Published online Nov 7, 2020. doi: 10.3748/wjg.v26.i41.6335
Peer-review started: July 22, 2020
First decision: August 8, 2020
Revised: August 10, 2020
Accepted: September 25, 2020
Article in press: September 25, 2020
Published online: November 7, 2020
Processing time: 106 Days and 15.3 Hours
The emergence of coronavirus disease-2019 induced by a newly identified b-coronavirus, namely severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has constituted a public health emergency. Even though it was considered a zoonotic disease, the virus has also spread among humans via respiratory secretions. The expression and distribution of angiotensin converting enzyme type 2 (ACE2) in various human organs might also show other possible infection routes. High ACE2 ribonucleic acid expression has been identified in the gastrointestinal tract (GI) indicating its importance as a possible infection pathway of SARS-CoV-2. ACE2 induces viral entry into the host and most importantly has been found to be associated with the function of the gut. Its deficiency has been implicated in several pathologies such as colorectal inflammation. The renin-angiotensin system (RAS) is an essential regulatory cascade operating both at a local tissue level and at the systemic or circulatory level. The RAS may be important in the pathogenesis of chronic liver disease and is associated with the up-regulation of ACE2. Thus, the aim of this review is firstly, the analysis of some important general and genome characteristics of SARS-CoV-2 and secondly, and most importantly, to focus on the utility of ACE2 receptors in both SARS-CoV-2 replication and pathogenesis, especially in the GI tract.
Core Tip: Although the main route of spread and symptomatology of coronavirus disease 2019 is closely related to the respiratory tract, the gastrointestinal tract could be an alternative and potential route of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) transmission and manifestation, most probably as a result of the overexpression of angiotensin converting enzyme type 2 (ACE2) receptors in the digestive system. Thus, this review primarily focuses on the analysis of some critical elements regarding the SARS-CoV-2 genome and its general characteristics, but most importantly, intends to highlight the role of ACE2 receptors in both SARS-CoV-2 replication and pathogenesis, especially in the gastrointestinal tract.
- Citation: Galanopoulos M, Doukatas A, Gazouli M. Origin and genomic characteristics of SARS-CoV-2 and its interaction with angiotensin converting enzyme type 2 receptors, focusing on the gastrointestinal tract. World J Gastroenterol 2020; 26(41): 6335-6345
- URL: https://www.wjgnet.com/1007-9327/full/v26/i41/6335.htm
- DOI: https://dx.doi.org/10.3748/wjg.v26.i41.6335
Emerging coronaviruses (CoVs), such as the severe acute respiratory syndrome coronavirus (SARS-CoV) and the Middle east respiratory syndrome coronavirus (MERS-CoV), caused, and continue to induce infections in humans, resulting in serious threats to Health Care Systems[1]. In December 2019, a group of patients with pneumonia of an unknown cause was reported in Wuhan, China. On January 7, 2020 the new identified coronavirus was initially named 2019 novel coronavirus by the World Health Organization (WHO)[2]. The virus was later renamed severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the disease was known as coronavirus disease-2019 (COVID-19)[3]. On March 11, 2020 the WHO stated that this disease was a pandemic[4]. As of July 15 2020, 13150645 cases have been registered worldwide leading to 574464 deaths[5].
The distribution and expression of angiotensin converting enzyme type 2 (ACE2) in the human body may be a possible infection route of SARS-CoV-2[6]. Through the developed single-cell transcriptomes and RNA sequencing, the ACE2 RNA expression profile was analyzed at the single-cell level[7]. Increased ACE2 levels were found in various organs such as type II alveolar cells of the lung, but most importantly in the epithelial cells of the stomach and bowel[8,9]. These findings suggested that organs with high ACE2 expression, especially the gastrointestinal tract (GI) should be regarded as a possible target for COVID-19[8].
Thus, this review focuses on the analysis of some key elements regarding the SARS-CoV-2 genome and its general characteristics, in addition to the utility of ACE2 receptors in SARS-CoV-2 replication and pathogenesis, especially in the GI tract.
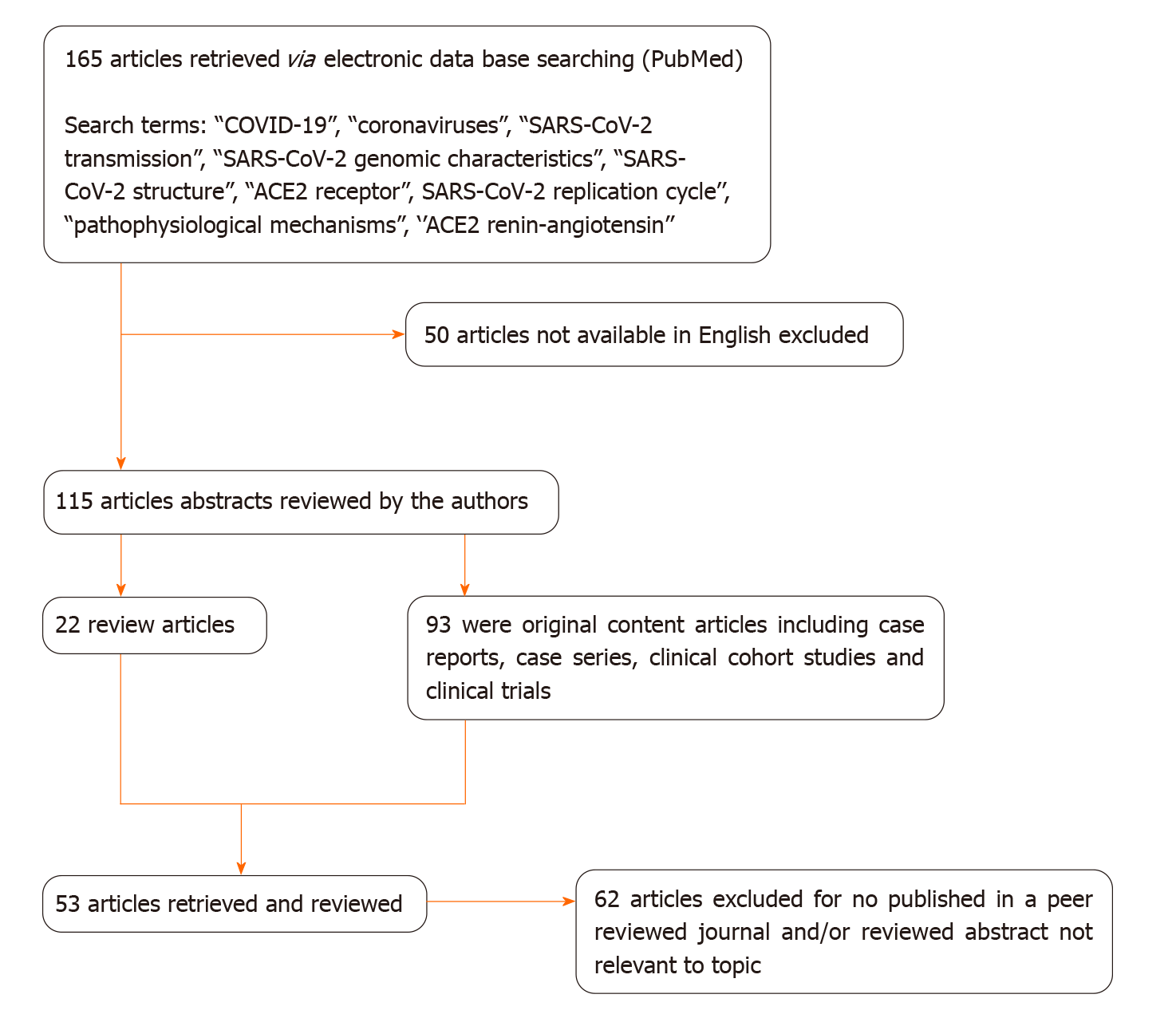
We conducted a literature search in the PubMed database, including most of the published studies in English and Chinese language since the emergence of the pandemic. The following keywords were used for the literature search: “COVID-19”, “coronaviruses”, “SARS-CoV-2 transmission”, “SARS-CoV-2 genomic characteristics”, “SARS-CoV-2 structure”, “ACE2 receptor”, SARS-CoV-2 replication cycle’’, “pathophysiological mechanisms”, and ‘’ACE2 renin-angiotensin’’. One hundred and sixty five publications fulfilled the search requirements. Of these, 115 articles were chosen due to the English language criteria. From these 115 articles, 93 were original articles and 22 were review articles, which included case series, case reports, clinical trials and clinical cohort studies. We reviewed all the abstracts and found 53 full-text articles appropriate for the current study (Figure 1). Two authors (DA and GM) independently reviewed the abstract and full text of the related articles.
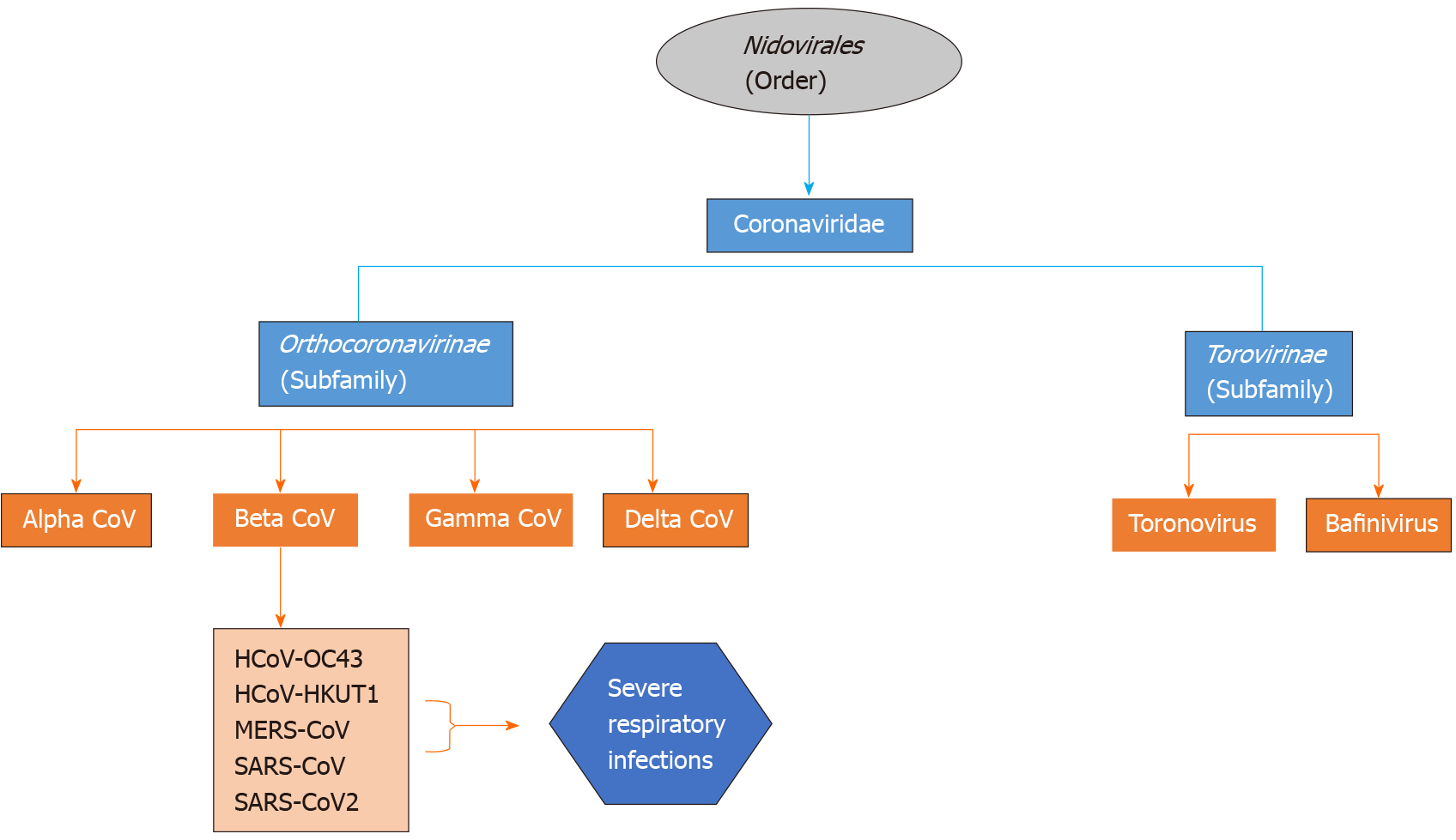
Coronaviruses are enveloped non-segmented positive-sense RNA viruses being members of the family of Coronaviridae, which is the largest class within the order Nidovirales, incorporating two subfamilies: Torovirinae and Orthocoronavirinae (Figure 2). The subfamily Orthocoronavirinae is classified into four genera: alphacoronavirus (alphaCoV), betacoronavirus (betaCoV), gammacoronavirus (gammaCoV) and deltacoronavirus (deltaCoV)[10,11]. CoVs mainly reside in birds and mammals and are frequently encountered in bats, camels and others. Genomic characterization has demonstrated that the most probable gene sources of alphaCoVs and betaCoVs are bats and rodents, and avian species for gammaCoVs and deltaCoVs. Regarding the manifestations of this group of viruses, they are able to cause enteric, neurological, hepatic and respiratory diseases in various animals[1,4].
It is estimated that approximately 2% of the population are asymptomatic carriers of a coronavirus, which in turn is responsible for 5% to 10% of respiratory infections[1]. The first recognized CoVs were Infectious Bronchitis Viruses causing respiratory disorders in chickens and the human CoVs (HCoVs), HCoV-OC43 and HCoV-229E, are responsible for the common cold. Since the emergence of HCoV-OC43 and HCoV-229E, seven other HCoVs have been recognized including HCoV-229E, HCoV-OC43, HCoV-NL63 and HCoV-HKU1[4,10]. The other two known betaCoVs, the SARS-CoV and the MERS-CoV are implicated in human respiratory infections with a case fatality rate of 10% and 35%, respectively[12,13].
Genetic sequence analysis has shown that the genome sequence of SARS-CoV-2 has a 79.0% similarity to SARS-CoV and 51.8% to MERS-CoV. This was the primary reason that the newly identified coronavirus was renamed SARS-CoV-2. In addition, it was shown that SARS-CoV-2 is 96% identical to the bat coronavirus (RaTG13) genome. Thus, bats have been considered the natural hosts of SARS-CoV-2. Nevertheless, the potential amplifying mammalian host, intermediate between human and host remains unidentified[14,15].
The emergence of multiple cases of unidentified respiratory tract infection, which appeared initially in China, was assumed to be caused by direct exposure to a seafood market. Zoonotic transmission was considered to be the primary mechanism. Several studies have indicated that bats could be a potential reservoir of SARS-CoV-2. Nevertheless, it remains unclear whether SARS-CoV-2 emerged from the seafood market. Therefore, it was concluded that human-to-human interactions were also a potential source of transmission of the virus[10,16].
SARS-CoV-2 human-to-human transmission takes place primarily among individuals closely related (i.e., family and friends), after encountering COVID-19 patients or carriers. According to the medical records of case series in Wuhan city and data from the local CDCs, it was reported that among non-residents of Wuhan, 72.3% exhibited a preceding contact with Wuhan inhabitants, and 31.3% had recently travelled to this city[17]. Moreover, it was reported that the incubation time of the virus might generally be within 3 to 7 d and up to 2 wk as the greatest time period from infection to symptomatology was 12.5 d[16,18]. As with other respiratory pathogens such as rhinovirus and influenza, it is mainly assumed that transmission occurs via respiratory droplets due to coughing and sneezing[1]. Fomites are also regarded as the main source of infectious particles, but there is some doubt regarding this. Fomites include both porous and nonporous surfaces or objects, which can be contaminated with pathogenic microorganisms that might be used as vehicles in the transmission process[19]. Additionally, in a symptomatic male, SARS-CoV-2 RNA was discovered in a stool specimen while the serum specimen was negative[18]. Chinese researchers were able to isolate SARS-CoV-2 from a swab sample of confirmed patient’s feces, demonstrating the potential for fecal-oral transmission[18]. Finally, it seems that possible COVID-19 sources are also asymptomatic people[20,21].
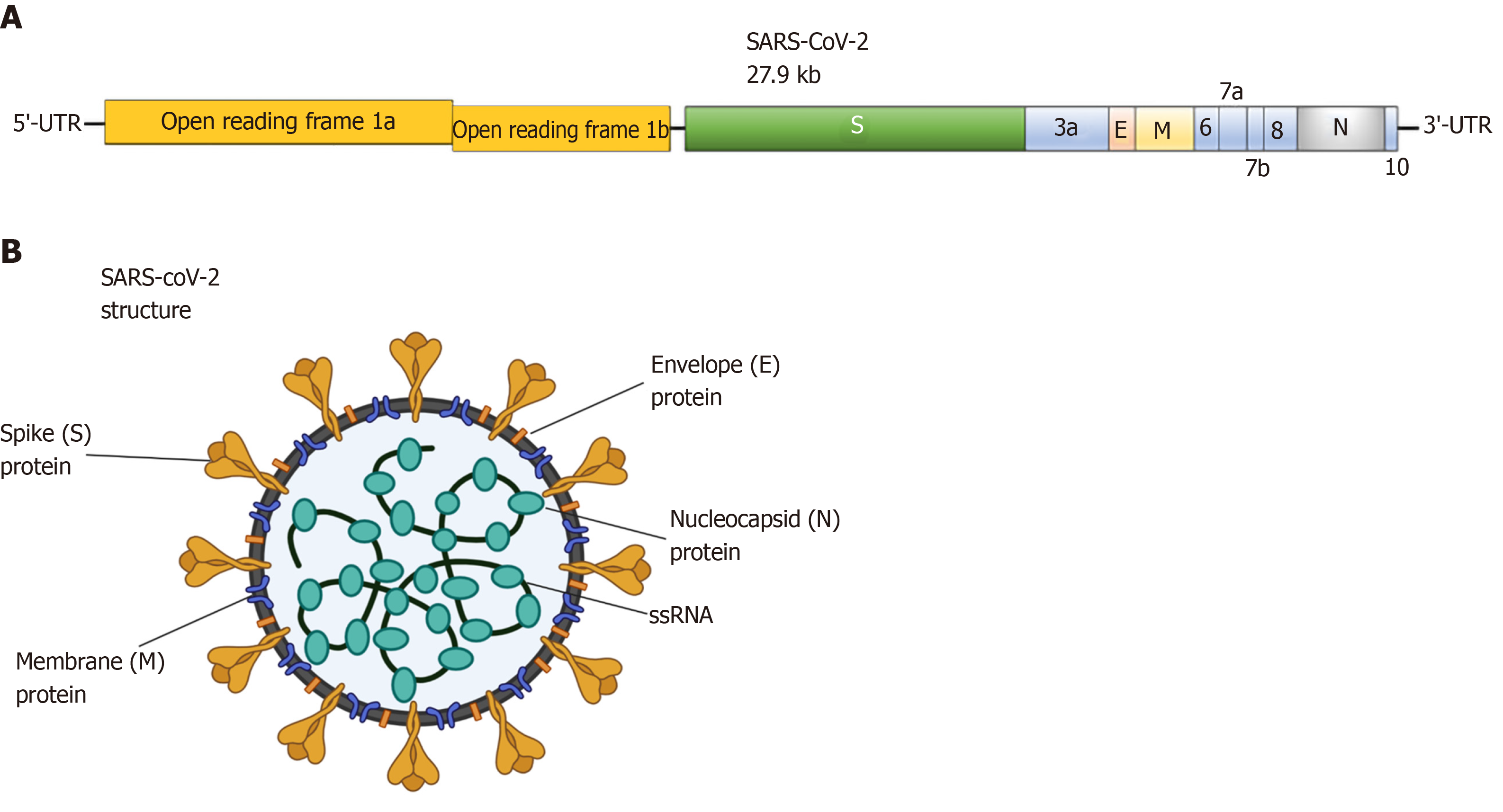
Coronaviruses are spherical and have spike protein projections on their surface resulting in a crown-like morphology, leading to their nomenclature[22,23]. SARS-CoV-2 has a round form with a diameter of approximately 60-140 nm[1]. In general, CoVs are viruses enveloped by a lipid bilayer. Their structure is composed of spike (S) glycoprotein, protein matrix (M), nucleocapsid (N) protein and small envelope (E) protein (Figure 3). The M, S and E proteins are enclosed in the viral envelope, while N protein binds with the viral RNA forming the nucleocapsid[13,15]. The M protein is responsible for the virus outline and with the assistance of E protein leads to the formation of the mature viral envelope, orchestrating in parallel, the assembly of the virus[24]. The S protein is responsible for the formation of homotrimeric spikes of SARS-CoV-2 surface inducing the entry into host cells[25,26].
The genomic size of CoVs ranges from 26 to 32 kilobases (kb)[27]. More specifically, according to genomic analysis of a new RNA virus strain initially named Wuhan-hu-1 coronavirus (WHCV) it was found that one strain of SARS-CoV-2 is 29.9 kb in size[28]. However, MERS-CoV and SARS-CoV have positive RNA genomes of 30.1 kb and 27.9 kb in size. Viral RNA codes for structural and nonstructural proteins. The structural proteins are formed within the 3’ end of the RNA genome. On the contrary, the 5΄ two-thirds of the genome codes for nonstructural proteins (nsps), essential for replication[11,27].
The aforementioned genomic content is initially utilized as a template for the synthesis of polyprotein 1a/b (pp1a/ppqab), encoding for nsps for the formation of the replication-transcription complex organized in double-membrane vesicles resulting in the production of subgenomic RNAs (sgRNAs). These subgenomic messenger RNAs (mRNAs) share the same leader sequence consisting of 70-90 nucleotides at their 5’ ends and the same 3’ ends. Transcription termination and the resulting acquisition of a leader RNA appears at transcription sequences, found between the open reading frames (ORFs). These particular minus-strand sgRNAs act as templates for the formation of subgenomic mRNAs[15,29].
The genome and the subgenomes of a typical CoV include at least six ORFs. Two-thirds of the viral RNA, primarily found in the first ORFs (ORF1a/b) encodes 16 nsps, except gammaCoV which lacks nsp1. Two polypeptides, pp1a and pp1b, are formed from the 1- frameshift between ORF1a and ORF1b. The two polypeptides are further processed by the virally encoded main protease or chymotrypsin-like protease. The remaining ORFs are responsible for encoding structural and accessory proteins. The remaining genome encodes the four critical structural proteins, S, M, E and N, as well as a number of additional proteins interfering with the host innate immune response (Figure 3)[10,15,29].
Based on primary data regarding the WHCV virus strain previously mentioned, it was shown that WHCV shares a phylogenetic and genomic resemblance to SARS-CoV, especially in the S-glycoprotein gene, emphasizing the ability of direct person spread[28]. In line with this, most proteins of SARS-CoV-2 are almost identical to SARS-CoVs with slight differences. Another study demonstrated that the mutation in both nsp2 and nsp3 are essential in the differentiation mechanism of the virus[30,31]. Therefore, further investigations are anticipated regarding the difference in transmission and host tropism between SARS-CoV and SARS-CoV-2. Finally, according to another Chinese research study analyzing the SARS-CoV-2 genotypes of patients from different areas, an increased mutation rate of the virus in many individuals was observed[32].
Various cellular receptors have been described as receptors of CoVs, such as ACE2[13,33]. ACE2 is a zinc-dependent carboxypeptidase and is expressed in vascular endothelial, cardiovascular and renal tissue, testes and epithelia of the small intestine[34]. The ACE2 genes codes for an 805 amino acid type I transmembrane glycoprotein and spans 18 exons. Furthermore, ACE2 includes a short intracellular cytoplasmic tail and a longer extracellular area exhibiting carboxyl-monopeptidase activity. The carboxyl-terminal domain of this particular receptor is approximately 48% homologous with collectrin, a protein expressed in the kidney[35].
ACE2 is a recognized cellular receptor for SARS-CoV regulating both human-to-human and cross-species spread[10]. Different studies have confirmed that SARS-CoV-2 exhibits similar and even, greater affinity to ACE2 receptors than SARS-CoV S protein[36,37]. Since the interaction of ACE2 receptor and S glycoprotein from SARS-CoV-2 is essential for virus entry, virus-receptor binding activity is undergoing exhaustive investigation via various methods. More specifically, the methodical identification of β-CoV receptors demonstrated that ACE2 positive human cells not expressing human aminopeptidase N or dipeptidyl peptidase-4 improved viral entry[10].
The virion S-glycoprotein binds to the human ACE2 receptor. After membrane infusion, the viral genome is liberated into the cytoplasm inducing translation of the two polyproteins, followed by the transcription of sgRNAs and lastly, its replication[13]. Newly formed envelope glycoproteins are directed in the Golgi apparatus or endoplasmic reticulum membranes leading to the formation of nucleocapsid proteins, envelope glycoproteins and genomic RNA, which in turn are accumulated and form viral unit buds. Finally, virion-containing vesicles release the virus[10,13].
ACE2 deficiency has been implicated in several pathologies such as acute lung injury or cardiovascular and renal disease due to the activation of the renin-angiotensin system (RAS)[38]. The RAS is an important regulatory cascade operating both at a systemic or circulatory level and at a local tissue level.
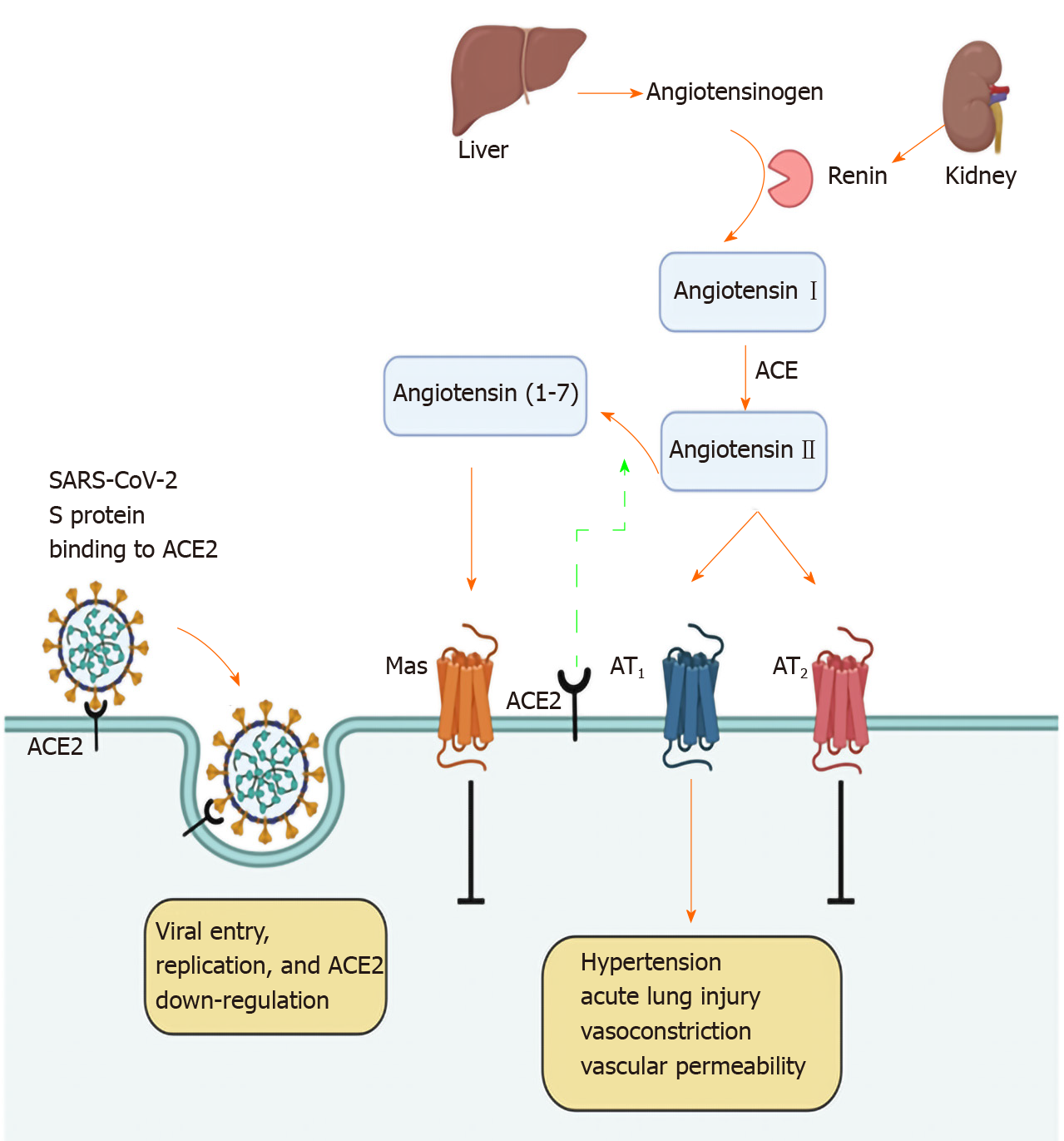
Within the RAS, regulation is carried out via protease cascades generating various bioactive peptides (Figure 4). The angiotensinogen originating from the liver and cleaved by renin is converted to angiotensin I (Ang I)[38]. The latter is subsequently, converted to angiotensin II (Ang II) by ACE, which is expressed in endothelial cells of vessels and in organs such as the heart, kidney, brain or the lung[39]. Ang II displays a major bioactive peptide within the RAS, which mediates its effects via the interaction with two G protein-coupled receptors, angiotensin II receptor type 1 and 2 (AT1 and AT2), activating downstream signaling[38,40].
Regarding the respiratory tract, elevated levels of Ang II stimulate the production of pulmonary fibrosis and hypertension. Local tissue-based RAS exacerbates lung injury, and worsens acute respiratory distress syndrome[41]. ACE inhibitors, along with AT1 antagonists have been associated with reduced pulmonary hypertension and reduced SARS-Co-V spike protein-induced lung injury.
ACE2 has also been found to be associated with the function of the gut, since it is expressed in intestinal epithelial cells. ACE2 function is required for the maintenance of antimicrobial peptide expression, ecology of the gut microbiome and amino acid homeostasis[42,43]. ACE2, as mentioned before, is a chimeric protein, which is homologous with collectrin. The discontinuance of collectrin function in mice is linked with near-complete downregulation of renal neutral amino acid transporters such as B0AT1; therefore, affecting renal amino acid re-absorption[42]. However, studies have shown that ACE2 can bind and stabilize B0AT1 on the surface of intestinal cells[38]. Moreover, both of these proteins are responsible for regulation of the recruitment of neutral amino acids in the bowel where the expression of collectrin is not found[38]. Furthermore, mutations in the slc6a encoding B0AT1 result in Hartnup’s disease[42]. The primary characteristic of this disease is the neutral aminoaciduria resembling the findings from collectrin-deficient mice. Most patients are commonly asymptomatic, even though certain conditions such as infections or malnutrition lead to pellagra-like symptoms[38]. Interestingly, dietary tryptophan is, mainly taken up via the B0AT1/ACE2 transport route on the cell surface located in the small gut. Tryptophan must be acquired from food, as is needed for the in vivo formation of nicotinamide preventing its deficiency (namely pellagra), by promoting the activation of mammalian target of rapamycin (mTOR), either directly via the tryptophan-nicotinamide pathway or through nutrient sensing, inducing the secretion of antimicrobial peptides such as a-defensins[42,44].
In an ongoing investigation of single cell-RNA sequencing information from control subjects and subjects with colitis or inflammatory bowel disease (IBD), it was indicated that ACE2 presence in colon cells was emphatically linked with genes inducing immunity, as well as viral infection, but was adversely connected with functions such as viral transcription, phagocytosis etc[45].
Furthermore, in another recent study, the potential pathogen transmission route of the SARS-CoV-2 infection in the GI tract was investigated. Colon tissue from healthy adults and colorectal cancer (CRC) patients was analyzed in terms of RNA expression of ACE2. The data showed that ACE2 is highly expressed in colonic cells, showing a gradual increase from healthy control individuals to CRC patients. Thus, it was suggested that SARS-CoV-2 may be more detrimental to patients with colorectal malignancy than individuals without CRC[46].
Finally, there has been important clinical evidence highlighting an ACE2 imbalance as a major player in poor outcomes (higher mortality rate and disease severity) in COVID-19 patients with pre-existing age-related comorbidities (renal, metabolic and cardiovascular disorders) addressing a possible role of gut microbiota dysbiosis in the disease prognosis[47].
Research has provided proof that RAS also plays an essential role in the pathophysiological mechanisms of chronic hepatobiliary disorders. It was shown in animal experiments that a noticeable up-regulation of intrahepatic RAS components exists, and the inhibition of RAS proved to prevent liver fibrogenetic changes[48,49]. RAS may be considered a dual function system in which the proliferative and antiproliferative functions are, mainly orchestrated by the ACE-ACE2 balance. High levels of Ang II due to up-regulation of ACE, result in the catabolism of Ang 1-7 inducing vasoconstriction and thus, liver injury and pulmonary fibrosis[50-52].
Different studies have shown that an opposite ratio in rat injured liver models and cirrhotic human livers is connected with up-regulation of ACE2 (gene and protein levels) and conversion of Ang II to Ang 1-7, resulting in elevated Ang 1-7 levels in the circulation facilitating vasodilation, and thus, antifibrotic activity in hepatic disorders[39,50,51]. Thus, the antagonism between Ang 1-7/Mas receptor and Ang II may represent an overall counter-regulatory action in the RAS system (Figure 4)[52]. Finally, but importantly, experimental studies have shown that ACE2 might also be upregulated due to oxygen reduction in the tissue. There is accumulating evidence that liver fibrosis is linked with advanced damage of the hepatocellular oxygenation mechanism with the most probable cause encompassing vasoconstriction, intrahepatic shunting and sinusoidal capillarization. Based on this hypothesis, it has been shown that in HepG2 cells, ACE2 mRNA progressively rose during increasing periods of hypoxia[50].
Finally, it has been shown that ACE2 up-regulation might also contribute to the vasodilation of cirrhosis via the degradation of Ang II. This observation might help understand why vasodilation continues to persist in individuals with progressive hepatic disorder, in spite of the stimulation of systemic RAS and why vasoconstrictor responses to Ang II are impaired[50,53].
Several studies have indicated the importance of the digestive system as an infection pathway of SARS-CoV-2. ACE2 receptors, highly expressed in the GI tract, have a significant contribution in the pathogenesis and replication of COVID-19, and consequently, in the GI system, as shown in this minireview. Further research is needed to clarify ACE cell function and its activity regarding SARS-CoV-2, as well as its utility concerning the pathophysiology of gastrointestinal and hepatobiliary diseases.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Greece
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B, B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Huerta-Franco MR, Lalmuanawma S S-Editor: Zhang H L-Editor: Webster JR P-Editor: Ma YJ
| 1. | Cascella M, Rajnik M, Cuomo A, Dulebohn SC, Di Napoli R. Features, Evaluation, and Treatment of Coronavirus (COVID-19). StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2020; Jan. [PubMed] |
| 2. | Wang C, Horby PW, Hayden FG, Gao GF. A novel coronavirus outbreak of global health concern. Lancet. 2020;395:470-473. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4848] [Cited by in RCA: 4383] [Article Influence: 876.6] [Reference Citation Analysis (1)] |
| 3. | Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W; China Novel Coronavirus Investigating and Research Team. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020;382:727-733. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18987] [Cited by in RCA: 17613] [Article Influence: 3522.6] [Reference Citation Analysis (0)] |
| 4. | Chang L, Yan Y, Wang L. Coronavirus Disease 2019: Coronaviruses and Blood Safety. Transfus Med Rev. 2020;34:75-80. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 447] [Cited by in RCA: 387] [Article Influence: 77.4] [Reference Citation Analysis (0)] |
| 5. | WHO. WHO Coronavirus Disease (COVID-19) Dashboard|WHO Coronavirus Disease (COVID-19) Dashboard [Internet]. 2020 [cited 2020 Jul 16]. Available from: URL: https://covid19.who.int/. |
| 6. | Xu H, Zhong L, Deng J, Peng J, Dan H, Zeng X, Li T, Chen Q. High expression of ACE2 receptor of 2019-nCoV on the epithelial cells of oral mucosa. Int J Oral Sci. 2020;12:8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1709] [Cited by in RCA: 1729] [Article Influence: 345.8] [Reference Citation Analysis (0)] |
| 7. | Zou X, Chen K, Zou J, Han P, Hao J, Han Z. Single-cell RNA-seq data analysis on the receptor ACE2 expression reveals the potential risk of different human organs vulnerable to 2019-nCoV infection. Front Med. 2020;14:185-192. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1286] [Cited by in RCA: 1526] [Article Influence: 305.2] [Reference Citation Analysis (0)] |
| 8. | Zhang H, Kang Z, Gong H, Xu D, Wang J, Li Z, Cui X, Xiao J, Meng T, Zhou W, Liu J, Xu H. The digestive system is a potential route of 2019-nCov infection: a bioinformatics analysis based on single-cell transcriptomes. bioRxiv. 2020;2020.01.30.927806. [DOI] [Full Text] |
| 9. | Xiao X, Chakraborti S, Dimitrov AS, Gramatikoff K, Dimitrov DS. The SARS-CoV S glycoprotein: expression and functional characterization. Biochem Biophys Res Commun. 2003;312:1159-1164. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 283] [Cited by in RCA: 301] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 10. | Guo YR, Cao QD, Hong ZS, Tan YY, Chen SD, Jin HJ, Tan KS, Wang DY, Yan Y. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak - an update on the status. Mil Med Res. 2020;7:11. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1854] [Cited by in RCA: 2017] [Article Influence: 403.4] [Reference Citation Analysis (0)] |
| 11. | Ashour HM, Elkhatib WF, Rahman MM, Elshabrawy HA. Insights into the Recent 2019 Novel Coronavirus (SARS-CoV-2) in Light of Past Human Coronavirus Outbreaks. Pathogens. 2020;9:186. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 389] [Cited by in RCA: 356] [Article Influence: 71.2] [Reference Citation Analysis (1)] |
| 12. | Ding Y, Wang H, Shen H, Li Z, Geng J, Han H, Cai J, Li X, Kang W, Weng D, Lu Y, Wu D, He L, Yao K. The clinical pathology of severe acute respiratory syndrome (SARS): a report from China. J Pathol. 2003;200:282-289. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 582] [Cited by in RCA: 576] [Article Influence: 26.2] [Reference Citation Analysis (0)] |
| 13. | de Wit E, van Doremalen N, Falzarano D, Munster VJ. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol. 2016;14:523-534. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2157] [Cited by in RCA: 2277] [Article Influence: 253.0] [Reference Citation Analysis (0)] |
| 14. | Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270-273. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15248] [Cited by in RCA: 14097] [Article Influence: 2819.4] [Reference Citation Analysis (1)] |
| 15. | Chen Y, Liu Q, Guo D. Emerging coronaviruses: Genome structure, replication, and pathogenesis. J Med Virol. 2020;92:418-423. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1873] [Cited by in RCA: 1872] [Article Influence: 374.4] [Reference Citation Analysis (1)] |
| 16. | Li Q, Guan X, Wu P, Wang X, Zhou L, Tong Y, Ren R, Leung KSM, Lau EHY, Wong JY, Xing X, Xiang N, Wu Y, Li C, Chen Q, Li D, Liu T, Zhao J, Liu M, Tu W, Chen C, Jin L, Yang R, Wang Q, Zhou S, Wang R, Liu H, Luo Y, Liu Y, Shao G, Li H, Tao Z, Yang Y, Deng Z, Liu B, Ma Z, Zhang Y, Shi G, Lam TTY, Wu JT, Gao GF, Cowling BJ, Yang B, Leung GM, Feng Z. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus-Infected Pneumonia. N Engl J Med. 2020;382:1199-1207. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11224] [Cited by in RCA: 9310] [Article Influence: 1862.0] [Reference Citation Analysis (0)] |
| 17. | Guan WJ, Ni ZY, Hu Y, Liang WH, Ou CQ, He JX, Liu L, Shan H, Lei CL, Hui DSC, Du B, Li LJ, Zeng G, Yuen KY, Chen RC, Tang CL, Wang T, Chen PY, Xiang J, Li SY, Wang JL, Liang ZJ, Peng YX, Wei L, Liu Y, Hu YH, Peng P, Wang JM, Liu JY, Chen Z, Li G, Zheng ZJ, Qiu SQ, Luo J, Ye CJ, Zhu SY, Zhong NS; China Medical Treatment Expert Group for Covid-19. Clinical Characteristics of Coronavirus Disease 2019 in China. N Engl J Med. 2020;382:1708-1720. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19202] [Cited by in RCA: 18850] [Article Influence: 3770.0] [Reference Citation Analysis (7)] |
| 18. | Jiang F, Deng L, Zhang L, Cai Y, Cheung CW, Xia Z. Review of the Clinical Characteristics of Coronavirus Disease 2019 (COVID-19). J Gen Intern Med. 2020;35:1545-1549. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 869] [Cited by in RCA: 703] [Article Influence: 140.6] [Reference Citation Analysis (0)] |
| 19. | Boone SA, Gerba CP. Significance of fomites in the spread of respiratory and enteric viral disease. Appl Environ Microbiol. 2007;73:1687-1696. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 484] [Cited by in RCA: 418] [Article Influence: 23.2] [Reference Citation Analysis (0)] |
| 20. | Rothe C, Schunk M, Sothmann P, Bretzel G, Froeschl G, Wallrauch C, Zimmer T, Thiel V, Janke C, Guggemos W, Seilmaier M, Drosten C, Vollmar P, Zwirglmaier K, Zange S, Wölfel R, Hoelscher M. Transmission of 2019-nCoV Infection from an Asymptomatic Contact in Germany. N Engl J Med. 2020;382:970-971. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2799] [Cited by in RCA: 2489] [Article Influence: 497.8] [Reference Citation Analysis (0)] |
| 21. | Hu Z, Song C, Xu C, Jin G, Chen Y, Xu X, Ma H, Chen W, Lin Y, Zheng Y, Wang J, Hu Z, Yi Y, Shen H. Clinical characteristics of 24 asymptomatic infections with COVID-19 screened among close contacts in Nanjing, China. Sci China Life Sci. 2020;63:706-711. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 802] [Cited by in RCA: 858] [Article Influence: 171.6] [Reference Citation Analysis (0)] |
| 22. | Neuman BW, Adair BD, Yoshioka C, Quispe JD, Orca G, Kuhn P, Milligan RA, Yeager M, Buchmeier MJ. Supramolecular architecture of severe acute respiratory syndrome coronavirus revealed by electron cryomicroscopy. J Virol. 2006;80:7918-7928. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 278] [Cited by in RCA: 266] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 23. | Bárcena M, Oostergetel GT, Bartelink W, Faas FG, Verkleij A, Rottier PJ, Koster AJ, Bosch BJ. Cryo-electron tomography of mouse hepatitis virus: Insights into the structure of the coronavirion. Proc Natl Acad Sci USA. 2009;106:582-587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 183] [Cited by in RCA: 194] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 24. | Siu YL, Teoh KT, Lo J, Chan CM, Kien F, Escriou N, Tsao SW, Nicholls JM, Altmeyer R, Peiris JS, Bruzzone R, Nal B. The M, E, and N structural proteins of the severe acute respiratory syndrome coronavirus are required for efficient assembly, trafficking, and release of virus-like particles. J Virol. 2008;82:11318-11330. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 337] [Cited by in RCA: 355] [Article Influence: 20.9] [Reference Citation Analysis (0)] |
| 25. | Bosch BJ, van der Zee R, de Haan CA, Rottier PJ. The coronavirus spike protein is a class I virus fusion protein: structural and functional characterization of the fusion core complex. J Virol. 2003;77:8801-8811. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1045] [Cited by in RCA: 1083] [Article Influence: 49.2] [Reference Citation Analysis (0)] |
| 26. | Simmons G, Reeves JD, Rennekamp AJ, Amberg SM, Piefer AJ, Bates P. Characterization of severe acute respiratory syndrome-associated coronavirus (SARS-CoV) spike glycoprotein-mediated viral entry. Proc Natl Acad Sci USA. 2004;101:4240-4245. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 428] [Cited by in RCA: 437] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 27. | Chen Y, Su C, Ke M, Jin X, Xu L, Zhang Z, Wu A, Sun Y, Yang Z, Tien P, Ahola T, Liang Y, Liu X, Guo D. Biochemical and structural insights into the mechanisms of SARS coronavirus RNA ribose 2'-O-methylation by nsp16/nsp10 protein complex. PLoS Pathog. 2011;7:e1002294. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 275] [Cited by in RCA: 260] [Article Influence: 18.6] [Reference Citation Analysis (0)] |
| 28. | Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG, Hu Y, Tao ZW, Tian JH, Pei YY, Yuan ML, Zhang YL, Dai FH, Liu Y, Wang QM, Zheng JJ, Xu L, Holmes EC, Zhang YZ. A new coronavirus associated with human respiratory disease in China. Nature. 2020;579:265-269. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6893] [Cited by in RCA: 7479] [Article Influence: 1495.8] [Reference Citation Analysis (0)] |
| 29. | Perlman S, Netland J. Coronaviruses post-SARS: update on replication and pathogenesis. Nat Rev Microbiol. 2009;7:439-450. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1309] [Cited by in RCA: 1160] [Article Influence: 72.5] [Reference Citation Analysis (0)] |
| 30. | Wu A, Peng Y, Huang B, Ding X, Wang X, Niu P, Meng J, Zhu Z, Zhang Z, Wang J, Sheng J, Quan L, Xia Z, Tan W, Cheng G, Jiang T. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe. 2020;27:325-328. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1413] [Cited by in RCA: 1500] [Article Influence: 300.0] [Reference Citation Analysis (0)] |
| 31. | Angeletti S, Benvenuto D, Bianchi M, Giovanetti M, Pascarella S, Ciccozzi M. COVID-2019: The role of the nsp2 and nsp3 in its pathogenesis. J Med Virol. 2020;92:584-588. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 231] [Cited by in RCA: 228] [Article Influence: 45.6] [Reference Citation Analysis (0)] |
| 32. | Zhang L, Shen FM, Chen F, Lin Z. Origin and Evolution of the 2019 Novel Coronavirus. Clin Infect Dis. 2020;71:882-883. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 97] [Cited by in RCA: 95] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 33. | He Y, Li J, Du L, Yan X, Hu G, Zhou Y, Jiang S. Identification and characterization of novel neutralizing epitopes in the receptor-binding domain of SARS-CoV spike protein: revealing the critical antigenic determinants in inactivated SARS-CoV vaccine. Vaccine. 2006;24:5498-5508. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 52] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 34. | Jia HP, Look DC, Shi L, Hickey M, Pewe L, Netland J, Farzan M, Wohlford-Lenane C, Perlman S, McCray PB. ACE2 receptor expression and severe acute respiratory syndrome coronavirus infection depend on differentiation of human airway epithelia. J Virol. 2005;79:14614-14621. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 620] [Cited by in RCA: 644] [Article Influence: 33.9] [Reference Citation Analysis (0)] |
| 35. | Zhang H, Wada J, Hida K, Tsuchiyama Y, Hiragushi K, Shikata K, Wang H, Lin S, Kanwar YS, Makino H. Collectrin, a collecting duct-specific transmembrane glycoprotein, is a novel homolog of ACE2 and is developmentally regulated in embryonic kidneys. J Biol Chem. 2001;276:17132-17139. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 182] [Cited by in RCA: 185] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 36. | Ge XY, Li JL, Yang XL, Chmura AA, Zhu G, Epstein JH, Mazet JK, Hu B, Zhang W, Peng C, Zhang YJ, Luo CM, Tan B, Wang N, Zhu Y, Crameri G, Zhang SY, Wang LF, Daszak P, Shi ZL. Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature. 2013;503:535-538. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1165] [Cited by in RCA: 1309] [Article Influence: 109.1] [Reference Citation Analysis (0)] |
| 37. | Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565-574. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8473] [Cited by in RCA: 7587] [Article Influence: 1517.4] [Reference Citation Analysis (0)] |
| 38. | Perlot T, Penninger JM. ACE2 - from the renin-angiotensin system to gut microbiota and malnutrition. Microbes Infect. 2013;15:866-873. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 151] [Cited by in RCA: 181] [Article Influence: 15.1] [Reference Citation Analysis (0)] |
| 39. | Moreira de Macêdo S, Guimarães TA, Feltenberger JD, Sousa Santos SH. The role of renin-angiotensin system modulation on treatment and prevention of liver diseases. Peptides. 2014;62:189-196. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 76] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 40. | Lambert DW, Clarke NE, Turner AJ. Not just angiotensinases: new roles for the angiotensin-converting enzymes. Cell Mol Life Sci. 2010;67:89-98. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 59] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 41. | Marshall RP, Gohlke P, Chambers RC, Howell DC, Bottoms SE, Unger T, McAnulty RJ, Laurent GJ. Angiotensin II and the fibroproliferative response to acute lung injury. Am J Physiol Lung Cell Mol Physiol. 2004;286:L156-L164. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 187] [Cited by in RCA: 200] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 42. | Hashimoto T, Perlot T, Rehman A, Trichereau J, Ishiguro H, Paolino M, Sigl V, Hanada T, Hanada R, Lipinski S, Wild B, Camargo SM, Singer D, Richter A, Kuba K, Fukamizu A, Schreiber S, Clevers H, Verrey F, Rosenstiel P, Penninger JM. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature. 2012;487:477-481. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 787] [Cited by in RCA: 984] [Article Influence: 75.7] [Reference Citation Analysis (0)] |
| 43. | Vuille-dit-Bille RN, Camargo SM, Emmenegger L, Sasse T, Kummer E, Jando J, Hamie QM, Meier CF, Hunziker S, Forras-Kaufmann Z, Kuyumcu S, Fox M, Schwizer W, Fried M, Lindenmeyer M, Götze O, Verrey F. Human intestine luminal ACE2 and amino acid transporter expression increased by ACE-inhibitors. Amino Acids. 2015;47:693-705. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 249] [Article Influence: 22.6] [Reference Citation Analysis (0)] |
| 44. | Yisireyili M, Uchida Y, Yamamoto K, Nakayama T, Cheng XW, Matsushita T, Nakamura S, Murohara T, Takeshita K. Angiotensin receptor blocker irbesartan reduces stress-induced intestinal inflammation via AT1a signaling and ACE2-dependent mechanism in mice. Brain Behav Immun. 2018;69:167-179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 61] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 45. | Wang J, Zhao S, Liu M, Zhao Z, Xu Y, Wang P, Lin M, Xu Y, Huang B, Zuo X, Chen Z, Bai F, Cui J, Lew AM, Zhao J, Zhang Y, Luo H, Zhang Y. ACE2 expression by colonic epithelial cells is associated with viral infection, immunity and energy metabolism. MedRxiv. 2020;. [DOI] [Full Text] |
| 46. | Chen H, Xuan B, Yan Y, Zhu X, Shen C, Zhao G, Ji L, Xu D, Xiong H, Yu T, Li X, Liu Q, Chen Y, Cui Y, Hong J, Fang J-Y. Profiling ACE2 expression in colon tissue of healthy adults and colorectal cancer patients by single-cell transcriptome analysis. MedRxiv. 2020;. [DOI] [Full Text] |
| 47. | Viana SD, Nunes S, Reis F. ACE2 imbalance as a key player for the poor outcomes in COVID-19 patients with age-related comorbidities - Role of gut microbiota dysbiosis. Ageing Res Rev. 2020;62:101123. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 77] [Cited by in RCA: 121] [Article Influence: 24.2] [Reference Citation Analysis (0)] |
| 48. | Bataller R, Sancho-Bru P, Ginès P, Lora JM, Al-Garawi A, Solé M, Colmenero J, Nicolás JM, Jiménez W, Weich N, Gutiérrez-Ramos JC, Arroyo V, Rodés J. Activated human hepatic stellate cells express the renin-angiotensin system and synthesize angiotensin II. Gastroenterology. 2003;125:117-125. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 258] [Cited by in RCA: 251] [Article Influence: 11.4] [Reference Citation Analysis (0)] |
| 49. | Paizis G, Cooper ME, Schembri JM, Tikellis C, Burrell LM, Angus PW. Up-regulation of components of the renin-angiotensin system in the bile duct-ligated rat liver. Gastroenterology. 2002;123:1667-1676. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 154] [Cited by in RCA: 145] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 50. | Paizis G, Tikellis C, Cooper ME, Schembri JM, Lew RA, Smith AI, Shaw T, Warner FJ, Zuilli A, Burrell LM, Angus PW. Chronic liver injury in rats and humans upregulates the novel enzyme angiotensin converting enzyme 2. Gut. 2005;54:1790-1796. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 236] [Cited by in RCA: 281] [Article Influence: 14.1] [Reference Citation Analysis (0)] |
| 51. | Liu J, Gong H, Zhang ZT, Wang Y. Effect of angiotensin II and angiotensin II type 1 receptor antagonist on the proliferation, contraction and collagen synthesis in rat hepatic stellate cells. Chin Med J (Engl). 2008;121:161-165. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 16] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 52. | Santos RA, Ferreira AJ, Simões E Silva AC. Recent advances in the angiotensin-converting enzyme 2-angiotensin(1-7)-Mas axis. Exp Physiol. 2008;93:519-527. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 326] [Cited by in RCA: 340] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 53. | Newby DE, Jalan R, Masumori S, Hayes PC, Boon NA, Webb DJ. Peripheral vascular tone in patients with cirrhosis: role of the renin-angiotensin and sympathetic nervous systems. Cardiovasc Res. 1998;38:221-228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 1.2] [Reference Citation Analysis (0)] |