Published online Jun 15, 1998. doi: 10.3748/wjg.v4.i3.234
Revised: April 20, 1998
Accepted: May 10, 1998
Published online: June 15, 1998
AIM: To search for bacterial DNA sequences in cholesterol gallstones with negative bacterial culture.
METHODS: DNA was extracted from cholesterol gallstones in gallbladders and nested primers polymerase chain reaction (NP-PCR) was used to amplify bacterial gene fragments for identifying the existence of bacteria. The samples of bacterial DNA extracted from potentially causative or unrelated living bacteria were amplified in vitro as the standard markers and comparative 16S ribosomal RNA sequence analysis was made for bacterial identification.
RESULTS: The gallbladder gallstones of 30 patients were analyzed and bacterial DNA was found in 26 patients. Among them, gallstones with cholesterol content between 30%-69% were seen in 5 (5/5) patients, 70%-90% in 11 (11/14) patients, and more than 90% in 10 (10/11) patients. There was no difference either in cholesterol and water content of gallstones or in harboring bacterial DNA of gallstones. E. coli-related DNA fragments appeared in the stones of 8 (26.67%) patients; propionibacteria type DNA in 7 (23.33%); and harbored bacterial gene fragments in 2 patients, similar to Streptococcus pyogenes. A more heterogenous sequence collection was found in 7 (23.33%) patients, which could belong to multiple bacterial infections. Two (6.67%) patients had bacterial DNA with low molecular weight which might be related to some unidentified bacteria.
CONCLUSION: Most cholesterol gallstones harbor bacterial DNA. It is important to determine whether these microorganisms are innocent bystanders or active participants in cholesterol gallstone formation.
- Citation: Wu XT, Xiao LJ, Li XQ, Li JS. Detection of bacterial DNA from cholesterol gallstones by nested primers polymerase chain reaction. World J Gastroenterol 1998; 4(3): 234-237
- URL: https://www.wjgnet.com/1007-9327/full/v4/i3/234.htm
- DOI: https://dx.doi.org/10.3748/wjg.v4.i3.234
Although several factors are known to trigger nucleation and/or growth of cholesterol crystals, the pathogenesis of cholesterol gallstones so far is still unclear. Most studies suggested that cholesterol gallstones may originate mainly from the disturbances of cholesterol metabolism, especially involving the balance between biliary pronucleating and antinucleating proteins[1]. Unfortunately, the role of bacterial etiology was usually neglected. We searched for bacterial DNA sequence in cholesteral gallbladder gallstones with negative bacterial culture by highly sensitive and specific nested primers polymerase chain reaction (NP-PCR) as the molecular genetic evidence of bacterial colonization of cholesterol gallstones.
The gallbladder gallstones were consecutively collected from October to December 1995. Patients with positive aerobic or anaerobic bile cultures were excluded. The group for analysis consisted of 5 men and 25 women with a mean age of 54.5 years, ranging from 27-73 years. All of them had symptomatic gallstone disease and underwent selective surgery. Fresh gallbladder bile samples were obtained by needle puncture aspiration during surgery. All stones had diameters of more than 5 mm and stored at -20 °C.
DNA extraction. DNA (3.1 μg-26.2 μg) was obtained from stone samples by the special purification methods using an UV-240 spectrophotometer (Shimadzu Corporation, Japan). Stone samples of about 300 mg were crushed in a 5 mL Eppendorf tube with a glass rod and incubated with 600 μL 1% sodium dodecyl sulfate rotating overnight at 20 °C. Lithium chloride solution (7 mol/L) was added to a final concentration of 1.5 mol/L for precipitating the interfering substances in gallstones. Bacterial DNA was extracted from potentially causative or unrelated living bacteria including E.coli, Propionibacterium acnes, Streptococcus pyogenes, Pseudomonas aeruginosa, Staphylococcus aureus and Serratia odorifera. All of them were ATCC strains. The DNA was then extracted by the conventional purification methods, resuspended in 50 μL TE buffer (Tris/EDTA), and stored at -4 °C.
NP-PCR amplification According to the literature[2], primers sequences were designed for amplifying bacterial 16S ribosomal RNA gene in vitro, including a generic 5’primer set p1: 5’-AGAGTTTGAT (c/t) (c/a) TGGCTCAG-3’, P2: 5’-ACTAC (c/t) (a/c/g) GGGTATCTAA (g/t) CC-3’ and a new nested 3’primer P3: 5’-ACCGC (g/t) (a/g) CTGCTGGCAC (Institute of Cellular Biology, Chinese academy of Sciences, Shanghai).
The PCR was done in 0.5 mL polyproylene microcentrifuge tube with a PTC-100TM Programmable Thermal Controller (MJ Research, INC., USA). The total incubation volume was 25 μL, containing 100 mmol/L “tris”-hydrochloric acid, pH 8.3, 500 mmol/L potassium chloride, 15 mmol/L magnesium chloride, 100 μmol/L of each dNTP, 0.2 μg-1.0 μg template DNA and 1 unit of Taq DNA polymerase (Sino-American Biotec Engineering, China). The reaction mixture was covered with 1 drop of mineral oil. In the first round of amplification, 0.2 pmol each of the outer primers (P1, P2), was used then the reaction tubes were supplemented with 0.2 pmol each of the inner primers (P1, P3), and 1 μL amplified DNA and 1 unit of Taq DNA polymerase were used in the second round. The PCR cycle took place at 94 °C for 30 s, 56 °C for 1 min, and 72 °C for 1 min 40 s. During the last cycle, the 72 °C step was extended to 8 min. Thirty-five cycles were carried out for each reaction.
NP-RCR product comparative analysis The sequence of 16S ribosomal RNA gene fragments amplified by NP-PCR was approximately 500-base pairs. Lambda DNA/EcoR I + Hind III (Promega Corporation, USA) was used as molecular standard. Five μg reamplified DNA was applied to a 1.4% agarose gel containing 0.8 g/L ethidium bromide and after electrophoresis 4.0 V/cm, 40 min was visualised with an ultraviolet light source. The reamplified DNA of ATCC bacteria was included in every run. The agarose gels from the NP-PCR procedure were then subjected to computer image system for further bands comparative analysis. If the NP-PCR of the first gallstone (chosen at random) was positive, the rest of the stones of a given patient were usually not analyzed. Otherwise, all gallstones or at least five stones would be analyzed.
Cholesterol and water content analysis DNA isolated from a part of a same gallstone was used for NP-PCR amplification and another part for cholesterol and water content analysis. Accurate 100 mg of nativeg allstone material was dried at 105 °C for 3 h to a constant weight. The cholesterol was extracted by a modified chemical method and underwent photometric test using a 722 Spectrophotometer (Shanghai Third Analysis Instrument Factory). The cholesterol and water content was analyzed according to the manufacturer’s instructions.
Statistical analysis All values were expressed as x-±s. The relationship between cholesterol or water content and NP-PCR results were analyzed with Wilcoxon rank sum test.
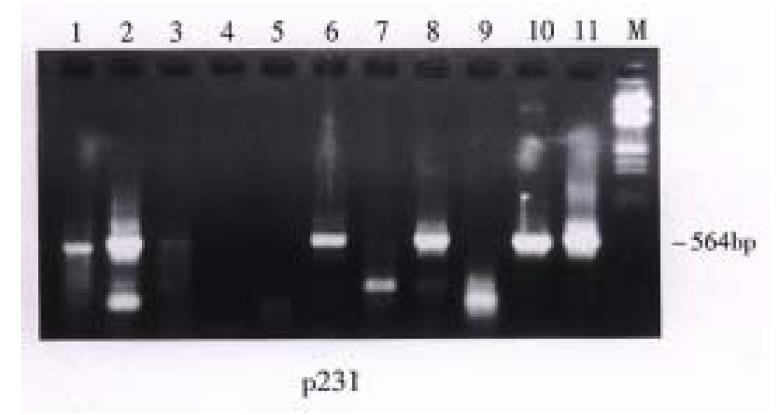
The gallstones of 30 patients were analyzed. Bacterial 16S ribosomal RNA gene fragments were amplified in the stones of 26 patients. Amplification was not possible in the remaining 4 patients whose stones had no detectable bacterial DNA (Figure 1). In the 26 patients, the positive gallstones contained cholesterol content between 30%-69% in 5 (5/5) patients, 70%-90% in 11 (11/14) patients, and more than 90% in 10 (10/11) patients. The water content of gallstones changed from 10.3% to 52%. The group with bacterial DNA in the gallstones had water content of 26.5% ± 10.39% while the group without bacterial DNA had water content of 19.18% ± 4.88%. There was no difference either in cholesterol and water content or in harboring bacterial DNA of gallstones (P > 0.05) (Table 1, Table 2). E.coli related DNA fragments were found in the stones of 8 (26.67%) patients. Propionibacteria-type DNA was obtained in 7 (23.33%) patients. Stones of 2 (6.67%) patients harbored bacterial gene fragments similar to Streptococcus pyogenes. Simultaneously, multiple heterogenous sequence collections were found in 7 (23.33%) patients, which could belong to the mixed infections including E.coli, Propionibacterum acnes and Streptococcus pyogenes or some other unidentified bacteria such as Clostridium difficile. The stones of the other 2 (6.67%) patients had bacterial DNA with lower molecular weight and could be related to some unidentified bacteria.
| Cholesterol content (%) | Harbor bacterial DNA | No harbor bacterial DNA | Total | |||
| Case | % | Case | % | Case | % | |
| 30-69 | 5 | 16.67 | 0 | 0 | 5 | 16.67 |
| 70-90 | 11 | 36.66 | 3 | 10.0 | 14 | 46.67 |
| > 90 | 10 | 33.33 | 1 | 33.33 | 11 | 36.66 |
| Group | Case | Water content (%) |
| Harbor bacterial DNA | 26 | 26.57 ± 10.39 |
| No harbor bacterial DNA | 4 | 19.18 ± 4.88 |
Few previous studies considered the role of bacteria in the pathogenesis of cholesterol gallstone as there was no direct evidence of bacterial existence, because of the failure in the bacteria culture from cholesterol stones. However, the process in gallbladder gallstone formation may take a longer time, and the embedded bacteria may be destroyed or killed. PCR technique has a high sensitivity and specificity as compared with the bacterial routine culture. Theoretically, PCR can detect a copy gene of a bacterium or cell. The nested primer approach increases the specificity as well as sensitivity of PCR[3]. To prevent artifacts resulting from contamination, gallbladder stones were collected and prepared with stern sterilization, control experiments omitting template DNA were made in parallel, and pre and post-PCR procedures were performed in different rooms . Furthermore, primers were designed according to bacterial genes coding for 16S ribosomal RNA . The structure of these ribosomal RNA genes is highly specific for different bacteria.
Bacteria can invade the biliary tract by ascending from the duodenum and via the hematogenous route from the hepatic portal venous blood[4]. Many aerobe, facultative aerobe and anaerobe species such as E.coli, P.vulgaris, B.fragilis, Clostridium perfringens, can produce β-glucaronidase that catalyzes the hydrolysis of bilirubin conjugates in bile, leading to increased amounts of unconjugated bilirubin, which precipitates as calcium bilirubinate stones . Bacteria could also disturb gallbladder wall secretion and cause acute or chronic inflammation with resultant precipitation of bile. Bacteria have adhesive properties and can produce glycocalyx or other matrices within which the pigment or cholesterol coalesces. In fact, most of cholesterol gallstones have a pigment nucleus or section. It seems likely that bacterial infection serves as an initiating factor or plays other important roles in the development of gallstones. Besides most bacteria that cause biliary infection can produce palmitate via a phospholipase, and result in precipitation of calcium palmitate. Meanwhile, the reduction of lecithin in bile leads to a lower solubility of bilirubin and promotes banding of bilirubin and calcium. Bacterial phospholipase, on the other hand, degrades lecithin which will reduce the solubility of cholesterol and promote cholesterol nucleation in bile.
Propionibacterium acnes is a gram-positive anaerobic rod bacterium which is a normal bacterial colony usually located on the skin and mucosa. The role of Propionibacterium acnes in gallstone formation has not been clarified. Up to the present, the possible pathogenic mechanisms are[5]: ① This bacterium produces a nonspecific extracellular lipase that catalyses the total hydrolysis of triacylglycerols to free fatty acids and glycerol and that is able to use lecithin as a substrate. ② Propionibacterium acens- also produces a phospholipase C, which is a promoter of cholesterol procipitation[6]. ③ Propionibacterium can bind to lipids[7], and the binding is increased as lipid hydrophobicity increases. This ability may permit Propionibacterium acnes to bind biliary lipid vesicles and degrade them with the extracellular lipolytic enzymes. The nonspecific lipase is a chemotactic agent for neutrophils, and activation of neutrophils would result in the release of oxygen radicals capable of causing phospholipid perioxidation, which reduces the stability of mixed micellar systems containing cholesterol, lecithin, and cholate. ④ Propionibacterium acnes has specific affinity for the calcium containing regions of gallstones and can promote calcium salt procipitation. Once the calcium bilirubinate salt and calcium palmitate salt were formed, the bacterial colony would secrete sufficient lipolytic enzymes to promote cholesterol nucleation.
We found that most cholesterol gallstones harbor bacterial DNA with the highly sensitive, specific and reproducible NP-PCR technique. Bacteria and their metabolites might play an important role in the formation of cholesterol gallstones with different mechanisms. Obviously, the study searching for molecular genetic evidence of bacterial conolization of cholesterol gallstones is only at the initial stage. A lot of work remains to be done before it can be decided whether these microorganisms have a cause and effect relationship in cholesterol gallstone formation and an infectious etiology can be established. We are now determining heterologous sequence collections with the dideoxy chain termination technique.
Project supported by the State Education Commission Research Foundation for Scientists Returning from Abroad (1997)436.
| 1. | Holzbach RT, Busch N. Nucleation and growth of cholesterol crystals. Kinetic determinants in supersaturated native bile. Gastroenterol Clin North Am. 1991;20:67-84. [PubMed] |
| 2. | Swidsinski A, Ludwig N, Pahlig H et al. Molecular genetic evidence of bac-terial colonizations of cholesterol gallstones. Gastroenterology. 1995;108:860-864. [RCA] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 54] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 3. | Aurelius E, Johansson B, Sköldenberg B, Staland A, Forsgren M. Rapid diagnosis of herpes simplex encephalitis by nested polymerase chain reaction assay of cerebrospinal fluid. Lancet. 1991;337:189-192. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 398] [Cited by in RCA: 372] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 4. | Wu XT, Xiao LJ. The progress of studies on the role of bacteria in the forma-tion of gallstone. Foreign Medicine (Surgery). 1996;23:277-279. |
| 5. | Soloway RD, Crowther RS. Bacteria and cholesterol gallstones: molecular biology comes to gallstone pathogenesis. Gastroenterology. 1995;108:934-936. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 6. | Pattinson NR, Willis KE. Effect of phospholipase C on cholesterol solubilization in model bile. A concanavalin A-binding nucleation-promoting factor from human gallbladder bile. Gastroenterology. 1991;101:1339-1344. [PubMed] |
| 7. | Gribbon EM, Cunliffe WJ, Holland KT. Interaction of Propionibacterium acnes with skin lipids in vitro. J Gen Microbiol. 1993;139:1745-1751. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 103] [Cited by in RCA: 96] [Article Influence: 3.0] [Reference Citation Analysis (0)] |