Published online Nov 7, 2018. doi: 10.3748/wjg.v24.i41.4652
Peer-review started: July 19, 2018
First decision: August 25, 2018
Revised: September 28, 2018
Accepted: October 16, 2018
Article in press: October 16, 2018
Published online: November 7, 2018
Processing time: 112 Days and 10.7 Hours
To investigate the adhesion and anti-inflammatory effects of Lactobacillus rhamnosus GG (LGG) in the colonic mucosa of healthy and ulcerative colitis (UC) patients, both in vivo and ex vivo in an organ culture model.
For the ex vivo experiment, a total of 98 patients (68 UC patients and 30 normal subjects) were included. Endoscopic biopsies were collected and incubated with and without LGG or LGG-conditioned media to evaluate the mucosal adhesion and anti-inflammatory effects [reduction of tumor necrosis factor alpha (TNFα) and interleukin (IL)-17 expression] of the bacteria, and extraction of DNA and RNA for quantification by real-time (RT)-PCR occurred after the incubation. A dose-response study was performed by incubating biopsies at “regular”, double and 5 times higher doses of LGG. For the in vivo experiment, a total of 42 patients (20 UC patients and 22 normal controls) were included. Biopsies were taken from the colons of normal subjects who consumed a commercial formulation of LGG for 7 d prior to the colonoscopy, and the adhesion of the bacteria to the colonic mucosa was evaluated by RT-PCR and compared with that of control biopsies from patients who did not consume the formulation. LGG adhesion and TNFα and IL-17 expression were compared between UC patients who consumed a regular or double dose of LGG supplementation prior to colonoscopy.
In the ex vivo experiment, LGG showed consistent adhesion to the distal and proximal colon in normal subjects and UC patients, with a trend towards higher concentrations in the distal colon, and in UC patients, adhesion was similar in biopsies with active and quiescent inflammation. In addition, bioptic samples from UC patients incubated with LGG conditioned media (CM) showed reduced expression of TNFα and IL-17 compared with the corresponding expression in controls (P < 0.05). Incubation with a double dose of LGG increased mucosal adhesion and the anti-inflammatory effects (P < 0.05). In the in vivo experiment, LGG was detectable only in the colon of patients who consumed the LGG formulation, and bowel cleansing did not affect LGG adhesion. UC patients who consumed the double LGG dose had increased mucosal concentrations of the bacteria and reduced TNFα and IL-17 expression compared with patients who consumed the regular dose (48% and 40% reduction, respectively, P < 0.05).
In an ex vivo organ culture model, LGG showed consistent adhesion and anti-inflammatory effects. Colonization by LGG after consumption for a week was demonstrated in vivo in the human colon. Increasing the administered dose increased the adhesion and effectiveness of the bacteria. For the first time, we demonstrated that LGG effectively adheres to the colonic mucosa and exerts anti-inflammatory effects, both ex vivo and in vivo.
Core tip: Since probiotic utilization is often driven by unproven health claims, we intended to explore the feasibility and effectiveness of a safe and well-characterized probiotic bacterial species, the Lactobacillus rhamnosus GG (LGG), in ulcerative colitis (UC) patients, through a preclinical proof-of-concept study. We demonstrated for the first time effective mucosal colonization and the anti-inflammatory effect of LGG, both ex vivo and in vivo, by quantifying bacterial DNA and cytokine RNA expression directly at the mucosal site using genomic techniques. Further translational and clinical studies would confirm the utility and optimize the therapeutic administration of LGG in UC patients.
- Citation: Pagnini C, Corleto VD, Martorelli M, Lanini C, D’Ambra G, Di Giulio E, Delle Fave G. Mucosal adhesion and anti-inflammatory effects of Lactobacillus rhamnosus GG in the human colonic mucosa: A proof-of-concept study. World J Gastroenterol 2018; 24(41): 4652-4662
- URL: https://www.wjgnet.com/1007-9327/full/v24/i41/4652.htm
- DOI: https://dx.doi.org/10.3748/wjg.v24.i41.4652
Recent research has shown a relevant role for the intestinal microbiota in health maintenance and disease occurrences, and consequently, microbiota manipulation by probiotic bacteria has been proposed as a therapeutic option in different pathologic conditions[1]. Nonetheless, the clinical utilization of probiotic bacteria still lacks robust, specific scientific evidence, and it is instead driven by commercial suggestions and generic, unproven health benefits. In particular, some probiotic strains have shown anti-inflammatory effects in vitro and in experimental models, suggesting a potential clinical utilization in intestinal inflammatory pathologies such as inflammatory bowel disease (IBD). In fact, IBD is a chronic condition of unknown aetiology and immunologic pathogenesis, characterized by persistent, deregulated inflammation in the intestine that clinically manifests with cyclic flares and remission states[2]. Therapeutic options include aminosalicylates, corticosteroids, and immunosuppressant and biologic drugs, but a consistent set of patients still do not respond adequately, so the optimization of conventional therapy and the development of novel drugs are desirable. In this scenario, the proposal and scientific investigation of probiotics as a therapeutic option in IBD patients may represent a relevant improvement in clinical management. Of the two main categories of that disease, namely, Crohn’s disease and ulcerative colitis (UC), the latter appears more sensitive to a possible clinical utilization of probiotic bacteria, and indeed, a number of clinical studies have explored that possibility[3]. Nonetheless, the positive results of those bacteria in experimental and in vitro systems still need to be translated with the same efficacy to the clinical setting. A possible explanation for the lack of consistent clinical data on the therapeutic efficacy of probiotic bacteria in UC patients may be accounted for by two major limitations in available studies. First, clinical studies showed a large variability in terms of the probiotic species used, dosages, duration of therapy and end points analysed[4,5]. Second, in those studies, the same baseline clinical condition shows consistent heterogeneity in terms of extension, severity and duration of disease, with evident implications for the evaluation of probiotic efficacy in specific situations. In this regard, a reduction in heterogeneity through methods and protocols standardization may be the key to the improvement of the results of probiotic therapeutic application in UC. In particular, preclinical studies investigating specific properties of probiotics in the human colonic mucosa, such as mucosal adhesion and cytokine modulation, are scarce. Although the mechanism of action of probiotic bacteria is still not completely clear and likely involves multiple pathways, the capacity of the bacteria to adhere to the target site (i.e., the colonic mucosa) and to promote a shift in the cytokine balance towards a regulatory pattern appears to be of particular relevance[6]. In fact, in UC patients, a relative increase in proinflammatory cytokines, such as tumor necrosis factor alpha (TNFα) and interleukin (IL)-17, with a concomitant reduction in regulatory mediators [i.e., transforming growth factor beta (TGFβ) and IL-10] has been described[7,8]. The demonstration of adhesion and mucosal anti-inflammatory effects by a well-characterized probiotic bacterium in the human colonic mucosa may be of great impact for the proposal of selected bacteria for the clinical treatment of UC. Then, to appropriately address and implement probiotic utilization in specific diseases, it appears to be of great importance to perform a methodological evaluation of a specific bacterial species as a real “biotherapeutic” agent, evaluating the characteristics of the bacterial species and its specific mechanisms of action in the same way as drugs, as this would have particular relevance for clinical and preclinical studies that include appropriate experimental models, dose and therapeutic scheme evaluations, and potential specific effects for the single pathologic condition. In addition, since probiotics are generally considered safe, but some important adverse events may still occur, especially in critically ill patients[9], safety evaluations should be included as well.
Lactobacillus rhamnosus GG ATCC 53103 (LGG) represents an ideal probiotic for such a study, since it is the bacterium most intensely investigated in the literature, with documented anti-inflammatory effects in vitro and solid safety data[10-13]. Preliminary clinical studies support the possible utilization of LGG preparations for maintaining remission in UC patients[14].
The aim of the present study was to explore the feasibility and potential efficacy of the clinical utilization of LGG in UC patients, testing the adhesion and mucosal anti-inflammatory effects of probiotic bacteria in an ex vivo experimental model and in a proof-of-concept in vivo study.
Patients: We included patients with UC and normal controls who underwent a rectosigmoidoscopy or complete colonoscopy for different clinical indications, and all participants gave informed consent for the study. The study was approved by the S. Andrea Hospital ethics committee. Patients and controls did not consume antibiotics, probiotics or laxatives in the month before the endoscopy. Standard cleansing preparation by the participants has been assumed. For UC patients, we included only patients in remission or with mild to moderate disease (clinical Mayo score < 7) who were not treated with immunosuppressants, biologic drugs or corticosteroids during the course of the study. A total of 68 UC patients and 30 normal subjects were included in the study for the ex vivo experiment.
Bioptic samples: Biopsies from UC patients and normal controls were collected during colonoscopy. With an ex vivo experimental model, we intended to test mucosal adhesion and cytokine modulation by LGG (ATCC 53103) in normal and inflamed colonic mucosa. To test adhesion, bioptic samples were collected from the proximal (caecum) and distal (rectum) colon of normal subjects (n = 20 samples per group) and UC patients (n = 15 per group). Moreover, bioptic samples from the rectum of UC patients with quiescent (defined as Mayo endoscopic score 0-1) and active (Mayo endoscopic score = 2) inflammation were collected (n = 15 per group). Bioptic specimens were placed in Roswell Park Memorial Institute medium (RPMI 1640, Sigma-Aldrich, St Louis, MO, United States), a standard medium commonly used for cell culture composed of a bicarbonate buffering system and different amounts of amino acids and vitamins, with the addition of LGG at a concentration of 6 × 106 CFU in each sample. After incubation at 37 °C for 2 h, the bioptic samples were collected and washed two times in fresh PBS and finally put in RNAlater solution (Qiagen, Valencia, CA, United States) for future utilization. For cytokine evaluation, biopsies from the caecum (n = 15) and rectum (n = 20) of UC patients and normal controls (n = 10) were collected and incubated in RPMI 1640 medium for 6 hours at 37 °C with or without LGG-conditioned media (CM) prepared according to a protocol modified from Petrof et al[15] and used at a 1:10 ratio in each sample. After incubation, samples were stored in RNAlater solution until they were processed. The incubation time for the organ cultures was optimized by evaluating the expression and degradation of DNA/RNA indirectly by the quantification of the expression of the housekeeping genes (β-actin and GADPH) after incubation at different time points (2, 6, 12, and 24 h).
Additionally, a dose-finding study was performed, and a total of 18 UC patients with quiescent inflammation (Mayo endoscopic score 0-1) were included. For adhesion evaluation, bioptic samples from the rectum were incubated with a regular dose (6 × 106 CFU), a double dose (1.2 × 107 CFU) or a 5-fold higher dose (3 × 107 CFU) of LGG, as previously described (n = 8 per group). For cytokine expression quantification, bioptic samples from the rectum were incubated with a regular dose (1:10), a double dose (2:10) or a 5-fold higher dose (5:10) of LGG CM (n = 10 per group), as previously described. After incubation, samples were stored in RNAlater solution for processing.
For adhesion evaluation, total DNA was extracted and mucosa-adherent LGG were quantified by real-time (RT)-PCR with specific primers already described in the literature[16]. A standard curve was obtained by 10 × serial dilutions of a previously amplified sample. For the evaluation of cytokine expression, total RNA was extracted, and TNFα and IL-17 concentrations were quantified by real time (RT)-PCR with specific primers and expressed in relation to the expression of the internal β-actin control. Primers for TNFα, β-actin, and IL-17 were ordered from Qiagen (Valencia, CA, United States).
Patients: UC patients (n = 20) and control subjects (n = 22) who had a scheduled colonoscopy for disease follow-up or for screening purposes, respectively, were enrolled in the study after informed consent was obtained. Only UC patients with no symptoms of active disease (clinical Mayo score 0-1); no on-going immunosuppressant, biologic or corticosteroid therapy; no consumption of probiotic formulations, antibiotics or laxatives in the last month; and quiescent mucosal inflammation (Mayo endoscopic score 0-1) at endoscopic examination were included in the study. Control subjects did not have major comorbidities and did not consume probiotic formulations, antibiotics or laxatives in the month prior to their inclusion in the study. Normal subjects (n = 12) were asked to consume a commercially available formulation of LGG (Dicoflor 60®, AG Pharma, Rome, Italy) at the “regular” dose of 1.2 × 1010 CFU/d (2 packets/d) for seven days prior to their scheduled colonoscopies. As negative controls, we included normal subjects who did not consume LGG (n = 10). UC patients were asked to consume either a “regular” (1.2 × 1010 CFU/d) or a double (2.4 × 1010 CFU/d) dose of LGG preparation (Dicoflor 60®, AG Pharma, Rome, Italy) for seven days prior to their scheduled colonoscopies (n = 10 each group). All patients underwent regular bowel preparation. Since the standard dose of LGG was not set by specific studies, we arbitrarily defined the “regular” dose of LGG as the standard dose that is most commonly suggested in clinical practice, namely, 2 packets per day of Dicoflor 60®, which is equivalent to 12 × 109 UFC/die of LGG.
At the time of colonoscopy, biopsies from the caecum and rectum were collected, washed and placed in RNAlater solution. Total DNA was extracted, and the amount of mucosa-adherent LGG was quantified by RT-PCR. Total RNA was extracted, and TNFα and IL-17 were quantified by RT-PCR with specific primers.
For DNA and RNA extraction, QIAmp mini DNeasy and RNeasy miniprep kits (Qiagen, Valencia, CA, United States), respectively, were used according to the manufacturer’s instructions. The concentration and purity of extracted nucleic acids were measured with the GeneQuant pro RNA/DNA calculator (Amersham Pharmacia Biotech, Uppsala, Sweden) spectrophotometer. Before utilization, RNA was reverse-transcribed into cDNA using the GeneAmp RNA PCR kit (Applied Biosystems, Foster City, CA, United States) according to the manufacturer’s instructions.
PCR reactions were performed in a total volume of 20 μL, including 16 μL SYBR Green PCR Supermix (Bio-Rad Laboratories, Inc., Hercules, CA, United States) and 4 μL target DNA, with an iCyclerQ detection system (Bio-Rad Laboratories, Inc., Hercules, CA, United States). For every set of primers, the original cycle conditions were maintained. A total of 35 cycles per reaction were performed. Data were expressed as a ratio relative to the lowest value detected after normalization to the internal control beta-actin. After reactions, PCR products were run on a 2% agarose gel with ethidium-bromide staining for visualization.
Data are expressed as the mean ± standard error (SE). t tests for paired data and for independent samples were used for statistical comparisons. A P value < 0.05 was considered significant. MedCalc statistical software version 12.5 (MedCalc Software, Ostend, Belgium) was used for analysis.
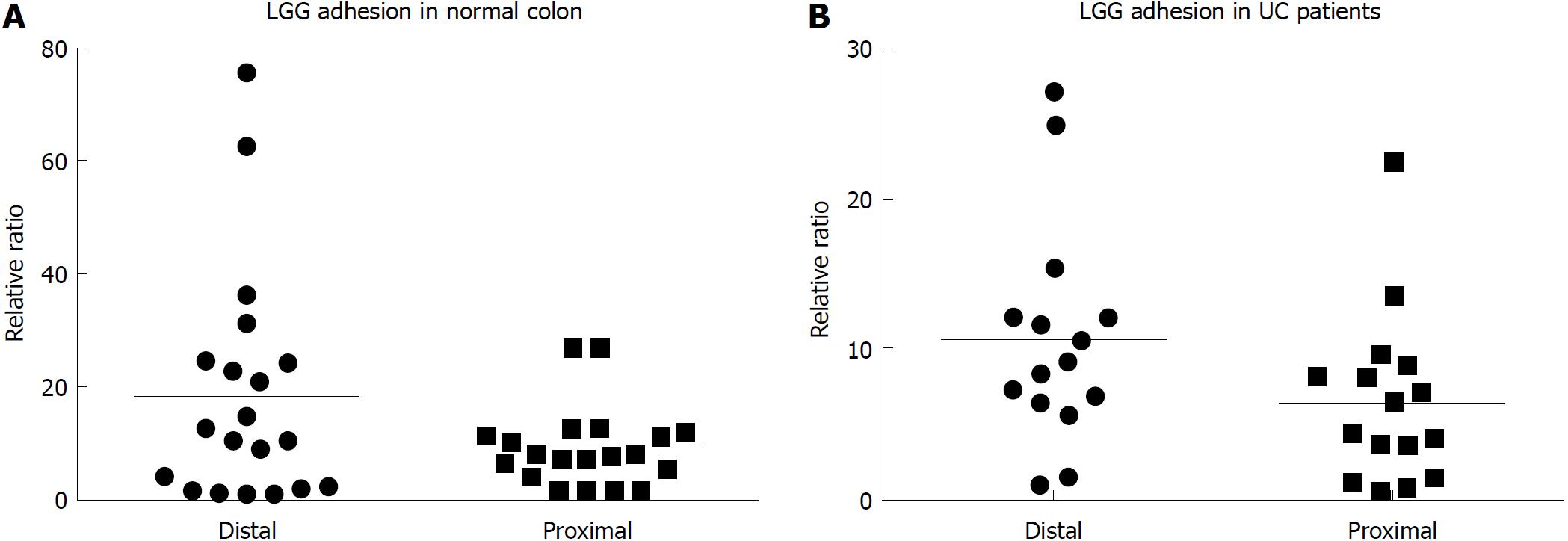
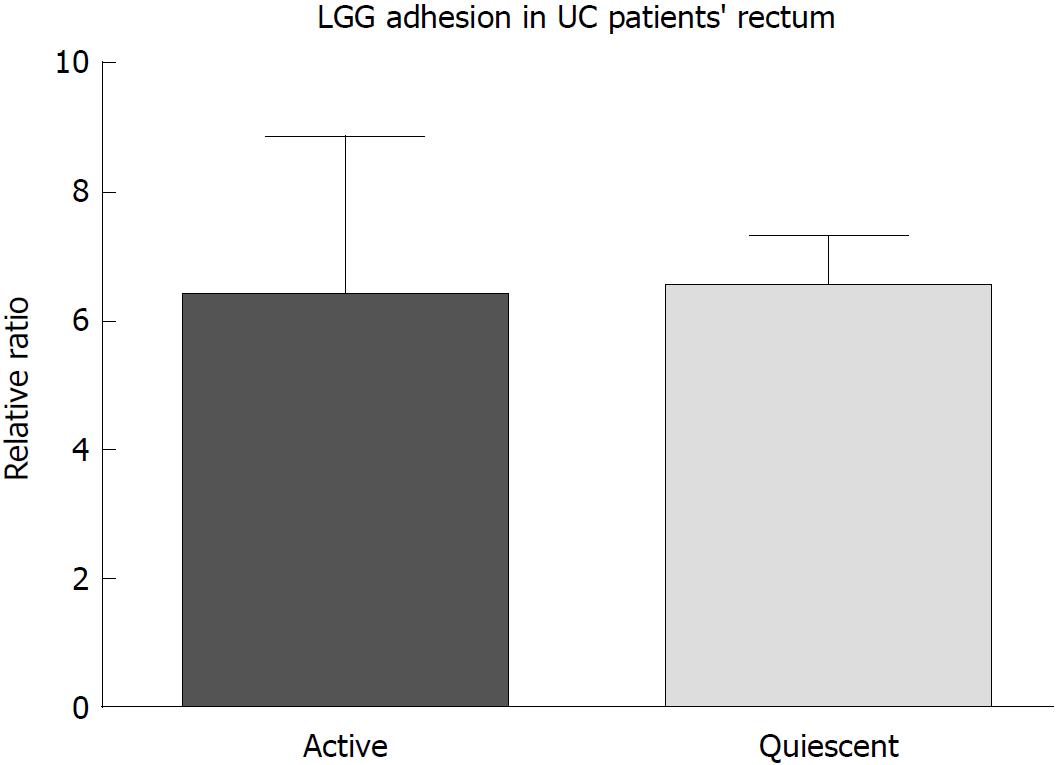
After incubation, LGG effectively adhered to the colonic mucosa in the ex vivo organ culture experimental model. In specimens from the normal colon, LGG showed consistent adhesion to the mucosa of bioptic samples from both the distal and proximal colon (20.4 ± 6.5 vs 9.5 ± 2.3, P = NS) (Figure 1A). Considering relative segmental adhesion, there was a trend for higher concentrations of LGG adhered to the distal vs proximal colon (2-fold relative increment), but the difference was not statistically significant. Similarly, in UC patients, LGG showed remarkable adhesion in bioptic samples from the proximal and distal colon, with a trend towards higher concentrations in the rectum (10.9 ± 2.9 vs 6.5 ± 2.5, P = NS) (Figure 1B). Moreover, LGG showed similar adhesion in mucosal biopsies with endoscopically active and quiescent inflammation, indicating that inflammatory activity does not impair the local adhesion of the bacteria (6.2 ± 2.8 vs 6.6 ± 1.2, P = NS) (Figure 2).
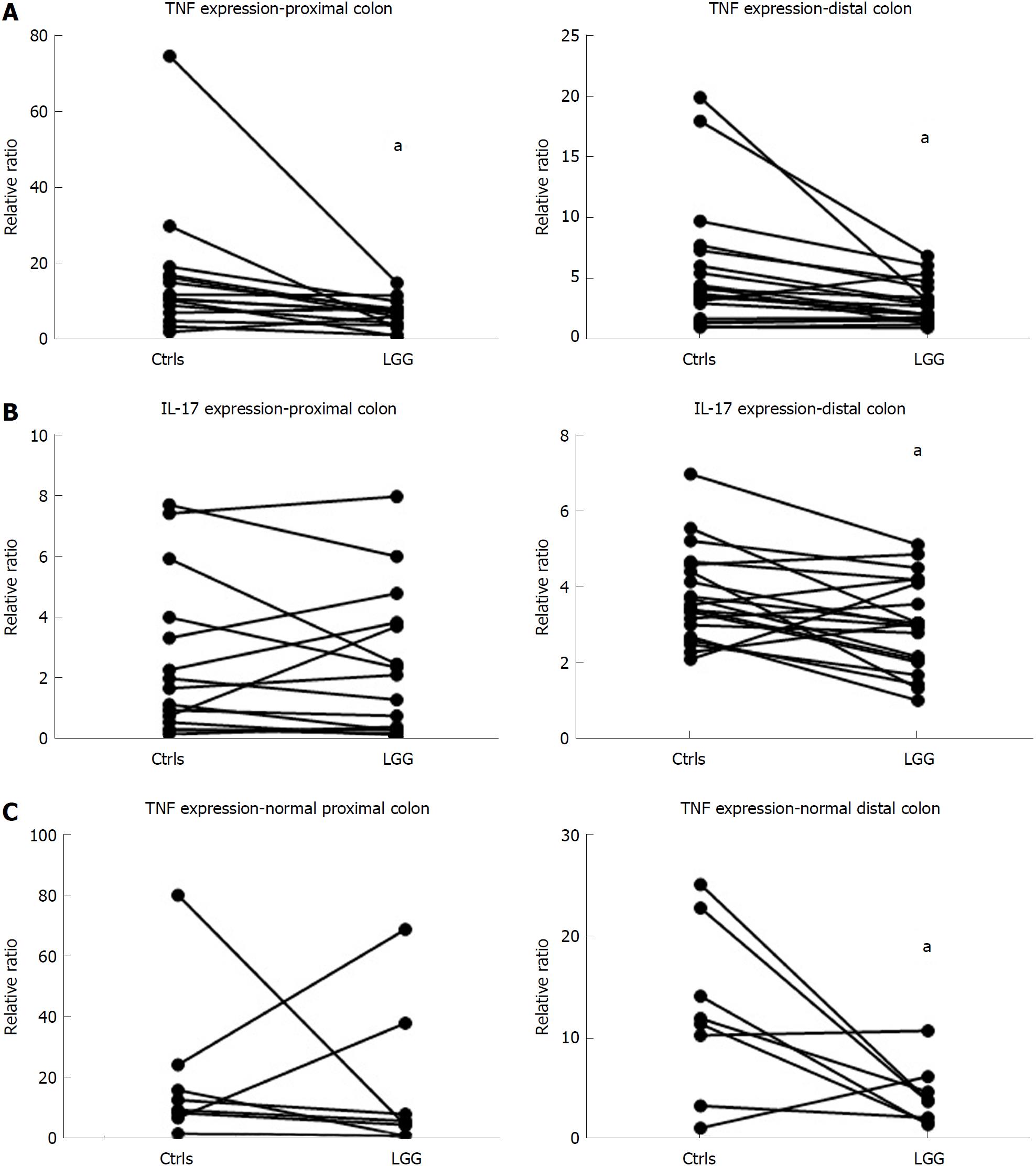
Considering cytokine modulation, incubation of mucosal samples from UC patients with LGG induced a reduction in proinflammatory cytokines. In fact, the mucosal expression of TNFα was significantly reduced in bioptic samples from both the proximal and distal colon incubated with LGG CM compared with biopsies incubated without LGG CM (proximal: 6.8 ± 1.0 vs 16.3 ± 4.7, distal: 3.0 ± 0.4 vs 5.6 ± 1.2, P < 0.05 for paired samples for both) (Figure 3A). Similarly, IL-17 expression was reduced in LGG CM-incubated distal colon bioptic samples (proximal: 2.4 ± 0.6 vs 2.6 ± 0.7, P = NS; distal: 3.0 ± 0.3 vs 3.7 ± 0.3, P < 0.05 for paired samples) (Figure 3B). Similar results were observed when LGG CM was incubated with normal mucosa specimens (Figure 3C).
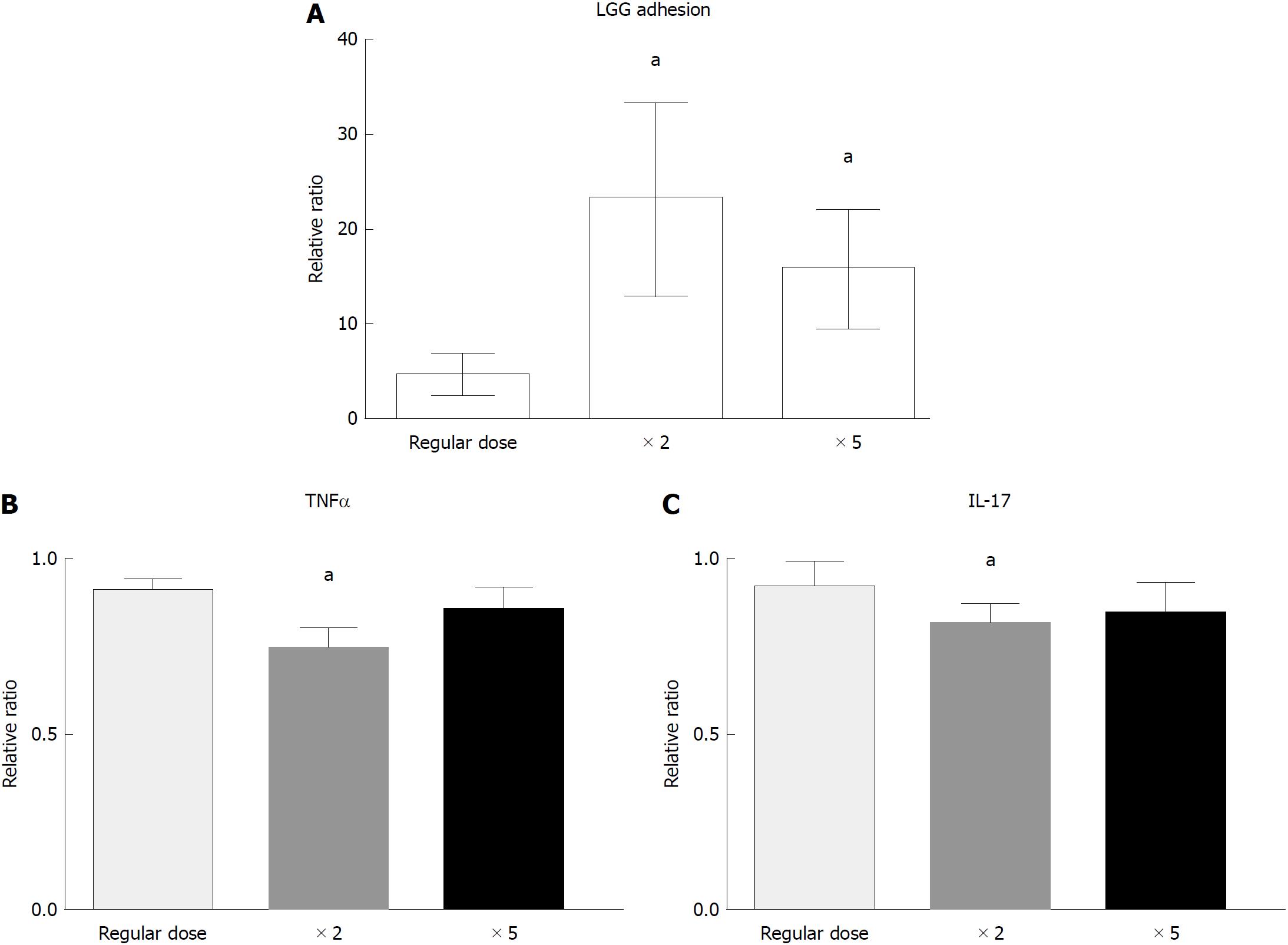
In the dose-response experiment, incubation with a double dose of LGG increased the bacterial DNA content of the bioptic specimens compared to that of the regular dose group (4.7 ± 2.0 vs 23.3 ± 10.0, P < 0.05), while incubation with a higher dose of LGG (× 5) did not further increase the mucosal concentration (15.9 ± 6.2, P < 0.05 vs regular dose, P = NS vs double dose) (Figure 4A). Similarly, incubation with a double dose of LGG CM significantly reduced TNFα and IL-17 expression compared with that of the normal dose (TNFα: 0.91 ± 0.03 vs 0.75 ± 0.05, IL-17: 0.92 ± 0.07 vs 0.82 ± 0.05; P < 0.05 for both), and a higher dose of LGG CM (× 5) did not further reduce TNFα and IL-17 expression (TNFα: 0.87 ± 0.05, IL-17: 0.86 ± 0.07, P = NS). The highest reduction was observed with the double LGG dose (25% and 18% reduction for TNFα and IL-17, respectively) (Figure 4B and C).
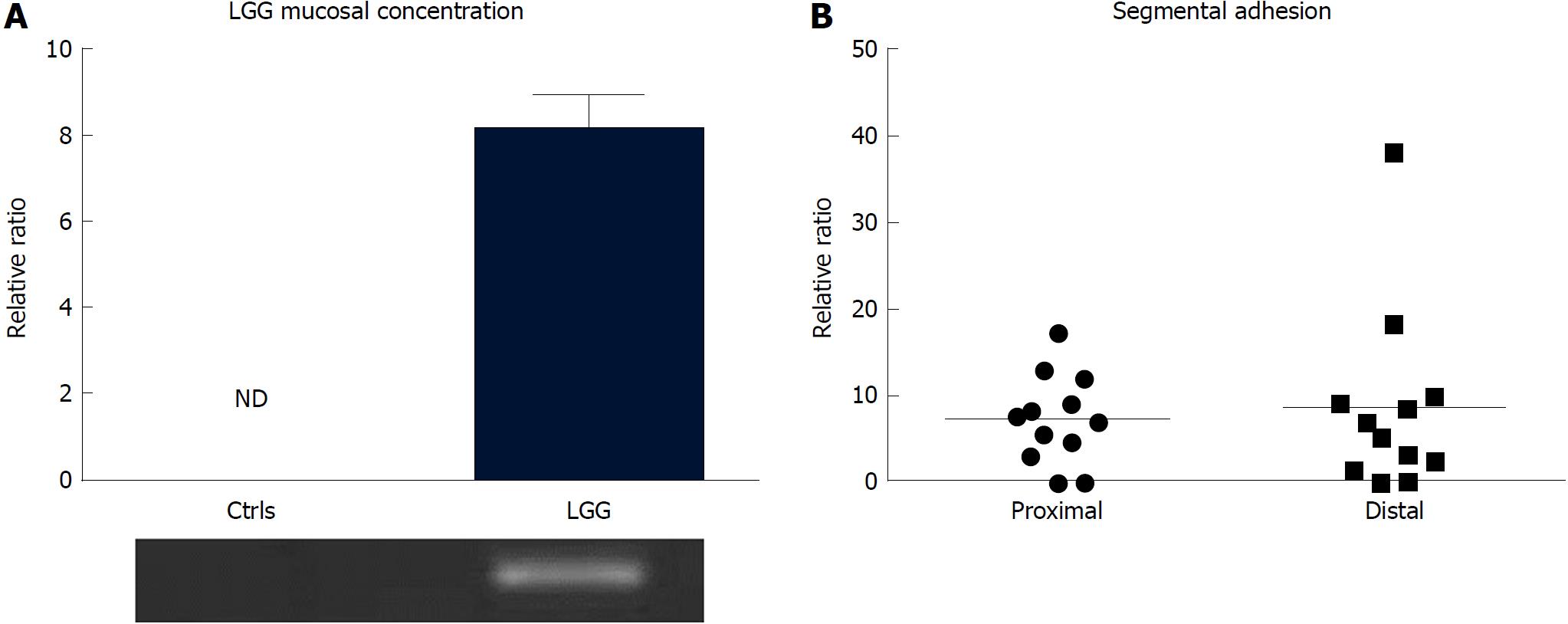
After 7 d of consumption of the regular dose of LGG, adherent bacteria were consistently detectable in bioptic samples, notwithstanding the bowel cleansing for the endoscopic procedure, and not in control biopsies from the colon of subjects who did not consume the probiotic (Figure 5A). LGG adhesion was consistent in both proximal and distal colon biopsies (7.4 ± 1.7 vs 9.0 ± 3.2) (Figure 5B).
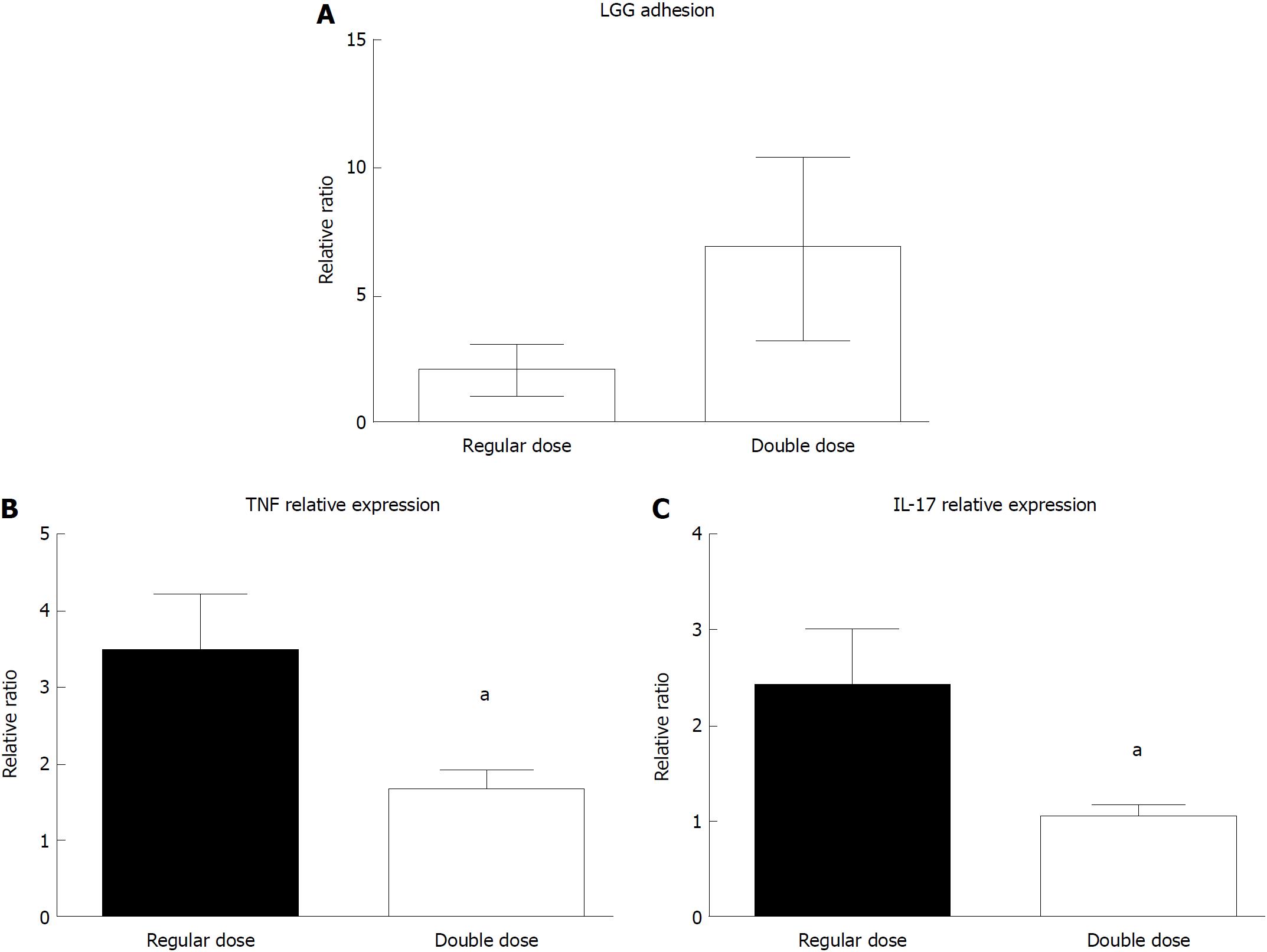
Despite variability among the samples, the LGG mucosal concentration was higher in UC patients who consumed the double dose of LGG compared to those who consumed the regular dose (6.83 ± 2.97 vs 2.02 ± 1.12) (Figure 6A). In line with that result, UC patients who consumed a double LGG dose had a significant reduction in mucosal TNFα and IL-17 expression compared with that of regular dose patients (TNFα: 1.68 ± 0.22 vs 3.48 ± 0.73, 48% reduction, IL-17: 1.05 ± 0.11 vs 2.43 ± 0.57, 43% reduction; P < 0.05 for both) (Figure 6B and C).
The present study shows for the first time that LGG effectively adheres to the colonic mucosa and exerts local anti-inflammatory effects, both in an experimental model and in vivo. Experimental studies have already highlighted the unique adhesion of LGG to the intestinal mucus[17]. Genetic studies have revealed that LGG has pili that produce a mucus-binding protein (SpaC) that enhances the adhesive features of the bacteria[18,19]. Many years ago, pivotal studies demonstrated the presence of cultivable LGG in bioptic samples from patients after consumption of a high dose of LGG[20,21]. In the present study, although a higher dose was associated with increased mucosal adhesion, we have demonstrated for the first time effective mucosal colonization by LGG after consumption of a standard dose, which is the mean dose commonly suggested for the treatment of digestive disorders with the LGG formulation used in the present work. In fact, in the studies by Alander et al[20,21], subjects consumed a preparation containing a dose of LGG approximately 10-20 times higher than that used in the present study, and the evaluation of adherent LGG was performed by traditional culture technique, which has some limitations compared with the molecular method (i.e., RT-PCR) used in the current study that allows direct quantification of bacterial DNA content in the mucosal site. Despite these limitations, such studies already highlighted the limit of exclusive evaluation of faecal samples and the importance of evaluating local mucosal flora in bioptic samples. In fact, even though systemic effects could be present, the adhesion of the probiotic bacteria to the intestinal mucosa appears to be of paramount importance, at least for some of the specific, crucial aspects of the multiple possible mechanisms of action of LGG, namely, pathogen antagonism, stimulation of toll-like receptor (TLR)-mediated pathways, restoration of intestinal bacterial function, and local mucosal cytokine modulation[22]. Accordingly, the preventive effect of a multiprobiotic compound on the development of inflammatory disease in a spontaneous model of Crohn’s disease (SAMP/YitFc mouse), was obtained only with a higher dose of probiotic that effectively induced an increase in the mucosal probiotic concentration in the ileal mucosa[23].
In this study, an experimental ex vivo model was used for the evaluation of the properties of LGG[24]. The utilization of models that are more representative of the in vivo situation is becoming increasingly relevant to developing translational studies with consistent clinical application. This would ideally lead to more in-depth characterization of probiotic species and to the tailored utilization of bacteria with specific features in pathologies where those properties may be specifically relevant for therapeutic purposes. The trend towards higher LGG adhesion in the distal colon, observed both in vivo and in the ex vivo model, may offer an indirect confirmation of the validity of such a representative model, although the observed trends may be simply due to a type two statistical error. In a previous study, we reported correspondence in the data on bacterial mucosal adhesion from both the experimental model and the in vivo setting. In fact, consistent reduction in adherent mucosal bacteria loads has been found in adenomatous polyps compared with normal mucosa and is due to hyperproduction of antibacterial molecules (defensins). Similarly, when bioptic samples were incubated with a probiotic formulation, mucosal adherent bacteria loads were significantly lower in the polyps than in the normal mucosa[25]. The utilization of short-term organ cultures in the present study represents an effective, reproducible and inexpensive method for the evaluation of probiotic adhesion and effects, and it requires materials (i.e., bioptic samples) that are easily available compared with for instance, surgical specimens. Moreover, ex vivo bioptic organ cultures may reflect a situation closer to the in vivo setting compared with other pure in vitro systems, such as primary cell cultures or cell lines. On the other hand, organ cultures may be more easily contaminated, and manipulations and long-term incubation are difficult to perform. In addition, in our method, unlike the in vivo setting where the epithelial cells represent the main interface between the luminal content and subepithelial compartment, the polarization of the stimulus (i.e., LGG CM) is not controlled, and direct paracellular stimulation may occur.
For genetic evaluations in the present paper, we used a SYBR Green-based dye detection system, which represents an accurate and relatively inexpensive method for DNA and RNA quantification. Since, to our knowledge, RT-PCR primers for SYBR Green-based systems that specifically target LGG have not been described, we used primers that were already developed for general Lactobacillus species[16]. Considering that LGG administration may affect the general composition of faecal microbiota[26,27], we cannot exclude the possibility that other Lactobacillus species might have been stimulated and unspecifically detected in our analysis. Nonetheless, the facts that we focused our analysis on the intestinal mucosa and not on the faeces, that the incubation time of the ex vivo experiments was short and that the consumption time for the in vivo experiment (a week) was relatively short indicate that concomitant significant alterations in the intestinal microbiota composition is unlikely and that the increased concentration observed with real-time PCR is specifically ascribable to LGG. To confirm our findings and to further characterize the LGG adhesion to the mucosa, investigations with different methods [i.e., fluorescence in situ hybridization (FISH)] are currently being performed.
Together with consistent colonization of the colonic mucosa, LGG demonstrated effective cytokine modulation in the present study. Previous in vitro studies have already shown consistent anti-inflammatory properties of LGG. In fact, lipopolysaccharide (LPS)-activated macrophages derived from BALB/c mice showed reduced production of TNFα after incubation with LGG CM but not with CM from other probiotic bacteria[28]. In peripheral blood mononuclear cells (PBMCs) from humans, incubation with LGG and other probiotic bacteria stimulated the modulation of cytokine production towards a regulatory pattern[11]. The modulation of the cytokine profile is likely dependent on the induction of nuclear factor (NF)-κB and STAT DNA-binding activities, as demonstrated in human primary macrophages[29]. Moreover, LGG displayed anti-inflammatory effects and protection in several experimental models of colitis. In fact, LGG administration ameliorated inflammation both in DSS-induced colitis and in human leukocyte antigen (HLA) B-27 transgenic rats[30,31]. Yan et al[32] demonstrated that a soluble protein derived from LGG, namely, p40, confers protection against intestinal inflammation through the prevention of cytokine-induced apoptosis in epithelial cells by means of an epithelial growth factor receptor (EGFR)-dependent mechanism. Recently, the same group demonstrated that LGG neonatal colonization in mice promotes intestinal development and protects from intestinal inflammation induced by DSS administration in adult mice[33], and LGG-mediated protection against inflammation and carcinogenesis has been demonstrated in the dimethyl hydrazine (DMH) model of colitis-associated cancer[34]. More evidence about the potential protective role of LGG in the intestinal mucosa was generated in an in vitro study by Priciandaro et al[35] that demonstrated a reduction in apoptosis and in the loss of intestinal barrier function in enterocytes pretreated with LGG supernatant; the described study is a further indication of the positive effect of the metabolic products of LGG. The confirmation of cytokine modulation by LGG, which we demonstrated for the first time in the colonic mucosa in the present study, may be relevant for proposing the use of LGG in the clinical setting as a therapeutic option in inflammatory pathologic conditions. The identification of a specific molecular target of the probiotic bacteria was not addressed in our investigation. In fact, although a direct effect on mucosal cytokines associated with acquired immunity appears most likely, alternative effects, such as permeability modification, interaction with other commensal bacteria, or concomitant action on the innate immune compartment, cannot be excluded. Moreover, in this proof-of-concept study, we intended to preliminarily demonstrate the potential rationale for LGG utilization in UC patients by analysing mucosal adhesion and the anti-inflammatory effect of the bacteria. We limited our investigation to the expression of two cytokines that are upregulated in the mucosa of UC patients and that are likely to play an important role in inflammation initiation and maintenance. Nonetheless, since chronic inflammation is most likely driven by multiple alterations in the complex cytokine network, further studies are needed to better characterize the molecular effect of LGG, and comparison with other probiotic species will help clarify the potential specificity of the effect of LGG on inflammation.
We included in the present study, for both the ex vivo and in vivo experiments, only UC patients with mild-moderate disease activity. In fact, according to current clinical knowledge and experimental observations, the clinical scenario for probiotic utilization in UC patient management appears to be the induction of remission in mild-moderate flares and overall remission maintenance (disease prevention). Despite some encouraging clinical results, solid evidence for the indication and correct utilization of probiotics in UC patient management is still to come, and many questions are currently still unanswered[36]. In this study, we demonstrated a clear anti-inflammatory effect in the experimental model. We also tested the effect on cytokines in the mucosa of patients consuming two different doses of LGG. In this “proof–of-concept” study, experimental and in vivo dose responses were observed after LGG administration. Since no analogous studies have been performed to date, in our experimental model, we set an LGG concentration of 6 × 106 CFU as the “regular dose” and an LGG CM concentration of 1:10 as the baseline dose for the reaction, according to the concentrations previously used in the probiotic literature for in vitro studies. When administering a double dose of LGG or LGG CM, we observed an increase in mucosal adherent bacteria and a decrease in proinflammatory cytokine expression, while no further increase in adhesion or any effect on the cytokines was observed with a fivefold increase in LGG dose. A possible explanation is that doubling the dose of LGG in the reaction reduces the quantity of culture media (i.e., in a 1:10 LGG reaction culture medium is 180 μL, while in a 1:50 LGG CM reaction it is 100 μL) so that bioptic samples may not have adequate culture conditions. Nonetheless, the identification of a real threshold effect was beyond the purpose of the study and was not investigated. We limited our investigation to the administration of two doses of oral LGG (regular and double) for the in vivo study, which allowed the preliminary confirmation of the ex vivo finding; the lack of safety and tolerability data for a very high dose of LGG supplement administration in a clinical setting also limited our dosing scheme. In fact, although LGG and other probiotic species are generally considered safe, adverse events are described[37], and a thorough safety evaluation is needed.
The novel findings of the present investigation preliminarily confirm and support the feasibility and the potential effectiveness of the application of LGG administration in UC patients. Since the variability of methods and protocols has been a major limitation of previously published studies on probiotic evaluation, we intended to restrict the patient population (i.e., mild-moderate active UC patients) and to standardize the methodological evaluation to provide the basis for a rigorous evaluation of LGG efficacy in this clinical scenario and eventually in other similar inflammatory conditions, with the ultimate goals of contributing to overcoming inappropriate and nonspecific use of probiotic bacteria and promoting more selective, evidence-based applications of those bacteria in specific clinical conditions.
Ulcerative colitis (UC) is one of the two major forms of Inflammatory bowel disease (IBD), and it is characterized by a deregulated inflammation of the intestinal mucosa clinically presenting with symptoms of diarrhea and bloody stools. Despite different therapies are available, still many patients do not adequately respond. The therapeutic application of probiotic bacteria in this setting could represent an efficient and attractive option for UC patients.
The research of probiotic application in IBD has been largely focused on in vitro systems and experimental models, and pre-clinical and clinical studies are characterized by an extreme dishomogeneity. Therefore, no specific indications for therapeutic application of probiotics (which probiotic species, doses, duration of therapy, and kind of UC disease severity/extension to treat) can be extrapolated from the literature.
In order to overcome the consistent dishomogeneity of previous studies, the purpose of the present work was to restrict the investigation of a well characterized probiotic species, Lactobacillus rhamnosus GG (LGG), in a specific clinical setting (mild- moderate UC patients), to measurable parameters correlated with the potential therapeutic effect (mucosal adhesion and anti-inflammatory effect).
We intended to investigate the adhesion and effect of LGG directly at the mucosal site by using modern genomic culture-independent techniques. An ex-vivo experimental model based on short-term biopsies organ culture was developed. Concentration and mucosal effect of LGG in colonic mucosa was then confirmed in vivo in UC patients and normal subjects after consumption of oral formulation containing LGG.
LGG affectively adhered to the colonic mucosa, both in the experimental model and in vivo. Moreover, the probiotc administration reduced expression of two important pro-inflammatory cytokines (tumor necrosis factor alpha and interleukin-17). Higher mucosal concentration of the bacteria and more consistent reduction of pro-inflammatory cytokines has been observed increasing the dose of LGG.
The present study demonstrated for the first time that LGG affectively adheres to the colonic mucosa and exerts anti-inflammatory effect, both ex vivo and in vivo. Moreover, we demonstrated a potential increasing of effect with higher dose of probiotic. Considering that probiotic utilization in clinical setting is often empirical and not evidence-based, the present study can put the basis for the evaluation of LGG in the setting of UC with specific clinical trials.
Utilization of appropriate experimental models appears crucial for the design of pre-clinical studies. Translational medicine studies, with both experimental and in vivo experiments, may contribute to correctly investigate potential novel treatments, and in particular to evaluate the potential application of LGG and other probiotic bacteria for the treatment of UC and other inflammatory conditions.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: Italy
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Antonelli A, Howarth GS, Sherman PM S- Editor: Ma RY L- Editor: A E- Editor: Yin SY
| 1. | Marchesi JR, Adams DH, Fava F, Hermes GD, Hirschfield GM, Hold G, Quraishi MN, Kinross J, Smidt H, Tuohy KM. The gut microbiota and host health: a new clinical frontier. Gut. 2016;65:330-339. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1341] [Cited by in F6Publishing: 1417] [Article Influence: 177.1] [Reference Citation Analysis (0)] |
| 2. | Abraham C, Cho JH. Inflammatory bowel disease. N Engl J Med. 2009;361:2066-2078. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1967] [Cited by in F6Publishing: 2077] [Article Influence: 138.5] [Reference Citation Analysis (6)] |
| 3. | Pagnini C, Delle Fave G, Bamias G. Probiotics in inflammatory bowel disease: Pathophysiological background and clinical applications. World J Immunol. 2013;3:31-43. [DOI] [Cited in This Article: ] [Cited by in CrossRef: 8] [Cited by in F6Publishing: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 4. | Mallon P, McKay D, Kirk S, Gardiner K. Probiotics for induction of remission in ulcerative colitis. Cochrane Database Syst Rev. 2007;CD005573. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 74] [Cited by in F6Publishing: 81] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 5. | Naidoo K, Gordon M, Fagbemi AO, Thomas AG, Akobeng AK. Probiotics for maintenance of remission in ulcerative colitis. Cochrane Database Syst Rev. 2011;CD007443. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 60] [Cited by in F6Publishing: 76] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 6. | Siciliano RA, Mazzeo MF. Molecular mechanisms of probiotic action: a proteomic perspective. Curr Opin Microbiol. 2012;15:390-396. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 44] [Cited by in F6Publishing: 48] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 7. | Bamias G, Cominelli F. Immunopathogenesis of inflammatory bowel disease: current concepts. Curr Opin Gastroenterol. 2007;23:365-369. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 79] [Cited by in F6Publishing: 76] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 8. | Jiang W, Su J, Zhang X, Cheng X, Zhou J, Shi R, Zhang H. Elevated levels of Th17 cells and Th17-related cytokines are associated with disease activity in patients with inflammatory bowel disease. Inflamm Res. 2014;63:943-950. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 120] [Cited by in F6Publishing: 128] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 9. | Didari T, Solki S, Mozaffari S, Nikfar S, Abdollahi M. A systematic review of the safety of probiotics. Expert Opin Drug Saf. 2014;13:227-239. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 187] [Cited by in F6Publishing: 187] [Article Influence: 18.7] [Reference Citation Analysis (0)] |
| 10. | Donato KA, Gareau MG, Wang YJ, Sherman PM. Lactobacillus rhamnosus GG attenuates interferon-{gamma} and tumour necrosis factor-alpha-induced barrier dysfunction and pro-inflammatory signalling. Microbiology. 2010;156:3288-3297. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 132] [Cited by in F6Publishing: 142] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 11. | Donkor ON, Ravikumar M, Proudfoot O, Day SL, Apostolopoulos V, Paukovics G, Vasiljevic T, Nutt SL, Gill H. Cytokine profile and induction of T helper type 17 and regulatory T cells by human peripheral mononuclear cells after microbial exposure. Clin Exp Immunol. 2012;167:282-295. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 69] [Cited by in F6Publishing: 70] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 12. | Braat H, van den Brande J, van Tol E, Hommes D, Peppelenbosch M, van Deventer S. Lactobacillus rhamnosus induces peripheral hyporesponsiveness in stimulated CD4+ T cells via modulation of dendritic cell function. Am J Clin Nutr. 2004;80:1618-1625. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 145] [Cited by in F6Publishing: 151] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 13. | Hibberd PL, Kleimola L, Fiorino AM, Botelho C, Haverkamp M, Andreyeva I, Poutsiaka D, Fraser C, Solano-Aguilar G, Snydman DR. No evidence of harms of probiotic Lactobacillus rhamnosus GG ATCC 53103 in healthy elderly-a phase I open label study to assess safety, tolerability and cytokine responses. PLoS One. 2014;9:e113456. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 34] [Cited by in F6Publishing: 34] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 14. | Zocco MA, dal Verme LZ, Cremonini F, Piscaglia AC, Nista EC, Candelli M, Novi M, Rigante D, Cazzato IA, Ojetti V. Efficacy of Lactobacillus GG in maintaining remission of ulcerative colitis. Aliment Pharmacol Ther. 2006;23:1567-1574. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 323] [Cited by in F6Publishing: 283] [Article Influence: 15.7] [Reference Citation Analysis (0)] |
| 15. | Petrof EO, Kojima K, Ropeleski MJ, Musch MW, Tao Y, De Simone C, Chang EB. Probiotics inhibit nuclear factor-kappaB and induce heat shock proteins in colonic epithelial cells through proteasome inhibition. Gastroenterology. 2004;127:1474-1487. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 244] [Cited by in F6Publishing: 235] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 16. | Furet JP, Quénée P, Tailliez P. Molecular quantification of lactic acid bacteria in fermented milk products using real-time quantitative PCR. Int J Food Microbiol. 2004;97:197-207. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 139] [Cited by in F6Publishing: 140] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 17. | Tuomola E, Crittenden R, Playne M, Isolauri E, Salminen S. Quality assurance criteria for probiotic bacteria. Am J Clin Nutr. 2001;73:393S-398S. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 254] [Cited by in F6Publishing: 232] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 18. | Kankainen M, Paulin L, Tynkkynen S, von Ossowski I, Reunanen J, Partanen P, Satokari R, Vesterlund S, Hendrickx AP, Lebeer S. Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human- mucus binding protein. Proc Natl Acad Sci USA. 2009;106:17193-17198. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 540] [Cited by in F6Publishing: 526] [Article Influence: 35.1] [Reference Citation Analysis (0)] |
| 19. | von Ossowski I, Reunanen J, Satokari R, Vesterlund S, Kankainen M, Huhtinen H, Tynkkynen S, Salminen S, de Vos WM, Palva A. Mucosal adhesion properties of the probiotic Lactobacillus rhamnosus GG SpaCBA and SpaFED pilin subunits. Appl Environ Microbiol. 2010;76:2049-2057. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 150] [Cited by in F6Publishing: 154] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 20. | Alander M, Korpela R, Saxelin M, Vilpponen-Salmela T, Mattila-Sandholm T, von Wright A. Recovery of Lactobacillus rhamnosus GG from human colonic biopsies. Lett Appl Microbiol. 1997;24:361-364. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 111] [Cited by in F6Publishing: 113] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 21. | Alander M, Satokari R, Korpela R, Saxelin M, Vilpponen-Salmela T, Mattila-Sandholm T, von Wright A. Persistence of colonization of human colonic mucosa by a probiotic strain, Lactobacillus rhamnosus GG, after oral consumption. Appl Environ Microbiol. 1999;65:351-354. [PubMed] [Cited in This Article: ] |
| 22. | Kemgang TS, Kapila S, Shanmugam VP, Kapila R. Cross-talk between probiotic lactobacilli and host immune system. J Appl Microbiol. 2014;117:303-319. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 66] [Cited by in F6Publishing: 71] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 23. | Pagnini C, Saeed R, Bamias G, Arseneau KO, Pizarro TT, Cominelli F. Probiotics promote gut health through stimulation of epithelial innate immunity. Proc Natl Acad Sci USA. 2010;107:454-459. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 242] [Cited by in F6Publishing: 254] [Article Influence: 16.9] [Reference Citation Analysis (0)] |
| 24. | Pagnini C, Martorelli M, Lanini C, Delle Fave G. Development of an Ex Vivo Organ Culture Technique to Evaluate Probiotic Utilization in IBD. J Clin Gastroenterol. 2016;50:S179-S182. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2] [Cited by in F6Publishing: 2] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 25. | Pagnini C, Corleto VD, Mangoni ML, Pilozzi E, Torre MS, Marchese R, Carnuccio A, Giulio ED, Delle Fave G. Alteration of local microflora and α-defensins hyper-production in colonic adenoma mucosa. J Clin Gastroenterol. 2011;45:602-610. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 38] [Cited by in F6Publishing: 38] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 26. | Ni Y, Wong VH, Tai WC, Li J, Wong WY, Lee MM, Fong FL, El-Nezami H, Panagiotou G. A metagenomic study of the preventive effect of Lactobacillus rhamnosus GG on intestinal polyp formation in ApcMin/+mice. J Appl Microbiol. 2017;122:770-784. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 27. | Zhang W, Zhu YH, Yang GY, Liu X, Xia B, Hu X, Su JH, Wang JF. Lactobacillus rhamnosus GG Affects Microbiota and Suppresses Autophagy in the Intestines of Pigs Challenged with Salmonella Infantis. Front Microbiol. 2018;8:2705. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 46] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 28. | Peña JA, Versalovic J. Lactobacillus rhamnosus GG decreases TNF-alpha production in lipopolysaccharide-activated murine macrophages by a contact-independent mechanism. Cell Microbiol. 2003;5:277-285. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 134] [Cited by in F6Publishing: 135] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 29. | Miettinen M, Lehtonen A, Julkunen I, Matikainen S. Lactobacilli and Streptococci activate NF-kappa B and STAT signaling pathways in human macrophages. J Immunol. 2000;164:3733-3740. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 157] [Cited by in F6Publishing: 147] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 30. | Yoda K, Miyazawa K, Hosoda M, Hiramatsu M, Yan F, He F. Lactobacillus GG-fermented milk prevents DSS-induced colitis and regulates intestinal epithelial homeostasis through activation of epidermal growth factor receptor. Eur J Nutr. 2014;53:105-115. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 56] [Cited by in F6Publishing: 61] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 31. | Dieleman LA, Goerres MS, Arends A, Sprengers D, Torrice C, Hoentjen F, Grenther WB, Sartor RB. Lactobacillus GG prevents recurrence of colitis in HLA-B27 transgenic rats after antibiotic treatment. Gut. 2003;52:370-376. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 167] [Cited by in F6Publishing: 157] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 32. | Yan F, Cao H, Cover TL, Washington MK, Shi Y, Liu L, Chaturvedi R, Peek RM Jr, Wilson KT, Polk DB. Colon-specific delivery of a probiotic-derived soluble protein ameliorates intestinal inflammation in mice through an EGFR-dependent mechanism. J Clin Invest. 2011;121:2242-2253. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 240] [Cited by in F6Publishing: 254] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 33. | Yan F, Liu L, Cao H, Moore DJ, Washington MK, Wang B, Peek RM, Acra SA, Polk DB. Neonatal colonization of mice with LGG promotes intestinal development and decreases susceptibility to colitis in adulthood. Mucosal Immunol. 2017;10:117-127. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 62] [Cited by in F6Publishing: 75] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 34. | Gamallat Y, Meyiah A, Kuugbee ED, Hago AM, Chiwala G, Awadasseid A, Bamba D, Zhang X, Shang X, Luo F. Lactobacillus rhamnosus induced epithelial cell apoptosis, ameliorates inflammation and prevents colon cancer development in an animal model. Biomed Pharmacother. 2016;83:536-541. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 119] [Cited by in F6Publishing: 128] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 35. | Prisciandaro LD, Geier MS, Chua AE, Butler RN, Cummins AG, Sander GR, Howarth GS. Probiotic factors partially prevent changes to caspases 3 and 7 activation and transepithelial electrical resistance in a model of 5-fluorouracil-induced epithelial cell damage. Support Care Cancer. 2012;20:3205-3210. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 34] [Cited by in F6Publishing: 36] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 36. | Derwa Y, Gracie DJ, Hamlin PJ, Ford AC. Systematic review with meta-analysis: the efficacy of probiotics in inflammatory bowel disease. Aliment Pharmacol Ther. 2017;46:389-400. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 224] [Cited by in F6Publishing: 238] [Article Influence: 34.0] [Reference Citation Analysis (1)] |
| 37. | Meini S, Laureano R, Fani L, Tascini C, Galano A, Antonelli A, Rossolini GM. Breakthrough Lactobacillus rhamnosus GG bacteremia associated with probiotic use in an adult patient with severe active ulcerative colitis: case report and review of the literature. Infection. 2015;43:777-781. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 57] [Cited by in F6Publishing: 62] [Article Influence: 6.9] [Reference Citation Analysis (0)] |