Published online Aug 26, 2020. doi: 10.4252/wjsc.v12.i8.761
Peer-review started: March 12, 2020
First decision: April 25, 2020
Revised: April 28, 2020
Accepted: June 14, 2020
Article in press: June 14, 2020
Published online: August 26, 2020
Processing time: 166 Days and 16.3 Hours
The developmental origins of health and diseases (DOHaD) is a concept stating that adverse intrauterine environments contribute to the health risks of offspring. Since the theory emerged more than 30 years ago, many epidemiological and animal studies have confirmed that in utero exposure to environmental insults, including hyperglycemia and chemicals, increased the risk of developing noncommunicable diseases (NCDs). These NCDs include metabolic syndrome, type 2 diabetes, and complications such as diabetic cardiomyopathy. Studying the effects of different environmental insults on early embryo development would aid in understanding the underlying mechanisms by which these insults promote NCD development. Embryonic stem cells (ESCs) have also been utilized by researchers to study the DOHaD. ESCs have pluripotent characteristics and can be differentiated into almost every cell lineage; therefore, they are excellent in vitro models for studying early developmental events. More importantly, human ESCs (hESCs) are the best alternative to human embryos for research because of ethical concerns. In this review, we will discuss different maternal conditions associated with DOHaD, focusing on the complications of maternal diabetes. Next, we will review the differentiation protocols developed to generate different cell lineages from hESCs. Additionally, we will review how hESCs are utilized as a model for research into the DOHaD. The effects of environmental insults on hESC differentiation and the possible involvement of epigenetic regulation will be discussed.
Core tip: The study of the mechanisms by which the intrauterine environment regulates offspring health is important. In this review, we will discuss the use of human embryonic stem cells as an in vitro model for understanding the developmental origins of diseases such as type 2 diabetes.
- Citation: Chen ACH, Lee KF, Yeung WSB, Lee YL. Human embryonic stem cells as an in vitro model for studying developmental origins of type 2 diabetes. World J Stem Cells 2020; 12(8): 761-775
- URL: https://www.wjgnet.com/1948-0210/full/v12/i8/761.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v12.i8.761
The increasing prevalence of diabetes is a serious global public health concern. According to the latest report from the International Diabetes Federation (Diabetes Atlas 2019), more than 400 million adults are thought to have diabetes[1]. More astonishingly, approximately half of them have not been diagnosed. The proportion of people with type 2 diabetes (T2D) has been increasing in most countries, including China. Indeed, the prevalence rate of diabetes in China has increased sharply in recent decades, from 1% in 1980[2] to 9.7% in 2008[3] and further to 10.9% in 2013[4]. Another report suggested that only one-fourth of the diabetes patients in China were diagnosed and treated, and among those treated, less than half of them had adequate glycemic control[5]. Diabetes is one of the biggest health issues in many countries. There is an urgent need for both national and international entities to tackle this problem.
T2D can be attributed to both genetic and environmental factors. For genetic factors, over 100 loci have been found to be associated with T2D. The susceptibility loci of T2D vary among ethnic groups. For instance, single nucleotide polymorphisms (SNPs) in KCNQ1 are associated with T2D in both East Asian and European people[6]. ARHGEF11 variants increase T2D risks in Pima Indian people[7]. On the other hand, SNPs of some loci (TSPAN8-LGR5, THADA, and ADAMTS9) are correlated with T2D susceptibility in Caucasian individuals but not in Chinese individuals[8]. Association studies suggested that genes such as TCF7L2 and KCNQ1 were related to pancreatic β-cell function and insulin secretion[9,10]. However, the causal relationship between genetic variants and disease phenotypes remains largely unclear. For environmental factors, in addition to personal lifestyle, maternal hyperglycemia also contributes to T2D risks. Approximately one-sixth of live births are affected by hyperglycemia during pregnancy[1]. Developmental epidemiological[11-13] and animal studies[14,15] indicated that in utero exposure to maternal diabetes increased the risks of developing T2D and insulin resistance in offspring. However, mechanistic studies on the inductive action of maternal hyperglycemic conditions on the development of T2D have been confined to animal models or pancreatic cell lines[16,17]. With the introduction of human embryonic stem cells (hESCs) in 1999[18], early human embryo development can be studied in vitro. We and others have used hESCs as models for studying the in utero effects of maternal diabetes on early embryo development, which was previously not possible in other pancreatic cell lines. In this review, we will discuss the long-term health consequences of fetal exposure to maternal diabetes and update the use of hESCs for studying the developmental origins of T2D.
The concept of developmental origins of health and diseases (DOHaD) was first proposed by Barker et al[19-21] more than 30 years ago; therefore, it is also known as “Barker’s hypothesis”. The epidemiological studies by Barker et al[21] revealed a high correlation between infant mortality rate and the incidence of ischemic heart disease later in life. Additionally, fetal malnutrition was associated with the risk of developing heart disease in adulthood[19]. Based on their observations, it was suggested that an adverse intrauterine environment would affect fetal programming. These changes permanently shaped the offspring’s organ function and metabolism, which would contribute to the adult onset of noncommunicable diseases (NCDs). Birthweight is the first and most common parameter predicting the health status of individuals at childhood and adulthood. Low birthweight is associated with many NCDs, including heart disease and T2D[19,22].
Early studies of DOHaD focused on maternal malnutrition. A famous example of this was the Dutch famine study. The offspring cohort who had prenatal exposure to Dutch famine (1944-1945) was traced. Studies have revealed a strong association between prenatal exposure to famine and glucose intolerance[23], obesity[24], heart disease[25], and even breast cancer[26]. A follow-up study demonstrated a trans-generational effect leading to neonatal adiposity in the F2 generation from the famine offspring cohort[27]. On the other hand, high birthweight, which has become more prevalent recently due to maternal obesity and overnutrition, is correlated with obesity[28], T1D[29], breast cancer, and pancreatic cancer[30].
In utero exposure to chemicals was found to be detrimental to long-term health in offspring. Animal studies have demonstrated that in utero exposure to endocrine disrupting chemicals (EDCs), such as bisphenol A (BPA), alters the development of the mammary gland, increasing the risk of breast cancer[31]. Prenatal exposure to BPA and diethylstilbestrol reduces the fertility of female mice, and the effect is transgenerational through the F3 generation[32]. In addition to affecting the reproductive system, in utero exposure to chemicals also contributes to an increase in T2D risk. Prenatal exposure to BPA induced leptin levels in female infants, and elevated leptin levels are correlated with insulin resistance[33]. A similar finding was observed in mice, where the administration of low-dose BPA (10 μg/kg) led to the development of chronic hyperinsulinemia and impaired glucose tolerance[34]. Another study traced the offspring born from individuals exposed to 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) due to explosion incidence in Italy in the 1970s. They found that in utero exposure to TCDD increased the risk for metabolic syndrome in male offspring[35]. To date, many maternal conditions have been identified to be associated with DOHaD, including maternal stress, hypertension, obesity, diabetes, smoking, infection, malnutrition, and even overnutrition[36].
One-sixth of live births worldwide are affected by hyperglycemia during pregnancy, among which approximately 80% are related to gestational diabetes (GDM)[1]. It is therefore apparent that maternal obesity, T2D, and GDM have long-term impacts on offspring health. GDM is defined as women without previously diagnosed diabetes who exhibit high blood glucose levels during pregnancy, especially during the third trimester. The prevalence of GDM ranges from 7%-10% of all pregnancies[37,38]. There are several risk factors contributing to GDM, which include obesity and personal or family history of T2D or GDM. Severely obese women have an 8-fold higher risk of developing GDM than pregnant women with a healthy weight[39]. It should be noted that GDM not only increased the risks of insulin resistance and T2D in offspring but also in mothers[40]. With the increasing number of pregnancies complicated by diabetes, it is important to understand the long-term impacts on offspring health through epidemiological studies. We will discuss the possible mechanisms in the context of epigenetics.
Epidemiological and animal studies: Maternal diabetes is often characterized by increased glucose transport from the placenta to the developing fetus; therefore, fetal macrosomia is the most obvious outcome that is studied[40,41]. Macrosomia is defined as birthweight of infants above 90th percentile of relative gestational age. More than 40% of infants born from diabetic pregnancy develop macrosomia[42], which is associated with increased neonatal morbidity rates. Macrosomic infants have an approximately 5-fold higher risk of glucose infusion and a 2-fold higher risk of neonatal jaundice than healthy infants[43]. A similar observation was found in an animal model in which rat offspring born from streptozotocin (STZ)-induced diabetic mothers developed macrosomia[44]. The mechanisms by which in utero hyperglycemia leads to macrosomia are not completely known. It has been suggested that GDM causes downregulation of adiponectin and upregulation of leptin. Macrosomic development has been linked to the modulation of cytokines[45].
The pathologies of macrosomia and maternal diabetes are associated with metabolic defects in infants. Macrosomic infants, and those born from diabetic pregnancies, have altered lipid metabolism. Compared with healthy babies, macrosomic infants have elevated plasma cholesterol and triglyceride levels[46]. In an STZ-induced diabetic rat model, the resulting offspring have increased lipid contents in serum and the liver[47]. These findings suggest alteration of lipid metabolism in the fetus, which contributes to risks of obesity and T2D in adulthood. Another important metabolic defect in the fetus is insulin secretion. Fetal development in the diabetic environment is accompanied by increased insulin secretion. Hyperinsulinemia has been found in cord blood in mothers with T2D or GDM[48]. Increased insulin secretion leads to overstimulation and exhaustion of fetal pancreatic β-cells. There is evidence of degranulation of fetal insulin-producing β-cells in the hyperglycemic intrauterine environment[49].
In addition to metabolic defects, abnormal organ development frequently occurs in offspring exposed to an intrauterine diabetic environment. At the beginning of gestation, impaired gene expression resulting from oxidative stress in the hyperglycemic environment can lead to embryopathy and an increased risk of cardiac, renal, and gastrointestinal malformations[50,51]. Early fetal exposure to a diabetic environment is correlated with higher risks of congenital abnormalities than what is observed when analyzing other exposure periods[52]. Reduced organ mass is another abnormality observed during development in hyperglycemic in utero environments. In rats born to diabetic mothers, there is a reduction in Igf2 expression levels in pancreatic β-cells and a decreased β-cell mass in the fetus[53]. In addition, in utero exposure to hyperglycemia is associated with a reduction in the number of nephrons and alterations of Igf expression in the fetal kidney[54,55].
Epigenetic mechanisms: It has long been suggested that epigenetic changes act as mediators between the early life exposure to environmental insults and the later onset of diseases. Epigenetic changes, such as DNA methylation and histone modifications, are actively involved in the course of embryo development. For example, there is global demethylation after fertilization, and DNA methylation is reestablished upon lineage specification[56]. Therefore, the early fetal development period is highly susceptible to epigenomic dysregulation with long-term implications for the health of the offspring[57].
The relationship between dysregulation of the DNA methylome and the risk of T2D has been extensively studied. In rats, offspring born from intrauterine growth retardation have increased risks of T2D in adulthood. In these offspring, Pdx1 transcription in pancreatic β cells is silenced due to DNA hypermethylation[58]. In humans, the PDX1 promoter is hypermethylated in the islets of T2D patients and is associated with lowered PDX1 expression in islet cells[59]. Pdx1 is important for early pancreatic specification in mouse embryos[60]. Peroxisome proliferator activated receptor gamma coactivator-1 alpha (PPARGC1A), which regulates ATP production, is also hypermethylated in human islet cells from T2D patients, and knockdown of PPARGC1A decreased insulin secretion[61].
Two independent studies utilized DNA methylation profiling on islet cells from T2D patients to determine the global dysregulation of the DNA methylome in diabetic pathology. Volkmar et al[62] and Dayeh et al[63] reported 254 and 853 differentially methylated genes, respectively, between T2D and normal samples, among which most were hypomethylated in T2D patients. Their studies also indicated that the differentially methylated genes were related to β-cell function, insulin secretion, and T2D pathogenesis. Another report also showed that GDM altered the placental DNA methylome of genes related to insulin signaling and endocrine disorders in both humans and rats[64].
Dysregulation of chromatin modifications is also closely associated with diabetes. High glucose conditions induce p300 acetyltransferase in primary human endothelial cells. The elevated p300 level increases histone acetylation, which results in induced gene expression of vasoactive factors and extracellular matrix proteins such as endothelin-1 (ET-1), vascular endothelial growth factor (VEGF), and fibronectin, leading to functional alterations in endothelial cells mimicking diabetic conditions[65]. Histone methylation of the H3K4 active mark and H3K9 repressive mark is responsible for gene expression regulation. In rats, offspring born under diabetic conditions exhibit dysregulated histone modification of the Pdx1 promoter; there is a progressive loss of H3K4 methylation but a gain of H3K9 methylation on the Pdx1 promoter, leading to silencing of this gene during development[58].
hESCs have pluripotent characteristics. They can spontaneously differentiate into three germ layers (mesoderm, endoderm, and ectoderm) during embryoid body (EB) formation[18]. Directed differentiation protocols of hESCs into specific cell types have been developed. These differentiated cells are excellent in vitro models for studying early human embryo development. With the introduction of induced pluripotent stem cells (iPSCs) by Yamanaka et al[66] in 2007, advancements were made to the regenerative medicine field, as patient iPSCs could be used to produce specific functional cell types to be used in replacement therapy. Indeed, iPSC technology-based regenerative therapy for diabetes has been vigorously studied in the past 10 years (reviewed in[67]).
Environmental insults such as maternal diabetes have been shown to affect neuronal, cardiac, and pancreatic development in offspring[68,69]. There is also evidence indicating the transgenerational epigenetic effects of environmental insults through germ cells. The specific cell lineages differentiated from pluripotent stem cells not only are of benefit for therapeutic purposes but also provide excellent in vitro models for studying DOHaD and the underlying mechanisms. In this section, we will update the differentiation protocols of those related cell lineages from pluripotent stem cells. The use of the models, in particular the pancreatic cell lineage, for studying the mechanism of DOHaD will also be discussed.
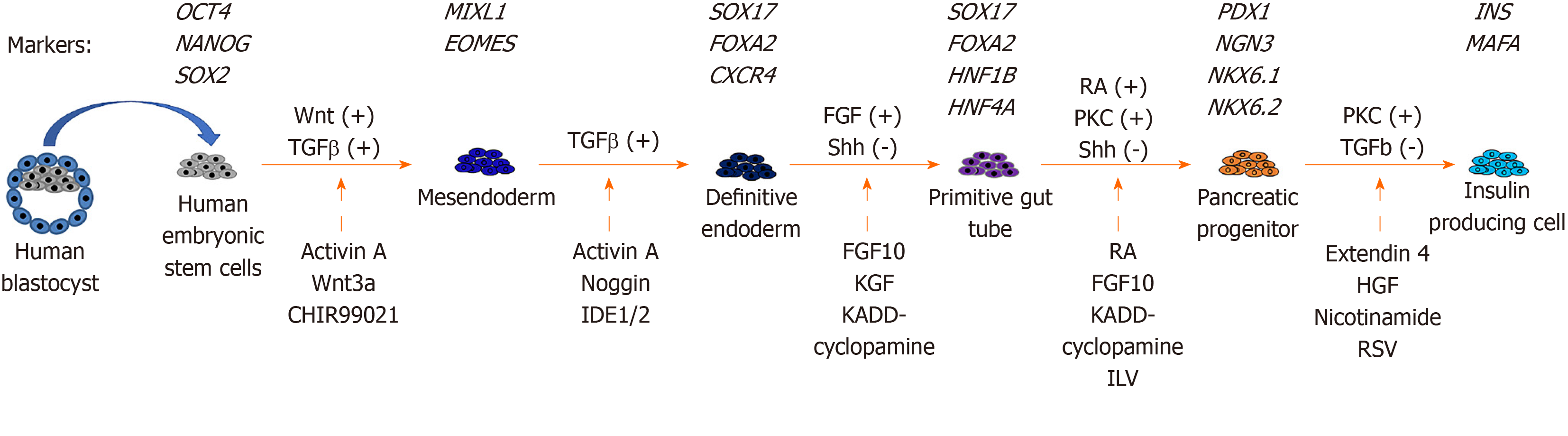
Pancreatic differentiation from hESCs: Since hESCs were first established from human embryos in 1998[18], there have been many studies on the production of glucose-responsive pancreatic β cells from hESCs for therapeutic purposes. The in vitro derivation of pancreatic β cells from hESCs involves stepwise inductions of cells representing mesendoderm (ME), definitive endoderm (DE), primitive gut tube (PGT), pancreatic progenitor (PP), and insulin-producing cell (IPC).
The stepwise differentiation of ESCs along the pancreatic lineage requires the activation of different signaling pathways (Figure 1). ME cells are bipotent in nature and are able to give rise to both the mesoderm and endoderm lineages during development[70]. In an early study of mouse embryonic development, ME cells were found to emerge from the anterior end of the primitive streak (APS)[71]. Brachyury (T)[72], goosecoid (GSC)[73], eomesodermin (EOMES)[74], and MIXL1[75] are valuable mesendoderm markers. Activation of the Wnt and TGFβ pathways is important for the derivation of ME cells from hESCs in vitro[72,76]. Therefore, the differentiation of ME includes the use of recombinant activin A (AA), which mimics the action of Nodal as the ligand for the TGFβ signaling pathway[77]. In addition, treatment with recombinant Wnt3a or a glycogen synthase kinase 3β inhibitor (CHIR-99021) can be used to activate the Wnt pathway[76,78].
DE can give rise to different endodermal cells, such as hepatocytes, epithelial cells of the respiratory tract, and the pancreas[79]. The efficient formation of DE cells is essential for subsequent differentiation into functional pancreatic cells[80]. The formation of the DE is marked by the expression of several transcription factors, including SRY (sex determining region Y)-box 17 (SOX17)[81], forkhead box A2 (FOXA2), and chemokine (C-X-C Motif) receptor 4 (CXCR4)[82]. Similar to ME formation, activation of the TGFβ pathway is important for the induction of DE markers. Recombinant AA and noggin, which acts as a bone morphogenic protein (BMP) antagonist, are used for DE induction[83]. Small molecules, including induction of definitive endoderm 1/2 (IDE1/IDE2), can mimic the effects of AA. Treatment of hESCs and mESCs with IDE1/2 induces DE formation, which is accompanied by an increase in SOX17 expression[84]. Using a commercially available DE differentiation kit (STEMdiff DE kit), we have shown that ME cells can be induced after 2 d of differentiation with T and MIXL1, and we have shown the efficient generation of DE cells with SOX17, FOXA2, and CXCR4 expression after 5 d of differentiation[85].
The formation of a PGT follows after DE induction[86]. Growth factors, including FGF10 and keratinocyte growth factor, enhance the efficiency of PGT formation[76,87]. Inhibiting the sonic hedgehog (Shh) signaling pathway by cyclopamine-KAAD treatment efficiently induces PGT specification[76]. The action is concordant with inhibition of cells entering an intestinal differentiation pathway following knockout of Shh signaling during mouse pancreatic bud formation[88]. Further specification into PP cells requires the continuous activation of FGF and inhibition of Shh signaling. The addition of retinoic acid together with FGF10 and cyclopamine-KAAD enhances PP formation[76]. In addition, activation of the protein kinase C (PKC) signaling pathway aids the formation of PP cells from the DE stage. A small molecule, indolactam V, which activates PKC signaling, was found to induce PP differentiation from hESCs[89]. The PP cells expressed several markers, including PDX1, SOX9, NKX6.1, and NKX6.2[76,90].
For the final step of producing IPCs from hESCs, there are two major approaches. One approach is to transplant PP cells into the mouse kidney capsule and allow them to mature in vivo[91]. The other approach is the in vitro differentiation of IPCs from PP cells. Treatment with extendin 4, hepatocyte growth factor, BMP4, and nicotinamide increased insulin secretion by PP cells in response to high glucose levels. However, the in vitro differentiation protocols are not efficient, and only approximately 10% of cells are insulin positive[76,92]. Pagliuca et al[93] reported the use of Alk5 receptor inhibitor II, PKC signaling activator, and thyroid hormone in the formation of β cells. Their results demonstrated that the β cells that formed were functional, as transplantation into diabetic mice successfully restored blood glucose to normal levels[93].
Three-dimensional organoid culture methods have recently been developed for the differentiation of hESCs. The organoids formed were reported to be structurally and functionally similar to their native tissue counterparts. For instance, pancreatic organoids were formed by aggregating hESC-derived PP cells in a novel hydrogel system named Amikagel. The resulting cells in the organoids closely mimicked pancreatic islet cells[94].
Pancreatic differentiation from hESCs as a model for studying DOHaD: Diabetic pregnancy is known to increase the risks of insulin resistance and T2D in offspring in adulthood. Epigenetic dysregulation is associated with disease phenotypes. For instance, mice born from diabetic pregnancies exhibit hypermethylation of pdx1 promoter DNA[58]. Diabetic pregnancies induce global changes in the DNA methylome related to insulin signaling in the human placenta[64]. However, studies on the effects of environmental insults on human fetal pancreas development are very limited. We used hESCs as an in vitro model to study the developmental origins of diabetes. Early pancreatic differentiation is mainly modulated by histone methylation[95,96]. We confirmed that the promoters of DE markers (SOX17, FOXA2, and CXCR4) were marked bivalently by both the activating mark H3K4me3 and the repressive mark H3K27me3 at the pluripotent stage. Upon differentiation into DE, the repressive mark H3K27me3 was removed, leading to active expression of DE markers. More importantly, our study was the first to discover that a hyperglycemic environment disrupted histone methylation patterns, resulting in retention of repressive H3K27me3 marks at DE promoters and a significant reduction in their expression compared to the control. The inhibition of DE specification is also observed in mice upon in utero exposure to hyperglycemia[85] (Table 1). Recently, studies have demonstrated active DNA methylation and hydroxymethylation during different stages of in vitro pancreatic differentiation from hESCs. DNA hydroxymethylation has been associated with chromatin accessibility, therefore allowing the binding of transcription factors for efficient pancreatic differentiation[97]. The above studies suggest the important roles of DNA methylation and hydroxymethylation in pancreatic development.
| Environmental insult(s) | Type of pluripotent stem cells | Cell lineage | Effects on differentiation | Ref. |
| Hyperglycemia (25-50 mmol/L) | hESC | Pancreatic | (1) Inhibited differentiation into DE; and (2) retained repressive H3K27me3 mark on DE marker promoters | [85] |
| Hyperglycemia (25-50 mmol/L) | mESC | Pancreatic | Inhibited differentiation into DE | [85] |
| TCDD (10 pmol/L) | hESC | Pancreatic | Dysregulated DNA methylome of genes related to diabetes | [99] |
| TCDD (10-100 pmol/L) | hESC | Pancreatic | (1) Dysregulated DNA methylome of genes related to insulin signaling and diabetes; (2) inhibited differentiation into pancreatic progenitor; and (3) promoted DNA hypermethylation of PRKAG1 | [100] |
| Hyperglycemia (10 mmol/L), endothelin-1 (ET-1) (10 nmol/L), and cortisol (1 μmol/L) | hiPSC | Cardiac | (1) Inhibited cardiomyocyte differentiation; and (2) elevated oxidative stress in cardiomyocytes formed | [114] |
| Hyperglycemia (25 mmol/L) | mESC | Cardiac | Inhibited mesoderm and subsequent cardiomyocyte differentiation | [117] |
| Hyperglycemia (25 mmol/L) | mESC | Cardiac | Enhanced cardiomyocyte differentiation | [118] |
| TCDD (1 nmol/L) | mESC | Cardiac | Inhibited cardiomyocyte differentiation | [136] |
| TCDD (10-100 pmol/L) | hESC | Cardiac | Dysregulated DNA methylome of genes related to cardiomyopathy | [100] |
| BPA (1-8 μg/mL); PFOS (5-40 μg/mL); PFOA (10-80 μg/mL) | mESC | Cardiac | Inhibited cardiomyocyte differentiation | [137] |
| Hyperglycemia (25 mmol/L) | mESC | Neural | Inhibited neural differentiation | [125] |
In addition to maternal diabetes, the effects of in utero exposure to chemicals such as EDCs have also been extensively studied in the later development of offspring. For instance, an animal study showed that in utero exposure to BPA increased glucagon secretion in fetal islets by affecting the α-to-β cell ratio[98]. Recently, we conducted transcriptomic and methylomic analyses on hESCs upon low-dose (10 pM) TCDD treatment. Our results revealed that the expression and DNA methylation status of a number of genes were dysregulated upon TCDD treatments. Among them, some of the genes, such as adenosine A1 receptor (ADORA1), ADORA2A, inhibin beta A subunit, and hemopexin, were associated with the pathogenesis of diabetes[99]. Low-dose TCDD (10-100 pM) treatment of hESCs also induced hypermethylation of a number of genes that are related to insulin signaling and T2D. Among them, PRKAG1 remained hypermethylated even upon PP differentiation. PRKAG1 knockdown in the pancreatic cell line INS-1E resulted in elevated levels of secreted insulin[100] (Table 1). In addition, our findings suggested that the dysregulated DNA methylation patterns induced by early chemical exposure might be maintained during early embryonic development. These changes might lead to pathology, such as insulin resistance and diabetes, in offspring.
Cardiac differentiation from hESCs: The human heart is often considered a nonregenerative organ due to the limited proliferative ability of adult cardiomyocytes (CMs). Following the first reports of hESCs[18] and iPSCs[66], several approaches have been developed to differentiate these cells into functional CMs. This section will discuss the transcription factors and cell signaling pathways essential for CM development. We will also introduce various CM differentiation protocols that have been developed.
The heart is one of the organs that develops early in embryos. In the human embryo, the primordial heart begins to develop at 20 d after fertilization. Cardiac cell lineage emerges from the mesoderm. The induction of mesoderm formation is mainly controlled by three cell signaling pathways: the FGF, Wnt and TGFβ pathways. Mesoderm development can be marked with the expression of markers such as T-box transcription factor brachyury (T) and EOMES[101]. The mesodermal cell population expressing mesoderm posterior 1 (MESP1) further develop into cardiac progenitor cells via inhibition of the Wnt/β-catenin pathway[102,103]. The subsequent specification into CMs requires the action of signaling pathways such as retinoic acid (RA) and FGF pathways, where MESP1 is the upstream regulator of cardiac-specific transcription factors, such as GATA binding protein 4 (GATA4) and NK2 homeobox 5 (NKX2.5)[104]. The in vitro differentiation of hESCs into CMs therefore involves stepwise manipulation of cell signaling pathways.
The successful derivation of CMs from hESCs was first reported through spontaneous differentiation of EBs. However, the efficiency was low, with 8.1% of the area exhibiting spontaneous beating and only 29.4% of cells expressing cardiac troponin I (cTnI) after 20 d of differentiation[105]. Several modified protocols have been subsequently reported. These reports also adopted the EB approach, but instead of spontaneous differentiation, and they mimicked in vivo signaling for directed differentiation. For instance, BMP4, Activin A, and bFGF were supplemented in culture for mesoderm induction. VEGF and DKK1 recombinant proteins were then added as Wnt/β-catenin inhibitors. The cells were further treated with VEGF, bFGF, and DKK1 to induce expansion and differentiation into CMs. With improved protocols, the efficiency of CM differentiation was increased (> 80% cTnI+ve cells), and it was achieved in a shorter period of time (8-10 d)[106,107]. Subsequently, different EB culturing tools were developed for scaling up CM production for therapeutic purposes. For instance, microwells allow the production of a large number of uniformly sized EBs[108]. On the other hand, researchers developed microcarriers that promoted the expansion of differentiating hESCs in spinner flasks and bioreactors for large-scale CM production[109,110].
Cardiac differentiation from hESCs as a model for studying DOHaD: Diabetic cardiomyopathy (DCM) is a complication of T2D. Maternal diabetes also increases the risk of cardiomyopathy in infants[111,112]. An early animal study using streptozotocin-induced diabetic mice demonstrated a high rate of apoptosis in cardiomyocytes. An in vitro study using adult CMs also exhibited reduced myofibrillar formation under high glucose treatment[113]. However, the underlying mechanisms of the developmental origins of cardiomyopathy remain largely unknown. hESC-derived CMs can therefore serve as an excellent in vitro model for recapitulating major events during embryonic heart development.
Diabetic conditions, including high glucose (10 mmol/L), ET-1 (10 nmol/L), and cortisol (1 μmol/L) treatments, induce hypertrophic stress with elevated expression of hypertrophic markers (NPPA, NPPB, ACTA1, and MYH7) during CM differentiation from hiPSCs. The treated CMs exhibit cardiomyopathy phenotypes such as disorganized sarcomere structures, accumulation of lipid contents, and oxidative stress[114] (Table 1). Defects in embryonic CM formation might lead to an increased risk of DCM in adulthood[114]. hESC-CMs are not extensively used as a DOHaD model for cardiomyopathy. This could be attributed to the fact that hESC-CMs do not represent fully mature CMs. The contractile function and cardiac marker expression of hESC-CMs are not comparable to those of fetal or adult CMs[115,116]. Notwithstanding, similar studies have been performed in a mESC model to understand the effects of in utero hyperglycemia on cardiac development. It was demonstrated that high glucose conditions (25 mmol/L) impaired cardiac differentiation from mESCs compared with what was observed in cells treated with physiological levels of glucose (5 mmol/L). There was a significant reduction in contracting CMs under high glucose levels. In addition, a significant reduction in the expression of mesoderm markers (T and Mixl1) and cardiac markers (Gata4 and Nkx2.5)[117] was observed upon hyperglycemia treatment. However, opposite results from another study showed that CM formation from EBs was more efficient under high glucose treatment[118] (Table 1). The effects of hyperglycemia and other environmental insults on human CM differentiation require further investigation.
A recent epigenomic study on human CMs revealed that prenatal and postnatal heart development were regulated by DNA methylation and histone modifications. More importantly, active histone marks (H3K27ac, H3K4me3, H3K9ac, and H3K36me3) were found in the promoters of pathology-related genes such as connective tissue growth factor (CTGF) and natriuretic peptides A and B (NPPA and NPPB) in diseased CMs[119]. Another recent study demonstrated distinct DNA methylation patterns in atrial and ventricular subtypes of hiPSC-derived CMs[120]. These findings reveal that epigenetic regulation not only occurs during prenatal heart development but also is responsible for cardiomyopathy. The study of DOHaD in relation to cardiomyopathy in an epigenetic context warrants further investigation.
Neural lineage: There is a strong clinical association between maternal diabetes and neural tube defects (NTDs). Maternal diabetes increases the risk of central nervous system malformation in fetuses by 10-15-fold over that of nondiabetic mothers[121,122]. Similarly, mouse offspring born from diabetic mothers have an approximately 10% chance of developing NTDs. A high level of oxidative stress leads to neural cell apoptosis in the affected offspring[123]. Maternal hyperglycemia also results in the activation of apoptosis signal-regulating kinase 1 (Ask1) in the developing neural tubes of mouse embryos. The activation of Ask1 is related to an increase in caspase 8 protein levels and apoptosis[124].
Studies in animal models provide information on the effects of the in utero environment on early neural development. However, further studies remain challenging because of the limited number of cells in fetal neural tissues. Nevertheless, high glucose treatment (25 mmol/L) in vitro impedes neural differentiation, resulting in the downregulation of neuronal markers (Sox1, Nestin, and Pax6)[125] (Table 1). Folate deficiency was shown to induce inhibition of the DNA methylation cycle, leading to NTDs in animals[126]. Knockout of histone modifiers such as Sirt1 and histone deacetylase 4 also causes NTDs in developing mouse embryos[127,128]. It should be noted that the effects of environmental insults on human neural development may be different from those observed in mice. Further mechanistic studies using hESCs as cell models can improve our understanding in the context of DOHaD. Indeed, treatment with noggin, which inhibits the BMP pathway, successfully enabled derivation of neuronal cells from hESCs. The neurospheres formed could further differentiate into mature neurons and glia[129], providing a good cellular research model.
Germ cell lineage: Growing evidence suggests that the negative impacts of adverse intrauterine environments on offspring might be transgenerational, meaning that the disease phenotypes will be expressed in the F2 generation. Such transgenerational effects are evidenced in animal models. For example, vinclozolin (VCZ; 3-(3,5-dichlorophenyl)-5-methyl-5-vinyl-oxazolidine-2,4-dione), one of the EDCs widely used as a fungicide, dysregulates the epigenome of primordial germ cells (PGCs) in mice from F1 to F3; the microRNA pattern in F1-F3 PGCs is disrupted following F0 animal exposure to VCZ[130]. The downregulation of miR-23b and miR-21 in the treated mice disrupts the let7 pathway, leading to increased apoptosis of embryonic PGCs. Another EDC, TCDD, alters transcriptomes in the gonads of F1 and F2 zebrafish[131]. The genes with altered expression were related to lipid and glucose metabolism, oxidative stress, and sperm cell development.
In addition to EDC exposure, the transgenerational effects of maternal hyperglycemia have been extensively studied in animal models. Ding et al[132] reported that the mating of F1 male mice from diabetic pregnancies with normal female mice resulted in F2 mice with increased birth weight and impaired glucose tolerance. They associated the above observations with DNA hypermethylation of imprinted genes Igf2 and H19 in the pancreatic islets of F1 and F2 mice[132]. A recent report revealed that maternal diabetes dysregulated the DNA methylome of embryonic F1 PGCs. The differentially methylated genes were related to obesity, insulin resistance, and T2D. More importantly, the same pattern was also observed in F2 somatic cells[133]. These studies demonstrated that environmental insults, such as chemicals or hyperglycemia, could be transmitted transgenerationally by changing the epigenomes of germ cells.
In vitro germ cell differentiation from ESCs has only recently been reported. Haploid germ cells can be generated by coculturing mESC-derived PGC-like cells with neonatal testicular somatic cells. In vitro-derived haploid spermatids are able to generate offspring when injected into oocytes[134]. In human culture systems, PGCs can be successfully derived from hESCs. The derivation protocol adopted a stepwise approach recapitulating in vivo developmental events, where Wnt and BMP pathways were activated to drive the formation of premesodermal cells. The specifications of PGCs were then induced by treatment with growth factors such as BMP2, stem cell factor, and epidermal growth factor[135]. Advances in germ cell differentiation using a human cell model also provide an opportunity for the study of the transgenerational effects of DOHaD.
Much evidence supporting the idea of DOHaD has been obtained from animal models and observational studies of human. The mechanisms behind the long-term health consequences of fetal exposure to adverse maternal conditions are largely unknown in humans. Accumulating data from both hESCs and mESCs suggested that early cell lineage differentiation might be one of the vulnerable embryonic windows through which early exposure to adverse maternal conditions could exert its diabetogenic effects. While protocols for the differentiation of different cell types from ESCs still require further improvement to better mimic physiological development, it is expected that the information obtained from these cell models will provide valuable mechanistic insight into the mechanisms underlying the DOHaD.
Manuscript source: Invited manuscript
Specialty type: Cell and tissue engineering
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): A
Grade B (Very good): 0
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: El-Sayyad HIH S-Editor: Wang JL L-Editor: Wang TQ P-Editor: Xing YX
| 1. | International Diabetes Federation. IDF Diabetes Atlas, 2019. 9th ed. Brussels: International Diabetes Federation, 2019. |
| 2. | [A mass survey of diabetes mellitus in a population of 300,000 in 14 provinces and municipalities in China (author's transl)]. Zhonghua Nei Ke Za Zhi. 1981;20:678-683. [PubMed] |
| 3. | Yang SH, Dou KF, Song WJ. Prevalence of diabetes among men and women in China. N Engl J Med. 2010;362:2425-6; author reply 2426. [PubMed] |
| 4. | Wang L, Gao P, Zhang M, Huang Z, Zhang D, Deng Q, Li Y, Zhao Z, Qin X, Jin D, Zhou M, Tang X, Hu Y, Wang L. Prevalence and Ethnic Pattern of Diabetes and Prediabetes in China in 2013. JAMA. 2017;317:2515-2523. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1099] [Cited by in RCA: 1357] [Article Influence: 169.6] [Reference Citation Analysis (0)] |
| 5. | Xu Y, Wang L, He J, Bi Y, Li M, Wang T, Wang L, Jiang Y, Dai M, Lu J, Xu M, Li Y, Hu N, Li J, Mi S, Chen CS, Li G, Mu Y, Zhao J, Kong L, Chen J, Lai S, Wang W, Zhao W, Ning G; 2010 China Noncommunicable Disease Surveillance Group. Prevalence and control of diabetes in Chinese adults. JAMA. 2013;310:948-959. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1961] [Cited by in RCA: 2169] [Article Influence: 180.8] [Reference Citation Analysis (0)] |
| 6. | Unoki H, Takahashi A, Kawaguchi T, Hara K, Horikoshi M, Andersen G, Ng DP, Holmkvist J, Borch-Johnsen K, Jørgensen T, Sandbaek A, Lauritzen T, Hansen T, Nurbaya S, Tsunoda T, Kubo M, Babazono T, Hirose H, Hayashi M, Iwamoto Y, Kashiwagi A, Kaku K, Kawamori R, Tai ES, Pedersen O, Kamatani N, Kadowaki T, Kikkawa R, Nakamura Y, Maeda S. SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat Genet. 2008;40:1098-1102. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 531] [Cited by in RCA: 531] [Article Influence: 31.2] [Reference Citation Analysis (0)] |
| 7. | Ma L, Hanson RL, Que LN, Cali AM, Fu M, Mack JL, Infante AM, Kobes S; International Type 2 Diabetes 1q Consortium, Bogardus C, Shuldiner AR, Baier LJ. Variants in ARHGEF11, a candidate gene for the linkage to type 2 diabetes on chromosome 1q, are nominally associated with insulin resistance and type 2 diabetes in Pima Indians. Diabetes. 2007;56:1454-1459. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 27] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 8. | Hu C, Zhang R, Wang C, Wang J, Ma X, Lu J, Qin W, Hou X, Wang C, Bao Y, Xiang K, Jia W. PPARG, KCNJ11, CDKAL1, CDKN2A-CDKN2B, IDE-KIF11-HHEX, IGF2BP2 and SLC30A8 are associated with type 2 diabetes in a Chinese population. PLoS One. 2009;4:e7643. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 125] [Cited by in RCA: 141] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 9. | Müssig K, Staiger H, Machicao F, Kirchhoff K, Guthoff M, Schäfer SA, Kantartzis K, Silbernagel G, Stefan N, Holst JJ, Gallwitz B, Häring HU, Fritsche A. Association of type 2 diabetes candidate polymorphisms in KCNQ1 with incretin and insulin secretion. Diabetes. 2009;58:1715-1720. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 84] [Cited by in RCA: 87] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 10. | Gloyn AL, Braun M, Rorsman P. Type 2 diabetes susceptibility gene TCF7L2 and its role in beta-cell function. Diabetes. 2009;58:800-802. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 58] [Cited by in RCA: 47] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 11. | Meigs JB, Cupples LA, Wilson PW. Parental transmission of type 2 diabetes: the Framingham Offspring Study. Diabetes. 2000;49:2201-2207. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 416] [Cited by in RCA: 426] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 12. | Hunter WA, Cundy T, Rabone D, Hofman PL, Harris M, Regan F, Robinson E, Cutfield WS. Insulin sensitivity in the offspring of women with type 1 and type 2 diabetes. Diabetes Care. 2004;27:1148-1152. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 39] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 13. | Gautier JF, Wilson C, Weyer C, Mott D, Knowler WC, Cavaghan M, Polonsky KS, Bogardus C, Pratley RE. Low acute insulin secretory responses in adult offspring of people with early onset type 2 diabetes. Diabetes. 2001;50:1828-1833. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 91] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 14. | Aerts L, Holemans K, Van Assche FA. Maternal diabetes during pregnancy: consequences for the offspring. Diabetes Metab Rev. 1990;6:147-167. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 147] [Cited by in RCA: 137] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 15. | Boloker J, Gertz SJ, Simmons RA. Gestational diabetes leads to the development of diabetes in adulthood in the rat. Diabetes. 2002;51:1499-1506. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 119] [Cited by in RCA: 114] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 16. | Ishihara H, Asano T, Tsukuda K, Katagiri H, Inukai K, Anai M, Kikuchi M, Yazaki Y, Miyazaki JI, Oka Y. Pancreatic beta cell line MIN6 exhibits characteristics of glucose metabolism and glucose-stimulated insulin secretion similar to those of normal islets. Diabetologia. 1993;36:1139-1145. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 319] [Cited by in RCA: 339] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 17. | Merglen A, Theander S, Rubi B, Chaffard G, Wollheim CB, Maechler P. Glucose sensitivity and metabolism-secretion coupling studied during two-year continuous culture in INS-1E insulinoma cells. Endocrinology. 2004;145:667-678. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 458] [Cited by in RCA: 482] [Article Influence: 23.0] [Reference Citation Analysis (0)] |
| 18. | Thomson JA, Itskovitz-Eldor J, Shapiro SS, Waknitz MA, Swiergiel JJ, Marshall VS, Jones JM. Embryonic stem cell lines derived from human blastocysts. Science. 1998;282:1145-1147. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11399] [Cited by in RCA: 10432] [Article Influence: 386.4] [Reference Citation Analysis (0)] |
| 19. | Barker DJ, Gluckman PD, Godfrey KM, Harding JE, Owens JA, Robinson JS. Fetal nutrition and cardiovascular disease in adult life. Lancet. 1993;341:938-941. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1895] [Cited by in RCA: 1843] [Article Influence: 57.6] [Reference Citation Analysis (0)] |
| 20. | Barker DJ, Winter PD, Osmond C, Margetts B, Simmonds SJ. Weight in infancy and death from ischaemic heart disease. Lancet. 1989;2:577-580. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2264] [Cited by in RCA: 2195] [Article Influence: 61.0] [Reference Citation Analysis (0)] |
| 21. | Barker DJ, Osmond C. Infant mortality, childhood nutrition, and ischaemic heart disease in England and Wales. Lancet. 1986;1:1077-1081. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1782] [Cited by in RCA: 1678] [Article Influence: 43.0] [Reference Citation Analysis (0)] |
| 22. | Whincup PH, Kaye SJ, Owen CG, Huxley R, Cook DG, Anazawa S, Barrett-Connor E, Bhargava SK, Birgisdottir BE, Carlsson S, de Rooij SR, Dyck RF, Eriksson JG, Falkner B, Fall C, Forsén T, Grill V, Gudnason V, Hulman S, Hyppönen E, Jeffreys M, Lawlor DA, Leon DA, Minami J, Mishra G, Osmond C, Power C, Rich-Edwards JW, Roseboom TJ, Sachdev HS, Syddall H, Thorsdottir I, Vanhala M, Wadsworth M, Yarbrough DE. Birth weight and risk of type 2 diabetes: a systematic review. JAMA. 2008;300:2886-2897. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 774] [Cited by in RCA: 693] [Article Influence: 40.8] [Reference Citation Analysis (0)] |
| 23. | Ravelli AC, van der Meulen JH, Michels RP, Osmond C, Barker DJ, Hales CN, Bleker OP. Glucose tolerance in adults after prenatal exposure to famine. Lancet. 1998;351:173-177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1102] [Cited by in RCA: 991] [Article Influence: 36.7] [Reference Citation Analysis (0)] |
| 24. | Ravelli AC, van Der Meulen JH, Osmond C, Barker DJ, Bleker OP. Obesity at the age of 50 y in men and women exposed to famine prenatally. Am J Clin Nutr. 1999;70:811-816. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 820] [Cited by in RCA: 741] [Article Influence: 28.5] [Reference Citation Analysis (0)] |
| 25. | Roseboom TJ, van der Meulen JH, Osmond C, Barker DJ, Ravelli AC, Schroeder-Tanka JM, van Montfrans GA, Michels RP, Bleker OP. Coronary heart disease after prenatal exposure to the Dutch famine, 1944-45. Heart. 2000;84:595-598. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 471] [Cited by in RCA: 429] [Article Influence: 17.2] [Reference Citation Analysis (0)] |
| 26. | Painter RC, De Rooij SR, Bossuyt PM, Osmond C, Barker DJ, Bleker OP, Roseboom TJ. A possible link between prenatal exposure to famine and breast cancer: a preliminary study. Am J Hum Biol. 2006;18:853-856. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 70] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 27. | Painter RC, Osmond C, Gluckman P, Hanson M, Phillips DI, Roseboom TJ. Transgenerational effects of prenatal exposure to the Dutch famine on neonatal adiposity and health in later life. BJOG. 2008;115:1243-1249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 432] [Cited by in RCA: 381] [Article Influence: 22.4] [Reference Citation Analysis (0)] |
| 28. | Rugholm S, Baker JL, Olsen LW, Schack-Nielsen L, Bua J, Sørensen TI. Stability of the association between birth weight and childhood overweight during the development of the obesity epidemic. Obes Res. 2005;13:2187-2194. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 97] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 29. | Harder T, Roepke K, Diller N, Stechling Y, Dudenhausen JW, Plagemann A. Birth weight, early weight gain, and subsequent risk of type 1 diabetes: systematic review and meta-analysis. Am J Epidemiol. 2009;169:1428-1436. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 144] [Cited by in RCA: 142] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 30. | Ahlgren M, Wohlfahrt J, Olsen LW, Sørensen TI, Melbye M. Birth weight and risk of cancer. Cancer. 2007;110:412-419. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 104] [Cited by in RCA: 110] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 31. | Markey CM, Luque EH, Munoz De Toro M, Sonnenschein C, Soto AM. In utero exposure to bisphenol A alters the development and tissue organization of the mouse mammary gland. Biol Reprod. 2001;65:1215-1223. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 303] [Cited by in RCA: 258] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 32. | Ziv-Gal A, Wang W, Zhou C, Flaws JA. The effects of in utero bisphenol A exposure on reproductive capacity in several generations of mice. Toxicol Appl Pharmacol. 2015;284:354-362. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 89] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 33. | Ashley-Martin J, Dodds L, Arbuckle TE, Ettinger AS, Shapiro GD, Fisher M, Morisset AS, Taback S, Bouchard MF, Monnier P, Dallaire R, Fraser WD. A birth cohort study to investigate the association between prenatal phthalate and bisphenol A exposures and fetal markers of metabolic dysfunction. Environ Health. 2014;13:84. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 65] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 34. | Alonso-Magdalena P, Morimoto S, Ripoll C, Fuentes E, Nadal A. The estrogenic effect of bisphenol A disrupts pancreatic beta-cell function in vivo and induces insulin resistance. Environ Health Perspect. 2006;114:106-112. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 450] [Cited by in RCA: 453] [Article Influence: 23.8] [Reference Citation Analysis (0)] |
| 35. | Warner M, Rauch S, Ames J, Mocarelli P, Brambilla P, Signorini S, Eskenazi B. In utero dioxin exposure and cardiometabolic risk in the Seveso Second Generation Study. Int J Obes (Lond). 2019;43:2233-2243. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 12] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 36. | Mandy M, Nyirenda M. Developmental Origins of Health and Disease: the relevance to developing nations. Int Health. 2018;10:66-70. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 159] [Cited by in RCA: 157] [Article Influence: 22.4] [Reference Citation Analysis (0)] |
| 37. | American Diabetes Association. Gestational diabetes mellitus. Diabetes Care. 2004;27 Suppl 1:S88-S90. [PubMed] |
| 38. | Ross G. Gestational diabetes. Aust Fam Physician. 2006;35:392-396. [PubMed] |
| 39. | Chu SY, Callaghan WM, Kim SY, Schmid CH, Lau J, England LJ, Dietz PM. Maternal obesity and risk of gestational diabetes mellitus. Diabetes Care. 2007;30:2070-2076. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 673] [Cited by in RCA: 692] [Article Influence: 38.4] [Reference Citation Analysis (0)] |
| 40. | Ben-Haroush A, Yogev Y, Hod M. Epidemiology of gestational diabetes mellitus and its association with Type 2 diabetes. Diabet Med. 2004;21:103-113. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 537] [Cited by in RCA: 554] [Article Influence: 26.4] [Reference Citation Analysis (0)] |
| 41. | Van Assche FA, Holemans K, Aerts L. Long-term consequences for offspring of diabetes during pregnancy. Br Med Bull. 2001;60:173-182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 97] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 42. | Kc K, Shakya S, Zhang H. Gestational diabetes mellitus and macrosomia: a literature review. Ann Nutr Metab. 2015;66 Suppl 2:14-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 329] [Cited by in RCA: 553] [Article Influence: 55.3] [Reference Citation Analysis (0)] |
| 43. | Hunter DJ, Burrows RF, Mohide PT, Whyte RK. Influence of maternal insulin-dependent diabetes mellitus on neonatal morbidity. CMAJ. 1993;149:47-52. [PubMed] |
| 44. | Gauguier D, Bihoreau MT, Picon L, Ktorza A. Insulin secretion in adult rats after intrauterine exposure to mild hyperglycemia during late gestation. Diabetes. 1991;40 Suppl 2:109-114. [PubMed] |
| 45. | Atègbo JM, Grissa O, Yessoufou A, Hichami A, Dramane KL, Moutairou K, Miled A, Grissa A, Jerbi M, Tabka Z, Khan NA. Modulation of adipokines and cytokines in gestational diabetes and macrosomia. J Clin Endocrinol Metab. 2006;91:4137-4143. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 259] [Cited by in RCA: 272] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 46. | Grissa O, Atègbo JM, Yessoufou A, Tabka Z, Miled A, Jerbi M, Dramane KL, Moutairou K, Prost J, Hichami A, Khan NA. Antioxidant status and circulating lipids are altered in human gestational diabetes and macrosomia. Transl Res. 2007;150:164-171. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 68] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 47. | Merzouk H, Madani S, Hichami A, Prost J, Belleville J, Khan NA. Age-related changes in fatty acids in obese offspring of streptozotocin-induced diabetic rats. Obes Res. 2002;10:703-714. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 19] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 48. | Westgate JA, Lindsay RS, Beattie J, Pattison NS, Gamble G, Mildenhall LF, Breier BH, Johnstone FD. Hyperinsulinemia in cord blood in mothers with type 2 diabetes and gestational diabetes mellitus in New Zealand. Diabetes Care. 2006;29:1345-1350. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 56] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 49. | Van Assche FA, Aerts L, de Prins F. Degranulation of the insulin-producing beta cells in an infant of a diabetic mother. Case report. Br J Obstet Gynaecol. 1983;90:182-185. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 22] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 50. | Zhao Z, Reece EA. Experimental mechanisms of diabetic embryopathy and strategies for developing therapeutic interventions. J Soc Gynecol Investig. 2005;12:549-557. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 46] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 51. | Loeken MR. Advances in understanding the molecular causes of diabetes-induced birth defects. J Soc Gynecol Investig. 2006;13:2-10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 62] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 52. | Farrell T, Neale L, Cundy T. Congenital anomalies in the offspring of women with type 1, type 2 and gestational diabetes. Diabet Med. 2002;19:322-326. [PubMed] |
| 53. | Serradas P, Goya L, Lacorne M, Gangnerau MN, Ramos S, Alvarez C, Pascual-Leone AM, Portha B. Fetal insulin-like growth factor-2 production is impaired in the GK rat model of type 2 diabetes. Diabetes. 2002;51:392-397. [PubMed] |
| 54. | Amri K, Freund N, Vilar J, Merlet-Bénichou C, Lelièvre-Pégorier M. Adverse effects of hyperglycemia on kidney development in rats: in vivo and in vitro studies. Diabetes. 1999;48:2240-2245. [PubMed] |
| 55. | Amri K, Freund N, Duong Van Huyen JP, Merlet-Bénichou C, Lelièvre-Pégorier M. Altered nephrogenesis due to maternal diabetes is associated with increased expression of IGF-II/mannose-6-phosphate receptor in the fetal kidney. Diabetes. 2001;50:1069-1075. [PubMed] |
| 56. | Messerschmidt DM, Knowles BB, Solter D. DNA methylation dynamics during epigenetic reprogramming in the germline and preimplantation embryos. Genes Dev. 2014;28:812-828. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 487] [Cited by in RCA: 482] [Article Influence: 43.8] [Reference Citation Analysis (0)] |
| 57. | Perera F, Herbstman J. Prenatal environmental exposures, epigenetics, and disease. Reprod Toxicol. 2011;31:363-373. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 487] [Cited by in RCA: 428] [Article Influence: 30.6] [Reference Citation Analysis (0)] |
| 58. | Park JH, Stoffers DA, Nicholls RD, Simmons RA. Development of type 2 diabetes following intrauterine growth retardation in rats is associated with progressive epigenetic silencing of Pdx1. J Clin Invest. 2008;118:2316-2324. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 273] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 59. | Yang BT, Dayeh TA, Volkov PA, Kirkpatrick CL, Malmgren S, Jing X, Renström E, Wollheim CB, Nitert MD, Ling C. Increased DNA methylation and decreased expression of PDX-1 in pancreatic islets from patients with type 2 diabetes. Mol Endocrinol. 2012;26:1203-1212. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 223] [Article Influence: 17.2] [Reference Citation Analysis (0)] |
| 60. | Oliver-Krasinski JM, Kasner MT, Yang J, Crutchlow MF, Rustgi AK, Kaestner KH, Stoffers DA. The diabetes gene Pdx1 regulates the transcriptional network of pancreatic endocrine progenitor cells in mice. J Clin Invest. 2009;119:1888-1898. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 111] [Cited by in RCA: 127] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 61. | Ling C, Del Guerra S, Lupi R, Rönn T, Granhall C, Luthman H, Masiello P, Marchetti P, Groop L, Del Prato S. Epigenetic regulation of PPARGC1A in human type 2 diabetic islets and effect on insulin secretion. Diabetologia. 2008;51:615-622. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 354] [Cited by in RCA: 343] [Article Influence: 20.2] [Reference Citation Analysis (0)] |
| 62. | Volkmar M, Dedeurwaerder S, Cunha DA, Ndlovu MN, Defrance M, Deplus R, Calonne E, Volkmar U, Igoillo-Esteve M, Naamane N, Del Guerra S, Masini M, Bugliani M, Marchetti P, Cnop M, Eizirik DL, Fuks F. DNA methylation profiling identifies epigenetic dysregulation in pancreatic islets from type 2 diabetic patients. EMBO J. 2012;31:1405-1426. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 309] [Cited by in RCA: 305] [Article Influence: 23.5] [Reference Citation Analysis (0)] |
| 63. | Dayeh T, Volkov P, Salö S, Hall E, Nilsson E, Olsson AH, Kirkpatrick CL, Wollheim CB, Eliasson L, Rönn T, Bacos K, Ling C. Genome-wide DNA methylation analysis of human pancreatic islets from type 2 diabetic and non-diabetic donors identifies candidate genes that influence insulin secretion. PLoS Genet. 2014;10:e1004160. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 330] [Cited by in RCA: 358] [Article Influence: 32.5] [Reference Citation Analysis (0)] |
| 64. | Petropoulos S, Guillemin C, Ergaz Z, Dimov S, Suderman M, Weinstein-Fudim L, Ornoy A, Szyf M. Gestational Diabetes Alters Offspring DNA Methylation Profiles in Human and Rat: Identification of Key Pathways Involved in Endocrine System Disorders, Insulin Signaling, Diabetes Signaling, and ILK Signaling. Endocrinology. 2015;156:2222-2238. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 56] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 65. | Chen S, Feng B, George B, Chakrabarti R, Chen M, Chakrabarti S. Transcriptional coactivator p300 regulates glucose-induced gene expression in endothelial cells. Am J Physiol Endocrinol Metab. 2010;298:E127-E137. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 120] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 66. | Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861-872. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14327] [Cited by in RCA: 14321] [Article Influence: 842.4] [Reference Citation Analysis (0)] |
| 67. | Kondo Y, Toyoda T, Inagaki N, Osafune K. iPSC technology-based regenerative therapy for diabetes. J Diabetes Investig. 2018;9:234-243. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 59] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 68. | Mills JL. Malformations in infants of diabetic mothers. Teratology 25:385-94. 1982. Birth Defects Res A Clin Mol Teratol. 2010;88:769-778. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 44] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 69. | Gabbay-Benziv R, Reece EA, Wang F, Yang P. Birth defects in pregestational diabetes: Defect range, glycemic threshold and pathogenesis. World J Diabetes. 2015;6:481-488. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 98] [Cited by in RCA: 102] [Article Influence: 10.2] [Reference Citation Analysis (2)] |
| 71. | Wells JM, Melton DA. Vertebrate endoderm development. Annu Rev Cell Dev Biol. 1999;15:393-410. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 402] [Cited by in RCA: 369] [Article Influence: 14.2] [Reference Citation Analysis (0)] |
| 72. | Kubo A, Shinozaki K, Shannon JM, Kouskoff V, Kennedy M, Woo S, Fehling HJ, Keller G. Development of definitive endoderm from embryonic stem cells in culture. Development. 2004;131:1651-1662. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 648] [Cited by in RCA: 630] [Article Influence: 30.0] [Reference Citation Analysis (0)] |
| 73. | Blum M, Gaunt SJ, Cho KW, Steinbeisser H, Blumberg B, Bittner D, De Robertis EM. Gastrulation in the mouse: the role of the homeobox gene goosecoid. Cell. 1992;69:1097-1106. [PubMed] |
| 74. | Teo AK, Arnold SJ, Trotter MW, Brown S, Ang LT, Chng Z, Robertson EJ, Dunn NR, Vallier L. Pluripotency factors regulate definitive endoderm specification through eomesodermin. Genes Dev. 2011;25:238-250. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 311] [Cited by in RCA: 277] [Article Influence: 19.8] [Reference Citation Analysis (0)] |
| 75. | Hart AH, Hartley L, Sourris K, Stadler ES, Li R, Stanley EG, Tam PP, Elefanty AG, Robb L. Mixl1 is required for axial mesendoderm morphogenesis and patterning in the murine embryo. Development. 2002;129:3597-3608. [PubMed] |
| 76. | D'Amour KA, Bang AG, Eliazer S, Kelly OG, Agulnick AD, Smart NG, Moorman MA, Kroon E, Carpenter MK, Baetge EE. Production of pancreatic hormone-expressing endocrine cells from human embryonic stem cells. Nat Biotechnol. 2006;24:1392-1401. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1447] [Cited by in RCA: 1413] [Article Influence: 74.4] [Reference Citation Analysis (0)] |
| 77. | Champeris Tsaniras S, Jones PM. Generating pancreatic beta-cells from embryonic stem cells by manipulating signaling pathways. J Endocrinol. 2010;206:13-26. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 40] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 78. | Mfopou JK, Geeraerts M, Dejene R, Van Langenhoven S, Aberkane A, Van Grunsven LA, Bouwens L. Efficient definitive endoderm induction from mouse embryonic stem cell adherent cultures: a rapid screening model for differentiation studies. Stem Cell Res. 2014;12:166-177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 29] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 79. | Lu CC, Brennan J, Robertson EJ. From fertilization to gastrulation: axis formation in the mouse embryo. Curr Opin Genet Dev. 2001;11:384-392. [PubMed] |
| 80. | Jaramillo M, Mathew S, Task K, Barner S, Banerjee I. Potential for pancreatic maturation of differentiating human embryonic stem cells is sensitive to the specific pathway of definitive endoderm commitment. PLoS One. 2014;9:e94307. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 14] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 81. | Wang P, Rodriguez RT, Wang J, Ghodasara A, Kim SK. Targeting SOX17 in human embryonic stem cells creates unique strategies for isolating and analyzing developing endoderm. Cell Stem Cell. 2011;8:335-346. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 123] [Cited by in RCA: 112] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 82. | D'Amour KA, Agulnick AD, Eliazer S, Kelly OG, Kroon E, Baetge EE. Efficient differentiation of human embryonic stem cells to definitive endoderm. Nat Biotechnol. 2005;23:1534-1541. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1288] [Cited by in RCA: 1288] [Article Influence: 64.4] [Reference Citation Analysis (0)] |
| 83. | Sumi T, Tsuneyoshi N, Nakatsuji N, Suemori H. Defining early lineage specification of human embryonic stem cells by the orchestrated balance of canonical Wnt/beta-catenin, Activin/Nodal and BMP signaling. Development. 2008;135:2969-2979. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 244] [Cited by in RCA: 247] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 84. | Borowiak M, Maehr R, Chen S, Chen AE, Tang W, Fox JL, Schreiber SL, Melton DA. Small molecules efficiently direct endodermal differentiation of mouse and human embryonic stem cells. Cell Stem Cell. 2009;4:348-358. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 355] [Cited by in RCA: 327] [Article Influence: 20.4] [Reference Citation Analysis (0)] |
| 85. | Chen ACH, Lee YL, Fong SW, Wong CCY, Ng EHY, Yeung WSB. Hyperglycemia impedes definitive endoderm differentiation of human embryonic stem cells by modulating histone methylation patterns. Cell Tissue Res. 2017;368:563-578. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 19] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 86. | Guney MA, Gannon M. Pancreas cell fate. Birth Defects Res C Embryo Today. 2009;87:232-248. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 55] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 87. | Kroon E, Martinson LA, Kadoya K, Bang AG, Kelly OG, Eliazer S, Young H, Richardson M, Smart NG, Cunningham J, Agulnick AD, D'Amour KA, Carpenter MK, Baetge EE. Pancreatic endoderm derived from human embryonic stem cells generates glucose-responsive insulin-secreting cells in vivo. Nat Biotechnol. 2008;26:443-452. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1331] [Cited by in RCA: 1304] [Article Influence: 76.7] [Reference Citation Analysis (0)] |
| 88. | Apelqvist A, Ahlgren U, Edlund H. Sonic hedgehog directs specialised mesoderm differentiation in the intestine and pancreas. Curr Biol. 1997;7:801-804. [PubMed] |
| 89. | Chen S, Borowiak M, Fox JL, Maehr R, Osafune K, Davidow L, Lam K, Peng LF, Schreiber SL, Rubin LL, Melton D. A small molecule that directs differentiation of human ESCs into the pancreatic lineage. Nat Chem Biol. 2009;5:258-265. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 384] [Cited by in RCA: 360] [Article Influence: 22.5] [Reference Citation Analysis (0)] |
| 90. | Rezania A, Bruin JE, Xu J, Narayan K, Fox JK, O'Neil JJ, Kieffer TJ. Enrichment of human embryonic stem cell-derived NKX6.1-expressing pancreatic progenitor cells accelerates the maturation of insulin-secreting cells in vivo. Stem Cells. 2013;31:2432-2442. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 196] [Cited by in RCA: 203] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 91. | Haller C, Piccand J, De Franceschi F, Ohi Y, Bhoumik A, Boss C, De Marchi U, Jacot G, Metairon S, Descombes P, Wiederkehr A, Palini A, Bouche N, Steiner P, Kelly OG, R-C Kraus M. Macroencapsulated Human iPSC-Derived Pancreatic Progenitors Protect against STZ-Induced Hyperglycemia in Mice. Stem Cell Reports. 2019;12:787-800. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 31] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 92. | Zhang D, Jiang W, Liu M, Sui X, Yin X, Chen S, Shi Y, Deng H. Highly efficient differentiation of human ES cells and iPS cells into mature pancreatic insulin-producing cells. Cell Res. 2009;19:429-438. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 429] [Cited by in RCA: 415] [Article Influence: 25.9] [Reference Citation Analysis (0)] |
| 93. | Pagliuca FW, Millman JR, Gürtler M, Segel M, Van Dervort A, Ryu JH, Peterson QP, Greiner D, Melton DA. Generation of functional human pancreatic β cells in vitro. Cell. 2014;159:428-439. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1321] [Cited by in RCA: 1526] [Article Influence: 152.6] [Reference Citation Analysis (1)] |
| 94. | Takahashi Y, Sekine K, Kin T, Takebe T, Taniguchi H. Self-Condensation Culture Enables Vascularization of Tissue Fragments for Efficient Therapeutic Transplantation. Cell Rep. 2018;23:1620-1629. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 100] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 95. | Xie R, Everett LJ, Lim HW, Patel NA, Schug J, Kroon E, Kelly OG, Wang A, D'Amour KA, Robins AJ, Won KJ, Kaestner KH, Sander M. Dynamic chromatin remodeling mediated by polycomb proteins orchestrates pancreatic differentiation of human embryonic stem cells. Cell Stem Cell. 2013;12:224-237. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 179] [Cited by in RCA: 190] [Article Influence: 15.8] [Reference Citation Analysis (0)] |
| 96. | Astro V, Adamo A. Epigenetic Control of Endocrine Pancreas Differentiation in vitro: Current Knowledge and Future Perspectives. Front Cell Dev Biol. 2018;6:141. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 8] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 97. | Li J, Wu X, Zhou Y, Lee M, Guo L, Han W, Mo W, Cao WM, Sun D, Xie R, Huang Y. Decoding the dynamic DNA methylation and hydroxymethylation landscapes in endodermal lineage intermediates during pancreatic differentiation of hESC. Nucleic Acids Res. 2018;46:2883-2900. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 56] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 98. | Whitehead R, Guan H, Arany E, Cernea M, Yang K. Prenatal exposure to bisphenol A alters mouse fetal pancreatic morphology and islet composition. Horm Mol Biol Clin Investig. 2016;25:171-179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 15] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 99. | Lai KP, Li JW, Chan TF, Chen A, Lee CYL, Yeung WSB, Wong CKC. Transcriptomic and methylomic analysis reveal the toxicological effect of 2,3,7,8-Tetrachlorodibenzodioxin on human embryonic stem cell. Chemosphere. 2018;206:663-673. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 7] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 100. | Kubi JA, Chen ACH, Fong SW, Lai KP, Wong CKC, Yeung WSB, Lee KF, Lee YL. Effects of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) on the differentiation of embryonic stem cells towards pancreatic lineage and pancreatic beta cell function. Environ Int. 2019;130:104885. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 15] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 101. | Später D, Hansson EM, Zangi L, Chien KR. How to make a cardiomyocyte. Development. 2014;141:4418-4431. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 110] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 102. | Evans SM, Yelon D, Conlon FL, Kirby ML. Myocardial lineage development. Circ Res. 2010;107:1428-1444. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 243] [Cited by in RCA: 210] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 103. | Foley AC, Mercola M. Heart induction by Wnt antagonists depends on the homeodomain transcription factor Hex. Genes Dev. 2005;19:387-396. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 164] [Cited by in RCA: 156] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 104. | Bondue A, Lapouge G, Paulissen C, Semeraro C, Iacovino M, Kyba M, Blanpain C. Mesp1 acts as a master regulator of multipotent cardiovascular progenitor specification. Cell Stem Cell. 2008;3:69-84. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 293] [Cited by in RCA: 286] [Article Influence: 16.8] [Reference Citation Analysis (0)] |
| 105. | Kehat I, Kenyagin-Karsenti D, Snir M, Segev H, Amit M, Gepstein A, Livne E, Binah O, Itskovitz-Eldor J, Gepstein L. Human embryonic stem cells can differentiate into myocytes with structural and functional properties of cardiomyocytes. J Clin Invest. 2001;108:407-414. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 460] [Article Influence: 19.2] [Reference Citation Analysis (0)] |
| 106. | Yoon BS, Yoo SJ, Lee JE, You S, Lee HT, Yoon HS. Enhanced differentiation of human embryonic stem cells into cardiomyocytes by combining hanging drop culture and 5-azacytidine treatment. Differentiation. 2006;74:149-159. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 126] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 107. | Kattman SJ, Witty AD, Gagliardi M, Dubois NC, Niapour M, Hotta A, Ellis J, Keller G. Stage-specific optimization of activin/nodal and BMP signaling promotes cardiac differentiation of mouse and human pluripotent stem cell lines. Cell Stem Cell. 2011;8:228-240. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 868] [Cited by in RCA: 869] [Article Influence: 62.1] [Reference Citation Analysis (0)] |
| 108. | Mohr JC, Zhang J, Azarin SM, Soerens AG, de Pablo JJ, Thomson JA, Lyons GE, Palecek SP, Kamp TJ. The microwell control of embryoid body size in order to regulate cardiac differentiation of human embryonic stem cells. Biomaterials. 2010;31:1885-1893. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 157] [Cited by in RCA: 167] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 109. | Ting S, Chen A, Reuveny S, Oh S. An intermittent rocking platform for integrated expansion and differentiation of human pluripotent stem cells to cardiomyocytes in suspended microcarrier cultures. Stem Cell Res. 2014;13:202-213. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 59] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 110. | Dahlmann J, Kensah G, Kempf H, Skvorc D, Gawol A, Elliott DA, Dräger G, Zweigerdt R, Martin U, Gruh I. The use of agarose microwells for scalable embryoid body formation and cardiac differentiation of human and murine pluripotent stem cells. Biomaterials. 2013;34:2463-2471. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 103] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 111. | Hornberger LK. Maternal diabetes and the fetal heart. Heart. 2006;92:1019-1021. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 60] [Cited by in RCA: 58] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 112. | Narchi H, Kulaylat N. Heart disease in infants of diabetic mothers. Images Paediatr Cardiol. 2000;2:17-23. [PubMed] |
| 113. | Dyntar D, Sergeev P, Klisic J, Ambühl P, Schaub MC, Donath MY. High glucose alters cardiomyocyte contacts and inhibits myofibrillar formation. J Clin Endocrinol Metab. 2006;91:1961-1967. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 44] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 114. | Drawnel FM, Boccardo S, Prummer M, Delobel F, Graff A, Weber M, Gérard R, Badi L, Kam-Thong T, Bu L, Jiang X, Hoflack JC, Kiialainen A, Jeworutzki E, Aoyama N, Carlson C, Burcin M, Gromo G, Boehringer M, Stahlberg H, Hall BJ, Magnone MC, Kolaja K, Chien KR, Bailly J, Iacone R. Disease modeling and phenotypic drug screening for diabetic cardiomyopathy using human induced pluripotent stem cells. Cell Rep. 2014;9:810-821. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 164] [Cited by in RCA: 196] [Article Influence: 17.8] [Reference Citation Analysis (0)] |
| 115. | Karakikes I, Ameen M, Termglinchan V, Wu JC. Human induced pluripotent stem cell-derived cardiomyocytes: insights into molecular, cellular, and functional phenotypes. Circ Res. 2015;117:80-88. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 292] [Cited by in RCA: 357] [Article Influence: 35.7] [Reference Citation Analysis (0)] |
| 116. | Lundy SD, Zhu WZ, Regnier M, Laflamme MA. Structural and functional maturation of cardiomyocytes derived from human pluripotent stem cells. Stem Cells Dev. 2013;22:1991-2002. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 484] [Cited by in RCA: 567] [Article Influence: 47.3] [Reference Citation Analysis (0)] |
| 117. | Yang P, Chen X, Kaushal S, Reece EA, Yang P. High glucose suppresses embryonic stem cell differentiation into cardiomyocytes : High glucose inhibits ES cell cardiogenesis. Stem Cell Res Ther. 2016;7:187. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 26] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 118. | Mochizuki H, Ohnuki Y, Kurosawa H. Effect of glucose concentration during embryoid body (EB) formation from mouse embryonic stem cells on EB growth and cell differentiation. J Biosci Bioeng. 2011;111:92-97. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 19] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 119. | Gilsbach R, Schwaderer M, Preissl S, Grüning BA, Kranzhöfer D, Schneider P, Nührenberg TG, Mulero-Navarro S, Weichenhan D, Braun C, Dreßen M, Jacobs AR, Lahm H, Doenst T, Backofen R, Krane M, Gelb BD, Hein L. Distinct epigenetic programs regulate cardiac myocyte development and disease in the human heart in vivo. Nat Commun. 2018;9:391. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 178] [Cited by in RCA: 175] [Article Influence: 25.0] [Reference Citation Analysis (0)] |
| 120. | Hoff K, Lemme M, Kahlert AK, Runde K, Audain E, Schuster D, Scheewe J, Attmann T, Pickardt T, Caliebe A, Siebert R, Kramer HH, Milting H, Hansen A, Ammerpohl O, Hitz MP. DNA methylation profiling allows for characterization of atrial and ventricular cardiac tissues and hiPSC-CMs. Clin Epigenetics. 2019;11:89. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 12] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 121. | Becerra JE, Khoury MJ, Cordero JF, Erickson JD. Diabetes mellitus during pregnancy and the risks for specific birth defects: a population-based case-control study. Pediatrics. 1990;85:1-9. [PubMed] |
| 122. | Correa A, Gilboa SM, Besser LM, Botto LD, Moore CA, Hobbs CA, Cleves MA, Riehle-Colarusso TJ, Waller DK, Reece EA. Diabetes mellitus and birth defects. Am J Obstet Gynecol. 2008;199:237.e1-237.e9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 416] [Cited by in RCA: 478] [Article Influence: 28.1] [Reference Citation Analysis (0)] |
| 123. | Wu Y, Wang F, Fu M, Wang C, Quon MJ, Yang P. Cellular Stress, Excessive Apoptosis, and the Effect of Metformin in a Mouse Model of Type 2 Diabetic Embryopathy. Diabetes. 2015;64:2526-2536. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 62] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 124. | Yang P, Li X, Xu C, Eckert RL, Reece EA, Zielke HR, Wang F. Maternal hyperglycemia activates an ASK1-FoxO3a-caspase 8 pathway that leads to embryonic neural tube defects. Sci Signal. 2013;6:ra74. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 76] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 125. | Yang P, Shen WB, Reece EA, Chen X, Yang P. High glucose suppresses embryonic stem cell differentiation into neural lineage cells. Biochem Biophys Res Commun. 2016;472:306-312. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 19] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 126. | Burren KA, Savery D, Massa V, Kok RM, Scott JM, Blom HJ, Copp AJ, Greene ND. Gene-environment interactions in the causation of neural tube defects: folate deficiency increases susceptibility conferred by loss of Pax3 function. Hum Mol Genet. 2008;17:3675-3685. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 97] [Cited by in RCA: 91] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 127. | Vega RB, Matsuda K, Oh J, Barbosa AC, Yang X, Meadows E, McAnally J, Pomajzl C, Shelton JM, Richardson JA, Karsenty G, Olson EN. Histone deacetylase 4 controls chondrocyte hypertrophy during skeletogenesis. Cell. 2004;119:555-566. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 589] [Cited by in RCA: 605] [Article Influence: 28.8] [Reference Citation Analysis (0)] |
| 128. | Cheng HL, Mostoslavsky R, Saito S, Manis JP, Gu Y, Patel P, Bronson R, Appella E, Alt FW, Chua KF. Developmental defects and p53 hyperacetylation in Sir2 homolog (SIRT1)-deficient mice. Proc Natl Acad Sci U S A. 2003;100:10794-10799. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 863] [Cited by in RCA: 916] [Article Influence: 41.6] [Reference Citation Analysis (0)] |
| 129. | Dottori M, Pera MF. Neural differentiation of human embryonic stem cells. Methods Mol Biol. 2008;438:19-30. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 41] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 130. | Brieño-Enríquez MA, García-López J, Cárdenas DB, Guibert S, Cleroux E, Děd L, Hourcade Jde D, Pěknicová J, Weber M, Del Mazo J. Exposure to endocrine disruptor induces transgenerational epigenetic deregulation of microRNAs in primordial germ cells. PLoS One. 2015;10:e0124296. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 76] [Cited by in RCA: 71] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 131. | Meyer DN, Baker BB, Baker TR. Ancestral TCDD Exposure Induces Multigenerational Histologic and Transcriptomic Alterations in Gonads of Male Zebrafish. Toxicol Sci. 2018;164:603-612. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 132. | Ding GL, Wang FF, Shu J, Tian S, Jiang Y, Zhang D, Wang N, Luo Q, Zhang Y, Jin F, Leung PC, Sheng JZ, Huang HF. Transgenerational glucose intolerance with Igf2/H19 epigenetic alterations in mouse islet induced by intrauterine hyperglycemia. Diabetes. 2012;61:1133-1142. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 195] [Cited by in RCA: 209] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 133. | Ren J, Cheng Y, Ming ZH, Dong XY, Zhou YZ, Ding GL, Pang HY, Rahman TU, Akbar R, Huang HF, Sheng JZ. Intrauterine hyperglycemia exposure results in intergenerational inheritance via DNA methylation reprogramming on F1 PGCs. Epigenetics Chromatin. 2018;11:20. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 35] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 134. | Zhou Q, Wang M, Yuan Y, Wang X, Fu R, Wan H, Xie M, Liu M, Guo X, Zheng Y, Feng G, Shi Q, Zhao XY, Sha J, Zhou Q. Complete Meiosis from Embryonic Stem Cell-Derived Germ Cells In Vitro. Cell Stem Cell. 2016;18:330-340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 313] [Cited by in RCA: 268] [Article Influence: 29.8] [Reference Citation Analysis (0)] |
| 135. | Kobayashi T, Zhang H, Tang WWC, Irie N, Withey S, Klisch D, Sybirna A, Dietmann S, Contreras DA, Webb R, Allegrucci C, Alberio R, Surani MA. Principles of early human development and germ cell program from conserved model systems. Nature. 2017;546:416-420. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 233] [Cited by in RCA: 228] [Article Influence: 28.5] [Reference Citation Analysis (0)] |
| 136. | Wang Q, Kurita H, Carreira V, Ko CI, Fan Y, Zhang X, Biesiada J, Medvedovic M, Puga A. Ah Receptor Activation by Dioxin Disrupts Activin, BMP, and WNT Signals During the Early Differentiation of Mouse Embryonic Stem Cells and Inhibits Cardiomyocyte Functions. Toxicol Sci. 2016;149:346-357. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 52] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 137. | Zhou R, Cheng W, Feng Y, Wei H, Liang F, Wang Y. Interactions between three typical endocrine-disrupting chemicals (EDCs) in binary mixtures exposure on myocardial differentiation of mouse embryonic stem cell. Chemosphere. 2017;178:378-383. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 37] [Article Influence: 4.6] [Reference Citation Analysis (0)] |