Published online Mar 26, 2012. doi: 10.4252/wjsc.v4.i3.17
Revised: February 28, 2012
Accepted: March 10, 2012
Published online: March 26, 2012
Cancer chemotherapy efficacy is frequently impaired by either intrinsic or acquired tumor resistance. A fundamental problem in cancer research is identifying the cell type that is capable of sustaining neoplastic growth and its origin from normal tissue cells. In recent years, the cancer stem cell (CSC) theory has changed the classical view of tumor growth and therefore the therapeutic perspective. Overcoming intrinsic and acquired resistance of cancer stem/progenitor cells to current clinical treatments represents a major challenge in treating and curing the most aggressive and metastatic cancers. On the other hand, the identification of CSCs in vivo and in vitro relies on specific surface markers that should allow the sorting cancer cells into phenotypically distinct subpopulations. In the present review, recent papers published on CSCs in solid tumors (breast, prostate, brain and melanoma) are discussed, highlighting critical points such as the choice of markers to sort CSCs and mouse models to demonstrate that CSCs are able to replicate the original tumor. A discussion of the possible role of aldehyde dehydrogenase and CXCR6 biomarkers as signaling molecules in CSCs and normal stem cells is also discussed. The author believes that efforts have to be made to investigate the functional and biological properties of putative CSCs in cancer. Developing diagnostic/prognostic tools to follow cancer development is also a challenge. In this connection it would be useful to develop a multidisciplinary approach combining mathematics, physics and biology which merges experimental approaches and theory. Biological models alone are probably unable to resolve the problem completely.
- Citation: La Porta CA. Thoughts about cancer stem cells in solid tumors. World J Stem Cells 2012; 4(3): 17-20
- URL: https://www.wjgnet.com/1948-0210/full/v4/i3/17.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v4.i3.17
Cancer stem cells (CSCs) are a subpopulation of tumor cells that possess the stem cell properties of self renewal and differentiation, generating the heterogeneous lineages of cancer cells that comprise the tumor. Thus, CSCs can only be defined experimentally by their ability to replicate the generation of a continuously growing tumor. Contrary to normal stem cells which are notable for the vigilance with which their proliferation is controlled and the care with which their genomic integrity is maintained, CSCs are frequently distinguished by their lack of control of such processes. Identifying differences between normal stem cells and CSCs is important for understanding how cancers progress and for translating advances in CSC biology into therapies that help patients.
There are many issues related to the identification of CSCs that must be considered. The first is the use of serial transplantation to validate a candidate CSC subpopulation, monitoring its capability to reproduce the heterogeneity of the primary tumor. Both xeno- and syngeneic transplantation may lose the intricate network of interactions with diverse support, such as those involving fibroblasts, endothelial cells, macrophages, mesenchymal stem cells, as well many of the cytokines and receptors involved in these interactions (for a more comprehensive discussion reed[1]). Another important problem is determining the best markers to identify CSCs. As discussed below this problem is still open. In light of recent findings reported for solid tumors like brain, prostate, breast and melanoma, I give my point of view on this issue. Moreover I discuss how to dissipate the shadows from the CSCs debate.
Tumor growth can be described either by the conventional model or by the CSC theory. According to the first model, cells are homogeneous and all are tumorigenic, while the CSC theory states that in the tumor there is a subpopulation sustaining tumor growth[1]. The first evidence of CSCs came from hematological tumors such as acute myeloid leukemia[2]. Later, CSCs were detected in breast, prostate, brain cancer and melanoma. In breast cancer the first evidence of a subpopulation with a specific cell-surface antigen profile (CD44+/CD24-) that can successfully establish itself as tumor xenograft was published in 2003[3]. More recently, aldehyde dehydrogenase (ALDH) was used as stem cell marker in 33 human breast cell lines[4]. ALDH is a detoxifying enzyme that oxidizes intracellular aldehydes and it is thought to play a role in the differentiation of stem cells via the metabolism of retinal to retinoic acid[5]. Interestingly, ALDH activity can be used to sort a subpopulation of cells that display stem cell properties from normal breast tissue and breast cancer[6]. ALDH activity, assessed by ALDEFLUOR assay, has been successfully used to isolate CSCs from multiple myeloma and acute leukemia as well as from brain tumors[7,8]. However in melanoma the ALDH phenotype was not associated with more aggressive subpopulations, arguing against ALDH as a “universal” marker[9].
Another interesting pathway that has been extensively studied is the Notch receptor signaling pathway (for a recent review see[10]). An important issue is the toxicity of potential treatments against these proteins. Even if the Notch pathway appears promising, it is also active in normal tissues, thus inhibition of Notch may have severe side effects. Therefore, as suggested by Harrison and colleagues, it seems important to study the complexity of the Notch pathway to target CSCs more successfully[10].
On the other hand, in a recent study, 275 patients with primary breast cancers of different subtypes and histological stages were analyzed for CD44+CD24- putative stem cell marker as well as for other markers (vimentin, osteonectin, connexin 43, ADLH, CK18, GATA3, MUC1). This study revealed a high degree of diversity in the expression of several of the selected markers in different tumor subtypes and histological stages[11]. I would like to point out that the latter findings could be explained by the fact that none of these markers are really specific for CSCs. In glioblastoma multiforme there is evidence for the existence of a more aggressive subpopulation of cancer cells and several markers have been identified[12-14]. Similarly, several candidate populations of prostate stem/progenitor cells have been reported including those expressing high levels of CD44, integrin α2β1, or CD133[15]. Interestingly, two recent independent studies in the mouse prostate have identified two different populations of stem cells (SCs). One, marked by CD117 (c-Kit), seems to be localized in the basal layer[16] and the other, called castration-resistant Nkx3.1-expressing cells, in the luminal layer[17]. Identification and characterization of normal prostate SCs is clearly relevant to understanding the origin of human prostate cancer, as suggested by recent reviews[14,18]. In fact, it is difficult to ascertain the potential overlap and the lineage relationships of the various candidate stem cells that have been identified[19]. This is due, in part, to the distinct methodologies and assays employed[19]. In melanoma seven papers were published from 2005-2008 showing that a CSCs subpopulation exists[20-25]. However, in 2008 one paper argued against the existence of CSCs, based on the following observations: a relatively large fraction of melanoma cells (up to about 25%) was shown to initiate tumors in severely immunocompromised NOD/SCID IL2Rγnull mice; the fraction of tumor-inducing cells depends upon assay conditions; several putative CSC markers appear to be reversibly expressed[26]. This paper, therefore, suggests that the detection of CSCs depends on how severely immunocompromised the mice are. The authors analyzed the expression of more than 50 surface markers on melanoma cells derived from several patients (A2B5, cKIT, CD44, CD49B, CD49D, CD49F, CD133, CD166) but focused on CD133 and CD166[26]. Using these markers they did not find any enrichment of tumor-initiating cells, but always found a high frequency of tumorigenic cells. However, in a recent paper it was shown that CD133 is highly expressed in melanoma cells and it is not a good marker for sorting CSCs[21]. Moreover, in 2010 Boiko and colleagues using the same immunocompromised mice could not confirm Quintana data[26]. Boiko et al[27] used CD271, a nerve growth factor receptor, as marker to identify CSCs.
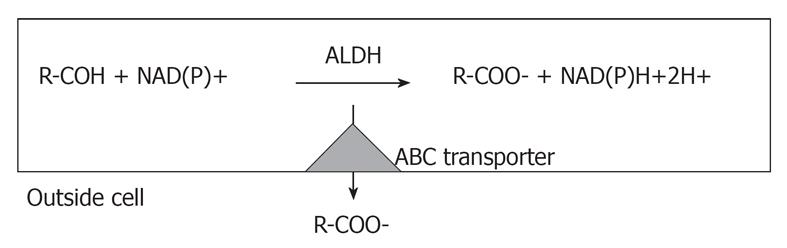
In my view it is quite clear that those involved in the CSC field must keep in mind that the only way to show that sorted putative CSCs are actually CSCs, is to replicate the heterogeneity of the tumor in syngeneic or immunodeficient mice. While it is possible that more severely immunocompromised mice are better than NOD/SCID mice, both are models, where the intricate interactions with the environment, such as mesenchymal cells, endothelial cells, and fibroblasts, are lost[1]. Moreover, another important issue is the choice markers for sorting CSCs. The most common strategy is to use markers that are expressed in normal stem cells. However, most of the time the functional role of these markers in stem cells is unknown and their role in stem cell biology is unclear. There are two exceptions reported in the literature. One is ALDH, which seems to play a specific functional role in stem cells[6-8] and the other is CXCR6[28]. ALDH actually determines cell survival through the ability to detoxify many potentially cytotoxic molecules and contributing to drug resistance (Figure 1) Several stem cell types, including some CSCs, reside in and have metabolic pathways attuned to a hypoxic environment and the increase in ALDH activity may reflect the demands of surviving in such niches[29]. A recent review summarizes the physiological role of ALDH[30].
The chemokine receptor CXCR6, known as Bonzo, STRL33 or TYMSTR is selectively expressed on the surface of CD4+ T cells, CD8+ T cells[31], NKT cells[32], natural killer cells[33] and plasma cells[34]. Moreover, CXCR6/CXCL16 is overexpressed in many cancer cells such as breast cancer[35]. CXCR6 is therefore expressed in stem cells when they grow asymmetrically and is down regulated when they switch to grow symmetrically[28]. Moreover, CXCR6 was recently shown to be expressed in a subpopulation of melanoma cells with higher self renewal capability[28]. The latest discoveries concerning melanocyte stem cells, such as their localization in the hair follicle, is discussed in a recent review[36].
An interesting recent paper shows that overexpressing Oct4 cells in human melanoma acquire a stem cell phenotype, increasing the expression of CSC markers[37]. This paper raises a number of new questions that will probably be studied in the next few years. In my opinion, one of the most important questions is whether this effect is caused by all the cells or by a subpopulation, such as CSCs.
I believe that much effort will be required to investigate the functional and biological properties of putative CSCs in cancer. Moreover it would be useful to confirm the expression of proposed markers in human biopsy samples. The development of diagnostic/prognostic tools to follow cancer development is also a challenge. It would be useful to develop a multidisciplinary approach combining mathematics, physics and biology, merging experimental approaches and theory. In fact, biological models alone are not probably able to resolve the problem completely.
Peer reviewer: Rasmus Freter, PhD, Ludwig Institute for Cancer Research, Nuffield Department of Clinical Medicine, University of Oxford, Old Road Campus Research Building (off Roosevelt Drive), Oxford OX3 7DQ, United Kingdom
S- Editor Wang JL L- Editor Hughes D E- Editor Zheng XM
| 1. | La Porta C. Cancer stem cells: lessons from melanoma. Stem Cell Rev. 2009;5:61-65. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 38] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 2. | Bonnet D, Dick JE. Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med. 1997;3:730-737. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4851] [Cited by in RCA: 4861] [Article Influence: 173.6] [Reference Citation Analysis (1)] |
| 3. | Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA. 2003;100:3983-3988. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7830] [Cited by in RCA: 7728] [Article Influence: 351.3] [Reference Citation Analysis (0)] |
| 4. | Charafe-Jauffret E, Ginestier C, Iovino F, Wicinski J, Cervera N, Finetti P, Hur MH, Diebel ME, Monville F, Dutcher J. Breast cancer cell lines contain functional cancer stem cells with metastatic capacity and a distinct molecular signature. Cancer Res. 2009;69:1302-1313. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 923] [Cited by in RCA: 914] [Article Influence: 57.1] [Reference Citation Analysis (0)] |
| 5. | Chute JP, Muramoto GG, Whitesides J, Colvin M, Safi R, Chao NJ, McDonnell DP. Inhibition of aldehyde dehydrogenase and retinoid signaling induces the expansion of human hematopoietic stem cells. Proc Natl Acad Sci USA. 2006;103:11707-11712. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 325] [Cited by in RCA: 348] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 6. | Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, Jacquemier J, Viens P, Kleer CG, Liu S. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 2007;1:555-567. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3173] [Cited by in RCA: 3064] [Article Influence: 170.2] [Reference Citation Analysis (0)] |
| 7. | Cheung AM, Wan TS, Leung JC, Chan LY, Huang H, Kwong YL, Liang R, Leung AY. Aldehyde dehydrogenase activity in leukemic blasts defines a subgroup of acute myeloid leukemia with adverse prognosis and superior NOD/SCID engrafting potential. Leukemia. 2007;21:1423-1430. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 171] [Cited by in RCA: 172] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 8. | Corti S, Locatelli F, Papadimitriou D, Donadoni C, Salani S, Del Bo R, Strazzer S, Bresolin N, Comi GP. Identification of a primitive brain-derived neural stem cell population based on aldehyde dehydrogenase activity. Stem Cells. 2006;24:975-985. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 210] [Cited by in RCA: 202] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 9. | Prasmickaite L, Engesaeter BØ, Skrbo N, Hellenes T, Kristian A, Oliver NK, Suo Z, Maelandsmo GM. Aldehyde dehydrogenase (ALDH) activity does not select for cells with enhanced aggressive properties in malignant melanoma. PLoS One. 2010;5:e10731. [PubMed] |
| 10. | Harrison H, Farnie G, Brennan KR, Clarke RB. Breast cancer stem cells: something out of notching. Cancer Res. 2010;70:8973-8976. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 60] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 11. | Park SY, Lee HE, Li H, Shipitsin M, Gelman R, Polyak K. Heterogeneity for stem cell-related markers according to tumor subtype and histologic stage in breast cancer. Clin Cancer Res. 2010;16:876-887. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 290] [Cited by in RCA: 315] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 12. | Gürsel DB, Shin BJ, Burkhardt JK, Kesavabhotla K, Schlaff CD, Boockvar JA. Glioblastoma Stem-Like Cells-Biology and Therapeutic Implications. Cancers (Basel). 2011;3:2655-2666. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 31] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 13. | Salmaggi A, Boiardi A, Gelati M, Russo A, Calatozzolo C, Ciusani E, Sciacca FL, Ottolina A, Parati EA, La Porta C. Glioblastoma-derived tumorospheres identify a population of tumor stem-like cells with angiogenic potential and enhanced multidrug resistance phenotype. Glia. 2006;54:850-860. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 196] [Cited by in RCA: 191] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 14. | Chen SY, Huang YC, Liu SP, Tsai FJ, Shyu WC, Lin SZ. An overview of concepts for cancer stem cells. Cell Transplant. 2011;20:113-120. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 32] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 15. | Tang DG, Patrawala L, Calhoun T, Bhatia B, Choy G, Schneider-Broussard R, Jeter C. Prostate cancer stem/progenitor cells: identification, characterization, and implications. Mol Carcinog. 2007;46:1-14. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 160] [Cited by in RCA: 164] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 16. | Leong KG, Wang BE, Johnson L, Gao WQ. Generation of a prostate from a single adult stem cell. Nature. 2008;456:804-808. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 321] [Cited by in RCA: 329] [Article Influence: 19.4] [Reference Citation Analysis (0)] |
| 17. | Wang X, Kruithof-de Julio M, Economides KD, Walker D, Yu H, Halili MV, Hu YP, Price SM, Abate-Shen C, Shen MM. A luminal epithelial stem cell that is a cell of origin for prostate cancer. Nature. 2009;461:495-500. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 522] [Cited by in RCA: 564] [Article Influence: 35.3] [Reference Citation Analysis (0)] |
| 18. | Li H, Tang DG. Prostate cancer stem cells and their potential roles in metastasis. J Surg Oncol. 2011;103:558-562. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 50] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 19. | Shen MM, Abate-Shen C. Molecular genetics of prostate cancer: new prospects for old challenges. Genes Dev. 2010;24:1967-2000. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 659] [Cited by in RCA: 717] [Article Influence: 47.8] [Reference Citation Analysis (0)] |
| 20. | Dou J, Pan M, Wen P, Li Y, Tang Q, Chu L, Zhao F, Jiang C, Hu W, Hu K. Isolation and identification of cancer stem-like cells from murine melanoma cell lines. Cell Mol Immunol. 2007;4:467-472. [PubMed] |
| 21. | Monzani E, Facchetti F, Galmozzi E, Corsini E, Benetti A, Cavazzin C, Gritti A, Piccinini A, Porro D, Santinami M. Melanoma contains CD133 and ABCG2 positive cells with enhanced tumourigenic potential. Eur J Cancer. 2007;43:935-946. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 404] [Cited by in RCA: 407] [Article Influence: 22.6] [Reference Citation Analysis (0)] |
| 22. | Schatton T, Murphy GF, Frank NY, Yamaura K, Waaga-Gasser AM, Gasser M, Zhan Q, Jordan S, Duncan LM, Weishaupt C. Identification of cells initiating human melanomas. Nature. 2008;451:345-349. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1140] [Cited by in RCA: 1034] [Article Influence: 60.8] [Reference Citation Analysis (0)] |
| 23. | Fang D, Nguyen TK, Leishear K, Finko R, Kulp AN, Hotz S, Van Belle PA, Xu X, Elder DE, Herlyn M. A tumorigenic subpopulation with stem cell properties in melanomas. Cancer Res. 2005;65:9328-9337. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 923] [Cited by in RCA: 934] [Article Influence: 46.7] [Reference Citation Analysis (0)] |
| 24. | Klein WM, Wu BP, Zhao S, Wu H, Klein-Szanto AJ, Tahan SR. Increased expression of stem cell markers in malignant melanoma. Mod Pathol. 2007;20:102-107. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 232] [Cited by in RCA: 230] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 25. | Hadnagy A, Gaboury L, Beaulieu R, Balicki D. SP analysis may be used to identify cancer stem cell populations. Exp Cell Res. 2006;312:3701-3710. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 225] [Cited by in RCA: 217] [Article Influence: 11.4] [Reference Citation Analysis (0)] |
| 26. | Quintana E, Shackleton M, Foster HR, Fullen DR, Sabel MS, Johnson TM, Morrison SJ. Phenotypic heterogeneity among tumorigenic melanoma cells from patients that is reversible and not hierarchically organized. Cancer Cell. 2010;18:510-523. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 497] [Cited by in RCA: 448] [Article Influence: 29.9] [Reference Citation Analysis (0)] |
| 27. | Boiko AD, Razorenova OV, van de Rijn M, Swetter SM, Johnson DL, Ly DP, Butler PD, Yang GP, Joshua B, Kaplan MJ. Human melanoma-initiating cells express neural crest nerve growth factor receptor CD271. Nature. 2010;466:133-137. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 28. | Taghizadeh R, Noh M, Huh YH, Ciusani E, Sigalotti L, Maio M, Arosio B, Nicotra MR, Natali P, Sherley JL. CXCR6, a newly defined biomarker of tissue-specific stem cell asymmetric self-renewal, identifies more aggressive human melanoma cancer stem cells. PLoS One. 2010;5:e15183. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 54] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 29. | Mohyeldin A, Garzón-Muvdi T, Quiñones-Hinojosa A. Oxygen in stem cell biology: a critical component of the stem cell niche. Cell Stem Cell. 2010;7:150-161. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1249] [Cited by in RCA: 1146] [Article Influence: 76.4] [Reference Citation Analysis (0)] |
| 30. | Balber AE. Concise review: aldehyde dehydrogenase bright stem and progenitor cell populations from normal tissues: characteristics, activities, and emerging uses in regenerative medicine. Stem Cells. 2011;29:570-575. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 92] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 31. | Kim CH, Kunkel EJ, Boisvert J, Johnston B, Campbell JJ, Genovese MC, Greenberg HB, Butcher EC. Bonzo/CXCR6 expression defines type 1-polarized T-cell subsets with extralymphoid tissue homing potential. J Clin Invest. 2001;107:595-601. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 261] [Cited by in RCA: 277] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 32. | Kim CH, Johnston B, Butcher EC. Trafficking machinery of NKT cells: shared and differential chemokine receptor expression among V alpha 24(+)V beta 11(+) NKT cell subsets with distinct cytokine-producing capacity. Blood. 2002;100:11-16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 221] [Cited by in RCA: 222] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 33. | Unutmaz D, Xiang W, Sunshine MJ, Campbell J, Butcher E, Littman DR. The primate lentiviral receptor Bonzo/STRL33 is coordinately regulated with CCR5 and its expression pattern is conserved between human and mouse. J Immunol. 2000;165:3284-3292. [PubMed] |
| 34. | Nakayama T, Hieshima K, Izawa D, Tatsumi Y, Kanamaru A, Yoshie O. Cutting edge: profile of chemokine receptor expression on human plasma cells accounts for their efficient recruitment to target tissues. J Immunol. 2003;170:1136-1140. [PubMed] |
| 35. | Matsumura S, Wang B, Kawashima N, Braunstein S, Badura M, Cameron TO, Babb JS, Schneider RJ, Formenti SC, Dustin ML. Radiation-induced CXCL16 release by breast cancer cells attracts effector T cells. J Immunol. 2008;181:3099-3107. [PubMed] |
| 36. | Nishimura EK. Melanocyte stem cells: a melanocyte reservoir in hair follicles for hair and skin pigmentation. Pigment Cell Melanoma Res. 2011;24:401-410. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 198] [Cited by in RCA: 199] [Article Influence: 14.2] [Reference Citation Analysis (0)] |
| 37. | Kumar SM, Liu S, Lu H, Zhang H, Zhang PJ, Gimotty PA, Guerra M, Guo W, Xu X. Acquired cancer stem cell phenotypes through Oct4-mediated dedifferentiation. Oncogene. 2012;Epub ahead of print. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 225] [Cited by in RCA: 256] [Article Influence: 19.7] [Reference Citation Analysis (0)] |