Published online Feb 27, 2018. doi: 10.4254/wjh.v10.i2.213
Peer-review started: November 23, 2017
First decision: December 18, 2017
Revised: December 28, 2017
Accepted: February 5, 2018
Article in press: February 5, 2018
Published online: February 27, 2018
Processing time: 102 Days and 22.5 Hours
There is wide agreement that cell fusion is a physiological process in cells in mammalian bone, muscle and placenta. In other organs, such as the cerebellum, cell fusion is controversial. The liver contains a considerable number of polyploid cells: They are commonly believed to originate by genome endoreplication, although the contribution of cell fusion to polyploidization has not been excluded. Here, we address the topic of cell fusion in the liver from a historical point of view. We discuss experimental evidence clearly supporting the hypothesis that cell fusion occurs in the liver, specifically when bone marrow cells were injected into mice and shown to rescue genetic hepatic degenerative defects. Those experiments-carried out in the latter half of the last century-were initially interpreted to show “transdifferentiation”, but are now believed to demonstrate fusion between donor macrophages and host hepatocytes, raising the possibility that physiologically polyploid cells, such as hepatocytes, could originate, at least partially, through homotypic cell fusion. In support of the homotypic cell fusion hypothesis, we present new data generated using a chimera-based model, a much simpler model than those previously used. Cell fusion as a road to polyploidization in the liver has not been extensively investigated, and its contribution to a variety of conditions, such as viral infections, carcinogenesis and aging, remains unclear.
Core tip: About 70% of hepatocytes are polyploid, arising either from genome duplication without division (endoreplication) or from cell fusion. Experiments with chimeric mice containing two cell populations each bearing a different genetic marker had shown that some liver cells express markers of both genomes, suggesting that cell fusion occurred. Here, we review the data in the literature and describe new experiments using a chimeric model that confirms that cell fusion contributes to liver polyploidy. We argue that the role of cell fusion in pathological conditions, such as viral hepatitis and neoplastic transformation, is worth further study.
- Citation: Lizier M, Castelli A, Montagna C, Lucchini F, Vezzoni P, Faggioli F. Cell fusion in the liver, revisited. World J Hepatol 2018; 10(2): 213-221
- URL: https://www.wjgnet.com/1948-5182/full/v10/i2/213.htm
- DOI: https://dx.doi.org/10.4254/wjh.v10.i2.213
Mammalian cells are usually diploid, with the exception of mature gametes, which are haploid. Interestingly, a few tissues contain polyploid cells, such as muscle cells, osteoclasts, hepatocytes, megakaryocytes and trophoblasts[1]. A common feature in all these tissues is the presence of a diploid progenitor cell that at some point during the differentiation/maturation process becomes polyploid. Polyploidization can be explained by two main mechanisms: endoreplication and cell fusion. Endoreplication occurs when the genome is duplicated without cell division, whereas cell fusion occurs between two different cells, either of the same or of a different identity. In the latter case, the genomes from two different cell types-which can come even from different species-fuse within the same membrane and, therefore, coexist within the same cell[2]. For both mechanisms, the outcome is a polyploid cell.
It is generally recognized that polyploidization in normal organisms is adaptive since it helps specialized cells acquire the ability to perform new, specific functions[3]. For example, osteoclasts are large multinucleated cells that perform the difficult task of resorbing bone matrix. This specialized function cannot be accomplished, for example, by mononucleated TRAP+ osteoclasts as this leads to osteopetrosis, a disease in which bone is not degraded[4]. Among the other types of polyploid cells, there is agreement that muscle cells and trophoblasts are products of cell fusion. In contrast, megakaryocytes are polyploid cells generated by genome duplication followed by aborted cytokinesis[5].
Polyploid liver cells are usually considered to be formed by endoreplication of the genome[6,7]. This conclusion was originally based on the seminal work of Mintz and colleagues, who pioneered the use of chimeric mice to study gene expression[8]. In the chimeric mouse, cells with two different genomes coexist in a single organism. If a cell fuses with another having a different genome, the resultant cell will contain markers of both. In their studies, Mintz and colleagues concluded that fusion occurred in muscle but not in the liver[8-10]. However, they had to rely mainly on the analysis of isoforms expressed by tissue-specific enzymes because single-cell markers were not yet available. Contrary to osteoclasts, trophoblasts and muscle cells, whose nuclei maintain their individuality, polyploid liver cells can be bi- or mono-nuclear.
Specific cell cycle genes, such as Cdk1, are involved in polyploidy formation and maintenance in liver[11]. Interestingly, after partial hepatectomy, quiescent hepatocytes can start proliferating again: deletion of cyclin-dependent kinase 1 (Cdk1), 2 (Cdk2), cyclin E1 or E2 individual genes does not limit liver regeneration, but concomitant ablation of Cdk2 and Cyclin E1 reduces liver regeneration, suggesting partial overlapping function of some cell cycle genes[11,12].
The study of cell fusion in the liver began while investigating the existence of lineage transdifferentiation. The possibility of cell transdifferentiation was raised at the end of the last century in a paper published in Science[13] in which the authors, after having transplanted neural stem cells transgenic for the beta-galactosidase (βgal) gene into wild-type mice, detected βgal+ cells in peripheral blood. Their interpretation was that neural cells had transdifferentiated into cells of the hematological lineage. This plasticity was surprising to many but, because the report was published just after the birth of Dolly the sheep, looked plausible[14]. We must emphasize here that that work had nothing to do with the reprogramming approach reported several years later by Yamanaka, who, in contrast, obtained reprogramming by forced expression of intracellular transcription factors[15]. The plasticity of neuronal stem cells was claimed by Bjornson et al[13] to occur by simple exposure to endogenous factors present in vivo.
Essentially, studies on plasticity were performed by transplanting cells from a mouse transgenic for an easily detectable marker gene (e.g., βgal) into a non-transgenic animal. The appearance of βgal-marked cells in an organ different from the tissue of origin was interpreted to be the result of transdifferentiation. The Science paper in which cells of the nervous system were suggested to acquire a hematological fate, was rapidly followed by other examples of transdifferentiation involving cells of several other lineages, including blood, brain, muscle, kidney and heart[16-25].
Due to the high potential for translation to the clinic, bone marrow cells (BMCs) escalated to center stage. BMCs are easily obtained, extensively investigated, routinely transplanted and well-characterized in humans. If simple transplantation protocols allowed the rescue of degenerative defects in organs such as brain, kidney or liver, we would have a sort of panacea in hand. Unfortunately, although many clinics-mostly in the United States-still advertise these kinds of treatments[26], transdifferentiation as originally proposed in the Science paper has not been confirmed by subsequent, more controlled studies[27-32]. Indeed, although a limited transdifferentiation capacity of some cells cannot be completely ruled out, more-recent studies have shown that transdifferentiation is often an experimental artifact. As stated above, transdifferentiation was claimed to occur if, after a given lineage (for example, hematopoietic cells) expressing a reporter gene was transplanted into a wild type mouse, cells of other lineages (for example, brain) were found to coexpress the reporter gene with accepted markers of their lineage. While βgal was initially used as a marker, most subsequent papers exploited fluorescent reporter genes that could be easily traced in vivo. It was assumed that all fluorescent cells found in a normal, non-transgenic, mouse had to be the progeny of transgenic donor cells: Hence, if they were found in other organs, they must have derived from original cells that had acquired a new fate by transdifferentiation.
In addition to trivial technical artifacts, cell fusion was raised to explain some of these results: in the experimental design discussed above, fusion between any cells of the host with transplanted donor cells could have provided the former with the reporter gene. It is difficult to discriminate between the two possibilities-transdifferentiation and cell fusion-with simple marker analysis.
As mentioned above, transdifferentiation of BMCs would be an attractive approach for regenerative medicine. Heart, brain and liver are heavily affected by degenerative genetic diseases that have a huge impact on human health; they would all greatly benefit from cell fusion-based therapies using exogenous cells, if that mechanism indeed occurs in vivo. Certainly, exogenous cells could provide defective endogenous ones with the missing genetic component while maintaining the differentiation status of the mature cell.
With regard to the liver, several reports in which BMCs were transplanted into recipient mice in the hope of inducing hepatocyte transdifferentiation showed that cells bearing donor-derived cellular markers could be found in host livers[18-21]. These “transdifferentiated” cells increased in number when the host livers were either injured (partial hepatectomy) or affected by a chronic degenerative genetic defect. However, the results were challenged by scientists who were unable to reproduce the transdifferentiation of hematological cells into non-hematological ones[33-36]. Cell fusion was shown to occur in vivo, so several reports investigated fusion events in a variety of other models. In the liver, the most spectacular experiments were performed by Grompe’s group on the classical model of fumarylacetoacetate hydrolase (Fah) deficiency[37]. Mice recessive for a Fah mutation are models for tyrosinemia type I, a severe genetic disease leading to liver failure in humans. Grompe and coworkers showed that bone marrow transplants in these mice led to the generation of liver cells bearing the donor marker, and demonstrated that this event was not due to transdifferentiation of hematological into hepatic lineage cells. Instead, these marker-carrier liver cells originated from cell fusion between donor bone marrow and resident hepatocytes, leading to polyploid cells that were not easily distinguishable from true hepatocytes in that the latter could also be polyploid. Due to the growth advantage shown by normal hepatocytes over diseased ones, the approach was so efficient that several mice were essentially cured. Results were confirmed by further studies[38-40], which also pointed to macrophages as the hematological cell responsible for fusion[41,42]. These results are in agreement with macrophages being physiologically prone to cell fusion[43]. In a review of 77 published studies on the generation of hepatocytes by hematopoietic cells transplanted in liver, the authors concluded that cell fusion was the mechanism involved[44]. Cell fusion is enhanced by the presence of liver injury or chronic disease, such as in the Fah model, since in a well-controlled study in which BMCs were injected into normal recipients, only 7 out of 470000 liver cells examined bore donor markers as a result of cell fusion[34]. In addition to BMCs, other types of cells, such as mesenchymal or amniotic stem cells and cells differentiated from pluripotent stem cells, can fuse with cells in injured livers, even when injected into a different species[45]. Human umbilical cord blood cells have also been reported to fuse with hepatocytes of immunocompromised mice[46], although no evidence of cell fusion was reported in other studies[47-49]. Moreover, cell fusion and transdifferentiation have been claimed to coexist[50].
The cell fusion-based explanation was found to hold also in other similar experimental settings[34,37,39,51,52] (reviewed in[27,53,54]). However, the possibility that at least in some cases, especially when an injury is applied to the recipient organ, bone marrow donor cells could be directed toward a different fate has not been completely ruled out, since several reports of well controlled differentiation have been published[55-64].
The discovery that cell fusion can cure a degenerative disease of the liver prompted Grompe’s group to investigate whether cell fusion occurs also in the disease-free state. The experimental plan to address this was as follows: they transplanted 1 × 105 wild-type (Fah+/+) hepatocytes into each of four Fah−/−/βgal+ recipients. After more than 80% of the liver was repopulated, 1 × 105 hepatocytes were serially transplanted into each of two Fah−/− recipients and the liver was again repopulated to a donor contribution of more than 80%. Then they analyzed 3 × 107 Fah+ hepatocytes, but were unable to find a single Fah+/βgal+ cell. They concluded that the frequency of cell fusion, if any, was very low[41].
It must be taken into considerations that the protocol involved damaged livers and injections of adult cells. However, although complex, the approach looks suitable to address the question of cell fusion in the disease-free liver. The only caveat is that, if the originally transplanted wild-type cells were mature hepatocytes (the age of the mice used was not specified), then it is possible that they represent polyploid cells that were already fully differentiated and functional and, therefore, less prone to fuse. This is because at this stage they have already achieved the benefits of being large cells with multiple genomes.
Apart from the original studies by Mintz and colleagues already cited, other studies investigating whether cell fusion occurs in the normal liver are lacking. Cell fusion has occasionally been reported to occur in hepatocytes or in hepatic tumor lines cultured in vitro[65-67]. Yet, this does not prove that the mechanism is physiologically relevant in vivo. For these reasons, Faggioli and coworkers devised and implemented a relatively simple but straightforward protocol based on chimeric mice, as originally proposed by Mintz’s group[8-10]. Embryo-derived mouse chimeras are mice born from embryonal cells carrying different genomes[68]. They can be created either by morula aggregation or by injection of embryonic stem cells (ESCs) into blastocysts, and they can be exploited for the study of cell fusion. If each of the two aggregated morulae contains a different reporter gene, then cells positive for both reporters will definitively be fused cells.
Faggioli et al[71] reasoned that by aggregating morulae from two different strains of transgenic mice expressing either green fluorescent protein (GFP)[69], or the βgal protein (Rosa 26 mouse[70]), the outcome would be animals that display two genetically distinct liver cell populations, each bearing a single marker (either GFP or βgal); any cell displaying both markers must be the result of cell fusion. With the appropriate controls, they identified three populations: GFP+/βgal−; GFP−/βgal+; and GFP+/βgal+. The percentage of double-positive cells in the chimeric samples was estimated to be about 25%.
The authors confirmed their results with two other independent strategies. Briefly, they performed PCR amplification on single hepatocytes with primers specific for each reporter gene, finding cells displaying both markers only in chimeric mice, in a percentage close to 10% of cells bearing at least one marker. In addition, the authors used fluorescent in situ hybridization (FISH) to investigate the sex chromosome content of hepatocytes in XY◄═►XX chimeric mice. They reasoned that, if fusion occurred between a female and a male cell, some binucleated cells containing Y chromosome(s) only in one of the two nuclei would be detected. Similarly, if mononucleated polyploid hepatocytes were analyzed, they should contain only X chromosomes in various numbers in the case they derived from the XX component of the chimeric mouse, or as many X as Y chromosomes if derived from an XY cell. In contrast, if the mononucleated polyploid hepatocytes were products of a cell fusion event between a female and a male cell, an unbalanced complement of X and Y chromosomes would be found. In the end, sex chromosome patterns were detected that were clearly indicative of cell fusion in binucleated as well as mononucleated hepatocytes in about 5%-10% of cells[71].
These results are at odds with those presented by Willenbring et al[41]. This discrepancy could be explained by the different approaches used, since that of Faggioli et al[71] mimics normal liver development, while the one used by Willenbring et al[41] involves the injection of exogenous hepatocytes into damaged liver and complex transplantation experiments. As mentioned before, this could ultimately lead to underestimation of fusion events in the latter study.
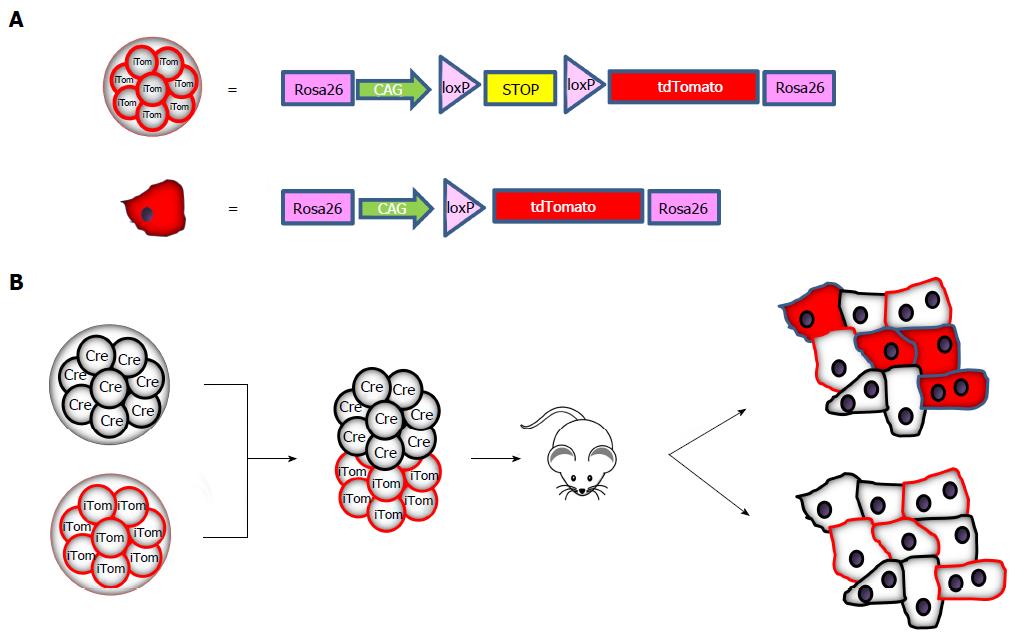
We are not aware of recent studies investigating cell fusion in normal liver, although replication of the chimera studies would not be too time consuming. Apparently, fusion is neither considered to occur frequently nor to be of physiological relevance. For this reason, while performing a study on the role of cell fusion in cancer[72], we addressed cell fusion with an even simpler approach based on the production of chimeric Cre: tdTomato mice. Morulae derived from mice transgenic for Cre recombinase under the control of a constitutive promoter were fused to morulae from mice transgenic for an inactive floxable tdTomato gene that is activated only if Cre recombinase is expressed in the same cell (Figure 1A). Cells from the two morulae will develop independently and no cell will be tdTomato-positive unless fusion with a Cre-containing cell has occurred (see the schematic representation in Figure 1B).
This approach-which has been widely used for lineage and transplantation studies-has the advantages of having an undetectable background if cell fusion does not occur, no interference between the two fluorescent reporter genes, and simple assessment in liver sections with well-validated tdTomato-specific antibodies. In addition, leakiness of the promoter, which sometimes occurs in Cre-based conditional mice, does not affect this model.
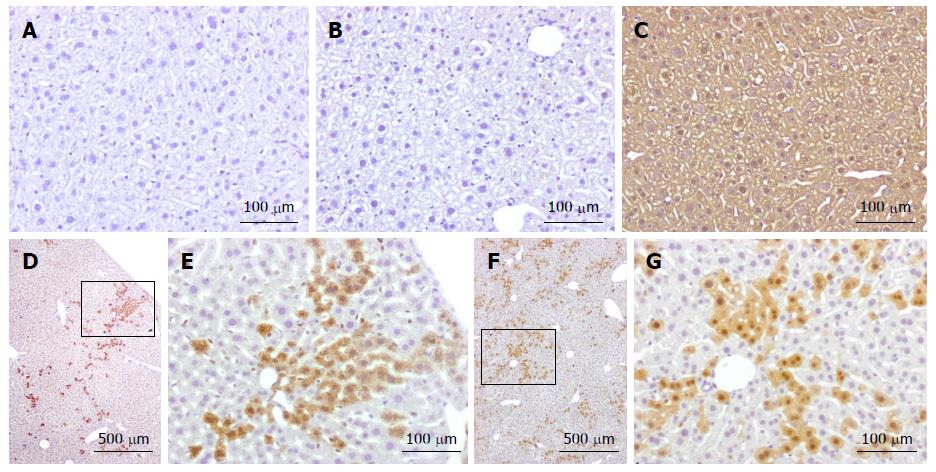
Analysis performed to date on two chimeric mice has clearly identified the presence of tdTomato-positive cells in the liver (Figure 2). Positivity was not detected in wild-type mice or in inactive tdTomato mice. As expected, the progeny of tdTomato-Cre mice crosses were positive in all tissues.
Fused cells are distributed all over the liver parenchyma, but are often found in clusters. This is in keeping with cell fusion occurring in cells maintaining their proliferative capacity, giving rise to a progeny that expands but remains in close proximity to their original location. This is in agreement with other studies showing hepatocytes originating form clonally derived clusters in postnatal liver[73,74].
However, the devil is in the detail and we are always at risk of artifacts[75]. In the chimeric experimental design, coexpression in the same cell of two reporter genes originally expressed independently by two distinct cells is commonly accepted as proof of a fusion event. This assumption was used in our original work on cell fusion[71]. However, the detection of fluorescence is prone to artifacts caused by endogenous background fluorescence, a phenomenon especially marked in liver; in the case of βgal, endogenous enzymatic activity can also lead to misinterpretation. In addition, it has become increasingly appreciated over the last ten years that transfer of materials-including RNA and proteins-between cells via extracellular vesicles is a frequent phenomenon[76-78]. Therefore, it cannot be excluded that in the Cre-tdTomato approach aforementioned, RNA encoding Cre recombinase or tdTomato could have been transferred from the Cre+ cell to the tdTomato one, and thus activating the reporter locus leading to expression of the reporter protein. Even the transfer of a few RNA or protein molecules over a very short period of time can activate the tdTomato gene, which then would become permanently expressed. However, the Cre-Lox and GFP systems have been widely used, in general giving consistent results for expression and expected specificity. Unfortunately, with the technologies available to date there is no way of discriminating fusion events from vesicle-mediated transfer in vivo while maintaining physiological conditions. In this regard, it is worth mentioning that several recent papers analyzing the fate of GFP+ cells transplanted into mouse retina have reported the detection of GFP+ cells that did not originate from the donor[79-81]. This suggests that GFP activity was leaked into the intracellular space and absorbed by endogenous cells or was transferred to them by extracellular vesicles–fusion can be excluded since retinal cells were normal in size and not polyploid. This is troubling if true, and some lineage or transplantation studies based on the detection of reporter genes should be carefully re-examined.
Techniques based on in situ hybridization with probes specific for sex chromosomes can be used to demonstrate cell fusion[71], since the presence of an XY nucleus as well as an XX one in a binucleated cell should definitively be due to cell fusion. This technique-which does not allow the analysis of live cells-has been used in studies on the ploidy of hepatocytes, with the caveat that the analysis might be complicated by the aneuploidy shown by some normal human and murine liver cells[82-85]. In any case, it will be difficult to investigate cell fusion in man: in theory, transplantation of male hepatocytes in female hosts performed for regenerative liver diseases could detect cell fusion, but this is a very rare occurrence and would require biopsies or post-mortem examination.
Cell fusion in the liver is still controversial. Thus, replication of previous studies with appropriate mouse chimeras is welcomed. Endoreplication and cell fusion are not mutually exclusive, as suggested by Gentric and Desdouets[86]. We strongly believe that fusion in the liver should be studied in order to confirm and explain this phenomenon. If established, this will open several new lines of investigation. For example, is cell fusion or endoreplication preferred in different contexts, or are they interchangeable? What is the fusion potential of hepatocytes with a DNA content higher than 4n? Are there hepatocytes with unbalanced or uneven-n chromosome numbers, and are there fusion products between one diploid and one tetraploid cell? Does cell fusion occur in species other than rodents, and particularly in man? Can fused cells participate in the ploidy reduction occurring after partial hepatectomy? Are HBV or HCV infections, which are themselves fusogenic viruses, able to change hepatocyte ploidy and binuclearity[87], or do other metabolic stresses[88] affect endoreplication or fusion? Does cell fusion play a role in HCV-mediated liver carcinogenesis[89]?
We thank Dr. Anna Villa, for useful discussion; Mr. Juan Pablo Casado for technical assistance; Dr. Elena Fontana and Dr. Stefano Mantero for help and assistance with immunohistochemical analysis.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: Italy
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): D
Grade E (Poor): 0
P- Reviewer: Gassler N, Tomizawa M S- Editor: Cui LJ L- Editor: A E- Editor: Li RF
| 1. | Brodsky WY, Uryvaeva IV. Cell polyploidy: its relation to tissue growth and function. Int Rev Cytol. 1977;50:275-332. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 287] [Cited by in RCA: 252] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 2. | Ogle BM, Cascalho M, Platt JL. Biological implications of cell fusion. Nat Rev Mol Cell Biol. 2005;6:567-575. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 225] [Cited by in RCA: 235] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 3. | Pandit SK, Westendorp B, de Bruin A. Physiological significance of polyploidization in mammalian cells. Trends Cell Biol. 2013;23:556-566. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 119] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 4. | Sobacchi C, Schulz A, Coxon FP, Villa A, Helfrich MH. Osteopetrosis: genetics, treatment and new insights into osteoclast function. Nat Rev Endocrinol. 2013;9:522-536. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 391] [Cited by in RCA: 398] [Article Influence: 33.2] [Reference Citation Analysis (0)] |
| 5. | Ravid K, Lu J, Zimmet JM, Jones MR. Roads to polyploidy: the megakaryocyte example. J Cell Physiol. 2002;190:7-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 185] [Cited by in RCA: 190] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 6. | Gupta S. Hepatic polyploidy and liver growth control. Semin Cancer Biol. 2000;10:161-171. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 194] [Cited by in RCA: 174] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 7. | Guidotti JE, Brégerie O, Robert A, Debey P, Brechot C, Desdouets C. Liver cell polyploidization: a pivotal role for binuclear hepatocytes. J Biol Chem. 2003;278:19095-19101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 227] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 8. | Mintz B. Gene control of mammalian differentiation. Annu Rev Genet. 1974;8:411-470. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 103] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 9. | Mintz B. Genetic Mosaicism in Adult Mice of Quadriparental Lineage. Science. 1965;148:1232-1233. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 110] [Cited by in RCA: 104] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 10. | Mintz B, Baker WW. Normal mammalian muscle differentiation and gene control of isocitrate dehydrogenase synthesis. Proc Natl Acad Sci USA. 1967;58:592-598. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 111] [Cited by in RCA: 102] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 11. | Diril MK, Ratnacaram CK, Padmakumar VC, Du T, Wasser M, Coppola V, Tessarollo L, Kaldis P. Cyclin-dependent kinase 1 (Cdk1) is essential for cell division and suppression of DNA re-replication but not for liver regeneration. Proc Natl Acad Sci USA. 2012;109:3826-3831. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 241] [Cited by in RCA: 311] [Article Influence: 23.9] [Reference Citation Analysis (0)] |
| 12. | Hu W, Nevzorova YA, Haas U, Moro N, Sicinski P, Geng Y, Barbacid M, Trautwein C, Liedtke C. Concurrent deletion of cyclin E1 and cyclin-dependent kinase 2 in hepatocytes inhibits DNA replication and liver regeneration in mice. Hepatology. 2014;59:651-660. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 41] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 13. | Bjornson CR, Rietze RL, Reynolds BA, Magli MC, Vescovi AL. Turning brain into blood: a hematopoietic fate adopted by adult neural stem cells in vivo. Science. 1999;283:534-537. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1022] [Cited by in RCA: 924] [Article Influence: 35.5] [Reference Citation Analysis (0)] |
| 14. | Theise ND, Wilmut I. Cell plasticity: flexible arrangement. Nature. 2003;425:21. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 34] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 15. | Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663-676. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17989] [Cited by in RCA: 18205] [Article Influence: 958.2] [Reference Citation Analysis (0)] |
| 16. | Gussoni E, Soneoka Y, Strickland CD, Buzney EA, Khan MK, Flint AF, Kunkel LM, Mulligan RC. Dystrophin expression in the mdx mouse restored by stem cell transplantation. Nature. 1999;401:390-394. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 904] [Cited by in RCA: 1004] [Article Influence: 38.6] [Reference Citation Analysis (0)] |
| 17. | Galli R, Borello U, Gritti A, Minasi MG, Bjornson C, Coletta M, Mora M, De Angelis MG, Fiocco R, Cossu G. Skeletal myogenic potential of human and mouse neural stem cells. Nat Neurosci. 2000;3:986-991. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 354] [Cited by in RCA: 318] [Article Influence: 12.7] [Reference Citation Analysis (0)] |
| 18. | Theise ND, Badve S, Saxena R, Henegariu O, Sell S, Crawford JM, Krause DS. Derivation of hepatocytes from bone marrow cells in mice after radiation-induced myeloablation. Hepatology. 2000;31:235-240. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 752] [Cited by in RCA: 696] [Article Influence: 27.8] [Reference Citation Analysis (0)] |
| 19. | Krause DS, Theise ND, Collector MI, Henegariu O, Hwang S, Gardner R, Neutzel S, Sharkis SJ. Multi-organ, multi-lineage engraftment by a single bone marrow-derived stem cell. Cell. 2001;105:369-377. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2032] [Cited by in RCA: 1889] [Article Influence: 78.7] [Reference Citation Analysis (0)] |
| 20. | Petersen BE, Bowen WC, Patrene KD, Mars WM, Sullivan AK, Murase N, Boggs SS, Greenberger JS, Goff JP. Bone marrow as a potential source of hepatic oval cells. Science. 1999;284:1168-1170. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1795] [Cited by in RCA: 1669] [Article Influence: 64.2] [Reference Citation Analysis (0)] |
| 21. | Lagasse E, Connors H, Al-Dhalimy M, Reitsma M, Dohse M, Osborne L, Wang X, Finegold M, Weissman IL, Grompe M. Purified hematopoietic stem cells can differentiate into hepatocytes in vivo. Nat Med. 2000;6:1229-1234. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1764] [Cited by in RCA: 1631] [Article Influence: 65.2] [Reference Citation Analysis (0)] |
| 22. | Orlic D, Kajstura J, Chimenti S, Jakoniuk I, Anderson SM, Li B, Pickel J, McKay R, Nadal-Ginard B, Bodine DM. Bone marrow cells regenerate infarcted myocardium. Nature. 2001;410:701-705. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3891] [Cited by in RCA: 3550] [Article Influence: 147.9] [Reference Citation Analysis (0)] |
| 23. | Alison MR, Poulsom R, Jeffery R, Dhillon AP, Quaglia A, Jacob J, Novelli M, Prentice G, Williamson J, Wright NA. Hepatocytes from non-hepatic adult stem cells. Nature. 2000;406:257. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 779] [Cited by in RCA: 749] [Article Influence: 30.0] [Reference Citation Analysis (0)] |
| 24. | Jackson KA, Majka SM, Wang H, Pocius J, Hartley CJ, Majesky MW, Entman ML, Michael LH, Hirschi KK, Goodell MA. Regeneration of ischemic cardiac muscle and vascular endothelium by adult stem cells. J Clin Invest. 2001;107:1395-1402. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1439] [Cited by in RCA: 1307] [Article Influence: 54.5] [Reference Citation Analysis (0)] |
| 25. | Mezey E, Chandross KJ, Harta G, Maki RA, McKercher SR. Turning blood into brain: cells bearing neuronal antigens generated in vivo from bone marrow. Science. 2000;290:1779-1782. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1353] [Cited by in RCA: 1250] [Article Influence: 50.0] [Reference Citation Analysis (0)] |
| 26. | Goff ZD, Kichura AB, Chibnall JT, Hauptman PJ. A Survey of Unregulated Direct-to-Consumer Treatment Centers Providing Stem Cells for Patients With Heart Failure. JAMA Intern Med. 2017;177:1387-1388. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 27. | Wagers AJ, Weissman IL. Plasticity of adult stem cells. Cell. 2004;116:639-648. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 856] [Cited by in RCA: 770] [Article Influence: 36.7] [Reference Citation Analysis (0)] |
| 28. | Ying QL, Nichols J, Evans EP, Smith AG. Changing potency by spontaneous fusion. Nature. 2002;416:545-548. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1123] [Cited by in RCA: 1036] [Article Influence: 45.0] [Reference Citation Analysis (0)] |
| 29. | Nygren JM, Jovinge S, Breitbach M, Säwén P, Röll W, Hescheler J, Taneera J, Fleischmann BK, Jacobsen SE. Bone marrow-derived hematopoietic cells generate cardiomyocytes at a low frequency through cell fusion, but not transdifferentiation. Nat Med. 2004;10:494-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 766] [Cited by in RCA: 721] [Article Influence: 34.3] [Reference Citation Analysis (0)] |
| 30. | Terada N, Hamazaki T, Oka M, Hoki M, Mastalerz DM, Nakano Y, Meyer EM, Morel L, Petersen BE, Scott EW. Bone marrow cells adopt the phenotype of other cells by spontaneous cell fusion. Nature. 2002;416:542-545. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1499] [Cited by in RCA: 1375] [Article Influence: 59.8] [Reference Citation Analysis (0)] |
| 31. | Duffield JS, Park KM, Hsiao LL, Kelley VR, Scadden DT, Ichimura T, Bonventre JV. Restoration of tubular epithelial cells during repair of the postischemic kidney occurs independently of bone marrow-derived stem cells. J Clin Invest. 2005;115:1743-1755. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 454] [Cited by in RCA: 428] [Article Influence: 21.4] [Reference Citation Analysis (0)] |
| 32. | Rizvi AZ, Swain JR, Davies PS, Bailey AS, Decker AD, Willenbring H, Grompe M, Fleming WH, Wong MH. Bone marrow-derived cells fuse with normal and transformed intestinal stem cells. Proc Natl Acad Sci USA. 2006;103:6321-6325. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 211] [Cited by in RCA: 197] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 33. | Castro RF, Jackson KA, Goodell MA, Robertson CS, Liu H, Shine HD. Failure of bone marrow cells to transdifferentiate into neural cells in vivo. Science. 2002;297:1299. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 323] [Cited by in RCA: 297] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 34. | Wagers AJ, Sherwood RI, Christensen JL, Weissman IL. Little evidence for developmental plasticity of adult hematopoietic stem cells. Science. 2002;297:2256-2259. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1136] [Cited by in RCA: 1027] [Article Influence: 44.7] [Reference Citation Analysis (0)] |
| 35. | Choi JB, Uchino H, Azuma K, Iwashita N, Tanaka Y, Mochizuki H, Migita M, Shimada T, Kawamori R, Watada H. Little evidence of transdifferentiation of bone marrow-derived cells into pancreatic beta cells. Diabetologia. 2003;46:1366-1374. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 177] [Cited by in RCA: 164] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 36. | Ono K, Yoshihara K, Suzuki H, Tanaka KF, Takii T, Onozaki K, Sawada M. Preservation of hematopoietic properties in transplanted bone marrow cells in the brain. J Neurosci Res. 2003;72:503-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 30] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 37. | Wang X, Willenbring H, Akkari Y, Torimaru Y, Foster M, Al-Dhalimy M, Lagasse E, Finegold M, Olson S, Grompe M. Cell fusion is the principal source of bone-marrow-derived hepatocytes. Nature. 2003;422:897-901. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1230] [Cited by in RCA: 1114] [Article Influence: 50.6] [Reference Citation Analysis (0)] |
| 38. | Quintana-Bustamante O, Alvarez-Barrientos A, Kofman AV, Fabregat I, Bueren JA, Theise ND, Segovia JC. Hematopoietic mobilization in mice increases the presence of bone marrow-derived hepatocytes via in vivo cell fusion. Hepatology. 2006;43:108-116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 82] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 39. | Alvarez-Dolado M, Pardal R, Garcia-Verdugo JM, Fike JR, Lee HO, Pfeffer K, Lois C, Morrison SJ, Alvarez-Buylla A. Fusion of bone-marrow-derived cells with Purkinje neurons, cardiomyocytes and hepatocytes. Nature. 2003;425:968-973. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1199] [Cited by in RCA: 1160] [Article Influence: 52.7] [Reference Citation Analysis (0)] |
| 40. | Zhou P, Hohm S, Olusanya Y, Hess DA, Nolta J. Human progenitor cells with high aldehyde dehydrogenase activity efficiently engraft into damaged liver in a novel model. Hepatology. 2009;49:1992-2000. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 41] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 41. | Willenbring H, Bailey AS, Foster M, Akkari Y, Dorrell C, Olson S, Finegold M, Fleming WH, Grompe M. Myelomonocytic cells are sufficient for therapeutic cell fusion in liver. Nat Med. 2004;10:744-748. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 303] [Cited by in RCA: 299] [Article Influence: 14.2] [Reference Citation Analysis (0)] |
| 42. | Camargo FD, Finegold M, Goodell MA. Hematopoietic myelomonocytic cells are the major source of hepatocyte fusion partners. J Clin Invest. 2004;113:1266-1270. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 165] [Cited by in RCA: 171] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 43. | Vignery A. Macrophage fusion: are somatic and cancer cells possible partners? Trends Cell Biol. 2005;15:188-193. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 83] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 44. | Thorgeirsson SS, Grisham JW. Hematopoietic cells as hepatocyte stem cells: a critical review of the evidence. Hepatology. 2006;43:2-8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 155] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 45. | Okamura K, Asahina K, Fujimori H, Ozeki R, Shimizu-Saito K, Tanaka Y, Teramoto K, Arii S, Takase K, Kataoka M. Generation of hybrid hepatocytes by cell fusion from monkey embryoid body cells in the injured mouse liver. Histochem Cell Biol. 2006;125:247-257. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 20] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 46. | Fujino H, Hiramatsu H, Tsuchiya A, Niwa A, Noma H, Shiota M, Umeda K, Yoshimoto M, Ito M, Heike T. Human cord blood CD34+ cells develop into hepatocytes in the livers of NOD/SCID/gamma(c)null mice through cell fusion. FASEB J. 2007;21:3499-3510. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 41] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 47. | Newsome PN, Johannessen I, Boyle S, Dalakas E, McAulay KA, Samuel K, Rae F, Forrester L, Turner ML, Hayes PC. Human cord blood-derived cells can differentiate into hepatocytes in the mouse liver with no evidence of cellular fusion. Gastroenterology. 2003;124:1891-1900. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 221] [Cited by in RCA: 200] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 48. | Chamberlain J, Yamagami T, Colletti E, Theise ND, Desai J, Frias A, Pixley J, Zanjani ED, Porada CD, Almeida-Porada G. Efficient generation of human hepatocytes by the intrahepatic delivery of clonal human mesenchymal stem cells in fetal sheep. Hepatology. 2007;46:1935-1945. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 99] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 49. | Zeng F, Chen MJ, Baldwin DA, Gong ZJ, Yan JB, Qian H, Wang J, Jiang X, Ren ZR, Sun D. Multiorgan engraftment and differentiation of human cord blood CD34+ Lin- cells in goats assessed by gene expression profiling. Proc Natl Acad Sci USA. 2006;103:7801-7806. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 38] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 50. | Hao NB, Li CZ, Lü MH, Tang B, Wang SM, Wu YY, Liang GP, Yang SM. SDF-1/CXCR4 Axis Promotes MSCs to Repair Liver Injury Partially through Trans-Differentiation and Fusion with Hepatocytes. Stem Cells Int. 2015;2015:960387. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 24] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 51. | Kanazawa Y, Verma IM. Little evidence of bone marrow-derived hepatocytes in the replacement of injured liver. Proc Natl Acad Sci USA. 2003;100 Suppl 1:11850-11853. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 121] [Cited by in RCA: 107] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 52. | Vassilopoulos G, Wang PR, Russell DW. Transplanted bone marrow regenerates liver by cell fusion. Nature. 2003;422:901-904. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1005] [Cited by in RCA: 898] [Article Influence: 40.8] [Reference Citation Analysis (0)] |
| 53. | Goodell MA. Stem-cell “plasticity”: befuddled by the muddle. Curr Opin Hematol. 2003;10:208-213. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 68] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 54. | Vassilopoulos G, Russell DW. Cell fusion: an alternative to stem cell plasticity and its therapeutic implications. Curr Opin Genet Dev. 2003;13:480-485. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 74] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 55. | Jang YY, Collector MI, Baylin SB, Diehl AM, Sharkis SJ. Hematopoietic stem cells convert into liver cells within days without fusion. Nat Cell Biol. 2004;6:532-539. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 430] [Cited by in RCA: 415] [Article Influence: 19.8] [Reference Citation Analysis (0)] |
| 56. | Sato Y, Araki H, Kato J, Nakamura K, Kawano Y, Kobune M, Sato T, Miyanishi K, Takayama T, Takahashi M. Human mesenchymal stem cells xenografted directly to rat liver are differentiated into human hepatocytes without fusion. Blood. 2005;106:756-763. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 449] [Cited by in RCA: 441] [Article Influence: 22.1] [Reference Citation Analysis (0)] |
| 57. | Theise ND, Krause DS, Sharkis S. Comment on “Little evidence for developmental plasticity of adult hematopoietic stem cells”. Science. 2003;299:1317; author reply 1317. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 61] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 58. | Harris RG, Herzog EL, Bruscia EM, Grove JE, Van Arnam JS, Krause DS. Lack of a fusion requirement for development of bone marrow-derived epithelia. Science. 2004;305:90-93. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 312] [Cited by in RCA: 302] [Article Influence: 14.4] [Reference Citation Analysis (0)] |
| 59. | Herzog EL, Chai L, Krause DS. Plasticity of marrow-derived stem cells. Blood. 2003;102:3483-3493. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 561] [Cited by in RCA: 532] [Article Influence: 24.2] [Reference Citation Analysis (0)] |
| 60. | Marongiu M, Serra MP, Contini A, Sini M, Strom SC, Laconi E, Marongiu F. Rat-derived amniotic epithelial cells differentiate into mature hepatocytes in vivo with no evidence of cell fusion. Stem Cells Dev. 2015;24:1429-1435. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 16] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 61. | Yadav N, Kanjirakkuzhiyil S, Kumar S, Jain M, Halder A, Saxena R, Mukhopadhyay A. The therapeutic effect of bone marrow-derived liver cells in the phenotypic correction of murine hemophilia A. Blood. 2009;114:4552-4561. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 33] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 62. | Amado LC, Saliaris AP, Schuleri KH, St John M, Xie JS, Cattaneo S, Durand DJ, Fitton T, Kuang JQ, Stewart G. Cardiac repair with intramyocardial injection of allogeneic mesenchymal stem cells after myocardial infarction. Proc Natl Acad Sci USA. 2005;102:11474-11479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 851] [Cited by in RCA: 777] [Article Influence: 38.9] [Reference Citation Analysis (0)] |
| 63. | Sekiya S, Suzuki A. Direct conversion of mouse fibroblasts to hepatocyte-like cells by defined factors. Nature. 2011;475:390-393. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 642] [Cited by in RCA: 646] [Article Influence: 46.1] [Reference Citation Analysis (0)] |
| 64. | Oh K, Shon SY, Seo MW, Lee HM, Oh JE, Choi EY, Lee DS, Park KS. Murine Sca1(+)Lin(-) bone marrow contains an endodermal precursor population that differentiates into hepatocytes. Exp Mol Med. 2015;47:e187. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 65. | Gómez-Lechón MJ, Barberá E, Gil R, Báguena J. Evolutive changes of ploidy and polynucleation in adult rat hepatocytes in culture. Cell Mol Biol Incl Cyto Enzymol. 1981;27:695-701. [PubMed] |
| 66. | Simic D, Euler C, Thurby C, Peden M, Tannehill-Gregg S, Bunch T, Sanderson T, Van Vleet T. Assessing cell fusion and cytokinesis failure as mechanisms of clone 9 hepatocyte multinucleation in vitro. Curr Protoc Toxicol. 2012;Chapter 14:Unit 14.9.1-Unit 14.917. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 67. | Yu Y, Duan J, Geng W, Li Q, Jiang L, Li Y, Yu Y, Sun Z. Aberrant cytokinesis and cell fusion result in multinucleation in HepG2 cells exposed to silica nanoparticles. Chem Res Toxicol. 2015;28:490-500. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 12] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 68. | Eckardt S, McLaughlin KJ, Willenbring H. Mouse chimeras as a system to investigate development, cell and tissue function, disease mechanisms and organ regeneration. Cell Cycle. 2011;10:2091-2099. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 26] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 69. | Okabe M, Ikawa M, Kominami K, Nakanishi T, Nishimune Y. ‘Green mice’ as a source of ubiquitous green cells. FEBS Lett. 1997;407:313-319. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2079] [Cited by in RCA: 2173] [Article Influence: 77.6] [Reference Citation Analysis (0)] |
| 70. | Friedrich G, Soriano P. Promoter traps in embryonic stem cells: a genetic screen to identify and mutate developmental genes in mice. Genes Dev. 1991;5:1513-1523. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1044] [Cited by in RCA: 1084] [Article Influence: 31.9] [Reference Citation Analysis (0)] |
| 71. | Faggioli F, Sacco MG, Susani L, Montagna C, Vezzoni P. Cell fusion is a physiological process in mouse liver. Hepatology. 2008;48:1655-1664. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 27] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 72. | Lizier M, Anselmo A, Mantero S, Ficara F, Paulis M, Vezzoni P, Lucchini F, Pacchiana G. Fusion between cancer cells and macrophages occurs in a murine model of spontaneous neu+ breast cancer without increasing its metastatic potential. Oncotarget. 2016;7:60793-60806. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 17] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 73. | Kennedy S, Rettinger S, Flye MW, Ponder KP. Experiments in transgenic mice show that hepatocytes are the source for postnatal liver growth and do not stream. Hepatology. 1995;22:160-168. [PubMed] |
| 74. | Shiojiri N, Imai H, Goto S, Ohta T, Ogawa K, Mori M. Mosaic pattern of ornithine transcarbamylase expression in spfash mouse liver. Am J Pathol. 1997;151:413-421. [PubMed] |
| 75. | Krause D, Cantley LG. Bone marrow plasticity revisited: protection or differentiation in the kidney tubule? J Clin Invest. 2005;115:1705-1708. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 64] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 76. | Lo Cicero A, Stahl PD, Raposo G. Extracellular vesicles shuffling intercellular messages: for good or for bad. Curr Opin Cell Biol. 2015;35:69-77. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 305] [Cited by in RCA: 381] [Article Influence: 38.1] [Reference Citation Analysis (0)] |
| 77. | Szabo G, Momen-Heravi F. Extracellular vesicles in liver disease and potential as biomarkers and therapeutic targets. Nat Rev Gastroenterol Hepatol. 2017;14:455-466. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 162] [Cited by in RCA: 219] [Article Influence: 27.4] [Reference Citation Analysis (0)] |
| 78. | Hirsova P, Ibrahim SH, Verma VK, Morton LA, Shah VH, LaRusso NF, Gores GJ, Malhi H. Extracellular vesicles in liver pathobiology: Small particles with big impact. Hepatology. 2016;64:2219-2233. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 185] [Cited by in RCA: 187] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 79. | Santos-Ferreira T, Llonch S, Borsch O, Postel K, Haas J, Ader M. Retinal transplantation of photoreceptors results in donor-host cytoplasmic exchange. Nat Commun. 2016;7:13028. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 212] [Cited by in RCA: 205] [Article Influence: 22.8] [Reference Citation Analysis (0)] |
| 80. | Pearson RA, Gonzalez-Cordero A, West EL, Ribeiro JR, Aghaizu N, Goh D, Sampson RD, Georgiadis A, Waldron PV, Duran Y. Donor and host photoreceptors engage in material transfer following transplantation of post-mitotic photoreceptor precursors. Nat Commun. 2016;7:13029. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 179] [Cited by in RCA: 224] [Article Influence: 24.9] [Reference Citation Analysis (0)] |
| 81. | Ortin-Martinez A, Tsai EL, Nickerson PE, Bergeret M, Lu Y, Smiley S, Comanita L, Wallace VA. A Reinterpretation of Cell Transplantation: GFP Transfer From Donor to Host Photoreceptors. Stem Cells. 2017;35:932-939. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 94] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 82. | Faggioli F, Vezzoni P, Montagna C. Single-cell analysis of ploidy and centrosomes underscores the peculiarity of normal hepatocytes. PLoS One. 2011;6:e26080. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 83. | Duncan AW, Taylor MH, Hickey RD, Hanlon Newell AE, Lenzi ML, Olson SB, Finegold MJ, Grompe M. The ploidy conveyor of mature hepatocytes as a source of genetic variation. Nature. 2010;467:707-710. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 429] [Cited by in RCA: 379] [Article Influence: 25.3] [Reference Citation Analysis (0)] |
| 84. | Duncan AW, Hanlon Newell AE, Smith L, Wilson EM, Olson SB, Thayer MJ, Strom SC, Grompe M. Frequent aneuploidy among normal human hepatocytes. Gastroenterology. 2012;142:25-28. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 159] [Cited by in RCA: 150] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 85. | Duncan AW. Aneuploidy, polyploidy and ploidy reversal in the liver. Semin Cell Dev Biol. 2013;24:347-356. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 135] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 86. | Gentric G, Desdouets C. Polyploidization in liver tissue. Am J Pathol. 2014;184:322-331. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 133] [Cited by in RCA: 145] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 87. | Toyoda H, Bregerie O, Vallet A, Nalpas B, Pivert G, Brechot C, Desdouets C. Changes to hepatocyte ploidy and binuclearity profiles during human chronic viral hepatitis. Gut. 2005;54:297-302. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 78] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 88. | Gentric G, Maillet V, Paradis V, Couton D, L’Hermitte A, Panasyuk G, Fromenty B, Celton-Morizur S, Desdouets C. Oxidative stress promotes pathologic polyploidization in nonalcoholic fatty liver disease. J Clin Invest. 2015;125:981-992. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 147] [Cited by in RCA: 187] [Article Influence: 18.7] [Reference Citation Analysis (0)] |
| 89. | Duelli D, Lazebnik Y. Cell-to-cell fusion as a link between viruses and cancer. Nat Rev Cancer. 2007;7:968-976. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 149] [Cited by in RCA: 150] [Article Influence: 8.3] [Reference Citation Analysis (0)] |