Published online May 15, 2003. doi: 10.3748/wjg.v9.i5.894
Revised: July 23, 2002
Accepted: August 2, 2002
Published online: May 15, 2003
AIM: To investigate the expression level of ZNRD1 gene in gastric cancer cells SGC7901 and gastric cancer MDR (multidrug resistant) cells SGC7901/VCR, and to observe the drug sensitizing and proliferation effect of ZNRD1 antisense nucleic acid transduction on SGC7901/VCR cells.
METHODS: Amplification of sequences encoding ZNRD1 from SGC7901/VCR cDNA by PCR. The levels of ZNRD1 mRNA expression were demonstrated using semiquantitative reverse transcription polymerase chain reaction (RT-PCR). Eukaryotic expression vector pcDNA3.1-anti ZNRD1 was constructed and transfected into SGC7901/VCR cells by lipofectamine. Immunochemical method was used to detect the expression of protein in SGC7901/VCR cells and transfectants. The cell cycle alteration and the intracellular adriamycin (ADM) accumulation were observed by FACS. Growth curve and drug sensitization of cells for vincristine (VCR) were analyzed with MTT assay.
RESULTS: We cloned the open reading frame of full-length ZNRD1. The expression of ZNRD1 showed higher in SGC7901/VCR than in SGC7901 cells. The antisense ZNRD1 drug-resistant clones were selected after gene transfection. Immunochemical results showed that the expression level of ZNRD1 protein was lower in anti ZNRD1-SGC7901/VCR cells than that in non-transfectants. Comparing to SGC7901/VCR and pcDNA3.1-SGC7901/VCR, anti ZNRD1-SGC7901/VCR showed gradually accumulated in G1 phase, with a concomitant decrease of cell population in S phase. FACS also suggested intracellular ADM accumulation increased 2fold in SGC7901/VCR cells after transfected with antisense ZNRD1. MTT assay showed that transfectants cells proliferation was lagged and more sensitive to VCR than non-transfectants.
CONCLUSION: ZNRD1 gene displayed highly expression in VCR resistant gastric cancer cells. Expression of ZNRD1 protein was effectively blocked in anti ZNRD1-SGC7901/VCR cells by gene transfection. ZNRD1 antisense nucleic acid transfection sensitized drug resistant gastric cancer cells to VCR, increased ADM accumulation and inhibited the cells proliferation. ZNRD1 antisense RNA transduction could reverse the MDR of human drug-resistant gastric cancer cell SGC7901/VCR to a degree.
- Citation: Zhang YM, Zhao YQ, Pan YL, Shi YQ, Jin XH, Yi H, Fan DM. Effect of ZNRD1 gene antisense RNA on drug resistant gastric cancer cells. World J Gastroenterol 2003; 9(5): 894-898
- URL: https://www.wjgnet.com/1007-9327/full/v9/i5/894.htm
- DOI: https://dx.doi.org/10.3748/wjg.v9.i5.894
Chemotherapy is one of the major methods in tumor treatment, but it often does not work due to MDR. In previous studies, we examined the expression of ZNRD1 is higher in VCR-resistant (SGC7901/VCR) cells than in parental cells SGC7901 by Subtractive Hybridization. The purpose of this study was to investigate the expression and effect of ZNRD1. We transfected ZNRD1 antisense nucleic acid, which is a Homo sapiens transcription-associated zinc ribbon protein, into VCR-resistant gastric cancer cells, and observed the expression of target protein in transfectants and the sensitivity of transfectants to chemotherapeutic agents in order to find the ground for reversing gastric cancer MDR.
Gastric cancer cell line SGC7901, VCR-resistant gastric cancer SGC7901/VCR cells, DH5α bacterial strain and rat anti human ZNRD1 polyclonal antibody were kept in our department. The expression vector pcDNA3.1+ was from our department. EcoR V, Xba I, BamH I, cloning vector pUCm-T and T4 DNA ligase were purchased from Takara; MTT, DEPC from Sigma. TRIZOL, M-MuLV RT enzyme, Taq DNA polymerase, DMEM, Lipofectamine and G418 were products of Gibco BRL, and Primer from Sangon, Shanghai. Avidin biotin peroxidase complex (ABC) kit was obtained from Vector. VCR from Shanghai Hualian Pharmaceutical Co. Ltd, and ADM from Meiji Pharmaceutical Co. Ltd.
Cell culture The MDR subline SGC7901/VCR was developed by exposing the parental SGC7901 cells to stepwise increasing concentrations of anticancer drugs VCR. All cells were cultivated in DMEM supplemented with 10% heat-inactivated fetal bovine serum at 37 °C in 5% CO2 atmosphere. Medium for SGC7901/VCR cells was further supplemented with VCR (1.0 μg/ml). Before use in experiments, SGC7901/VCR cells were cultured in drug-free medium for 2 weeks. Cell stocks were cultivated in these conditions until nearly confluent and subcultured at 1:20 dilution or plated for experiments.
RT-PCR of ZNRD1 Total cellular RNA was extracted from SGC7901 and SGC7901/VCR cells using TRIZOL reagent, cDNA was transcripted from mRNA by RT-PCR. Total RNA (5 μg) and random primer (2 μg) in DEPC water were denatured at 65 °C for 15 min, then 4 μL 5 × reverse transcriptase buffer, 3 μL 10 mmol·L-1 dNTP, 1 μL M-MuLV reverse transcriptase (200 U) and DEPC water were added to have the total volume of 20 μL. The reaction was performed at 25 °C for 10 min, then 42 °C for 1 hour, and 70 °C for 10 min to inactivate the reverse transcriptase. PCR was performed in a 50 µl reaction mixture containing 2 µL reactant, 2.5 U Taq DNA polymerase, deoxynucleoside triphosphates (0.2 mM each), and 1.5 mM MgCl2. The sense primer (5’-ccaactccctcctcagacccg-3’) and antisense primer (5’-cctgggcaaatatacagtcc-3’) designed according to the DNA sequence of ZNRD1 (GenBank accession no. AF024617). The PCR consisted of initial denaturation at 94 °C for 5 min, followed by 30 reaction cycles (50 s at 94 °C and 60 °C, and 30 s at 70 °C) and a final cycle at 72 °C for 10 min. The β2-microglobulin was used to internal standard. The amplified fragments were detected by agarose gel electrophoresise.
Construction of recombinant pUCm-T-ZNRD1 and sequencing of ZNRD1 The PCR product of about 434 bp was isolated and ligated with pUCm-T vector by T4 DNA ligase. E. coli DH5α was transformed with the resulted recombinants pUCm-T-ZNRD1 and the positive bacteria colonies were screened by ampicillin resistant and blue-white screening with X-gal and IPTG. The cDNA of ZNRD1 cloned in pUCm-T was sequenced by the dideoxynucleotide termination procedure of Sanger.
Construction and identification of antisense nucleic acid The ZNRD1 cDNA was generated by digestion of pUCm-T-ZNRD1 plasmid with EcoR V and BamH I restriction enzyme site. Under T4 DNA ligase, cDNA fragment was inserted in the reverse orientation into the pcDNA3.1+ expression vector that was linearized with the same enzymes. The recombinant plasmid was identified through enzyme digestion and agarose gel electrophoresis.
Gene transfection and selection The pcDNA3.1-anti ZNRD1 (1 μg) diluted in 100 μl DMEM was mixed with 2 μl lipofectamine, and placed at room temperature for 10 min. The mixture was then transfected into 1 × 105 SGC7901/VCR cells. After 24 h incubation at 37 °C, the culture was rinsed and re-fed with fresh growth medium. After transfection 48 h, the culture was split and then selected in the culture medium containing the neomycin analogue G418 (400 mg/L). At the same time, pcDNA3.1+ vector that lacked target genes transfected cells and non-transfectants served as negative controls.
Immunocytochemistry The expression of ZNRD1 was detected by immunocytochemical staining with the ABC kit. Cells were seeded in glass slides in DMEM at 37 °C for further 24 h, slides were washed with phosphate-buffered saline (PBS) for 3-5 min, then fixed in cold acetone for 5 min; 50 mL·L-1 H2O2 was added and incubated at room temperature for 10-15 min then added 3 g·L-1 TritonX-100 for another 15 min. The rabbit-anti-human polyclonal antibody ZNRD1 and normal rabbit serum were used at dilution of 1:100, and incubated at 4 °C over night; Biotinylated goat anti-rabbit IgG was added and incubated at 37 °C for 1 h, then incubated with avidin DH-Biotinylated peroxidase for 45 min. Finally, color was developed by immersion of the sections in a peroxidase substrate solution DAB.
Cell cycle Cells were harvested, washed with PBS twice and fixed with ethanol at 4 °C overnight. Fixed cells were washed twice with PBS and stained with propidium iodide. The fluorescence indensity of propidium iodide-stained nuclei was determined by flow cytometric analysis.
Adriamycin accumulation According to described previously[1], cells were cultured in 6-well culture plates at 37 °C for 48 h, ADM was added to the final concentration of 5 mg·L-1. After further culture for 1 h, PBS was used as negative controls. Cells were harvested and suspended in cold PBS; intracellular adriamycin fluorescence intensity was determined by flow cytometric analysis with the stimulative and acceptant wave length at 488 nm and 575 nm, respectively.
Growth curve Exponentially growing cells (103) were trypsinized and inoculated in 96-well plates. The cells were allowed to attach overnight and then incubated for eight days. Cell number and viability were determined by MTT cytotoxic assay. The cells in three wells were supplemented with 20 μl MTT solution (5 g/L) every day. After 4 h, culture media were discarded followed by addition of 150 μl DMSO to melt crystal. The absorbance (A) was measured at 490nm using a microplate reader.
Drug sensitivity assay Dose-survival curves defining the sensitivity of SGC7901/VCR, pcDNA3.1-SGC7901/VCR and anti ZNRD1-SGC7901/VCR transfectants to VCR was obtained using MTT cytotoxic assay. Cells (103-104) diluted with 200 μl 10% DMEM were seeded into 96 well plates, after culturing for 12 h, added VCR (0.05, 0.5, 5 and 50 μg/mL) according to the clinically established plasma peak concentration. Three days later, the absorbance (A) was measured as previously described above. The percentage of viable cells was calculated as follow: (A of experimental group/A of control group) × 100%. The IC50 was obtained by Nosa soft.
Student’s t test was used to assess statistical significance of differences. If P < 0.05, the difference was considered significant.
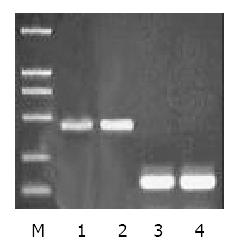
To examine ZNRD1 gene expression in MDR subline SGC7901/VCR and SGC7901 cell line with Semiquantitative RT-PCR. Two fragments were amplified, ZNRD1 cDNA (434 bp) and endogenous control gene β2-microglobulin (118 bp). It is revealed significantly higher ZNRD1 expression in SGC7901/VCR than in SGC7901 cell line (Figure 1).
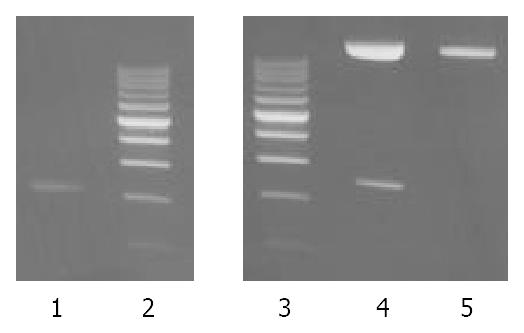
The recombinant of pUCm-T-ZNRD1 was constructed with the human ZNRD1 inserted into the cloning site of vector pUCm-T. Selection and identification of the recombinant was carried out by EcoR V/Xba I endonuclease digestion and agarose electorphoresis (Figure 2). The cloned DNA segments in selected recombinants were sequenced completely. According to the results of DNA sequencing, the cDNA in a selected recombinant was identical to the DNA sequence of ZNRD1 (GenBank accession no. AF024617) in the reading frame.
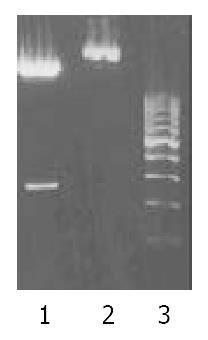
The EcoR V/BamH I fragment containing the complete ZNRD1 cDNA was subcloned into the pcDNA3.1+ expression vector. Selection and identification of the recombinants were carried out by EcoR V/BamH I endonuclease digestion and agarose electrophoresis (Figure 3). The pcDNA3.1+ vector carrying the neomycin-resistance gene for the selection of transfected cells.
When transfected cells were cultured selectively by G418 for 4-5 weeks, resistant clones formed gradually. In contrast, all nontransfectants died two weeks after G418 selection. Resistant clones were further incubated in the presence of low dose G418 for 40-50 days. We got resistant clones, anti ZNRD1-SGC7901/VCR cells.
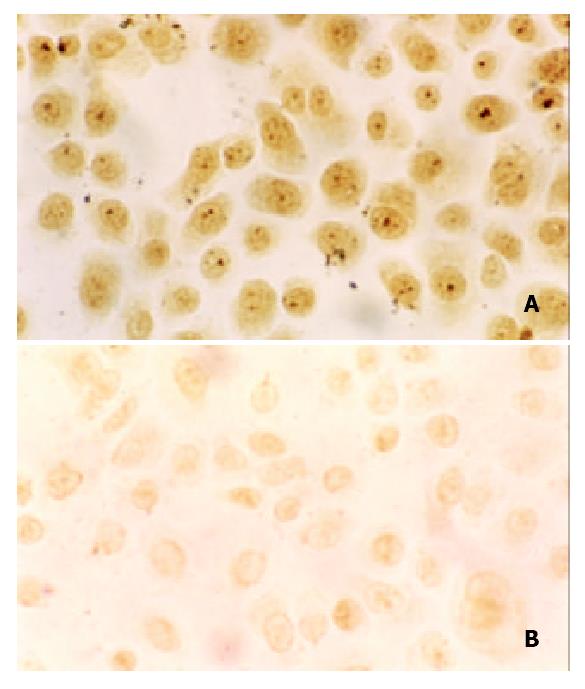
We detected ZNRD1 protein expression in transfectants by immunocytochemistry. All cells had positive staining, staining on anti ZNRD1-SGC7901/VCR cells was weaker than that on SGC7901/VCR, ZNRD1 protein was expressed mostly in the nucleus (Figure 4).
The effects of ZNRD1 antisense nucleic acid on ADM accumulation in SGC7901/VCR cells were determined by flow cytometric analysis. When cells were cultured in drug-DMEM, intracellular drug concentration would increase and finally stabilized at the highest plateau value, which was called adriamycin accumulation. The fluorescence intensity of SGC7901/VCR, pcDNA3.1-SGC7901/VCR and anti ZNRD1-SGC7901/VCR cells, mean fluorescence of 104 cells, is 3.68, 5.70, and 7.24 respectively. The intracellular ADR accumulation marked increase 2fold.
Cell cycle When transfected SGC7901/VCR cells with ZNRD1 antisense nucleic acid, the cell cycle obtained by FCM was as shown in Table 1. The anti ZNRD1-SGC7901/VCR cells gradually accumulated in G1 phase, with a concomitant decrease of cell population in S phase.
| Cell line | G1 | G2 | S | G1/G2 |
| SGC7901/VCR | 0.572 | 0.111 | 0.317 | 1.900 |
| pcDNA3.1-SGC7901/VCR | 0.652 | 0.091 | 0.256 | 1.887 |
| Anti ZNRD1-SGC7901/VCR | 0.744 | 0.118 | 0.138 | 1.881 |
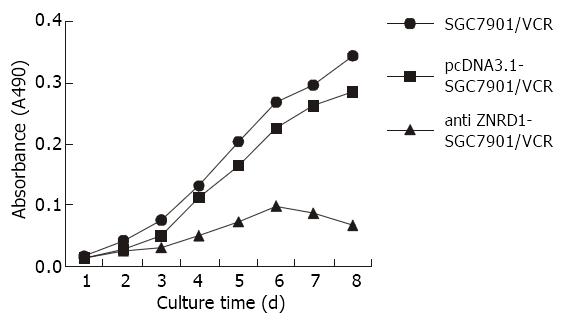
Growth curves The cells viability were daily measured by MTT proliferation analysis. The growth curve showed that cell growth was suppressed after transfected with ZNRD1 antisense nucleic acid (Figure 5).
The IC50 of SGC7901/VCR, pcDNA3.1-SGC7901/VCR and anti ZNRD1-SGC7901/VCR cells treated with VCR is 50.51 ± 0.3, 42.68 ± 0.79, 10.87 ± 0.25 (μg/mL), respectively. The anti ZNRD1-SGC7901/VCR cells were more sensitive to VCR (P < 0.01).
Multidrug resistance (MDR), the principal mechanism by which many cancers develop resistance to chemotherapeutic drugs, is a major factor in the failure of many forms of chemotherapy. There are several ways for cancer cells to develop resistance or defense mechanisms against cytotoxic drugs[2-4]. Several molecules have been found to confer MDR phenotype of tumor cells, such as P-gp, MRP, LRP, BCRP, GSH/GSH, TopoII, apoptosis associated proteins and antiapoptosis proteins. We found the MDR markers are differently over-expressed and no coexpression exists in gastric cancer, MGr1-Ag was a novel MDR protein[5-7]. However, these molecules could not interpret completely how tumor cells develop MDR. The key to the clinical use of reversal agents therefore lies in searching for agents with low toxicity and high reversal activity. For gene therapy has very low toxicity, it might therefore be suitable for clinical use.
SGC7901/VCR is a vincristine-resistant SGC7901 cell line, and it overexpresses P-gp and is cross-resistant to several anti-cancer drugs. In previous studies, we have identified that 54 cDNA fragments were preferentially expressed in SGC7901/VCR cells by DD-PCR[8]. We isolated differentially expressed genes from drug-resistant human gastric adenocarcinoma cell lines using a polymerase chain reaction-based subtractive hybridization technique, and the expression of ZNRD1 is higher in SGC7901/VCR cells than in parental cells SGC7901[9]. ZNRD1 is a new zinc ribbon gene that cloned from the human MHC class I region, named the gene ZNRD1 for zinc ribbon domain-containing 1 protein[10]. Alignment with genomic DNA demonstrates that this gene spans 3.6 kb and consists of four exons and three introns. The full-length cDNA contains an open reading frame of 378 bp. Within the putative polypeptide of 126 amino acids, two zinc-ribbon domains were identified: Cx2Cx15Cx2C at the N-terminal and Cx2Cx24Cx2C at the C-terminal. The C-terminal domain is actually folded as three β-sheets stabilized by a zinc ion instead of finger like helices, compared to TFIIS zinc ribbon folds, they fit each other very well. The conserved amino acid sequence, CxRCx6Yx3QxRSADEx2TxFxCx2C is highly homologous to the yeast RNA polymerase A subunit 9 and transcription-associated proteins.
Transcriptional initiation and elongation provide control points in gene expression. The highly conserved eukaryotic transcriptional elongation factor TFIIS helps overcome elongation barriers and enhances proofreading by RNA polymerase II (RNAPII). The nucleic acid binding domain of TFIIS contains a Cys4 Zn(2+)-binding site with no homology to previously characterized Cys4, Cys6, or Cys2-His2 Zn fingers. Its solution structure exhibits a novel three-stranded antiparallel beta-sheet (designated the Zn ribbon), extends the repertoire of Zn-mediated peptide architectures and highlights the growing recognition of the beta-sheet as a motif of nucleic-acid recognition. The TFIIS functions may be modulated by the Zn ribbon domain through interactions with nucleic acids in the elongation complex. The RNAPII contain a Zn ribbon, in as much as the polymerase’s 15-kDa subunit contains a sequence that aligns well with the TFIIS Zn ribbon sequence, including a similarly placed pair of acidic residues. It has demonstrated that TFIIS Zn ribbon is critical for stimulation of both elongation and RNA cleavage activities of RNAPII. Eukaryotic RNA polymerase II subunit 9 (RPB9) contains Zn ribbon motifs homologous to TFIIS; it regulates start-site selection and elongational arrest. RNAPII lacking the Rpb9 subunit uses alternate transcription initiation sites in vitro and in vivo and is unable to respond to the transcription elongation factor TFIIS in vitro. Alanine substitutions in the C-terminal zinc ribbon domain of Rpb9, like amino acid substitutions in the homologous part of TFIIS, completely eliminated elongation activity. It has show that RPB9 has a synthetic phenotype with the TFIIS gene[11].
DNA primases are enzymes whose continual activity is required at the DNA replication fork. They catalyze the synthesis of short RNA molecules used as primers for DNA polymerases. All DNA primases contain a metal binding site, which is composed of four conserved Cys or His residues that could potentially coordinate zinc[12]. Motif IV in the small subunit contains four cysteines that could form a zinc ribbon motif as a member of the zinc ribbon subfamily of zinc binding motifs.
The ability to selectively regulate the expression of genes implicated in cancer could have important ramifications for both basic research and for therapy. Using peptide combinatorial libraries expressed in yeast, Bartsevich[13] has screened for a novel zinc finger proteins that selectively bind to an overlapping EGR1/SP1/WT1 regulatory site in the promoter of the MDR1 multidrug resistance gene. The novel proteins were only moderately effective in blocking transcription by simple masking of the target site. However, when coupled to mammalian transactivator or repressor domains, they could selectively modulate the expression of reporter genes having promoters containing the MDR1 target site. Moreover, they could also regulate transcription of the chromosomal MDR1 gene. These studies potentially provide a novel alternative approach to the control of multidrug resistance. They also provide important insights into strategies for developing selective regulators of gene expression.
Thus, zinc ribbon is related to transcription and DNA replication, and the construction is same to zinc finger. It may regulate the expression of genes to control MDR.
Semiquantitative RT-PCR analysis is a sensitive and specific method; it is revealed that the RNA expression levels of ZNRD1 were significantly higher in SGC7901/VCR cells than that in SGC7901 cells. We transducted ZNRD1 antisense RNA into SGC7901/VCR cells, then found that intrinsic ZNRD1 protein was decreased, and transfected cells were more sensitive to VCR and cell growth was suppressed, the cell cycle was notably changed, the G1 phase cells were gradually increased, with a concomitant decrease of cell population in S phase, these observations suggest that ZNRD1 antisense RNA may be involved in the action of the G1 checkpoint and inhibition of DNA replication. ADR can emit fluorescence; its intensity represents its accumulation, which was markedly enhanced in transfectants. The results suggested that the formation of MDR in SGC7901/VCR cells was associated with over expression of ZNRD1. Because gastric cancer is common in China and some areas in the world[14-33], these results may be important for further study.
Edited by Bo XN
| 1. | Han Y, Han ZY, Zhou XM, Shi R, Zheng Y, Shi YQ, Miao JY, Pan BR, Fan DM. Expression and function of classical protein kinase C isoenzymes in gastric cancer cell line and its drug-resistant sublines. World J Gastroenterol. 2002;8:441-445. [PubMed] |
| 2. | Monden N, Abe S, Hishikawa Y, Yoshimura H, Kinugasa S, Dhar DK, Tachibana M, Nagasue N. The role of P-glycoprotein in human gastric cancer xenografts in response to chemotherapy. Int J Surg Investig. 1999;1:3-10. [PubMed] |
| 3. | Liu B, Staren E, Iwamura T, Appert H, Howard J. Effects of Taxotere on invasive potential and multidrug resistance phenotype in pancreatic carcinoma cell line SUIT-2. World J Gastroenterol. 2001;7:143-148. [PubMed] |
| 4. | Yin F, Shi YQ, Zhao WP, Xiao B, Miao JY, Fan DM. Suppression of P-gp induced multiple drug resistance in a drug resistant gastric cancer cell line by overexpression of Fas. World J Gastroenterol. 2000;6:664-670. [PubMed] |
| 5. | Fan K, Fan D, Cheng LF, Li C. Expression of multidrug resistance-related markers in gastric cancer. Anticancer Res. 2000;20:4809-4814. [PubMed] |
| 6. | Shi Y, Han Y, Wang X, Zhao Y, Ning X, Xiao B, Fan D. MGr1-Ag is associated with multidrug-resistant phenotype of gastric cancer cells. Gastric Cancer. 2002;5:154-159. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 24] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 7. | Shi Y, Zhai H, Wang X, Wu H, Ning X, Han Y, Zhang D, Xiao B, Wu K, Fan D. Multidrug-resistance-associated protein MGr1-Ag is identical to the human 37-kDa laminin receptor precursor. Cell Mol Life Sci. 2002;59:1577-1583. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 33] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 8. | Wang X, Lan M, Shi YQ, Lu J, Zhong YX, Wu HP, Zai HH, Ding J, Wu KC, Pan BR. Differential display of vincristine-resistance-related genes in gastric cancer SGC7901 cell. World J Gastroenterol. 2002;8:54-59. [PubMed] |
| 9. | Zhao Y, You H, Liu F, An H, Shi Y, Yu Q, Fan D. Differentially expressed gene profiles between multidrug resistant gastric adenocarcinoma cells and their parental cells. Cancer Lett. 2002;185:211-218. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 98] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 10. | Fan W, Wang Z, Kyzysztof F, Prange C, Lennon G. A new zinc ribbon gene (ZNRD1) is cloned from the human MHC class I region. Genomics. 2000;63:139-141. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 18] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 11. | Hemming SA, Jansma DB, Macgregor PF, Goryachev A, Friesen JD, Edwards AM. RNA polymerase II subunit Rpb9 regulates transcription elongation in vivo. J Biol Chem. 2000;275:35506-35511. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 73] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 12. | Frick DN, Richardson CC. DNA primases. Annu Rev Biochem. 2001;70:39-80. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 298] [Cited by in RCA: 296] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 13. | Bartsevich VV, Juliano RL. Regulation of the MDR1 gene by transcriptional repressors selected using peptide combinatorial libraries. Mol Pharmacol. 2000;58:1-10. [PubMed] |
| 14. | Xu L, Zhang SM, Wang YP, Zhao FK, Wu DY, Yan X. Relationship between DNA ploidy,expression of ki-67 antigen and gastric cancer metastasis. World J Gastroenterol. 1999;5:10-11. [PubMed] |
| 15. | Wu YA, Lu B, Liu J, Li J, Chen JR, Hu SX. Consequence alimentary reconstruction in nutritional status after total gastrectomy for gastric cancer. World J Gastroenterol. 1999;5:34-37. [PubMed] |
| 16. | Ji F, Peng QB, Zhan JB, Li YM. Study of differential polymerase chain reaction of C-erbB-2 oncogene amplification in gastric cancer. World J Gastroenterol. 1999;5:152-155. [PubMed] |
| 17. | Zhan WH, Ma JP, Peng JS, Gao JS, Cai SR, Wang JP, Zheng ZQ, Wang L. Telomerase activity in gastric cancer and its clinical implications. World J Gastroenterol. 1999;5:316-319. [PubMed] |
| 18. | Ji F, Wang WL, Yang ZL, Li YM, Huang HD, Chen WD. Study on the expression of matrix metallo proteinase-2 mRNA in human gastric cancer. World J Gastroenterol. 1999;5:455-457. [PubMed] |
| 19. | Wang H, Zheng MH, Zhang HB, Zhu J, He JR, Lu AG, Ji YB, Zhang MJ, Jiang Y, Yu BM. Study on incisional implantation of tumor cells by carbon dioxide pneumo peritoneum in gastric cancer of a murine model. World J Gastroenterol. 1999;5:544-546. [PubMed] |
| 20. | Zou SC, Qiu HS, Zhang CW, Tao HQ. A clinical and long-term follow-up study of peri-operative sequential triple therapy for gastric cancer. World J Gastroenterol. 2000;6:284-286. [PubMed] |
| 21. | Cai L, Yu SZ, Zhang ZF. Helicobacter pylori infection and risk of gastric cancer in Changle County,Fujian Province,China. World J Gastroenterol. 2000;6:374-376. [PubMed] |
| 22. | Zhang FX, Zhang XY, Fan DM, Deng ZY, Yan Y, Wu HP, Fan JJ. Antisense telomerase RNA induced human gastric cancer cell apoptosis. World J Gastroenterol. 2000;6:430-432. [PubMed] |
| 23. | Gu QL, Li NL, Zhu ZG, Yin HR, Lin YZ. A study on arsenic trioxide inducing in vitro apoptosis of gastric cancer cell lines. World J Gastroenterol. 2000;6:435-437. [PubMed] |
| 24. | Wang ZN, Xu HM. Relationship between collagen IV expression and biological behavior of gastric cancer. World J Gastroenterol. 2000;6:438-439. [PubMed] |
| 25. | Tu SP, Zhong J, Tan JH, Jiang XH, Qiao MM, Wu YX, Jiang SH. Induction of apoptosis by arsenic trioxide and hydroxy camptothecin in gastriccancer cells in vitro. World J Gastroenterol. 2000;6:532-539. [PubMed] |
| 26. | Jiang BJ, Sun RX, Lin H, Gao YF. Study on the risk factors of lymphatic metastasis and the indications of less in vasive operations in early gastric cancer. World J Gastroenterol. 2000;6:553-556. [PubMed] |
| 27. | Deng DJ. progress of gastric cancer etiology: N-nitrosamides 1999s. World J Gastroenterol. 2000;6:613-618. [PubMed] |
| 28. | Gao HJ, Yu LZ, Bai JF, Peng YS, Sun G, Zhao HL, Miu K, L XZ, Zhang XY, Zhao ZQ. Multiple genetic alterations and behavior of cellular biology in gastric cancer and other gastric mucosal lesions: H.pylori infection, histological types and staging. World J Gastroenterol. 2000;6:848-854. [PubMed] |
| 29. | Miehlke S, Kirsch C, Dragosics B, Gschwantler M, Oberhuber G, Antos D, Dite P, Läuter J, Labenz J, Leodolter A. Helicobacter pylori and gastric cancer: current status of the Austrain Czech German gastric cancer prevention trial (PRISMA Study). World J Gastroenterol. 2001;7:243-247. [PubMed] |
| 30. | Xu AG, Li SG, Liu JH, Gan AH. Function of apoptosis and expression of the proteins Bcl-2, p53 and C-myc in the development of gastric cancer. World J Gastroenterol. 2001;7:403-406. [PubMed] |
| 31. | Cai L, Yu SZ, Zhang ZF. Glutathione S-transferases M1, T1 genotypes and the risk of gastric cancer: a case-control study. World J Gastroenterol. 2001;7:506-509. [PubMed] |
| 32. | He XS, Su Q, Chen ZC, He XT, Long ZF, Ling H, Zhang LR. Expression, deletion [was deleton] and mutation of p16 gene in human gastric cancer. World J Gastroenterol. 2001;7:515-521. [PubMed] |
| 33. | Fang DC, Yang SM, Zhou XD, Wang DX, Luo YH. Telomere erosion is independent of microsatellite instability but related to loss of heterozygosity in gastric cancer. World J Gastroenterol. 2001;7:522-526. [PubMed] |