leading causes of mortality, but it has also become a habitual cause of living with a disability[2].
Acute liver injury is characterized by an abrupt decline in hepatocyte function. Unlike liver cirrhosis, acute liver failure (ALF) typically has no underlying liver problem and worsens rapidly in days or weeks. Regarding etiology, hepatitis B viral infection and medication toxicity, particularly from acetaminophen (APAP), are the primary contributors to ALF. However, other types of hepatitis, autoimmune disorders, Wilson’s disease, and cardiovascular diseases are less common suspects for ALF[3]. On the contrary, there are two classes of chronic liver injuries: Cholestatic conditions that block the bile flow and persistent hepatotoxicity. Various factors can lead to hepatotoxicity, such as hepatitis B viruses (HBV), hepatitis D viruses, and hepatitis C viruses (HCV), alcohol abuse, or non-alcoholic steatohepatitis (NASH). At the same time, biliary cholangitis, atresia of bile ducts, and primary scl
Different physiological mechanisms have been involved in liver injury, including autophagy and their different types, microRNAs (miRNAs) and their crucial effect, inflammation, hepatic cell regulation role, and the main effects of transcription factors and inflammatory cytokines. Considering the therapeutic interventions for liver diseases, there are specific treatments that are basically dependent on the cause of the disease. For instance, alcohol cessation, acetylcysteine for APAP toxicity, antiviral medication for hepatitis viruses, and immunosuppressants for autoimmune hepatitis are considered[3].
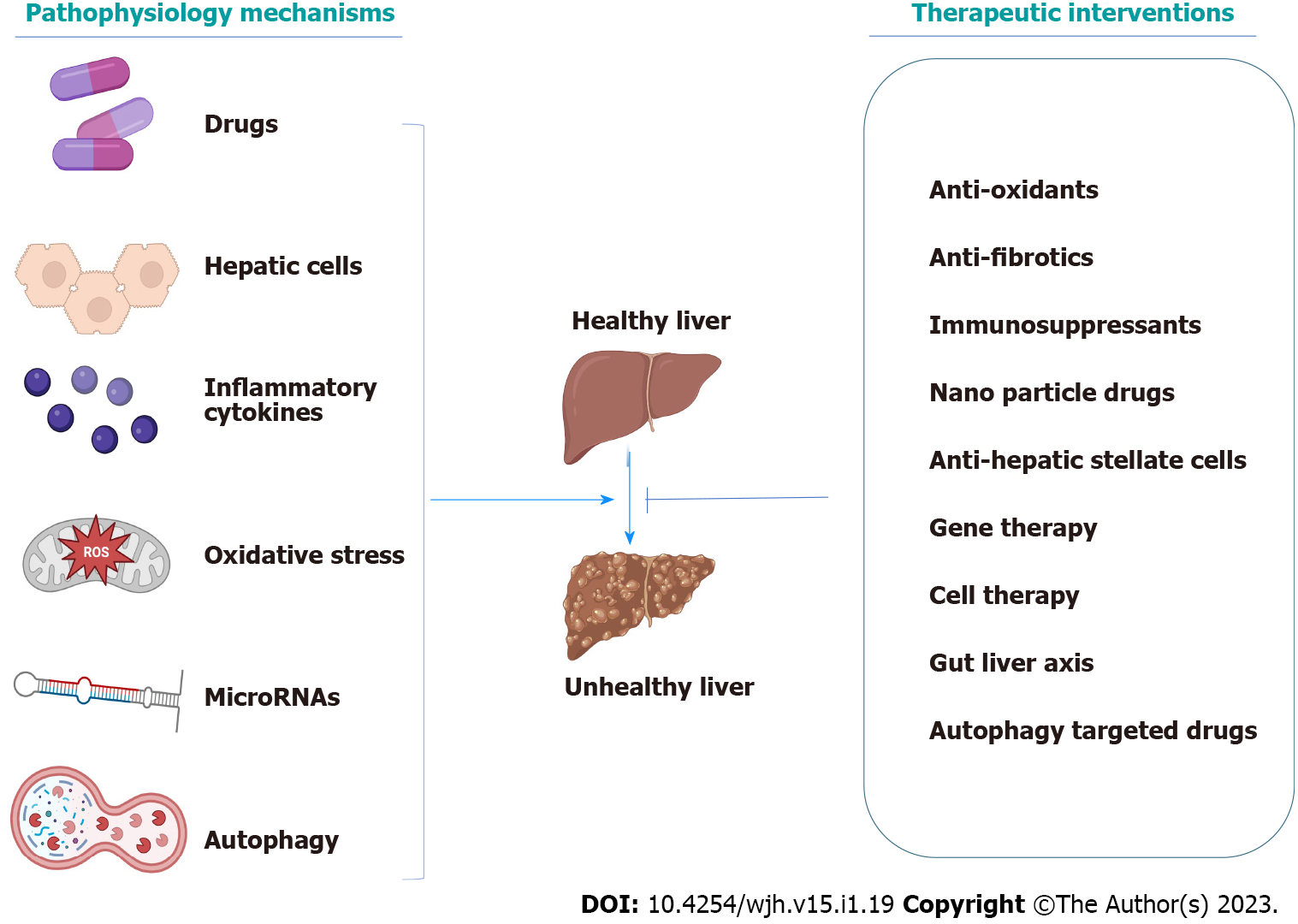
Recent studies have discussed various interventions for liver disorders, such as antifibrotic agents, cell-based therapies, gut microbiota, different nanoparticle systems, gene therapy, and much more. Consequently, we aim to discuss the newly characterized pathophysiological mechanisms and the most appropriate and recent therapy discovered to be effective on acute and chronic liver disorders (Figure 1).
Both initial liver damage and subsequent multiple organ failure (MOF) can be classified as parts of the pathophysiology of ALF. The mechanism of APAP-induced ALF is the most well-known in terms of the first liver injury. Glucuronidation and sulfation of APAP create harmless chemicals that are eliminated through the urine in nontoxic doses (4 g/d)[5]. The residual APAP is transformed into the hazardous metabolite N-acetyl-p-benzoquinone (NAPQI) by cytochrome P450 enzymes (CYPs), which is then detoxified by bringing it to glutathione (GSH)[6]. Interestingly, after overdosing on APAP, GSH is depleted after its conjugation with NAPQI, and the extra NAPQI binds to hepatocellular proteins causing mitochondrial oxidative stress and necrosis[7]. NAPQI amount is enhanced during the decrease of GSH availability which will exacerbate the toxic effects of an APAP overdose. Antibiotics, antiepileptic medications, and ethanol activate CYPs and increase NAPQI production. Reduced GSH production is a result of fasting and malnutrition[6].
Moreover, the pathogenesis of secondary MOF appears to have several characteristics in common with severe sepsis. The innate immune response is triggered early in the course of a disease. It can be a response to heterotropic viruses’ pathogen-specific molecular patterns (PAMPs) or to damage-associated molecular patterns (DAMPs), which include histones, DNA, and high mobility group box molecules-1 proteins produced from wounded cells following hepatocyte apoptosis as a result of toxic causes[8]. It is well known that the innate response involves a wide range of immune cells, such as monocytes, macrophages, dendritic cells, leukocytes, and natural killer cells. PAMPs and DAMPs are recognized by these cells, which then react and generate proinflammatory mediators like tumor necrosis factor α (TNF-α), interleukin (IL)-1, and IL-6, as well as reactive oxygen species (ROS), which trigger a systemic inflammatory response.
Additionally, IL-17 and IL-10 contribute to the overall inflammatory response[8]. Afterward, MOF is produced, and liver damage is still being brought on by ROS and cytokines. Proinflammatory cytokines entice neutrophils and encourage extravasation into the parenchyma of the liver. They begin to emit ROS and proteases once they are inside the parenchyma, which causes hepatocyte destruction. Promoting neutrophil extravasation into the hepatic parenchyma is greatly aided by mediators released from dying or dead hepatocytes and CXC chemokines. By releasing reactive oxygen intermediates and proteases once they have reached the hepatic parenchyma, neutrophils cause intracellular hepatocyte stress and oncotic necrosis[9]. The vasodilatation of the peripheral microcirculatory leads to inefficient pulmonary oxygen exchange, decreased peripheral tissue oxygen supply, and subsequently, lactic acidosis, which finally causes hypotension. The most severe effects are on cerebral and renovascular tone, which results in hemorrhage, cerebral hyper-perfusion, and functional renal failure[8]. The most common pathological mechanisms of acute and chronic liver disease are summarized in Table 1.
| Disease | Mechanism | Model | Findings | Ref. |
| ALF | N-acetyl-p-APAP | Mice model | The hazardous metabolite N-acetyl-p-benzoquinone depleted GSH and caused mitochondrial oxidative stress and necrosis | [6] |
| Innate immunity, apoptosis, and cytokine release | Bio-samples from roughly 2000 patients with ALF | Generated pro-inflammatory mediators and oxidative stress, vasodilatation of the peripheral microcirculatory, hypoxia, lactic acidosis, and hypotension | [8] | |
| MiR-122 and miR-192 | APAP in mice | Increased miR-122 and miR-192 levels after acute hepatic poisoning with acetaminophen in mice before transaminases | [82] | |
| MiRNAs | ALF in mice | Up-regulated miR-155, miR-146a, miR-125a, miR-15b, and miR-16 | [83] | |
| Down-regulated miR-1187 | ||||
| Acute liver injury | MiRNAs | Acetaminophen or carbon tetrachloride in male rats | Down-regulated miR-29c_AS, miR298, miR327, miR342, miR370, miR376c, miR494, and miR503 | [66] |
| Upregulated miR-153, miR-302b AS, miR-337, miR-363, miR-409-5p, and miR-542-3p | ||||
| MiR-122 | I/R mouse model | Elevated miR-122 level | [67] | |
| MiR-192 | APAP induced liver injury in mouse | Dose- and exposure-dependent elevation of miR-192 level | [79] | |
| HBV | MiRNAs | Pooled sera obtained from HBV patients | Up-regulated miR-122 level. miR-122 could inhibit HBV replication in Huh7 and HepG2 cells | [84] |
| MiR-155 | Human hepatoma cells | MiR-155 enhances innate antiviral immunity by promoting JAK/STAT signaling pathway by targeting SOCS1 | [86] | |
| HCV | MiR-122 | Human hepatoma Huh-7.5 cells | MiR-122 is the predominant miRNA in the liver tissue. 2’-O-methyl antisense oligonucleotide depletion of miR-122 also inhibits HCV genotype 2a replication and infectious virus production | [89] |
| MiRNAs | Human hepatoma cells | MiR-24, miR-149, miR-638, and miR-1181 were identified to be involved in HCV entry, replication, and propagation | [90] | |
| Alcoholic steatohepatitis | MiRNAs | In vitro (RAW 264.7 macrophage) and in vivo (Kupffer cells of alcohol-fed mice) | Up-regulated miR-155 expression both in vitro and in vivo | [94] |
| Increased TNF alpha production in response to miR-155 induction | ||||
| Increased expression of miR-155 and miR-132 in the total liver | ||||
| MiRNAs | Bile duct ligation rat model | Down-regulated miR-150 and miR-194 expression | [98] | |
| MiRNAs | Human stellate cell line | Up-regulated miR-199 and miR-200 led to higher expression of fibrosis-related genes in an HSC cell line | [97] | |
| NAFLD and alcoholic liver disease | Autophagy | In-vivo | Activation of macroautophagy and CMA eliminated damaged mitochondria, lessens oxidative stress, and promotes regeneration | [136] |
| Liver cancer | Autophagy | Oncogene-driven cancer models | Protein kinase C promotes autophagy and oxidative phosphorylation | |
| ROS generation, which through Nrf2 drives HCC through cell-autonomous and non-autonomous mechanisms | ||||
| Liver cirrhosis | Hepatocyte | In-vivo | Activation of hepatic stellate cells by damaged hepatocytes | [18] |
| Hepatic stellate cell | In-vivo | The activated hepatic stellate cells produce endothelin-1, TGF-β, and cytoglobin that share in the process of fibrogenesis | [24] | |
| Sinusoidal endothelial cells SECs | Co-culture with freshly isolated SECs | Differentiated SECs prevent HSC activation and promote reversion of activated HSCs to quiescence through VEGF-stimulated NO production | [32] | |
| Kupffer cells | Mouse model | Enhanced death ligand expression | [35] | |
| Inhibition of hepatocyte apoptosis with a caspase inhibitor prevented Kupffer cell activation | ||||
| Hepatic stellate cell activation |
The liver is composed of two types of cells; hepatocytes, known as parenchymal cells, which constitute most of the liver and non-parenchymal cells. Around 10% of the liver’s mass comprises non-parenchymal cells that include liver sinusoidal endothelial cells (SECs), hepatic stellate cells (HSCs), biliary cells, Kupffer cells (KCs), and immune cells such as neutrophils, natural killer cells, and infiltrating macrophages[10]. Whenever the liver is exposed to a harmful substance, both parenchymal and non-parenchymal hepatic cells take a role in the onset of liver fibrosis and cirrhosis.
Understanding the etiology of chronic liver disorders is essential for their prevention, slowing their progress, and advancing different treatment options. There are various etiologies to chronic liver disease, from alcoholic liver disease (ALD), non-alcoholic fatty liver (NAFLD), steatohepatitis, and chronic viral hepatitis, to other genetic, autoimmune, drugs, or cryptogenic liver diseases. Among the different etiologies of liver disorders, alcohol abuse is the most common cause. As a result of excessive alcohol consumption, the condition of alcoholic liver worsens to fatty liver and chronic steatohepatitis, which in turn triggers liver fibrosis, cirrhosis, or even HCC[11]. The non-ALD shares the same fate as the ALD but is correlated with metabolic syndrome[12]. Also, different types of hepatitis viruses can result in chronic liver disease, especially hepatitis B and C; hence, they are considered a major concern for cirrhosis and liver cancer[13]. Concerning the less common causes of liver disorders, genetic factors such as hemochromatosis, alpha-1 antitrypsin deficiency, Wilson’s disease, and autoimmune hepatitis can all contribute to irreversible cirrhosis[14]. Additionally, hepatotoxic drugs, primarily APAP, followed by idiosyncratic drugs inducing liver injuries, such as antibiotics, nonsteroidal anti-inflammatory drugs, herbal remedies, and statins, can cause the liver to progress to liver fibrosis and cirrhosis[15]. When the liver is exposed to any of the above-mentioned destructive agents, liver cells undergo a remodeling process to compensate for the damage.
Hepatocytes play a complex role in the progression of cirrhosis since they are the main constituent of the liver and are particularly susceptible to harm from hepatotoxic substances[16]. Hepatocytes produce most of the matrix metalloproteinases and tissue inhibitors of matrix metalloproteinases, which regulate the extracellular matrix deposition and thus participate in the process of liver cirrhosis[17]. Damaged hepatocytes activate HSCs, increase the ability of myofibroblasts to synthesize fibrous tissue, and produce ROS and other fibrogenic mediators[18]. The persistence of fibrosis induces hepatocytes to become hypoxic and produce large amounts of tumor growth factor-beta (TGF-β), a powerful stimulator of fibrogenesis[19]. Additionally, recent studies showed that hepatocyte telomere shortening and aging is a possible factor contributing to fibrosis and is thus implicated in the pathogenesis of cirrhosis[20].
HSCs are primarily in charge of regulating and storing vitamin A or retinol, and they are found in the subendothelial space between hepatocytes and SECs. ROS, cytokines, and growth factors, such as TNF-α and TGF-β, respectively, can activate these quiescent cells, causing them to synthesize a lot of extra
SECs are an important type of liver cells, surrounded by the bloodstream from one side and hepatocytes from the other side[28]. Morphologically, liver SECs are characterized by transcellular pores known as fenestrae, which are essential for transporting nutrients and other components from the blood to the hepatocytes and vice versa. Fenestrae is important for normal liver function and plays a great role in maintaining liver homeostasis and regeneration[29]. In pathological conditions, SECs lose their fenestrae and become capillarized, impairing proper liver function[30]. Furthermore, they encourage fibrogenesis by activating HSCs by releasing IL-33[31]. In contrast, several studies have documented that differentiated liver SECs can encourage the reversion of activated HSCs to the quiescent form and thus stop the progress of fibrosis via modulating vascular endothelial growth factor (VEGF)-stimulated NO release[32].
KCs are liver macrophage cells that comprise an average of 85% of body macrophages and are present in hepatic sinusoids. KCs are necessary for innate and adaptive immunity as they deal with detrimental pathogens entering the liver from the portal vein[33]. As a result of liver injury, KCs get activated and respond by producing various cytokines, ILs, and chemokines[16]. Additionally, NO produced by KCs, together with TNF-α, TGF-β, and platelet-derived growth factors (PDGFs), activate HSCs, causing an excess of extracellular matrix to be produced[34]. Although KCs produce death ligands and contribute to liver fibrogenesis and fibrosis[35], they are not a suitable target for therapeutic interventions due to their crucial host defense function.
Cytokines are bioactive molecules made by several types of liver cells that are essential in the progression of liver cirrhosis[36]. They consist of TNF-α, PDGF, interferons (IFNs), ILs, TGF-β, chemokines, and adipokines. Several important biological processes, such as hematopoiesis, immunology, inflammation, and body development, are mediated by cytokines. However, they are also linked to several illnesses, including liver disorders, rheumatoid arthritis, and atherosclerosis[37]. A significant coordinated program of cellular and molecular alterations in liver cirrhosis results in a potent fibrotic response. Cytokines are involved in the combative signaling pathways that regulate the activation of HSCs and fibrogenesis[38].
PDGF is the most powerful HSCs activator concerning all polypeptide growth factors. According to the degree of fibrosis, it seems to be overexpressed, enhancing its receptors and their activity in fibrous tissue[39]. Mainly in reaction to diverse stimuli, including viruses, chemicals, or mechanical injury, KCs manufacture and release PDGF[40]. When PDGF is released, it attaches to a particular receptor on the HSCs’ membrane, activating transcription factors and matching signal molecules involved in the process[41]. This causes the activation of its target genes, which are downstream of the receptor, as well as the activation of HSCs. It has been shown that PDGF (P38-MAPK) increases the activity of C-Jun N-terminal kinase, extracellular signal-regulated kinase (ERK) 1/2, MMP, TIMP, protein kinase B/AKT pathways, and P38 mitogen-activated protein kinase[39].
Transforming growth factor-beta is the strongest known fibrogenic inducer during liver cirrhosis[42]. It is released by all types of hepatic cells in response to unpleasant stimuli and is essential for developing and spreading cirrhosis and liver fibrosis. In fibrotic diseases, TGF-β is abundantly expressed and reaches its peak in cirrhosis[43]. TGF-β pro-fibrogenic impact is carried out by boosting the production of HSCs and ECM while inhibiting MMPs, which results in an excessive buildup of collagen fibers and aids in the progression of liver fibrosis[44]. Additionally, it has been demonstrated that TGF-β causes hepatocyte death and inhibits DNA synthesis[38].
TNF-α is a pro-inflammatory cytokine generated during inflammation and oversees various cell signaling processes. HSCs, KCs, monocytes, and macrophages secrete it[45]. According to a study showing that TNF-α is a mediator of hepatotoxicity and inflammation in many liver diseases, hepatocellular injury followed by inflammation and activation of the innate immune system leads to early-stage liver fibrosis, which in turn causes HSC activation and ECM deposition[46]. In addition, TNF-α contributes to ECM deposition by enhancing the expression of TIMP-1 in HSCs[47]. TNF-α has complex and sometimes conflicting effects on HSCs and fibrosis. TNF-α, on the other hand, has also been demonstrated to have an anti-fibrogenic impact in rat’s HSCs by lowering GSH and decreasing pro-collagen 1 expression. TNF’s function in fibrogenesis is debatable, and it is unknown exactly how TNF receptors contribute to the activation of HSCs. Researchers demonstrate that loss of both TNF receptors decreased pro-collagen 1 expression, slowed HSC proliferation, and impaired PDGF-induced pro-mitogenic signaling in HSC from wild-type, TNF-receptor-1 (TNFR1) knockout, TNFR2 knockout, or TNFR1/R2 double knockout (TNFR-DKO) mice. In response to PDGF, TNFR-DKO HSC showed decreased AKT phosphorylation and in vitro proliferation. However, these effects were not replicated in TNFR2 knockout HSC. Additionally, in primary mouse HSC, TNF binding to TNFR1 was necessary for MMP-9 expression. Neutralizing antibodies against TNFR1 and TNFR2 confirmed these findings in the human HSC cell line LX2. Additionally, compared to wild-type or TNFR2 knockout mice, TNFR-DKO and TNFR1 knockout animals showed less in vivo liver damage and fibrogenesis after bile duct ligation (BDL)[48].
Oxidative stress is frequently described as a general imbalance between oxidizing and reducing substances in the cell. The signaling transduction pathways are governed by these redox states. Numerous human disorders, particularly chronic liver diseases, have been linked to the development of ROS[49]. The production of ROS is crucial in causing liver injury and kicking off hepatic fibrogenesis. Oxidative stress alters lipids, proteins, and DNA, causing hepatocytes to necrotize and apoptosis and escalating the inflammatory response[50].
Additionally, ROS directly activates HSCs and encourages the synthesis of profibrogenic mediators from KCs and circulating inflammatory cells, which leads to the beginning of fibrosis[51]. Regardless of their underlying causes, almost all liver illnesses have been found to exhibit oxidative stress[52]. Prooxidants are ROS that can harm liver cells and whose levels may be raised by some medications, infections, environmental exposures, tissue damage, and other factors. Oxidative stress can be caused by increased prooxidant production, a reduction in antioxidant levels, or a shortage of antioxidants. Signaling, regulation, and redox balance of the liver system are biased by molecular redox switches, oxygen detection by the thiol redox proteome, NAD/NADP, and phosphorylation/dephosphorylation systems. ROS rapidly interact with all biological macromolecules due to their reactivity. The phosphodiester bonds that keep the bases in RNA and DNA together are cleaved by ROS, causing RNA and DNA to lose their chain structure. In a process known as lipid peroxidation, polyunsaturated fatty acids are another important target for oxidation by ROS. This process disturbs the normal structure of the membrane and results in necrosis. Additionally, since cysteine is necessary for the function of enzymes, ROS, particularly the hydroxyl radical, oxidizes cysteine residues in proteins to form disulfides, sulfoxides, or sulfonic acids. Additionally, oxidative stress promotes fibrogenesis by raising toxic cytokines such as TNF-α, IL-6, and TGF-β[53].
ROS generated by the NADP/NADPH oxidase system can control the cellular redox environment in hepatocytes and KCs. NADPH oxidase activation is the main ROS source in myofibroblasts and the stimulation of profibrogenic pathways[54]. It is regarded as the main producer of superoxide anion and hydrogen peroxide, the two most damaging ROS contributing to liver damage from oxidative stress[55]. NADPH oxidase inhibition is emerging as a target for antifibrotic treatment since NADPH oxidase activation may constitute a central mechanism in fibrosis[56].
The activities of various antioxidant enzymes, whose expression is controlled by several redox-sensitive transcription factors like nuclear factor kappa-light-chain-enhancer of activated B cells [nuclear factor-kappaB (NF-κB)] and nuclear factor erythroid 2-related factor 2 (Nrf2), may have an impact on the generation of ROS[57]. Quiescent HSCs lack NF-κB in contrast to activated HSCs, which suggests that a redox-sensitive activation of NF-B might govern the expression of NF-B-targeted genes and provide a suitable cellular redox threshold for quiescent HSCs to enter the proliferative cycle[58]. In support of this theory, it has been shown that blocking NF-κB activity shields rats from the onset of hepatic fibrosis. The suppression of Nrf-2 may also change the expression of antioxidant enzymes, disrupting the cellular redox environment and impacting HSC proliferation, cell death, and collagen formation, all of which contribute to liver fibrosis[49].
Additionally, ROS-sensitive cytokines help activate HSCs during inflammation by receiving paracrine cues from immune cells. Hepatic fibrosis progresses more quickly due to the activated HSCs’ increased receptivity to PDGF and TGF-β[40]. TGF-β boosts the generation of ROS while lowering the level of reduced GSH. The production of the collagen I protein is increased when lipid peroxidation is increased, and anti-oxidant defenses like GSH, catalase, or superoxide dismutase are decreased[59].
MiRNAs are a group of tiny, non-coding endogenous RNA molecules with a high degree of chemical stability (22 nucleotides). MiRNAs have been thoroughly investigated since their discovery in 1993[60] because of their function in RNA-induced posttranscriptional gene silencing. One of the most prevalent adult hepatic miRNAs, miR-122, controls several important gene networks, including lipid metabolism, cell differentiation, and the hepatic circadian rhythm[61]. Recently, miR-223 is thought to interfere with the development and homeostasis of the immune system as well as it has an important role in inflammatory disorders and other liver disorders[62]. Moreover, MiR-223 also controls the nucleotide-binding oligomerization domain-like receptor (NLR) inflammasome by targeting the NLR protein 3 (NLRP3) 3′-untranslated regions[63]. Notably, different cell types require NLRP3 inflammasome to start the inflammatory reaction and the production of ILs. Accordingly, overexpression of miR-223 reduces IL-1 production from the inflammasome and prevents NLRP3 protein formation. Additionally, miR-223 may prevent macrophage hyperactivation[64].
Recent evidence showed that numerous liver disorders, including viral hepatitis, alcohol-induced liver damage, drug-induced liver injury, NAFLD, cirrhosis, and HCC, have dysregulated the expression of the miR-223 gene. Markedly, Weseslindtner et al[65] revealed that, the elevation of miR-106a, miR-122, and miR-197 levels in patients with severe acute viral hepatitis. Interestingly, Fukushima et al[66] made a thorough comparative microarray study and looked at how different miRNAs changed in rats after receiving APAP and CCL4 and discovered that eight miRNAs were downregulated while six miRNAs (miR-153, miR-337, miR-363, miR-302b AS, miR-409-5p, and miR-542-3p) were upregulated in both hepatotoxicity models.
Since miR-122 is very liver-specific and makes up around three-quarters of the entire miRNAs that the liver expresses, it has been extensively studied concerning liver damage[67-69]. It is highly expressed in hepatocytes because of liver-specific transcriptional regulation under the effect of hepatic transcription factors[70]. Further, it seems to be elevated in the majority of liver disorders, including HCV and HBV, in addition to ALD, drug-induced liver damage, NAFLD, and HCC[71-74]. Along with this, loss of miR-122 is seen during hepatocellular carcinogenesis due to hepatic cell dedifferentiation[75,76]. In acute and chronic liver disorders, miR-122 serum/plasma levels are correlated to hepatic necro-inflammation, elevated aminotransferase levels, liver injury, and cell death[77].
Not only miR-122 but miR-192 as well was elevated in the mice sera after APAP administration compared to controls in a dose-dependent and exposure-dependent manner. In this context, the levels of those miRNAs were enhanced sooner than the levels of serum transferases[78], highlighting that they can be used diagnostically superior to the conventional ALF indicators[79]. Similarly, the serum levels of miR-122 were also elevated in the I/R mice models, and they were connected to both aspartate aminotransferase (AST)/alanine aminotransferase (ALT) levels and the hepatic cell death identified by terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling. As in vitro studies showed that miR-122 levels increase in the supernatant after hepatocyte injury. The presence data imply that miR-122 may replace hepatocyte mortality in liver damage[67]. Additionally, the elevation of miR-122 and miR-192 in the sera of patients with APAP-induced ALF could be confirmed, and these findings concur with results from high-throughput sequencing of patients who had taken too much APAP[69]. Krauskopf et al[69] showed that, compared to controls, the serum levels of 36 types of miRNAs were higher in these individuals. Additionally, following APAP overdose, miR-122, miR-192, miR-194, miR-210, and miR-483 were shown to be reinforced in the liver.
In ALF, a considerable downregulation of miR-122 is seen in the injured liver in both acute and chronic liver injury, and it showed an inverse correlation between hepatic damage and ALT levels, suggesting that it may play a role in human ALF. When paraquat was administered to humans, miR-122 was noticeably upregulated, whereas miR-483 and miR-711 were concurrently downregulated. This is consistent with what was shown in the rats given an APAP overdose[78,80]. Zhang et al[81] discovered that blood levels of miR-122 and miR-192 were increased after acute hepatic poisoning with APAP in mice before transaminases, particularly ALT, were raised. However, it was shown that the miRNA levels in liver tissue were lower. Since these miRNAs may be detected before the liver experiences apparent cell death, they may serve as a more accurate indicator of liver failure than liver enzymes[82]. Recent studies showed that miR-15b, miR-16, miR-125a, miR-146a, and miR-155 were considerably up-regulated during ALF in mice, while miR-1187 showed a significant down-regulation[83].
Hepatitis B e antigen (HBeAg) positive patients had much greater blood levels of miRs than those with HBeAg negative, especially miR-122 and miR-194, which showed the greatest differential expression[84]. Additionally, it has been shown that the expression of miR-122, miR-638, miR-572, miR-575, miR-638, and miR-744 was dysregulated in chronic HBV patients; these miRs were significantly more abundant in HBV than AST or ALT. MiR-122, miR-572, miR-575, and miR-638 were more abundant than miR-744[85]. In human hepatoma cells, HepG2, miR-155 has been shown to contribute to antiviral immunity against HBV infection[86]. An initial therapeutic response to IFN (independent relationship with early virologic response) may be predicted in HBV patients using a miR profile of 11 miRs for example, hsa-let-7a, hsa-miR-30a, hsa-miR-106b, hsa-miR-198, hsa-miR-1224-5p, and hsa-miR-1290. It has been demonstrated that certain miRs might play a function in the HBV life cycle[87]. According to studies, specific miRs have been shown to affect HCV infection or be affected by the virus. There is still much to learn about how miR-122 interacts with the HCV genome[88].
However, miR-122 expression is unaffected by viral infection or replication. Recently, Randall et al[89] looked at miR-21 and miR-122 expression in the liver biopsy samples from patients infected with HCV and controls. They established that miR-122 levels were inversely linked to the fibrotic stage, ALT, and AST but that miR-21 levels were positively linked. It was suggested that rather than levels of expression, fibrosis might be brought on by dysregulation of miR-21 and miR-122. MiRs 24, 149, 638, and 1182, among others, share in HCV entrance, replication, and spread[90]. The tumor suppressor “deleted in liver cell-1” protein was shown to be highly dependent on miR-141 activation, miR-141-targeted downregulation, and depletion for sustained HCV propagation. According to research on the association between HCV and the levels of miR-29 in both HSC and hepatocytes, HSC stimulation results in miR-29 down-regulation[91]. The overexpression of miR-29 in infected cells reduced HCV replication by 70% and inhibited the growth of HSCs and collagen synthesis. When comparing the livers of HCV with non-SVR, miR-29a, b, and c levels were higher[92], indicating a potential function for these biomarkers in monitoring the effectiveness of anti-HCV therapy.
In alcoholic steatohepatitis, miRs are crucial immune response regulators and activators of the innate immune system[93]. Alcohol-induced gut leakiness, which permits endotoxin to enter the blood and begin liver damage, has been shown to play a critical role in ALD and to increase miR-122 expression. It is shown that inducing miR-155 and -132 causes KCs to release higher TNF-α in response to lipopolysaccharide (LPS)[94]. Hepatic miRs 182, 183, 705, 1224, and 199a-3p are modulated by endotoxemia and alcohol use directly[95]. Alcohol specifically targets and upregulates the miR-155 gene in macrophages, which controls the production of TNF-α[96]. Prolonged alcohol exposure also stimulates the miR-155 gene in KCs and RAW264.7 macrophages. As a result, miR-155 upregulation might be engaged in the oxidative stress and LPS pathways, thus promoting the development of ALD[94].
There is mounting evidence that miRs, namely via controlling gene expression in HSCs, are important regulators of hepatic fibrogenesis. The advancement of liver fibrosis has been linked to the miR-199 and miR-200 family’s expression. Patients with fibrotic livers had higher levels of the miR-199 and miR-200 families, and upregulation of these miRs led to considerably higher levels of the genes related to fibrosis in a cell line of HSC. In a fibrosis model of BDL in rats, miR-150 and miR-194 levels were significantly lower than in animals with a sham procedure. Furthermore, in a human stellate cell line called LX2, it has been shown that overexpressing miR-150 or miR-194 through the reduction of c-myb and rac1 expression can reverse the activated stellate cells (i.e., expression of collagen and alpha-smooth muscle actin genes). Therefore, miR-150 and miR-194 may represent promising therapeutic targets for fibrosis treatment[97,98].
Autophagy is a self-eating catabolic mechanism in eukaryotic cells that ends in the lysosome[99,100]. In addition to its anti-aging function, autophagy plays a significant role in immune response and organ homeostasis[101,102]. Numerous pathological disorders, such as obesity and type 2 diabetes, inflammatory and viral diseases, neurodegenerative diseases, and cancer, exhibit autophagy dysregulation[103,104]. There are distinct phases of autophagy; induction, phagophore development, autophagosome creation, autolysosome formation, and destruction[105,106]. Atg molecules participate in several complexes crucial for triggering autophagy and creating autophagosomes[107]. The unc-51-like kinase 1 complex (Atg1 in yeast) is activated first, then beclin 1 (Atg6 in yeast), and followed by a series of Atg proteins that result in the production of autophagosomes, with LC3 (Atg8 in yeast), being one of them[108]. Further processing of LC3 results in the formation of LC3-I and LC3-II[109]. As soon as the autophagosome gets created, a blockade of autophagic flux at later stages will suppress the autophagosome’s ability to be cleared, ultimately leading to autophagy-dependent cell death[110]. To date, macroautophagy, microautophagy, and chaperone-mediated autophagy (CMA) are the three main types of autophagy that have been characterized[111,112].
Lately, researchers have investigated the relationship between autophagy and the immune system[113,114]. There have been documented non-canonical macroautophagic processes that create lysosome-fusing autophagosomes[115]. Only a portion of the Atgs equipment is utilized. Due to its significance in immunological modulation, LC3-associated phagocytosis (LAP) has received the most attention[116,117]. LAP draws LC3-II to the phagosomal membrane via innate immune receptors, such as toll-like receptors, where macrophages consume it. The crucial part that CMA plays is antigen presentation and aging, which has also garnered attention[118]. Innate immunity’s ability to hinder macrophage autophagy is also associated with autophagy. Innate immunity and autophagy interact because IFN-α stimulates autophagy in macrophages[119].
In some circumstances, autophagy can either serve as a defense mechanism or contribute to cellular death[120,121]. The main way autophagy contributes to cellular death is through its influence on apoptosis. Apoptosis and autophagy are linked, and these two cellular destructing processes influence one another[122,123]. This is crucial in the demise of liver cells[124]. Autophagy generally prevents caspase-dependent apoptosis from being induced, whereas apoptosis-related caspase activation halts the autophagic process.
Along with these results, Ni et al[125] documented that necrosis and necroptosis are caused by caspase-independent cell death, which is closely linked to autophagy. Cells harmed by the tumor suppressor gene p53 are removed by the induction of apoptosis[126]. In addition to being engaged in autophagy, the mechanistic target of rapamycin (mTOR)/AKT pathway also inhibits apoptosis. For the destiny of damaged cells, p53 and AKT/mTOR must coexist in equilibrium[127]. Numerous proteins linked to autophagy, including Atgs and BECN1, also played a role in ferroptosis. Additionally, erastin, an activator of ferroptosis, caused the formation of autophagosomes, and activation of autophagy resulted in ferroptotic cell death, maybe because of the ferritin being broken down by ferritinophagy[128].
Autophagy and the liver’s inflammatory response are tightly related. The same inhibitory mechanisms govern autophagy and inflammasome but are regulated by various input pathways. Procaspase-1 activation results from the activation of the NLRP3 inflammasome, which is often triggered by pathogen- or danger-associated molecular patterns[129], which will further stimulate the synthesis of IL-1 and IL-18 that causes pyroptotic cell death. Moreover, the activation of autophagy by caspase-1 prevents these occurrences. Additionally, autophagy decreases inflammasome activation by destroying inflammasomes in autophagosomes and removing damaged cytoplasmic organelles that, in the absence of autophagy, would otherwise create DAMPS and increase inflammasome activation[130]. On the other hand, when autophagy is diminished, the pro-inflammatory IL-1 is produced more often due to the negative association between inflammasomes and autophagy[131,132]. Although the connection between NLRP3 and autophagy is not entirely understood, recent research has indicated that NF-κB activation can similarly modify NLRP3 and autophagy[133].
Given the preceding, it is not surprising that many autophagy reviews emphasize the contrasting impacts that autophagy may have on the same biological process by using the phrase “double-edged sword”[134]. Cancer[135] and viral infections[101] are prominent fundamental paradigms. The fact that autophagy exhibits Jekyll-like and Hyde-like characteristics depending on the cells involved is another trait exclusive to the liver. Hepatocytes in NAFLD and ALD exhibit protective macroautophagy and CMA (in NAFLD). It eliminates damaged mitochondria, lessens oxidative stress, and promotes regeneration. In macrophages, macroautophagy reduces liver fibrosis and inflammation while promoting fibrosis-activated stellate cells. It is preventative in the early stages of HCC but might be damaging in the later stages[136]. Both the non-parenchymal sinusoidal cells of the liver and the hepatocytes depend on autophagy for proper liver function[137], and autophagy abnormalities are linked to most liver illnesses’ pathogenesis[138]. Autophagy disorders are linked to both common conditions like alcoholic and NAFLD or viral hepatitis and uncommon conditions like Wilson disease and alpha 1 antitrypsin deficiency[139,140]. Due to the 6-12 mo half-life of hepatocytes, impaired autophagy also contributes to the accumulation of toxic hepatocyte byproducts. A large number of xenobiotics must also be processed by the liver, and autophagy is a cytoprotective mechanism[141].
Cirrhosis of the liver, the end stage of liver fibrosis after chronic liver damage, used to be cured by nearly liver transplantation only. That is why researchers used to focus on preventing liver cirrhosis by eradicating the cause and reversion of fibrosis. However, if liver cirrhosis develops, treatment is restricted to preventing the progression of the complications and avoiding the need for liver transplantation[142-144]. Besides removing the cause, various categories of treatments have proven to be beneficial in preventing fibrosis progression or regression, such as antioxidants, and antifibrotic agents, including phyto drugs[144-147]. Via understanding the process of fibrogenesis, various mechanisms implicated in this process would be potential for the reversion of fibrosis and cirrhosis. Here in, we discuss several conventional and novel therapeutic interventions that showcased, by recent data, the ability to modulate liver fibrosis and cirrhosis. The recent therapeutic interventions are summarized in Table 2.
| Therapeutic intervention | Drugs | Main findings | Ref. |
| Antioxidants | Silymarin | Possesses free radical scavenging activity and inhibits lipid peroxidation thus improving chronic liver diseases | [149] |
| Selenium | Decrease DNA damage, hepatocyte necrosis, oxidative stress biomarkers, and liver toxicity | ||
| Vitamin E | Reduces inflammation and protects from hepatocellular damage | [155,157,160] | |
| N acetylcysteine | Increasing GSH peroxidase and decreasing oxidative stress in liver fibrosis | ||
| MitoQ | Reduces lipid peroxidation and cultured hepatic stellate cell activation | [162] | |
| Antifibrotic agents | Pirfenidone | Pirfenidone is effective at diminishing liver fibrosis as it suppresses TGF-β1 and NF-κB and decreases inflammatory cell infiltration and excess matrix deposition | [166-168] |
| Statins, and anti- NADPH oxidases | PPAR-α modulators might decrease inflammation and fibrosis in cases of primary sclerosing cholangitis | ||
| Immunosuppressants | Corticosteroids, and azathioprine | The first line of treatment for autoimmune hepatitis | [169] |
| Anti-HSC therapy | Imatinib and sorafenib | Respectively act as PDGF and angiogenesis inhibitors thus they modulate fibrogenesis and fibrosis in autoimmune hepatitis | [173] |
| Paclitaxel, ferulic acid and methyl ferulic acid | Can inhibit hepatic stellate cell activation through TGF-β/Smad pathway modulation | [175-177] | |
| Curcumin | Can interrupt the PDGF-β/ERK pathway and inhibit hepatic stellate cell angiogenesis through activation of PPAR-γ. Curcumin can also activate autophagy and thus inhibit the TGF-β/Smad pathway thus reducing epithelial-mesenchymal transition | [178-180] | |
| Gene therapy | HGF | Decreases the expression of TGF-β1, suppresses hepatocyte apoptosis, and improves fibrosis | [181] |
| Matrix metalloproteinase-1 | Enhances the proliferation of hepatocytes and diminishes fibrosis | [183] | |
| siRNA | By silencing CTGF, TGF-β, NF-κB target gene A, galectin-3, and αvβ3 integrin, siRNA effectively stops fibrogenesis by preventing HSCs activation and/or promoting their apoptosis | [184] | |
| Cell therapy | MSCs | Inhibit hepatocyte degeneration, promote liver regeneration, and suppress fibrosis through differentiation into hepatocytes and production of various growth factors | [187] |
| BMSCs | Decrease serum markers of liver injury and mRNA expression of TNF-α, IFN-γ, and FasL, and increase IL-10 mRNA expression in acute liver failure | [189] | |
| Matrix metalloproteinase 2, tissue inhibitor of metalloproteinase 1, and growth arrest-specific 6 | Promote hepatocytes regeneration, neovascularization, and extracellular matrix remodeling all contributing to liver regeneration | [191] | |
| Gut liver axis | Baicalin | Modulates FXR and G-protein-coupled bile acid receptor TGR5 thus modulating the levels of TNF-α, NF-kβ, and TGF-β. It also inhibits inflammation, autophagy, and necrosis of parenchymal liver cells | [195-198] |
| Probiotics | Modulate gut dysbiosis and bile acid dysregulation thus aiding in the treatment of NAFLD. Probiotics also modulate inflammation and fibrosis in NASH | [199-201] | |
| Nanoparticle drug delivery | Gold | Enhances the antifibrotic activity of silymarin through increasing the expression of protective microRNAs and suppression of inflammatory mediators in the TGF-β1/smad pathway | [204] |
| Phosphatidylserine-decorated nanoparticles | Enhances curcumin efficacy in fibrosis reduction | [205] | |
| Liposome nanoparticles | Can be specifically delivered to integrins of activated hepatic stellate cells, in addition to facilitating gene therapy using siRNAs and mRNAs to modulate gene expression of hepatocytes | [208] | |
| Autophagy inhibition | Becn1 knockdown | Autophagy suppression and inhibition of T lymphocyte infiltration, HSCs proliferation, as well as production of TNF-α, IFN-γ, and TGF-β1 | [209] |
| Carvedilol | Increased p62 protein levels and inhibited autophagic flux by increasing lysosomal pH | [210] | |
| Doxazosin | Inhibited HSC proliferation and migration, blocked autophagic flux and induced HSCs apoptosis | [211] | |
| Resolvin D1 | Modulated AKT/mTOR signaling pathway resulting in the inhibition of autophagy and suppression of hepatic stellate cell activation | [212,213] |
Oxidative stress is well known to play a detrimental role in developing liver cirrhosis. When ROS production exceeds antioxidants level, cellular signaling pathways alterations eventually result in liver damage[148]. For this reason, antioxidants received much attention and extensive study to prevent and treat various liver disorders. Silymarin is an herbal extract that consists mainly of silybin, which is responsible for the activity of silymarin. Free radical scavenging activity and inhibition of lipid peroxidation have been exhibited as reasons for the antioxidant activity of silybin[149]. Selenium is an essential element for the GSH antioxidant system in our bodies that has been extensively studied for its antioxidant activity in various cases of liver damage[150]. Selenium showed the ability to decrease DNA damage and hepatocyte necrosis against cyclophosphamide-induced oxidative stress[151]. In cadmium-induced acute liver injury, selenium nanoparticles decreased liver toxicity by boosting the Nrf2 pathway[152]. In chronic liver injury, selenium is reported to mitigate lipid peroxidation and decrease other oxidative stress biomarkers, especially when combined with the natural antioxidant gum arabic[153]. A study investigating the effect of curcumin, selenium, and silymarin showed that the combination of selenium, curcumin, and silymarin ameliorates the oxidant/antioxidant status in lipopolysaccharide and diclofenac-induced liver damage[154]. Vitamin E is a fat-soluble vitamin and one of the most potent antioxidants. This action is attributed to the ability of the hydroxyl group to scavenge free radicals and restoration of GSH levels and hence the improvement of oxidant/antioxidant status. Accordingly, in addition to other mechanisms, vitamin E effectively reduces inflammation[155] but not fibrosis[156].
Nevertheless, a recent in vivo study by Aljuhr et al[157] showed that using vitamins E and C loaded on selenium nanoparticles effectively reduces the induced hepatocellular damage, making it a potent combination for preventing and treating HCC. Acute hepatotoxicity induced by APAP overdose is typically countered by N acetyl cysteine via its antioxidant activity and increasing the level of GSH in the liver[158]. In cases of APAP-induced acute liver injury, N acetylcysteine is the antidote for hepatotoxicity as it can preserve GSH stores and counteract the toxic metabolite NAPQI[159]. In addition, N acetyl cysteine exerted favorable effects at increasing GSH peroxidase and decreasing oxidative stress in liver fibrosis induced by carbon tetra chloride[160]. Mitoquinone (MitoQ), mitochondrial-targeted coenzyme Q, is a recent advance in antioxidant therapy that delivers coenzyme Q directly to the mitochondria[161]. In carbon tetrachloride-induced liver fibrosis, MitoQ showed a reduction in lipid peroxidation marker, 4-hydroxynonenal, in vivo and inhibition of cultured HSC activation[162]. Accordingly, MitoQ seems promising in mitigating liver fibrosis, but further studies are needed to confirm its efficacy.
Both acute and chronic liver disorders involve a series of cytokine and chemokine production and inflammatory cell infiltration that promote fibrogenesis[163,164]. This emphasizes the importance of using anti-fibrotic and anti-inflammatory drugs to modulate fibrogenesis and reduce the progression of liver fibrosis. Pirfenidone is a pyridone derivative with antifibrotic and anti-inflammatory properties and is mainly used for pulmonary fibrosis[165]. These actions are attributed to the ability of pirfenidone to suppress TGF-β and NF-κB activation and thus decrease inflammatory cell infiltration and excess matrix deposition. Along with the antioxidant activity of pirfenidone, it effectively diminishes liver fibrosis[166], as shown in a two-year in vivo study on CHC virus-infected patients[167]. Statins and anti-NADPH oxidases as anti-fibrotic classes, peroxisome proliferator-activated receptor alpha modulators, and timolumab as immunomodulators have been recently investigated and declared promising for decreasing inflammation and fibrosis in cases of primary sclerosing cholangitis[168].
Autoimmune hepatitis is a chronic inflammatory liver disease that occurs when helper T cells and cytotoxic T cells attack the liver causing inflammation that may progress to liver cirrhosis. Autoimmune hepatitis can be acute, fulminant, or chronic and, like other autoimmune disorders, require immunosuppressive therapy to suppress the disease progression. The treatment of autoimmune hepatitis involves using corticosteroids as antifibrotic agents and azathioprine, and when this line of management is insufficient, mycophenolate mofetil and calcineurin inhibitors are used[169,170]. In contrast, these drugs require more investigation for their use in other autoimmune liver disorders, such as primary sclerosing cholangitis and primary biliary cirrhosis[171].
One of the most important mitogens in profibrogenic HSC activation following liver damage is PDGF[172]. A recent study using PDGF and angiogenesis inhibitors as imatinib and sorafenib, respectively, concluded that they were able to modulate fibrogenesis and fibrosis in induced autoimmune hepatitis models[173]. Silymarin possesses antioxidant activity and antifibrotic properties through inhibition of KCs activation, decreasing extracellular matrix deposition, and inhibiting the production of IL-1 and IL-8 on HSCs[174]. As TGF-β is a crucial cytokine for HSC fibrogenesis and hence liver fibrosis progression[172], different studies have been conducted to study the effect of various substances to obstruct TGF-β/Smad signals. In vitro studies on paclitaxel, ferulic acid, and methyl ferulic acid were encouraging for inhibition of HSC activation via TGF-β/Smad pathway modulation[175-177]. Curcumin is a natural antioxidant, anti-inflammatory, and antifibrotic agent that can modulate different apoptotic pathways during tissue injury. Recent studies showed that curcumin could interrupt the PDGF-β/ERK signaling pathway and inhibit HSC angiogenesis by activating PPAR-γ[178,179]. Furthermore, curcumin can activate autophagy and thus inhibit the TGF-β/Smad pathway, which reduces epithelial-mesenchymal transition[180]. Accordingly, curcumin is considered a good candidate for treating liver fibrosis.
Acute liver injury is usually reversible; however, chronic liver damage is a progressive condition that usually progresses from inflammation and fibrosis to cirrhosis. That is why extensive investigations on gene therapy have been conducted with various genes and delivering vectors to modulate liver fibrosis and cirrhosis. Hepatocyte growth factor (HGF) is an essential antiapoptotic and hepatoprotective factor for hepatocytes and an antifibrogenic agent in liver fibrosis models. HGF gene therapy has been studied for liver cirrhosis in rats and was shown to decrease the expression of TGF-β, suppress hepatocyte apoptosis, and improve fibrosis in dimethyl nitrosamine-induced cirrhosis[181]. Due to the ability of HGF to suppress TGF-β, it exhibits immunomodulatory action that is promising in cases of autoimmune disorders, but further investigations are still required[182]. As HSCs generate abundant amounts of extracellular matrix during fibrogenesis, matrix metalloproteinase-1 delivered by adenovirus to fibrotic livers enhances the proliferation of hepatocytes and diminishes fibrosis[183]. Another mechanism involves the use of small interfering RNA (siRNA) to silence the genes that are essential for the process of fibrosis, such as connective tissue growth factor, TGF-β, NF-κB target gene A, galectin-3, and αvβ3 integrin. Silencing these genes stops fibrogenesis effectively by preventing HSCs activation and promoting their apoptosis[184].
Stem cells are a category of cells that can replicate and differentiate into numerous types of specialized cells in the body[185]. During the last two decades, stem cell-based therapy has been extensively investigated and appears promising for liver regeneration. Thus, it is a considerable alternative for liver transplantation and overcoming its demerits like the shortage of liver donors, high cost, and surgical complications. Various types of stem cells have been studied in acute and chronic liver disorders, including embryonic stem cells, induced pluripotent, and adult stem cells composed of the liver, mesenchymal, and hematopoietic stem cells[186]. Mesenchymal stem cells (MSCs) are a suitable alternative for liver transplantation because they inhibit hepatocyte degeneration, promote liver regeneration, suppress fibrosis via differentiation into hepatocytes, and produce various growth factors[187].
Moreover, combining MSCs with induced bone marrow-derived macrophages showed stronger antifibrotic activity and hence better improvement of the cirrhotic liver than monotherapy[188]. An in vivo study investigating cell therapy in mice used four types of cells; mature hepatocytes, fetal liver cells, bone marrow-derived mesenchymal stromal cells (BMSCs), and induced hepatic stem cells for concanavalin A-induced fulminant hepatitis causing ALF and fumarylacetoacetate hydrolase-deficient induced chronic liver failure. Remission of concanavalin A-induced ALF was only noticed with BMSCs as they decreased serum markers of liver injury and mRNA expression of some inflammatory cytokines, including TNF-α, IFN-γ, and FasL, and increased IL-10 mRNA expression. In the chronic liver failure model, mature hepatocytes in the adult liver were the most effective for liver regeneration compared to other cell types. However, these hepatocytes are not common in clinical applications due to their limited sources[189]. In acute liver injury induced by carbon tetra chloride, using hepatocyte-like cells derived from embryonic stem cells showed the potential for attenuation of liver injury and the remission of induced liver fibrosis[190]. Furthermore, rather than cell transplantation, using trophic factors such as matrix metalloproteinase 2, tissue inhibitor of metalloproteinase 1, and growth arrest-specific 6, released from embryonic-derived hepatocyte-like cells, promoted hepatocytes regeneration, neovascularization, and extracellular matrix remodeling, all of which contribute to liver regeneration[191]. Despite the numerous advantages of stem cell therapy, safety concerns such as ethical approval of embryonic stem cell use, lack of knowledge of appropriate transmission methods, enhancement of tumor growth, and incomplete prediction of tissue response are limiting their use nowadays[192].
The relationship between the gut and the liver involves the delivery of intestinal contents to the liver through the portal vein and the transport of bile acids and immunoglobulins from the liver back to the intestines. Any disruption of the homeostasis of this axis through altering gut microbiota (gut dysbiosis), bile acid composition, or intestinal barrier damage will result in the exposure of the liver to these microbes and their metabolites which is critical in the pathogenesis of the ALD, NAFLD and even liver cirrhosis[193,194]. That is why various experiments and clinical trials targeting the gut-liver axis are being studied to treat liver disorders, including NASH, NAFLD, and chronic hepatitis B and C[195]. The farnesoid X receptor (FXR) is a nuclear receptor highly expressed in the gut-liver axis and regulates bile acid production, detoxification, maintenance of triglyceride homeostasis, and enhancement of the function of the intestinal epithelial barrier[194,196,197]. Baicalin is a natural flavonoid studied on various liver disorders and exhibited favorable effects such as inhibition of inflammation and autophagy and necrosis of parenchymal liver cells, thus decreasing liver injury. One of the pathways involved in baicalin effects is FXR and G-protein-coupled bile acid receptor, as they can modulate TNF-α, NF-kβ, and TGF-β levels[195,198]. As gut dysbiosis and bile acid dysregulation are directly related to NAFLD’s pathogenesis, using various probiotics, prebiotics, and synbiotics has been proven to be promising for treating NAFLD[199,200]. In addition to the role of probiotics in NAFLD and their ability to modulate inflammation and fibrosis in NASH, probiotics are an attractive target for gut-liver-related disorders as they are also cost-efficient, with mild adverse effects and nearly no long-term adverse reactions[201].
Recently, nanomedicine gained much attention as an innovative way for effective drug delivery in various resistant types of diseases. Numerous nanoparticle types are used in liver fibrosis treatment: Inorganic oxides and metals[202] or organic micelles and liposomes[203]. Gold, an inert inorganic widely used material, is formulated in nanoparticle form to deliver silymarin to fibrotic livers induced by carbon tetrachloride. This process enhanced the antifibrotic activity of silymarin, attributed to increased expression of protective miRNAs and suppression of inflammatory mediators in the TGF-β/Smad pathway[204]. As we previously mentioned, the anti-fibrotic action of curcumin, enhancing drug delivery and bioavailability of curcumin using phosphatidylserine-decorated nanoparticles, further enhances curcumin efficacy in fibrosis reduction[205].
Interestingly, nanoparticles can also target different liver cells involved in liver fibrosis. As the expression of c-x-c chemokine receptor 4 (CXCR4) and VEGF is associated with HSCs activation and hence liver fibrosis progression, combining CXCR4 antagonist in nanoparticles with siRNA against VEGF provided significant inhibition of the process of angiogenesis making it auspicious treatment for liver fibrosis[206]. Considering liposome nanoparticles, the use of liposomes to be specifically delivered to integrins of activated HSCs rather than any other type of liver cells has been conducted, making it available to deliver therapeutic drugs to special sites overcoming their complications[207]. A novel advantage of liposome nanoparticles is that they facilitate gene therapy using siRNAs and mRNAs to modulate gene expression of hepatocytes instead of using viruses as carriers[208].
As we have mentioned, the BECN1 protein has been involved in autophagy, resulting in ferroptotic cell death. A study on knocking down BECN1 showed inhibition of autophagy and its consequent inflammation in addition to increasing prostaglandin E2 (PGE2) levels. Modulation of the prostaglandin-endoperoxide synthase 2/PGE2 pathway may cause suppression of HSC proliferation and lymphocyte infiltration, all contributing to MSCs’ enhanced antifibrotic activity[209]. That is why inhibition of autophagy is a potential target for liver fibrosis treatment. Carvedilol, a non-selective B-blocker, has been thought to possess antifibrotic activity. Testing this theory in vitro revealed that carvedilol can alleviate liver fibrosis by inhibiting the autophagy of HSCs and enhancing their apoptosis[210]. Doxazosin, an alpha-1 adrenergic receptor agonist, has also been studied in vitro and in vivo and showed similar action to carvedilol on activating apoptosis of HSCs and inhibiting autophagy through the PI3K/Akt/mTOR signaling pathway[211]. Resolvin D1 is a polyunsaturated fatty acid that has been proven effective in various liver disorders, such as acute liver injury and liver fibrosis, due to its antioxidant, anti-inflammatory, and antifibrotic effects. Further investigations on resolvin D1 on CCL4-induced liver fibrosis demonstrated its ability to modulate the AKT/mTOR signaling pathway, resulting in inhibition of autophagy and suppression of HSC activation, which further intensifies resolvin D1 liver protective effect[212,213].
Collectively, acute and chronic liver diseases are worldwide problems with multifactorial pathogenesis. The exact pathological mechanism of several liver disorders is still unclear. However, many suggested mechanisms are involved, including but not limited to oxidative stress, inflammation, autophagy, and miRNA. The role of autophagy and miRNA is still unclear and requires more clarification. Besides, it may be a new way to find new therapy for hepatic disorders. Recent therapeutic strategies like gene the
Provenance and peer review: Invited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Egypt
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): 0
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Lu H, China; Yang YZ, China S-Editor: Wang JJ L-Editor: A P-Editor: Wang JJ
| 1. | Cheemerla S, Balakrishnan M. Global Epidemiology of Chronic Liver Disease. Clin Liver Dis (Hoboken). 2021;17:365-370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 321] [Article Influence: 80.3] [Reference Citation Analysis (0)] |
| 2. | Asrani SK, Devarbhavi H, Eaton J, Kamath PS. Burden of liver diseases in the world. J Hepatol. 2019;70:151-171. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1382] [Cited by in RCA: 2290] [Article Influence: 381.7] [Reference Citation Analysis (0)] |
| 3. | Sowa JP, Gerken G, Canbay A. Acute Liver Failure - It's Just a Matter of Cell Death. Dig Dis. 2016;34:423-428. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 21] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 4. | Sun M, Kisseleva T. Reversibility of liver fibrosis. Clin Res Hepatol Gastroenterol. 2015;39 Suppl 1:S60-S63. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 154] [Cited by in RCA: 195] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 5. | Yan M, Huo Y, Yin S, Hu H. Mechanisms of acetaminophen-induced liver injury and its implications for therapeutic interventions. Redox Biol. 2018;17:274-283. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 468] [Cited by in RCA: 434] [Article Influence: 62.0] [Reference Citation Analysis (0)] |
| 6. | Lancaster EM, Hiatt JR, Zarrinpar A. Acetaminophen hepatotoxicity: an updated review. Arch Toxicol. 2015;89:193-199. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 168] [Cited by in RCA: 204] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 7. | Jaeschke H, McGill MR, Ramachandran A. Oxidant stress, mitochondria, and cell death mechanisms in drug-induced liver injury: lessons learned from acetaminophen hepatotoxicity. Drug Metab Rev. 2012;44:88-106. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 563] [Cited by in RCA: 686] [Article Influence: 52.8] [Reference Citation Analysis (0)] |
| 8. | Chung RT, Stravitz RT, Fontana RJ, Schiodt FV, Mehal WZ, Reddy KR, Lee WM. Pathogenesis of liver injury in acute liver failure. Gastroenterology. 2012;143:e1-e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 80] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 9. | Antoniades CG, Berry PA, Wendon JA, Vergani D. The importance of immune dysfunction in determining outcome in acute liver failure. J Hepatol. 2008;49:845-861. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 237] [Cited by in RCA: 267] [Article Influence: 15.7] [Reference Citation Analysis (0)] |
| 10. | Damm G, Pfeiffer E, Burkhardt B, Vermehren J, Nüssler AK, Weiss TS. Human parenchymal and non-parenchymal liver cell isolation, culture and characterization. Hepatol Int. 2013;7:951-958. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 22] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 11. | Seitz HK, Bataller R, Cortez-Pinto H, Gao B, Gual A, Lackner C, Mathurin P, Mueller S, Szabo G, Tsukamoto H. Alcoholic liver disease. Nat Rev Dis Primers. 2018;4:16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 655] [Cited by in RCA: 809] [Article Influence: 115.6] [Reference Citation Analysis (0)] |
| 12. | Younossi ZM. Non-alcoholic fatty liver disease - A global public health perspective. J Hepatol. 2019;70:531-544. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 943] [Cited by in RCA: 1458] [Article Influence: 243.0] [Reference Citation Analysis (1)] |
| 13. | Schwarz KB, Balistreri W. Viral hepatitis. J Pediatr Gastroenterol Nutr. 2002;35 Suppl 1:S29-S32. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 7] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 14. | Patel D, Teckman JH. Alpha-1-Antitrypsin Deficiency Liver Disease. Clin Liver Dis. 2018;22:643-655. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 46] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 15. | David S, Hamilton JP. Drug-induced Liver Injury. US Gastroenterol Hepatol Rev. 2010;6:73-80. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 264] [Cited by in RCA: 470] [Article Influence: 78.3] [Reference Citation Analysis (0)] |
| 16. | Koyama Y, Brenner DA. Liver inflammation and fibrosis. J Clin Invest. 2017;127:55-64. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 548] [Cited by in RCA: 912] [Article Influence: 114.0] [Reference Citation Analysis (0)] |
| 17. | Calabro SR, Maczurek AE, Morgan AJ, Tu T, Wen VW, Yee C, Mridha A, Lee M, d'Avigdor W, Locarnini SA, McCaughan GW, Warner FJ, McLennan SV, Shackel NA. Hepatocyte produced matrix metalloproteinases are regulated by CD147 in liver fibrogenesis. PLoS One. 2014;9:e90571. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 37] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 18. | Bataller R, Brenner DA. Liver fibrosis. J Clin Invest. 2005;115:209-218. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3381] [Cited by in RCA: 4119] [Article Influence: 206.0] [Reference Citation Analysis (3)] |
| 19. | Jeong WI, Do SH, Yun HS, Song BJ, Kim SJ, Kwak WJ, Yoo SE, Park HY, Jeong KS. Hypoxia potentiates transforming growth factor-beta expression of hepatocyte during the cirrhotic condition in rat liver. Liver Int. 2004;24:658-668. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 43] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 20. | Wiemann SU, Satyanarayana A, Tsahuridu M, Tillmann HL, Zender L, Klempnauer J, Flemming P, Franco S, Blasco MA, Manns MP, Rudolph KL. Hepatocyte telomere shortening and senescence are general markers of human liver cirrhosis. FASEB J. 2002;16:935-942. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 374] [Cited by in RCA: 382] [Article Influence: 16.6] [Reference Citation Analysis (0)] |
| 21. | Senoo H, Kojima N, Sato M. Vitamin A-storing cells (stellate cells). Vitam Horm. 2007;75:131-159. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 106] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 22. | She H, Xiong S, Hazra S, Tsukamoto H. Adipogenic transcriptional regulation of hepatic stellate cells. J Biol Chem. 2005;280:4959-4967. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 213] [Cited by in RCA: 250] [Article Influence: 11.9] [Reference Citation Analysis (1)] |
| 23. | Böttcher K, Pinzani M. Pathophysiology of liver fibrosis and the methodological barriers to the development of anti-fibrogenic agents. Adv Drug Deliv Rev. 2017;121:3-8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 127] [Article Influence: 15.9] [Reference Citation Analysis (0)] |
| 24. | Higashi T, Friedman SL, Hoshida Y. Hepatic stellate cells as key target in liver fibrosis. Adv Drug Deliv Rev. 2017;121:27-42. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 530] [Cited by in RCA: 1059] [Article Influence: 132.4] [Reference Citation Analysis (0)] |
| 25. | Dewidar B, Meyer C, Dooley S, Meindl-Beinker AN. TGF-β in Hepatic Stellate Cell Activation and Liver Fibrogenesis-Updated 2019. Cells. 2019;8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 214] [Cited by in RCA: 539] [Article Influence: 89.8] [Reference Citation Analysis (0)] |
| 26. | Sato-Matsubara M, Matsubara T, Daikoku A, Okina Y, Longato L, Rombouts K, Thuy LTT, Adachi J, Tomonaga T, Ikeda K, Yoshizato K, Pinzani M, Kawada N. Fibroblast growth factor 2 (FGF2) regulates cytoglobin expression and activation of human hepatic stellate cells via JNK signaling. J Biol Chem. 2017;292:18961-18972. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 34] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 27. | Lam PL, Kok SHL, Gambari R, Kok TW, Leung HY, Choi KL, Wong CS, Hau DKP, Wong WY, Lam KH, Bian ZX, Lee KKH, Chui CH. Evaluation of berberine/bovine serum albumin nanoparticles for liver fibrosis therapy. Green Chem. 2014;17. [DOI] [Full Text] |
| 28. | Gracia-Sancho J, Caparrós E, Fernández-Iglesias A, Francés R. Role of liver sinusoidal endothelial cells in liver diseases. Nat Rev Gastroenterol Hepatol. 2021;18:411-431. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 235] [Article Influence: 58.8] [Reference Citation Analysis (0)] |
| 29. | Li P, Zhou J, Li W, Wu H, Hu J, Ding Q, Lü S, Pan J, Zhang C, Li N, Long M. Characterizing liver sinusoidal endothelial cell fenestrae on soft substrates upon AFM imaging and deep learning. Biochim Biophys Acta Gen Subj. 2020;1864:129702. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 12] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 30. | Xie G, Wang X, Wang L, Atkinson RD, Kanel GC, Gaarde WA, Deleve LD. Role of differentiation of liver sinusoidal endothelial cells in progression and regression of hepatic fibrosis in rats. Gastroenterology. 2012;142:918-927.e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 279] [Cited by in RCA: 298] [Article Influence: 22.9] [Reference Citation Analysis (0)] |
| 31. | Marvie P, Lisbonne M, L'helgoualc'h A, Rauch M, Turlin B, Preisser L, Bourd-Boittin K, Théret N, Gascan H, Piquet-Pellorce C, Samson M. Interleukin-33 overexpression is associated with liver fibrosis in mice and humans. J Cell Mol Med. 2010;14:1726-1739. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 167] [Cited by in RCA: 202] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 32. | Deleve LD, Wang X, Guo Y. Sinusoidal endothelial cells prevent rat stellate cell activation and promote reversion to quiescence. Hepatology. 2008;48:920-930. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 321] [Cited by in RCA: 295] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 33. | Li P, He K, Li J, Liu Z, Gong J. The role of Kupffer cells in hepatic diseases. Mol Immunol. 2017;85:222-229. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 165] [Article Influence: 20.6] [Reference Citation Analysis (0)] |
| 34. | López-Navarrete G, Ramos-Martínez E, Suárez-Álvarez K, Aguirre-García J, Ledezma-Soto Y, León-Cabrera S, Gudiño-Zayas M, Guzmán C, Gutiérrez-Reyes G, Hernández-Ruíz J, Camacho-Arroyo I, Robles-Díaz G, Kershenobich D, Terrazas LI, Escobedo G. Th2-associated alternative Kupffer cell activation promotes liver fibrosis without inducing local inflammation. Int J Biol Sci. 2011;7:1273-1286. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 36] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 35. | Canbay A, Feldstein AE, Higuchi H, Werneburg N, Grambihler A, Bronk SF, Gores GJ. Kupffer cell engulfment of apoptotic bodies stimulates death ligand and cytokine expression. Hepatology. 2003;38:1188-1198. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 364] [Cited by in RCA: 352] [Article Influence: 16.0] [Reference Citation Analysis (1)] |
| 36. | Noor MT, Manoria P. Immune Dysfunction in Cirrhosis. J Clin Transl Hepatol. 2017;5:50-58. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 84] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 37. | Tayal V, Kalra BS. Cytokines and anti-cytokines as therapeutics--an update. Eur J Pharmacol. 2008;579:1-12. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 123] [Cited by in RCA: 123] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 38. | Tsochatzis EA, Bosch J, Burroughs AK. Liver cirrhosis. Lancet. 2014;383:1749-1761. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1139] [Cited by in RCA: 1310] [Article Influence: 119.1] [Reference Citation Analysis (0)] |
| 39. | Ying HZ, Chen Q, Zhang WY, Zhang HH, Ma Y, Zhang SZ, Fang J, Yu CH. PDGF signaling pathway in hepatic fibrosis pathogenesis and therapeutics (Review). Mol Med Rep. 2017;16:7879-7889. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 188] [Cited by in RCA: 170] [Article Influence: 21.3] [Reference Citation Analysis (0)] |
| 40. | Mekala S, Tulimilli SV, Geesala R, Manupati K, Dhoke NR, Das A. Cellular crosstalk mediated by platelet-derived growth factor BB and transforming growth factor β during hepatic injury activates hepatic stellate cells. Can J Physiol Pharmacol. 2018;96:728-741. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 20] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 41. | Kikuchi A, Pradhan-Sundd T, Singh S, Nagarajan S, Loizos N, Monga SP. Platelet-Derived Growth Factor Receptor α Contributes to Human Hepatic Stellate Cell Proliferation and Migration. Am J Pathol. 2017;187:2273-2287. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 42] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 42. | Meng XM, Nikolic-Paterson DJ, Lan HY. TGF-β: the master regulator of fibrosis. Nat Rev Nephrol. 2016;12:325-338. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1512] [Cited by in RCA: 2546] [Article Influence: 282.9] [Reference Citation Analysis (0)] |
| 43. | Cai J, Hu M, Chen Z, Ling Z. The roles and mechanisms of hypoxia in liver fibrosis. J Transl Med. 2021;19:186. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 54] [Cited by in RCA: 66] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 44. | Cui Q, Wang Z, Jiang D, Qu L, Guo J, Li Z. HGF inhibits TGF-β1-induced myofibroblast differentiation and ECM deposition via MMP-2 in Achilles tendon in rat. Eur J Appl Physiol. 2011;111:1457-1463. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 45] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 45. | Zhou WC, Zhang QB, Qiao L. Pathogenesis of liver cirrhosis. World J Gastroenterol. 2014;20:7312-7324. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 335] [Cited by in RCA: 406] [Article Influence: 36.9] [Reference Citation Analysis (10)] |
| 46. | Connolly MK, Bedrosian AS, Mallen-St Clair J, Mitchell AP, Ibrahim J, Stroud A, Pachter HL, Bar-Sagi D, Frey AB, Miller G. In liver fibrosis, dendritic cells govern hepatic inflammation in mice via TNF-alpha. J Clin Invest. 2009;119:3213-3225. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 137] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 47. | Osawa Y, Hoshi M, Yasuda I, Saibara T, Moriwaki H, Kozawa O. Tumor necrosis factor-α promotes cholestasis-induced liver fibrosis in the mouse through tissue inhibitor of metalloproteinase-1 production in hepatic stellate cells. PLoS One. 2013;8:e65251. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 72] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 48. | Varela-Rey M, Fontán-Gabás L, Blanco P, López-Zabalza MJ, Iraburu MJ. Glutathione depletion is involved in the inhibition of procollagen alpha1(I) mRNA levels caused by TNF-alpha on hepatic stellate cells. Cytokine. 2007;37:212-217. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 17] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 49. | Sánchez-Valle V, Chávez-Tapia NC, Uribe M, Méndez-Sánchez N. Role of oxidative stress and molecular changes in liver fibrosis: a review. Curr Med Chem. 2012;19:4850-4860. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 452] [Cited by in RCA: 415] [Article Influence: 31.9] [Reference Citation Analysis (0)] |
| 50. | Masarone M, Rosato V, Dallio M, Gravina AG, Aglitti A, Loguercio C, Federico A, Persico M. Role of Oxidative Stress in Pathophysiology of Nonalcoholic Fatty Liver Disease. Oxid Med Cell Longev. 2018;2018:9547613. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 496] [Cited by in RCA: 482] [Article Influence: 68.9] [Reference Citation Analysis (0)] |
| 51. | Tsuchida T, Friedman SL. Mechanisms of hepatic stellate cell activation. Nat Rev Gastroenterol Hepatol. 2017;14:397-411. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1221] [Cited by in RCA: 1995] [Article Influence: 249.4] [Reference Citation Analysis (0)] |
| 52. | Ezhilarasan D. Oxidative stress is bane in chronic liver diseases: Clinical and experimental perspective. Arab J Gastroenterol. 2018;19:56-64. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 88] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 53. | Muriel P, Gordillo KR. Role of Oxidative Stress in Liver Health and Disease. Oxid Med Cell Longev. 2016;2016:9037051. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 49] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 54. | Liang S, Kisseleva T, Brenner DA. The Role of NADPH Oxidases (NOXs) in Liver Fibrosis and the Activation of Myofibroblasts. Front Physiol. 2016;7:17. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 106] [Cited by in RCA: 158] [Article Influence: 17.6] [Reference Citation Analysis (0)] |
| 55. | Crosas-Molist E, Fabregat I. Role of NADPH oxidases in the redox biology of liver fibrosis. Redox Biol. 2015;6:106-111. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 88] [Cited by in RCA: 108] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 56. | Aoyama T, Paik YH, Watanabe S, Laleu B, Gaggini F, Fioraso-Cartier L, Molango S, Heitz F, Merlot C, Szyndralewiez C, Page P, Brenner DA. Nicotinamide adenine dinucleotide phosphate oxidase in experimental liver fibrosis: GKT137831 as a novel potential therapeutic agent. Hepatology. 2012;56:2316-2327. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 230] [Cited by in RCA: 268] [Article Influence: 20.6] [Reference Citation Analysis (0)] |
| 57. | Alfadda AA, Sallam RM. Reactive oxygen species in health and disease. J Biomed Biotechnol. 2012;2012:936486. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 398] [Cited by in RCA: 479] [Article Influence: 36.8] [Reference Citation Analysis (0)] |
| 58. | Rani V, Deep G, Singh RK, Palle K, Yadav UC. Oxidative stress and metabolic disorders: Pathogenesis and therapeutic strategies. Life Sci. 2016;148:183-193. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 549] [Cited by in RCA: 759] [Article Influence: 84.3] [Reference Citation Analysis (0)] |
| 59. | Torok NJ. Dysregulation of redox pathways in liver fibrosis. Am J Physiol Gastrointest Liver Physiol. 2016;311:G667-G674. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 38] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 60. | Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843-854. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8672] [Cited by in RCA: 8876] [Article Influence: 277.4] [Reference Citation Analysis (0)] |
| 61. | Bandiera S, Pfeffer S, Baumert TF, Zeisel MB. miR-122--a key factor and therapeutic target in liver disease. J Hepatol. 2015;62:448-457. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 399] [Cited by in RCA: 490] [Article Influence: 49.0] [Reference Citation Analysis (0)] |
| 62. | Haneklaus M, Gerlic M, O'Neill LA, Masters SL. miR-223: infection, inflammation and cancer. J Intern Med. 2013;274:215-226. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 289] [Cited by in RCA: 328] [Article Influence: 27.3] [Reference Citation Analysis (0)] |
| 63. | Bauernfeind F, Rieger A, Schildberg FA, Knolle PA, Schmid-Burgk JL, Hornung V. NLRP3 inflammasome activity is negatively controlled by miR-223. J Immunol. 2012;189:4175-4181. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 309] [Cited by in RCA: 380] [Article Influence: 29.2] [Reference Citation Analysis (0)] |
| 64. | Li T, Morgan MJ, Choksi S, Zhang Y, Kim YS, Liu ZG. MicroRNAs modulate the noncanonical transcription factor NF-kappaB pathway by regulating expression of the kinase IKKalpha during macrophage differentiation. Nat Immunol. 2010;11:799-805. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 295] [Cited by in RCA: 280] [Article Influence: 18.7] [Reference Citation Analysis (0)] |
| 65. | Weseslindtner L, Macheleidt I, Eischeid H, Strassl R, Hofer H, Popow-Kraupp T, Dienes HP, Holzmann H, Odenthal M. Micro RNAs mir-106a, mir-122 and mir-197 are increased in severe acute viral hepatitis with coagulopathy. Liver Int. 2016;36:353-360. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 66. | Fukushima T, Hamada Y, Yamada H, Horii I. Changes of micro-RNA expression in rat liver treated by acetaminophen or carbon tetrachloride--regulating role of micro-RNA for RNA expression. J Toxicol Sci. 2007;32:401-409. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 67] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 67. | Roderburg C, Benz F, Vargas Cardenas D, Koch A, Janssen J, Vucur M, Gautheron J, Schneider AT, Koppe C, Kreggenwinkel K, Zimmermann HW, Luedde M, Trautwein C, Tacke F, Luedde T. Elevated miR-122 serum levels are an independent marker of liver injury in inflammatory diseases. Liver Int. 2015;35:1172-1184. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 100] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 68. | Starkey Lewis PJ, Dear J, Platt V, Simpson KJ, Craig DG, Antoine DJ, French NS, Dhaun N, Webb DJ, Costello EM, Neoptolemos JP, Moggs J, Goldring CE, Park BK. Circulating microRNAs as potential markers of human drug-induced liver injury. Hepatology. 2011;54:1767-1776. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 413] [Cited by in RCA: 408] [Article Influence: 29.1] [Reference Citation Analysis (0)] |
| 69. | Krauskopf J, Caiment F, Claessen SM, Johnson KJ, Warner RL, Schomaker SJ, Burt DA, Aubrecht J, Kleinjans JC. Application of high-throughput sequencing to circulating microRNAs reveals novel biomarkers for drug-induced liver injury. Toxicol Sci. 2015;143:268-276. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 80] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 70. | Xu H, He JH, Xiao ZD, Zhang QQ, Chen YQ, Zhou H, Qu LH. Liver-enriched transcription factors regulate microRNA-122 that targets CUTL1 during liver development. Hepatology. 2010;52:1431-1442. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 227] [Article Influence: 15.1] [Reference Citation Analysis (0)] |
| 71. | Cheung O, Puri P, Eicken C, Contos MJ, Mirshahi F, Maher JW, Kellum JM, Min H, Luketic VA, Sanyal AJ. Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression. Hepatology. 2008;48:1810-1820. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 583] [Cited by in RCA: 551] [Article Influence: 32.4] [Reference Citation Analysis (0)] |
| 72. | Girard M, Jacquemin E, Munnich A, Lyonnet S, Henrion-Caude A. miR-122, a paradigm for the role of microRNAs in the liver. J Hepatol. 2008;48:648-656. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 281] [Cited by in RCA: 295] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 73. | Negrini M, Gramantieri L, Sabbioni S, Croce CM. microRNA involvement in hepatocellular carcinoma. Anticancer Agents Med Chem. 2011;11:500-521. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 82] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 74. | Ura S, Honda M, Yamashita T, Ueda T, Takatori H, Nishino R, Sunakozaka H, Sakai Y, Horimoto K, Kaneko S. Differential microRNA expression between hepatitis B and hepatitis C leading disease progression to hepatocellular carcinoma. Hepatology. 2009;49:1098-1112. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 299] [Cited by in RCA: 306] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 75. | Coulouarn C, Factor VM, Andersen JB, Durkin ME, Thorgeirsson SS. Loss of miR-122 expression in liver cancer correlates with suppression of the hepatic phenotype and gain of metastatic properties. Oncogene. 2009;28:3526-3536. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 585] [Cited by in RCA: 590] [Article Influence: 36.9] [Reference Citation Analysis (0)] |
| 76. | Burchard J, Zhang C, Liu AM, Poon RT, Lee NP, Wong KF, Sham PC, Lam BY, Ferguson MD, Tokiwa G, Smith R, Leeson B, Beard R, Lamb JR, Lim L, Mao M, Dai H, Luk JM. microRNA-122 as a regulator of mitochondrial metabolic gene network in hepatocellular carcinoma. Mol Syst Biol. 2010;6:402. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 141] [Cited by in RCA: 165] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 77. | Waidmann O, Bihrer V, Pleli T, Farnik H, Berger A, Zeuzem S, Kronenberger B, Piiper A. Serum microRNA-122 levels in different groups of patients with chronic hepatitis B virus infection. J Viral Hepat. 2012;19:e58-e65. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 106] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 78. | Wang K, Zhang S, Marzolf B, Troisch P, Brightman A, Hu Z, Hood LE, Galas DJ. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc Natl Acad Sci U S A. 2009;106:4402-4407. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 891] [Cited by in RCA: 946] [Article Influence: 59.1] [Reference Citation Analysis (0)] |
| 79. | Bala S, Petrasek J, Mundkur S, Catalano D, Levin I, Ward J, Alao H, Kodys K, Szabo G. Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology. 2012;56:1946-1957. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 480] [Cited by in RCA: 525] [Article Influence: 40.4] [Reference Citation Analysis (0)] |
| 80. | An FM, Yu DS, Xie Q, Gong BD, Wang H, Guo Q, Yu H. [The role of miRNA-122 expression during the acute liver failure in mice induced by D-GalN/LPS]. Zhonghua Gan Zang Bing Za Zhi. 2010;18:527-532. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 81. | Zhang Y, Jia Y, Zheng R, Guo Y, Wang Y, Guo H, Fei M, Sun S. Plasma microRNA-122 as a biomarker for viral-, alcohol-, and chemical-related hepatic diseases. Clin Chem. 2010;56:1830-1838. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 307] [Cited by in RCA: 323] [Article Influence: 21.5] [Reference Citation Analysis (0)] |
| 82. | Wang S, Qiu L, Yan X, Jin W, Wang Y, Chen L, Wu E, Ye X, Gao GF, Wang F, Chen Y, Duan Z, Meng S. Loss of microRNA 122 expression in patients with hepatitis B enhances hepatitis B virus replication through cyclin G(1) -modulated P53 activity. Hepatology. 2012;55:730-741. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 213] [Cited by in RCA: 209] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 83. | Yu DS, An FM, Gong BD, Xiang XG, Lin LY, Wang H, Xie Q. The regulatory role of microRNA-1187 in TNF-α-mediated hepatocyte apoptosis in acute liver failure. Int J Mol Med. 2012;29:663-668. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 32] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 84. | Ji F, Yang B, Peng X, Ding H, You H, Tien P. Circulating microRNAs in hepatitis B virus-infected patients. J Viral Hepat. 2011;18:e242-e251. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 124] [Cited by in RCA: 122] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 85. | Zhang H, Li QY, Guo ZZ, Guan Y, Du J, Lu YY, Hu YY, Liu P, Huang S, Su SB. Serum levels of microRNAs can specifically predict liver injury of chronic hepatitis B. World J Gastroenterol. 2012;18:5188-5196. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 34] [Reference Citation Analysis (0)] |
| 86. | Su C, Hou Z, Zhang C, Tian Z, Zhang J. Ectopic expression of microRNA-155 enhances innate antiviral immunity against HBV infection in human hepatoma cells. Virol J. 2011;8:354. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 93] [Cited by in RCA: 109] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 87. | Zhang X, Chen C, Wu M, Chen L, Zhang J, Zhang X, Zhang Z, Wu J, Wang J, Chen X, Huang T, Yuan Z. Plasma microRNA profile as a predictor of early virological response to interferon treatment in chronic hepatitis B patients. Antivir Ther. 2012;17:1243-1253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 34] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 88. | Jopling CL, Yi M, Lancaster AM, Lemon SM, Sarnow P. Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science. 2005;309:1577-1581. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1993] [Cited by in RCA: 1986] [Article Influence: 99.3] [Reference Citation Analysis (0)] |
| 89. | Randall G, Panis M, Cooper JD, Tellinghuisen TL, Sukhodolets KE, Pfeffer S, Landthaler M, Landgraf P, Kan S, Lindenbach BD, Chien M, Weir DB, Russo JJ, Ju J, Brownstein MJ, Sheridan R, Sander C, Zavolan M, Tuschl T, Rice CM. Cellular cofactors affecting hepatitis C virus infection and replication. Proc Natl Acad Sci U S A. 2007;104:12884-12889. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 449] [Cited by in RCA: 438] [Article Influence: 24.3] [Reference Citation Analysis (0)] |
| 90. | Liu X, Wang T, Wakita T, Yang W. Systematic identification of microRNA and messenger RNA profiles in hepatitis C virus-infected human hepatoma cells. Virology. 2010;398:57-67. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 108] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 91. | Bandyopadhyay S, Friedman RC, Marquez RT, Keck K, Kong B, Icardi MS, Brown KE, Burge CB, Schmidt WN, Wang Y, McCaffrey AP. Hepatitis C virus infection and hepatic stellate cell activation downregulate miR-29: miR-29 overexpression reduces hepatitis C viral abundance in culture. J Infect Dis. 2011;203:1753-1762. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 118] [Cited by in RCA: 131] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 92. | Murakami Y, Tanaka M, Toyoda H, Hayashi K, Kuroda M, Tajima A, Shimotohno K. Hepatic microRNA expression is associated with the response to interferon treatment of chronic hepatitis C. BMC Med Genomics. 2010;3:48. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 47] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 93. | Baltimore D, Boldin MP, O'Connell RM, Rao DS, Taganov KD. MicroRNAs: new regulators of immune cell development and function. Nat Immunol. 2008;9:839-845. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 824] [Cited by in RCA: 889] [Article Influence: 52.3] [Reference Citation Analysis (0)] |
| 94. | Bala S, Szabo G. MicroRNA Signature in Alcoholic Liver Disease. Int J Hepatol. 2012;2012:498232. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 106] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 95. | Tang Y, Forsyth CB, Farhadi A, Rangan J, Jakate S, Shaikh M, Banan A, Fields JZ, Keshavarzian A. Nitric oxide-mediated intestinal injury is required for alcohol-induced gut leakiness and liver damage. Alcohol Clin Exp Res. 2009;33:1220-1230. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 88] [Cited by in RCA: 94] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 96. | Miranda RC, Pietrzykowski AZ, Tang Y, Sathyan P, Mayfield D, Keshavarzian A, Sampson W, Hereld D. MicroRNAs: master regulators of ethanol abuse and toxicity? Alcohol Clin Exp Res. 2010;34:575-587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 129] [Cited by in RCA: 135] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 97. | Murakami Y, Toyoda H, Tanaka M, Kuroda M, Harada Y, Matsuda F, Tajima A, Kosaka N, Ochiya T, Shimotohno K. The progression of liver fibrosis is related with overexpression of the miR-199 and 200 families. PLoS One. 2011;6:e16081. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 226] [Cited by in RCA: 238] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 98. | Venugopal SK, Jiang J, Kim TH, Li Y, Wang SS, Torok NJ, Wu J, Zern MA. Liver fibrosis causes downregulation of miRNA-150 and miRNA-194 in hepatic stellate cells, and their overexpression causes decreased stellate cell activation. Am J Physiol Gastrointest Liver Physiol. 2010;298:G101-G106. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 167] [Cited by in RCA: 182] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 99. | Boya P, Reggiori F, Codogno P. Emerging regulation and functions of autophagy. Nat Cell Biol. 2013;15:713-720. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 767] [Cited by in RCA: 969] [Article Influence: 80.8] [Reference Citation Analysis (0)] |
| 100. | Mizushima N, Levine B, Cuervo AM, Klionsky DJ. Autophagy fights disease through cellular self-digestion. Nature. 2008;451:1069-1075. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5421] [Cited by in RCA: 5293] [Article Influence: 311.4] [Reference Citation Analysis (0)] |
| 101. | Deretic V, Saitoh T, Akira S. Autophagy in infection, inflammation and immunity. Nat Rev Immunol. 2013;13:722-737. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1278] [Cited by in RCA: 1521] [Article Influence: 126.8] [Reference Citation Analysis (0)] |
| 102. | Mizushima N, Yoshimori T, Ohsumi Y. The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol. 2011;27:107-132. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2082] [Cited by in RCA: 2350] [Article Influence: 167.9] [Reference Citation Analysis (0)] |
| 103. | Choi AM, Ryter SW, Levine B. Autophagy in human health and disease. N Engl J Med. 2013;368:651-662. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1792] [Cited by in RCA: 1914] [Article Influence: 159.5] [Reference Citation Analysis (0)] |
| 104. | Jiang P, Mizushima N. Autophagy and human diseases. Cell Res. 2014;24:69-79. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 562] [Cited by in RCA: 632] [Article Influence: 52.7] [Reference Citation Analysis (0)] |
| 105. | Tooze SA, Yoshimori T. The origin of the autophagosomal membrane. Nat Cell Biol. 2010;12:831-835. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 405] [Cited by in RCA: 440] [Article Influence: 29.3] [Reference Citation Analysis (0)] |
| 106. | Yu L, Chen Y, Tooze SA. Autophagy pathway: Cellular and molecular mechanisms. Autophagy. 2018;14:207-215. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 911] [Cited by in RCA: 1073] [Article Influence: 134.1] [Reference Citation Analysis (0)] |
| 107. | Dikic I, Elazar Z. Mechanism and medical implications of mammalian autophagy. Nat Rev Mol Cell Biol. 2018;19:349-364. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1271] [Cited by in RCA: 2041] [Article Influence: 340.2] [Reference Citation Analysis (0)] |
| 108. | Tanida I. Autophagosome formation and molecular mechanism of autophagy. Antioxid Redox Signal. 2011;14:2201-2214. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 342] [Cited by in RCA: 385] [Article Influence: 27.5] [Reference Citation Analysis (0)] |
| 109. | Nakamura S, Yoshimori T. New insights into autophagosome-lysosome fusion. J Cell Sci. 2017;130:1209-1216. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 256] [Cited by in RCA: 326] [Article Influence: 40.8] [Reference Citation Analysis (0)] |
| 110. | Sarkar C, Zhao Z, Aungst S, Sabirzhanov B, Faden AI, Lipinski MM. Impaired autophagy flux is associated with neuronal cell death after traumatic brain injury. Autophagy. 2014;10:2208-2222. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 188] [Cited by in RCA: 263] [Article Influence: 26.3] [Reference Citation Analysis (0)] |
| 111. | Mizushima N. A brief history of autophagy from cell biology to physiology and disease. Nat Cell Biol. 2018;20:521-527. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 363] [Cited by in RCA: 509] [Article Influence: 72.7] [Reference Citation Analysis (0)] |
| 112. | Yin XM, Ding WX, Gao W. Autophagy in the liver. Hepatology. 2008;47:1773-1785. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 192] [Cited by in RCA: 211] [Article Influence: 12.4] [Reference Citation Analysis (0)] |
| 113. | Virgin HW, Levine B. Autophagy genes in immunity. Nat Immunol. 2009;10:461-470. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 368] [Cited by in RCA: 350] [Article Influence: 21.9] [Reference Citation Analysis (0)] |
| 114. | Liu K, Zhao E, Ilyas G, Lalazar G, Lin Y, Haseeb M, Tanaka KE, Czaja MJ. Impaired macrophage autophagy increases the immune response in obese mice by promoting proinflammatory macrophage polarization. Autophagy. 2015;11:271-284. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 251] [Cited by in RCA: 395] [Article Influence: 43.9] [Reference Citation Analysis (0)] |
| 115. | Codogno P, Mehrpour M, Proikas-Cezanne T. Canonical and non-canonical autophagy: variations on a common theme of self-eating? Nat Rev Mol Cell Biol. 2011;13:7-12. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 383] [Cited by in RCA: 443] [Article Influence: 31.6] [Reference Citation Analysis (0)] |
| 116. | Heckmann BL, Boada-Romero E, Cunha LD, Magne J, Green DR. LC3-Associated Phagocytosis and Inflammation. J Mol Biol. 2017;429:3561-3576. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 155] [Cited by in RCA: 205] [Article Influence: 25.6] [Reference Citation Analysis (0)] |
| 117. | Münz C. Non-canonical Functions of Macroautophagy Proteins During Endocytosis by Myeloid Antigen Presenting Cells. Front Immunol. 2018;9:2765. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 15] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 118. | Schneider JL, Cuervo AM. Liver autophagy: much more than just taking out the trash. Nat Rev Gastroenterol Hepatol. 2014;11:187-200. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 132] [Cited by in RCA: 150] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 119. | Gutierrez MG, Master SS, Singh SB, Taylor GA, Colombo MI, Deretic V. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell. 2004;119:753-766. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1625] [Cited by in RCA: 1750] [Article Influence: 83.3] [Reference Citation Analysis (0)] |
| 120. | Kroemer G, Mariño G, Levine B. Autophagy and the integrated stress response. Mol Cell. 2010;40:280-293. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2886] [Cited by in RCA: 2759] [Article Influence: 183.9] [Reference Citation Analysis (0)] |
| 121. | Yonekawa T, Thorburn A. Autophagy and cell death. Essays Biochem. 2013;55:105-117. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 167] [Article Influence: 15.2] [Reference Citation Analysis (0)] |
| 122. | Boya P, González-Polo RA, Casares N, Perfettini JL, Dessen P, Larochette N, Métivier D, Meley D, Souquere S, Yoshimori T, Pierron G, Codogno P, Kroemer G. Inhibition of macroautophagy triggers apoptosis. Mol Cell Biol. 2005;25:1025-1040. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1289] [Cited by in RCA: 1320] [Article Influence: 66.0] [Reference Citation Analysis (0)] |
| 123. | Nikoletopoulou V, Markaki M, Palikaras K, Tavernarakis N. Crosstalk between apoptosis, necrosis and autophagy. Biochim Biophys Acta. 2013;1833:3448-3459. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 844] [Cited by in RCA: 998] [Article Influence: 83.2] [Reference Citation Analysis (0)] |
| 124. | Wang K. Autophagy and apoptosis in liver injury. Cell Cycle. 2015;14:1631-1642. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 299] [Article Influence: 33.2] [Reference Citation Analysis (0)] |
| 125. | Ni HM, Bockus A, Boggess N, Jaeschke H, Ding WX. Activation of autophagy protects against acetaminophen-induced hepatotoxicity. Hepatology. 2012;55:222-232. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 314] [Cited by in RCA: 355] [Article Influence: 27.3] [Reference Citation Analysis (0)] |
| 126. | Lockshin RA, Zakeri Z. Cell death in health and disease. J Cell Mol Med. 2007;11:1214-1224. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 143] [Cited by in RCA: 142] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 127. | Strozyk E, Kulms D. The role of AKT/mTOR pathway in stress response to UV-irradiation: implication in skin carcinogenesis by regulation of apoptosis, autophagy and senescence. Int J Mol Sci. 2013;14:15260-15285. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 114] [Cited by in RCA: 113] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 128. | Gao M, Monian P, Pan Q, Zhang W, Xiang J, Jiang X. Ferroptosis is an autophagic cell death process. Cell Res. 2016;26:1021-1032. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 612] [Cited by in RCA: 1326] [Article Influence: 147.3] [Reference Citation Analysis (2)] |
| 129. | Guo H, Callaway JB, Ting JP. Inflammasomes: mechanism of action, role in disease, and therapeutics. Nat Med. 2015;21:677-687. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1797] [Cited by in RCA: 2484] [Article Influence: 248.4] [Reference Citation Analysis (0)] |
| 130. | Shibutani ST, Saitoh T, Nowag H, Münz C, Yoshimori T. Autophagy and autophagy-related proteins in the immune system. Nat Immunol. 2015;16:1014-1024. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 348] [Cited by in RCA: 430] [Article Influence: 43.0] [Reference Citation Analysis (0)] |
| 131. | Sanjuan MA, Dillon CP, Tait SW, Moshiach S, Dorsey F, Connell S, Komatsu M, Tanaka K, Cleveland JL, Withoff S, Green DR. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature. 2007;450:1253-1257. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 968] [Cited by in RCA: 1090] [Article Influence: 64.1] [Reference Citation Analysis (0)] |
| 132. | Schroder K, Tschopp J. The inflammasomes. Cell. 2010;140:821-832. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3880] [Cited by in RCA: 4547] [Article Influence: 303.1] [Reference Citation Analysis (0)] |
| 133. | Wang Z, Zhang S, Xiao Y, Zhang W, Wu S, Qin T, Yue Y, Qian W, Li L. NLRP3 Inflammasome and Inflammatory Diseases. Oxid Med Cell Longev. 2020;2020:4063562. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 64] [Cited by in RCA: 156] [Article Influence: 31.2] [Reference Citation Analysis (0)] |
| 134. | Thorburn A. Autophagy and its effects: making sense of double-edged swords. PLoS Biol. 2014;12:e1001967. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 73] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 135. | White E. Deconvoluting the context-dependent role for autophagy in cancer. Nat Rev Cancer. 2012;12:401-410. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1226] [Cited by in RCA: 1380] [Article Influence: 106.2] [Reference Citation Analysis (0)] |
| 136. | Codogno P, Meijer AJ. Autophagy in the liver. J Hepatol. 2013;59:389-391. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 34] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 137. | Ueno T, Komatsu M. Autophagy in the liver: functions in health and disease. Nat Rev Gastroenterol Hepatol. 2017;14:170-184. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 281] [Cited by in RCA: 398] [Article Influence: 49.8] [Reference Citation Analysis (0)] |
| 138. | Allaire M, Rautou PE, Codogno P, Lotersztajn S. Autophagy in liver diseases: Time for translation? J Hepatol. 2019;70:985-998. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 181] [Cited by in RCA: 289] [Article Influence: 48.2] [Reference Citation Analysis (0)] |
| 139. | Madrigal-Matute J, Cuervo AM. Regulation of Liver Metabolism by Autophagy. Gastroenterology. 2016;150:328-339. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 242] [Cited by in RCA: 247] [Article Influence: 27.4] [Reference Citation Analysis (0)] |
| 140. | Hazari Y, Bravo-San Pedro JM, Hetz C, Galluzzi L, Kroemer G. Autophagy in hepatic adaptation to stress. J Hepatol. 2020;72:183-196. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 125] [Article Influence: 25.0] [Reference Citation Analysis (0)] |
| 141. | Youle RJ, Narendra DP. Mechanisms of mitophagy. Nat Rev Mol Cell Biol. 2011;12:9-14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2666] [Cited by in RCA: 2518] [Article Influence: 179.9] [Reference Citation Analysis (0)] |
| 142. | Schuppan D, Afdhal NH. Liver cirrhosis. Lancet. 2008;371:838-851. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1686] [Cited by in RCA: 1565] [Article Influence: 92.1] [Reference Citation Analysis (0)] |
| 143. | Smith A, Baumgartner K, Bositis C. Cirrhosis: Diagnosis and Management. Am Fam Physician. 2019;100:759-770. [PubMed] |
| 144. | Roehlen N, Crouchet E, Baumert TF. Liver Fibrosis: Mechanistic Concepts and Therapeutic Perspectives. Cells. 2020;9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 198] [Cited by in RCA: 741] [Article Influence: 148.2] [Reference Citation Analysis (0)] |
| 145. | Turkseven S, Bolognesi M, Brocca A, Pesce P, Angeli P, Di Pascoli M. Mitochondria-targeted antioxidant mitoquinone attenuates liver inflammation and fibrosis in cirrhotic rats. Am J Physiol Gastrointest Liver Physiol. 2020;318:G298-G304. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 41] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 146. | Schuppan D, Ashfaq-Khan M, Yang AT, Kim YO. Liver fibrosis: Direct antifibrotic agents and targeted therapies. Matrix Biol. 2018;68-69:435-451. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 207] [Cited by in RCA: 331] [Article Influence: 47.3] [Reference Citation Analysis (0)] |
| 147. | Sohrabpour AA, Mohamadnejad M, Malekzadeh R. Review article: the reversibility of cirrhosis. Aliment Pharmacol Ther. 2012;36:824-832. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 48] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 148. | Li S, Wang N, Feng YG, Li HY, Feng Y. Antioxidants in the Prevention and Treatment of Liver Diseases. In: Al-Gubory KH, Laher I. Nutritional Antioxidant Therapies: Treatments and Perspectives. Cham: Springer International Publishing; 2017: 467-491. |
| 149. | Federico A, Dallio M, Loguercio C. Silymarin/Silybin and Chronic Liver Disease: A Marriage of Many Years. Molecules. 2017;22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 203] [Cited by in RCA: 290] [Article Influence: 36.3] [Reference Citation Analysis (0)] |
| 150. | Tóth RJ, Csapó J. The role of selenium in nutrition - A review. Acta Univ Sapientiae Aliment. 2018;11:128-144. [RCA] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 17] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 151. | Li B, Li W, Tian Y, Guo S, Qian L, Xu D, Cao N. Selenium-Alleviated Hepatocyte Necrosis and DNA Damage in Cyclophosphamide-Treated Geese by Mitigating Oxidative Stress. Biol Trace Elem Res. 2020;193:508-516. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 19] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 152. | Du H, Zheng Y, Zhang W, Tang H, Jing B, Li H, Xu F, Lin J, Fu H, Chang L, Shu G. Nano-Selenium Alleviates Cadmium-Induced Acute Hepatic Toxicity by Decreasing Oxidative Stress and Activating the Nrf2 Pathway in Male Kunming Mice. Front Vet Sci. 2022;9:942189. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 8] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 153. | Hamid M, Abdulrahim Y, Liu D, Qian G, Khan A, Huang K. The Hepatoprotective Effect of Selenium-Enriched Yeast and Gum Arabic Combination on Carbon Tetrachloride-Induced Chronic Liver Injury in Rats. J Food Sci. 2018;83:525-534. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 34] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 154. | Al-Dossari MH, Fadda LM, Attia HA, Hasan IH, Mahmoud AM. Curcumin and Selenium Prevent Lipopolysaccharide/Diclofenac-Induced Liver Injury by Suppressing Inflammation and Oxidative Stress. Biol Trace Elem Res. 2020;196:173-183. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 47] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 155. | Nagashimada M, Ota T. Role of vitamin E in nonalcoholic fatty liver disease. IUBMB Life. 2019;71:516-522. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 71] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 156. | Sanyal AJ, Chalasani N, Kowdley KV, McCullough A, Diehl AM, Bass NM, Neuschwander-Tetri BA, Lavine JE, Tonascia J, Unalp A, Van Natta M, Clark J, Brunt EM, Kleiner DE, Hoofnagle JH, Robuck PR; NASH CRN. Pioglitazone, vitamin E, or placebo for nonalcoholic steatohepatitis. N Engl J Med. 2010;362:1675-1685. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2642] [Cited by in RCA: 2469] [Article Influence: 164.6] [Reference Citation Analysis (2)] |
| 157. | Aljuhr SA, Abdelaziz G, Essa BM, Zaghary WA, Sakr TM. Hepatoprotective, antioxidant and anti-inflammatory potentials of Vit-E/C@SeNPs in rats: Synthesis, characterization, biochemical, radio-biodistribution, molecular and histopathological studies. Bioorg Chem. 2021;117:105412. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 12] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 158. | Millea PJ. N-acetylcysteine: multiple clinical applications. Am Fam Physician. 2009;80:265-269. [PubMed] |
| 159. | Fisher ES, Curry SC. Evaluation and treatment of acetaminophen toxicity. Adv Pharmacol. 2019;85:263-272. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 114] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 160. | Pereira-Filho G, Ferreira C, Schwengber A, Marroni C, Zettler C, Marroni N. Role of N-acetylcysteine on fibrosis and oxidative stress in cirrhotic rats. Arq Gastroenterol. 2008;45:156-162. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 49] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 161. | Bond ST, Kim J, Calkin AC, Drew BG. The Antioxidant Moiety of MitoQ Imparts Minimal Metabolic Effects in Adipose Tissue of High Fat Fed Mice. Front Physiol. 2019;10:543. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 28] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 162. | Rehman H, Liu Q, Krishnasamy Y, Shi Z, Ramshesh VK, Haque K, Schnellmann RG, Murphy MP, Lemasters JJ, Rockey DC, Zhong Z. The mitochondria-targeted antioxidant MitoQ attenuates liver fibrosis in mice. Int J Physiol Pathophysiol Pharmacol. 2016;8:14-27. [PubMed] |
| 163. | Dong V, Nanchal R, Karvellas CJ. Pathophysiology of Acute Liver Failure. Nutr Clin Pract. 2020;35:24-29. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 99] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 164. | Seki E, Schwabe RF. Hepatic inflammation and fibrosis: functional links and key pathways. Hepatology. 2015;61:1066-1079. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 548] [Cited by in RCA: 741] [Article Influence: 74.1] [Reference Citation Analysis (0)] |
| 165. | Taniguchi H, Ebina M, Kondoh Y, Ogura T, Azuma A, Suga M, Taguchi Y, Takahashi H, Nakata K, Sato A, Takeuchi M, Raghu G, Kudoh S, Nukiwa T; Pirfenidone Clinical Study Group in Japan. Pirfenidone in idiopathic pulmonary fibrosis. Eur Respir J. 2010;35:821-829. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 650] [Cited by in RCA: 729] [Article Influence: 45.6] [Reference Citation Analysis (0)] |
| 166. | Ruwanpura SM, Thomas BJ, Bardin PG. Pirfenidone: Molecular Mechanisms and Potential Clinical Applications in Lung Disease. Am J Respir Cell Mol Biol. 2020;62:413-422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 167] [Article Influence: 33.4] [Reference Citation Analysis (0)] |
| 167. | Flores-Contreras L, Sandoval-Rodríguez AS, Mena-Enriquez MG, Lucano-Landeros S, Arellano-Olivera I, Alvarez-Álvarez A, Sanchez-Parada MG, Armendáriz-Borunda J. Treatment with pirfenidone for two years decreases fibrosis, cytokine levels and enhances CB2 gene expression in patients with chronic hepatitis C. BMC Gastroenterol. 2014;14:131. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 62] [Cited by in RCA: 70] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 168. | Floreani A, De Martin S. Treatment of primary sclerosing cholangitis. Dig Liver Dis. 2021;53:1531-1538. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 23] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 169. | Komori A. Recent updates on the management of autoimmune hepatitis. Clin Mol Hepatol. 2021;27:58-69. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 81] [Cited by in RCA: 96] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 170. | Montano-Loza AJ, Thandassery RB, Czaja AJ. Targeting Hepatic Fibrosis in Autoimmune Hepatitis. Dig Dis Sci. 2016;61:3118-3139. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 26] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 171. | Abhyankar A, Tapper E, Bonder A. Immunosuppressive therapy in immune-mediated liver disease in the non-transplanted patient. Pharmaceuticals (Basel). 2013;7:18-28. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 172. | Puche JE, Saiman Y, Friedman SL. Hepatic stellate cells and liver fibrosis. Compr Physiol. 2013;3:1473-1492. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 421] [Cited by in RCA: 570] [Article Influence: 51.8] [Reference Citation Analysis (0)] |
| 173. | Pesce A, Ciurleo R, Bramanti A, Armeli Iapichino EC, Petralia MC, Magro GG, Fagone P, Bramanti P, Nicoletti F, Mangano K. Effects of Combined Admistration of Imatinib and Sorafenib in a Murine Model of Liver Fibrosis. Molecules. 2020;25. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 174. | Trappoliere M, Caligiuri A, Schmid M, Bertolani C, Failli P, Vizzutti F, Novo E, di Manzano C, Marra F, Loguercio C, Pinzani M. Silybin, a component of sylimarin, exerts anti-inflammatory and anti-fibrogenic effects on human hepatic stellate cells. J Hepatol. 2009;50:1102-1111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 177] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 175. | Zhou J, Zhong DW, Wang QW, Miao XY, Xu XD. Paclitaxel ameliorates fibrosis in hepatic stellate cells via inhibition of TGF-beta/Smad activity. World J Gastroenterol. 2010;16:3330-3334. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 46] [Cited by in RCA: 49] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 176. | Mu M, Zuo S, Wu RM, Deng KS, Lu S, Zhu JJ, Zou GL, Yang J, Cheng ML, Zhao XK. Ferulic acid attenuates liver fibrosis and hepatic stellate cell activation via inhibition of TGF-β/Smad signaling pathway. Drug Des Devel Ther. 2018;12:4107-4115. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 145] [Article Influence: 20.7] [Reference Citation Analysis (0)] |
| 177. | Cheng Q, Li C, Yang CF, Zhong YJ, Wu D, Shi L, Chen L, Li YW, Li L. Methyl ferulic acid attenuates liver fibrosis and hepatic stellate cell activation through the TGF-β1/Smad and NOX4/ROS pathways. Chem Biol Interact. 2019;299:131-139. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 92] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 178. | Zhang SS, Gong ZJ, Li WH, Wang X, Ling TY. Antifibrotic effect of curcumin in TGF-β 1-induced myofibroblasts from human oral mucosa. Asian Pac J Cancer Prev. 2012;13:289-294. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 35] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 179. | Zhang F, Zhang Z, Chen L, Kong D, Zhang X, Lu C, Lu Y, Zheng S. Curcumin attenuates angiogenesis in liver fibrosis and inhibits angiogenic properties of hepatic stellate cells. J Cell Mol Med. 2014;18:1392-1406. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 100] [Cited by in RCA: 113] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 180. | Kong D, Zhang Z, Chen L, Huang W, Zhang F, Wang L, Wang Y, Cao P, Zheng S. Curcumin blunts epithelial-mesenchymal transition of hepatocytes to alleviate hepatic fibrosis through regulating oxidative stress and autophagy. Redox Biol. 2020;36:101600. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 64] [Cited by in RCA: 170] [Article Influence: 34.0] [Reference Citation Analysis (0)] |
| 181. | Fujimoto J. Gene therapy for liver cirrhosis. J Gastroenterol Hepatol. 2000;15 Suppl:D33-D36. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 21] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 182. | Wang LS, Wang H, Zhang QL, Yang ZJ, Kong FX, Wu CT. Hepatocyte Growth Factor Gene Therapy for Ischemic Diseases. Hum Gene Ther. 2018;29:413-423. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 37] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 183. | Iimuro Y, Nishio T, Morimoto T, Nitta T, Stefanovic B, Choi SK, Brenner DA, Yamaoka Y. Delivery of matrix metalloproteinase-1 attenuates established liver fibrosis in the rat. Gastroenterology. 2003;124:445-458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 179] [Cited by in RCA: 190] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 184. | Cheng K, Mahato RI. Gene modulation for treating liver fibrosis. Crit Rev Ther Drug Carrier Syst. 2007;24:93-146. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 37] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 185. | Alessandrini M, Preynat-Seauve O, De Bruin K, Pepper MS. Stem cell therapy for neurological disorders. S Afr Med J. 2019;109:70-77. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 75] [Article Influence: 12.5] [Reference Citation Analysis (0)] |
| 186. | Wang J, Sun M, Liu W, Li Y, Li M. Stem Cell-Based Therapies for Liver Diseases: An Overview and Update. Tissue Eng Regen Med. 2019;16:107-118. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 50] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 187. | Eom YW, Shim KY, Baik SK. Mesenchymal stem cell therapy for liver fibrosis. Korean J Intern Med. 2015;30:580-589. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 140] [Cited by in RCA: 162] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 188. | Watanabe Y, Tsuchiya A, Seino S, Kawata Y, Kojima Y, Ikarashi S, Starkey Lewis PJ, Lu WY, Kikuta J, Kawai H, Yamagiwa S, Forbes SJ, Ishii M, Terai S. Mesenchymal Stem Cells and Induced Bone Marrow-Derived Macrophages Synergistically Improve Liver Fibrosis in Mice. Stem Cells Transl Med. 2019;8:271-284. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 73] [Cited by in RCA: 111] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 189. | Sun K, Xie X, Xie J, Jiao S, Chen X, Zhao X, Wang X, Wei L. Cell-based therapy for acute and chronic liver failures: distinct diseases, different choices. Sci Rep. 2014;4:6494. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 27] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 190. | Park S, In Hwang S, Kim J, Hwang S, Kang S, Yang S, Kang W, Kim KH, Han DW, Paik YH. The therapeutic potential of induced hepatocyte-like cells generated by direct reprogramming on hepatic fibrosis. Stem Cell Res Ther. 2019;10:21. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 17] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 191. | Woo DH, Kim SK, Lim HJ, Heo J, Park HS, Kang GY, Kim SE, You HJ, Hoeppner DJ, Kim Y, Kwon H, Choi TH, Lee JH, Hong SH, Song KW, Ahn EK, Chenoweth JG, Tesar PJ, McKay RD, Kim JH. Direct and indirect contribution of human embryonic stem cell-derived hepatocyte-like cells to liver repair in mice. Gastroenterology. 2012;142:602-611. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 107] [Cited by in RCA: 109] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 192. | Nadi A, Moradi L, Ai J, Asadpour S. Stem Cells and Hydrogels for Liver Tissue Engineering: Synergistic Cure for Liver Regeneration. Stem Cell Rev Rep. 2020;16:1092-1104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 193. | Lambrecht J, van Grunsven LA, Tacke F. Current and emerging pharmacotherapeutic interventions for the treatment of liver fibrosis. Expert Opin Pharmacother. 2020;21:1637-1650. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 46] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 194. | Albillos A, de Gottardi A, Rescigno M. The gut-liver axis in liver disease: Pathophysiological basis for therapy. J Hepatol. 2020;72:558-577. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 542] [Cited by in RCA: 1261] [Article Influence: 252.2] [Reference Citation Analysis (1)] |
| 195. | Milosevic I, Vujovic A, Barac A, Djelic M, Korac M, Radovanovic Spurnic A, Gmizic I, Stevanovic O, Djordjevic V, Lekic N, Russo E, Amedei A. Gut-Liver Axis, Gut Microbiota, and Its Modulation in the Management of Liver Diseases: A Review of the Literature. Int J Mol Sci. 2019;20. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 393] [Cited by in RCA: 352] [Article Influence: 58.7] [Reference Citation Analysis (0)] |
| 196. | Godoy P, Hewitt NJ, Albrecht U, Andersen ME, Ansari N, Bhattacharya S, Bode JG, Bolleyn J, Borner C, Böttger J, Braeuning A, Budinsky RA, Burkhardt B, Cameron NR, Camussi G, Cho CS, Choi YJ, Craig Rowlands J, Dahmen U, Damm G, Dirsch O, Donato MT, Dong J, Dooley S, Drasdo D, Eakins R, Ferreira KS, Fonsato V, Fraczek J, Gebhardt R, Gibson A, Glanemann M, Goldring CE, Gómez-Lechón MJ, Groothuis GM, Gustavsson L, Guyot C, Hallifax D, Hammad S, Hayward A, Häussinger D, Hellerbrand C, Hewitt P, Hoehme S, Holzhütter HG, Houston JB, Hrach J, Ito K, Jaeschke H, Keitel V, Kelm JM, Kevin Park B, Kordes C, Kullak-Ublick GA, LeCluyse EL, Lu P, Luebke-Wheeler J, Lutz A, Maltman DJ, Matz-Soja M, McMullen P, Merfort I, Messner S, Meyer C, Mwinyi J, Naisbitt DJ, Nussler AK, Olinga P, Pampaloni F, Pi J, Pluta L, Przyborski SA, Ramachandran A, Rogiers V, Rowe C, Schelcher C, Schmich K, Schwarz M, Singh B, Stelzer EH, Stieger B, Stöber R, Sugiyama Y, Tetta C, Thasler WE, Vanhaecke T, Vinken M, Weiss TS, Widera A, Woods CG, Xu JJ, Yarborough KM, Hengstler JG. Recent advances in 2D and 3D in vitro systems using primary hepatocytes, alternative hepatocyte sources and non-parenchymal liver cells and their use in investigating mechanisms of hepatotoxicity, cell signaling and ADME. Arch Toxicol. 2013;87:1315-1530. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1026] [Cited by in RCA: 977] [Article Influence: 81.4] [Reference Citation Analysis (0)] |
| 197. | Parola M, Pinzani M. Liver fibrosis: Pathophysiology, pathogenetic targets and clinical issues. Mol Aspects Med. 2019;65:37-55. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 286] [Cited by in RCA: 783] [Article Influence: 111.9] [Reference Citation Analysis (0)] |
| 198. | Hu Q, Zhang W, Wu Z, Tian X, Xiang J, Li L, Li Z, Peng X, Wei S, Ma X, Zhao Y. Baicalin and the liver-gut system: Pharmacological bases explaining its therapeutic effects. Pharmacol Res. 2021;165:105444. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 138] [Article Influence: 34.5] [Reference Citation Analysis (0)] |
| 199. | Chen J, Vitetta L. Gut Microbiota Metabolites in NAFLD Pathogenesis and Therapeutic Implications. Int J Mol Sci. 2020;21. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 48] [Cited by in RCA: 206] [Article Influence: 41.2] [Reference Citation Analysis (0)] |
| 200. | Paolella G, Mandato C, Pierri L, Poeta M, Di Stasi M, Vajro P. Gut-liver axis and probiotics: their role in non-alcoholic fatty liver disease. World J Gastroenterol. 2014;20:15518-15531. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 122] [Cited by in RCA: 151] [Article Influence: 13.7] [Reference Citation Analysis (3)] |
| 201. | Frasinariu OE, Ceccarelli S, Alisi A, Moraru E, Nobili V. Gut-liver axis and fibrosis in nonalcoholic fatty liver disease: an input for novel therapies. Dig Liver Dis. 2013;45:543-551. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 75] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 202. | Tee JK, Peng F, Ho HK. Effects of inorganic nanoparticles on liver fibrosis: Optimizing a double-edged sword for therapeutics. Biochem Pharmacol. 2019;160:24-33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 20] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 203. | Wakaskar RR. General overview of lipid-polymer hybrid nanoparticles, dendrimers, micelles, liposomes, spongosomes and cubosomes. J Drug Target. 2018;26:311-318. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 112] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 204. | Abdullah AS, El Sayed IET, El-Torgoman AMA, Alghamdi NA, Ullah S, Wageh S, Kamel MA. Preparation and Characterization of Silymarin-Conjugated Gold Nanoparticles with Enhanced Anti-Fibrotic Therapeutic Effects against Hepatic Fibrosis in Rats: Role of MicroRNAs as Molecular Targets. Biomedicines. 2021;9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 29] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 205. | Wang J, Pan W, Wang Y, Lei W, Feng B, Du C, Wang XJ. Enhanced efficacy of curcumin with phosphatidylserine-decorated nanoparticles in the treatment of hepatic fibrosis. Drug Deliv. 2018;25:1-11. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 34] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 206. | Liu CH, Chan KM, Chiang T, Liu JY, Chern GG, Hsu FF, Wu YH, Liu YC, Chen Y. Dual-Functional Nanoparticles Targeting CXCR4 and Delivering Antiangiogenic siRNA Ameliorate Liver Fibrosis. Mol Pharm. 2016;13:2253-2262. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 21] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 207. | Li Y, Pu S, Liu Q, Li R, Zhang J, Wu T, Chen L, Li H, Yang X, Zou M, Xiao J, Xie W, He J. An integrin-based nanoparticle that targets activated hepatic stellate cells and alleviates liver fibrosis. J Control Release. 2019;303:77-90. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 70] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 208. | Cullis PR, Hope MJ. Lipid Nanoparticle Systems for Enabling Gene Therapies. Mol Ther. 2017;25:1467-1475. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 364] [Cited by in RCA: 760] [Article Influence: 95.0] [Reference Citation Analysis (0)] |
| 209. | Wang HY, Li C, Liu WH, Deng FM, Ma Y, Guo LN, Kong H, Hu KA, Liu Q, Wu J, Sun J, Liu YL. Autophagy inhibition via Becn1 downregulation improves the mesenchymal stem cells antifibrotic potential in experimental liver fibrosis. J Cell Physiol. 2020;235:2722-2737. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 30] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 210. | Meng D, Li Z, Wang G, Ling L, Wu Y, Zhang C. Carvedilol attenuates liver fibrosis by suppressing autophagy and promoting apoptosis in hepatic stellate cells. Biomed Pharmacother. 2018;108:1617-1627. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 96] [Article Influence: 13.7] [Reference Citation Analysis (0)] |
| 211. | Xiu AY, Ding Q, Li Z, Zhang CQ. Doxazosin Attenuates Liver Fibrosis by Inhibiting Autophagy in Hepatic Stellate Cells via Activation of the PI3K/Akt/mTOR Signaling Pathway. Drug Des Devel Ther. 2021;15:3643-3659. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 48] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 212. | Yang M, Song XQ, Han M, Liu H. The role of Resolvin D1 in liver diseases. Prostaglandins Other Lipid Mediat. 2022;160:106634. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 15] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 213. | Li J, Deng X, Wang S, Jiang Q, Xu K. Resolvin D1 attenuates CCl4 Induced Liver Fibrosis by Inhibiting Autophagy-Mediated HSC activation via AKT/mTOR Pathway. Front Pharmacol. 2021;12:792414. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 27] [Article Influence: 6.8] [Reference Citation Analysis (0)] |