Published online Apr 14, 2021. doi: 10.3748/wjg.v27.i14.1406
Peer-review started: February 1, 2021
First decision: February 27, 2021
Revised: March 13, 2021
Accepted: March 19, 2021
Article in press: March 19, 2021
Published online: April 14, 2021
Processing time: 66 Days and 20.6 Hours
Gastrointestinal (GI) symptoms have been described in a conspicuous percentage of coronavirus disease 2019 (COVID-19) patients. This clinical evidence is supported by the detection of viral RNA in stool, which also supports the hypothesis of a possible fecal-oral transmission route. The involvement of GI tract in severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection is corroborated by the theoretical assumption that angiotensin converting enzyme 2, which is a SARS-CoV-2 target receptor, is present along the GI tract. Studies have pointed out that gut dysbiosis may occur in COVID-19 patients, with a possible correlation with disease severity and with complications such as multisystem inflammatory syndrome in children. However, the question to be addressed is whether dysbiosis is a consequence or a contributing cause of SARS-CoV-2 infection. In such a scenario, pharmacological therapies aimed at decreasing GI permeability may be beneficial for COVID-19 patients. Considering the possibility of a fecal-oral transmission route, water and environmental sanitation play a crucial role for COVID-19 containment, especially in developing countries.
Core Tip: Coronavirus disease 2019 (COVID-19) patients may suffer from gastrointestinal symptoms that are associated with gastrointestinal dysbiosis. Even though the exact role of gut microbiome perturbation as a either a cause or a consequence of the disease is still to be elucidated, pharmacological interventions aimed at containing intestinal permeability may be of support in COVID-19 patients.
- Citation: Troisi J, Venutolo G, Pujolassos Tanyà M, Delli Carri M, Landolfi A, Fasano A. COVID-19 and the gastrointestinal tract: Source of infection or merely a target of the inflammatory process following SARS-CoV-2 infection? World J Gastroenterol 2021; 27(14): 1406-1418
- URL: https://www.wjgnet.com/1007-9327/full/v27/i14/1406.htm
- DOI: https://dx.doi.org/10.3748/wjg.v27.i14.1406
In December 2019, in Wuhan, one of the largest cities in the Chinese province of Hubei, a new coronavirus appeared that was responsible for a pneumonia epidemic that was named by the World Health Organization (WHO) on February 11, 2020 as coronavirus disease 2019 (COVID-19), and declared as a dangerous threat to public health and an international health emergency[1,2]. Indeed, COVID-19 infections rapidly spread and was declared a pandemic by the WHO one month later on March 11, 2020. In a short time, the pandemic involved all continents, and at the time of writing this review, accounts for more than 100 million confirmed cases worldwide, also becoming a serious threat to the global economy[3,4].
At the beginning of January 2020, the new coronavirus responsible for this pandemic was named “severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)” by the International Committee on Taxonomy of Viruses[5]. It is a single-stranded RNA virus with positive polarity and is closely related to the B-line of beta-coronaviruses, which are typically responsible for severe and lethal respiratory syndromes[2,6]. Phylogenetic analysis showed that the viral sequence of this new coronavirus has a strong similarity (89%) with two other coronaviruses similar to SARS and deriving from bats, namely bat-SL-CoVZC45 (GenBank access n. MG772933.1) and bat-SL-CoVZXC21 (GenBank access n. MG772934.1). Similarities were also found with SARS-CoV, the virus that caused the SARS pandemic in 2002/2003, and the Middle East Respiratory Syndrome (MERS-CoV) pandemic of 2012, with which SARS-CoV-2, shares sequence homologies (79% with SARS-CoV and 50% with MERS-CoV)[7-9] as well as zoonotic transmission and several clinical features.
Studies carried out so far suggest that the human angiotensin converting enzyme 2 (ACE2) is the main target receptor for the virus to gain access into human host cells. Binding occurs between the ACE2 binding domain located at the C-terminal end of the viral spike protein and the ACE2 target receptor located on the surface of the host cell membrane[10]. After binding to ACE2 receptors on human cells[11], the SARS-CoV-2 spike protein is cleaved by two mucosa-specific serine proteases, namely TMPRSS2 and TMPRSS4, to facilitate viral cellular infection[12].
Tissues with the highest expression of ACE2, and therefore a high sensitivity to infection are present in both the respiratory and the gastrointestinal (GI) tracts. The receptor is present in the pulmonary epithelium, in type 2 pneumocytes that are present in the alveoli where the exchange between oxygen and carbon dioxide takes place, in the nose, mouth, stratified epithelial cells in the upper esophagus, and absorbent enterocytes of the ileum and colon. Other receptors that are known to allow entry of SARS-CoV-2 are DC-SIGN (CD209), CD147 and L-SIGN (CD209L)[13-15]. Transmission of the virus can take place through direct contact with respiratory droplets from infected subjects or indirectly from contaminated objects and surfaces[16].
Clinical manifestations associated with COVID-19 infection range from complete absence of symptoms (asymptomatic subjects) to serious and even fatal infections. In most cases, the disease is mild to moderate, so that symptoms can be managed safely at home without the need for hospitalization. The main target of the infection is the respiratory tract, and therefore, the clinical manifestations associated with the disease are mostly respiratory. The most common symptoms are similar to those of seasonal flu and include fever, dry cough, dyspnea, sore throat, headache, muscle aches, and asthenia. Severe respiratory distress, chest pain, pneumonia, and pulmonary insufficiency can also be found in severe cases[17-19]. In SARS-CoV-2 pneumonia, lung computed tomography scans shows bilateral, subpleural ground-glass opacity lesions[20-22]. In addition to pulmonary clinical manifestations, heart damage can also occur in infected patients. Shi et al[23] in a study conducted in Wuhan on 400 patients, found that about one-fifth experienced heart disease as a risk factor for worse outcome. Heart damage essentially consists of sudden and severe myocardial inflammation leading to heart failure and arrhythmias[24]. For this reason, hypertensive patients, and in general all individuals with a positive history of cardiovascular disease, are more at risk of experiencing a poor prognosis than other patients[25]. SARS-CoV-2 can also damage the central nervous system, with symptoms that range from mild (anosmia, dysgeusia) to severe (viral encephalitis). Moreover, COVID-19 infection, like other pulmonary infectious diseases, may represent a risk factor for acute cerebrovascular events. The observed D-dimer elevation in COVID-19 patients may indicate additional risk of cerebrovascular events[26]. Kidney and liver damage have also been reported. Proteinuria, hematuria, increased aspartate transaminase (AST, also known as aspartate aminotransferase or glutamic oxaloacetic transaminase), alanine aminotransferase (ALT, also known as alanine transaminase or glutamic-pyruvic transaminase), and bilirubin have been found in about half of infected patients[27,28].
The virulence of microorganisms determines the damage to host cells before the microorganisms are cleared by host immunity[29]. Yet, the virulence of the infective microorganism does not fully account for the virulence of the disease, the majority of which results from inflammation produced by the immune response[29]. That has been demonstrated for many infective diseases and is particularly true for SARS-CoV-2. Most viral and bacterial infections in humans are self-limiting[29,30], and infecting microorganisms become commensal if they can coexist with the human host. The SARS-CoV-2 virus infection is no different[31], and that could explain why most COVID-19 cases are asymptomatic or mild.
Although, as reported, SARS-CoV-2 infection mainly affects the respiratory system[32], leading to breathing difficulties, dry cough, and nasal congestion to respiratory failure, this novel coronavirus can be found in the GI tract as well[33]. Furthermore, SARS-CoV-2 RNA has been isolated from stools. Viral nucleocapsid protein can be observed in duodenal and rectal glandular epithelial cells by laser scanning confocal microscopy. The available results provide evidence of the activity of this virus in the GI tract[34].
GI symptoms are present in approximately 50% of COVID-19 patients, consisting mainly of diarrhea, nausea, vomiting, and abdominal discomfort[35]. Moreover, several studies have shown that SARS-CoV-2 is able to interact with ACE2 receptors on ileal enterocytes and colon epithelial cells, so that a trophism for the GI tract has been hypothesized as well[27,36]. Further evidence supporting the possibility that both the small and large intestines may be susceptible to SARS-CoV-2 infection, was the isolation of viral RNA from the GI epithelium and staining of the viral nucleocapsid protein in these cells[34]. In addition, viral RNA was also isolated in stool samples from COVID-19 patients, and it was presumed that the viral components originated from infected enterocytes[34,36]. Considering the overall evidence, several reports have expressed concerns of possible fecal-oral transmission of SARS-CoV-2[37,38]. Nevertheless, the findings were not unexpected as other well-known coronaviruses, including those causing the MERS and SARS epidemics, have shown to infect enterocytes, leading to several GI symptoms[39].
Several reports have focused on the GI symptomatology in COVID-19 patients, underlining clinical outcomes that varied widely in terms of frequency. Nine of those studies that included a total of 4177 patients found that the incidence of GI symptoms varied from 4.9% to 61.3%. The most frequent symptoms were diarrhea (8.4%) and lack of appetite (7.4%); 4.1% of patients complained about nausea-related disorders, while vomiting was reported in 2.4% and abdominal pain in 1.7% of the cases[40-48]. Interestingly, the frequency of diarrhea, nausea, and abdominal pain in COVID-19 patients was shown to be higher among women than men[49]. Finally, Yang et al[50] examined 50 COVID-19 patients by dividing them in two groups by the presence of either initial pulmonary or initial GI symptoms. They found that patients with GI manifestations as initial symptoms had more severe clinical outcomes, leading to longer hospitalization.
As stated above, and similar to SARS-CoV-1, SARS-CoV-2 infection is mediated by ACE2, which allows the virus to enter the cells and replicate by taking advantage of the host synthesis apparatus[51]. A recent study has shown that the binding affinity of the spike protein of SARS-CoV-2 is about 10-20 times higher than that of SARS-CoV-1[52]. ACE2 receptors are widely expressed in the epithelia of both the respiratory and GI tracts. A recent multiomic in vitro study has shown that expression of these membrane enzymes is higher in intestinal cell lines than lung cell lines[36].
The pathophysiology of GI symptoms is not entirely clear, but it seems that they originate as a result of several phenomena. ACE2 performs an essential function in the intestine by regulating amino acid homeostasis and microbiome balance[53]. It is possible that the binding of SARS-CoV-2 to these membrane receptors reduces their availability, giving rise to an alteration of physiological function that induces a dysbiosis that leads to diarrhea, one of the most frequent symptoms reported during SARS-CoV2 infection[54]. The development of a cytokine storm could be indeed the cause of damage to the GI tract. Autopsies of SARS patients found increased expression of proinflammatory cytokines in ACE2-expressing cells compared with those that do not express this enzyme on their membranes[55]. It is fair to think that this happens following SARS-CoV-2 infection. Inflammation could also stimulate the gut microbiota to release molecules that increase the inflammatory state of the intestine and subsequently spread from the intestine into the circulatory system to cause systemic damage with more severe consequences than the viral infection itself[56]. In addition to locally occurring phenomena, the gut-lung axis seems to have a pivotal role in the development of GI symptoms as well. This axis represents a link between two organs that share a common mucous immune system[57]. Its influence has been hypothesized as a result of studies that have described COVID-19 patients with GI symptoms but without SARS-CoV-2 RNA-positive stool samples[54].
The impact of GI comorbidities on the development of symptoms in COVID-19 patients is yet to be defined. While the relationship between disorders such as diabetes, cardiovascular diseases, and hypertension and COVID-19-related severity was proven by several studies[58]. There are not yet sufficient studies to demonstrate a causal link between the presence of previous GI conditions and the onset of COVID-19-related symptomatology.
Indeed, existing studies do not provide indisputable results. A study carried out in China found that pre-existing GI disorders represent a third comorbidity in COVID-19 patients[59]. Another study carried out in Wuhan reported a relatively high percentage of COVID-19 patients with GI disorders[60]. Conversely, other studies carried out in the United States[61] and in Italy[62] have not found a significant correlation between SARS-CoV-2 infection and the presence of inflammatory bowel disease. On the contrary, Papa et al[63] observed a better clinical outcome in patients presenting with digestive comorbidities, as presumably they do not hinder the immune response against SARS-CoV-2. However, the authors point out that the number of cases observed was small[63].
A potential fecal-oral transmission route of SARS-CoV-2, in addition to respiratory droplets, person-to-person contact, or indirect infection following contact with contaminated surfaces, has been proposed by several authors[37,38,64,65]. The presence of common GI symptoms in the COVID-19 patients mentioned above is evidence that the GI tract is involved in SARS-CoV-2 infection.
Viral RNA has been detected in esophagus, stomach, duodenum, and rectum tissue from patients with severe SARS-CoV-2 infections[66]. Notably, the first COVID-19 patient in the United States tested positive for SARS-CoV-2 in loose stool specimens by real-time reverse transcription polymerase chain reaction[67]. Several studies report detection of viral RNA in stool samples in adults[34,68-72] and children[73-75] diagnosed with COVID-19. Interestingly, two studies confirmed that SARS-CoV-2 was detectable in fecal samples even after negative testing of respiratory swabs[70,72], and in patients who did not present GI symptoms[66,69,71]. In addition to SARS-CoV-2 RNA detection in stool samples, positive staining of viral nucleocapsid protein in gastric, duodenal, and rectal epithelia might demonstrate the infection of GI glandular epithelial cells[34]. Wang et al[68] found a high content of viral RNA in fecal specimens from COVID-19 patients, and electron microscopy confirmed the presence of living viral particles in those samples. Wölfel et al[69] could not isolate living SARS-CoV-2 from stool samples despite the presence of high concentrations of viral RNA, but did detect cells in a few stools that contained subgenomic mRNA, which suggests active replication of the virus in the intestinal tract. The presence of viral subgenomic mRNA indicates actively infected cells because such mRNA is transcribed only in infected cells and is not found packaged into virions[69]. As in adults, SARS-CoV-2 in children might persist in the GI tract longer than in the respiratory system. Actually, a high percentage of children diagnosed of COVID-19 tested positive on rectal or anal swabs, and some were positive after nasopharyngeal or throat swabs became negative[73-75].
Other evidence that supports an alternative fecal-oral transmission of COVID-19 is the expression of SARS-CoV-2 entry-genes in the cells of intestinal tissues. Mucosal cells of the lower intestine can be infected by other coronaviruses, resulting in diarrhea and other enteric symptoms. In this case, SARS-CoV-2 could be carried by saliva and secretions into the digestive tract, where it would infect the ACE2-expressing enterocytes. It has been confirmed that SARS-CoV-2 can infect enterocytes present in human small intestine organoid models, which supports virus replication and a GI route of infection[76]. However, another study demonstrated that human GI secretions were able to inactivate SARS-CoV-2, which suggests that although virus infection and pathogenesis is enhanced in the intestine, it is not clear if it would be a primary site of infection and possible route of transmission[77]. An alternative route of transmission that acts together with the respiratory route might explain the rapid spread of disease.
COVID-19 symptomatology is strictly related to both a physiological and an aberrant activation of immune system, leading in very severe cases to a cytokine storm[78]. Based on this premise, it is not surprising that the gut microbiome, which is closely linked to the immune status of the host, may play a role in COVID-19 severity. A gut microbiome imbalance has been associated in COVID-19 patients with elevated concentrations of inflammatory cytokines, and blood markers, including C-reactive protein (CRP), lactate dehydrogenase, and aspartate aminotransferase[79]. Yeoh et al[79] examined blood and fecal samples from 100 young (mean age of about 36 years) confirmed positive COVID-19 patients in two Hong Kong hospitals. They also analyzed samples from patients for up to 30 days after clearance of SARS-CoV-2. All patients had slight or moderate symptomatology, with only 5% with severe and 3% with critical conditions. Compared with a group of healthy adult controls, COVID-19 patients had higher fecal populations of Ruminococcus gnavus, Ruminococcus torques, and Bacteroides dorei species. This imbalance seemed not to depend on the pharmacological therapy administered. The same investigators reported a stratification of gut microbiota composition associated with disease severity in hospitalized COVID-19 patients. Cytokine (IL-10 and tumor necrosis factor-), enzyme (AST, -glutamyl transferase, lactate dehydrogenase), CRP, N-terminal proB-type natriuretic peptide levels, and erythrocyte sedimentation rate were previously reported as associated with increased COVID-19-related symptomatology[80-82]. Those parameters were also significantly associated with microbiota composition in COVID-19 patients. Remarkably, the levels of those molecules increased in parallel with changes in the microbiota composition, representing more severe disease states[79]. Overall, the results suggest that the gut microbiota composition might be linked to the extent of the immune response to COVID-19 and subsequent tissue damage, thus playing a role in regulating disease severity. The persistence of gut microbiota dysbiosis even 30 days after clearance of SARS-CoV-2 might also be linked to the persistence of COVID-19 related symptoms, such as fatigue, dyspnea, and joint pain.
Moreover, Yu[83] suggested that nutritional disorders impair immunity, causing hyperinflammation and leading to an overload of cytokines in COVID-19 patients. Autophagy induced by restrictive eating could be an efficient strategy to prevent COVID-19-related symptomatology. Indeed, it inhibits overgrowth of the microbiota and partially reduces excessive nutrition for replication. Autophagy also attenuates inflammation. On that basis, Yu[83] recommended restoration of good nutritional status together with restriction of food intake, especially in older subjects, to reduce the risk of developing severe symptomatology if infected by COVID-19.
On the other hand, as inflammation can be considered as a physiological response of the immune system to damaged tissues[84], it is a protective reaction to remove the injurious stimuli and to initiate the healing of damaged tissues[84]. Nutrition excess slows tissue healing, indeed nutrition derived from the degradation of the damaged tissue coupled with excess nutrition intake will be mostly turned into lipid intermediates and deposited in nonadipose tissue, lipotoxicity and further tissue damage. Thus, in a state of overnutrition, inflammation can become chronic. In this regard, undernutrition might be beneficial in fighting viral infection, as anorexia nervosa patients seem to be free of common viral infections[85,86].
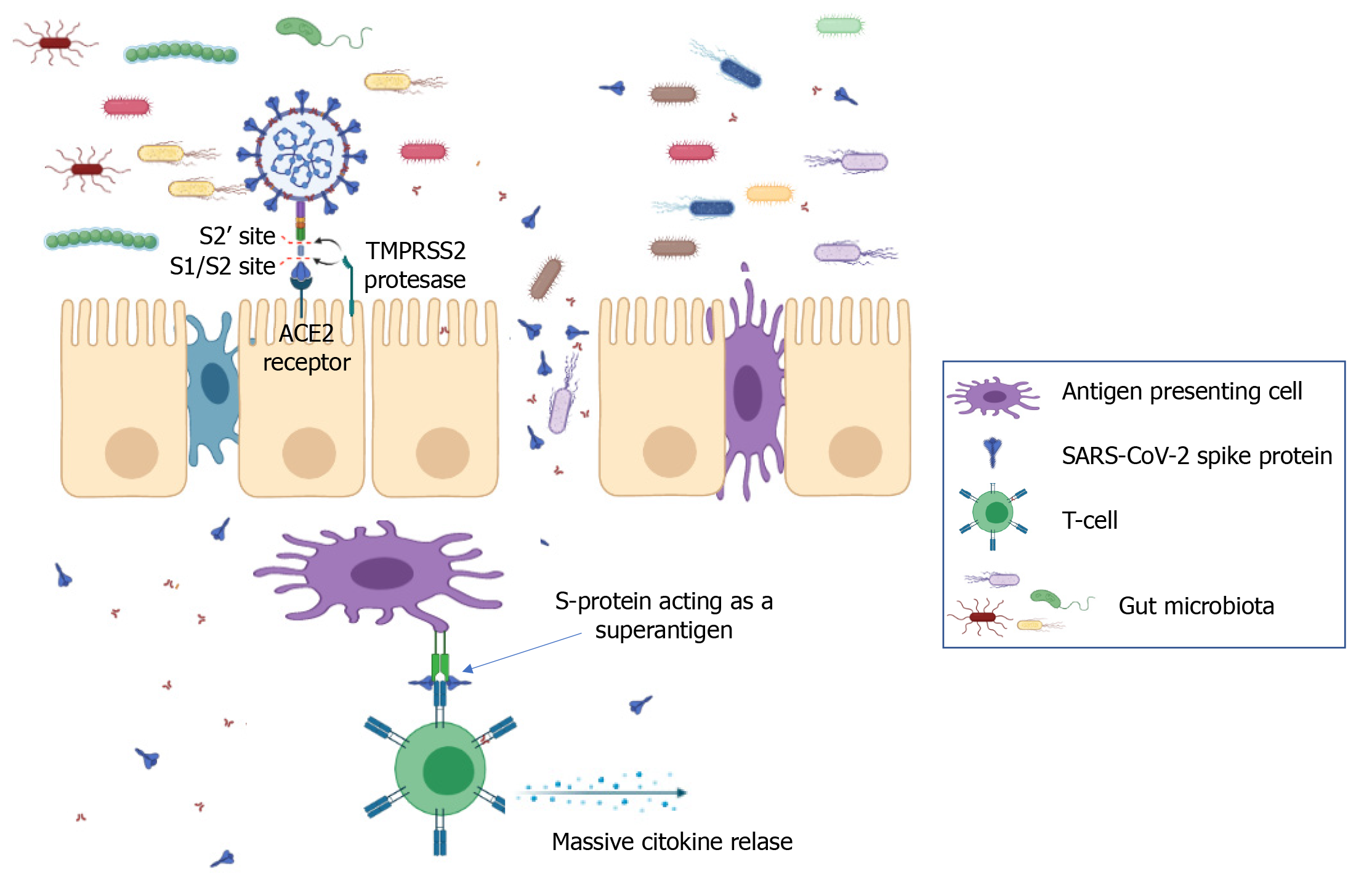
Multisystem inflammatory syndrome in children (MIS-C) is defined by the Centers for Disease Control and Prevention[87] as a severe illness requiring hospitalization that occurs in individuals < 21 years of age with evidence of current or recent SARS-CoV-2 infection, or recent exposure to an individual with COVID-19. Children with MIS-C have fever, laboratory evidence of inflammation, and multiorgan involvement without an alternative plausible diagnosis. Ninety-two percent of children with MIS-C report GI symptoms[88] and 80% develop cardiac pathology. MIS-C is characterized by a superantigen-mediated hyperinflammatory response with a unique T-cell profile and multiple autoantigens[89-91]. It has been recently shown that the spike protein of SARS-CoV-2 contains sequence and structural motifs similar to a Staphylococcal Enterotoxin B superantigen[92], and superantigen-induced skewing of the T-cell repertoire in MIS-C patients with expansion of the T-cell receptor beta variable gene correlates with the cytokine storm and hyperinflammation seen in MIS-C[91]. Inflamed expansion of antibodies for multiple non-COVID-19 pathogens, including common coronaviruses, influenza and respiratory syncytial virus[93] plus numerous self-antigens[89,90] also correlates with the severe cardiac complications of MIS-C. Immune complexes stimulate monocytes/macrophages, eliciting phagocytosis, macrophage activation and cytokine storm[94,95] (Figure 1). The instigating factor for MIS-C has not been elucidated, but prominent GI symptoms suggest an intestinal source. The upper or lower respiratory tract are unlikely sources of antigens triggering this hyperinflammatory response, given the relative lack of respiratory symptoms in MIS-C, low/undetectable viral load in respiratory secretions[93], and the 2-4 wk delay between initial infection/exposure and development of MIS-C. Rather, ongoing antigen exposure is much more likely to be from the GI tract as has been shown in Kawasaki disease (KD), a similar illness with a hyperinflammatory profile leading to fever, rash, elevated inflammatory markers resulting in vasculitis, coronary aneurysms, and cardiac pathology. In KD, increased intestinal permeability secondary to upregulation of zonulin, a molecule that regulates paracellular molecule trafficking, has been associated with increased circulating immunoglobulin A and cardiac complications in children with KD[96].
A study by Yeoh et al[79] was not designed to determine whether dysbiosis is the consequence of patient immune response rather than a contributing cause in disease onset and severity. Nevertheless, the reported association seems to be sufficiently robust to support the benefit of pharmacological intervention aimed at decreasing gut permeability for COVID-19 GI symptoms management, and potentially to reduce the risk of its complications, including MIS-C. Such a strategy could both reduce GI tract inflammation and help accelerate naturally occurring dysbiosis correction. Larazotide acetate, a synthetic octapeptide designed as a specific zonulin receptor antagonist, was successfully and safely used to modulate gut permeability and the related inflammation[97,98]. Di Micco et al[99] recently proposed an additional role for larazotide in COVID-19 patients. Based on an in-silico evaluation, they reported the ability of larazotide to inhibit the N3 protease of SARS-CoV-2. As larazotide was specifically designed as an oral drug able to reach the lower GI tract, it might be useful both for its direct anti-SARS-CoV-2 effect and to prevent antigen trafficking of SARS-CoV-2-derived molecules, including spike protein from the gut lumen to the bloodstream, which could trigger a cytokine storm leading to MIS-C.
In summary, whether digestive symptoms are a direct response to SARS-CoV-2 infection of the GI tract, or a secondary outcome of COVID-19, is still to be completely understood. The presence of viral RNA in the GI tract might partially explain GI symptoms even if it does not mean that there is infection in the intestine. In either case, robust evidence supports GI tract involvement in COVID-19 as well as a potential fecal-oral transmission route. The GI seems to be not only a reservoir of SARS-CoV-2, but also an active battlefield in which virus, innate and adaptive immune system responses, epithelial cells, and gut microbiota play critical roles. The interventions for managing GI-related issues should be twofold. From a patient perspective, pharmacological treatment to reduce the severity of GI symptoms and support a quick re-establishment of eubiosis should be evaluated. Moreover, interventions to ensure water sanitation should be added to the current policies for COVID-19 pandemic control. Considering that many people worldwide lack access to clean drinking water and good sanitation, the possible containment of COVID-19 through water treatment should be carefully taken into account to reduce the actual and future magnitude of the pandemic.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Italy
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Merrett ND S-Editor: Gao CC L-Editor: Filipodia P-Editor: Wang LL
| 1. | Wu YC, Chen CS, Chan YJ. The outbreak of COVID-19: An overview. J Chin Med Assoc. 2020;83:217-220. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 878] [Cited by in RCA: 719] [Article Influence: 143.8] [Reference Citation Analysis (0)] |
| 2. | Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W; China Novel Coronavirus Investigating and Research Team. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020;382:727-733. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18987] [Cited by in RCA: 17616] [Article Influence: 3523.2] [Reference Citation Analysis (0)] |
| 3. | Jin Y, Yang H, Ji W, Wu W, Chen S, Zhang W, Duan G. Virology, Epidemiology, Pathogenesis, and Control of COVID-19. Viruses. 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1055] [Cited by in RCA: 847] [Article Influence: 169.4] [Reference Citation Analysis (0)] |
| 4. | World Health Organization. Coronavirus Disease (COVID-19) Pandemic, 2020. [cited January 26, 2021]. In: World Health Organization [Internet]. Available from: https://www.who.int/emergencies/diseases/novel-coronavirus-2019. |
| 5. | Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol. 2020;5:536-544. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5202] [Cited by in RCA: 4628] [Article Influence: 925.6] [Reference Citation Analysis (0)] |
| 6. | Chan JF, Kok KH, Zhu Z, Chu H, To KK, Yuan S, Yuen KY. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microbes Infect. 2020;9:221-236. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1813] [Cited by in RCA: 1949] [Article Influence: 389.8] [Reference Citation Analysis (0)] |
| 7. | Jiang S, Du L, Shi Z. An emerging coronavirus causing pneumonia outbreak in Wuhan, China: calling for developing therapeutic and prophylactic strategies. Emerg Microbes Infect. 2020;9:275-277. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 237] [Cited by in RCA: 220] [Article Influence: 44.0] [Reference Citation Analysis (0)] |
| 8. | Ren LL, Wang YM, Wu ZQ, Xiang ZC, Guo L, Xu T, Jiang YZ, Xiong Y, Li YJ, Li XW, Li H, Fan GH, Gu XY, Xiao Y, Gao H, Xu JY, Yang F, Wang XM, Wu C, Chen L, Liu YW, Liu B, Yang J, Wang XR, Dong J, Li L, Huang CL, Zhao JP, Hu Y, Cheng ZS, Liu LL, Qian ZH, Qin C, Jin Q, Cao B, Wang JW. Identification of a novel coronavirus causing severe pneumonia in human: a descriptive study. Chin Med J (Engl). 2020;133:1015-1024. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 642] [Cited by in RCA: 725] [Article Influence: 145.0] [Reference Citation Analysis (0)] |
| 9. | Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565-574. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8473] [Cited by in RCA: 7588] [Article Influence: 1517.6] [Reference Citation Analysis (0)] |
| 10. | Wan Y, Shang J, Graham R, Baric RS, Li F. Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J Virol. 2020;94. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3085] [Cited by in RCA: 2910] [Article Influence: 582.0] [Reference Citation Analysis (0)] |
| 11. | Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020; 181: 271-280. e8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11946] [Cited by in RCA: 14241] [Article Influence: 2848.2] [Reference Citation Analysis (0)] |
| 12. | Zou X, Chen K, Zou J, Han P, Hao J, Han Z. Single-cell RNA-seq data analysis on the receptor ACE2 expression reveals the potential risk of different human organs vulnerable to 2019-nCoV infection. Front Med. 2020;14:185-192. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1286] [Cited by in RCA: 1527] [Article Influence: 305.4] [Reference Citation Analysis (0)] |
| 13. | Hamming I, Timens W, Bulthuis ML, Lely AT, Navis G, van Goor H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J Pathol. 2004;203:631-637. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3643] [Cited by in RCA: 4147] [Article Influence: 197.5] [Reference Citation Analysis (0)] |
| 14. | Xiao HL, Zhao LX, Yang J, Tong N, An L, Liu QT, Xie MR, Li CS. Association between ACE2/ACE balance and pneumocyte apoptosis in a porcine model of acute pulmonary thromboembolism with cardiac arrest. Mol Med Rep. 2018;17:4221-4228. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 6] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 15. | Zhang H, Kang Z, Gong H, Xu D, Wang J, Li Z, Cui X, Xiao J, Meng T, Zhou W, Liu J, Xu H. The digestive system is a potential route of 2019-nCov infection: a bioinformatics analysis based on single-cell transcriptomes. 2020 Preprint. bioRxiv. . [DOI] [Full Text] |
| 16. | Lotfi M, Hamblin MR, Rezaei N. COVID-19: Transmission, prevention, and potential therapeutic opportunities. Clin Chim Acta. 2020;508:254-266. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 525] [Cited by in RCA: 461] [Article Influence: 92.2] [Reference Citation Analysis (0)] |
| 17. | Grasselli G, Zangrillo A, Zanella A, Antonelli M, Cabrini L, Castelli A, Cereda D, Coluccello A, Foti G, Fumagalli R, Iotti G, Latronico N, Lorini L, Merler S, Natalini G, Piatti A, Ranieri MV, Scandroglio AM, Storti E, Cecconi M, Pesenti A; COVID-19 Lombardy ICU Network. Baseline Characteristics and Outcomes of 1591 Patients Infected With SARS-CoV-2 Admitted to ICUs of the Lombardy Region, Italy. JAMA. 2020;323:1574-1581. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3537] [Cited by in RCA: 3824] [Article Influence: 764.8] [Reference Citation Analysis (0)] |
| 18. | Lovato A, de Filippis C, Marioni G. Upper airway symptoms in coronavirus disease 2019 (COVID-19). Am J Otolaryngol. 2020;41:102474. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 76] [Cited by in RCA: 86] [Article Influence: 17.2] [Reference Citation Analysis (0)] |
| 20. | Song F, Shi N, Shan F, Zhang Z, Shen J, Lu H, Ling Y, Jiang Y, Shi Y. Emerging 2019 Novel Coronavirus (2019-nCoV) Pneumonia. Radiology. 2020;295:210-217. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 903] [Cited by in RCA: 784] [Article Influence: 156.8] [Reference Citation Analysis (0)] |
| 21. | Cavallo AU, Troisi J, Forcina M, Mari PV, Forte V, Sperandio M, Pagano S, Cavallo P, Floris R, Garaci F. Texture Analysis in the Evaluation of Covid-19 Pneumonia in Chest X-Ray Images: a Proof of Concept Study. Curr Med Imaging. 2021;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 11] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 22. | Cui N, Zou X, Xu L. Preliminary CT findings of coronavirus disease 2019 (COVID-19). Clin Imaging. 2020;65:124-132. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 27] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 23. | Shi S, Qin M, Shen B, Cai Y, Liu T, Yang F, Gong W, Liu X, Liang J, Zhao Q, Huang H, Yang B, Huang C. Association of Cardiac Injury With Mortality in Hospitalized Patients With COVID-19 in Wuhan, China. JAMA Cardiol. 2020;5:802-810. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2428] [Cited by in RCA: 3005] [Article Influence: 601.0] [Reference Citation Analysis (1)] |
| 24. | Zhu H, Rhee JW, Cheng P, Waliany S, Chang A, Witteles RM, Maecker H, Davis MM, Nguyen PK, Wu SM. Cardiovascular Complications in Patients with COVID-19: Consequences of Viral Toxicities and Host Immune Response. Curr Cardiol Rep. 2020;22:32. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 115] [Cited by in RCA: 124] [Article Influence: 24.8] [Reference Citation Analysis (0)] |
| 25. | Rizzo P, Vieceli Dalla Sega F, Fortini F, Marracino L, Rapezzi C, Ferrari R. COVID-19 in the heart and the lungs: could we "Notch" the inflammatory storm? Basic Res Cardiol. 2020;115:31. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 154] [Cited by in RCA: 136] [Article Influence: 27.2] [Reference Citation Analysis (0)] |
| 26. | Wu Y, Xu X, Chen Z, Duan J, Hashimoto K, Yang L, Liu C, Yang C. Nervous system involvement after infection with COVID-19 and other coronaviruses. Brain Behav Immun. 2020;87:18-22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1219] [Cited by in RCA: 1253] [Article Influence: 250.6] [Reference Citation Analysis (0)] |
| 27. | Cha MH, Regueiro M, Sandhu DS. Gastrointestinal and hepatic manifestations of COVID-19: A comprehensive review. World J Gastroenterol. 2020;26:2323-2332. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 124] [Cited by in RCA: 107] [Article Influence: 21.4] [Reference Citation Analysis (2)] |
| 28. | Larsen CP, Bourne TD, Wilson JD, Saqqa O, Sharshir MA. Collapsing Glomerulopathy in a Patient With COVID-19. Kidney Int Rep. 2020;5:935-939. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 197] [Cited by in RCA: 261] [Article Influence: 52.2] [Reference Citation Analysis (0)] |
| 29. | Levin BR, Antia R. Why we don't get sick: the within-host population dynamics of bacterial infections. Science. 2001;292:1112-1115. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 77] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 30. | Levin BR, Baquero F, Ankomah PP, McCall IC. Phagocytes, Antibiotics, and Self-Limiting Bacterial Infections. Trends Microbiol. 2017;25:878-892. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 36] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 31. | Azkur AK, Akdis M, Azkur D, Sokolowska M, van de Veen W, Brüggen MC, O'Mahony L, Gao Y, Nadeau K, Akdis CA. Immune response to SARS-CoV-2 and mechanisms of immunopathological changes in COVID-19. Allergy. 2020;75:1564-1581. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 820] [Cited by in RCA: 741] [Article Influence: 148.2] [Reference Citation Analysis (0)] |
| 32. | Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497-506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35178] [Cited by in RCA: 30080] [Article Influence: 6016.0] [Reference Citation Analysis (3)] |
| 33. | Patel KP, Patel PA, Vunnam RR, Hewlett AT, Jain R, Jing R, Vunnam SR. Gastrointestinal, hepatobiliary, and pancreatic manifestations of COVID-19. J Clin Virol. 2020;128:104386. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 144] [Cited by in RCA: 120] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 34. | Xiao F, Tang M, Zheng X, Liu Y, Li X, Shan H. . Gastroenterology 2020; 158: 1831-1833. e3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1985] [Cited by in RCA: 1993] [Article Influence: 398.6] [Reference Citation Analysis (1)] |
| 35. | Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA. 2020;323:1061-1069. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14113] [Cited by in RCA: 14753] [Article Influence: 2950.6] [Reference Citation Analysis (0)] |
| 36. | Du M, Cai G, Chen F, Christiani DC, Zhang Z, Wang M. Multiomics Evaluation of Gastrointestinal and Other Clinical Characteristics of COVID-19. Gastroenterology 2020; 158: 2298-2301. e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 113] [Cited by in RCA: 111] [Article Influence: 22.2] [Reference Citation Analysis (0)] |
| 37. | Gu J, Han B, Wang J. COVID-19: Gastrointestinal Manifestations and Potential Fecal-Oral Transmission. Gastroenterology. 2020;158:1518-1519. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 954] [Cited by in RCA: 949] [Article Influence: 189.8] [Reference Citation Analysis (1)] |
| 38. | Tian Y, Rong L, Nian W, He Y. Review article: gastrointestinal features in COVID-19 and the possibility of faecal transmission. Aliment Pharmacol Ther. 2020;51:843-851. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 594] [Cited by in RCA: 583] [Article Influence: 116.6] [Reference Citation Analysis (0)] |
| 39. | Leung WK, To KF, Chan PK, Chan HL, Wu AK, Lee N, Yuen KY, Sung JJ. Enteric involvement of severe acute respiratory syndrome-associated coronavirus infection. Gastroenterology. 2003;125:1011-1017. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 295] [Article Influence: 13.4] [Reference Citation Analysis (1)] |
| 40. | Ai JW, Zi H, Wang Y, Huang Q, Wang N, Li LY, Pei B, Ji J, Zeng XT. Clinical Characteristics of COVID-19 Patients With Gastrointestinal Symptoms: An Analysis of Seven Patients in China. Front Med (Lausanne). 2020;7:308. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 41. | Cao C, Chen M, He L, Xie J, Chen X. Clinical features and outcomes of COVID-19 patients with gastrointestinal symptoms. Crit Care. 2020;24:340. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 24] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 42. | Ferm S, Fisher C, Pakala T, Tong M, Shah D, Schwarzbaum D, Cooley V, Hussain S, Kim SH. Analysis of Gastrointestinal and Hepatic Manifestations of SARS-CoV-2 Infection in 892 Patients in Queens, NY. Clin Gastroenterol Hepatol 2020; 18: 2378-2379. e1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 60] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 43. | Jin X, Lian JS, Hu JH, Gao J, Zheng L, Zhang YM, Hao SR, Jia HY, Cai H, Zhang XL, Yu GD, Xu KJ, Wang XY, Gu JQ, Zhang SY, Ye CY, Jin CL, Lu YF, Yu X, Yu XP, Huang JR, Xu KL, Ni Q, Yu CB, Zhu B, Li YT, Liu J, Zhao H, Zhang X, Yu L, Guo YZ, Su JW, Tao JJ, Lang GJ, Wu XX, Wu WR, Qv TT, Xiang DR, Yi P, Shi D, Chen Y, Ren Y, Qiu YQ, Li LJ, Sheng J, Yang Y. Epidemiological, clinical and virological characteristics of 74 cases of coronavirus-infected disease 2019 (COVID-19) with gastrointestinal symptoms. Gut. 2020;69:1002-1009. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 944] [Cited by in RCA: 868] [Article Influence: 173.6] [Reference Citation Analysis (0)] |
| 44. | Redd WD, Zhou JC, Hathorn KE, McCarty TR, Bazarbashi AN, Thompson CC, Shen L, Chan WW. Prevalence and Characteristics of Gastrointestinal Symptoms in Patients With Severe Acute Respiratory Syndrome Coronavirus 2 Infection in the United States: A Multicenter Cohort Study. Gastroenterology 2020; 159: 765-767. e2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 241] [Cited by in RCA: 283] [Article Influence: 56.6] [Reference Citation Analysis (0)] |
| 45. | Remes-Troche JM, Ramos-de-la-Medina A, Manríquez-Reyes M, Martínez-Pérez-Maldonado L, Lara EL, Solís-González MA. Initial Gastrointestinal Manifestations in Patients With Severe Acute Respiratory Syndrome Coronavirus 2 Infection in 112 Patients From Veracruz in Southeastern Mexico. Gastroenterology. 2020;159:1179-1181. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 25] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 46. | Zhang H, Liao YS, Gong J, Liu J, Xia X, Zhang H. Clinical characteristics of coronavirus disease (COVID-19) patients with gastrointestinal symptoms: A report of 164 cases. Dig Liver Dis. 2020;52:1076-1079. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 38] [Article Influence: 7.6] [Reference Citation Analysis (3)] |
| 47. | Zhang L, Mei Q, Li L, Ye C, Huang Y, Wang Y, Tong F, Gao Y, Pan A. [Analysis of gastrointestinal symptoms in 80 patients with coronavirus disease 2019]. Zhonghua Wei Zhong Bing Ji Jiu Yi Xue. 2020;32:412-416. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 48. | Zheng T, Yang C, Wang HY, Chen X, Yu L, Wu ZL, Sun H. Clinical characteristics and outcomes of COVID-19 patients with gastrointestinal symptoms admitted to Jianghan Fangcang Shelter Hospital in Wuhan, China. J Med Virol. 2020;92:2735-2741. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 36] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 49. | Sierpiński R, Pinkas J, Jankowski M, Zgliczyński WS, Wierzba W, Gujski M, Szumowski Ł. Sex differences in the frequency of gastrointestinal symptoms and olfactory or taste disorders in 1942 nonhospitalized patients with coronavirus disease 2019 (COVID-19). Pol Arch Intern Med. 2020;130:501-505. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 28] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 50. | Yang TY, Li YC, Wang SC, Dai QQ, Jiang XS, Zuo S, Jia L, Zheng JB, Wang HL. Clinical characteristics of patients with COVID-19 presenting with gastrointestinal symptoms as initial symptoms: Retrospective case series. World J Clin Cases. 2020;8:2950-2958. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 8] [Cited by in RCA: 8] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 51. | Kotfis K, Skonieczna-Żydecka K. COVID-19: gastrointestinal symptoms and potential sources of SARS-CoV-2 transmission. Anaesthesiol Intensive Ther. 2020;52:171-172. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 48] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 52. | Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367:1260-1263. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5792] [Cited by in RCA: 6468] [Article Influence: 1293.6] [Reference Citation Analysis (0)] |
| 53. | Hashimoto T, Perlot T, Rehman A, Trichereau J, Ishiguro H, Paolino M, Sigl V, Hanada T, Hanada R, Lipinski S, Wild B, Camargo SM, Singer D, Richter A, Kuba K, Fukamizu A, Schreiber S, Clevers H, Verrey F, Rosenstiel P, Penninger JM. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature. 2012;487:477-481. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 787] [Cited by in RCA: 984] [Article Influence: 75.7] [Reference Citation Analysis (0)] |
| 54. | Ye Q, Wang B, Zhang T, Xu J, Shang S. The mechanism and treatment of gastrointestinal symptoms in patients with COVID-19. Am J Physiol Gastrointest Liver Physiol. 2020;319:G245-G252. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 113] [Article Influence: 22.6] [Reference Citation Analysis (2)] |
| 55. | He L, Ding Y, Zhang Q, Che X, He Y, Shen H, Wang H, Li Z, Zhao L, Geng J, Deng Y, Yang L, Li J, Cai J, Qiu L, Wen K, Xu X, Jiang S. Expression of elevated levels of pro-inflammatory cytokines in SARS-CoV-infected ACE2+ cells in SARS patients: relation to the acute lung injury and pathogenesis of SARS. J Pathol. 2006;210:288-297. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 330] [Cited by in RCA: 323] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 56. | Villapol S. Gastrointestinal symptoms associated with COVID-19: impact on the gut microbiome. Transl Res. 2020;226:57-69. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 194] [Cited by in RCA: 217] [Article Influence: 43.4] [Reference Citation Analysis (0)] |
| 57. | Budden KF, Gellatly SL, Wood DL, Cooper MA, Morrison M, Hugenholtz P, Hansbro PM. Emerging pathogenic links between microbiota and the gut-lung axis. Nat Rev Microbiol. 2017;15:55-63. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 622] [Cited by in RCA: 1005] [Article Influence: 111.7] [Reference Citation Analysis (0)] |
| 58. | Emami A, Javanmardi F, Pirbonyeh N, Akbari A. Prevalence of Underlying Diseases in Hospitalized Patients with COVID-19: a Systematic Review and Meta-Analysis. Arch Acad Emerg Med. 2020;8:e35. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 158] [Cited by in RCA: 124] [Article Influence: 24.8] [Reference Citation Analysis (0)] |
| 59. | Wu J, Liu J, Zhao X, Liu C, Wang W, Wang D, Xu W, Zhang C, Yu J, Jiang B, Cao H, Li L. Clinical Characteristics of Imported Cases of Coronavirus Disease 2019 (COVID-19) in Jiangsu Province: A Multicenter Descriptive Study. Clin Infect Dis. 2020;71:706-712. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 393] [Cited by in RCA: 413] [Article Influence: 82.6] [Reference Citation Analysis (1)] |
| 60. | Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395:507-513. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14869] [Cited by in RCA: 12966] [Article Influence: 2593.2] [Reference Citation Analysis (1)] |
| 61. | CDC COVID-19 Response Team. Preliminary Estimates of the Prevalence of Selected Underlying Health Conditions Among Patients with Coronavirus Disease 2019 - United States, February 12–March 28, 2020. MMWR Morb Mortal Wkly Rep. 2020;69:382-386. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 878] [Cited by in RCA: 1052] [Article Influence: 210.4] [Reference Citation Analysis (0)] |
| 62. | Istituto Superiore di Sanità. Characteristics of COVID-19 patients dying in Italy. Epicentro. [cited January 26, 2021]. In: Istituto Superiore di Sanità [Internet]. Available from: https://www.epicentro.iss.it/en/coronavirus/sars-cov-2-analysis-of-deaths. |
| 63. | Papa A, Covino M, Pizzolante F, Miele L, Lopetuso LR, Bove V, Iorio R, Simeoni B, Vetrone LM, Tricoli L, Mignini I, Schepis T, D'Alessandro A, Coppola G, Nicoletti T, Visconti E, Rapaccini G. Gastrointestinal symptoms and digestive comorbidities in an Italian cohort of patients with COVID-19. Eur Rev Med Pharmacol Sci. 2020;24:7506-7511. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (0)] |
| 64. | Ding S, Liang TJ. Is SARS-CoV-2 Also an Enteric Pathogen With Potential Fecal-Oral Transmission? Gastroenterology. 2020;159:53-61. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 126] [Cited by in RCA: 138] [Article Influence: 27.6] [Reference Citation Analysis (0)] |
| 65. | Heller L, Mota CR, Greco DB. COVID-19 faecal-oral transmission: Are we asking the right questions? Sci Total Environ. 2020;729:138919. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 194] [Cited by in RCA: 157] [Article Influence: 31.4] [Reference Citation Analysis (0)] |
| 66. | Lin L, Jiang X, Zhang Z, Huang S, Fang Z, Gu Z, Gao L, Shi H, Mai L, Liu Y, Lin X, Lai R, Yan Z, Li X, Shan H. Gastrointestinal symptoms of 95 cases with SARS-CoV-2 infection. Gut. 2020;69:997-1001. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 630] [Cited by in RCA: 657] [Article Influence: 131.4] [Reference Citation Analysis (0)] |
| 67. | Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, Spitters C, Ericson K, Wilkerson S, Tural A, Diaz G, Cohn A, Fox L, Patel A, Gerber SI, Kim L, Tong S, Lu X, Lindstrom S, Pallansch MA, Weldon WC, Biggs HM, Uyeki TM, Pillai SK; Washington State 2019-nCoV Case Investigation Team. First Case of 2019 Novel Coronavirus in the United States. N Engl J Med. 2020;382:929-936. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4155] [Cited by in RCA: 3820] [Article Influence: 764.0] [Reference Citation Analysis (1)] |
| 68. | Wang W, Xu Y, Gao R, Lu R, Han K, Wu G, Tan W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA. 2020;323:1843-1844. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1894] [Cited by in RCA: 2658] [Article Influence: 531.6] [Reference Citation Analysis (0)] |
| 69. | Wölfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Müller MA, Niemeyer D, Jones TC, Vollmar P, Rothe C, Hoelscher M, Bleicker T, Brünink S, Schneider J, Ehmann R, Zwirglmaier K, Drosten C, Wendtner C. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020;581:465-469. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4682] [Cited by in RCA: 4805] [Article Influence: 961.0] [Reference Citation Analysis (0)] |
| 70. | Wu Y, Guo C, Tang L, Hong Z, Zhou J, Dong X, Yin H, Xiao Q, Tang Y, Qu X, Kuang L, Fang X, Mishra N, Lu J, Shan H, Jiang G, Huang X. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol Hepatol. 2020;5:434-435. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1050] [Cited by in RCA: 1150] [Article Influence: 230.0] [Reference Citation Analysis (0)] |
| 71. | Zhang J, Wang S, Xue Y. Fecal specimen diagnosis 2019 novel coronavirus-infected pneumonia. J Med Virol. 2020;92:680-682. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 278] [Cited by in RCA: 301] [Article Influence: 60.2] [Reference Citation Analysis (0)] |
| 72. | Zhang W, Du RH, Li B, Zheng XS, Yang XL, Hu B, Wang YY, Xiao GF, Yan B, Shi ZL, Zhou P. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg Microbes Infect. 2020;9:386-389. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1192] [Cited by in RCA: 1219] [Article Influence: 243.8] [Reference Citation Analysis (0)] |
| 73. | Xing YH, Ni W, Wu Q, Li WJ, Li GJ, Wang WD, Tong JN, Song XF, Wing-Kin Wong G, Xing QS. Prolonged viral shedding in feces of pediatric patients with coronavirus disease 2019. J Microbiol Immunol Infect. 2020;53:473-480. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 239] [Cited by in RCA: 240] [Article Influence: 48.0] [Reference Citation Analysis (0)] |
| 74. | Xu CLH, Raval M, Schnall JA, Kwong JC, Holmes NE. Duration of Respiratory and Gastrointestinal Viral Shedding in Children With SARS-CoV-2: A Systematic Review and Synthesis of Data. Pediatr Infect Dis J. 2020;39:e249-e256. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 53] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 75. | Xu Y, Li X, Zhu B, Liang H, Fang C, Gong Y, Guo Q, Sun X, Zhao D, Shen J, Zhang H, Liu H, Xia H, Tang J, Zhang K, Gong S. Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding. Nat Med. 2020;26:502-505. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 992] [Cited by in RCA: 1051] [Article Influence: 210.2] [Reference Citation Analysis (0)] |
| 76. | Lamers MM, Beumer J, van der Vaart J, Knoops K, Puschhof J, Breugem TI, Ravelli RBG, Paul van Schayck J, Mykytyn AZ, Duimel HQ, van Donselaar E, Riesebosch S, Kuijpers HJH, Schipper D, van de Wetering WJ, de Graaf M, Koopmans M, Cuppen E, Peters PJ, Haagmans BL, Clevers H. SARS-CoV-2 productively infects human gut enterocytes. Science. 2020;369:50-54. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1245] [Cited by in RCA: 1308] [Article Influence: 261.6] [Reference Citation Analysis (0)] |
| 77. | Zang R, Gomez Castro MF, McCune BT, Zeng Q, Rothlauf PW, Sonnek NM, Liu Z, Brulois KF, Wang X, Greenberg HB, Diamond MS, Ciorba MA, Whelan SPJ, Ding S. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci Immunol. 2020;5. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 809] [Cited by in RCA: 763] [Article Influence: 152.6] [Reference Citation Analysis (0)] |
| 78. | Ye Q, Wang B, Mao J. The pathogenesis and treatment of the `Cytokine Storm' in COVID-19. J Infect. 2020;80:607-613. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1858] [Cited by in RCA: 1958] [Article Influence: 391.6] [Reference Citation Analysis (0)] |
| 79. | Yeoh YK, Zuo T, Lui GC, Zhang F, Liu Q, Li AY, Chung AC, Cheung CP, Tso EY, Fung KS, Chan V, Ling L, Joynt G, Hui DS, Chow KM, Ng SSS, Li TC, Ng RW, Yip TC, Wong GL, Chan FK, Wong CK, Chan PK, Ng SC. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut. 2021;70:698-706. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 995] [Cited by in RCA: 861] [Article Influence: 215.3] [Reference Citation Analysis (0)] |
| 80. | Shen B, Yi X, Sun Y, Bi X, Du J, Zhang C, Quan S, Zhang F, Sun R, Qian L, Ge W, Liu W, Liang S, Chen H, Zhang Y, Li J, Xu J, He Z, Chen B, Wang J, Yan H, Zheng Y, Wang D, Zhu J, Kong Z, Kang Z, Liang X, Ding X, Ruan G, Xiang N, Cai X, Gao H, Li L, Li S, Xiao Q, Lu T, Zhu Y, Liu H, Guo T. Proteomic and Metabolomic Characterization of COVID-19 Patient Sera. Cell 2020; 182: 59-72. e15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 728] [Cited by in RCA: 1087] [Article Influence: 217.4] [Reference Citation Analysis (0)] |
| 81. | Gao L, Jiang D, Wen XS, Cheng XC, Sun M, He B, You LN, Lei P, Tan XW, Qin S, Cai GQ, Zhang DY. Prognostic value of NT-proBNP in patients with severe COVID-19. Respir Res. 2020;21:83. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 170] [Cited by in RCA: 192] [Article Influence: 38.4] [Reference Citation Analysis (0)] |
| 82. | Henry BM, Aggarwal G, Wong J, Benoit S, Vikse J, Plebani M, Lippi G. Lactate dehydrogenase levels predict coronavirus disease 2019 (COVID-19) severity and mortality: A pooled analysis. Am J Emerg Med. 2020;38:1722-1726. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 232] [Cited by in RCA: 354] [Article Influence: 70.8] [Reference Citation Analysis (0)] |
| 83. | Yu L. Restoring Good Health in Elderly with Diverse Gut Microbiome and Food Intake Restriction to Combat COVID-19. Indian J Microbiol. 2021: 1-4. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 7] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 84. | Costantini S. The Value of the Cytokinome Profile. In: Sharma A. Inflammatory Diseases. Rijeka: IntechOpen, 2011. [DOI] [Full Text] |
| 85. | Omodei D, Pucino V, Labruna G, Procaccini C, Galgani M, Perna F, Pirozzi D, De Caprio C, Marone G, Fontana L, Contaldo F, Pasanisi F, Matarese G, Sacchetti L. Immune-metabolic profiling of anorexic patients reveals an anti-oxidant and anti-inflammatory phenotype. Metabolism. 2015;64:396-405. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 36] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 86. | Nova E, Samartín S, Gómez S, Morandé G, Marcos A. The adaptive response of the immune system to the particular malnutrition of eating disorders. Eur J Clin Nutr. 2002;56 Suppl 3:S34-S37. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 71] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 87. | Centers for Disease Control and Prevention. Information for Healthcare Providers about Multisystem Inflammatory Syndrome in Children (MIS-C). [cited January 27, 2021]. In: Centers for Disease Control and Prevention [Internet]. Available from: https://www.cdc.gov/mis-c/hcp/. |
| 88. | Feldstein LR, Rose EB, Horwitz SM, Collins JP, Newhams MM, Son MBF, Newburger JW, Kleinman LC, Heidemann SM, Martin AA, Singh AR, Li S, Tarquinio KM, Jaggi P, Oster ME, Zackai SP, Gillen J, Ratner AJ, Walsh RF, Fitzgerald JC, Keenaghan MA, Alharash H, Doymaz S, Clouser KN, Giuliano JS Jr, Gupta A, Parker RM, Maddux AB, Havalad V, Ramsingh S, Bukulmez H, Bradford TT, Smith LS, Tenforde MW, Carroll CL, Riggs BJ, Gertz SJ, Daube A, Lansell A, Coronado Munoz A, Hobbs CV, Marohn KL, Halasa NB, Patel MM, Randolph AG; Overcoming COVID-19 Investigators; CDC COVID-19 Response Team. Multisystem Inflammatory Syndrome in U.S. Children and Adolescents. N Engl J Med. 2020;383:334-346. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1587] [Cited by in RCA: 1846] [Article Influence: 369.2] [Reference Citation Analysis (0)] |
| 89. | Consiglio CR, Cotugno N, Sardh F, Pou C, Amodio D, Rodriguez L, Tan Z, Zicari S, Ruggiero A, Pascucci GR, Santilli V, Campbell T, Bryceson Y, Eriksson D, Wang J, Marchesi A, Lakshmikanth T, Campana A, Villani A, Rossi P; CACTUS Study Team; Landegren N, Palma P, Brodin P. The Immunology of Multisystem Inflammatory Syndrome in Children with COVID-19. Cell 2020; 183: 968-981. e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 679] [Cited by in RCA: 653] [Article Influence: 130.6] [Reference Citation Analysis (0)] |
| 90. | Gruber CN, Patel RS, Trachtman R, Lepow L, Amanat F, Krammer F, Wilson KM, Onel K, Geanon D, Tuballes K, Patel M, Mouskas K, O'Donnell T, Merritt E, Simons NW, Barcessat V, Del Valle DM, Udondem S, Kang G, Gangadharan S, Ofori-Amanfo G, Laserson U, Rahman A, Kim-Schulze S, Charney AW, Gnjatic S, Gelb BD, Merad M, Bogunovic D. Mapping Systemic Inflammation and Antibody Responses in Multisystem Inflammatory Syndrome in Children (MIS-C). Cell 2020; 183: 982-995. e14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 422] [Cited by in RCA: 429] [Article Influence: 85.8] [Reference Citation Analysis (0)] |
| 91. | Porritt RA, Paschold L, Rivas MN, Cheng MH, Yonker LM, Chandnani H, Lopez M, Simnica D, Schultheiß C, Santiskulvong C, Van Eyk J, Fasano A, Bahar I, Binder M, Arditi M. Identification of a unique TCR repertoire, consistent with a superantigen selection process in Children with Multi-system Inflammatory Syndrome. bioRxiv. 2020;Preprint. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 6] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 92. | Cheng MH, Zhang S, Porritt RA, Noval Rivas M, Paschold L, Willscher E, Binder M, Arditi M, Bahar I. Superantigenic character of an insert unique to SARS-CoV-2 spike supported by skewed TCR repertoire in patients with hyperinflammation. Proc Natl Acad Sci USA. 2020;117:25254-25262. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 138] [Cited by in RCA: 235] [Article Influence: 47.0] [Reference Citation Analysis (0)] |
| 93. | Yonker LM, Neilan AM, Bartsch Y, Patel AB, Regan J, Arya P, Gootkind E, Park G, Hardcastle M, St John A, Appleman L, Chiu ML, Fialkowski A, De la Flor D, Lima R, Bordt EA, Yockey LJ, D'Avino P, Fischinger S, Shui JE, Lerou PH, Bonventre JV, Yu XG, Ryan ET, Bassett IV, Irimia D, Edlow AG, Alter G, Li JZ, Fasano A. Pediatric Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): Clinical Presentation, Infectivity, and Immune Responses. J Pediatr 2020; 227: 45-52. e5. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 223] [Cited by in RCA: 243] [Article Influence: 48.6] [Reference Citation Analysis (0)] |
| 94. | Carter MJ, Fish M, Jennings A, Doores KJ, Wellman P, Seow J, Acors S, Graham C, Timms E, Kenny J, Neil S, Malim MH, Tibby SM, Shankar-Hari M. Peripheral immunophenotypes in children with multisystem inflammatory syndrome associated with SARS-CoV-2 infection. Nat Med. 2020;26:1701-1707. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 299] [Cited by in RCA: 308] [Article Influence: 61.6] [Reference Citation Analysis (0)] |
| 95. | Pierce CA, Preston-Hurlburt P, Dai Y, Aschner CB, Cheshenko N, Galen B, Garforth SJ, Herrera NG, Jangra RK, Morano NC, Orner E, Sy S, Chandran K, Dziura J, Almo SC, Ring A, Keller MJ, Herold KC, Herold BC. Immune responses to SARS-CoV-2 infection in hospitalized pediatric and adult patients. Sci Transl Med. 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 203] [Cited by in RCA: 273] [Article Influence: 54.6] [Reference Citation Analysis (0)] |
| 96. | Noval Rivas M, Wakita D, Franklin MK, Carvalho TT, Abolhesn A, Gomez AC, Fishbein MC, Chen S, Lehman TJ, Sato K, Shibuya A, Fasano A, Kiyono H, Abe M, Tatsumoto N, Yamashita M, Crother TR, Shimada K, Arditi M. Intestinal Permeability and IgA Provoke Immune Vasculitis Linked to Cardiovascular Inflammation. Immunity 2019; 51: 508-521. e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 60] [Cited by in RCA: 107] [Article Influence: 17.8] [Reference Citation Analysis (0)] |
| 97. | Troisi J, Venutolo G, Terracciano C, Carri MD, Di Micco S, Landolfi A, Fasano A. The therapeutic use of the zonulin inhibitor AT-1001 (Larazotide) for a variety of acute and chronic inflammatory diseases. Curr Med Chem. 2021;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 20] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 98. | Fasano A. Zonulin and its regulation of intestinal barrier function: the biological door to inflammation, autoimmunity, and cancer. Physiol Rev. 2011;91:151-175. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 500] [Cited by in RCA: 631] [Article Influence: 45.1] [Reference Citation Analysis (0)] |
| 99. | Di Micco S, Musella S, Scala MC, Sala M, Campiglia P, Bifulco G, Fasano A. In silico Analysis Revealed Potential Anti-SARS-CoV-2 Main Protease Activity by the Zonulin Inhibitor Larazotide Acetate. Front Chem. 2020;8:628609. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 18] [Article Influence: 4.5] [Reference Citation Analysis (0)] |